PathoFusion: An Open-Source AI Framework for Recognition of Pathomorphological Features and Mapping of Immunohistochemical Data
Abstract
Simple Summary
Abstract
1. Introduction
2. Results
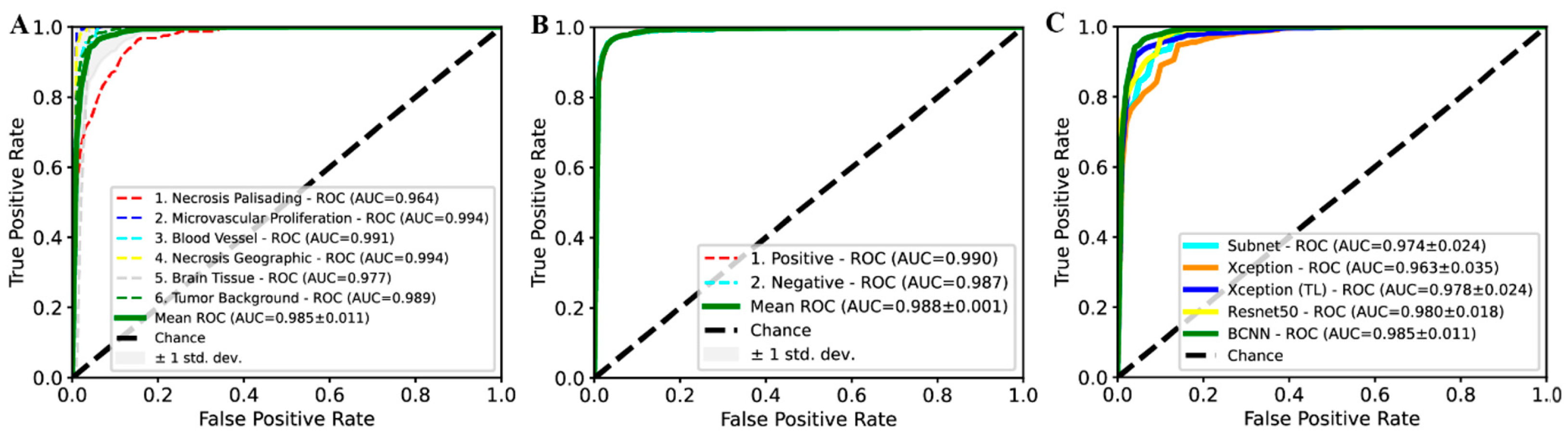
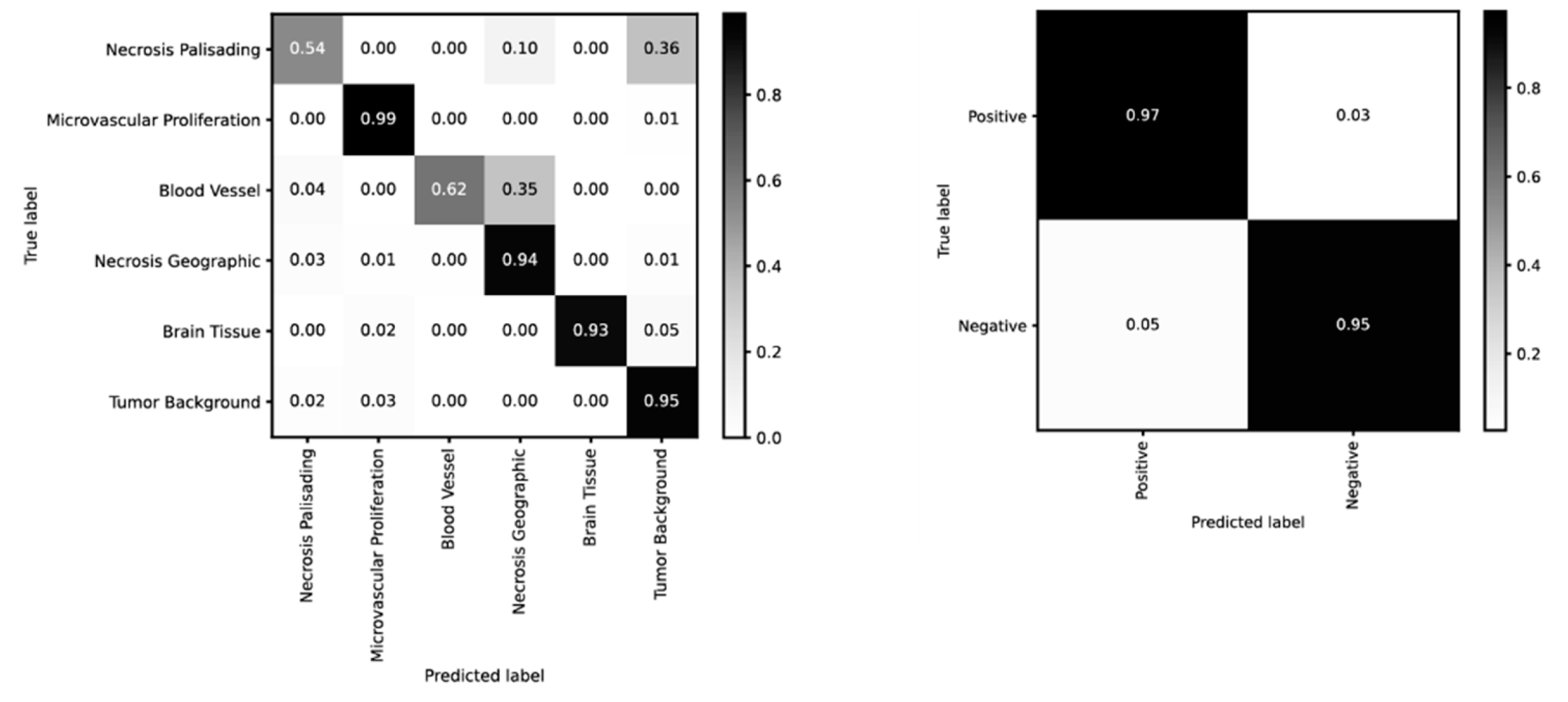
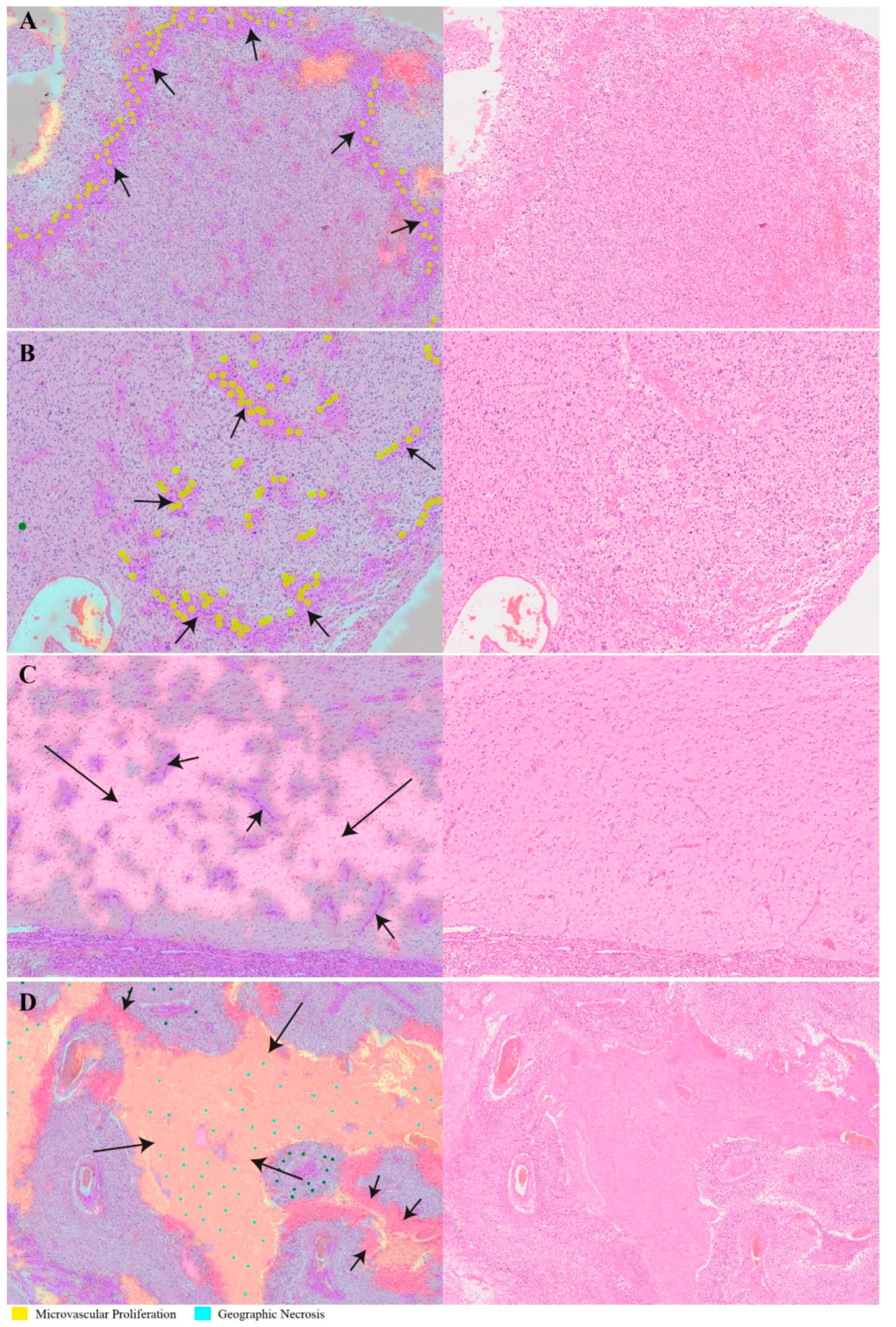
2.1. Recognition of Morphological Features and Associated Immunoreactivity
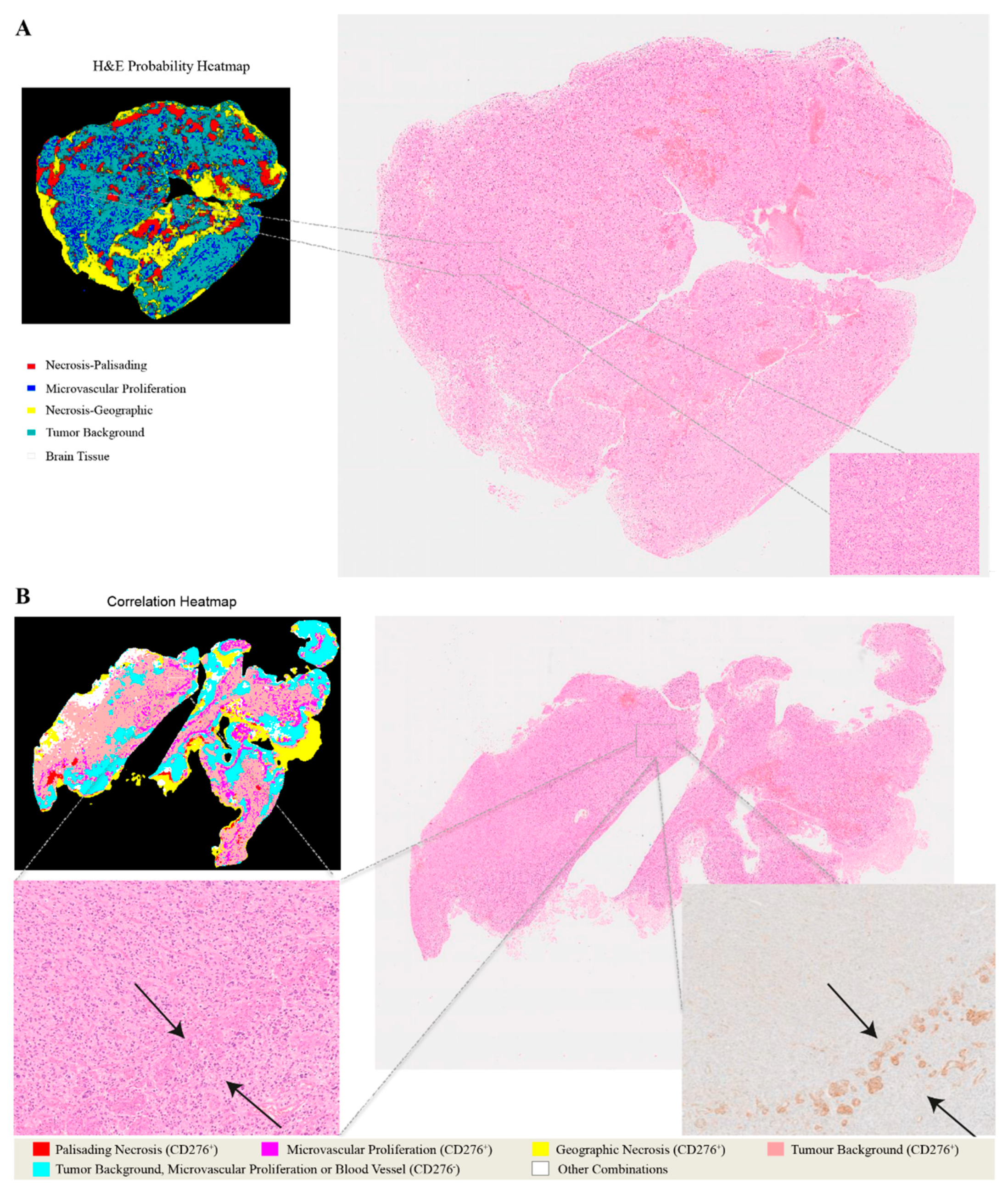
2.2. Analysis of Entire Histological Sections
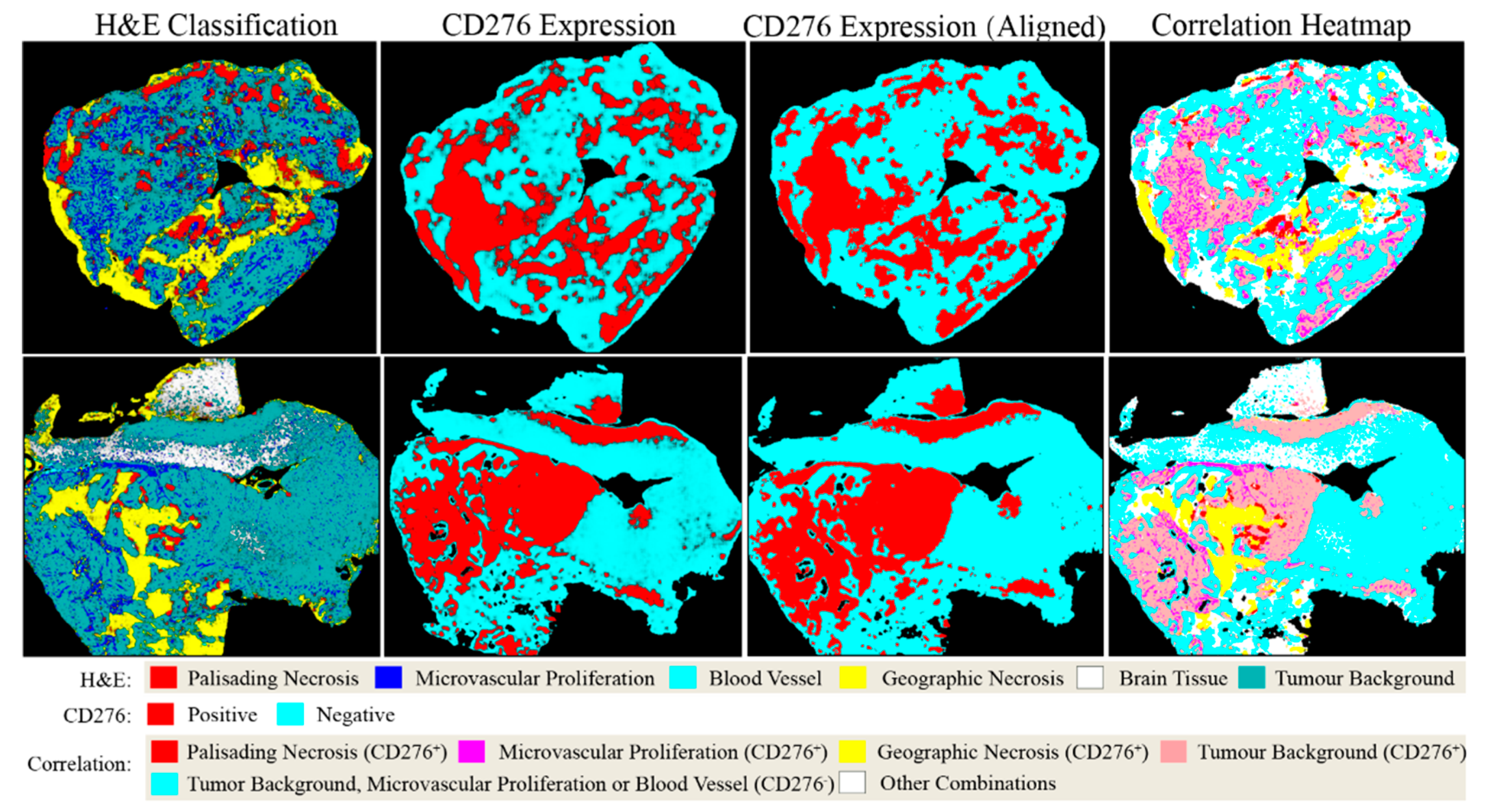
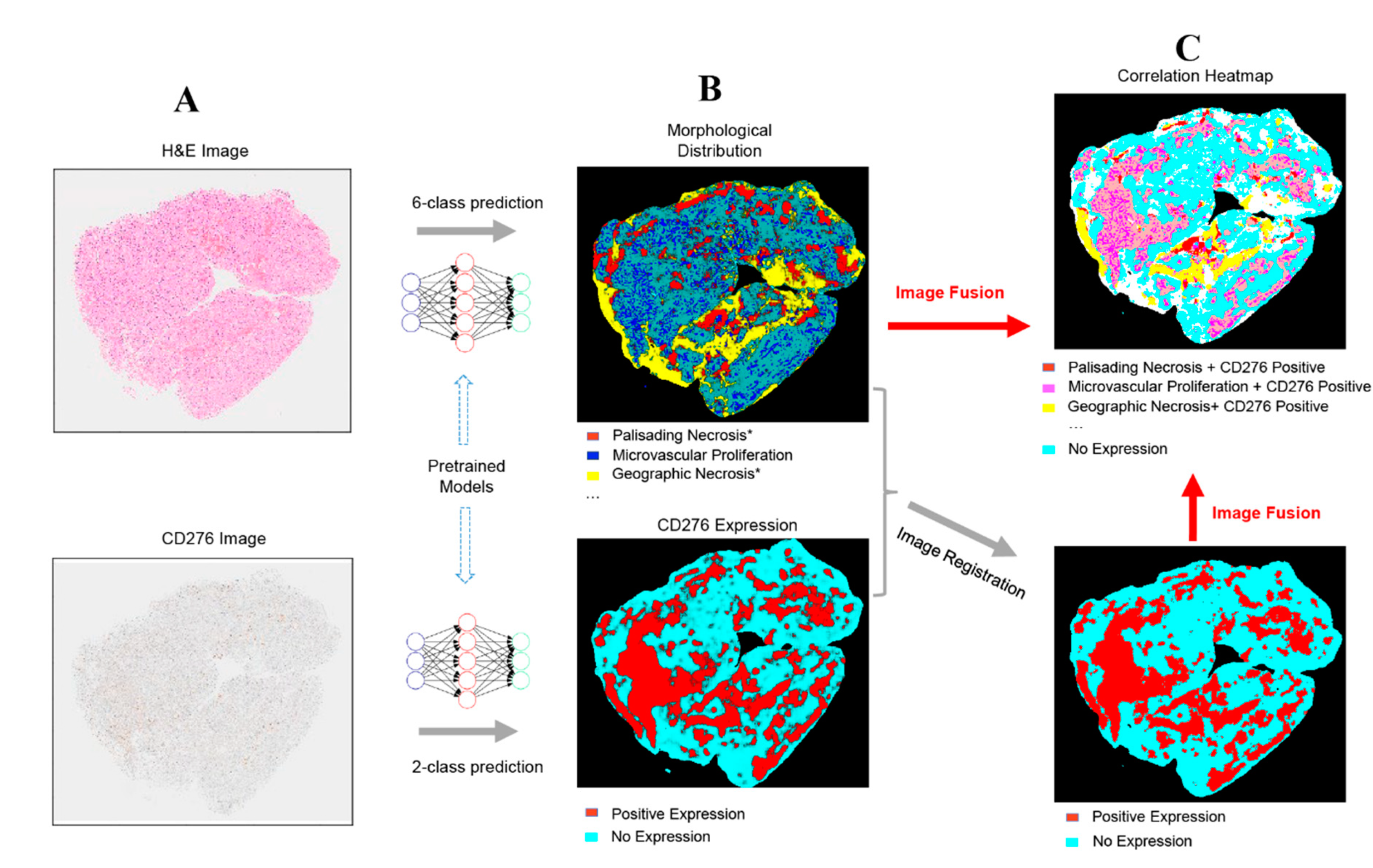
2.3. Fusion of Bimodal Neuropathological Images
3. Discussion
4. Materials and Methods
4.1. Clinical Cases
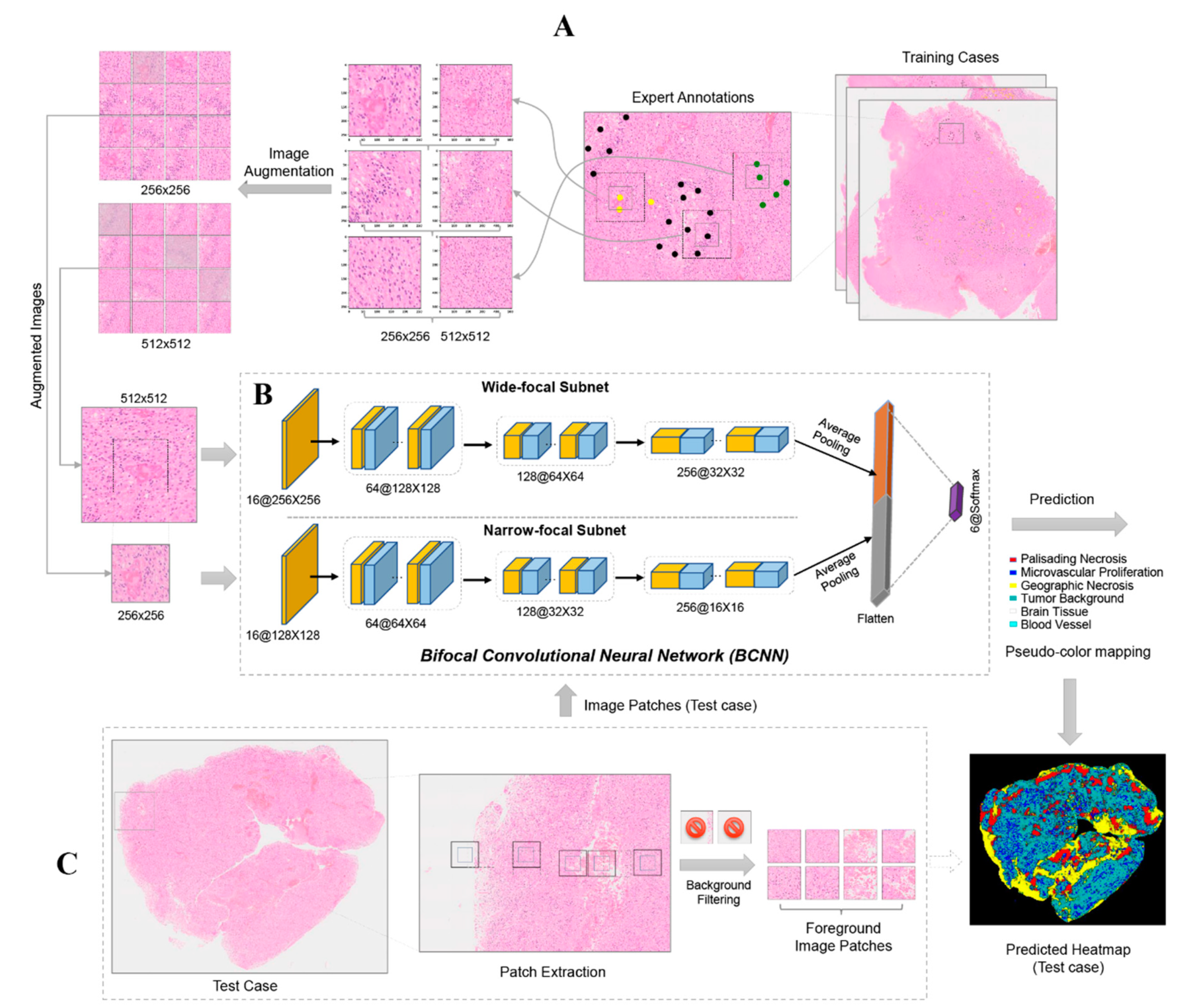
4.2. PathoFusion Framework
4.2.1. Expert Marking and Datasets
4.2.2. Bifocal Convolutional Neural Network (BCNN)
4.2.3. Recognition of Morphological Features and Associated Immunoreactivity
4.2.4. Method for Fusing Bimodal Histological Images
4.2.5. Quantitative Analysis
4.2.6. Data Availability and Experimental Reproducibility
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- LeCun, Y.; Bengio, Y. Convolutional networks for images, speech, and time series. In The Handbook of Brain Theory and Neural Networks; The MIT Press: Cambridge, MA, USA, 1995; Volume 3361, p. 1995. [Google Scholar]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. Imagenet classification with deep convolutional neural networks. In Advances in Neural Information Processing Systems; NIPS: Lake Tahoe, Nevada, USA, 2012; pp. 1097–1105. [Google Scholar]
- Niazi, M.; Parwani, A.; Gurcan, M. Digital pathology and artificial intelligence. Lancet Oncol. 2019, 20, e253–e261. [Google Scholar] [CrossRef]
- Signaevsky, M.; Prastawa, M.; Farrell, K.; Tabish, N.; Baldwin, E.; Han, N.; Iida, M.A.; Koll, J.; Bryce, C.; Purohit, D.; et al. Artificial intelligence in neuropathology: Deep learning-based assessment of tauopathy. Lab. Invest. 2019, 99, 1019–1029. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Wang, J.; Marzese, D.M.; Wang, X.; Yang, Z.; Li, C.; Zhang, H.; Zhang, J.; Chen, C.C.; Kelly, D.F.; et al. B7H3 regulates differentiation and serves as a potential biomarker and theranostic target for human glioblastoma. Lab. Invest. 2019, 99, 1117–1129. [Google Scholar] [CrossRef] [PubMed]
- Papanicolau-Sengos, A.; Yang, Y.; Pabla, S.; Lenzo, F.L.; Kato, S.; Kurzrock, R.; DePietro, P.; Nesline, M.; Conroy, J.; Glenn, S.; et al. Identification of targets for prostate cancer immunotherapy. Prostate 2019, 79, 498–505. [Google Scholar] [CrossRef] [PubMed]
- Majzner, R.G.; Theruvath, J.L.; Nellan, A.; Heitzeneder, S.; Cui, Y.; Mount, C.W.; Rietberg, S.P.; Linde, M.H.; Xu, P.; Rota, C.; et al. CAR T Cells Targeting B7-H3, a Pan-Cancer Antigen, Demonstrate Potent Preclinical Activity Against Pediatric Solid Tumors and Brain Tumors. Clin. Cancer Res. 2019, 25, 2560–2574. [Google Scholar] [CrossRef] [PubMed]
- Dong, P.; Xiong, Y.; Yue, J.; Hanley, S.J.B.; Watari, H. B7H3 As a Promoter of Metastasis and Promising Therapeutic Target. Front. Oncol. 2018, 8, 264. [Google Scholar] [CrossRef] [PubMed]
- Rogiers, A.; Boekhout, A.; Schwarze, J.K.; Awada, G.; Blank, C.U.; Neyns, B. Long-Term Survival, Quality of Life, and Psychosocial Outcomes in Advanced Melanoma Patients Treated with Immune Checkpoint Inhibitors. J. Oncol. 2019, 2019, 5269062. [Google Scholar] [CrossRef] [PubMed]
- Louis, D.N.; Ohgaki, H.; Wiestler, O.D.; Cavenee, W.K. World Health Organization Histological Classification of Tumours of the Central Nervous System; International Agency for Research on Cancer: Lyon, France, 2016. [Google Scholar]
- Rojianl, A.; Dorovini-Zis, K. Microvascular proliferation in glioblastoma multiforme. J. Neuropathol. Exp. Neurol. 1990, 49, 300. [Google Scholar] [CrossRef]
- Wesseling, P.; Schlingemann, R.O.; Rietveld, F.J.; Link, M.; Burger, P.C.; Ruiter, J.D. Early and extensive contribution of pericytes/vascular smooth muscle cells to microvascular proliferation in glioblastoma multiforme: An immuno-light and immuno-electron microscopic study. J. Neuropathol. Exp. Neurol. 1995, 54, 304–310. [Google Scholar] [CrossRef] [PubMed]
- Brat, D.J.; van Meir, E.G. Glomeruloid microvascular proliferation orchestrated by VPF/VEGF: A new world of angiogenesis research. Am. J. Pathol. 2001, 158, 789–796. [Google Scholar] [CrossRef]
- Takashima, Y.; Kawaguchi, A.; Hayano, A.; Yamanaka, R. CD276 and the gene signature composed of GATA3 and LGALS3 enable prognosis prediction of glioblastoma multiforme. PLoS ONE 2019, 14, e0216825. [Google Scholar] [CrossRef] [PubMed]
- Inamura, K.; Yokouchi, Y.; Kobayashi, M.; Sakakibara, R.; Ninomiya, H.; Subat, S.; Nagano, H.; Nomura, K.; Okumura, S.; Shibutani, T.; et al. Tumor B7-H3 (CD276) expression and smoking history in relation to lung adenocarcinoma prognosis. Lung Cancer 2017, 103, 44–51. [Google Scholar] [CrossRef] [PubMed]
- Benzon, B.; Zhao, S.G.; Haffner, M.C.; Takhar, M.; Erho, N.; Yousefi, K.; Hurley, P.; Bishop, J.L.; Tosoian, J.; Ghabili, K.; et al. Correlation of B7-H3 with androgen receptor, immune pathways and poor outcome in prostate cancer: An expression-based analysis. Prostate Cancer Prostatic Dis. 2017, 20, 28–35. [Google Scholar] [CrossRef]
- Lemke, D.; Pfenning, P.-N.; Sahm, F.; Klein, A.-C.; Kempf, T.; Warnken, U.; Schnölzer, M.; Tudoran, R.; Weller, M.; Platten, M.; et al. Costimulatory protein 4IgB7H3 drives the malignant phenotype of glioblastoma by mediating immune escape and invasiveness. Clin. Cancer Res. 2012, 18, 105–117. [Google Scholar] [CrossRef]
- Kraan, J.; Broek, P.V.D.; Verhoef, C.; Grunhagen, D.J.; Taal, W.; Gratama, J.W.; Sleijfer, S. Endothelial CD276 (B7-H3) expression is increased in human malignancies and distinguishes between normal and tumour-derived circulating endothelial cells. Br. J. Cancer 2014, 111, 149–156. [Google Scholar] [CrossRef]
- Gootjes, E.C.; Kraan, J.; Buffart, T.; Verhoef, C.; Verheul, H.M.; Sleijfer, S. ORCHESTRA Study Group CD276-positive circulating endothelial cells in advanced colorectal cancer. J. Clin. Oncol. 2019, 37, 572. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Chollet, F. Xception: Deep learning with depthwise separable convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 1251–1258. [Google Scholar]
- Mezirow, J. Transformative Dimensions of Adult Learning. In Proceedings of the ERIC, Alexandria, VA, USA, 30 October–1 November 1991. [Google Scholar]
- Raina, R.; Battle, A.; Lee, H.; Packer, B.; Ng, A.Y. Self-taught learning: Transfer learning from unlabeled data. In Proceedings of the 24th International Conference on Machine Learning, Corvalis, OR, USA, 20–24 June 2007; ACM: New York, NY, USA, 2007; pp. 759–766. [Google Scholar]
- Bengio, Y. Deep learning of representations for unsupervised and transfer learning. In Proceedings of the ICML Workshop on Unsupervised and Transfer Learning, Edinburgh, UK, 1–26 July 2012; pp. 17–36. [Google Scholar]
- Deng, J.; Dong, W.; Socher, R.; Li, L.-J.; Li, K.; Fei-Fei, L. Imagenet: A large-scale hierarchical image database. In Proceedings of the 2009 IEEE Conference on Computer Vision and Pattern Recognition, Miami, FL, USA, 20–25 June 2009; pp. 248–255. [Google Scholar]
- Bao, G.; Graeber, M.B.; Wang, X. A Bifocal Classification and Fusion Network for Multimodal Image Analysis in Histopathology. In Proceedings of the 16th International Conference on Control, Automation, Robotics and Vision (ICARCV 2020), Shenzhen, China, 13–15 December 2020. [Google Scholar]
- Shan, H.; Zhang, Y.; Yang, Q.; Kruger, U.; Kalra, M.K.; Sun, L.; Cong, W.; Wang, G. 3-D Convolutional Encoder-Decoder Network for Low-Dose CT via Transfer Learning From a 2-D Trained Network. IEEE Trans. Med Imaging 2018, 37, 1522–1534. [Google Scholar] [CrossRef] [PubMed]
- Samala, R.K.; Chan, H.; Hadjiiski, L.; Helvie, M.A.; Richter, C.D.; Cha, K.H. Breast Cancer Diagnosis in Digital Breast Tomosynthesis: Effects of Training Sample Size on Multi-Stage Transfer Learning Using Deep Neural Nets. IEEE Trans. Med Imaging 2019, 38, 686–696. [Google Scholar] [CrossRef] [PubMed]
- Weiss, K.; Khoshgoftaar, T.M.; Wang, D. A survey of transfer learning. J. Big Data 2016, 3, 9. [Google Scholar] [CrossRef]
- Kornblith, S.; Shlens, J.; Le, Q.V. Do better imagenet models transfer better? In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Long Beach, CA, USA, 15–20 June 2019; pp. 2661–2671. [Google Scholar]
- Raghu, M.; Zhang, C.; Kleinberg, J.; Bengio, S. Transfusion: Understanding transfer learning for medical imaging. In Advances in Neural Information Processing Systems; Vancouver Convention Center: Vancouver, BC, Canada, 2019; pp. 3342–3352. [Google Scholar]
- Dubrofsky, E. Homography Estimation. Master’s Thesis, Univerzita Britské Kolumbie, Vancouver, BC, Canada, 2009. [Google Scholar]
- Janowczyk, A.; Basavanhally, A.; Madabhushi, A. Stain normalization using sparse autoencoders (StaNoSA): Application to digital pathology. Comput. Med Imaging Graph. 2017, 57, 50–61. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bao, G.; Wang, X.; Xu, R.; Loh, C.; Adeyinka, O.D.; Pieris, D.A.; Cherepanoff, S.; Gracie, G.; Lee, M.; McDonald, K.L.; et al. PathoFusion: An Open-Source AI Framework for Recognition of Pathomorphological Features and Mapping of Immunohistochemical Data. Cancers 2021, 13, 617. https://doi.org/10.3390/cancers13040617
Bao G, Wang X, Xu R, Loh C, Adeyinka OD, Pieris DA, Cherepanoff S, Gracie G, Lee M, McDonald KL, et al. PathoFusion: An Open-Source AI Framework for Recognition of Pathomorphological Features and Mapping of Immunohistochemical Data. Cancers. 2021; 13(4):617. https://doi.org/10.3390/cancers13040617
Chicago/Turabian StyleBao, Guoqing, Xiuying Wang, Ran Xu, Christina Loh, Oreoluwa Daniel Adeyinka, Dula Asheka Pieris, Svetlana Cherepanoff, Gary Gracie, Maggie Lee, Kerrie L. McDonald, and et al. 2021. "PathoFusion: An Open-Source AI Framework for Recognition of Pathomorphological Features and Mapping of Immunohistochemical Data" Cancers 13, no. 4: 617. https://doi.org/10.3390/cancers13040617
APA StyleBao, G., Wang, X., Xu, R., Loh, C., Adeyinka, O. D., Pieris, D. A., Cherepanoff, S., Gracie, G., Lee, M., McDonald, K. L., Nowak, A. K., Banati, R., Buckland, M. E., & Graeber, M. B. (2021). PathoFusion: An Open-Source AI Framework for Recognition of Pathomorphological Features and Mapping of Immunohistochemical Data. Cancers, 13(4), 617. https://doi.org/10.3390/cancers13040617






