Zebrafish—A Suitable Model for Rapid Translation of Effective Therapies for Pediatric Cancers
Abstract
Simple Summary
Abstract
1. Introduction
2. In Vitro Models for Pediatric Cancers
3. In Vivo Models for Pediatric Cancers
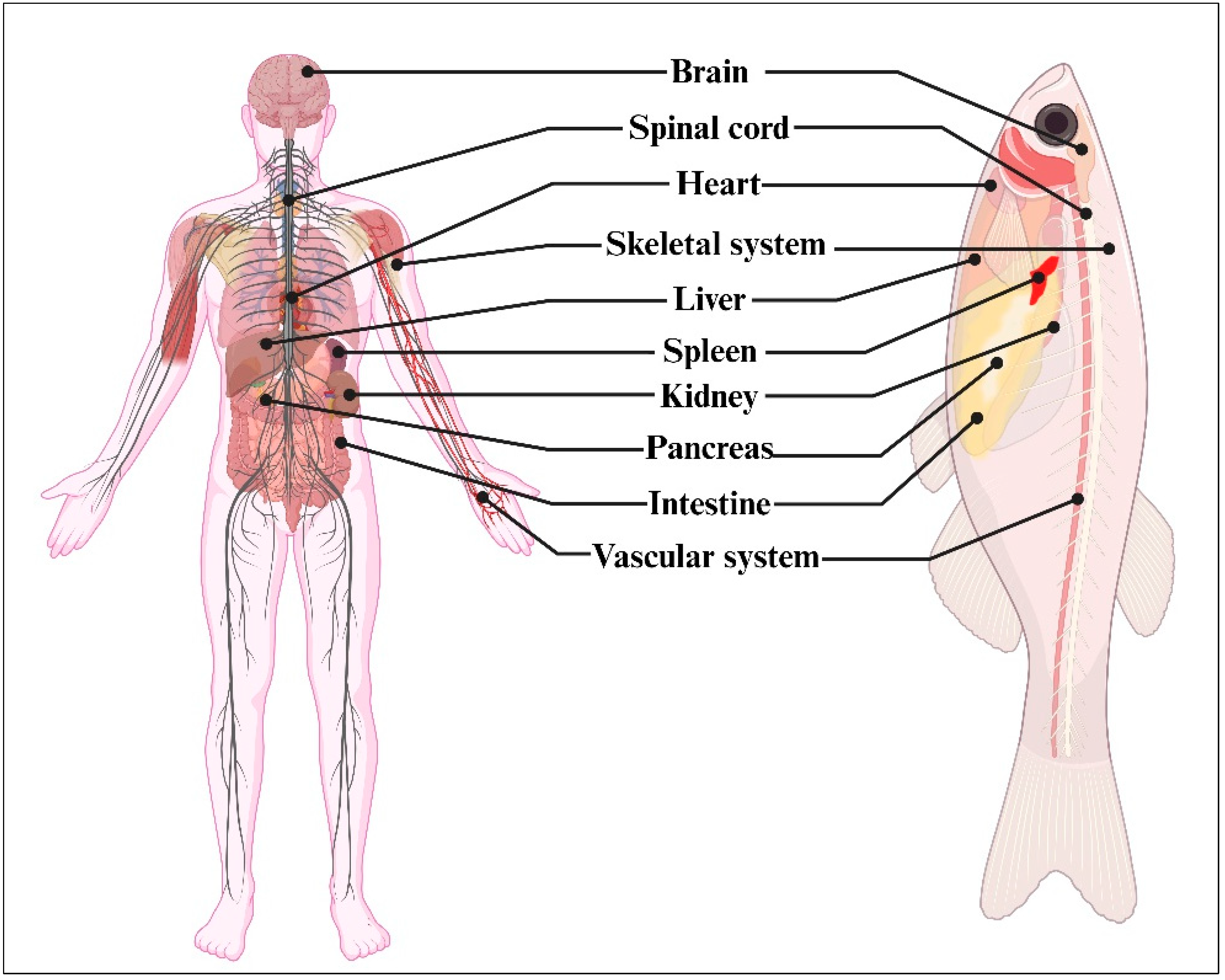
4. Zebrafish—A Tiny Human
5. Zebrafish Models for Human Disorders
6. Zebrafish Models for Cancer Research
7. Pediatric Cancer Models in Zebrafish
8. Non-CNS Pediatric Cancer Models in Zebrafish
9. Pediatric CNS Cancer Models in Zebrafish
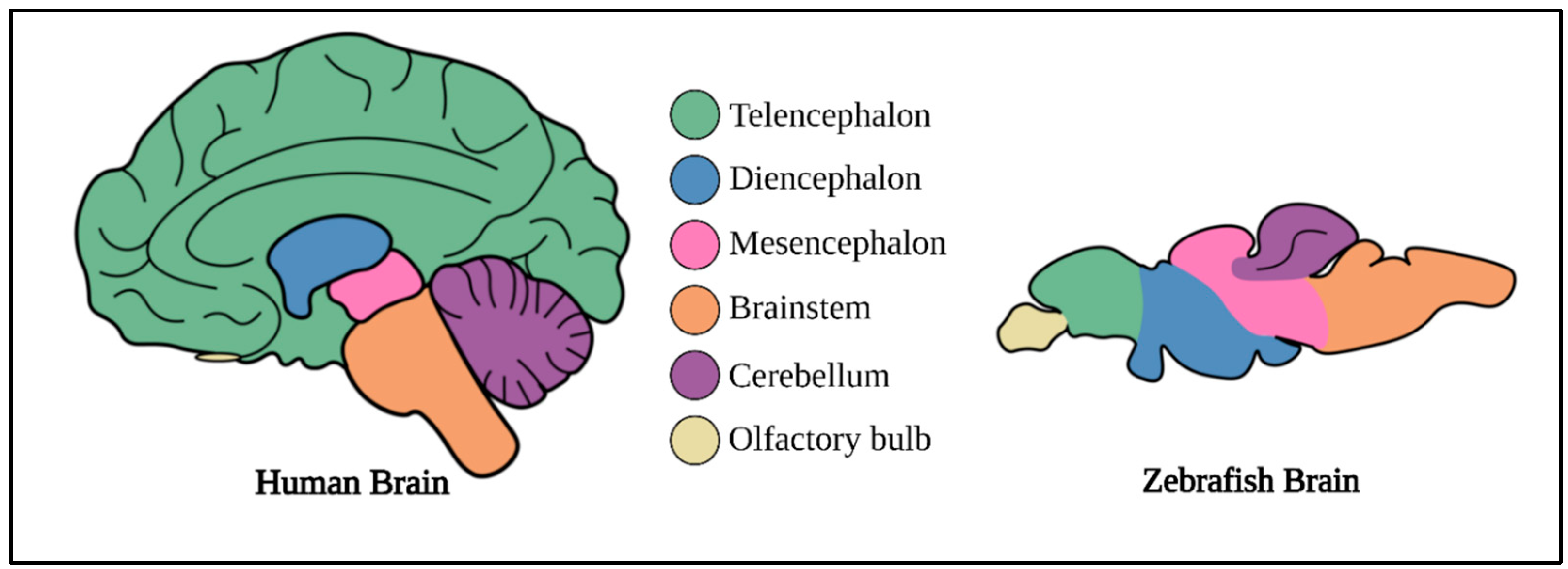
10. Zebrafish Brain
11. Zebrafish Models of Childhood CNS Cancer
12. Humanized Zebrafish
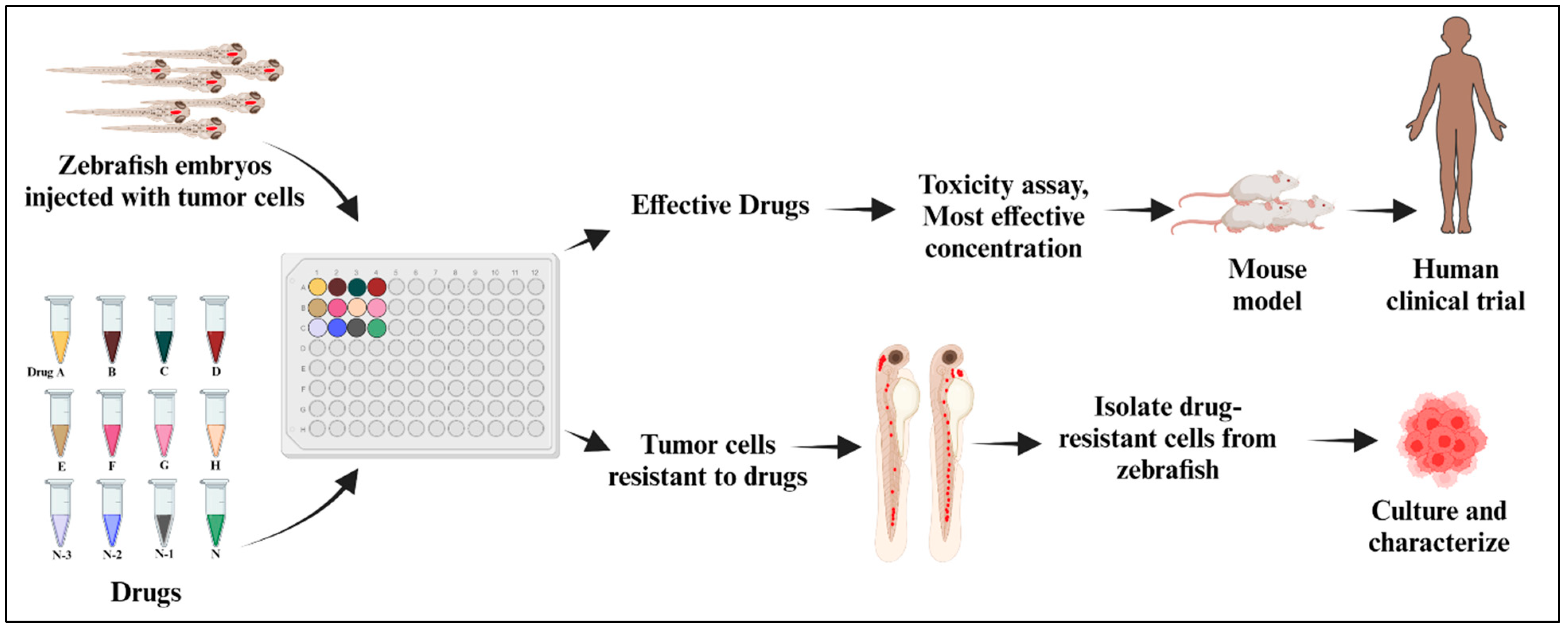
13. Zebrafish as a Tool for Cancer Drug Discovery
14. Zebrafish: A Rapid and Cost-Effective Pediatric Cancer Model Organism
15. Concluding Remarks and Future Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Siegel, D.A.; King, J.B.; Lupo, P.J.; Durbin, E.B.; Tai, E.; Mills, K.; Van Dyne, E.; Buchanan Lunsford, N.; Henley, S.J.; Wilson, R.J. Counts, Incidence Rates, and Trends of Pediatric Cancer in the United States, 2003–2019. JNCI J. Natl. Cancer Inst. 2023, 115, 1337–1354. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, S. Pediatric Cancers: Insights and Novel Therapeutic Approaches. Cancers 2023, 15, 3537. [Google Scholar] [CrossRef] [PubMed]
- Smeland, S.; Bielack, S.S.; Whelan, J.; Bernstein, M.; Hogendoorn, P.; Krailo, M.D.; Gorlick, R.; Janeway, K.A.; Ingleby, F.C.; Anninga, J.; et al. Survival and Prognosis with Osteosarcoma: Outcomes in More than 2000 Patients in the EURAMOS-1 (European and American Osteosarcoma Study) Cohort. Eur. J. Cancer 2019, 109, 36–50. [Google Scholar] [CrossRef] [PubMed]
- Ostrom, Q.T.; Cioffi, G.; Waite, K.; Kruchko, C.; Barnholtz-Sloan, J.S. CBTRUS Statistical Report: Primary Brain and Other Central Nervous System Tumors Diagnosed in the United States in 2014–2018. Neuro Oncol. 2021, 23 (Suppl. 3), iii1–iii105. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Percie du Sert, N.; Vollert, J.; Rice, A.S.C. General Principles of Preclinical Study Design. In Handbook of Experimental Pharmacology; Springer: Berlin/Heidelberg, Germany, 2020; Volume 257, pp. 55–69. [Google Scholar] [CrossRef]
- Hahn, W.C.; Counter, C.M.; Lundberg, A.S.; Beijersbergen, R.L.; Brooks, M.W.; Weinberg, R.A. Creation of Human Tumour Cells with Defined Genetic Elements. Nature 1999, 400, 464–468. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Kasai, T.; Li, Y.; Sugii, Y.; Jin, G.; Okada, M.; Vaidyanath, A.; Mizutani, A.; Satoh, A.; Kudoh, T.; et al. A Model of Cancer Stem Cells Derived from Mouse Induced Pluripotent Stem Cells. PLoS ONE 2012, 7, e33544. [Google Scholar] [CrossRef] [PubMed]
- Stackhouse, C.T.; Gillespie, G.Y.; Willey, C.D. Cancer Explant Models. In Three Dimensional Human Organotypic Models for Biomedical Research; Current Topics in Microbiology and Immunology; Springer: Cham, Switzerland, 2021; Volume 430, p. 131. [Google Scholar] [CrossRef]
- Choorapoikayil, S.; Overvoorde, J.; Den Hertog, J. Deriving Cell Lines from Zebrafish Embryos and Tumors. Zebrafish 2013, 10, 316. [Google Scholar] [CrossRef] [PubMed]
- Papapetrou, E.P. Patient-Derived Induced Pluripotent Stem Cells in Cancer Research and Precision Oncology. Nat. Med. 2016, 22, 1392. [Google Scholar] [CrossRef]
- Simeonova, I.; Huillard, E. In Vivo Models of Brain Tumors: Roles of Genetically Engineered Mouse Models in Understanding Tumor Biology and Use in Preclinical Studies. Cell. Mol. Life Sci. 2014, 71, 4007. [Google Scholar] [CrossRef]
- Webber, M.M.; Bello, D.; Quader, S. Immortalized and Tumorigenic Adult Human Prostatic Epithelial Cell Lines: Characteristics and Applications Part 2. Tumorigenic Cell Lines. Prostate 1997, 30, 58–64. [Google Scholar] [CrossRef]
- Lindström, H.J.G.; Friedman, R. Inferring Time-Dependent Population Growth Rates in Cell Cultures Undergoing Adaptation. BMC Bioinform. 2020, 21, 583. [Google Scholar] [CrossRef]
- Harrison, N.J.; Baker, D.; Andrews, P.W. The Significance of Culture Adaptation of Embryonic Stem Cells for Regenerative Medicine. In Advances in Stem Cell Research; Springer: Cham, Switzerland, 2012; pp. 17–27. [Google Scholar] [CrossRef]
- Jahanban-Esfahlan, R.; Seidi, K.; Monhemi, H.; Adli, A.D.F.; Minofar, B.; Zare, P.; Farajzadeh, D.; Farajnia, S.; Behzadi, R.; Abbasi, M.M.; et al. RGD Delivery of Truncated Coagulase to Tumor Vasculature Affords Local Thrombotic Activity to Induce Infarction of Tumors in Mice. Sci. Rep. 2017, 7, 8126. [Google Scholar] [CrossRef]
- Hanahan, D.; Coussens, L.M. Accessories to the Crime: Functions of Cells Recruited to the Tumor Microenvironment. Cancer Cell 2012, 21, 309–322. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.L.; Liang, C.Y.; Ritz, D.; Coelho, R.; Septiadi, D.; Estermann, M.; Cumin, C.; Rimmer, N.; Schötzau, A.; López, M.N.; et al. Collagen-Rich Omentum Is a Premetastatic Niche for Integrin A2-Mediated Peritoneal Metastasis. eLife 2020, 9, e59442. [Google Scholar] [CrossRef] [PubMed]
- Langhans, S.A. Three-Dimensional in Vitro Cell Culture Models in Drug Discovery and Drug Repositioning. Front. Pharmacol. 2018, 9, 334617. [Google Scholar] [CrossRef]
- Edmondson, R.; Broglie, J.J.; Adcock, A.F.; Yang, L. Three-Dimensional Cell Culture Systems and Their Applications in Drug Discovery and Cell-Based Biosensors. Assay Drug Dev. Technol. 2014, 12, 207–218. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Koo, B.K.; Knoblich, J.A. Human Organoids: Model Systems for Human Biology and Medicine. Nat. Rev. Mol. Cell Biol. 2020, 21, 571–584. [Google Scholar] [CrossRef]
- Huang, Y.; Huang, Z.; Tang, Z.; Chen, Y.; Huang, M.; Liu, H.; Huang, W.; Ye, Q.; Jia, B. Research Progress, Challenges, and Breakthroughs of Organoids as Disease Models. Front. Cell Dev. Biol. 2021, 9, 3259. [Google Scholar] [CrossRef]
- Krieger, T.G.; Tirier, S.M.; Park, J.; Jechow, K.; Eisemann, T.; Peterziel, H.; Angel, P.; Eils, R.; Conrad, C. Modeling Glioblastoma Invasion Using Human Brain Organoids and Single-Cell Transcriptomics. Neuro Oncol. 2020, 22, 1138. [Google Scholar] [CrossRef]
- Huang, M.; Xu, S.; Li, Y.; Shang, L.; Zhan, X.; Qin, C.; Su, J.; Zhao, Z.; He, Y.; Qin, L.; et al. Novel Human Meningioma Organoids Recapitulate the Aggressiveness of the Initiating Cell Subpopulations Identified by ScRNA-Seq. Adv. Sci. 2023, 10, 2205525. [Google Scholar] [CrossRef]
- Liu, J.; Huang, X.; Huang, L.; Huang, J.; Liang, D.; Liao, L.; Deng, Y.; Zhang, L.; Zhang, B.; Tang, W. Organoid: Next-Generation Modeling of Cancer Research and Drug Development. Front. Oncol. 2022, 11, 5897. [Google Scholar] [CrossRef] [PubMed]
- Kirienko, N.V.; Mani, K.; Fay, D.S. Cancer Models in Caenorhabditis Elegans. Dev. Dyn. 2010, 239, 1413–1448. [Google Scholar] [CrossRef]
- Mirzoyan, Z.; Sollazzo, M.; Allocca, M.; Valenza, A.M.; Grifoni, D.; Bellosta, P. Drosophila Melanogaster: A Model Organism to Study Cancer. Front. Genet. 2019, 10, 51. [Google Scholar] [CrossRef] [PubMed]
- Bayens-Simmonds, J.; Boisvert, D.P.J.; Castro, M.E.; Johnson, E.S. A Feline Model for Experimental Studies of Peritumor Brain Edema. J. Neurooncol. 1988, 6, 371–378. [Google Scholar] [CrossRef]
- Hicks, J.; Platt, S.; Kent, M.; Haley, A. Canine Brain Tumours: A Model for the Human Disease? Vet. Comp. Oncol. 2017, 15, 252–272. [Google Scholar] [CrossRef] [PubMed]
- Xia, H.J.; Chen, C.S. Progress of Non-Human Primate Animal Models of Cancers. Dongwuxue Yanjiu 2011, 32, 70–80. [Google Scholar] [CrossRef]
- Huszthy, P.C.; Daphu, I.; Niclou, S.P.; Stieber, D.; Nigro, J.M.; Sakariassen, P.O.; Miletic, H.; Thorsen, F.; Bjerkvig, R. In Vivo Models of Primary Brain Tumors: Pitfalls and Perspectives. Neuro Oncol. 2012, 14, 979–993. [Google Scholar] [CrossRef]
- Tratar, U.L.; Horvat, S.; Cemazar, M. Transgenic Mouse Models in Cancer Research. Front. Oncol. 2018, 8, 268. [Google Scholar] [CrossRef]
- Davies, B.; Morris, T. Physiological Parameters in Laboratory Animals and Humans. Pharm. Res. 1993, 10, 1093–1095. [Google Scholar] [CrossRef]
- Hetze, S.; Sure, U.; Schedlowski, M.; Hadamitzky, M.; Barthel, L. Rodent Models to Analyze the Glioma Microenvironment. ASN Neuro 2021, 13, 17590914211005074. [Google Scholar] [CrossRef]
- Terpstra, A.H.M. Differences between Humans and Mice in Efficacy of the Body Fat Lowering Effect of Conjugated Linoleic Acid: Role of Metabolic Rate. J. Nutr. 2001, 131, 2067–2068. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Li, Y.; Yao, T.; Jia, R. Benefits of Zebrafish Xenograft Models in Cancer Research. Front. Cell Dev. Biol. 2021, 9, 616551. [Google Scholar] [CrossRef] [PubMed]
- Choi, T.Y.; Choi, T.I.; Lee, Y.R.; Choe, S.K.; Kim, C.H. Zebrafish as an Animal Model for Biomedical Research. Exp. Mol. Med. 2021, 53, 310–317. [Google Scholar] [CrossRef] [PubMed]
- Richter-Pechańska, P.; Kunz, J.B.; Bornhauser, B.; von Knebel Doeberitz, C.; Rausch, T.; Erarslan-Uysal, B.; Assenov, Y.; Frismantas, V.; Marovca, B.; Waszak, S.M.; et al. PDX Models Recapitulate the Genetic and Epigenetic Landscape of Pediatric T-Cell Leukemia. EMBO Mol. Med. 2018, 10, e9443. [Google Scholar] [CrossRef] [PubMed]
- Roussel, M.F.; Stripay, J.L. Modeling Pediatric Medulloblastoma. Brain Pathol. 2020, 30, 703–712. [Google Scholar] [CrossRef] [PubMed]
- Olson, B.; Li, Y.; Lin, Y.; Liu, E.T.; Patnaik, A. Mouse Models for Cancer Immunotherapy Research. Cancer Discov. 2018, 8, 1358. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Z.; Wong, C.J.; Yang, L.; Ouardaoui, N.; Li, D.; Zhang, W.; Gu, S.; Zhang, Y.; Liu, Y.; Wang, X.; et al. TISMO: Syngeneic Mouse Tumor Database to Model Tumor Immunity and Immunotherapy Response. Nucleic Acids Res. 2022, 50, D1391–D1397. [Google Scholar] [CrossRef] [PubMed]
- Becher, O.J.; Holland, E.C. Genetically Engineered Models Have Advantages over Xenografts for Preclinical Studies. Cancer Res. 2006, 66, 3355–3359. [Google Scholar] [CrossRef] [PubMed]
- Sharpless, N.E.; DePinho, R.A. The Mighty Mouse: Genetically Engineered Mouse Models in Cancer Drug Development. Nat. Rev. Drug Discov. 2006, 5, 741–754. [Google Scholar] [CrossRef]
- Hamilton, F. An Account of the Fishes Found in the River Ganges and Its Branches; Archibald Constable and Company: Edinburgh, Scotland, 1822; Volume 1. [Google Scholar]
- Parichy, D.M. The Natural History of Model Organisms: Advancing Biology through a Deeper Understanding of Zebrafish Ecology and Evolution. eLife 2015. [Google Scholar] [CrossRef]
- Kalueff, A.V.; Stewart, A.M.; Gerlai, R. Zebrafish as an Emerging Model for Studying Complex Brain Disorders. Trends Pharmacol. Sci. 2014, 35, 63. [Google Scholar] [CrossRef] [PubMed]
- Hsu, C.-H.; Wen, Z.-H.; Lin, C.-S.; Chakraborty, C. The Zebrafish Model: Use in Studying Cellular Mechanisms for a Spectrum of Clinical Disease Entities. Curr. Neurovasc. Res. 2007, 4, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Howe, K.; Clark, M.D.; Torroja, C.F.; Torrance, J.; Berthelot, C.; Muffato, M.; Collins, J.E.; Humphray, S.; McLaren, K.; Matthews, L.; et al. The Zebrafish Reference Genome Sequence and Its Relationship to the Human Genome. Nature 2013, 496, 498–503. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Luan, Y.; Liu, T.; Lee, H.J.; Fang, L.; Wang, Y.; Wang, X.; Zhang, B.; Jin, Q.; Ang, K.C.; et al. A Map of Cis-Regulatory Elements and 3D Genome Structures in Zebrafish. Nature 2020, 588, 337–343. [Google Scholar] [CrossRef] [PubMed]
- Bernier, R.; Golzio, C.; Xiong, B.; Stessman, H.A.; Coe, B.P.; Penn, O.; Witherspoon, K.; Gerdts, J.; Baker, C.; Vulto-Van Silfhout, A.T.; et al. Disruptive CHD8 Mutations Define a Subtype of Autism Early in Development. Cell 2014, 158, 263–276. [Google Scholar] [CrossRef] [PubMed]
- Seth, A.; Stemple, D.L.; Barroso, I. The Emerging Use of Zebrafish to Model Metabolic Disease. Dis. Models Mech. 2013, 6, 1080. [Google Scholar] [CrossRef] [PubMed]
- de Abreu, M.S.; Genario, R.; Giacomini, A.C.V.V.; Demin, K.A.; Lakstygal, A.M.; Amstislavskaya, T.G.; Fontana, B.D.; Parker, M.O.; Kalueff, A.V. Zebrafish as a Model of Neurodevelopmental Disorders. Neuroscience 2020, 445, 3–11. [Google Scholar] [CrossRef]
- Meshalkina, D.A.; Kizlyk, M.N.; Kysil, E.V.; Collier, A.D.; Echevarria, D.J.; Abreu, M.S.; Barcellos, L.J.G.; Song, C.; Warnick, J.E.; Kyzar, E.J.; et al. Zebrafish Models of Autism Spectrum Disorder. Exp. Neurol. 2018, 299, 207–216. [Google Scholar] [CrossRef]
- Rea, V.; Van Raay, T.J. Using Zebrafish to Model Autism Spectrum Disorder: A Comparison of ASD Risk Genes Between Zebrafish and Their Mammalian Counterparts. Front. Mol. Neurosci. 2020, 13, 207. [Google Scholar] [CrossRef]
- Nozawa, K.; Lin, Y.; Kubodera, R.; Shimizu, Y.; Tanaka, H.; Ohshima, T. Zebrafish Mecp2 Is Required for Proper Axonal Elongation of Motor Neurons and Synapse Formation. Dev. Neurobiol. 2017, 77, 1101–1113. [Google Scholar] [CrossRef]
- Pietri, T.; Roman, A.C.; Guyon, N.; Romano, S.A.; Washbourne, P.; Moens, C.B.; de Polavieja, G.G.; Sumbre, G. The First Mecp2-Null Zebrafish Model Shows Altered Motor Behaviors. Front. Neural Circuits 2013, 7, 118. [Google Scholar] [CrossRef] [PubMed]
- Chia, K.; Klingseisen, A.; Sieger, D.; Priller, J. Zebrafish as a Model Organism for Neurodegenerative Disease. Front. Mol. Neurosci. 2022, 15, 376. [Google Scholar] [CrossRef] [PubMed]
- Blaser, R.E.; Chadwick, L.; McGinnis, G.C. Behavioral Measures of Anxiety in Zebrafish (Danio Rerio). Behav. Brain Res. 2010, 208, 56–62. [Google Scholar] [CrossRef] [PubMed]
- Norton, W.; Bally-Cuif, L. Adult Zebrafish as a Model Organism for Behavioural Genetics. BMC Neurosci. 2010, 11, 90. [Google Scholar] [CrossRef] [PubMed]
- Singh, J.; Patten, S.A. Modeling Neuromuscular Diseases in Zebrafish. Front. Mol. Neurosci. 2022, 15, 698. [Google Scholar] [CrossRef] [PubMed]
- Sztal, T.E.; Sonntag, C.; Hall, T.E.; Currie, P.D. Epistatic Dissection of Laminin-Receptor Interactions in Dystrophic Zebrafish Muscle. Hum. Mol. Genet. 2012, 21, 4718–4731. [Google Scholar] [CrossRef] [PubMed]
- Parsons, M.J.; Campos, I.; Hirst, E.M.A.; Stemple, D.L. Removal of Dystroglycan Causes Severe Muscular Dystrophy in Zebrafish Embryos. Development 2002, 129, 3505–3512. [Google Scholar] [CrossRef]
- Bolaos-Jiménez, F.; Bordais, A.; Behra, M.; Strähle, U.; Sahel, J.; Rendón, A. Dystrophin and Dp71, Two Products of the DMD Gene, Show a Different Pattern of Expression during Embryonic Development in Zebrafish. Mech. Dev. 2001, 102, 239–241. [Google Scholar] [CrossRef]
- Bassett, D.I.; Currie, P.D. The Zebrafish as a Model for Muscular Dystrophy and Congenital Myopathy. Hum. Mol. Genet. 2003, 12 (Suppl. 2), R265–R270. [Google Scholar] [CrossRef]
- Arbones, M.L.; Thomazeau, A.; Nakano-Kobayashi, A.; Hagiwara, M.; Delabar, J.M. DYRK1A and Cognition: A Lifelong Relationship. Pharmacol. Ther. 2019, 194, 199–221. [Google Scholar] [CrossRef]
- Liang, S.T.; Audira, G.; Juniardi, S.; Chen, J.R.; Lai, Y.H.; Du, Z.C.; Lin, D.S.; Hsiao, C. Der. Zebrafish Carrying Pycr1 Gene Deficiency Display Aging and Multiple Behavioral Abnormalities. Cells 2019, 8, 453. [Google Scholar] [CrossRef]
- Anchelin, M.; Murcia, L.; Alcaraz-Pérez, F.; García-Navarro, E.M.; Cayuela, M.L. Behaviour of Telomere and Telomerase during Aging and Regeneration in Zebrafish. PLoS ONE 2011, 6, e16955. [Google Scholar] [CrossRef]
- Gore, A.V.; Pillay, L.M.; Venero Galanternik, M.; Weinstein, B.M. The Zebrafish: A Fintastic Model for Hematopoietic Development and Disease. Wiley Interdiscip. Rev. Dev. Biol. 2018, 7, e312. [Google Scholar] [CrossRef] [PubMed]
- Rasighaemi, P.; Basheer, F.; Liongue, C.; Ward, A.C. Zebrafish as a Model for Leukemia and Other Hematopoietic Disorders. J. Hematol. Oncol. 2015, 8, 29. [Google Scholar] [CrossRef] [PubMed]
- Berman, J.; Hsu, K.; Look, A.T. Zebrafish as a Model Organism for Blood Diseases. Br. J. Haematol. 2003, 123, 568–576. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Yu, S.H.; Zhen, W.X.; Cheng, T.; Wang, D.; Lin, J.B.; Wu, Y.H.; Wang, Y.F.; Chen, Y.; Shu, L.P.; et al. Tanshinone I, a New EZH2 Inhibitor Restricts Normal and Malignant Hematopoiesis through Upregulation of MMP9 and ABCG2. Theranostics 2021, 11, 6891–6904. [Google Scholar] [CrossRef]
- Outtandy, P.; Russell, C.; Kleta, R.; Bockenhauer, D. Zebrafish as a Model for Kidney Function and Disease. Pediatr. Nephrol. 2019, 34, 751–762. [Google Scholar] [CrossRef]
- Poureetezadi, S.J.; Wingert, R.A. Little Fish, Big Catch: Zebrafish as a Model for Kidney Disease. Kidney Int. 2016, 89, 1204–1210. [Google Scholar] [CrossRef]
- Dang, L.; Cao, X.; Zhang, T.; Sun, Y.; Tian, S.; Gong, T.; Xiong, H.; Cao, P.; Li, Y.; Yu, S.; et al. Nuclear Condensation of CDYL Links Histone Crotonylation and Cystogenesis in Autosomal Dominant Polycystic Kidney Disease. J. Am. Soc. Nephrol. 2022, 33, 1708–1725. [Google Scholar] [CrossRef]
- Devane, J.; Ott, E.; Olinger, E.G.; Epting, D.; Decker, E.; Friedrich, A.; Bachmann, N.; Renschler, G.; Eisenberger, T.; Briem-Richter, A.; et al. Progressive Liver, Kidney, and Heart Degeneration in Children and Adults Affected by TULP3 Mutations. Am. J. Hum. Genet. 2022, 109, 928–943. [Google Scholar] [CrossRef]
- Ko, S.; Russell, J.O.; Tian, J.; Gao, C.; Kobayashi, M.; Feng, R.; Yuan, X.; Shao, C.; Ding, H.; Poddar, M.; et al. Hdac1 Regulates Differentiation of Bipotent Liver Progenitor Cells During Regeneration via Sox9b and Cdk8. Gastroenterology 2019, 156, 187–202.e14. [Google Scholar] [CrossRef] [PubMed]
- de Oliveira, S.; Houseright, R.A.; Graves, A.L.; Golenberg, N.; Korte, B.G.; Miskolci, V.; Huttenlocher, A. Metformin Modulates Innate Immune-Mediated Inflammation and Early Progression of NAFLD-Associated Hepatocellular Carcinoma in Zebrafish. J. Hepatol. 2019, 70, 710–721. [Google Scholar] [CrossRef] [PubMed]
- Goessling, W.; Sadler, K.C. Zebrafish: An Important Tool for Liver Disease Research. Gastroenterology 2015, 149, 1361. [Google Scholar] [CrossRef] [PubMed]
- Sehgal, P.; Mathew, S.; Sivadas, A.; Ray, A.; Tanwar, J.; Vishwakarma, S.; Ranjan, G.; Shamsudheen, K.V.; Bhoyar, R.C.; Pateria, A.; et al. LncRNA VEAL2 Regulates PRKCB2 to Modulate Endothelial Permeability in Diabetic Retinopathy. EMBO J. 2021, 40, e107134. [Google Scholar] [CrossRef] [PubMed]
- Wohlfart, D.P.; Lou, B.; Middel, C.S.; Morgenstern, J.; Fleming, T.; Sticht, C.; Hausser, I.; Hell, R.; Hammes, H.P.; Szendrödi, J.; et al. Accumulation of Acetaldehyde in Aldh2.1-/- Zebrafish Causes Increased Retinal Angiogenesis and Impaired Glucose Metabolism. Redox Biol. 2022, 50, 102249. [Google Scholar] [CrossRef] [PubMed]
- Ping, X.; Liang, J.; Shi, K.; Bao, J.; Wu, J.; Yu, X.; Tang, X.; Zou, J.; Shentu, X. Rapamycin Relieves the Cataract Caused by Ablation of Gja8b through Stimulating Autophagy in Zebrafish. Autophagy 2021, 17, 3323–3337. [Google Scholar] [CrossRef] [PubMed]
- Chhetri, J.; Jacobson, G.; Gueven, N. Zebrafish—On the Move towards Ophthalmological Research. Eye 2014, 28, 367–380. [Google Scholar] [CrossRef] [PubMed]
- Maleski, A.L.A.; Rosa, J.G.S.; Bernardo, J.T.G.; Astray, R.M.; Walker, C.I.B.; Lopes-Ferreira, M.; Lima, C. Recapitulation of Retinal Damage in Zebrafish Larvae Infected with Zika Virus. Cells 2022, 11, 1457. [Google Scholar] [CrossRef] [PubMed]
- Tyrkalska, S.D.; Candel, S.; Pedoto, A.; García-Moreno, D.; Alcaraz-Pérez, F.; Sánchez-Ferrer, Á.; Cayuela, M.L.; Mulero, V. Zebrafish Models of COVID-19. FEMS Microbiol. Rev. 2023, 47, fuac042. [Google Scholar] [CrossRef]
- Stanton, M.F. Diethylnitrosamine-Induced Hepatic Degeneration and Neoplasia in the Aquarium Fish, Brachydanio Rerio. JNCI J. Natl. Cancer Inst. 1965, 34, 117–130. [Google Scholar] [CrossRef]
- Haffter, P.; Granato, M.; Brand, M.; Mullins, M.C.; Hammerschmidt, M.; Kane, D.A.; Odenthal, J.; Van Eeden, F.J.M.; Jiang, Y.J.; Heisenberg, C.P.; et al. The Identification of Genes with Unique and Essential Functions in the Development of the Zebrafish, Danio rerio. Development 1996, 123, 1–36. [Google Scholar] [CrossRef] [PubMed]
- Beckwith, L.G.; Moore, J.L.; Tsao-Wu, G.S.; Harshbarger, J.C.; Cheng, K.C. Ethylnitrosourea Induces Neoplasia in Zebrafish (Danio rerio). Lab. Investig. 2000, 80, 379–385. [Google Scholar] [CrossRef] [PubMed]
- Spitsbergen, J.M.; Tsai, H.W.; Reddy, A.; Miller, T.; Arbogast, D.; Hendricks, J.D.; Bailey, G.S. Neoplasia in Zebrafish (Danio rerio) Treated with N-Methyl-N’nitro-N-Nitrosoguanidine by Three Exposure Routes at Different Developmental Stages. Toxicol. Pathol. 2000, 28, 716–725. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q. Cancer Predisposition Genes: Molecular Mechanisms and Clinical Impact on Personalized Cancer Care: Examples of Lynch and HBOC Syndromes. Acta Pharmacol. Sin. 2016, 37, 143. [Google Scholar] [CrossRef] [PubMed]
- Rahman, N. Realizing the Promise of Cancer Predisposition Genes. Nature 2014, 505, 302–308. [Google Scholar] [CrossRef] [PubMed]
- Kobar, K.; Collett, K.; Prykhozhij, S.V.; Berman, J.N. Zebrafish Cancer Predisposition Models. Front. Cell Dev. Biol. 2021, 9, 660069. [Google Scholar] [CrossRef] [PubMed]
- Gröbner, S.N.; Worst, B.C.; Weischenfeldt, J.; Buchhalter, I.; Kleinheinz, K.; Rudneva, V.A.; Johann, P.D.; Balasubramanian, G.P.; Segura-Wang, M.; Brabetz, S.; et al. The Landscape of Genomic Alterations across Childhood Cancers. Nature 2018, 555, 321–327. [Google Scholar] [CrossRef]
- Cunningham, R.M.; Walton, M.A.; Carter, P.M. The Major Causes of Death in Children and Adolescents in the United States. N. Engl. J. Med. 2018, 379, 2468. [Google Scholar] [CrossRef]
- Ward, E.; Desantis, C.; Robbins, A.; Kohler, B.; Jemal, A. Childhood and Adolescent Cancer Statistics, 2014. CA Cancer J. Clin. 2014, 64, 83–103. [Google Scholar] [CrossRef]
- Mariotto, E.; Corallo, D.; Pantile, M.; Giarin, E.; Pigazzi, M.; Basso, G.; Viola, G.; Aveic, S. BAG1 Down-Regulation Increases Chemo-Sensitivity of Acute Lymphoblastic Leukaemia Cells. J. Cell. Mol. Med. 2021, 25, 9060–9065. [Google Scholar] [CrossRef]
- Raetz, E.A.; Teachey, D.T. T-Cell Acute Lymphoblastic Leukemia. Hematol. Am. Soc. Hematol. Educ. Program 2016, 2016, 580. [Google Scholar] [CrossRef] [PubMed]
- Burns, M.A.; Liao, Z.W.; Yamagata, N.; Pouliot, G.P.; Stevenson, K.E.; Neuberg, D.S.; Thorner, A.R.; Ducar, M.; Silverman, E.A.; Hunger, S.P.; et al. Hedgehog Pathway Mutations Drive Oncogenic Transformation in High-Risk T-Cell Acute Lymphoblastic Leukemia. Leukemia 2018, 32, 2126. [Google Scholar] [CrossRef] [PubMed]
- Martin-Giacalone, B.A.; Weinstein, P.A.; Plon, S.E.; Lupo, P.J. Pediatric Rhabdomyosarcoma: Epidemiology and Genetic Susceptibility. J. Clin. Med. 2021, 10, 2028. [Google Scholar] [CrossRef] [PubMed]
- McEvoy, M.T.; Siegel, D.A.; Dai, S.; Okcu, M.F.; Zobeck, M.; Venkatramani, R.; Lupo, P.J. Pediatric Rhabdomyosarcoma Incidence and Survival in the United States: An Assessment of 5656 Cases, 2001–2017. Cancer Med. 2023, 12, 3644. [Google Scholar] [CrossRef] [PubMed]
- Campbell, B.B.; Galati, M.A.; Stone, S.C.; Riemenschneider, A.N.; Edwards, M.; Sudhaman, S.; Siddaway, R.; Komosa, M.; Nunes, N.M.; Nobre, L.; et al. Mutations in the RAS/MAPK Pathway Drive Replication Repair–Deficient Hypermutated Tumors and Confer Sensitivity to MEK Inhibition. Cancer Discov. 2021, 11, 1454. [Google Scholar] [CrossRef] [PubMed]
- Kahsay, A.; Rodriguez-Marquez, E.; López-Pérez, A.; Hörnblad, A.; von Hofsten, J. Pax3 Loss of Function Delays Tumour Progression in KRAS-Induced Zebrafish Rhabdomyosarcoma Models. Sci. Rep. 2022, 12, 17149. [Google Scholar] [CrossRef] [PubMed]
- Kent, M.R.; Calderon, D.; Silvius, K.M.; Kucinski, J.P.; LaVigne, C.A.; Cannon, M.V.; Kendall, G.C. Zebrafish Her3 Knockout Impacts Developmental and Cancer-Related Gene Signatures. Dev. Biol. 2023, 496, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Daakour, S.; Hajingabo, L.J.; Kerselidou, D.; Devresse, A.; Kettmann, R.; Simonis, N.; Dequiedt, F.; Twizere, J.C. Systematic Interactome Mapping of Acute Lymphoblastic Leukemia Cancer Gene Products Reveals EXT-1 Tumor Suppressor as a Notch1 and FBWX7 Common Interactor. BMC Cancer 2016, 16, 335. [Google Scholar] [CrossRef] [PubMed]
- Borga, C.; Park, G.; Foster, C.; Burroughs-Garcia, J.; Marchesin, M.; Shah, R.; Hasan, A.; Ahmed, S.T.; Bresolin, S.; Batchelor, L.; et al. Simultaneous B and T Cell Acute Lymphoblastic Leukemias in Zebrafish Driven by Transgenic MYC: Implications for Oncogenesis and Lymphopoiesis. Leukemia 2019, 33, 333–347. [Google Scholar] [CrossRef]
- Mandelbaum, J.; Shestopalov, I.A.; Henderson, R.E.; Chau, N.G.; Knoechel, B.; Wick, M.J.; Zon, L.I. Zebrafish Blastomere Screen Identifies Retinoic Acid Suppression of MYB in Adenoid Cystic Carcinoma. J. Exp. Med. 2018, 215, 2673–2685. [Google Scholar] [CrossRef]
- Xu, M.; Ye, Y.; Ye, Z.; Xu, S.; Liu, W.; Xu, J.; Zhang, Y.; Liu, Q.; Huang, Z.; Zhang, W. Human BCR/ABL1 Induces Chronic Myeloid Leukemia-like Disease in Zebrafish. Haematologica 2020, 105, 674. [Google Scholar] [CrossRef]
- Somasagara, R.R.; Huang, X.; Xu, C.; Haider, J.; Serody, J.S.; Armistead, P.M.; Leung, T.C. Targeted Therapy of Human Leukemia Xenografts in Immunodeficient Zebrafish. Sci. Rep. 2021, 11, 5715. [Google Scholar] [CrossRef] [PubMed]
- Rosenbluh, J.; Nijhawan, D.; Cox, A.G.; Li, X.; Neal, J.T.; Schafer, E.J.; Zack, T.I.; Wang, X.; Tsherniak, A.; Schinzel, A.C.; et al. β-Catenin Driven Cancers Require a YAP1 Transcriptional Complex for Survival and Tumorigenesis. Cell 2012, 151, 1457. [Google Scholar] [CrossRef] [PubMed]
- Meng, Y.; Du, M.; Gu, D.; Li, C.; Li, S.; Zhang, Q.; Ben, S.; Zhu, Q.; Xin, J.; Zhang, Z.; et al. Genome-Wide Association Analyses Identify CATSPERE as a Mediator of Colorectal Cancer Susceptibility and Progression. Cancer Res. 2022, 82, 986–997. [Google Scholar] [CrossRef]
- Maradonna, F.; Fontana, C.M.; Sella, F.; Giommi, C.; Facchinello, N.; Rampazzo, C.; Caichiolo, M.; Hoseinifar, S.H.; Dalla Valle, L.; Van Doan, H.; et al. A Zebrafish HCT116 Xenograft Model to Predict Anandamide Outcomes on Colorectal Cancer. Cell Death Dis. 2022, 13, 1069. [Google Scholar] [CrossRef] [PubMed]
- Noonan, H.R.; Metelo, A.M.; Kamei, C.N.; Peterson, R.T.; Drummond, I.A.; Iliopoulos, O. Loss of Vhl in the Zebrafish Pronephros Recapitulates Early Stages of Human Clear Cell Renal Cell Carcinoma. Dis. Models Mech. 2016, 9, 873–884. [Google Scholar] [CrossRef]
- Luo, J.; Lu, C.; Feng, M.; Dai, L.; Wang, M.; Qiu, Y.; Zheng, H.; Liu, Y.; Li, L.; Tang, B.; et al. Cooperation between Liver-Specific Mutations of Pten and Tp53 Genetically Induces Hepatocarcinogenesis in Zebrafish. J. Exp. Clin. Cancer Res. 2021, 40, 262. [Google Scholar] [CrossRef] [PubMed]
- Kaufman, C.K.; Mosimann, C.; Fan, Z.P.; Yang, S.; Thomas, A.J.; Ablain, J.; Tan, J.L.; Fogley, R.D.; Van Rooijen, E.; Hagedorn, E.J.; et al. A Zebrafish Melanoma Model Reveals Emergence of Neural Crest Identity during Melanoma Initiation. Science 2016, 351, aad2197. [Google Scholar] [CrossRef]
- van der Ent, W.; Burrello, C.; Teunisse, A.F.A.S.; Ksander, B.R.; van der Velden, P.A.; Jager, M.J.; Jochemsen, A.G.; Snaar-Jagalska, B.E. Modeling of Human Uveal Melanoma in Zebrafish Xenograft Embryos. Investig. Ophthalmol. Vis. Sci. 2014, 55, 6612–6622. [Google Scholar] [CrossRef]
- Travnickova, J.; Wojciechowska, S.; Khamseh, A.; Gautier, P.; Brown, D.V.; Lefevre, T.; Brombin, A.; Ewing, A.; Capper, A.; Spitzer, M.; et al. Zebrafish MITF-Low Melanoma Subtype Models Reveal Transcriptional Subclusters and MITF-Independent Residual Disease. Cancer Res. 2019, 79, 5769–5784. [Google Scholar] [CrossRef]
- Santoriello, C.; Gennaro, E.; Anelli, V.; Distel, M.; Kelly, A.; Köster, R.W.; Hurlstone, A.; Mione, M. Kita Driven Expression of Oncogenic HRAS Leads to Early Onset and Highly Penetrant Melanoma in Zebrafish. PLoS ONE 2010, 5, e15170. [Google Scholar] [CrossRef] [PubMed]
- Gjini, E.; Mansour, M.R.; Sander, J.D.; Moritz, N.; Nguyen, A.T.; Kesarsing, M.; Gans, E.; He, S.; Chen, S.; Ko, M.; et al. A Zebrafish Model of Myelodysplastic Syndrome Produced through Tet2 Genomic Editing. Mol. Cell. Biol. 2015, 35, 789. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Wu, M.; Huang, Z.; Lian, J.; Chen, J.; Wang, T.; Leung, A.Y.H.; Liao, Y.; Zhang, Z.; Liu, Q.; et al. C-Myb Hyperactivity Leads to Myeloid and Lymphoid Malignancies in Zebrafish. Leukemia 2017, 31, 222. [Google Scholar] [CrossRef] [PubMed]
- Nissim, S.; Leshchiner, I.; Mancias, J.D.; Greenblatt, M.B.; Maertens, O.; Cassa, C.A.; Rosenfeld, J.A.; Cox, A.G.; Hedgepeth, J.; Wucherpfennig, J.I.; et al. Mutations in RABL3 Alter KRAS Prenylation and Are Associated with Hereditary Pancreatic Cancer. Nat. Genet. 2019, 51, 1308. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.; Park, J.T. Zebrafish Model of KRAS-Initiated Pancreatic Endocrine Tumor. Anim. Cells Syst. 2019, 23, 209. [Google Scholar] [CrossRef] [PubMed]
- Barroso, M.T.; Costa, B.; de Almeida, C.R.; Martin, M.C.; Couto, N.; Carvalho, T.; Fior, R. Establishment of Pancreatobiliary Cancer Zebrafish Avatars for Chemotherapy Screening. Cells 2021, 10, 2077. [Google Scholar] [CrossRef] [PubMed]
- Brandt, Z.J.; North, P.N.; Link, B.A. Somatic Mutations of Lats2 Cause Peripheral Nerve Sheath Tumors in Zebrafish. Cells 2019, 8, 972. [Google Scholar] [CrossRef]
- Kochat, V.; Raman, A.T.; Landers, S.M.; Tang, M.; Schulz, J.; Terranova, C.; Landry, J.P. Enhancer Reprogramming in PRC2-Deficient Malignant Peripheral Nerve Sheath Tumors Induces a Targetable de-Differentiated State. Acta Neuropathol. 2021, 142, 565–591. [Google Scholar] [CrossRef]
- Asnaghi, L.; White, D.T.; Key, N.; Choi, J.; Mahale, A.; Alkatan, H.; Edward, D.P.; Elkhamary, S.M.; Al-Mesfer, S.; Maktabi, A.; et al. ACVR1C/SMAD2 Signaling Promotes Invasion and Growth in Retinoblastoma. Oncogene 2019, 38, 2056. [Google Scholar] [CrossRef]
- Storer, N.Y.; White, R.M.; Uong, A.; Price, E.; Nielsen, G.P.; Langenau, D.M.; Zon, L.I. Zebrafish Rhabdomyosarcoma Reflects the Developmental Stage of Oncogene Expression during Myogenesis. Development 2013, 140, 3040–3050. [Google Scholar] [CrossRef]
- Kendall, G.C.; Watson, S.; Xu, L.; Lavigne, C.A.; Murchison, W.; Rakheja, D.; Skapek, S.X.; Tirode, F.; Delattre, O.; Amatruda, J.F. PAX3-FOXO1 Transgenic Zebrafish Models Identify HES3 as a Mediator of Rhabdomyosarcoma Tumorigenesis. eLife 2018, 7, e33800. [Google Scholar] [CrossRef]
- Balci, T.B.; Prykhozhij, S.V.; Teh, E.M.; Da’as, S.I.; Mcbride, E.; Liwski, R.; Chute, I.C.; Leger, D.; Lewis, S.M.; Berman, J.N. A Transgenic Zebrafish Model Expressing KIT-D816V Recapitulates Features of Aggressive Systemic Mastocytosis. Br. J. Haematol. 2014, 167, 48–61. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, M.L.; Veloso, A.; Garcia, E.G.; Iyer, S.; Pereira, C.; Barreto, V.M.; Langenau, D.M.; Barata, J.T. Mutant IL7R Collaborates with MYC to Induce T-Cell Acute Lymphoblastic Leukemia. Leukemia 2022, 36, 1533. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.; Wang, J.; Yue, M.; Cai, X.; Wang, T.; Wu, C.; Su, H.; Wang, Y.; Han, M.; Zhang, Y.; et al. Direct Phosphorylation and Stabilization of MYC by Aurora B Kinase Promote T-Cell Leukemogenesis. Cancer Cell 2020, 37, 200. [Google Scholar] [CrossRef]
- Garcia, E.G.; Veloso, A.; Oliveira, M.L.; Allen, J.R.; Loontiens, S.; Brunson, D.; Do, D.; Yan, C.; Morris, R.; Iyer, S.; et al. PRL3 Enhances T-Cell Acute Lymphoblastic Leukemia Growth through Suppressing T-Cell Signaling Pathways and Apoptosis. Leukemia 2021, 35, 679–690. [Google Scholar] [CrossRef] [PubMed]
- Kindermann, B.; Valkova, C.; Krämer, A.; Perner, B.; Engelmann, C.; Behrendt, L.; Kritsch, D.; Jungnickel, B.; Kehlenbach, R.H.; Oswald, F.; et al. The Nuclear Pore Proteins Nup88/214 and T-Cell Acute Lymphatic Leukemia–Associated NUP214 Fusion Proteins Regulate Notch Signaling. J. Biol. Chem. 2019, 294, 11741. [Google Scholar] [CrossRef]
- Basten, S.G.; Davis, E.E.; Gillis, A.J.M.; van Rooijen, E.; Stoop, H.; Babala, N.; Logister, I.; Heath, Z.G.; Jonges, T.N.; Katsanis, N.; et al. Mutations in LRRC50 Predispose Zebrafish and Humans to Seminomas. PLoS Genet. 2013, 9, e1003384. [Google Scholar] [CrossRef] [PubMed]
- Anelli, V.; Villefranc, J.A.; Chhangawala, S.; Martinez-Mcfaline, R.; Riva, E.; Nguyen, A.; Verma, A.; Bareja, R.; Chen, Z.; Scognamiglio, T.; et al. Oncogenic BRAF Disrupts Thyroid Morphogenesis and Function via Twist Expression. eLife 2017, 6, e20728. [Google Scholar] [CrossRef] [PubMed]
- Pan, Z.; Xu, T.; Bao, L.; Hu, X.; Jin, T.; Chen, J.; Chen, J.; Qian, Y.; Lu, X.; Li, L.; et al. CREB3L1 Promotes Tumor Growth and Metastasis of Anaplastic Thyroid Carcinoma by Remodeling the Tumor Microenvironment. Mol. Cancer 2022, 21, 190. [Google Scholar] [CrossRef]
- Curtin, S.C.; Minino, A.M.; Anderson, R.N. Declines in Cancer Death Rates among Children and Adolescents in the United States, 1999–2014; Data Brief, no 257; National Center for Health Statistics: Hyattsville, MD, USA, 2016. [Google Scholar]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer Statistics, 2018. CA Cancer J. Clin. 2018, 68, 7–30. [Google Scholar] [CrossRef]
- Langova, V.; Vales, K.; Horka, P.; Horacek, J. The Role of Zebrafish and Laboratory Rodents in Schizophrenia Research. Front. Psychiatry 2020, 11, 703. [Google Scholar] [CrossRef] [PubMed]
- Ronneberger, O.; Liu, K.; Rath, M.; Rue, D.; Mueller, T.; Skibbe, H.; Drayer, B.; Schmidt, T.; Filippi, A.; Nitschke, R.; et al. ViBE-Z: A Framework for 3D Virtual Colocalization Analysis in Zebrafish Larval Brains. Nat. Methods 2012, 9, 735–742. [Google Scholar] [CrossRef] [PubMed]
- Friedrich, R.W.; Korsching, S.I. Combinatorial and Chemotopic Odorant Coding in the Zebrafish Olfactory Bulb Visualized by Optical Imaging. Neuron 1997, 18, 737–752. [Google Scholar] [CrossRef] [PubMed]
- Miyasaka, N.; Morimoto, K.; Tsubokawa, T.; Higashijima, S.I.; Okamoto, H.; Yoshihara, Y. From the Olfactory Bulb to Higher Brain Centers: Genetic Visualization of Secondary Olfactory Pathways in Zebrafish. J. Neurosci. 2009, 29, 4756–4767. [Google Scholar] [CrossRef] [PubMed]
- Teles, M.C.; Cardoso, S.D.; Oliveira, R.F. Social Plasticity Relies on Different Neuroplasticity Mechanisms across the Brain Social Decision-Making Network in Zebrafish. Front. Behav. Neurosci. 2016, 10, 16. [Google Scholar] [CrossRef] [PubMed]
- Stednitz, S.J.; McDermott, E.M.; Ncube, D.; Tallafuss, A.; Eisen, J.S.; Washbourne, P. Forebrain Control of Behaviorally Driven Social Orienting in Zebrafish. Curr. Biol. 2018, 28, 2445–2451.e3. [Google Scholar] [CrossRef]
- Lin, Q.; Jesuthasan, S. Masking of a Circadian Behavior in Larval Zebrafish Involves the Thalamo-Habenula Pathway. Sci. Rep. 2017, 7, 4104. [Google Scholar] [CrossRef]
- Redila, V.; Kinzel, C.; Jo, Y.S.; Puryear, C.B.; Mizumori, S.J.Y. A Role for the Lateral Dorsal Tegmentum in Memory and Decision Neural Circuitry. Neurobiol. Learn. Mem. 2015, 117, 93–108. [Google Scholar] [CrossRef]
- Lodge, D.J.; Grace, A.A. The Laterodorsal Tegmentum Is Essential for Burst Firing of Ventral Tegmental Area Dopamine Neurons. Proc. Natl. Acad. Sci. USA 2006, 103, 5167–5172. [Google Scholar] [CrossRef]
- Moens, C.B.; Prince, V.E. Constructing the Hindbrain: Insights from the Zebrafish. Dev. Dyn. 2002, 224, 1–17. [Google Scholar] [CrossRef]
- Oxtoby, E.; Jowett, T. Cloning of the Zebrafish Krox-20 Gene (Krx-20) and Its Expression during Hindbrain Development. Nucleic Acids Res. 1993, 21, 1087–1095. [Google Scholar] [CrossRef] [PubMed]
- Köster, R.W.; Fraser, S.E. Direct Imaging of in Vivo Neuronal Migration in the Developing Cerebellum. Curr. Biol. 2001, 11, 1858–1863. [Google Scholar] [CrossRef]
- Lalonde, R.; Botez, M.I. The Cerebellum and Learning Processes in Animals. Brain Res. Rev. 1990, 15, 325–332. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez, F.; Durán, E.; Gómez, A.; Ocaña, F.M.; Álvarez, E.; Jiménez-Moya, F.; Broglio, C.; Salas, C. Cognitive and Emotional Functions of the Teleost Fish Cerebellum. Brain Res. Bull. 2005, 66, 365–370. [Google Scholar] [CrossRef]
- Nakajima, R.; Hagihara, H.; Miyakawa, T. Similarities of Developmental Gene Expression Changes in the Brain between Human and Experimental Animals: Rhesus Monkey, Mouse, Zebrafish, and Drosophila. Mol. Brain 2021, 14, 135. [Google Scholar] [CrossRef]
- Barreiro-Iglesias, A.; Mysiak, K.S.; Adrio, F.; Rodicio, M.C.; Becker, C.G.; Becker, T.; Anadón, R. Distribution of Glycinergic Neurons in the Brain of Glycine Transporter-2 Transgenic Tg(Glyt2:Gfp) Adult Zebrafish: Relationship to Brain-Spinal Descending Systems. J. Comp. Neurol. 2013, 521, 389–425. [Google Scholar] [CrossRef] [PubMed]
- Mueller, T.; Vernier, P.; Wullimann, M.F. The Adult Central Nervous Cholinergic System of a Neurogenetic Model Animal, the Zebrafish Danio Rerio. Brain Res. 2004, 1011, 156–169. [Google Scholar] [CrossRef]
- Filippi, A.; Mueller, T.; Driever, W. Vglut2 and Gad Expression Reveal Distinct Patterns of Dual GABAergic versus Glutamatergic Cotransmitter Phenotypes of Dopaminergic and Noradrenergic Neurons in the Zebrafish Brain. J. Comp. Neurol. 2014, 522, 2019–2037. [Google Scholar] [CrossRef]
- Higashijima, S.I.; Mandel, G.; Fetcho, J.R. Distribution of Prospective Glutamatergic, Glycinergic, and GABAergic Neurons in Embryonic and Larval Zebrafish. J. Comp. Neurol. 2004, 480, 1–18. [Google Scholar] [CrossRef]
- Schweitzer, J.; Löhr, H.; Filippi, A.; Driever, W. Dopaminergic and Noradrenergic Circuit Development in Zebrafish. Dev. Neurobiol. 2012, 72, 256–268. [Google Scholar] [CrossRef]
- Kaslin, J.; Panula, P. Comparative Anatomy of the Histaminergic and Other Aminergic Systems in Zebrafish (Danio rerio). J. Comp. Neurol. 2001, 440, 342–377. [Google Scholar] [CrossRef] [PubMed]
- Tay, T.L.; Ronneberger, O.; Ryu, S.; Nitschke, R.; Driever, W. Comprehensive Catecholaminergic Projectome Analysis Reveals Single-Neuron Integration of Zebrafish Ascending and Descending Dopaminergic Systems. Nat. Commun. 2011, 2, 171. [Google Scholar] [CrossRef] [PubMed]
- McLean, D.L.; Fetcho, J.R. Ontogeny and Innervation Patterns of Dopaminergic, Noradrenergic, and Serotonergic Neurons in Larval Zebrafish. J. Comp. Neurol. 2004, 480, 38–56. [Google Scholar] [CrossRef] [PubMed]
- Lillesaar, C.; Stigloher, C.; Tannhäuser, B.; Wullimann, M.F.; Bally-Cuif, L. Axonal Projections Originating from Raphe Serotonergic Neurons in the Developing and Adult Zebrafish, Danio Rerio, Using Transgenics to Visualize Raphe-Specific Pet1 Expression. J. Comp. Neurol. 2009, 512, 158–182. [Google Scholar] [CrossRef] [PubMed]
- Biran, J.; Blechman, J.; Wircer, E.; Levkowitz, G. Development and Function of the Zebrafish Neuroendocrine System. In Model Animals in Neuroendocrinology: From Worm to Mouse to Man; John Wiley & Sons Ltd.: Hoboken, NJ, USA, 2018; pp. 101–131. [Google Scholar] [CrossRef]
- Ziv, L.; Muto, A.; Schoonheim, P.J.; Meijsing, S.H.; Strasser, D.; Ingraham, H.A.; Schaaf, M.J.M.; Yamamoto, K.R.; Baier, H. An Affective Disorder in Zebrafish with Mutation of the Glucocorticoid Receptor. Mol. Psychiatry 2012, 18, 681–691. [Google Scholar] [CrossRef]
- Piato, A.L.; Capiotti, K.M.; Tamborski, A.R.; Oses, J.P.; Barcellos, L.J.G.; Bogo, M.R.; Lara, D.R.; Vianna, M.R.; Bonan, C.D. Unpredictable Chronic Stress Model in Zebrafish (Danio Rerio): Behavioral and Physiological Responses. Prog. Neuropsychopharmacol. Biol. Psychiatry 2011, 35, 561–567. [Google Scholar] [CrossRef] [PubMed]
- Griffiths, B.; Schoonheim, P.J.; Ziv, L.; Voelker, L.; Baier, H.; Gahtan, E. A Zebrafish Model of Glucocorticoid Resistance Shows Serotonergic Modulation of the Stress Response. Front. Behav. Neurosci. 2012, 6, 68. [Google Scholar] [CrossRef] [PubMed]
- Haynes, E.M.; Ulland, T.K.; Eliceiri, K.W. A Model of Discovery: The Role of Imaging Established and Emerging Non-Mammalian Models in Neuroscience. Front. Mol. Neurosci. 2022, 15, 153. [Google Scholar] [CrossRef]
- Shin, J.; Padmanabhan, A.; De Groh, E.D.; Lee, J.S.; Haidar, S.; Dahlberg, S.; Guo, F.; He, S.; Wolman, M.A.; Granato, M.; et al. Zebrafish Neurofibromatosis Type 1 Genes Have Redundant Functions in Tumorigenesis and Embryonic Development. DMM Dis. Models Mech. 2012, 5, 881–894. [Google Scholar] [CrossRef]
- Graeber, M.B.; Scheithauer, B.W.; Kreutzberg, G.W. Microglia in Brain Tumors. Glia 2002, 40, 252–259. [Google Scholar] [CrossRef]
- Chia, K.; Mazzolini, J.; Mione, M.; Sieger, D. Tumor Initiating Cells Induce Cxcr4-Mediated Infiltration of pro-Tumoral Macrophages into the Brain. eLife 2018, 7, e31918. [Google Scholar] [CrossRef] [PubMed]
- Gualano, F.M.; Hassoun, P.; Carter, C.L.; Hanson, D. Embryonal Tumor with Multilayered Rosettes: Post-treatment Maturation and Implications for Future Therapy. Cancer Rep. 2023, 6, e1812. [Google Scholar] [CrossRef] [PubMed]
- Malchenko, S.; Sredni, S.T.; Boyineni, J.; Bi, Y.; Margaryan, N.V.; Guda, M.R.; Kostenko, Y.; Tomita, T.; Davuluri, R.V.; Velpula, K.; et al. Characterization of Brain Tumor Initiating Cells Isolated from an Animal Model of CNS Primitive Neuroectodermal Tumors. Oncotarget 2018, 9, 13733. [Google Scholar] [CrossRef]
- Louis, D.N.; Perry, A.; Reifenberger, G.; von Deimling, A.; Figarella-Branger, D.; Cavenee, W.K.; Ohgaki, H.; Wiestler, O.D.; Kleihues, P.; Ellison, D.W. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A Summary. Acta Neuropathol. 2016, 131, 803–820. [Google Scholar] [CrossRef]
- Louis, D.N.; Ohgaki, H.; Wiestler, O.D.; Cavenee, W.K.; Burger, P.C.; Jouvet, A.; Scheithauer, B.W.; Kleihues, P. The 2007 WHO Classification of Tumours of the Central Nervous System. Acta Neuropathol. 2007, 114, 97–109. [Google Scholar] [CrossRef]
- Schultz, L.E.; Haltom, J.A.; Almeida, M.P.; Wierson, W.A.; Solin, S.L.; Weiss, T.J.; Helmer, J.A.; Sandquist, E.J.; Shive, H.R.; McGrail, M. Epigenetic Regulators Rbbp4 and Hdac1 Are Overexpressed in a Zebrafish Model of RB1 Embryonal Brain Tumor, and Are Required for Neural Progenitor Survival and Proliferation. DMM Dis. Models Mech. 2018, 11, dmm034124. [Google Scholar] [CrossRef] [PubMed]
- Modzelewska, K.; Boer, E.F.; Mosbruger, T.L.; Picard, D.; Anderson, D.; Miles, R.R.; Kroll, M.; Oslund, W.; Pysher, T.J.; Schiffman, J.D.; et al. MEK Inhibitors Reverse Growth of Embryonal Brain Tumors Derived from Oligoneural Precursor Cells. Cell Rep. 2016, 17, 1255–1264. [Google Scholar] [CrossRef]
- Lam, S.H.; Chua, H.L.; Gong, Z.; Lam, T.J.; Sin, Y.M. Development and Maturation of the Immune System in Zebrafish, Danio Rerio: A Gene Expression Profiling, in Situ Hybridization and Immunological Study. Dev. Comp. Immunol. 2004, 28, 9–28. [Google Scholar] [CrossRef]
- Almstedt, E.; Rosen, E.; Gloger, M.; Stockgard, R.; Hekmati, N.; Koltowska, K.; Krona, C.; Nelander, S. Real-Time Evaluation of Glioblastoma Growth in Patient-Specific Zebrafish Xenografts. Neuro Oncol. 2022, 24, 726. [Google Scholar] [CrossRef]
- Hamilton, L.; Astell, K.R.; Velikova, G.; Sieger, D. A Zebrafish Live Imaging Model Reveals Differential Responses of Microglia Toward Glioblastoma Cells In Vivo. Zebrafish 2016, 13, 523. [Google Scholar] [CrossRef]
- Pudelko, L.; Rouhi, P.; Sanjiv, K.; Gad, H.; Kalderén, C.; Höglund, A.; Squatrito, M.; Schuhmacher, A.J.; Edwards, S.; Hägerstrand, D.; et al. Glioblastoma and Glioblastoma Stem Cells Are Dependent on Functional MTH1. Oncotarget 2017, 8, 84671. [Google Scholar] [CrossRef] [PubMed]
- Yan, C.; Yang, Q.; Do, D.; Brunson, D.C.; Langenau, D.M. Adult Immune Compromised Zebrafish for Xenograft Cell Transplantation Studies. eBioMedicine 2019, 47, 24. [Google Scholar] [CrossRef] [PubMed]
- Yan, C.; Brunson, D.C.; Tang, Q.; Do, D.; Iftimia, N.A.; Moore, J.C.; Hayes, M.N.; Welker, A.M.; Garcia, E.G.; Dubash, T.D.; et al. Visualizing Engrafted Human Cancer and Therapy Responses in Immunodeficient Zebrafish. Cell 2019, 177, 1903–1914.e14. [Google Scholar] [CrossRef] [PubMed]
- Yan, C.; Do, D.; Yang, Q.; Brunson, D.C.; Rawls, J.F.; Langenau, D.M. Single-Cell Imaging of Human Cancer Xenografts Using Adult Immunodeficient Zebrafish. Nat. Protoc. 2020, 15, 3105–3128. [Google Scholar] [CrossRef] [PubMed]
- Idilli, A.I.; Cusanelli, E.; Pagani, F.; Berardinelli, F.; Bernabé, M.; Cayuela, M.L.; Poliani, P.L.; Mione, M.C. Expression of Tert Prevents ALT in Zebrafish Brain Tumors. Front. Cell Dev. Biol. 2020, 8, 514067. [Google Scholar] [CrossRef] [PubMed]
- Gronseth, E.; Gupta, A.; Koceja, C.; Kumar, S.; Kutty, R.G.; Rarick, K.; Wang, L.; Ramchandran, R. Astrocytes Influence Medulloblastoma Phenotypes and CD133 Surface Expression. PLoS ONE 2020, 15, e0235852. [Google Scholar] [CrossRef] [PubMed]
- Yuan, M.; White, D.; Resar, L.; Bar, E.; Groves, M.; Cohen, A.; Jackson, E.; Bynum, J.; Rubens, J.; Mumm, J.; et al. Conditional Reprogramming Culture Conditions Facilitate Growth of Lower-Grade Glioma Models. Neuro Oncol. 2021, 23, 770–782. [Google Scholar] [CrossRef] [PubMed]
- Gatzweiler, C.; Ridinger, J.; Herter, S.; Gerloff, X.F.; Elharouni, D.; Berker, Y.; Imle, R.; Schmitt, L.; Kreth, S.; Stainczyk, S.; et al. Functional Therapeutic Target Validation Using Pediatric Zebrafish Xenograft Models. Cancers 2022, 14, 849. [Google Scholar] [CrossRef]
- Zhu, F.; Nair, R.R.; Fisher, E.M.C.; Cunningham, T.J. Humanising the Mouse Genome Piece by Piece. Nat. Commun. 2019, 10, 1845. [Google Scholar] [CrossRef]
- Rajan, V.; Melong, N.; Wong, W.H.; King, B.; Spencer Tong, R.; Mahajan, N.; Gaston, D.; Lund, T.; Rittenberg, D.; Dellaire, G.; et al. Humanized Zebrafish Enhance Human Hematopoietic Stem Cell Survival and Promote Acute Myeloid Leukemia Clonal Diversity. Haematologica 2020, 105, 2391–2399. [Google Scholar] [CrossRef]
- Häberlein, F.; Mingardo, E.; Merten, N.; Schulze Köhling, N.K.; Reinoß, P.; Simon, K.; Japp, A.; Nagarajan, B.; Schrage, R.; Pegurier, C.; et al. Humanized Zebrafish as a Tractable Tool for in Vivo Evaluation of Pro-Myelinating Drugs. Cell Chem. Biol. 2022, 29, 1541–1555. [Google Scholar] [CrossRef] [PubMed]
- Al-Hamaly, M.A.; Turner, L.T.; Rivera-Martinez, A.; Rodriguez, A.; Blackburn, J.S. Zebrafish Cancer Avatars: A Translational Platform for Analyzing Tumor Heterogeneity and Predicting Patient Outcomes. Int. J. Mol. Sci. 2023, 24, 2288. [Google Scholar] [CrossRef] [PubMed]
- Hoffman, L.M.; Van Zanten, S.E.M.V.; Colditz, N.; Baugh, J.; Chaney, B.; Hoffmann, M.; Lane, A.; Fuller, C.; Miles, L.; Hawkins, C.; et al. Clinical, Radiologic, Pathologic, and Molecular Characteristics of Long-Term Survivors of Diffuse Intrinsic Pontine Glioma (DIPG): A Collaborative Report from the International and European Society for Pediatric Oncology DIPG Registries. J. Clin. Oncol. 2018, 36, 1963. [Google Scholar] [CrossRef] [PubMed]
- Packer, R.J. Brain Tumors in Children. Arch. Neurol. 1999, 56, 421–425. [Google Scholar] [CrossRef] [PubMed]
- ’t Hart, E.; Bianco, J.; Besse, H.C.; Chin Joe Kie, L.A.; Cornet, L.; Eikelenboom, K.L.; van den Broek, T.J.M.; Derieppe, M.; Su, Y.; Hoving, E.W.; et al. Towards Standardisation of a Diffuse Midline Glioma Patient-Derived Xenograft Mouse Model Based on Suspension Matrices for Preclinical Research. Biomedicines 2023, 11, 527. [Google Scholar] [CrossRef]
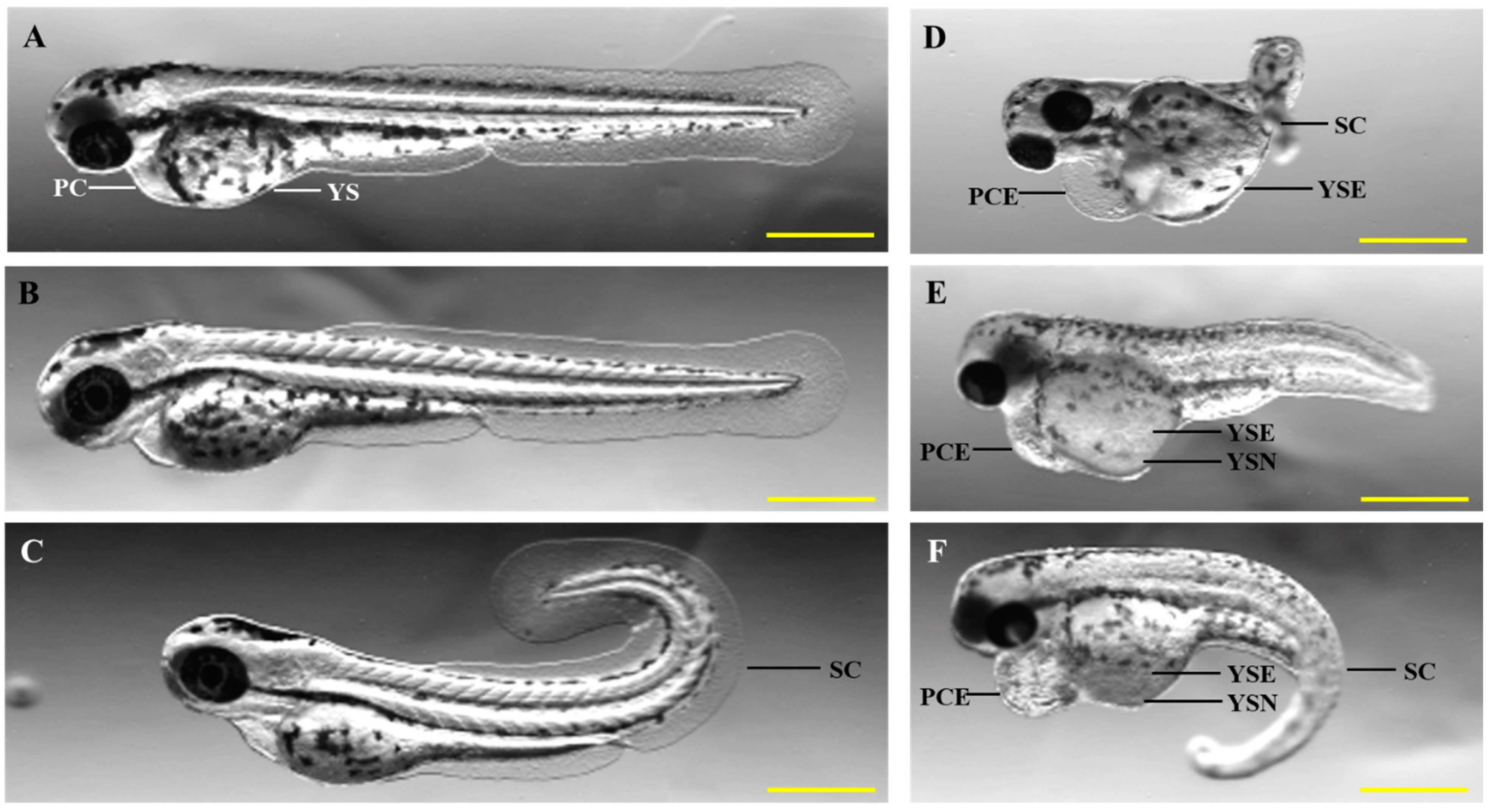
- Pelka, K.E.; Henn, K.; Keck, A.; Sapel, B.; Braunbeck, T. Size Does Matter–Determination of the Critical Molecular Size for the Uptake of Chemicals across the Chorion of Zebrafish (Danio rerio) Embryos. Aquat. Toxicol. 2017, 185, 1–10. [Google Scholar] [CrossRef]
| Model | Cost | Advantages | Disadvantages | Drug Screening Throughput |
|---|---|---|---|---|
| Cell culture | Low | Immortality, rapid growth, robustness, ease of genetic modifications, maintenance, and storage | Long-term culture can result in the development of cells that are genetically distinct from primary tumor cells. There are no tumor microenvironments. | Very High |
| Organoids | High | Similar tumor heterogeneity, characteristics, and microenvironments to human systems. | Technically difficult to develop, costly, and variable in growth. | High |
| Rodents | Very High | Replicate tumor microenvironment, genetic alterations, and pharmacodynamics as closely as possible to humans. | Time-consuming; lack immune interactions. PDX models primarily depend on tissue integrity; highly variable in nature. | Low |
| Zebrafish | Intermediate | External fertilization, large number of offspring, ease of transplantation, high efficiency in genetic manipulation, rapid tumor engraftment, and the development of tumors with histopathology similar to that of humans. Cell tracking in embryos and larvae is very easy due to their transparent bodies. | Difficulties in accurately measuring drug concentration in larval plasma, hindering drug absorption, distribution, metabolism, and excretion (ADME) studies, along with a lack of adaptive immune system in larvae, restrict the direct prediction of clinical dosage from zebrafish to humans. | High |
| Drosophila | Low | Short lifespan, large number of progenies, ease of genetic manipulation, drug screening, dissection of complex tissues. | Tumors can grow for a limited time and thus are not suitable for long-term studies. Angiogenesis and adaptive immunity cannot be studied. A few organ-specific tumors, such as pancreas, liver, and lung tumors, cannot be studied due to a lack of homologous organs. | High |
| C. elegans | Low | Conserve signaling pathways, genetic similarities, ease of genetic manipulation, transparent body. | They do not develop cancers in a similar way to humans. They lack adaptive immunity, angiogenesis, and organ systems comparable to those of humans. | High |
| Cancer | Studied Gene/Cell Line/Drug | Method | Zebrafish Line Used/Developed | Reference |
|---|---|---|---|---|
| Acute lymphoid leukemia | BAG1 | RS4;11 cell xenograft | Tg(fli1:GFP) | [94] |
| Ext1 and Ext 2 | Morpholino-based knockdown | Tg(Tp1bglob:eGFP) | [102] | |
| MYC | Transgenic | Tg(rag2:hMYC), Tg(lck:eGFP), Tg(hMYC;GFP) | [103] | |
| Adenoid Cystic Carcinoma | MYB | Transgenic | Zebrafish blastomeres, Tg(c-myb-GFP), Tg(c-myb:GFP; lyz:dsRed), Tg(c-myb:GFP; mpeg1:mCherry) | [104] |
| Chronic Myeloid Leukemia | BCR/ABL1 | Transgenic | WT (AB), Tg(lyz:DsRed), Tg(hsp70:p210BCR/ABL1) | [105] |
| ABL inhibitor imatinib, MEK inhibitor U0126, cytarabine, azacitidine, and arsenic trioxide | K562, CD34+ HPSC, MV4-11 and MOLM-13 cell xenograft | prkdc−/− in casper background (SCID zebrafish) | [106] | |
| Intestine | YES1, YAP1 | Morpholino-based knockdown, dasatinib treatment | WT, Tg(fabp2:RFP)as200, axin1tm213 | [107] |
| CATSPERE | Transgenic | WT (AB), Tg(ifabp:DsRed-P2A-CATSPERE; CATSPERE), Tg(ifabp:EGFP;WT), p53−/− | [108] | |
| HCT116, anandamide | Xenograft | WT (Tübingen), Tg(fli1:EGFP), Tg(mpeg1:EGFP) | [109] | |
| Renal cell carcinoma | VHL | Transgenic | Tg(ATPase1.a1A4:GFP), Tg(vhlhu2117+/−), Tg(ATPase1.a1A4:GFP:vhl−/−) | [110] |
| Liver | Tulp3 | CRISPR knockout | WT (AB, TL), Tg(wt1b:EGFP), homozygous mutant-tulpm/m | [74] |
| Pten and Tp53 | CRISPR knockout | WT (AB), Tg(fabp10:Cas9-mCherry);ptena−/−, Tg(fabp10:Cas9-mCherry); ptenb−/−, Tg(fabp10:Cas9-mCherry); tp53−/−, Tg(fabp10:Cas9-mCherry) | [111] | |
| Melanoma | BRAF | Transgenic | WT (AB), Tg(crestin:CreERt2;crystallin:YFP), Tg(−3.5ubi:loxP-GFP-loxP-mCherry), Tg(p53/BRAF/Na/MiniCoopR/crestin:EGFP) | [112] |
| 92.1 and Mel270 cell line | Xenograft | Tg(fli1:eGFP) | [113] | |
| MITF, BRAF | Transgenic | mitfavc7, tp53M214K, Tg(mitfa-BRAFV600E), Tg(mitfa:BRAFV600E); mitfavc7; p53M214K) | [114] | |
| kita promoter, HRAS | Transgenic | WT (AB), tg(UAS:GFP), tg(5XUAS:eGFP-HRASV12)io6, mitfaw2/w2, p53zdf1/zdf1, tg(mitfa:Gal4VP16;UAS:mCherry); Et(kita:GalTA4,UAS:mCherry)hzm1 | [115] | |
| Myelodysplastic syndrome | TET2 | Transgenic | WT (AB), tet2m/m, Tg(c-myb-GFP), Tg(cd41-GFP) | [116] |
| c-myb | Transgenic | WT (AB), Tg(c-myb:gfp), Tg(c-mybhkz3), Tg(rag2:dsRed), Tg(lyz:dsRed) | [117] | |
| Pancreatic cancer | Rabl3, KRAS | CRISPR transgenic | WT, tp53−/−, Tg(tp53−/−;rabl3-TR52), Tg(rabl3-TR41/+), Tg(rabl3-TR41/41) | [118] |
| KRAS | Transgenic | Tg(ubb:Lox-NucmCherry-stop-Lox-GFP::KRASG12D), Tg(elastase3I:CRE;cryaa:Venus), Tg(ela3I-CRE; LSL-KRASG12D) | [119] | |
| Panc-1 cells | Xenograft | Tg(fli1:eGFP), Nacre (mitfa−/−) | [120] | |
| Peripheral nerve sheath tumor (PNST) | lats1 and lats2 | CRISPR knockout | WT (AB) lats2mw87/mw87 | [121] |
| Suz12 | Morpholino-based knockdown | nf1a+/−:nf1b−/−:p53e7/e7 | [122] | |
| Retinoblastoma | ACVR1C, SMAD | Y79-GFP cell xenograft | WT (AB) | [123] |
| Rhabdomyosarcoma | Pax3 | Transgenic | Tg(pax3a:EGFP), pax3a−/− and pax3b−/− | [100] |
| HES3, her3 | CRISPR knockout | her3 null mutants (her3nch1, her3nc2, her3nch3) | [101] | |
| kRAS | Transgenic | WT (AB), Tg(myf5:GFP;mylz2:mCherry), Tg(cdh15:GFP), Tg(mylz2:mCherry), Tg(cdh15:KRASG12D), Tg(mylz2:KRASG12D) | [124] | |
| PAX3-FOXO1, HES3 | Transgenic | WT (AB), WIK, TL, AB/TL, tp53M214K, Tg(BetaActin:GFP2A:PAX3FOXO1), Tg(CMV:GFP2A:PAX3FOXO1), Tg(ubi:GFP2A:PAX3FOXO1) | [125] | |
| Systemic mastocytosis | KIT | Transgenic | Tg(actb2:KITD816V:2AeGFP) | [126] |
| T-cell acute lymphoid leukemia | ptch1 | CRISPR knockout | Tg(ptch1mutant) Tg (rag2-notch1aICD) | [96] |
| IL7R | Transgenic | CG1, Tg(rag2:RFP), Tg(rag2:IL7Rmut2), Tg(rag2:IL7Rmut2-tdTomato) | [127] | |
| AURKB and Myc | Transgenic | WT, Tg(rag2:AURKB;rag2:mCherry), Tg(rag2:Loxp-dsRED2-Loxp-EGFP-Myc;hsp70:Cre), Tg(rag2:EGFP;rag2:Myc), Tg(rag2:EGFP;rag2:MycS67A), Tg(rag2:EGFP;rag2:Myc;rag2:AURKB) | [128] | |
| prl3 and Myc | Transgenic | CG1, Tg(rag2:GFP;rag2:Myc), Tg(rag2:prl3;rag2:mCherry) | [129] | |
| NUP88/Nup214 | Morpholino-based knockdown | WT (AB, Tübingen) | [130] | |
| Lrrc50 | ENU mutagenesis, Morpholino-based knockdown | WT, lrrc50hu255h (+/−) | [131] | |
| Thyroid cancer | BRAF | Transgenic | Tg(TdTomato-pA),Tg(BRAFV600E-pA:TdTomato-pA) | [132] |
| CREB3L1 | 8505C cell xenograft | WT | [133] |
| Cancer | Studied Gene/Cell Line/Drug | Method | Zebrafish Line Used/Developed | Reference |
|---|---|---|---|---|
| Glioblastoma | nf1 | Transgenic | Tg(nf1a+/–; nf1b–/–; p53e7/e7) | [165] |
| MTH1, TH588 and TH1579 (MTH1 inhibitors) | GBM #18-CMV-LUC cell xenograft | Wild-type (TL) | [177] | |
| Microglial response toward GBM cells | U87 and U251 cell xenograft | Tg(mpeg1:EGFP), irf8−/− | [176] | |
| Role of tert in telomere stability | Transgenic | Tg(10xUAS:tert), Tg(10xUAS:terc), | [181] | |
| Cxcr4-mediated infiltration of pro-tumoral macrophages | Transgenic | Tg(NBT:∆LexPR-lexOP-pA; mpeg1:EGFP), Tg(mpeg1:mCherry; p2ry12:p2ry12-GFP), cxcr4b−/− | [167] | |
| Embryonal tumors (previously classified as CNS-PNETs) | rb1, rbbp4, and hdac1 | Transgenic | Tg(H2A.F/Z-GFP), rb1Δ7/Δ7 | [172] |
| SOX10 and OLIG2 | Transgenic, Tumor allograft from Tg(mitfaw2; p53M214K; Tg(sox10:mCherry-NRASWT) fish into mitfw2 fish | Tg(mitfaw2; p53M214K; Tg(sox10:mCherry-NRASWT), mitfw2 | [173] | |
| Medulloblastoma | CD133 | Daoy cell xenograft | Tg(flk:mCherry); Absolut+/+ (ednrbl−/− mitfa−/−) | [182] |
| Pilocytic astrocytoma | NF1 | JHH-NF1-PA1 cell xenograft | Wild-type (AB) | [183] |
| Rhabdoid tumor | SMARCB, PRKCD, DDR2 | INF_R_1288_r1, INF_R_1467_r1, and INF_R_359_r3 cell xenograft | Wild-type (AB) | [184] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Roy, D.; Subramaniam, B.; Chong, W.C.; Bornhorst, M.; Packer, R.J.; Nazarian, J. Zebrafish—A Suitable Model for Rapid Translation of Effective Therapies for Pediatric Cancers. Cancers 2024, 16, 1361. https://doi.org/10.3390/cancers16071361
Roy D, Subramaniam B, Chong WC, Bornhorst M, Packer RJ, Nazarian J. Zebrafish—A Suitable Model for Rapid Translation of Effective Therapies for Pediatric Cancers. Cancers. 2024; 16(7):1361. https://doi.org/10.3390/cancers16071361
Chicago/Turabian StyleRoy, Debasish, Bavani Subramaniam, Wai Chin Chong, Miriam Bornhorst, Roger J. Packer, and Javad Nazarian. 2024. "Zebrafish—A Suitable Model for Rapid Translation of Effective Therapies for Pediatric Cancers" Cancers 16, no. 7: 1361. https://doi.org/10.3390/cancers16071361
APA StyleRoy, D., Subramaniam, B., Chong, W. C., Bornhorst, M., Packer, R. J., & Nazarian, J. (2024). Zebrafish—A Suitable Model for Rapid Translation of Effective Therapies for Pediatric Cancers. Cancers, 16(7), 1361. https://doi.org/10.3390/cancers16071361





