Minimal Residual Disease in Acute Lymphoblastic Leukemia: Current Practice and Future Directions
Abstract
Simple Summary
Abstract
1. Introduction
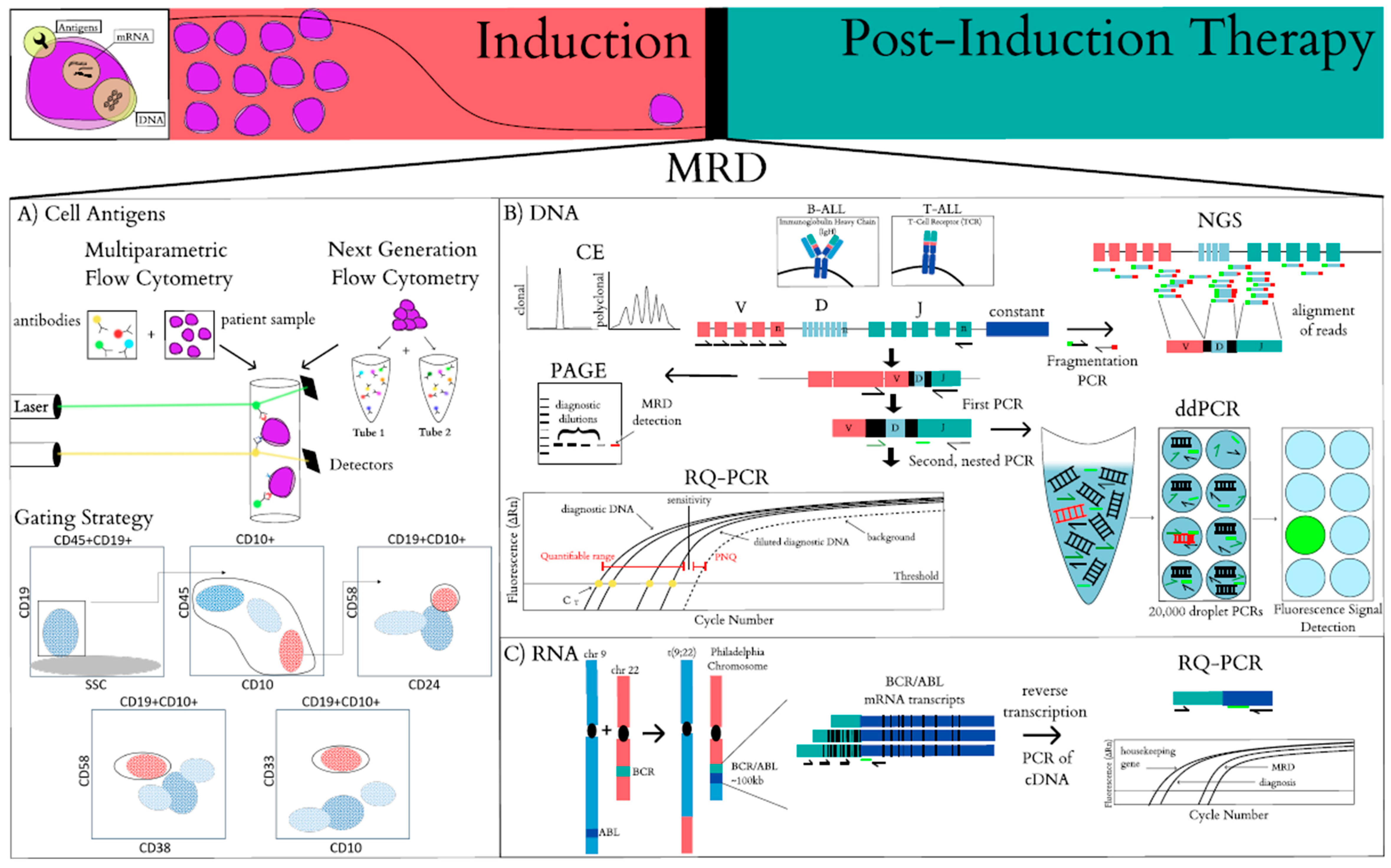
2. Methods Of Minimal Residual Disease Detection
2.1. Multiparametric Flow Cytometry for Surface Proteins
2.2. Polymerase Chain Reaction (PCR) Amplification of Antigen Receptor Rearrangements
2.3. PCR Amplification of Chromosomal Translocations
2.4. Next Generation Sequencing
3. Clinical Relevance Of Minimal Residual Disease
4. MRD Status Interaction with Other Genetic High Risk Groups
4.1. Infant B-ALL
4.2. Philadelphia Chromosome Positive B-ALL
4.3. Philadelphia (Ph)-Like B-ALL
4.4. Hypodiploid B-ALL
4.5. T-Cell Acute Lymphoblastic Leukemia (T-ALL)
5. MRD Prognostic Value Peri-Stem Cell Transplant
6. Current Recommendations and Dilemmas
Author Contributions
Funding
Conflicts of Interest
References
- Howlader, N.; Noone, A.M.; Krapcho, M.; Miller, D.; Brest, A.; Yu, M.; Ruhl, J.; Tatalovich, Z.; Mariotto, A.; Lewis, D.R.; et al. SEER Cancer Statistics Review, 1975–2017; National Cancer Institute: Bethesda, MD, USA, 2020. [Google Scholar]
- Pui, C.H.; Yang, J.J.; Hunger, S.P.; Pieters, R.; Schrappe, M.; Biondi, A.; Vora, A.; Baruchel, A.; Silverman, L.B.; Schmiegelow, K.; et al. Childhood Acute Lymphoblastic Leukemia: Progress Through Collaboration. J. Clin. Oncol. 2015, 33, 2938–2948. [Google Scholar] [CrossRef]
- Hunger, S.P.; Lu, X.; Devidas, M.; Camitta, B.M.; Gaynon, P.S.; Winick, N.J.; Reaman, G.H.; Carroll, W.L. Improved survival for children and adolescents with acute lymphoblastic leukemia between 1990 and 2005: A report from the children’s oncology group. J. Clin. Oncol. 2012, 30, 1663–1669. [Google Scholar] [CrossRef] [PubMed]
- Lavi, O.; Gottesman, M.M.; Levy, D. The dynamics of drug resistance: A mathematical perspective. Drug Resist. Updates 2012, 15, 90–97. [Google Scholar] [CrossRef] [PubMed]
- Bartram, J.; Patel, B.; Fielding, A.K. Monitoring MRD in ALL: Methodologies, technical aspects and optimal time points for measurement. Semin. Hematol. 2020, 57, 142–148. [Google Scholar] [CrossRef] [PubMed]
- McKinnon, K.M. Flow Cytometry: An Overview. Curr. Protoc. Immunol. 2018, 120, 5.1.1–5.1.11. [Google Scholar] [CrossRef]
- McKinnon, K.M. Multiparameter Conventional Flow Cytometry. In Flow Cytometry Protocols; Methods in Molecular Biology; Humana Press: New York, NY, USA, 2018; Volume 1678, pp. 139–150. [Google Scholar] [CrossRef]
- Brüggemann, M.; Gökbuget, N.; Kneba, M. Acute lymphoblastic leukemia: Monitoring minimal residual disease as a therapeutic principle. Semin. Oncol. 2012, 39, 47–57. [Google Scholar] [CrossRef]
- Dworzak, M.N.; Gaipa, G.; Ratei, R.; Veltroni, M.; Schumich, A.; Maglia, O.; Karawajew, L.; Benetello, A.; Pötschger, U.; Husak, Z.; et al. Standardization of flow cytometric minimal residual disease evaluation in acute lymphoblastic leukemia: Multicentric assessment is feasible. Cytom. Part B: Clin. Cytom. 2008, 74, 331–340. [Google Scholar] [CrossRef]
- Gaipa, G.; Basso, G.; Maglia, O.; Leoni, V.; Faini, A.; Cazzaniga, G.; Bugarin, C.; Veltroni, M.; Michelotto, B.; Ratei, R.; et al. Drug-induced immunophenotypic modulation in childhood ALL: Implications for minimal residual disease detection. Leukemia 2005, 19, 49–56. [Google Scholar] [CrossRef]
- Wood, B.L. Flow cytometric monitoring of residual disease in acute leukemia. Methods Mol. Biol. 2013, 999, 123–136. [Google Scholar]
- Della Starza, I.; Chiaretti, S.; De Propris, M.S.; Elia, L.; Cavalli, M.; De Novi, L.A.; Soscia, R.; Messina, M.; Vitale, A.; Guarini, A.; et al. Minimal Residual Disease in Acute Lymphoblastic Leukemia: Technical and Clinical Advances. Front. Oncol. 2019, 9, 726. [Google Scholar] [CrossRef]
- Chen, X.; Wood, B.L. How do we measure MRD in ALL and how should measurements affect decisions. Re: Treatment and prognosis? Best Pract. Res. Clin. Haematol. 2017, 30, 237–248. [Google Scholar] [CrossRef] [PubMed]
- Flores-Montero, J.; Sanoja-Flores, L.; Paiva, B.; Puig, N.; García-Sánchez, O.; Böttcher, S.; Van Der Velden, V.H.J.; Pérez-Morán, J.-J.; Vidriales, M.-B.; García-Sanz, R.; et al. Next Generation Flow for highly sensitive and standardized detection of minimal residual disease in multiple myeloma. Leukemia 2017, 31, 2094–2103. [Google Scholar] [CrossRef] [PubMed]
- Theunissen, P.; Mejstrikova, E.; Sedek, L.; Van Der Sluijs-Gelling, A.J.; Gaipa, G.; Bartels, M.; Da Costa, E.S.; Kotrová, M.; Novakova, M.; Sonneveld, E.; et al. Standardized flow cytometry for highly sensitive MRD measurements in B-cell acute lymphoblastic leukemia. Blood 2017, 129, 347–357. [Google Scholar] [CrossRef] [PubMed]
- Kruse, A.; Abdel-Azim, N.; Kim, Y.-M.; Ruan, Y.; Phan, V.; Ogana, H.; Wang, W.; Lee, R.; Gang, E.J.; Khazal, S.; et al. Minimal Residual Disease Detection in Acute Lymphoblastic Leukemia. Int. J. Mol. Sci. 2020, 21, 1054. [Google Scholar] [CrossRef] [PubMed]
- Roth, D.B. V(D)J Recombination: Mechanism, Errors, and Fidelity. Microbiol. Spectr. 2014, 2. [Google Scholar] [CrossRef]
- Van Dongen, J.J.; Van Der Velden, V.; Brüggemann, M.; Orfao, A. Minimal residual disease diagnostics in acute lymphoblastic leukemia: Need for sensitive, fast, and standardized technologies. Blood 2015, 125, 3996–4009. [Google Scholar] [CrossRef]
- Beaubier, N.T.; Hart, A.P.; Bartolo, C.; Willman, C.L.; Viswanatha, D.S. Comparison of capillary electrophoresis and polyacrylamide gel electrophoresis for the evaluation of T and B cell clonality by polymerase chain reaction. Diagn. Mol. Pathol. 2000, 9, 121–131. [Google Scholar] [CrossRef]
- Brisco, M.J.; Condon, J.; Sykes, P.J.; Neoh, S.H.; Morley, A.A. Detection and quantitation of neoplastic cells in acute lymphoblastic leukaemia, by use of the polymerase chain reaction. Br. J. Haematol. 1991, 79, 211–217. [Google Scholar] [CrossRef]
- d’Auriol, L.; MacIntyre, E.; Galibert, F.; Sigaux, F. In vitro amplification of T cell gamma gene rearrangements: A new tool for the assessment of minimal residual disease in acute lymphoblastic leukemias. Leukemia 1989, 3, 155–158. [Google Scholar]
- Voena, C.; Ladetto, M.; Astolfi, M.; Provan, D.; Gribben, J.G.; Boccadoro, M.; Pileri, A.; Corradini, P. A novel nested-PCR strategy for the detection of rearranged immunoglobulin heavy-chain genes in B cell tumors. Leukemia 1997, 11, 1793–1798. [Google Scholar] [CrossRef]
- Brisco, M.; Condon, J.; Hughes, E.; Neoh, S.-H.; Sykes, P.; Seshadri, R.; Morley, A.; Toogood, I.; Waters, K.; Tauro, G.; et al. Outcome prediction in childhood acute lymphoblastic leukaemia by molecular quantification of residual disease at the end of induction. Lancet 1994, 343, 196–200. [Google Scholar] [CrossRef]
- van Dongen, J.J.; Seriu, T.; Panzer-Grümayer, E.R.; Biondi, A.; Pongers-Willemse, M.J.; Corral, L.; Stolz, F.; Schrappe, M.; Masera, G.; Kamps, W.A.; et al. Prognostic value of minimal residual disease in acute lymphoblastic leukaemia in childhood. Lancet 1998, 352, 1731–1738. [Google Scholar] [CrossRef]
- Cavé, H.; Guidal, C.; Rohrlich, P.; Delfau, M.H.; Broyart, A.; Lescoeur, B.; Rahimy, C.; Fenneteau, O.; Monplaisir, N.; D’Auriol, L. Prospective monitoring and quantitation of residual blasts in childhood acute lymphoblastic leukemia by polymerase chain reaction study of delta and gamma T-cell receptor genes. Blood 1994, 83, 1892–1902. [Google Scholar] [CrossRef] [PubMed]
- Van Der Velden, V.H.; Hochhaus, A.; Cazzaniga, G.; Szczepanski, T.; Gabert, J.; Van Dongen, J.J.M. Detection of minimal residual disease in hematologic malignancies by real-time quantitative PCR: Principles, approaches, and laboratory aspects. Leukemia 2003, 17, 1013–1034. [Google Scholar] [CrossRef] [PubMed]
- Van Der Velden, V.H.; Cazzaniga, G.; Schrauder, A.; Hancock, J.F.; Bader, P.; Panzer-Grumayer, E.R.; Flohr, T.R.; Sutton, R.S.; Cave, H.; Madsen, H.O.; et al. Analysis of minimal residual disease by Ig/TCR gene rearrangements: Guidelines for interpretation of real-time quantitative PCR data. Leukemia 2007, 21, 604–611. [Google Scholar] [CrossRef] [PubMed]
- Coccaro, N.; Tota, G.; Anelli, L.; Zagaria, A.; Specchia, G.; Albano, F. Digital PCR: A Reliable Tool for Analyzing and Monitoring Hematologic Malignancies. Int. J. Mol. Sci. 2020, 21, 3141. [Google Scholar] [CrossRef]
- Della Starza, I.; Nunes, V.; Cavalli, M.; De Novi, L.A.; del Ilaria, G.; Apicella, V.; Vitale, A.; Testi, A.M.; Del Giudice, I.; Chiaretti, S.; et al. Comparative analysis between RQ-PCR and digital-droplet-PCR of immunoglobulin/T-cell receptor gene rearrangements to monitor minimal residual disease in acute lymphoblastic leukaemia. Br. J. Haematol. 2016, 174, 541–549. [Google Scholar] [CrossRef]
- Della Starza, I.; Nunes, V.; Lovisa, F.; Silvestri, D.; Cavalli, M.; Garofalo, A.; Campeggio, M.; De Novi, L.A.; Oggioni, C.; Biondi, A.; et al. Digital-Droplet PCR, an Accurate Method for IG/TR PCR-MRD Stratification in Childhood Acute Lymphoblastic Leukemia. Blood 2018, 132 (Suppl. S1), 1544. [Google Scholar] [CrossRef]
- Saliba, J.; Evensen, N.A.; Meyer, J.A.; Newman, D.; Nersting, J.; Narang, S.; Ma, X.; Schmiegelow, K.; Carroll, W.L. Feasibility of monitoring peripheral blood to detect emerging clones in children with acute lymphoblastic leukemia. Pediatr. Blood Cancer 2020, 67, e28306. [Google Scholar] [CrossRef]
- Bernt, K.M.; Hunger, S.P. Current concepts in pediatric Philadelphia chromosome—Positive acute lymphoblastic leukemia. Front. Oncol. 2014, 4, 54. [Google Scholar] [CrossRef]
- Lazic, J.; Tosic, N.; Dokmanovic, L.; Krstovski, N.; Rodic, P.; Pavlović, S.; Janic, D. Clinical features of the most common fusion genes in childhood acute lymphoblastic leukemia. Med. Oncol. 2010, 27, 449–453. [Google Scholar] [CrossRef] [PubMed]
- Cazzaniga, G.; Rossi, V.; Biondi, A. Monitoring minimal residual disease using chromosomal translocations in childhood ALL. Best Pract. Res. Clin. Haematol. 2002, 15, 21–35. [Google Scholar] [CrossRef]
- Zaliova, M.; Fronkova, E.; Krejcikova, K.; Muzikova, K.; Mejstrikova, E.; Stary, J.; Trka, J.; Zuna, J. Quantification of fusion transcript reveals a subgroup with distinct biological properties and predicts relapse in BCR/ABL-positive ALL: Implications for residual disease monitoring. Leukemia 2009, 23, 944–951. [Google Scholar] [CrossRef]
- Bruggemann, M.; Kotrová, M.; Knecht, H.; Bartram, J.; Boudjogrha, M.; Bystry, V.; Fazio, G.; Fronkova, E.; Giraud, M.; Grioni, A.; et al. Standardized next-generation sequencing of immunoglobulin and T-cell receptor gene recombinations for MRD marker identification in acute lymphoblastic leukaemia; A EuroClonality-NGS validation study. Leukemia 2019, 33, 2241–2253. [Google Scholar] [CrossRef] [PubMed]
- Coccaro, N.; Anelli, L.; Zagaria, A.; Specchia, G.; Albano, F. Next-Generation Sequencing in Acute Lymphoblastic Leukemia. Int. J. Mol. Sci. 2019, 20, 2929. [Google Scholar] [CrossRef] [PubMed]
- Kotrova, M.; Van Der Velden, V.H.J.; Van Dongen, J.J.M.; Formankova, R.; Sedlacek, P.; Brüggemann, M.; Zuna, J.; Stary, J.; Trka, J.; Fronkova, E. Next-generation sequencing indicates false-positive MRD results and better predicts prognosis after SCT in patients with childhood ALL. Bone Marrow Transplant. 2017, 52, 962–968. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Jiang, N.; Lim, E.H.; Ni Chin, W.H.; Lu, Y.; Chiew, K.H.; Kham, S.K.Y.; Yang, W.; Quah, T.C.; Lin, H.P.; et al. Identifying IGH disease clones for MRD monitoring in childhood B-cell acute lymphoblastic leukemia using RNA-Seq. Leukemia 2020, 34, 2418–2429. [Google Scholar] [CrossRef] [PubMed]
- Abel, H.J.; Al-Kateb, H.; Cottrell, C.E.; Bredemeyer, A.J.; Pritchard, C.C.; Grossmann, A.H.; Wallander, M.L.; Pfeifer, J.D.; Lockwood, C.M.; Duncavage, E.J. Detection of gene rearrangements in targeted clinical next-generation sequencing. J. Mol. Diagn. 2014, 16, 405–417. [Google Scholar] [CrossRef]
- Borowitz, M.J.; Wood, B.L.; Devidas, M.; Loh, M.L.; Raetz, E.A.; Salzer, W.L.; Nachman, J.B.; Carroll, A.J.; Heerema, N.A.; Gastier-Foster, J.M.; et al. Prognostic significance of minimal residual disease in high risk B-ALL: A report from Children’s Oncology Group study AALL0232. Blood 2015, 126, 964–971. [Google Scholar] [CrossRef]
- Sutton, R.; Venn, N.C.; Tolisano, J.; Bahar, A.Y.; Giles, J.E.; Ashton, L.J.; Teague, L.; Rigutto, G.; Waters, K.; Marshall, G.M.; et al. Clinical significance of minimal residual disease at day 15 and at the end of therapy in childhood acute lymphoblastic leukaemia. Br. J. Haematol. 2009, 146, 292–299. [Google Scholar] [CrossRef]
- Bartram, J.; Wade, R.; Vora, A.; Hancock, J.; Mitchell, C.; Kinsey, S.; Steward, C.; Moppett, J.; Goulden, N. Excellent outcome of minimal residual disease-defined low-risk patients is sustained with more than 10 years follow-up: Results of UK paediatric acute lymphoblastic leukaemia trials 1997–2003. Arch. Dis. Child. 2016, 101, 449–454. [Google Scholar] [CrossRef] [PubMed]
- Borowitz, M.J.; Devidas, M.; Hunger, S.P.; Bowman, W.P.; Carroll, A.J.; Carroll, W.L.; Linda, S.; Martin, P.L.; Pullen, D.J.; Viswanatha, D.; et al. Clinical significance of minimal residual disease in childhood acute lymphoblastic leukemia and its relationship to other prognostic factors: A Children’s Oncology Group study. Blood 2008, 111, 5477–5485. [Google Scholar] [CrossRef] [PubMed]
- Conter, V.; Bartram, C.R.; Valsecchi, M.G.; Schrauder, A.; Panzer-Grümayer, R.; Möricke, A.; Aricò, M.; Zimmermann, M.; Mann, G.; De Rossi, G.; et al. Molecular response to treatment redefines all prognostic factors in children and adolescents with B-cell precursor acute lymphoblastic leukemia: Results in 3184 patients of the AIEOP-BFM ALL 2000 study. Blood 2010, 115, 3206–3214. [Google Scholar] [CrossRef] [PubMed]
- Schrappe, M.; Valsecchi, M.G.; Bartram, C.R.; Schrauder, A.; Panzer-Grümayer, R.; Möricke, A.; Parasole, R.; Zimmermann, M.; Dworzak, M.; Buldini, B.; et al. Late MRD response determines relapse risk overall and in subsets of childhood T-cell ALL: Results of the AIEOP-BFM-ALL 2000 study. Blood 2011, 118, 2077–2084. [Google Scholar] [CrossRef] [PubMed]
- Borowitz, M.J.; Wood, B.L.; Devidas, M.; Loh, M.L.; Raetz, E.A.; Larsen, E.; Maloney, K.W.; Carroll, A.J.; Friedmann, A.M.; Gastier-Foster, J.M.; et al. Assessment of end induction minimal residual disease (MRD) in childhood B precursor acute lymphoblastic leukemia (ALL) to eliminate the need for day 14 marrow examination: A Children’s Oncology Group study. J. Clin. Oncol. 2013, 31 (Suppl. 15), 10001. [Google Scholar] [CrossRef]
- Stow, P.; Key, L.; Chen, X.; Pan, Q.; Neale, G.A.; Coustan-Smith, E.; Mullighan, C.G.; Zhou, Y.; Pui, C.-H.; Campana, D. Clinical significance of low levels of minimal residual disease at the end of remission induction therapy in childhood acute lymphoblastic leukemia. Blood 2010, 115, 4657–4663. [Google Scholar] [CrossRef]
- Berry, D.A.; Zhou, S.; Higley, H.; Mukundan, L.; Fu, S.; Reaman, G.H.; Wood, B.L.; Kelloff, G.J.; Jessup, J.M.; Radich, J.P. Association of Minimal Residual Disease with Clinical Outcome in Pediatric and Adult Acute Lymphoblastic Leukemia: A Meta-analysis. JAMA Oncol. 2017, 3, e170580. [Google Scholar] [CrossRef]
- Wood, B.; Wu, D.; Crossley, B.; Dai, Y.; Williamson, D.; Gawad, C.; Borowitz, M.J.; Devidas, M.; Maloney, K.W.; Larsen, E.; et al. Measurable residual disease detection by high-throughput sequencing improves risk stratification for pediatric B-ALL. Blood 2018, 131, 1350–1359. [Google Scholar] [CrossRef]
- Pui, C.H.; Pei, D.; Raimondi, S.C.; Coustan-Smith, E.; Jeha, S.; Cheng, C.; Bowman, W.P.; Sandlund, J.T.; Ribeiro, R.C.; Rubnitz, J.E.; et al. Clinical impact of minimal residual disease in children with different subtypes of acute lymphoblastic leukemia treated with Response-Adapted therapy. Leukemia 2017, 31, 333–339. [Google Scholar] [CrossRef] [PubMed]
- Pieters, R.; De Groot-Kruseman, H.; Van Der Velden, V.; Fiocco, M.; Berg, H.V.D.; De Bont, E.; Egeler, R.M.; Hoogerbrugge, P.; Kaspers, G.; Van Der Schoot, E.; et al. Successful Therapy Reduction and Intensification for Childhood Acute Lymphoblastic Leukemia Based on Minimal Residual Disease Monitoring: Study ALL10 From the Dutch Childhood Oncology Group. J. Clin. Oncol. 2016, 34, 2591–2601. [Google Scholar] [CrossRef]
- O’Connor, D.; Enshaei, A.; Bartram, J.; Hancock, J.; Harrison, C.J.; Hough, R.; Samarasinghe, S.; Schwab, C.; Vora, A.; Wade, R.; et al. Genotype-Specific Minimal Residual Disease Interpretation Improves Stratification in Pediatric Acute Lymphoblastic Leukemia. J. Clin. Oncol. 2018, 36, 34–43. [Google Scholar] [CrossRef]
- Coustan-Smith, E.; Ribeiro, R.C.; Stow, P.; Zhou, Y.; Pui, C.-H.; Rivera, G.K.; Pedrosa, F.; Campana, D. A simplified flow cytometric assay identifies children with acute lymphoblastic leukemia who have a superior clinical outcome. Blood 2006, 108, 97–102. [Google Scholar] [CrossRef][Green Version]
- Van Der Velden, V.H.; Corral, L.; Valsecchi, M.G.; Jansen, M.W.J.C.; De Lorenzo, P.; Cazzaniga, G.; Panzer-Grümayer, E.R.; Schrappe, M.; Schrauder, A.; Meyer, C.; et al. Prognostic significance of minimal residual disease in infants with acute lymphoblastic leukemia treated within the Interfant-99 protocol. Leukemia 2009, 23, 1073–1079. [Google Scholar] [CrossRef] [PubMed]
- Cazzaniga, G.; De Lorenzo, P.; Alten, J.; Röttgers, S.; Hancock, J.; Saha, V.; Castor, A.; Madsen, H.O.; Gandemer, V.; Cavé, H.; et al. Predictive value of minimal residual disease in Philadelphia-chromosome-positive acute lymphoblastic leukemia treated with imatinib in the European intergroup study of post-induction treatment of Philadelphia-chromosome-positive acute lymphoblastic leukemia, based on immunoglobulin/T-cell receptor and BCR/ABL1 methodologies. Haematologica 2018, 103, 107–115. [Google Scholar] [CrossRef] [PubMed]
- Hovorkova, L.; Zaliova, M.; Venn, N.C.; Bleckmann, K.; Trkova, M.; Potuckova, E.; Vaskova, M.; Linhartova, J.; Polakova, K.M.; Fronkova, E.; et al. Monitoring of childhood ALL using BCR-ABL1 genomic breakpoints identifies a subgroup with CML-like biology. Blood 2017, 129, 2771–2781. [Google Scholar] [CrossRef]
- Jeha, S.; Coustan-Smith, E.; Pei, D.; Sandlund, J.T.; Rubnitz, J.E.; Howard, S.C.; Inaba, H.; Bhojwani, D.; Metzger, M.L.; Cheng, C.; et al. Impact of tyrosine kinase inhibitors on minimal residual disease and outcome in childhood Philadelphia chromosome-positive acute lymphoblastic leukemia. Cancer 2014, 120, 1514–1519. [Google Scholar] [CrossRef]
- Schultz, K.R.; Carroll, A.J.; Heerema, N.A.; Bowman, W.P.; Aledo, A.; Slayton, W.B.; Sather, H.N.; Devidas, M.; Zheng, H.W.; Davies, S.M.; et al. Long-term follow-up of imatinib in pediatric Philadelphia chromosome-positive acute lymphoblastic leukemia: Children’s Oncology Group Study AALL0031. Leukemia 2014, 28, 1467–1471. [Google Scholar] [CrossRef]
- Slayton, W.B.; Schultz, K.R.; Kairalla, J.A.; Devidas, M.; Mi, X.; Pulsipher, M.A.; Chang, B.H.; Mullighan, C.; Iacobucci, I.; Silverman, L.B.; et al. Dasatinib Plus Intensive Chemotherapy in Children, Adolescents, and Young Adults with Philadelphia Chromosome–Positive Acute Lymphoblastic Leukemia: Results of Children’s Oncology Group Trial AALL0622. J. Clin. Oncol. 2018, 36, 2306–2314. [Google Scholar] [CrossRef] [PubMed]
- Biondi, A.; Schrappe, M.; De Lorenzo, P.; Castor, A.; Lucchini, G.; Gandemer, V.; Pieters, R.; Stary, J.; Escherich, G.; Campbell, M.; et al. Imatinib after induction for treatment of children and adolescents with Philadelphia-chromosome-positive acute lymphoblastic leukaemia (EsPhALL): A randomised, open-label, intergroup study. Lancet Oncol. 2012, 13, 936–945. [Google Scholar] [CrossRef]
- Schultz, K.R.; Devidas, M.; Bowman, W.P.; Aledo, A.; Slayton, W.B.; Sather, H.; Zheng, H.W.; Davies, S.M.; Gaynon, P.S.; Trigg, M.; et al. Philadelphia chromosome-negative very high-risk acute lymphoblastic leukemia in children and adolescents: Results from Children’s Oncology Group Study AALL0031. Leukemia 2014, 28, 964–967. [Google Scholar] [CrossRef]
- Xue, Y.J.; Cheng, Y.-F.; Lu, A.-D.; Wang, Y.; Zuo, Y.-X.; Yan, C.-H.; Wu, J.; Sun, Y.-Q.; Suo, P.; Chen, Y.-H.; et al. Allogeneic Hematopoietic Stem Cell Transplantation, Especially Haploidentical, May Improve Long-Term Survival for High-Risk Pediatric Patients with Philadelphia Chromosome–Positive Acute Lymphoblastic Leukemia in the Tyrosine Kinase Inhibitor Era. Biol. Blood Marrow Transpl. 2019, 25, 1611–1620. [Google Scholar] [CrossRef] [PubMed]
- Van Der Veer, A.; Waanders, E.; Pieters, R.; Willemse, M.; Van Reijmersdal, S.; Russell, L.; Harrison, C.; Evans, W.; Van Der Velden, V.; Hoogerbrugge, P.; et al. Independent prognostic value of BCR-ABL1-like signature and IKZF1 deletion, but not high CRLF2 expression, in children with B-cell precursor ALL. Blood 2013, 122, 2622–2629. [Google Scholar] [CrossRef] [PubMed]
- Kiyokawa, N.; Iijima, K.; Yoshihara, H.; Ohki, K.; Kato, M.; Fukushima, T.; Kikuchi, A.; Fujimoto, J.; Hayashi, Y.; Koh, K.; et al. An Analysis of Ph-Like ALL In Japanese Patients. Blood 2013, 122, 352. [Google Scholar] [CrossRef]
- Te Kronnie, G.; Silvestri, D.; Vendramini, E.; Fazio, G.; Locatelli, F.; Conter, V.; Borga, C.; Valsecchi, M.G.; Biondi, A.; Basso, G.; et al. Philadelphia-Like Signature in Childhood Acute Lymphoblastic Leukemia: The AIEOP Experience. Blood 2013, 122, 353. [Google Scholar] [CrossRef]
- Roberts, K.G.; Payne-Turner, D.; Pei, D.; Becksfort, J.; Harvey, R.C.; Li, Y.; Song, G.; Cheng, J.; Ma, J.; Zhang, J.; et al. Integrated Genomic and Mutational Profiling of Adolescent and Young Adult ALL Identifies a High Frequency of BCR-ABL1-Like ALL with Very Poor Outcome. Blood 2013, 122, 825. [Google Scholar] [CrossRef]
- Loh, M.L.; Zhang, J.; Harvey, R.C.; Roberts, K.; Payne-Turner, D.; Kang, H.; Wu, G.; Chen, X.; Becksfort, J.; Edmonson, M.; et al. Tyrosine kinome sequencing of pediatric acute lymphoblastic leukemia: A report from the Children’s Oncology Group TARGET Project. Blood 2013, 121, 485–488. [Google Scholar] [CrossRef] [PubMed]
- Mullighan, C.G.; Jeha, S.; Pei, D.; Payne-Turner, D.; Coustan-Smith, E.; Roberts, K.G.; Waanders, E.; Choi, J.K.; Ma, X.; Raimondi, S.C.; et al. Outcome of children with hypodiploid ALL treated with risk-directed therapy based on MRD levels. Blood 2015, 126, 2896–2899. [Google Scholar] [CrossRef] [PubMed]
- McNeer, J.L.; Devidas, M.; Dai, Y.; Carroll, A.J.; Heerema, N.A.; Gastier-Foster, J.M.; Kahwash, S.B.; Borowitz, M.J.; Wood, B.L.; Larsen, E.; et al. Hematopoietic Stem-Cell Transplantation Does Not Improve the Poor Outcome of Children with Hypodiploid Acute Lymphoblastic Leukemia: A Report from Children’s Oncology Group. J. Clin. Oncol. 2019, 37, 780–789. [Google Scholar] [CrossRef]
- Pui, C.-H.; Rebora, P.; Schrappe, M.; Attarbaschi, A.; Baruchel, A.; Basso, G.; Cavé, H.; Elitzur, S.; Koh, K.; Liu, H.-C.; et al. Outcome of Children with Hypodiploid Acute Lymphoblastic Leukemia: A Retrospective Multinational Study. J. Clin. Oncol. 2019, 37, 770–779. [Google Scholar] [CrossRef]
- Winter, S.S.; Devidas, M.; Chen, M.S.; Asselin, B.; Carroll, W.L.; Wood, B.L.; Esiashvili, N.; Nikki, B.J.; Hayashi, R.J.; Loh, M.L.; et al. Capizzi-Style Methotrexate with Pegasparagase (C-MTX) Is Superior to High-Dose Methotrexate (HDMTX) in T-Lineage Acute Lymphoblastic Leukemia (T-ALL): Results from Children’s Oncology Group (COG) AALL0434. Blood 2015, 126, 794. [Google Scholar] [CrossRef]
- Möricke, A.; Zimmermann, M.; Valsecchi, M.G.; Stanulla, M.; Biondi, A.; Mann, G.; Locatelli, F.; Cazzaniga, G.; Niggli, F.; Aricò, M.; et al. Dexamethasone vs. prednisone in induction treatment of pediatric ALL: Results of the randomized trial AIEOP-BFM ALL 2000. Blood 2016, 127, 2101–2112. [Google Scholar] [CrossRef] [PubMed]
- Patrick, K.; Wade, R.; Goulden, N.; Mitchell, C.; Moorman, A.V.; Rowntree, C.; Jenkinson, S.; Hough, R.; Vora, A. Outcome for children and young people with Early T-cell precursor acute lymphoblastic leukaemia treated on a contemporary protocol, UKALL 2003. Br. J. Haematol. 2014, 166, 421–424. [Google Scholar] [CrossRef]
- Place, A.E.; Stevenson, K.E.; Harris, M.H.; Weinberg, O.K.; Vrooman, L.M.; Hunt, S.K.; Ahmed, S.S.; Athale, U.H.; Clavell, L.A.; Cole, P.D.; et al. Outcome of childhood T-cell acute lymphoblastic leukemia (T-ALL): Results from DFCI protocol 05-001. J. Clin. Oncol. 2014, 32 (Suppl. 15), 10015. [Google Scholar] [CrossRef]
- Place, A.E.; Stevenson, K.E.; Vrooman, L.M.; Harris, M.H.; Hunt, S.K.; O’Brien, J.E.; Supko, J.G.; Asselin, B.L.; Athale, U.H.; Clavell, L.A.; et al. Intravenous pegylated asparaginase versus intramuscular native Escherichia coli l-asparaginase in newly diagnosed childhood acute lymphoblastic leukaemia (DFCI 05-001): A randomised, open-label phase 3 trial. Lancet Oncol. 2015, 16, 1677–1690. [Google Scholar] [CrossRef]
- Vora, A.; Goulden, N.; Wade, R.; Mitchell, C.; Hancock, J.; Hough, R.; Rowntree, C.; Richards, S. Treatment reduction for children and young adults with low-risk acute lymphoblastic leukaemia defined by minimal residual disease (UKALL 2003): A randomised controlled trial. Lancet Oncol. 2013, 14, 199–209. [Google Scholar] [CrossRef]
- Hayashi, R.J.; Winter, S.S.; Dunsmore, K.P.; Devidas, M.; Chen, Z.; Wood, B.L.; Hermiston, M.L.; Teachey, D.T.; Perkins, S.L.; Miles, R.R.; et al. Successful Outcomes of Newly Diagnosed T Lymphoblastic Lymphoma: Results from Children’s Oncology Group AALL0434. J. Clin. Oncol. 2020, 38, 3062–3070. [Google Scholar] [CrossRef]
- Coustan-Smith, E.; Mullighan, C.G.; Onciu, M.; Behm, F.G.; Raimondi, S.C.; Pei, D.; Cheng, C.; Su, X.; Rubnitz, J.E.; Basso, G.; et al. Early T-cell precursor leukaemia: A subtype of very high-risk acute lymphoblastic leukaemia. Lancet Oncol. 2009, 10, 147–156. [Google Scholar] [CrossRef]
- Inukai, T.; Kiyokawa, N.; Campana, D.; Coustan-Smith, E.; Kikuchi, A.; Kobayashi, M.; Takahashi, H.; Koh, K.; Manabe, A.; Kumagai, M.; et al. Clinical significance of early T-cell precursor acute lymphoblastic leukaemia: Results of the Tokyo Children’s Cancer Study Group Study L99-15. Br. J. Haematol. 2012, 156, 358–365. [Google Scholar] [CrossRef]
- Ma, M.; Wang, X.; Tang, J.; Xue, H.; Chen, J.; Pan, C.; Jiang, H.; Shen, S. Early T-cell precursor leukemia: A subtype of high risk childhood acute lymphoblastic leukemia. Front. Med. 2012, 6, 416–420. [Google Scholar] [CrossRef]
- Wood, B.L.; Winter, S.S.; Dunsmore, K.P.; Devidas, M.; Chen, S.; Asselin, B.; Esiashvili, N.; Loh, M.L.; Winick, N.J.; Carroll, W.L.; et al. T-Lymphoblastic Leukemia (T-ALL) Shows Excellent Outcome, Lack of Significance of the Early Thymic Precursor (ETP) Immunophenotype, and Validation of the Prognostic Value of End-Induction Minimal Residual Disease (MRD) in Children’s Oncology Group (COG) Study AALL0434. Blood 2014, 124, 1. [Google Scholar] [CrossRef]
- Knechtli, C.J.C.; Goulden, N.J.; Hancock, J.P.; Grandage, V.L.G.; Harris, E.L.; Garland, R.J.; Jones, C.G.; Rowbottom, A.W.; Hunt, L.P.; Green, A.F.; et al. Minimal Residual Disease Status Before Allogeneic Bone Marrow Transplantation Is an Important Determinant of Successful Outcome for Children and Adolescents with Acute Lymphoblastic Leukemia. Blood 1998, 92, 4072–4079. [Google Scholar] [CrossRef] [PubMed]
- Bader, P.; Kreyenberg, H.; Henze, G.H.; Eckert, C.; Reising, M.; Willasch, A.; Barth, A.; Borkhardt, A.; Peters, C.; Handgretinger, R.; et al. Prognostic value of minimal residual disease quantification before allogeneic stem-cell transplantation in relapsed childhood acute lymphoblastic leukemia: The ALL-REZ BFM Study Group. J. Clin. Oncol. 2009, 27, 377–384. [Google Scholar] [CrossRef] [PubMed]
- Leung, W.; Pui, C.-H.; Coustan-Smith, E.; Yang, J.; Pei, D.; Gan, K.; Srinivasan, A.; Hartford, C.; Triplett, B.M.; Dallas, M.; et al. Detectable minimal residual disease before hematopoietic cell transplantation is prognostic but does not preclude cure for children with very-high-risk leukemia. Blood 2012, 120, 468–472. [Google Scholar] [CrossRef] [PubMed]
- Umeda, K.; Hiramatsu, H.; Kawaguchi, K.; Iwai, A.; Mikami, M.; Nodomi, S.; Saida, S.; Heike, T.; Ohomori, K.; Adachi, S. Impact of pretransplant minimal residual disease on the post-transplant outcome of pediatric acute lymphoblastic leukemia. Pediatr. Transplant. 2016, 20, 692–696. [Google Scholar] [CrossRef]
- Pulsipher, M.A.; Carlson, C.; Langholz, B.; Wall, D.A.; Schultz, K.R.; Bunin, N.; Kirsch, I.; Gastier-Foster, J.M.; Borowitz, M.; Desmarais, C.; et al. IgH-V(D)J NGS-MRD measurement pre- and early post-allotransplant defines very low- and very high-risk ALL patients. Blood 2015, 125, 3501–3508. [Google Scholar] [CrossRef]
- Galimberti, S.; Devidas, M.; Lucenti, A.; Cazzaniga, G.; Möricke, A.; Bartram, C.R.; Mann, G.; Carroll, W.; Winick, N.; Borowitz, M.; et al. Validation of Minimal Residual Disease as Surrogate Endpoint for Event-Free Survival in Childhood Acute Lymphoblastic Leukemia. JNCI Cancer Spectr. 2018, 2, pky069. [Google Scholar] [CrossRef]
- Hilal, T.; Prasad, V. Eliminating MRD—FDA approval of blinatumomab for B-ALL in complete remission. Nat. Rev. Clin. Oncol. 2018, 15, 727–728. [Google Scholar] [CrossRef] [PubMed]
| Multiparametric Flow Cytometry | IgH/TCR RQ-PCR | Translocation RQ-PCR | Digital Droplet PCR | Next Generation Sequencing | |
|---|---|---|---|---|---|
| Requires patient specific design and optimization | Yes for LAIP No for newer approaches | Yes | No | Yes | No |
| Sensitivity | 4 color—10−4 8 color—10−5 8 colors with >4 × 106 cells—10−6 | 10−4–10−5 | 10−4–10−5 | 10−5 | 10−5–10−7 Depending on amount of DNA |
| Quantification | Absolute | Semi | Semi | Absolute | Absolute |
| Quantification based on diagnostic material | No | Yes | No | Yes | No |
| Applicability | >90% | 90–95% | 30–40% | 90–95% | >95% |
| Turnaround time | hours | weeks | days | hours | 1 week |
| Substrate -ease of use | Cell suspension -unstable | DNA -stable | RNA -unstable | DNA -stable | DNA -stable |
| Additional patient information | Identifies heterogeneity of whole population | None | None | None | -Identifies clonal repertoire -Identifies precise breakpoints |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Contreras Yametti, G.P.; Ostrow, T.H.; Jasinski, S.; Raetz, E.A.; Carroll, W.L.; Evensen, N.A. Minimal Residual Disease in Acute Lymphoblastic Leukemia: Current Practice and Future Directions. Cancers 2021, 13, 1847. https://doi.org/10.3390/cancers13081847
Contreras Yametti GP, Ostrow TH, Jasinski S, Raetz EA, Carroll WL, Evensen NA. Minimal Residual Disease in Acute Lymphoblastic Leukemia: Current Practice and Future Directions. Cancers. 2021; 13(8):1847. https://doi.org/10.3390/cancers13081847
Chicago/Turabian StyleContreras Yametti, Gloria Paz, Talia H. Ostrow, Sylwia Jasinski, Elizabeth A. Raetz, William L. Carroll, and Nikki A. Evensen. 2021. "Minimal Residual Disease in Acute Lymphoblastic Leukemia: Current Practice and Future Directions" Cancers 13, no. 8: 1847. https://doi.org/10.3390/cancers13081847
APA StyleContreras Yametti, G. P., Ostrow, T. H., Jasinski, S., Raetz, E. A., Carroll, W. L., & Evensen, N. A. (2021). Minimal Residual Disease in Acute Lymphoblastic Leukemia: Current Practice and Future Directions. Cancers, 13(8), 1847. https://doi.org/10.3390/cancers13081847






