Measurement of the Drug Sensitivity of Single Prostate Cancer Cells
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Reagents and Antibodies
2.2. Preparation of the Capture Membranes
2.3. Cell Lines
2.4. Preparation of the Microwell Array
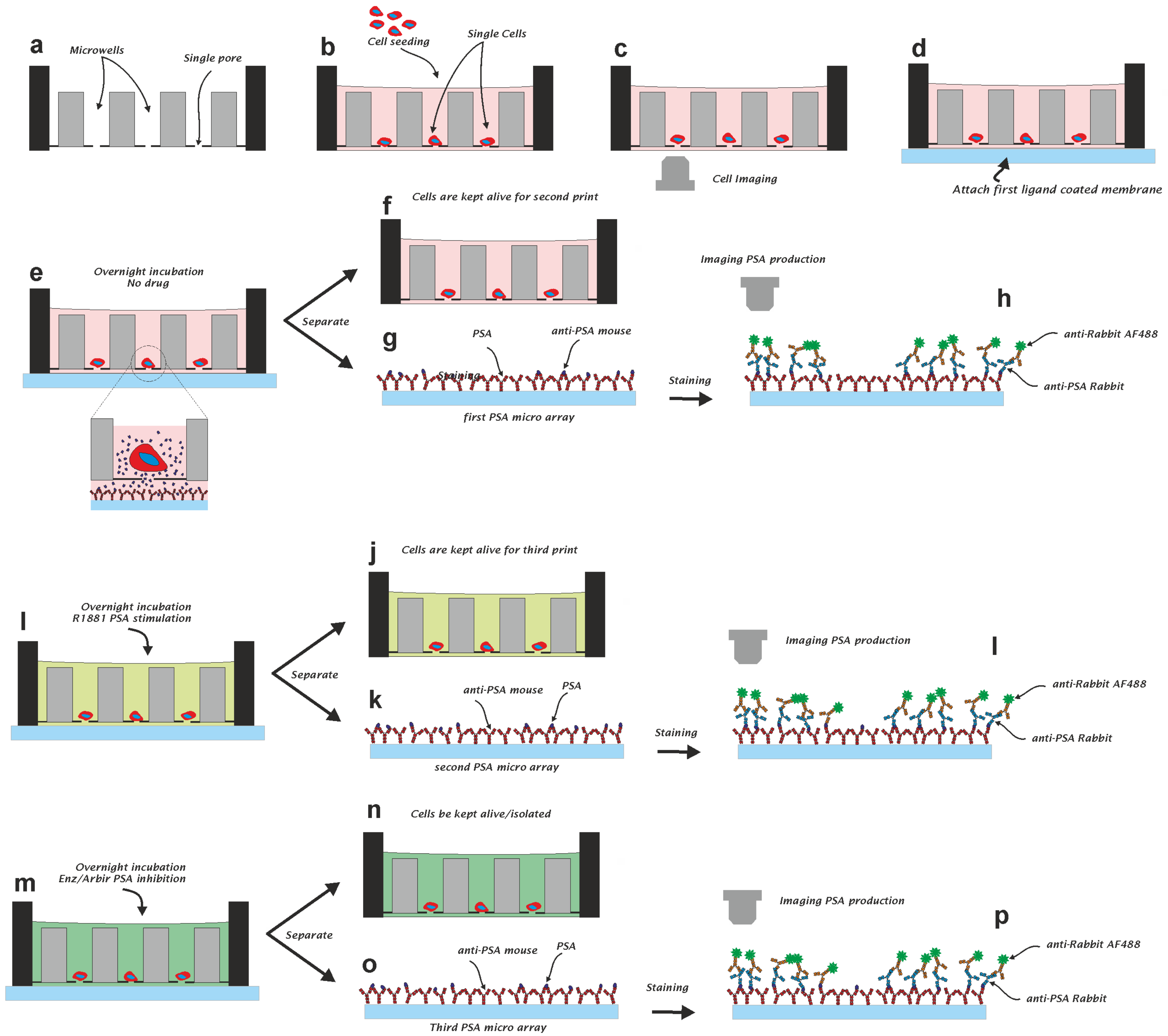
2.5. Capturing PSA from Single Prostate Cancer Cells
2.6. Detection of Printed Antibody Arrays
2.7. Calibration Curves
2.8. Image Acquisition
2.9. Image Analysis and Quantification of Fluorescent Signals
3. Results
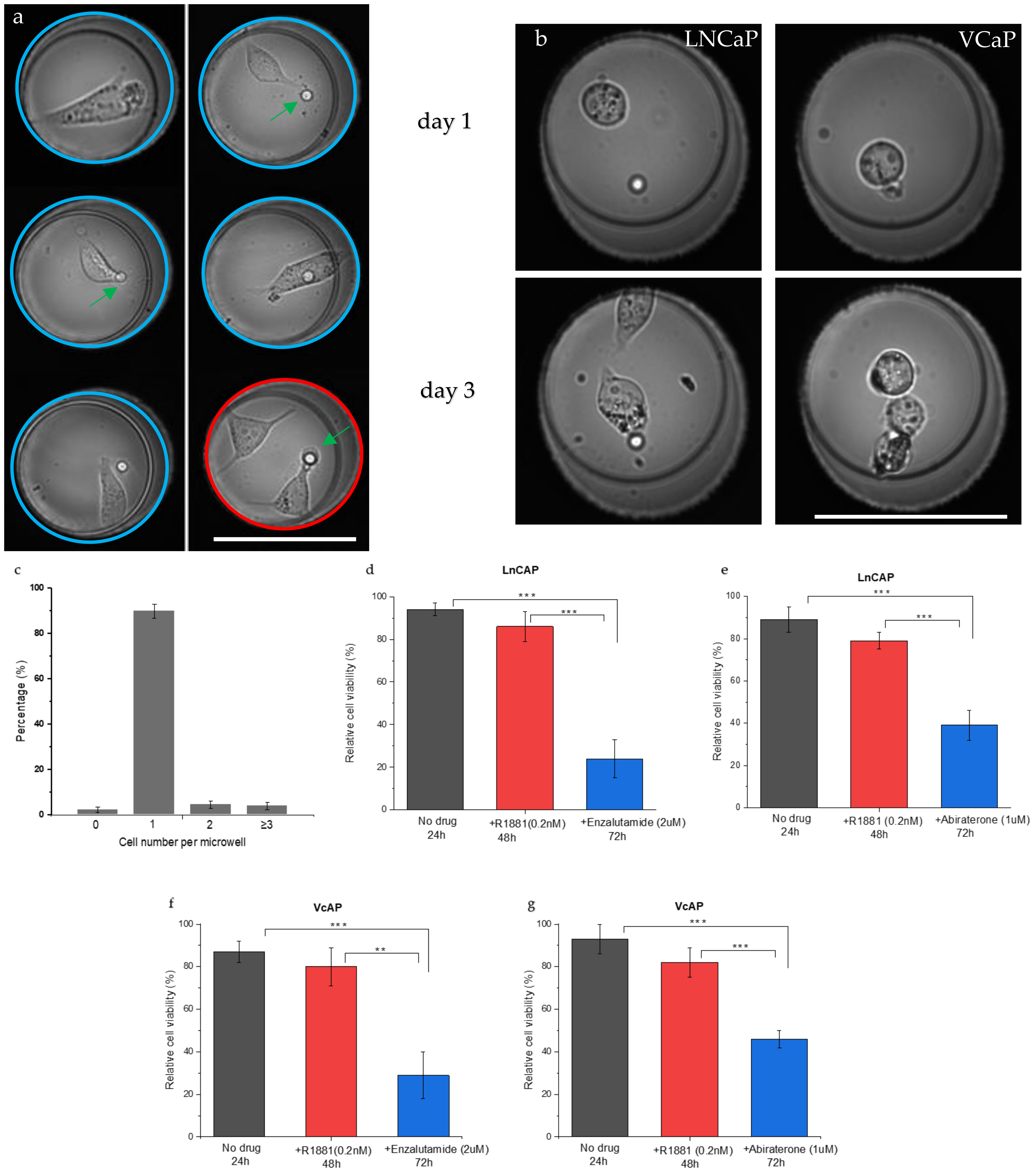
3.1. Single Cell Seeding and Viability
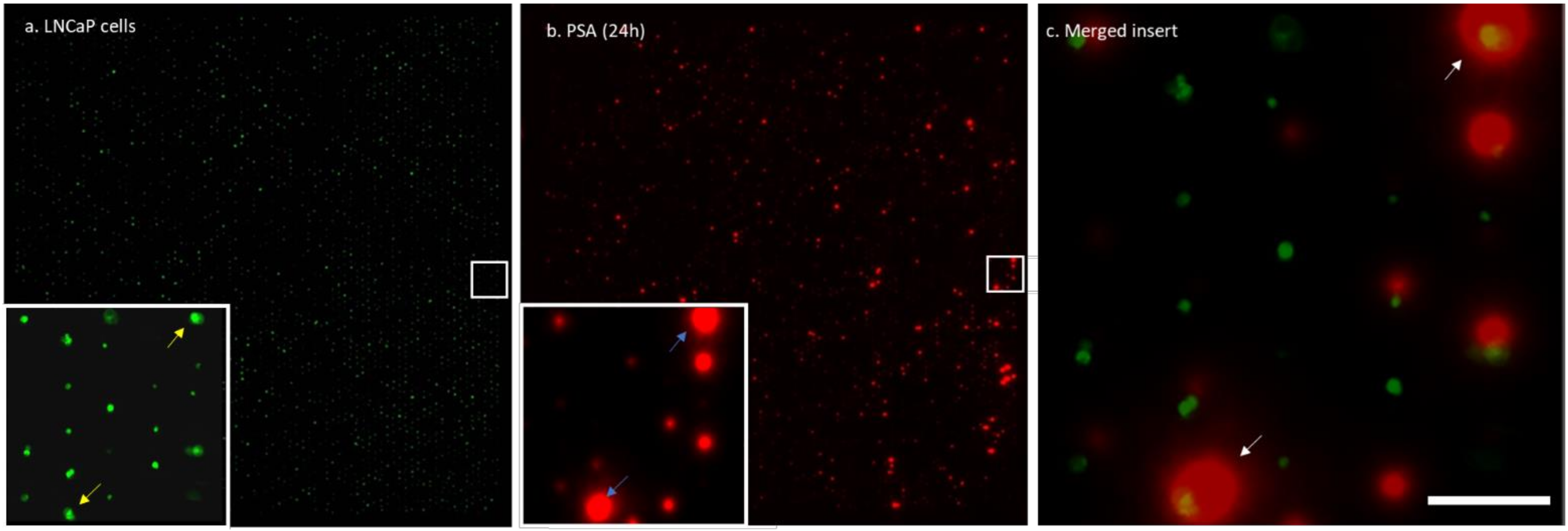
3.2. Single Cell PSA Secretion Assay
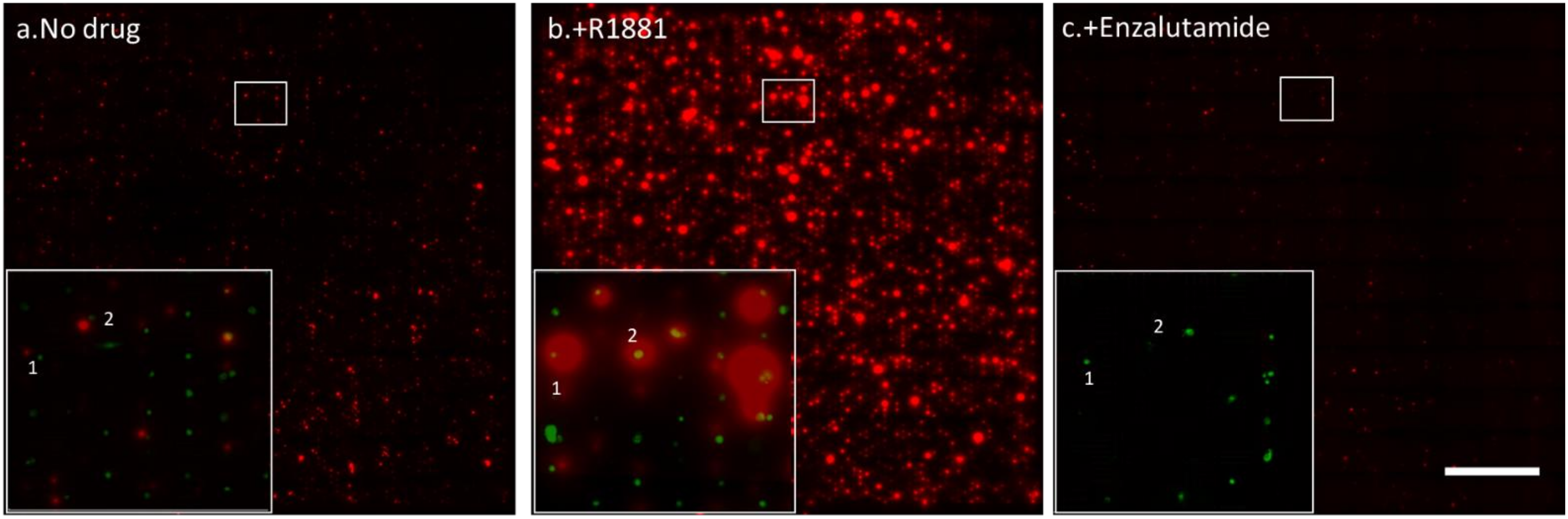
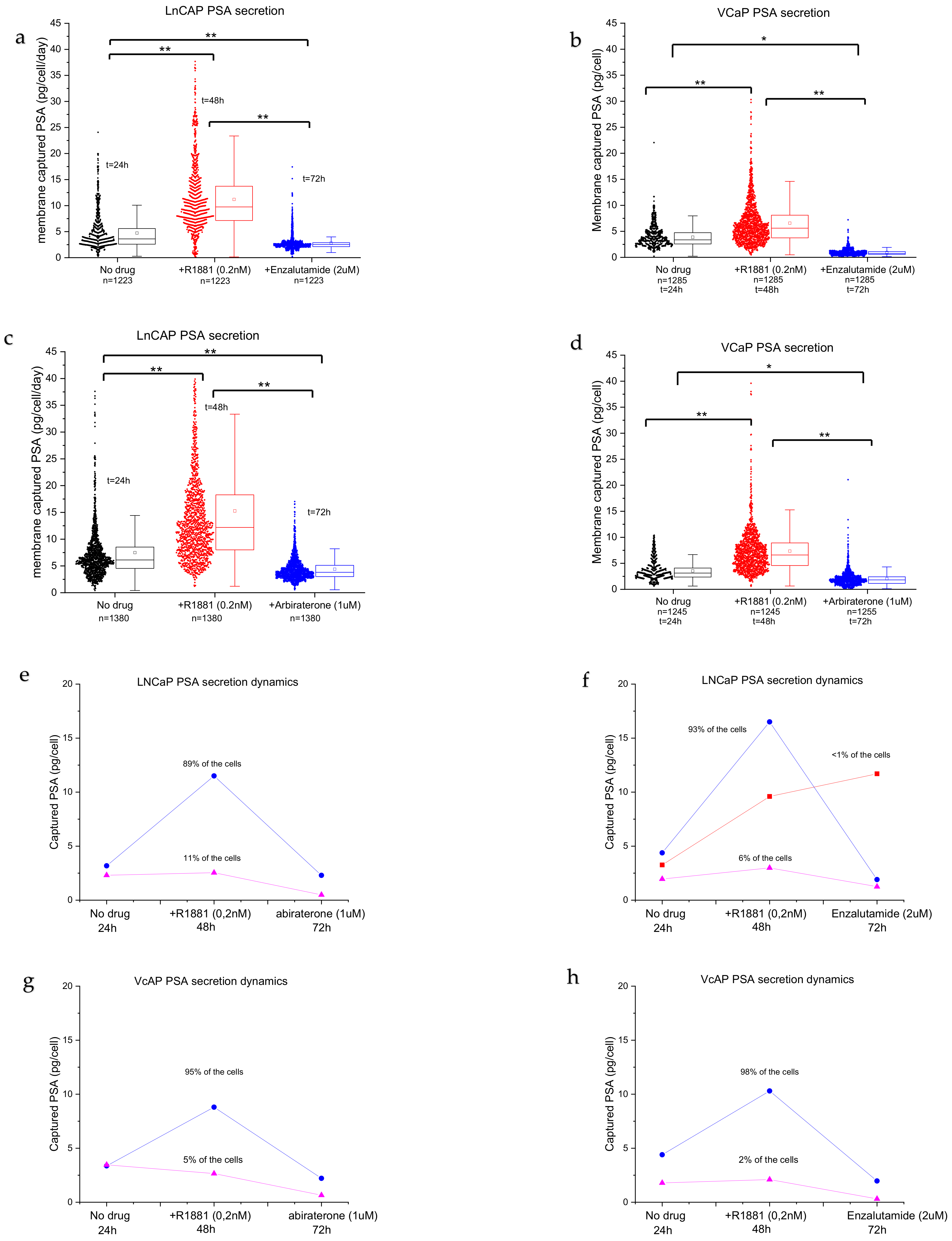
3.3. Drug Sensitivity of PC Cells
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tang, D.G. Understanding cancer stem cell heterogeneity and plasticity. Cell Res. 2012, 22, 457–472. [Google Scholar] [CrossRef] [PubMed]
- Chang, H.H.; Hemberg, M.; Barahona, M.; Ingber, D.E.; Huang, S. Transcriptome-wide noise controls lineage choice in mammalian progenitor cells. Nature 2008, 453, 544–547. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cohen, A.A.; Geva-Zatorsky, N.; Eden, E.; Frenkel-Morgenstern, M.; Issaeva, I.; Sigal, A.; Milo, R.; Cohen-Saidon, C.; Liron, Y.; Kam, Z.; et al. Dynamic proteomics of individual cancer cells in response to a drug. Science 2008, 322, 1511–1516. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Armbrecht, L.; Müller, R.S.; Nikoloff, J.; Dittrich, P.S. Single-cell protein profiling in microchambers with barcoded beads. Microsyst. Nanoeng. 2019, 5, 55. [Google Scholar] [CrossRef] [PubMed]
- Riley, R.S.; June, C.H.; Langer, R.; Mitchell, M.J. Delivery technologies for cancer immunotherapy. Nat. Rev. Drug Discov. 2019, 18, 175–196. [Google Scholar] [CrossRef] [PubMed]
- Scheetz, L.; Park, K.S.; Li, Q.; Lowenstein, P.R.; Castro, M.G.; Schwendeman, A.; Moon, J.J. Engineering patient-specific cancer immunotherapies. Nat. Biomed. Eng. 2019, 3, 768–782. [Google Scholar] [CrossRef] [PubMed]
- Lopez, J.S.; Banerji, U. Combine and conquer: Challenges for targeted therapy combinations in early phase trials. Nat. Rev. Clin. Oncol. 2016, 14, 57–66. [Google Scholar] [CrossRef] [PubMed]
- Van Dalum, G.; Holland, L.; Terstappen, L.W. Metastasis and circulating tumor cells. EJIFCC 2012, 23, 87–97. [Google Scholar] [PubMed]
- Allard, W.J.; Matera, J.; Miller, M.C.; Repollet, M.; Connelly, M.C.; Rao, C.; Tibbe, A.G.J.; Uhr, J.W.; Terstappen, L.W.M.M. Tumor cells circulate in the peripheral blood of all major carcinomas but not in healthy subjects or patients with nonmalignant diseases. Clin. Cancer Res. 2004, 10, 6897–6904. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fischer, J.; Niederacher, D.; Topp, S.A.; Honisch, E.; Schumacher, S.; Schmitz, N.; Fohrding, L.Z.; Vay, C.; Hoffmann, I.; Kasprowicz, N.S.; et al. Diagnostic leukapheresis enables reliable detection of circulating tumor cells of nonmetastatic cancer patients. Proc. Natl. Acad. Sci. USA 2013, 110, 16580–16585. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andree, K.C.; Mentink, A.; Zeune, L.L.; Terstappen, L.W.; Stoecklein, N.H.; Neves, R.P.; Driemel, C.; Lampignano, R.; Yang, L.; Neubauer, H.; et al. Toward a real liquid biopsy in metastatic breast and prostate cancer: Diagnostic LeukApheresis increases CTC yields in a European prospective multicenter study (CTCTrap). Int. J. Cancer 2018, 143, 2584–2591. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stoecklein, N.H.; Fischer, J.C.; Niederacher, D.; Terstappen, L.W.M.M. Challenges for CTC-based liquid biopsies: Low CTC frequency and diagnostic leukapheresis as a potential solution. Expert Rev. Mol. Diagn. 2015, 16, 147–164. [Google Scholar] [CrossRef] [PubMed]
- Swennenhuis, J.F.; Tibbe, A.G.J.; Stevens, M.; Katika, M.R.; van Dalum, J.; Tong, H.D.; van Rijn, C.J.M.; Terstappen, L.W.M.M. Self-seeding microwell chip for the isolation and characterization of single cells. Lab Chip 2015, 15, 3039–3046. [Google Scholar] [CrossRef]
- Abali, F.; Broekmaat, J.; Tibbe, A.; Schasfoort, R.B.M.; Zeune, L.; Terstappen, L.W.M.M. A microwell array platform to print and measure biomolecules produced by single cells. Lab Chip 2019, 19, 1850–1859. [Google Scholar] [CrossRef] [PubMed]
- Abali, F.; Stevens, M.; Tibbe, A.; Terstappen, L.; van der Velde, P.; Schasfoort, R. Isolation of single cells for protein therapeutics using microwell selection and Surface Plasmon Resonance imaging. Anal. Biochem. 2017, 531, 45–47. [Google Scholar] [CrossRef] [PubMed]
- Andree, K.C.; Abali, F.; Oomens, L.; Passanha, F.R.; Broekmaat, J.J.; Kraan, J.; Mendelaar, P.A.; Sleijfer, S.; Terstappen, L.W. Self-seeding microwells to isolate and assess the viability of single circulating tumor cells. Int. J. Mol. Sci. 2019, 20, 477. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nettikadan, S.; Radke, K.; Johnson, J.; Xu, J.; Lynch, M.; Mosher, C.; Henderson, E. Detection and quantification of protein biomarkers from fewer than 10 cells. Mol. Cell. Proteom. 2006, 5, 895–901. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alix-Panabières, C.; Rebillard, X.; Brouillet, J.-P.; Barbotte, E.; Iborra, F.; Segui, B.; Maudelonde, T.; Jolivet-Reynaud, C.; Vendrell, J.-P. Detection of circulating prostate-specific antigen–secreting cells in prostate cancer patients. Clin. Chem. 2005, 51, 1538–1541. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Rho, H.S.; Stevens, M.; Tibbe, A.G.J.; Gardeniers, H.; Terstappen, L.W.M.M. Microfluidic device for DNA amplification of single cancer cells isolated from whole blood by self-seeding microwells. Lab Chip 2015, 15, 4331–4337. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abali, F.; Baghi, N.; Mout, L.; Broekmaat, J.J.; Tibbe, A.G.J.; Terstappen, L.W.M.M. Measurement of the Drug Sensitivity of Single Prostate Cancer Cells. Cancers 2021, 13, 6083. https://doi.org/10.3390/cancers13236083
Abali F, Baghi N, Mout L, Broekmaat JJ, Tibbe AGJ, Terstappen LWMM. Measurement of the Drug Sensitivity of Single Prostate Cancer Cells. Cancers. 2021; 13(23):6083. https://doi.org/10.3390/cancers13236083
Chicago/Turabian StyleAbali, Fikri, Narges Baghi, Lisanne Mout, Joska J. Broekmaat, Arjan G. J. Tibbe, and Leon W. M. M. Terstappen. 2021. "Measurement of the Drug Sensitivity of Single Prostate Cancer Cells" Cancers 13, no. 23: 6083. https://doi.org/10.3390/cancers13236083
APA StyleAbali, F., Baghi, N., Mout, L., Broekmaat, J. J., Tibbe, A. G. J., & Terstappen, L. W. M. M. (2021). Measurement of the Drug Sensitivity of Single Prostate Cancer Cells. Cancers, 13(23), 6083. https://doi.org/10.3390/cancers13236083






