Differential Spatial Distribution of TSPO or Amino Acid PET Signal and MRI Contrast Enhancement in Gliomas
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients
2.2. Imaging
2.3. Normalization of Images
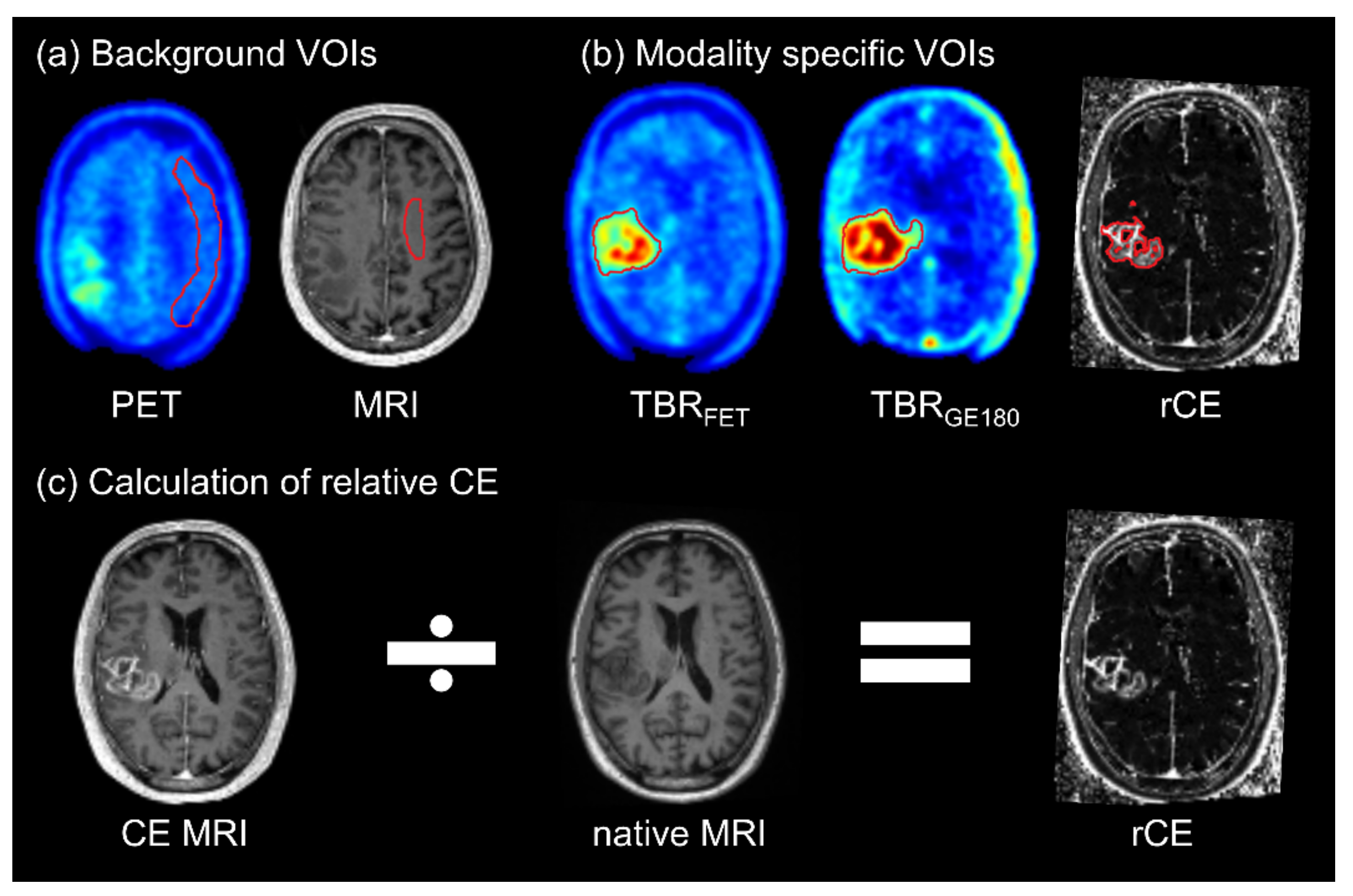
2.4. Delineation of Volumes for Voxel-Wise Analyses and Tumor Hotspots
2.5. Statistical Analyses
3. Results
3.1. Patients
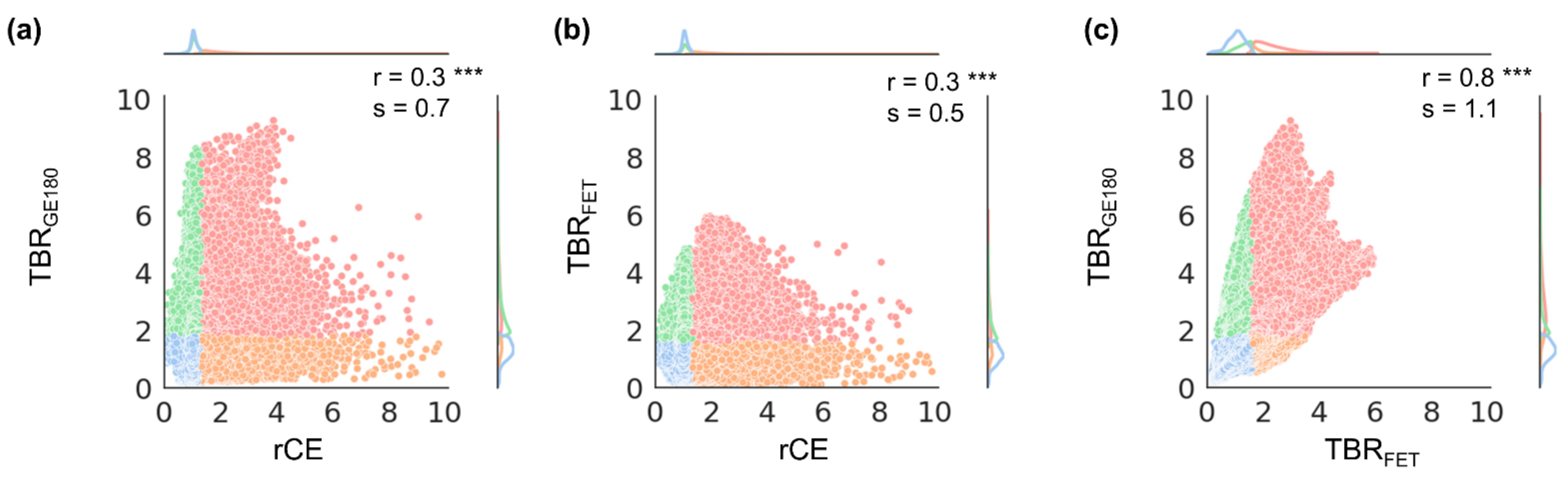
3.2. Voxel-Wise Correlation
3.3. Dice Coefficients and Hausdorff Distances for Comparison of Tumor Hotspots
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ostrom, Q.T.; Gittleman, H.; Liao, P.; Vecchione-Koval, T.; Wolinsky, Y.; Kruchko, C.; Barnholtz-Sloan, J.S. CBTRUS Statistical Report: Primary brain and other central nervous system tumors diagnosed in the United States in 2010–2014. Neuro Oncol. 2017, 19, v1–v88. [Google Scholar] [CrossRef]
- Villanueva-Meyer, J.E.; Mabray, M.C.; Cha, S. Current Clinical Brain Tumor Imaging. Neurosurgery 2017, 81, 397–415. [Google Scholar] [CrossRef]
- Albert, N.L.; Weller, M.; Suchorska, B.; Galldiks, N.; Soffietti, R.; Kim, M.M.; la Fougere, C.; Pope, W.; Law, I.; Arbizu, J.; et al. Response Assessment in Neuro-Oncology working group and European Association for Neuro-Oncology recommendations for the clinical use of PET imaging in gliomas. Neuro Oncol. 2016, 18, 1199–1208. [Google Scholar] [CrossRef] [PubMed]
- Lohmann, P.; Stavrinou, P.; Lipke, K.; Bauer, E.K.; Ceccon, G.; Werner, J.M.; Neumaier, B.; Fink, G.R.; Shah, N.J.; Langen, K.J.; et al. FET PET reveals considerable spatial differences in tumour burden compared to conventional MRI in newly diagnosed glioblastoma. Eur. J. Nucl. Med. Mol. Imaging 2019, 46, 591–602. [Google Scholar] [CrossRef] [PubMed]
- Law, I.; Albert, N.L.; Arbizu, J.; Boellaard, R.; Drzezga, A.; Galldiks, N.; la Fougere, C.; Langen, K.J.; Lopci, E.; Lowe, V.; et al. Joint EANM/EANO/RANO practice guidelines/SNMMI procedure standards for imaging of gliomas using PET with radiolabelled amino acids and [18F]FDG: Version 1.0. Eur. J. Nucl. Med. Mol. Imaging 2018. [Google Scholar] [CrossRef]
- Zinnhardt, B.; Roncaroli, F.; Foray, C.; Agushi, E.; Osrah, B.; Hugon, G.; Jacobs, A.H.; Winkeler, A. Imaging of the glioma microenvironment by TSPO PET. Eur. J. Nucl. Med. Mol. Imaging 2021. [Google Scholar] [CrossRef]
- Papadopoulos, V.; Baraldi, M.; Guilarte, T.R.; Knudsen, T.B.; Lacapere, J.J.; Lindemann, P.; Norenberg, M.D.; Nutt, D.; Weizman, A.; Zhang, M.R.; et al. Translocator protein (18kDa): New nomenclature for the peripheral-type benzodiazepine receptor based on its structure and molecular function. Trends Pharmacol. Sci. 2006, 27, 402–409. [Google Scholar] [CrossRef]
- Rupprecht, R.; Papadopoulos, V.; Rammes, G.; Baghai, T.C.; Fan, J.; Akula, N.; Groyer, G.; Adams, D.; Schumacher, M. Translocator protein (18 kDa)(TSPO) as a therapeutic target for neurological and psychiatric disorders. Nat. Rev. Drug Discov. 2010, 9, 971–988. [Google Scholar] [CrossRef]
- Boutin, H.; Murray, K.; Pradillo, J.; Maroy, R.; Smigova, A.; Gerhard, A.; Jones, P.A.; Trigg, W. 18F-GE-180: A novel TSPO radiotracer compared to 11C-R-PK11195 in a preclinical model of stroke. Eur. J. Nucl. Med. Mol. Imaging 2015, 42, 503–511. [Google Scholar] [CrossRef]
- Brendel, M.; Probst, F.; Jaworska, A.; Overhoff, F.; Korzhova, V.; Albert, N.L.; Beck, R.; Lindner, S.; Gildehaus, F.J.; Baumann, K.; et al. Glial Activation and Glucose Metabolism in a Transgenic Amyloid Mouse Model: A Triple-Tracer PET Study. J. Nucl. Med. 2016, 57, 954–960. [Google Scholar] [CrossRef] [PubMed]
- Russmann, V.; Brendel, M.; Mille, E.; Helm-Vicidomini, A.; Beck, R.; Gunther, L.; Lindner, S.; Rominger, A.; Keck, M.; Salvamoser, J.D.; et al. Identification of brain regions predicting epileptogenesis by serial [18F]GE-180 positron emission tomography imaging of neuroinflammation in a rat model of temporal lobe epilepsy. NeuroImage Clin. 2017, 15, 35–44. [Google Scholar] [CrossRef]
- Vomacka, L.; Albert, N.L.; Lindner, S.; Unterrainer, M.; Mahler, C.; Brendel, M.; Ermoschkin, L.; Gosewisch, A.; Brunegraf, A.; Buckley, C.; et al. TSPO imaging using the novel PET ligand [18F]GE-180: Quantification approaches in patients with multiple sclerosis. EJNMMI Res. 2017, 7, 89. [Google Scholar] [CrossRef]
- Unterrainer, M.; Mahler, C.; Vomacka, L.; Lindner, S.; Havla, J.; Brendel, M.; Boning, G.; Ertl-Wagner, B.; Kumpfel, T.; Milenkovic, V.M.; et al. TSPO PET with [(18)F]GE-180 sensitively detects focal neuroinflammation in patients with relapsing-remitting multiple sclerosis. Eur. J. Nucl. Med. Mol. Imaging 2018, 45, 1423–1431. [Google Scholar] [CrossRef]
- Mahler, C.; Schumacher, A.M.; Unterrainer, M.; Kaiser, L.; Hollbacher, T.; Lindner, S.; Havla, J.; Ertl-Wagner, B.; Patzig, M.; Seelos, K.; et al. TSPO PET imaging of natalizumab-associated progressive multifocal leukoencephalopathy. Brain 2021, 144, 2683–2695. [Google Scholar] [CrossRef]
- Miettinen, H.; Kononen, J.; Haapasalo, H.; Helen, P.; Sallinen, P.; Harjuntausta, T.; Helin, H.; Alho, H. Expression of peripheral-type benzodiazepine receptor and diazepam binding inhibitor in human astrocytomas: Relationship to cell proliferation. Cancer Res 1995, 55, 2691–2695. [Google Scholar]
- Unterrainer, M.; Fleischmann, D.F.; Vettermann, F.; Ruf, V.; Kaiser, L.; Nelwan, D.; Lindner, S.; Brendel, M.; Wenter, V.; Stocklein, S.; et al. TSPO PET, tumour grading and molecular genetics in histologically verified glioma: A correlative (18)F-GE-180 PET study. Eur. J. Nucl. Med. Mol. Imaging 2019, 47, 1368–1380. [Google Scholar] [CrossRef] [PubMed]
- Michelson, N.; Rincon-Torroella, J.; Quinones-Hinojosa, A.; Greenfield, J.P. Exploring the role of inflammation in the malignant transformation of low-grade gliomas. J. Neuroimmunol. 2016, 297, 132–140. [Google Scholar] [CrossRef]
- Pigeon, H.; Peres, E.A.; Truillet, C.; Jego, B.; Boumezbeur, F.; Caille, F.; Zinnhardt, B.; Jacobs, A.H.; Le Bihan, D.; Winkeler, A. TSPO-PET and diffusion-weighted MRI for imaging a mouse model of infiltrative human glioma. Neuro Oncol. 2019, 21, 755–764. [Google Scholar] [CrossRef] [PubMed]
- Zinnhardt, B.; Pigeon, H.; Theze, B.; Viel, T.; Wachsmuth, L.; Fricke, I.B.; Schelhaas, S.; Honold, L.; Schwegmann, K.; Wagner, S.; et al. Combined PET Imaging of the Inflammatory Tumor Microenvironment Identifies Margins of Unique Radiotracer Uptake. Cancer Res. 2017, 77, 1831–1841. [Google Scholar] [CrossRef]
- Jensen, P.; Feng, L.; Law, I.; Svarer, C.; Knudsen, G.M.; Mikkelsen, J.D.; de Nijs, R.; Larsen, V.A.; Dyssegaard, A.; Thomsen, G.; et al. TSPO Imaging in Glioblastoma Multiforme: A Direct Comparison Between 123I-CLINDE SPECT, 18F-FET PET, and Gadolinium-Enhanced MR Imaging. J. Nucl. Med. 2015, 56, 1386–1390. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Unterrainer, M.; Fleischmann, D.F.; Diekmann, C.; Vomacka, L.; Lindner, S.; Vettermann, F.; Brendel, M.; Wenter, V.; Ertl-Wagner, B.; Herms, J.; et al. Comparison of 18F-GE-180 and dynamic 18F-FET PET in high grade glioma: A double-tracer pilot study. Eur. J. Nucl. Med. Mol. Imaging 2019, 46, 580–590. [Google Scholar] [CrossRef] [PubMed]
- Zanotti-Fregonara, P.; Veronese, M.; Pascual, B.; Rostomily, R.C.; Turkheimer, F.; Masdeu, J.C. The validity of (18)F-GE180 as a TSPO imaging agent. Eur. J. Nucl. Med. Mol. Imaging 2019, 46, 1205–1207. [Google Scholar] [CrossRef] [PubMed]
- Zanotti-Fregonara, P.; Pascual, B.; Rostomily, R.C.; Rizzo, G.; Veronese, M.; Masdeu, J.C.; Turkheimer, F. Anatomy of 18F-GE180, a failed radioligand for the TSPO protein. Eur. J. Nucl. Med. Mol. Imaging 2020, 47, 2233–2236. [Google Scholar] [CrossRef]
- Ku, M.C.; Waiczies, S.; Niendorf, T.; Pohlmann, A. Assessment of Blood Brain Barrier Leakage with Gadolinium-Enhanced MRI. Methods Mol. Biol. 2018, 1718, 395–408. [Google Scholar] [CrossRef]
- Louis, D.N.; Perry, A.; Reifenberger, G.; von Deimling, A.; Figarella-Branger, D.; Cavenee, W.K.; Ohgaki, H.; Wiestler, O.D.; Kleihues, P.; Ellison, D.W. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A summary. Acta Neuropathol. 2016, 131, 803–820. [Google Scholar] [CrossRef]
- Owen, D.R.; Gunn, R.N.; Rabiner, E.A.; Bennacef, I.; Fujita, M.; Kreisl, W.C.; Innis, R.B.; Pike, V.W.; Reynolds, R.; Matthews, P.M.; et al. Mixed-affinity binding in humans with 18-kDa translocator protein ligands. J. Nucl. Med. 2011, 52, 24–32. [Google Scholar] [CrossRef]
- Guo, Q.; Owen, D.R.; Rabiner, E.A.; Turkheimer, F.E.; Gunn, R.N. Identifying improved TSPO PET imaging probes through biomathematics: The impact of multiple TSPO binding sites in vivo. NeuroImage 2012, 60, 902–910. [Google Scholar] [CrossRef]
- Owen, D.R.; Yeo, A.J.; Gunn, R.N.; Song, K.; Wadsworth, G.; Lewis, A.; Rhodes, C.; Pulford, D.J.; Bennacef, I.; Parker, C.A.; et al. An 18-kDa translocator protein (TSPO) polymorphism explains differences in binding affinity of the PET radioligand PBR28. J. Cereb. Blood Flow Metab. 2012, 32, 1–5. [Google Scholar] [CrossRef]
- Wickstrøm, T.; Clarke, A.; Gausemel, I.; Horn, E.; Jørgensen, K.; Khan, I.; Mantzilas, D.; Rajanayagam, T.; In’t Veld, D.J.; Trigg, W. The development of an automated and GMP compliant FASTlab™ Synthesis of [18F] GE-180; a radiotracer for imaging translocator protein (TSPO). J. Label. Comp. Radiopharm. 2014, 57, 42–48. [Google Scholar] [CrossRef] [PubMed]
- Albert, N.L.; Unterrainer, M.; Fleischmann, D.F.; Lindner, S.; Vettermann, F.; Brunegraf, A.; Vomacka, L.; Brendel, M.; Wenter, V.; Wetzel, C.; et al. TSPO PET for glioma imaging using the novel ligand (18)F-GE-180: First results in patients with glioblastoma. Eur. J. Nucl. Med. Mol. Imaging 2017, 44, 2230–2238. [Google Scholar] [CrossRef]
- Feeney, C.; Scott, G.; Raffel, J.; Roberts, S.; Coello, C.; Jolly, A.; Searle, G.; Goldstone, A.P.; Brooks, D.J.; Nicholas, R.S.; et al. Kinetic analysis of the translocator protein positron emission tomography ligand [(18)F]GE-180 in the human brain. Eur. J. Nucl. Med. Mol. Imaging 2016, 43, 2201–2210. [Google Scholar] [CrossRef]
- Unterrainer, M.; Vettermann, F.; Brendel, M.; Holzgreve, A.; Lifschitz, M.; Zahringer, M.; Suchorska, B.; Wenter, V.; Illigens, B.M.; Bartenstein, P.; et al. Towards standardization of (18)F-FET PET imaging: Do we need a consistent method of background activity assessment? EJNMMI Res. 2017, 7, 48. [Google Scholar] [CrossRef]
- Sourbron, S. Technical aspects of MR perfusion. Eur. J. Radiol. 2010, 76, 304–313. [Google Scholar] [CrossRef] [PubMed]
- Ingrisch, M.; Sourbron, S. Tracer-kinetic modeling of dynamic contrast-enhanced MRI and CT: A primer. J. Pharmacokinet. Pharmacodyn. 2013, 40, 281–300. [Google Scholar] [CrossRef]
- Albert, N.L.; Winkelmann, I.; Suchorska, B.; Wenter, V.; Schmid-Tannwald, C.; Mille, E.; Todica, A.; Brendel, M.; Tonn, J.C.; Bartenstein, P.; et al. Early static (18)F-FET-PET scans have a higher accuracy for glioma grading than the standard 20-40 min scans. Eur. J. Nucl. Med. Mol. Imaging 2016, 43, 1105–1114. [Google Scholar] [CrossRef] [PubMed]
- Pauleit, D.; Floeth, F.; Hamacher, K.; Riemenschneider, M.J.; Reifenberger, G.; Muller, H.W.; Zilles, K.; Coenen, H.H.; Langen, K.J. O-(2-[18F]fluoroethyl)-L-tyrosine PET combined with MRI improves the diagnostic assessment of cerebral gliomas. Brain 2005, 128, 678–687. [Google Scholar] [CrossRef] [PubMed]
- Wolf, I.; Vetter, M.; Wegner, I.; Bottger, T.; Nolden, M.; Schobinger, M.; Hastenteufel, M.; Kunert, T.; Meinzer, H.P. The medical imaging interaction toolkit. Med. Image Anal. 2005, 9, 594–604. [Google Scholar] [CrossRef] [PubMed]
- Nolden, M.; Zelzer, S.; Seitel, A.; Wald, D.; Muller, M.; Franz, A.M.; Maleike, D.; Fangerau, M.; Baumhauer, M.; Maier-Hein, L.; et al. The Medical Imaging Interaction Toolkit: Challenges and advances: 10 years of open-source development. Int. J. Comput. Assist Radiol. Surg. 2013, 8, 607–620. [Google Scholar] [CrossRef]
- Lowekamp, B.; Chen, D.; Ibanez, L.; Blezek, D. The Design of SimpleITK. Front. Neuroinformatics 2013, 7, 45. [Google Scholar] [CrossRef] [PubMed]
- Dice, L.R. Measures of the amount of ecologic association between species. Ecology 1945, 26, 297–302. [Google Scholar] [CrossRef]
- Dubuisson, M.-P.; Jain, A.K. A modified Hausdorff distance for object matching. In Proceedings of the 12th International Conference on Pattern Recognition, Jerusalem, Israel, 9–13 October 1994; pp. 566–568. [Google Scholar]
- Taha, A.A.; Hanbury, A. Metrics for evaluating 3D medical image segmentation: Analysis, selection, and tool. BMC Med. Imaging 2015, 15, 29. [Google Scholar] [CrossRef] [PubMed]
- Albert, N.L.; Unterrainer, M.; Brendel, M.; Kaiser, L.; Zweckstetter, M.; Cumming, P.; Bartenstein, P. In response to: The validity of (18)F-GE180 as a TSPO imaging agent. Eur. J. Nucl. Med. Mol. Imaging 2019, 46, 1208–1211. [Google Scholar] [CrossRef] [PubMed]
- Sridharan, S.; Raffel, J.; Nandoskar, A.; Record, C.; Brooks, D.J.; Owen, D.; Sharp, D.; Muraro, P.A.; Gunn, R.; Nicholas, R. Confirmation of Specific Binding of the 18-kDa Translocator Protein (TSPO) Radioligand [(18)F]GE-180: A Blocking Study Using XBD173 in Multiple Sclerosis Normal Appearing White and Grey Matter. Mol. Imaging Biol. 2019, 21, 935–944. [Google Scholar] [CrossRef] [PubMed]
- Zanotti-Fregonara, P.; Veronese, M.; Rizzo, G.; Pascual, B.; Masdeu, J.C.; Turkheimer, F.E. Letter to the Editor re: Confirmation of Specific Binding of the 18-kDa Translocator Protein (TSPO) Radioligand [(18)F]GE-180: A Blocking Study Using XBD173 in Multiple Sclerosis Normal Appearing White and Grey Matter. Mol. Imaging Biol. 2020, 22, 10–12. [Google Scholar] [CrossRef]
- Nicholas, R.; Brooks, D.; Owen, D. In Response to Letter from Fregonara et al. 2019. Mol. Imaging Biol. 2020, 22, 13–14. [Google Scholar] [CrossRef]
- Visi, E.; Hinz, R.; Punter, M.; Majid, A.; Gerhard, A.; Herholz, K. Positron emission tomography to image cerebral neuroinflammation in ischaemic stroke: A pilot study. In Positron Emission Tomography to Image Cerebral Neuroinflammation in Ischaemic Stroke: A Pilot Study; Efficacy and Mechanism Evaluation: Southampton, UK, 2020. [Google Scholar]
- Nicholas, R.; Brooks, D.; Owen, D. 18F-GE180, a radioligand for the TSPO protein: Not ready for clinical trials in multiple sclerosis. Eur. J. Nucl. Med. Mol. Imaging 2020, 47, 2242–2243. [Google Scholar] [CrossRef] [PubMed]
- Albert, N.L.; Unterrainer, M.; Kaiser, L.; Brendel, M.; Vettermann, F.J.; Holzgreve, A.; Bartenstein, P. In response to: Anatomy of 18F-GE180, a failed radioligand for the TSPO protein. Eur. J. Nucl. Med. Mol. Imaging 2020, 47, 2237–2241. [Google Scholar] [CrossRef]
- Stegmayr, C.; Oliveira, D.; Niemietz, N.; Willuweit, A.; Lohmann, P.; Galldiks, N.; Shah, N.J.; Ermert, J.; Langen, K.J. Influence of Bevacizumab on Blood-Brain Barrier Permeability and O-(2-18F-Fluoroethyl)-l-Tyrosine Uptake in Rat Gliomas. J. Nucl. Med. 2017, 58, 700–705. [Google Scholar] [CrossRef]
- Stegmayr, C.; Stoffels, G.; Kops, E.R.; Lohmann, P.; Galldiks, N.; Shah, N.J.; Neumaier, B.; Langen, K.J. Influence of Dexamethasone on O-(2-[18F]-Fluoroethyl)-L-Tyrosine Uptake in the Human Brain and Quantification of Tumor Uptake. Mol. Imaging Biol. 2019, 21, 168–174. [Google Scholar] [CrossRef]
- Fan, Z.; Calsolaro, V.; Atkinson, R.Á.; Femminella, G.D.; Waldman, A.; Buckley, C.; Trigg, W.; Brooks, D.J.; Hinz, R.; Edison, P. Flutriciclamide (18F-GE180) PET: First in human PET study of novel 3rd generation in vivo marker of human translator protein. J. Nucl. Med. 2016, 57, 1753–1759. [Google Scholar] [CrossRef]
- Thiele, F.; Ehmer, J.; Piroth, M.D.; Eble, M.J.; Coenen, H.H.; Kaiser, H.J.; Schaefer, W.M.; Buell, U.; Boy, C. The quantification of dynamic FET PET imaging and correlation with the clinical outcome in patients with glioblastoma. Phys. Med. Biol. 2009, 54, 5525–5539. [Google Scholar] [CrossRef] [PubMed]
- Vomacka, L.; Unterrainer, M.; Holzgreve, A.; Mille, E.; Gosewisch, A.; Brosch, J.; Ziegler, S.; Suchorska, B.; Kreth, F.W.; Tonn, J.C.; et al. Voxel-wise analysis of dynamic 18F-FET PET: A novel approach for non-invasive glioma characterisation. EJNMMI Res. 2018, 8, 91. [Google Scholar] [CrossRef] [PubMed]




| Overlap/Distance Measures | rCE vs. TBRGE180 | rCE vs. TBRFET | TBRFET vs. TBRGE180 |
|---|---|---|---|
| V1\V2 | 47 ± 6% | 47 ± 7% | 42 ± 10% |
| V2\V1 | 47 ± 6% | 47 ± 7% | 42 ± 10% |
| V1 ∩ V2 | 7 ± 11% | 7 ± 13% | 16 ± 20% |
| Dice coefficient | 11 ± 17% | 10 ± 19% | 23 ± 26% |
| Average Hausdorff distance | 12 ± 13 mm | 14 ± 12 mm | 9 ± 10 mm |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kaiser, L.; Holzgreve, A.; Quach, S.; Ingrisch, M.; Unterrainer, M.; Dekorsy, F.J.; Lindner, S.; Ruf, V.; Brosch-Lenz, J.; Delker, A.; et al. Differential Spatial Distribution of TSPO or Amino Acid PET Signal and MRI Contrast Enhancement in Gliomas. Cancers 2022, 14, 53. https://doi.org/10.3390/cancers14010053
Kaiser L, Holzgreve A, Quach S, Ingrisch M, Unterrainer M, Dekorsy FJ, Lindner S, Ruf V, Brosch-Lenz J, Delker A, et al. Differential Spatial Distribution of TSPO or Amino Acid PET Signal and MRI Contrast Enhancement in Gliomas. Cancers. 2022; 14(1):53. https://doi.org/10.3390/cancers14010053
Chicago/Turabian StyleKaiser, Lena, Adrien Holzgreve, Stefanie Quach, Michael Ingrisch, Marcus Unterrainer, Franziska J. Dekorsy, Simon Lindner, Viktoria Ruf, Julia Brosch-Lenz, Astrid Delker, and et al. 2022. "Differential Spatial Distribution of TSPO or Amino Acid PET Signal and MRI Contrast Enhancement in Gliomas" Cancers 14, no. 1: 53. https://doi.org/10.3390/cancers14010053
APA StyleKaiser, L., Holzgreve, A., Quach, S., Ingrisch, M., Unterrainer, M., Dekorsy, F. J., Lindner, S., Ruf, V., Brosch-Lenz, J., Delker, A., Böning, G., Suchorska, B., Niyazi, M., Wetzel, C. H., Riemenschneider, M. J., Stöcklein, S., Brendel, M., Rupprecht, R., Thon, N., ... Albert, N. L. (2022). Differential Spatial Distribution of TSPO or Amino Acid PET Signal and MRI Contrast Enhancement in Gliomas. Cancers, 14(1), 53. https://doi.org/10.3390/cancers14010053









