Identification of Key Genes Associated with Progression and Prognosis of Bladder Cancer through Integrated Bioinformatics Analysis
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Data Set
2.3. Data Processing
2.4. Functional and Pathway Enrichment Analysis
2.5. Quantitative RT-PCR
2.6. Identification of Prognostic Genes
2.7. Statistical Analysis
3. Results
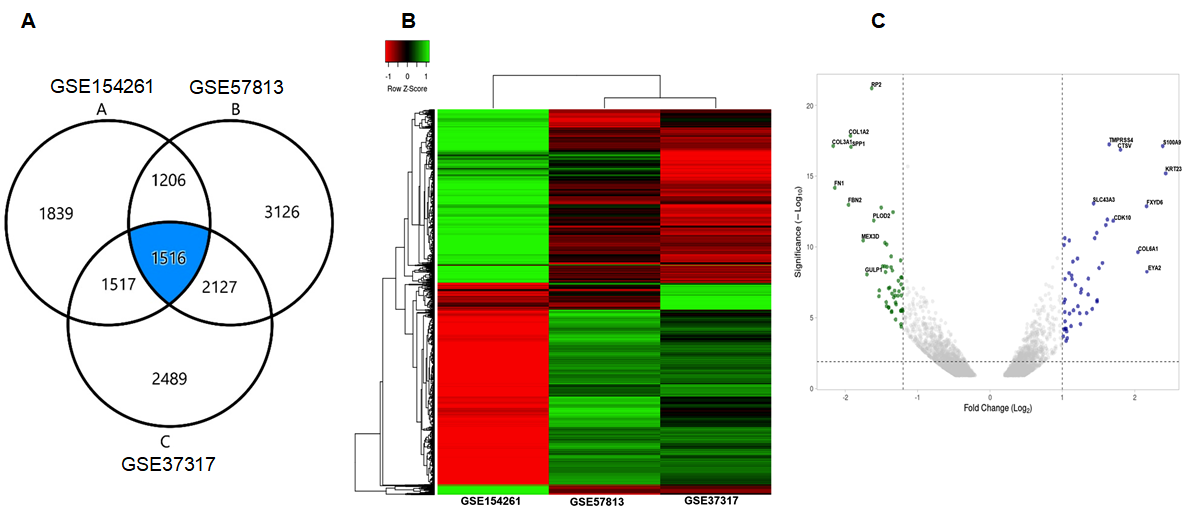
3.1. Identification of DEGs in NMIBC versus MIBC
3.2. GO and KEGG Pathway Analysis
3.3. Protein-Protein Interaction and Identification of Genes of Prognostic Significance
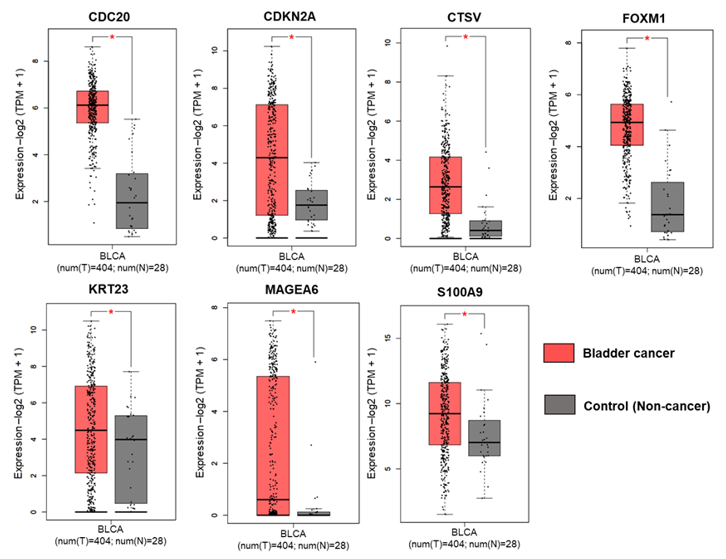
3.4. Differential Expression of Selected Genes in Bladder Cancer and Normal Bladder Patients
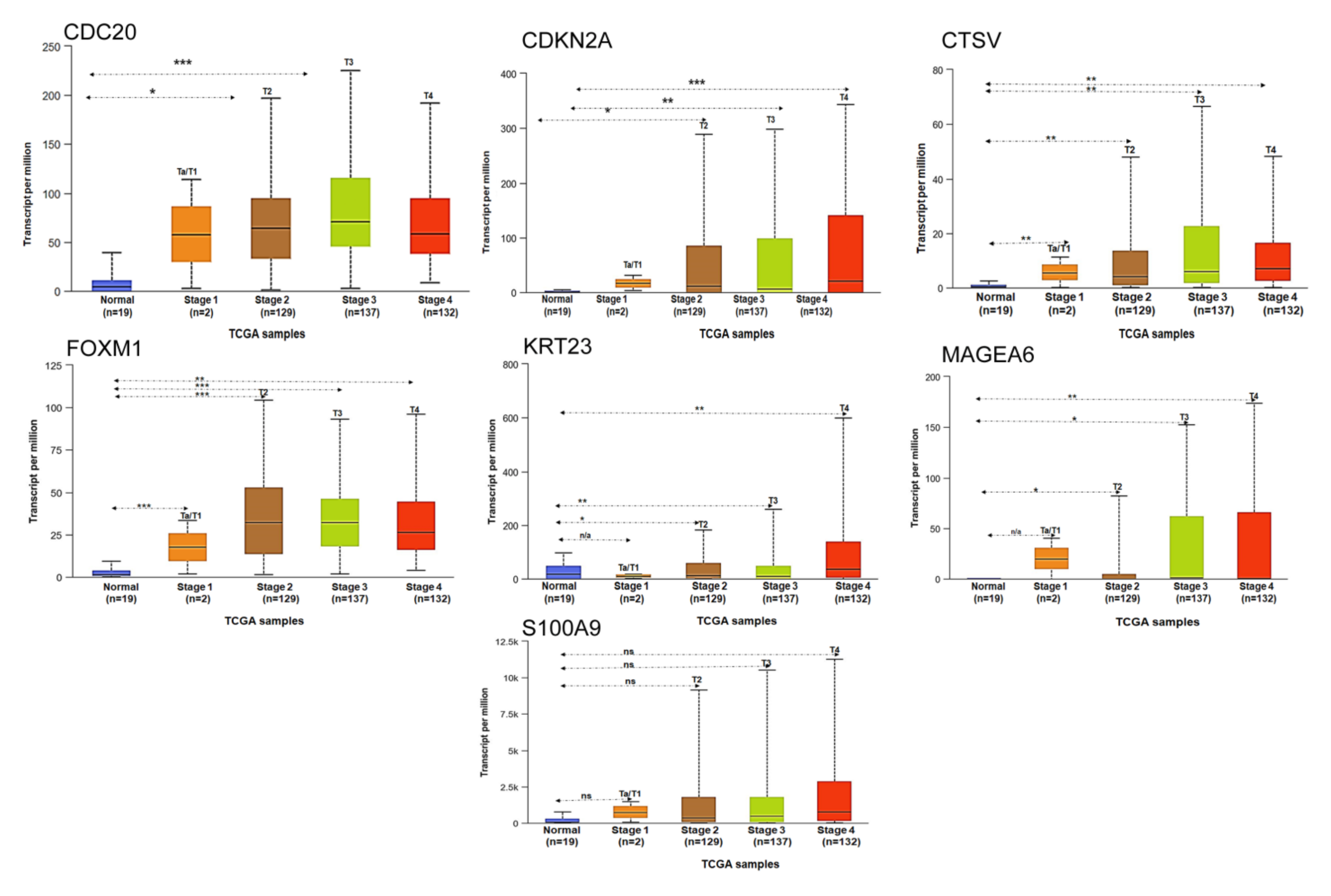
3.5. Stage-Specific Expression of Selected Genes in Bladder Cancer Progression
3.6. Survival Analysis of Selected Genes in Bladder Cancer
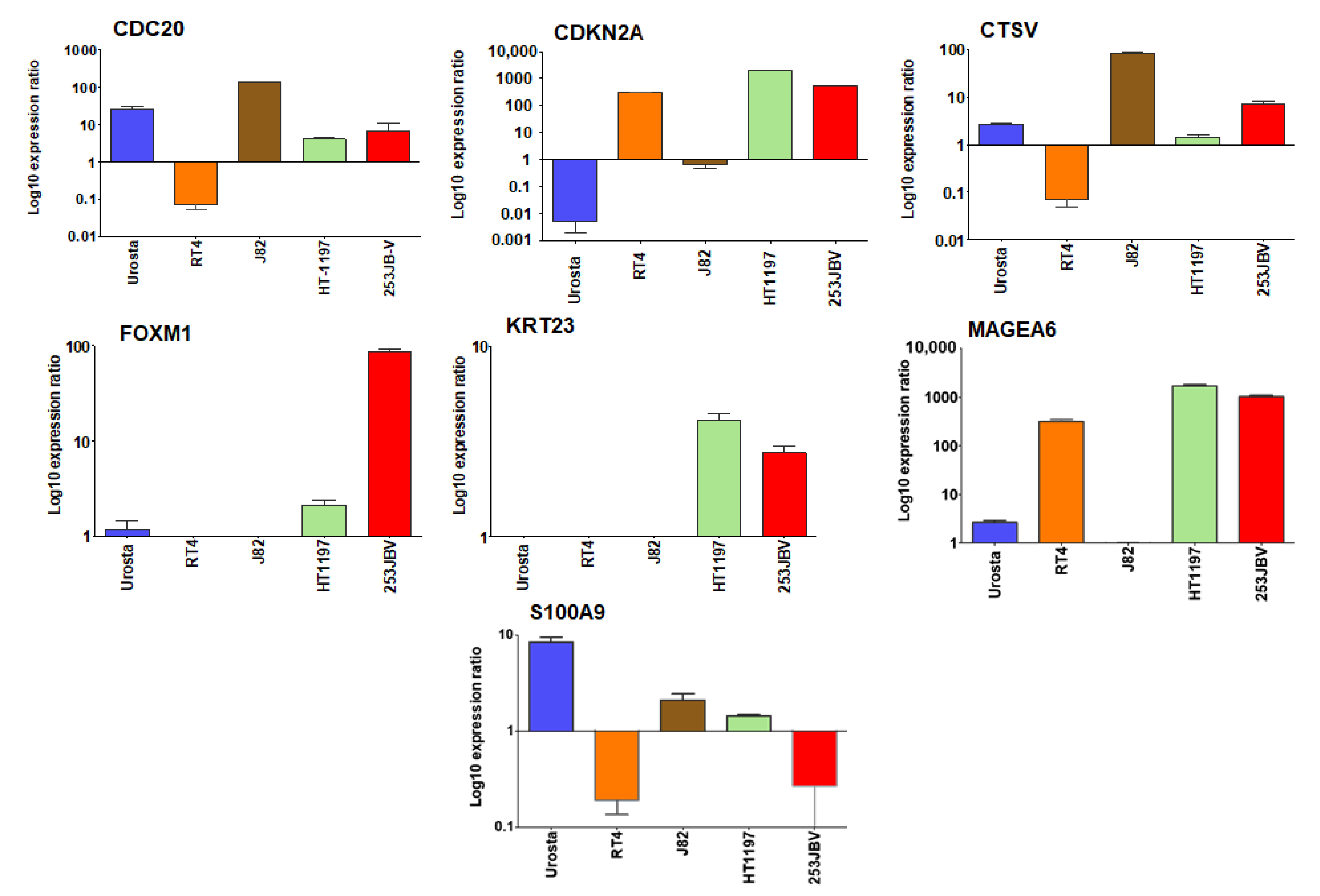
3.7. Differential mRNA Expression of Selected Genes in NMIBC and MIBC Cell Lines
3.8. Prognostic Role of Selected Genes in Bladder Cancer
3.9. Construction of Multiplex Model for Prognosis of Bladder Cancer
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kaufman, D.S.; Shipley, W.U.; Feldman, A.S. Bladder cancer. Lancet 2009, 374, 239–249. [Google Scholar] [CrossRef]
- Matulewicz, R.S.; Steinberg, G.D. Non-muscle-invasive Bladder Cancer: Overview and Contemporary Treatment Landscape of Neoadjuvant Chemoablative Therapies. Rev. Urol. 2020, 22, 43–51. [Google Scholar]
- Pasin, E.; Josephson, D.Y.; Mitra, A.P.; Cote, R.J.; Stein, J.P. Superficial bladder cancer: An update on etiology, molecular development, classification, and natural history. Rev. Urol. 2008, 10, 31–43. [Google Scholar]
- Balar, A.V.; Kamat, A.M.; Kulkarni, G.S.; Uchio, E.M.; Boormans, J.L.; Bajorin, D.F.; Roumiguié, M.; Singer, E.A.; Krieger, L.E.M.; Grivas, P.; et al. Pembrolizumab (pembro) for the treatment of patients with Bacillus Calmette-Guérin (BCG) unresponsive, high-risk (HR) non–muscle-invasive bladder cancer (NMIBC): Over two years follow-up of KEYNOTE-057. J. Clin. Oncol. 2020, 38, 5041. [Google Scholar] [CrossRef]
- Aldousari, S.; Kassouf, W. Update on the management of non-muscle invasive bladder cancer. Can. Urol. Assoc. J. 2010, 4, 56–64. [Google Scholar] [CrossRef]
- Fujii, Y. Prediction models for progression of non-muscle-invasive bladder cancer: A review. Int. J. Urol. 2018, 25, 212–218. [Google Scholar] [CrossRef]
- Macaluso, M.; Paggi, M.G.; Giordano, A. Genetic and epigenetic alterations as hallmarks of the intricate road to cancer. Oncogene 2003, 22, 6472–6478. [Google Scholar] [CrossRef]
- Oszczudlowski, M.; Dobruch, J. Prediction of progression to muscle-invasive disease in patients with high-risk bladder cancer. Transl. Androl. Urol. 2018, 7, 749–751. [Google Scholar] [CrossRef] [PubMed]
- Robertson, A.G.; Kim, J.; Al-Ahmadie, H.; Bellmunt, J.; Guo, G.; Cherniack, A.D.; Hinoue, T.; Laird, P.W.; Hoadley, K.A.; Akbani, R.; et al. Comprehensive Molecular Characterization of Muscle-Invasive Bladder Cancer. Cell 2017, 171, 540–556.e25. [Google Scholar] [CrossRef] [PubMed]
- Carrasco, R.; Izquierdo, L.; van der Heijden, A.G.; Lozano, J.J.; Franco, M.; Ingelmo-Torres, M.; Roldan, F.L.; Llorens, M.; Ribal, M.J.; Mengual, L.; et al. Differential gene expression profile between progressive and de novo muscle invasive bladder cancer and its prognostic implication. Sci. Rep. 2021, 11, 6132. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Zhou, B.; Pache, L.; Chang, M.; Khodabakhshi, A.H.; Tanaseichuk, O.; Benner, C.; Chanda, S.K. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 2019, 10, 1523. [Google Scholar] [CrossRef]
- Tang, Z.; Kang, B.; Li, C.; Chen, T.; Zhang, Z. GEPIA2: An enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 2019, 47, W556–W560. [Google Scholar] [CrossRef] [PubMed]
- Meeks, J.J.; Al-Ahmadie, H.; Faltas, B.M.; Taylor, J.A., 3rd; Flaig, T.W.; DeGraff, D.J.; Christensen, E.; Woolbright, B.L.; McConkey, D.J.; Dyrskjøt, L. Genomic heterogeneity in bladder cancer: Challenges and possible solutions to improve outcomes. Nat. Rev. Urol. 2020, 17, 259–270. [Google Scholar] [CrossRef] [PubMed]
- Doherty, S.C.; McKeown, S.R.; McKelvey-Martin, V.; Downes, C.S.; Atala, A.; Yoo, J.J.; Simpson, D.A.; Kaufmann, W.K. Cell Cycle Checkpoint Function in Bladder Cancer. JNCI J. Natl. Cancer Inst. 2003, 95, 1859–1868. [Google Scholar] [CrossRef]
- Xu, Y.; Wu, G.; Li, J.; Li, J.; Ruan, N.; Ma, L.; Han, X.; Wei, Y.; Li, L.; Zhang, H.; et al. Screening and Identification of Key Biomarkers for Bladder Cancer: A Study Based on TCGA and GEO Data. BioMed Res. Int. 2020, 2020, 8283401. [Google Scholar] [CrossRef] [PubMed]
- Sansregret, L.; Patterson, J.O.; Dewhurst, S.; López-García, C.; Koch, A.; McGranahan, N.; Chao, W.C.H.; Barry, D.J.; Rowan, A.; Instrell, R.; et al. APC/C Dysfunction Limits Excessive Cancer Chromosomal Instability. Cancer Discov. 2017, 7, 218. [Google Scholar] [CrossRef]
- Fraschini, R.; Beretta, A.; Sironi, L.; Musacchio, A.; Lucchini, G.; Piatti, S. Bub3 interaction with Mad2, Mad3 and Cdc20 is mediated by WD40 repeats and does not require intact kinetochores. EMBO J. 2001, 20, 6648–6659. [Google Scholar] [CrossRef] [PubMed]
- Vleugel, M.; Hoek, T.A.; Tromer, E.; Sliedrecht, T.; Groenewold, V.; Omerzu, M.; Kops, G.J. Dissecting the roles of human BUB1 in the spindle assembly checkpoint. J. Cell Sci. 2015, 128, 2975–2982. [Google Scholar] [CrossRef]
- Raaijmakers, J.A.; van Heesbeen, R.; Blomen, V.A.; Janssen, L.M.E.; van Diemen, F.; Brummelkamp, T.R.; Medema, R.H. BUB1 Is Essential for the Viability of Human Cells in which the Spindle Assembly Checkpoint Is Compromised. Cell Rep. 2018, 22, 1424–1438. [Google Scholar] [CrossRef]
- Tsang, Y.H.; Wang, Y.; Kong, K.; Grzeskowiak, C.; Zagorodna, O.; Dogruluk, T.; Lu, H.; Villafane, N.; Bhavana, V.H.; Moreno, D.; et al. Differential expression of MAGEA6 toggles autophagy to promote pancreatic cancer progression. eLife 2020, 9, e48963. [Google Scholar] [CrossRef]
- Endo, M.; Kanda, M.; Sawaki, K.; Shimizu, D.; Tanaka, C.; Kobayashi, D.; Hattori, N.; Hayashi, M.; Yamada, S.; Koike, M.; et al. Tissue Expression of Melanoma-associated Antigen A6 and Clinical Characteristics of Gastric Cancer. Anticancer Res. 2019, 39, 5903–5910. [Google Scholar] [CrossRef] [PubMed]
- Kidokoro, T.; Tanikawa, C.; Furukawa, Y.; Katagiri, T.; Nakamura, Y.; Matsuda, K. CDC20, a potential cancer therapeutic target, is negatively regulated by p53. Oncogene 2008, 27, 1562–1571. [Google Scholar] [CrossRef] [PubMed]
- Rinaldetti, S.; Wirtz, R.; Worst, T.S.; Hartmann, A.; Breyer, J.; Dyrskjot, L.; Erben, P. FOXM1 predicts disease progression in non-muscle invasive bladder cancer. J. Cancer Res. Clin. Oncol. 2018, 144, 1701–1709. [Google Scholar] [CrossRef]
- Yao, Z.; Zhang, J.; Chen, C.; Li, P.; Cao, J.; Han, D.; Li, J.; Ying, L.; Tian, J. High Expression of CTSV in Bladder Cancer as a Predictor of Poor Prognosis: A Study Based on TCGA and GEO Database. Res. Sq. 2021, 1, 1–17. [Google Scholar] [CrossRef]
- Marangos, P.; Carroll, J. Securin regulates entry into M-phase by modulating the stability of cyclin B. Nat. Cell Biol. 2008, 10, 445–451. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Ramsay, E.S.; Mock, B.A. Cdkn2a, the cyclin-dependent kinase inhibitor encoding p16INK4a and p19ARF, is a candidate for the plasmacytoma susceptibility locus, Pctr1. Proc. Natl. Acad. Sci. USA 1998, 95, 2429–2434. [Google Scholar] [CrossRef] [PubMed]
- Witkiewicz, A.K.; Knudsen, K.E.; Dicker, A.P.; Knudsen, E.S. The meaning of p16(ink4a) expression in tumors: Functional significance, clinical associations and future developments. Cell Cycle 2011, 10, 2497–2503. [Google Scholar] [CrossRef]
- Kollmann, K.; Heller, G.; Schneckenleithner, C.; Warsch, W.; Scheicher, R.; Ott, R.G.; Schäfer, M.; Fajmann, S.; Schlederer, M.; Schiefer, A.-I.; et al. A Kinase-Independent Function of CDK6 Links the Cell Cycle to Tumor Angiogenesis. Cancer Cell 2013, 24, 167–181. [Google Scholar] [CrossRef]
- Sherr, C.J.; Beach, D.; Shapiro, G.I. Targeting CDK4 and CDK6: From Discovery to Therapy. Cancer Discov. 2016, 6, 353–367. [Google Scholar] [CrossRef]
- Topacio, B.R.; Zatulovskiy, E.; Cristea, S.; Xie, S.; Tambo, C.S.; Rubin, S.M.; Sage, J.; Kõivomägi, M.; Skotheim, J.M. Cyclin D-Cdk4,6 Drives Cell-Cycle Progression via the Retinoblastoma Protein’s C-Terminal Helix. Mol. Cell 2019, 74, 758–770.e4. [Google Scholar] [CrossRef]
- Narasimha, A.M.; Kaulich, M.; Shapiro, G.S.; Choi, Y.J.; Sicinski, P.; Dowdy, S.F. Cyclin D activates the Rb tumor suppressor by mono-phosphorylation. eLife 2014, 3, e02872. [Google Scholar] [CrossRef]
- Weintraub, S.J.; Chow, K.N.B.; Luo, R.X.; Zhang, S.H.; He, S.; Dean, D.C. Mechanism of active transcriptional repression by the retinoblastoma protein. Nature 1995, 375, 812–816. [Google Scholar] [CrossRef]
- Chen, H.Z.; Tsai, S.Y.; Leone, G. Emerging roles of E2Fs in cancer: An exit from cell cycle control. Nat. Rev. Cancer 2009, 9, 785–797. [Google Scholar] [CrossRef] [PubMed]
- Møller, M.B.; Ino, Y.; Gerdes, A.M.; Skjødt, K.; Louis, D.N.; Pedersen, N.T. Aberrations of the p53 pathway components p53, MDM2 and CDKN2A appear independent in diffuse large B cell lymphoma. Leukemia 1999, 13, 453–459. [Google Scholar] [CrossRef][Green Version]
- Stott, F.J.; Bates, S.; James, M.C.; McConnell, B.B.; Starborg, M.; Brookes, S.; Palmero, I.; Ryan, K.; Hara, E.; Vousden, K.H.; et al. The alternative product from the human CDKN2A locus, p14(ARF), participates in a regulatory feedback loop with p53 and MDM2. EMBO J. 1998, 17, 5001–5014. [Google Scholar] [CrossRef] [PubMed]
- Knudsen, E.S.; Nambiar, R.; Rosario, S.R.; Smiraglia, D.J.; Goodrich, D.W.; Witkiewicz, A.K. Pan-cancer molecular analysis of the RB tumor suppressor pathway. Commun. Biol. 2020, 3, 158. [Google Scholar] [CrossRef] [PubMed]
- Italiano, A.; Bianchini, L.; Gjernes, E.; Keslair, F.; Ranchere-Vince, D.; Dumollard, J.M.; Haudebourg, J.; Leroux, A.; Mainguené, C.; Terrier, P.; et al. Clinical and biological significance of CDK4 amplification in well-differentiated and dedifferentiated liposarcomas. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2009, 15, 5696–5703. [Google Scholar] [CrossRef] [PubMed]
- Worst, T.S.; Weis, C.-A.; Stöhr, R.; Bertz, S.; Eckstein, M.; Otto, W.; Breyer, J.; Hartmann, A.; Bolenz, C.; Wirtz, R.M.; et al. CDKN2A as transcriptomic marker for muscle-invasive bladder cancer risk stratification and therapy decision-making. Sci. Rep. 2018, 8, 14383. [Google Scholar] [CrossRef] [PubMed]
- Sudhan, D.R.; Siemann, D.W. Cathepsin L inhibition by the small molecule KGP94 suppresses tumor microenvironment enhanced metastasis associated cell functions of prostate and breast cancer cells. Clin. Exp. Metastasis 2013, 30, 891–902. [Google Scholar] [CrossRef]
- Zhang, W.; Wang, S.; Wang, Q.; Yang, Z.; Pan, Z.; Li, L. Overexpression of cysteine cathepsin L is a marker of invasion and metastasis in ovarian cancer. Oncol. Rep. 2014, 31, 1334–1342. [Google Scholar] [CrossRef]
- Zhao, H.; Huang, C.; Luo, Y.; Yao, X.; Hu, Y.; Wang, M.; Chen, X.; Zeng, J.; Hu, W.; Wang, J.; et al. A Correlation Study of Prognostic Risk Prediction for Colorectal Cancer Based on Autophagy Signature Genes. Front. Oncol. 2021, 11, 595099. [Google Scholar] [CrossRef] [PubMed]
- Svatek, R.S.; Karam, J.; Karakiewicz, P.I.; Gallina, A.; Casella, R.; Roehrborn, C.G.; Shariat, S.F. Role of urinary cathepsin B and L in the detection of bladder urothelial cell carcinoma. J. Urol. 2008, 179, 478–484. [Google Scholar] [CrossRef] [PubMed]
- Bach, D.-H.; Long, N.P.; Luu, T.-T.-T.; Anh, N.H.; Kwon, S.W.; Lee, S.K. The Dominant Role of Forkhead Box Proteins in Cancer. Int. J. Mol. Sci. 2018, 19, 3279. [Google Scholar] [CrossRef] [PubMed]
- Barger, C.J.; Branick, C.; Chee, L.; Karpf, A.R. Pan-Cancer Analyses Reveal Genomic Features of FOXM1 Overexpression in Cancer. Cancers 2019, 11, 251. [Google Scholar] [CrossRef]
- Kim, S.-K.; Roh, Y.-G.; Park, K.; Kang, T.-H.; Kim, W.-J.; Lee, J.-S.; Leem, S.-H.; Chu, I.-S. Expression Signature Defined by FOXM1–CCNB1 Activation Predicts Disease Recurrence in Non–Muscle-Invasive Bladder Cancer. Clin. Cancer Res. 2014, 20, 3233–3243. [Google Scholar] [CrossRef]
- Breyer, J.; Wirtz, R.M.; Erben, P.; Rinaldetti, S.; Worst, T.S.; Stoehr, R.; Eckstein, M.; Sikic, D.; Denzinger, S.; Burger, M.; et al. FOXM1 overexpression is associated with adverse outcome and predicts response to intravesical instillation therapy in stage pT1 non-muscle-invasive bladder cancer. BJU Int. 2019, 123, 187–196. [Google Scholar] [CrossRef]
- Zhang, J.S.; Wang, L.; Huang, H.; Nelson, M.; Smith, D.I. Keratin 23 (K23), a novel acidic keratin, is highly induced by histone deacetylase inhibitors during differentiation of pancreatic cancer cells. Genes Chromosomes Cancer 2001, 30, 123–135. [Google Scholar] [CrossRef]
- Zhang, N.; Zhang, R.; Zou, K.; Yu, W.; Guo, W.; Gao, Y.; Li, J.; Li, M.; Tai, Y.; Huang, W.; et al. Keratin 23 promotes telomerase reverse transcriptase expression and human colorectal cancer growth. Cell Death Dis. 2017, 8, e2961. [Google Scholar] [CrossRef]
- Ren, M.; Gao, Y.; Chen, Q.; Zhao, H.; Zhao, X.; Yue, W. The Overexpression of Keratin 23 Promotes Migration of Ovarian Cancer via Epithelial-Mesenchymal Transition. BioMed Res. Int. 2020, 2020, 8218735. [Google Scholar] [CrossRef]
- Weon, J.L.; Potts, P.R. The MAGE protein family and cancer. Curr. Opin. Cell Biol. 2015, 37, 1–8. [Google Scholar] [CrossRef]
- Pineda, C.T.; Ramanathan, S.; Fon Tacer, K.; Weon, J.L.; Potts, M.B.; Ou, Y.-H.; White, M.A.; Potts, P.R. Degradation of AMPK by a cancer-specific ubiquitin ligase. Cell 2015, 160, 715–728. [Google Scholar] [CrossRef]
- Sang, M.; Gu, L.; Yin, D.; Liu, F.; Lian, Y.; Zhang, X.; Liu, S.; Huang, W.; Wu, Y.; Shan, B. MAGE-A family expression is correlated with poor survival of patients with lung adenocarcinoma: A retrospective clinical study based on tissue microarray. J. Clin. Pathol. 2017, 70, 533–540. [Google Scholar] [CrossRef] [PubMed]
- Donato, R.; Cannon, B.R.; Sorci, G.; Riuzzi, F.; Hsu, K.; Weber, D.J.; Geczy, C.L. Functions of S100 proteins. Curr. Mol. Med. 2013, 13, 24–57. [Google Scholar] [CrossRef]
- Hermani, A.; De Servi, B.; Medunjanin, S.; Tessier, P.A.; Mayer, D. S100A8 and S100A9 activate MAP kinase and NF-kappaB signaling pathways and trigger translocation of RAGE in human prostate cancer cells. Exp. Cell Res. 2006, 312, 184–197. [Google Scholar] [CrossRef]
- Markowitz, J.; Carson, W.E., 3rd. Review of S100A9 biology and its role in cancer. Biochim. Biophys. Acta 2013, 1835, 100–109. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Zhou, L.; Song, Z.; Xiong, M.; Zhang, Y.; Yang, Y.; Chen, K.; Chen, Z. The identification of new biomarkers for bladder cancer: A study based on TCGA and GEO datasets. J. Cell. Physiol. 2019, 234, 15607–15618. [Google Scholar] [CrossRef] [PubMed]







| GSE Dataset | Tumor Type | Tumor Type | DEGs | Platform |
|---|---|---|---|---|
| GSE154261 | Non-muscle invasive (Surveillance) (n = 73) | Non-muscle invasive (Surgery) (n = 26) | 6078 | GPL20301 (Illumina HiSeq 4000) (Homo sapiens) |
| GSE57813 | Non-muscle invasive (n = 9) | Muscle invasive (n = 12) | 7975 | GPL14951Illumina HumanHT-12 WG-DASL V4.0 R2 expression beadchip |
| GSE37317 | Non-muscle invasive (n=8) | Muscle invasive (n=11) | 7589 | GPL96 [HG-U133A] Affymetrix Human Genome U133A Array |
| Gene | Entrez Gene Name | Ensemble Gene ID | Molecule Type | Putative Biological Function | Location |
|---|---|---|---|---|---|
| CDKN2A | Cyclin Dependent Kinase Inhibitor 2A | ENSG00000147889 | transcription regulator | Tumor suppressor | Nucleus |
| CDC20 | Cell Division Cycle 20 | ENSG00000117399 | Enzyme | Cell cycle regulator | Nucleus |
| CTSV | Cathepsin V | ENSG00000136943 | Peptidase | Lysosomal protease | Cytoplasm |
| FOXM1 | Forkhead box M1 | ENSG00000111206 | Transcription regulator | Promotes oncogenesis | Nucleus |
| MAGEA6 | MAGE Family Member A6 | ENSG00000197172 | other | Tumor progression | Cytoplasm |
| KRT23 | Keratin 23 | ENSG00000108244 | other | Structural integrity | Cytoplasm |
| S100A9 | S100 calcium binding protein A9 | ENSG00000163220 | other | Cell cycle progression | Cytoplasm |
| Factor | Frequency or Mean (STD) (n = 37) |
|---|---|
| Age (year) | 65.9 (10.6) |
| Weight | 82.1 (20.8) |
| Race (Asian/Black/White) | 4/4/29 |
| Stage (1/2/3/4) | 2/13/12/10 |
| Tumor aggressiveness (NMIBC / MIBC) | 15/22 |
| Tumor grade (High/Low) | 34/3 |
| Lymph node present (No/Yes) | 5/30 |
| Sex (female/male) | 5/32 |
| Factors | Odds Ratio (95% CI) | p Value |
|---|---|---|
| CDKN2A (per unit increase) | 1.03 (1.01, 1.05) | 0.01 |
| CDC20 (per unit increase) | 1.08 (0.92, 1.25) | 0.355 |
| CTSV (per unit increase) | 0.84 (0.74, 0.96) | 0.008 |
| FOXM1 (per unit increase) | 0.97 (0.96, 0.99) | 0.0009 |
| KRT23 (per unit increase) | 5.07 (1.32, 19.44) | 0.018 |
| MAGEA6 (per unit increase) | 1.03 (0.97, 1.09) | 0.419 |
| S100A9 (per unit increase) | 2.07 (0.63, 6.81) | 0.233 |
| Factors | Hazard Ratio (95% CI) | p Value |
|---|---|---|
| CDKN2A (per unit increase) | 0.999 (0.999, 1) | 0.127 |
| CDC20 (per unit increase) | 1.12 (1.02, 1.23) | 0.014 |
| CTSV (per unit increase) | 1.01 (0.96, 1.05) | 0.747 |
| FOXM1 (per unit increase) | 1.004 (0.998, 1.01) | 0.213 |
| KRT23 (per unit increase) | 1.19 (0.86, 1.64) | 0.297 |
| MAGEA6 (per unit increase) | 1 (0.999, 1.001) | 0.987 |
| S100A9 (per unit increase) | 0.94 (0.53, 1.66) | 0.834 |
| Factors | Hazard Ratio (95% CI) | p Value |
|---|---|---|
| CDKN2A (per unit increase) | 0.999 (0.999, 1) | 0.11 |
| CDC20 (per unit increase) | 1.12 (1.02, 1.22) | 0.021 |
| CTSV (per unit increase) | 1.01 (0.98, 1.05) | 0.529 |
| FOXM1 (per unit increase) | 1.002 (0.997, 1.01) | 0.447 |
| KRT23 (per unit increase) | 1.08 (0.79, 1.47) | 0.625 |
| MAGEA6 (per unit increase) | 1 (0.999, 1.001) | 0.954 |
| S100A9 (per unit increase) | 1.27 (0.79, 2.06) | 0.329 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Verma, S.; Shankar, E.; Lin, S.; Singh, V.; Chan, E.R.; Cao, S.; Fu, P.; MacLennan, G.T.; Ponsky, L.E.; Gupta, S. Identification of Key Genes Associated with Progression and Prognosis of Bladder Cancer through Integrated Bioinformatics Analysis. Cancers 2021, 13, 5931. https://doi.org/10.3390/cancers13235931
Verma S, Shankar E, Lin S, Singh V, Chan ER, Cao S, Fu P, MacLennan GT, Ponsky LE, Gupta S. Identification of Key Genes Associated with Progression and Prognosis of Bladder Cancer through Integrated Bioinformatics Analysis. Cancers. 2021; 13(23):5931. https://doi.org/10.3390/cancers13235931
Chicago/Turabian StyleVerma, Shiv, Eswar Shankar, Spencer Lin, Vaibhav Singh, E. Ricky Chan, Shufen Cao, Pingfu Fu, Gregory T. MacLennan, Lee E. Ponsky, and Sanjay Gupta. 2021. "Identification of Key Genes Associated with Progression and Prognosis of Bladder Cancer through Integrated Bioinformatics Analysis" Cancers 13, no. 23: 5931. https://doi.org/10.3390/cancers13235931
APA StyleVerma, S., Shankar, E., Lin, S., Singh, V., Chan, E. R., Cao, S., Fu, P., MacLennan, G. T., Ponsky, L. E., & Gupta, S. (2021). Identification of Key Genes Associated with Progression and Prognosis of Bladder Cancer through Integrated Bioinformatics Analysis. Cancers, 13(23), 5931. https://doi.org/10.3390/cancers13235931








