A Novel Urinary miRNA Biomarker for Early Detection of Colorectal Cancer
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
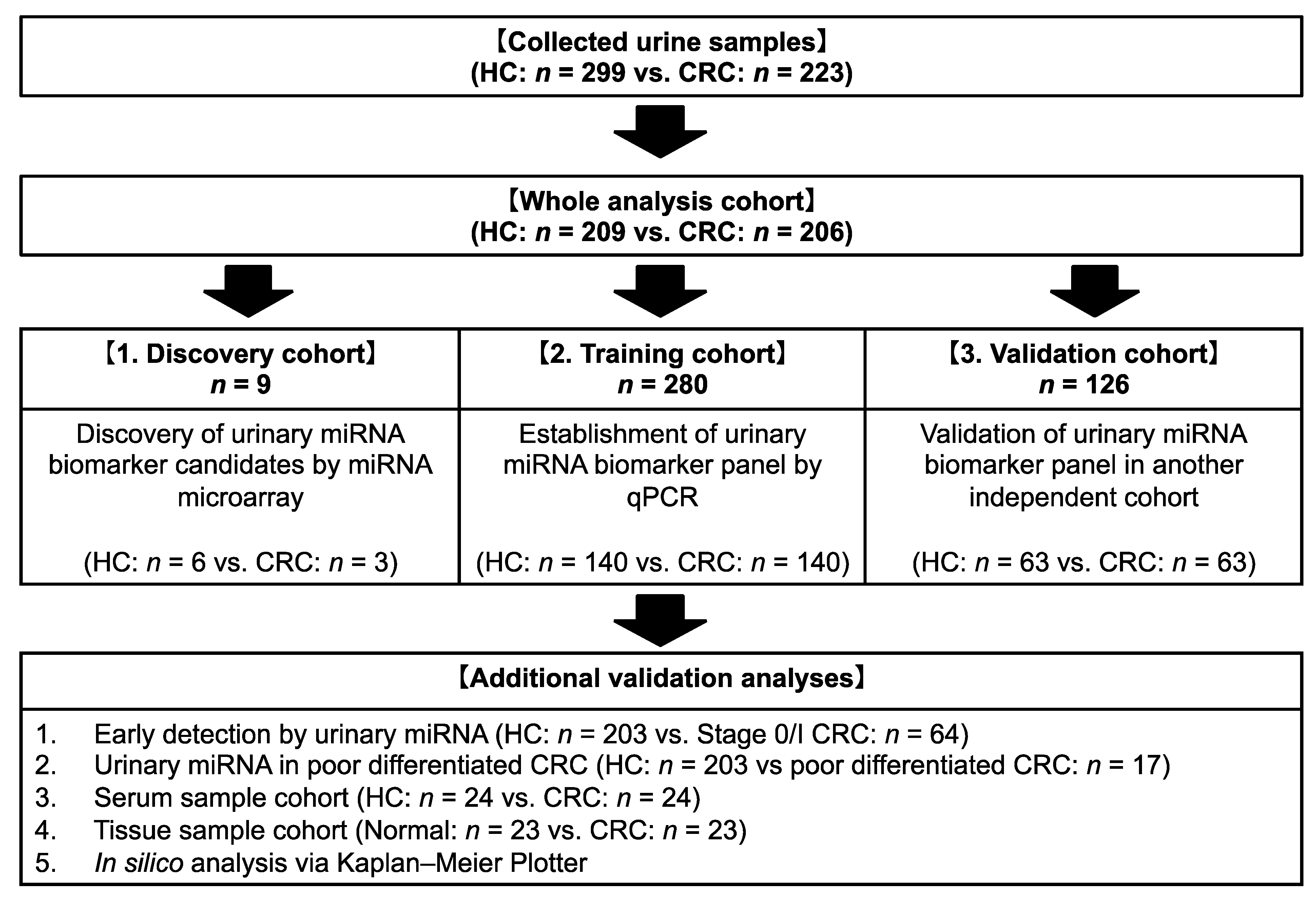
2.1. Patients and Study Design
2.2. Samples and Definition
2.3. miRNA Extraction
2.4. miRNA Microarray Assay
2.5. Quantitative Reverse Transcription-Polymerase Chain Reaction (qRT-PCR)
2.6. In Silico Analyses
2.7. Statistical Analyses
3. Results
3.1. Participants
3.2. Urinary miRNA Difference between HC and CRC Groups
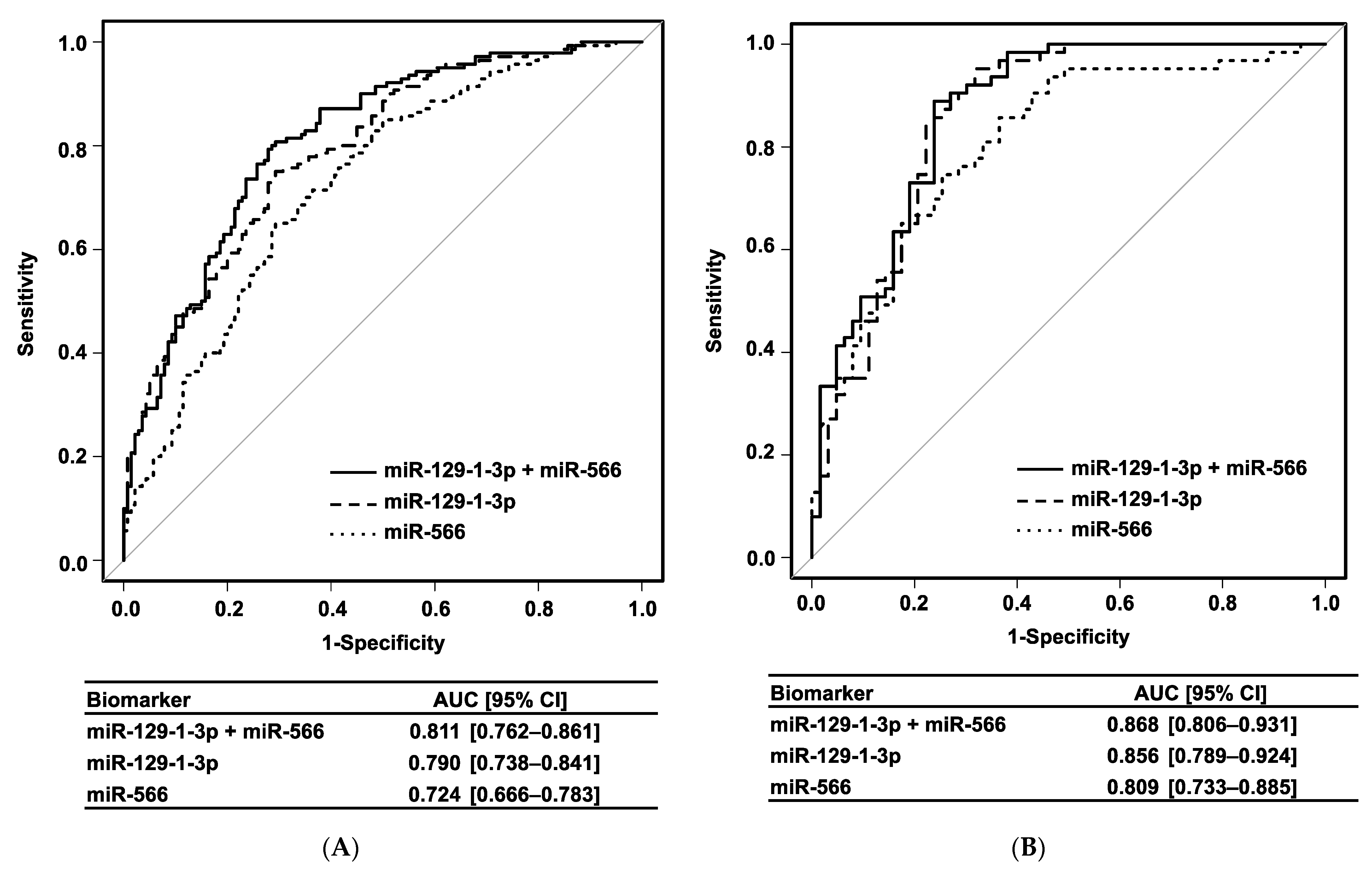
3.3. Development of Urinary miRNA Biomarker
3.4. Validation of Urinary miRNA Biomarker
3.5. Analysis Using Serum and Tissue Samples
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Fitzmaurice, C.; Akinyemiju, T.F.; al Lami, F.H.; Alam, T.; Alizadeh-Navaei, R.; Allen, C.; Alsharif, U.; Alvis-Guzman, N.; Amini, E.; Anderson, B.O.; et al. Global Burden of Disease Cancer Collaboration. Global, Regional, and National Cancer Incidence, Mortality, Years of Life Lost, Years Lived with Disability, and Disability-Adjusted Life-Years for 29 Cancer Groups, 1990 to 2016: A Systematic Analysis for the Global Burden of Disease Study. JAMA Oncol. 2018, 4, 1553–1568. [Google Scholar] [CrossRef] [PubMed]
- Bretthauer, M.; Kaminski, M.; Løberg, M.; Zauber, A.G.; Regula, J.; Kuipers, E.J.; Hernán, M.; McFadden, E.; Sunde, A.; Kalager, M.; et al. Population-Based Colonoscopy Screening for Colorectal Cancer: A Randomized Clinical Trial. JAMA Intern. Med. 2016, 176, 894–902. [Google Scholar] [CrossRef]
- Halloran, S.; Launoy, G.; Zappa, M. European guidelines for quality assurance in colorectal cancer screening and diagnosis.—First Edition Faecal occult blood testing. Endoscopy 2012, 44 (Suppl. S3), SE65–SE87. [Google Scholar] [CrossRef] [Green Version]
- US Preventive Services Task Force; Bibbins-Domingo, K.; Grossman, D.C.; Curry, S.J.; Davidson, K.W.; Epling, J.W., Jr.; García, F.A.R.; Gillman, M.W.; Harper, D.M.; Kemper, A.R.; et al. Screening for Colorectal Cancer: US Preventive Services Task Force Recommendation Statement. JAMA 2016, 315, 2564–2575. [Google Scholar] [CrossRef]
- van Dam, L.; Kuipers, E.J.; van Leerdam, M.E. Performance improvements of stool-based screening tests. Best Pr. Res. Clin. Gastroenterol. 2010, 24, 479–492. [Google Scholar] [CrossRef] [PubMed]
- Gies, A.; Cuk, K.; Schrotz-King, P.; Brenner, H. Direct Comparison of Diagnostic Performance of 9 Quantitative Fecal Immunochemical Tests for Colorectal Cancer Screening. Gastroenterology 2018, 154, 93–104. [Google Scholar] [CrossRef] [Green Version]
- de Klaver, W.; Wisse, P.H.A.; van Wifferen, F.; Bosch, L.J.W.; Jimenez, C.R.; van der Hulst, R.W.M.; Fijneman, R.J.A.; Kuipers, E.J.; Greuter, M.J.E.; Carvalho, B.; et al. Clinical Validation of a Multitarget Fecal Immunochemical Test for Colorectal Cancer Screening: A Diagnostic Test Accuracy Study. Ann. Intern. Med. 2021, 174, 1224–1231. [Google Scholar] [CrossRef]
- Imperiale, T.; Ransohoff, D.F.; Itzkowitz, S.H.; Levin, T.R.; Lavin, P.; Lidgard, G.P.; Ahlquist, D.A.; Berger, B.M. Multitarget Stool DNA Testing for Colorectal-Cancer Screening. N. Engl. J. Med. 2014, 370, 1287–1297. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhai, R.-L.; Xu, F.; Zhang, P.; Zhang, W.-L.; Wang, H.; Wang, J.-L.; Cai, K.-L.; Long, Y.-P.; Lu, X.-M.; Tao, K.-X.; et al. The Diagnostic Performance of Stool DNA Testing for Colorectal Cancer: A Systematic Review and Meta-Analysis. Medicine 2016, 95, e2129. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, R.H. Carcinoembryonic Antigen. Ann. Intern. Med. 1986, 104, 66–73. [Google Scholar] [CrossRef] [Green Version]
- Nicolini, A.; Ferrari, P.; Duffy, M.J.; Antonelli, A.; Rossi, G.; Metelli, M.R.; Fulceri, F.; Anselmi, L.; Conte, M.; Berti, P.; et al. Intensive Risk-Adjusted Follow-up With the CEA, TPA, CA19.9, and CA72.4 Tumor Marker Panel and Abdominal Ultrasonography to Diagnose Operable Colorectal Cancer Recurrences: Effect on survival. Arch. Surg. 2010, 145, 1177–1183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thomas, D.; Fourkala, E.-O.; Apostolidou, S.; Gunu, R.; Ryan, A.; Jacobs, I.; Menon, U.; Alderton, W.; Gentry-Maharaj, A.; Timms, J.F. Evaluation of serum CEA, CYFRA21-1 and CA125 for the early detection of colorectal cancer using longitudinal preclinical samples. Br. J. Cancer 2015, 113, 268–274. [Google Scholar] [CrossRef] [Green Version]
- Calin, G.A.; Croce, C.M. MicroRNA Signatures in Human Cancers. Nat. Rev. Cancer 2006, 6, 857–866. [Google Scholar] [CrossRef]
- Calin, G.; Croce, C.M. Chromosomal Rearrangements and MicroRNAs: A New Cancer Link with Clinical Implications. J. Clin. Investig. 2007, 117, 2059–2066. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arroyo, J.D.; Chevillet, J.R.; Kroh, E.M.; Ruf, I.K.; Pritchard, C.C.; Gibson, D.F.; Mitchell, P.S.; Bennett, C.F.; Pogosova-Agadjanyan, E.L.; Stirewalt, D.L.; et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl. Acad. Sci. USA 2011, 108, 5003–5008. [Google Scholar] [CrossRef] [Green Version]
- Egidi, M.G.; Cochetti, G.; Guelfi, G.; Zampini, D.; Diverio, S.; Poli, G.; Mearini, E. Stability Assessment of Candidate Reference Genes in Urine Sediment of Prostate Cancer Patients for miRNA Applications. Dis. Markers 2015, 2015, 973597. [Google Scholar] [CrossRef] [PubMed]
- Mall, C.; Rocke, D.M.; Durbin-Johnson, B.; Weiss, R.H. Stability of miRNA in human urine supports its biomarker potential. Biomark. Med. 2013, 7, 623–631. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andreasen, D.; Fog, J.U.; Biggs, W.; Salomon, J.; Dahslveen, I.K.; Baker, A.; Mouritzen, P. Improved microRNA quantification in total RNA from clinical samples. Methods 2010, 50, S6–S9. [Google Scholar] [CrossRef]
- Mraz, M.; Malinova, K.; Mayer, J.; Pospisilova, S. MicroRNA isolation and stability in stored RNA samples. Biochem. Biophys. Res. Commun. 2009, 390, 1–4. [Google Scholar] [CrossRef]
- Ng, E.K.-O.; Chong, W.W.S.; Jin, H.; Lam, E.K.Y.; Shin, V.Y.; Yu, J.; Poon, T.C.W.; Ng, S.S.M.; Sung, J.J.Y. Differential expression of microRNAs in plasma of patients with colorectal cancer: A potential marker for colorectal cancer screening. Gut 2009, 58, 1375–1381. [Google Scholar] [CrossRef] [Green Version]
- Imaoka, H.; Toiyama, Y.; Fujikawa, H.; Hiro, J.; Saigusa, S.; Tanaka, K.; Inoue, Y.; Mohri, Y.; Mori, T.; Kato, T.; et al. Circulating microRNA-1290 as a novel diagnostic and prognostic biomarker in human colorectal cancer. Ann. Oncol. 2016, 27, 1879–1886. [Google Scholar] [CrossRef]
- Iwasaki, H.; Shimura, T.; Kataoka, H. Current status of urinary diagnostic biomarkers for colorectal cancer. Clin. Chim. Acta 2019, 498, 76–83. [Google Scholar] [CrossRef]
- Shimura, T.; Dagher, A.; Sachdev, M.; Ebi, M.; Yamada, T.; Yamada, T.; Joh, T.; Moses, M.A. Urinary ADAM12 and MMP-9/NGAL Complex Detect the Presence of Gastric Cancer. Cancer Prev. Res. 2015, 8, 240–248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shimura, T.; Ebi, M.; Yamada, T.; Yamada, T.; Katano, T.; Nojiri, Y.; Iwasaki, H.; Nomura, S.; Hayashi, N.; Mori, Y.; et al. Urinary kallikrein 10 predicts the incurability of gastric cancer. Oncotarget 2017, 8, 29247–29257. [Google Scholar] [CrossRef] [Green Version]
- Shimura, T.; Iwasaki, H.; Kitagawa, M.; Ebi, M.; Yamada, T.; Yamada, T.; Katano, T.; Nisie, H.; Okamoto, Y.; Ozeki, K.; et al. Urinary Cysteine-Rich Protein 61 and Trefoil Factor 3 as Diagnostic Biomarkers for Colorectal Cancer. Transl. Oncol. 2019, 12, 539–544. [Google Scholar] [CrossRef] [PubMed]
- Shimura, T.; Dayde, D.; Wang, H.; Okuda, Y.; Iwasaki, H.; Ebi, M.; Kitagawa, M.; Yamada, T.; Yamada, T.; Hanash, S.M.; et al. Novel urinary protein biomarker panel for early diagnosis of gastric cancer. Br. J. Cancer 2020, 123, 1656–1664. [Google Scholar] [CrossRef]
- Iwasaki, H.; Shimura, T.; Yamada, T.; Okuda, Y.; Natsume, M.; Kitagawa, M.; Horike, S.-I.; Kataoka, H. A novel urinary microRNA biomarker panel for detecting gastric cancer. J. Gastroenterol. 2019, 54, 1061–1069. [Google Scholar] [CrossRef]
- Okuda, Y.; Shimura, T.; Iwasaki, H.; Fukusada, S.; Nishigaki, R.; Kitagawa, M.; Katano, T.; Okamoto, Y.; Yamada, T.; Horike, S.-I.; et al. Urinary microRNA biomarkers for detecting the presence of esophageal cancer. Sci. Rep. 2021, 11, 8508. [Google Scholar] [CrossRef]
- McShane, L.M.; Altman, D.G.; Sauerbrei, W.; Taube, S.E.; Gion, M.; Clark, G.M. Reporting Recommendations for Tumor Marker Prognostic Studies. J. Clin. Oncol. 2005, 23, 9067–9072. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vandenbroucke, J.P.; von Elm, E.; Altman, D.G.; Gøtzsche, P.C.; Mulrow, C.D.; Pocock, S.J.; Poole, C.; Schlesselman, J.J.; Egger, M. Strengthening the Reporting of Observational Studies in Epidemiology (STROBE). Epidemiology 2007, 18, 805–835. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fisseler-Eckhoff, A. New TNM classification of malignant lung tumors 2009 from a pathology perspective. Der Pathol. 2009, 30 (Suppl. S2), 193–199. [Google Scholar] [CrossRef]
- Mestdagh, P.; Van Vlierberghe, P.; De Weer, A.; Muth, D.; Westermann, F.; Speleman, F.; Vandesompele, J. A novel and universal method for microRNA RT-qPCR data normalization. Genome Biol. 2009, 10, 1–10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brenner, B.M.; Bohrer, M.P.; Baylis, C.; Deen, W.M. Determinants of glomerular permselectivity: Insights derived from observations in vivo. Kidney Int. 1977, 12, 229–237. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weber, J.A.; Baxter, D.H.; Zhang, S.; Huang, D.Y.; How Huang, K.; Jen Lee, M.; Galas, D.J.; Wang, K. The MicroRNA Spectrum in 12 Body Fluids. Clin. Chem. 2010, 56, 1733–1741. [Google Scholar] [CrossRef] [PubMed]
- Sanchon, S.D.; Moreno, L.; Augé, J.M.; Serra-Burriel, M.; Cuatrecasas, M.; Moreira, L.; Martín, A.; Serradesanferm, A.; Pozo, À.; Costa, R.; et al. Identification and Validation of MicroRNA Profiles in Fecal Samples for Detection of Colorectal Cancer. Gastroenterology 2020, 158, 947–957.e4. [Google Scholar] [CrossRef]
- Armstrong, D.A.; Dessaint, J.A.; Ringelberg, C.S.; Hazlett, H.F.; Howard, L.; Abdalla, M.A.; Barnaby, R.L.; Stanton, B.A.; Cervinski, M.; Ashare, A. Pre-Analytical Handling Conditions and Small RNA Recovery from Urine for miRNA Profiling. J. Mol. Diagn. 2018, 20, 565–571. [Google Scholar] [CrossRef]
- Dyrskjøt, L.; Ostenfeld, M.S.; Bramsen, J.B.; Silahtaroglu, A.N.; Lamy, P.; Ramanathan, R.; Fristrup, N.; Jensen, J.L.; Andersen, C.L.; Zieger, K.; et al. Genomic Profiling of MicroRNAs in Bladder Cancer: miR-129 Is Associated with Poor Outcome and Promotes Cell Death In vitro. Cancer Res. 2009, 69, 4851–4860. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bandres, E.; Agirre, X.; Bitarte, N.; Ramirez, N.; Zarate, R.; Roman-Gomez, J.; Prosper, F.; Garcia-Foncillas, J. Epigenetic regulation of microRNA expression in colorectal cancer. Int. J. Cancer 2009, 125, 2737–2743. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Sun, K.-X.; Liu, B.-L.; Zong, Z.-H.; Zhao, Y. The role of glycogen synthase kinase-3β (GSK-3β) in endometrial carcinoma: A carcinogenesis, progression, prognosis, and target therapy marker. Oncotarget 2016, 7, 27538–27551. [Google Scholar] [CrossRef] [Green Version]
- Okuzaki, D.; Yamauchi, T.; Mitani, F.; Miyata, M.; Ninomiya, Y.; Watanabe, R.; Akamatsu, H.; Oneyama, C. c-Src promotes tumor progression through downregulation of microRNA-129-1-3p. Cancer Sci. 2020, 111, 418–428. [Google Scholar] [CrossRef] [Green Version]
- Du, Y.; Wang, D.; Luo, L.; Guo, J. miR-129-1-3p Promote BGC-823 Cell Proliferation by Targeting PDCD2. Anat. Rec. Adv. Integr. Anat. Evol. Biol. 2014, 297, 2273–2279. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.-L.; Zhou, X.; Han, L.; Chen, L.-Y.; Chen, L.-C.; Shi, Z.-D.; Yang, M.; Ren, Y.; Yang, J.-X.; Frank, T.S.; et al. MicroRNA-566 activates EGFR signaling and its inhibition sensitizes glioblastoma cells to nimotuzumab. Mol. Cancer 2014, 13, 63. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pan, X.; Quan, J.; Li, Z.; Zhao, L.; Zhou, L.; Jinling, X.; Weijie, X.; Guan, X.; Li, H.; Yang, S.; et al. miR-566 functions as an oncogene and a potential biomarker for prognosis in renal cell carcinoma. Biomed. Pharmacother. 2018, 102, 718–727. [Google Scholar] [CrossRef] [PubMed]
- Drusco, A.; Nuovo, G.J.; Zanesi, N.; Di Leva, G.; Pichiorri, F.; Volinia, S.; Fernandez, C.; Antenucci, A.; Costinean, S.; Bottoni, A.; et al. MicroRNA Profiles Discriminate among Colon Cancer Metastasis. PLoS ONE 2014, 9, e96670. [Google Scholar] [CrossRef]
- Di Ruocco, F.; Basso, V.; Rivoire, M.; Mehlen, P.; Ambati, J.; De Falco, S.; Tarallo, V. Alu RNA accumulation induces epithelial-to-mesenchymal transition by modulating miR-566 and is associated with cancer progression. Oncogene 2017, 37, 627–637. [Google Scholar] [CrossRef] [Green Version]

| Item | HC | CRC | p Value | ||
|---|---|---|---|---|---|
| (n = 209) | (n = 206) | ||||
| Age (years) | Median (IQR) | 69 (63–74) | 69.5 (63–75) | 0.275 | |
| Gender, n | Male | 123 | 117 | 0.672 | |
| Female | 86 | 89 | |||
| Serum Cr (mg/dL) | Mean ± SD | 0.78 ± 0.18 | 0.77 ± 0.26 | 0.787 | |
| Histological grade, n, % | well to mod | 189 | (91.7) | ||
| por | 17 | (8.3) | |||
| Location, n, % | Cecum | 20 | (9.7) | ||
| Ascending | 29 | (14.1) | |||
| Transverse | 22 | (10.7) | |||
| Descending | 8 | (3.9) | |||
| Sigmoid | 54 | (26.2) | |||
| Rectum | 73 | (35.4) | |||
| Stage, n, % | 0 | 22 | (10.7) | ||
| I | 44 | (21.4) | |||
| II | 44 | (21.4) | |||
| III | 48 | (23.3) | |||
| IV | 48 | (23.3) | |||
| Variable | 2−ΔCt (Median, IQR) | Univariate | Multivariate | |||
|---|---|---|---|---|---|---|
| Analysis | Analysis | |||||
| HC | CRC | p Value | Odds Ratio (95% CI) | p Value | ||
| (n = 140) | (n = 140) | |||||
| miR-129-1-3p | 0.00054 | 0.00150 | <0.001 | 5.59 | (2.82–11.10) | <0.001 |
| (0.00019–0.00101) | (0.00082–0.00326) | |||||
| miR-566 | 0.050 | 0.184 | <0.001 | 1.64 | (1.09–2.45) | 0.017 |
| (0.029–0.163) | (0.071–0.438) | |||||
| miR-598-5p | 0.122 | 0.273 | <0.001 | |||
| (0.070–0.266) | (0.130–0.402) | |||||
| Variable | 2−ΔCt (Median, IQR) | Univariate | Multivariate | |||
|---|---|---|---|---|---|---|
| Analysis | Analysis | |||||
| HC | CRC | p Value | Odds Ratio (95% CI) | p Value | ||
| (n = 63) | (n = 63) | |||||
| miR-129-1-3p | 0.00025 | 0.00141 | <0.001 | 5.03 | (1.99–12.70) | <0.001 |
| (0.00015–0.00079) | (0.00095–0.00218) | |||||
| miR-566 | 0.040 | 0.222 | <0.001 | 2.99 | (1.13–7.89) | 0.027 |
| (0.017–0.105) | (0.100–0.526) | |||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Iwasaki, H.; Shimura, T.; Kitagawa, M.; Yamada, T.; Nishigaki, R.; Fukusada, S.; Okuda, Y.; Katano, T.; Horike, S.-i.; Kataoka, H. A Novel Urinary miRNA Biomarker for Early Detection of Colorectal Cancer. Cancers 2022, 14, 461. https://doi.org/10.3390/cancers14020461
Iwasaki H, Shimura T, Kitagawa M, Yamada T, Nishigaki R, Fukusada S, Okuda Y, Katano T, Horike S-i, Kataoka H. A Novel Urinary miRNA Biomarker for Early Detection of Colorectal Cancer. Cancers. 2022; 14(2):461. https://doi.org/10.3390/cancers14020461
Chicago/Turabian StyleIwasaki, Hiroyasu, Takaya Shimura, Mika Kitagawa, Tamaki Yamada, Ruriko Nishigaki, Shigeki Fukusada, Yusuke Okuda, Takahito Katano, Shin-ichi Horike, and Hiromi Kataoka. 2022. "A Novel Urinary miRNA Biomarker for Early Detection of Colorectal Cancer" Cancers 14, no. 2: 461. https://doi.org/10.3390/cancers14020461
APA StyleIwasaki, H., Shimura, T., Kitagawa, M., Yamada, T., Nishigaki, R., Fukusada, S., Okuda, Y., Katano, T., Horike, S.-i., & Kataoka, H. (2022). A Novel Urinary miRNA Biomarker for Early Detection of Colorectal Cancer. Cancers, 14(2), 461. https://doi.org/10.3390/cancers14020461







