HLA Expression in Relation to HLA Type in Classic Hodgkin Lymphoma Patients
Abstract
:Simple Summary
Abstract
1. Introduction
2. Results
2.1. Patient Characteristics and HLA Expression
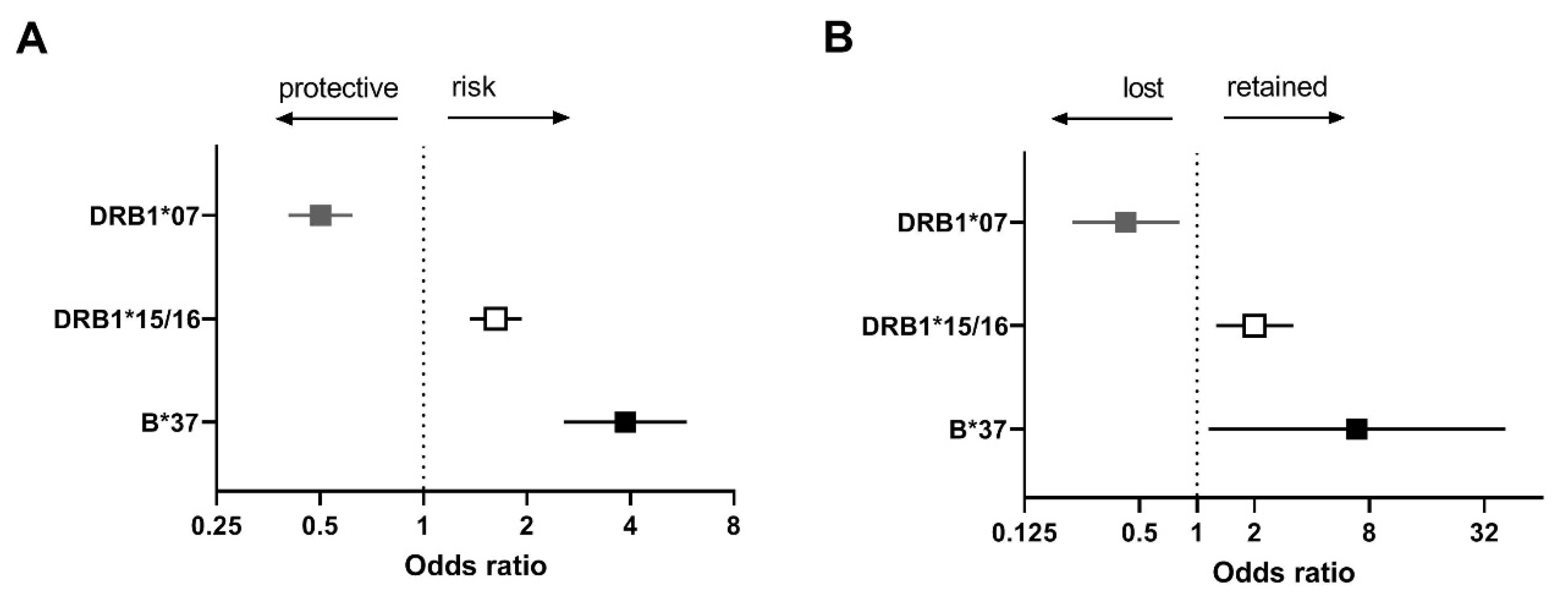
2.2. Identification of HLA Susceptibility Alleles in the Combined Dutch and UK cHL Cohorts
2.3. HLA Expression and Susceptibility in cHL Patients
3. Discussion
4. Materials and Methods
4.1. Study Outline
4.2. Patients
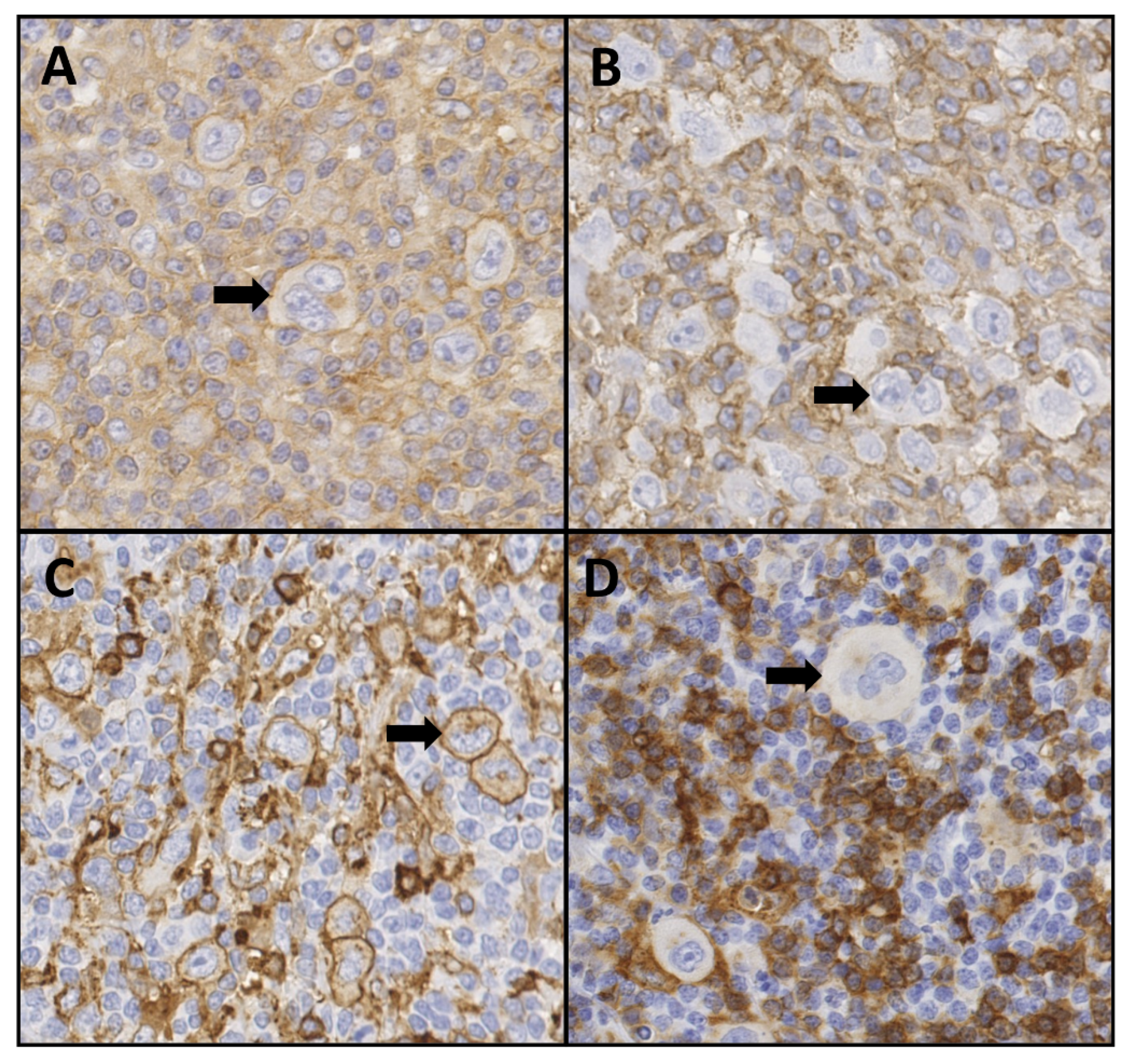
4.3. HLA Class I and Class II Expression
4.4. Meta-Analysis of Susceptibility Alleles
4.5. HLA Expression Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Engert, A.; Younes, A. Hodgkin Lymphoma A Comprehensive Overview; Springer: Amsterdam, The Netherlands, 2020. [Google Scholar]
- Deacon, E.M.; Pallesen, G.; Niedobitek, G.; Crocker, J.; Brooks, L.; Rickinson, A.B.; Young, L.S. Epstein-Barr virus and Hodgkin’s disease: Transcriptional analysis of virus latency in the malignant cells. J. Exp. Med. 1993, 177, 339–349. [Google Scholar] [CrossRef] [Green Version]
- Niedobitek, G.; Agathanggelou, A.; Herbst, H.; Whitehead, L.; Wright, D.H.; Young, L.S. Epstein-Barr virus (EBV) infection in infectious mononucleosis: Virus latency, replication and phenotype of EBV-infected cells. J. Pathol. 1997, 182, 151–159. [Google Scholar] [CrossRef]
- Long, H.M.; Taylor, G.S.; Rickinson, A.B. Immune defence against EBV and EBV-associated disease. Curr. Opin. Immunol. 2011, 23, 258–264. [Google Scholar] [CrossRef]
- Veldman, J.; Visser, L.; Huberts-Kregel, M.; Muller, N.; Hepkema, B.; van den Berg, A.; Diepstra, A. Rosetting T cells in Hodgkin lymphoma are activated by immunological synapse components HLA class II and CD58. Blood 2020, 136, 2437–2441. [Google Scholar] [CrossRef] [PubMed]
- Nijland, M.; Veenstra, R.N.; Visser, L.; Xu, C.; Kushekhar, K.; van Imhoff, G.W.; Kluin, P.M.; van den Berg, A.; Diepstra, A. HLA dependent immune escape mechanisms in B-cell lymphomas: Implications for immune checkpoint inhibitor therapy? Oncoimmunology 2017, 6, e1295202. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Diepstra, A.; van Imhoff, G.W.; Karim-Kos, H.E.; van den Berg, A.; te Meerman, G.J.; Niens, M.; Nolte, I.M.; Bastiaannet, E.; Schaapveld, M.; Vellenga, E.; et al. HLA class II expression by Hodgkin Reed-Sternberg cells is an independent prognostic factor in classical Hodgkin’s lymphoma. J. Clin. Oncol. 2007, 25, 3101–3108. [Google Scholar] [CrossRef]
- Sonmez, M.; Erkut, N.; Ucar, F.; Buruk, K.; Cobanoglu, U.; Bahce, M.; Ural, A.U. Familial Hodgkin’s lymphoma from the perspective of HLA. Intern. Med. 2010, 49, 607–610. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Diepstra, A.; Niens, M.; te Meerman, G.J.; Poppema, S.; van den Berg, A. Genetic susceptibility to Hodgkin’s lymphoma associated with the human leukocyte antigen region. Eur. J. Haematol. Suppl. 2005, 66, 34–41. [Google Scholar] [CrossRef]
- Diepstra, A.; Niens, M.; Vellenga, E.; van Imhoff, G.W.; Nolte, I.M.; Schaapveld, M.; van der Steege, G.; van den Berg, A.; Kibbelaar, R.E.; te Meerman, G.J.; et al. Association with HLA class I in Epstein-Barr-virus-positive and with HLA class III in Epstein-Barr-virus-negative Hodgkin’s lymphoma. Lancet 2005, 365, 2216–2224. [Google Scholar] [CrossRef]
- Urayama, K.Y.; Jarrett, R.F.; Hjalgrim, H.; Diepstra, A.; Kamatani, Y.; Chabrier, A.; Gaborieau, V.; Boland, A.; Nieters, A.; Becker, N.; et al. Genome-wide association study of classical Hodgkin lymphoma and Epstein-Barr virus status-defined subgroups. J. Natl. Cancer Inst. 2012, 104, 240–253. [Google Scholar] [CrossRef] [Green Version]
- Cozen, W.; Li, D.; Best, T.; Van Den Berg, D.J.; Gourraud, P.; Cortessis, V.K.; Skol, A.D.; Mack, T.M.; Glaser, S.L.; Weiss, L.M.; et al. A genome-wide meta-analysis of nodular sclerosing Hodgkin lymphoma identifies risk loci at 6p21.32. Blood 2012, 119, 469–475. [Google Scholar] [CrossRef] [PubMed]
- Frampton, M.; da Silva Filho, M.I.; Broderick, P.; Thomsen, H.; Forsti, A.; Vijayakrishnan, J.; Cooke, R.; Enciso-Mora, V.; Hoffmann, P.; Nothen, M.M.; et al. Variation at 3p24.1 and 6q23.3 influences the risk of Hodgkin’s lymphoma. Nat. Commun. 2013, 4, 2549. [Google Scholar] [CrossRef]
- Cozen, W.; Timofeeva, M.N.; Li, D.; Diepstra, A.; Hazelett, D.; Delahaye-Sourdeix, M.; Edlund, C.K.; Franke, L.; Rostgaard, K.; Van Den Berg, D.J.; et al. A meta-analysis of Hodgkin lymphoma reveals 19p13.3 TCF3 as a novel susceptibility locus. Nat. Commun. 2014, 5, 3856. [Google Scholar] [CrossRef] [Green Version]
- Niens, M.; Jarrett, R.F.; Hepkema, B.; Nolte, I.M.; Diepstra, A.; Platteel, M.; Kouprie, N.; Delury, C.P.; Gallagher, A.; Visser, L.; et al. HLA-A*02 is associated with a reduced risk and HLA-A*01 with an increased risk of developing EBV+ Hodgkin lymphoma. Blood 2007, 110, 3310–3315. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hjalgrim, H.; Rostgaard, K.; Johnson, P.C.; Lake, A.; Shield, L.; Little, A.M.; Ekstrom-Smedby, K.; Adami, H.O.; Glimelius, B.; Hamilton-Dutoit, S.; et al. HLA-A alleles and infectious mononucleosis suggest a critical role for cytotoxic T-cell response in EBV-related Hodgkin lymphoma. Proc. Natl. Acad. Sci. USA 2010, 107, 6400–6405. [Google Scholar] [CrossRef] [Green Version]
- Huang, X.; Kushekhar, K.; Nolte, I.; Kooistra, W.; Visser, L.; Bouwman, I.; Kouprie, N.; Veenstra, R.; van Imhoff, G.; Olver, B.; et al. HLA associations in classical Hodgkin lymphoma: EBV status matters. PLoS ONE 2012, 7, e39986. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Johnson, P.C.; McAulay, K.A.; Montgomery, D.; Lake, A.; Shield, L.; Gallagher, A.; Little, A.M.; Shah, A.; Marsh, S.G.; Taylor, G.M.; et al. Modeling HLA associations with EBV-positive and -negative Hodgkin lymphoma suggests distinct mechanisms in disease pathogenesis. Int. J. Cancer 2015, 137, 1066–1075. [Google Scholar] [CrossRef]
- Parham, P.; Ohta, T. Population biology of antigen presentation by MHC class I molecules. Science 1996, 272, 67–74. [Google Scholar] [CrossRef]
- Liu, Y.; l Razak, F.R.; Terpstra, M.; Chan, F.C.; Saber, A.; Nijland, M.; van Imhoff, G.; Visser, L.; Gascoyne, R.; Steidl, C.; et al. The mutational landscape of Hodgkin lymphoma cell lines determined by whole-exome sequencing. Leukemia 2014, 28, 2248–2251. [Google Scholar] [CrossRef]
- Wienand, K.; Chapuy, B.; Stewart, C.; Dunford, A.J.; Wu, D.; Kim, J.; Kamburov, A.; Wood, T.R.; Cader, F.Z.; Ducar, M.D.; et al. Genomic analyses of flow-sorted Hodgkin Reed-Sternberg cells reveal complementary mechanisms of immune evasion. Blood Adv. 2019, 3, 4065–4080. [Google Scholar] [CrossRef] [Green Version]
- Burr, M.L.; Sparbier, C.E.; Chan, K.L.; Chan, Y.C.; Kersbergen, A.; Lam, E.Y.N.; Azidis-Yates, E.; Vassiliadis, D.; Bell, C.C.; Gilan, O.; et al. An evolutionarily conserved function of polycomb silences the MHC class I antigen presentation pathway and enables immune evasion in cancer. Cancer Cell 2019, 36, 385–401.e8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- de Charette, M.; Houot, R. Hide or defend, the two strategies of lymphoma immune evasion: Potential implications for immunotherapy. Haematologica 2018, 103, 1256–1268. [Google Scholar] [CrossRef]
- Wilkinson, S.T.; Vanpatten, K.A.; Fernandez, D.R.; Brunhoeber, P.; Garsha, K.E.; Glinsmann-Gibson, B.J.; Grogan, T.M.; Teruya-Feldstein, J.; Rimsza, L.M. Partial plasma cell differentiation as a mechanism of lost major histocompatibility complex class II expression in diffuse large B-cell lymphoma. Blood 2012, 119, 1459–1467. [Google Scholar] [CrossRef] [Green Version]
- Jordanova, E.S.; Riemersma, S.A.; Philippo, K.; Giphart-Gassler, M.; Schuuring, E.; Kluin, P.M. Hemizygous deletions in the HLA region account for loss of heterozygosity in the majority of diffuse large B-cell lymphomas of the testis and the central nervous system. Genes Chromosomes Cancer 2002, 35, 38–48. [Google Scholar] [CrossRef]
- Jordanova, E.S.; Philippo, K.; Giphart, M.J.; Schuuring, E.; Kluin, P.M. Mutations in the HLA class II genes leading to loss of expression of HLA-DR and HLA-DQ in diffuse large B-cell lymphoma. Immunogenetics 2003, 55, 203–209. [Google Scholar] [CrossRef]
- Murray, P.G.; Constandinou, C.M.; Crocker, J.; Young, L.S.; Ambinder, R.F. Analysis of major histocompatibility complex class I, TAP expression, and LMP2 epitope sequence in Epstein-Barr virus-positive Hodgkin’s disease. Blood 1998, 92, 2477–2483. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.P.; Constandinou, C.M.; Thomas, W.A.; Croom-Carter, D.; Blake, N.W.; Murray, P.G.; Crocker, J.; Rickinson, A.B. Antigen presenting phenotype of Hodgkin Reed-Sternberg cells: Analysis of the HLA class I processing pathway and the effects of interleukin-10 on epstein-barr virus-specific cytotoxic T-cell recognition. Blood 1998, 92, 1020–1030. [Google Scholar] [CrossRef]
- Oudejans, J.J.; Jiwa, N.M.; Kummer, J.A.; Horstman, A.; Vos, W.; Baak, J.P.; Kluin, P.M.; van der Valk, P.; Walboomers, J.M.; Meijer, C.J. Analysis of major histocompatibility complex class I expression on Reed-Sternberg cells in relation to the cytotoxic T-cell response in Epstein-Barr virus-positive and -negative Hodgkin’s disease. Blood 1996, 87, 3844–3851. [Google Scholar] [CrossRef]
- Flavell, K.J.; Billingham, L.J.; Biddulph, J.P.; Gray, L.; Flavell, J.R.; Constandinou, C.M.; Young, L.S.; Murray, P.G. The effect of Epstein-Barr virus status on outcome in age- and sex-defined subgroups of patients with advanced Hodgkin’s disease. Ann. Oncol. 2003, 14, 282–290. [Google Scholar] [CrossRef] [PubMed]
- Rickinson, A.B.; Moss, D.J. Human cytotoxic T lymphocyte responses to Epstein-Barr virus infection. Annu. Rev. Immunol. 1997, 15, 405–431. [Google Scholar] [CrossRef]
- Brennan, R.M.; Burrows, S.R. A mechanism for the HLA-A*01-associated risk for EBV+ Hodgkin lymphoma and infectious mononucleosis. Blood 2008, 112, 2589–2590. [Google Scholar] [CrossRef]
- Lee, S.P.; Thomas, W.A.; Murray, R.J.; Khanim, F.; Kaur, S.; Young, L.S.; Rowe, M.; Kurilla, M.; Rickinson, A.B. HLA A2.1-restricted cytotoxic T cells recognizing a range of Epstein-Barr virus isolates through a defined epitope in latent membrane protein LMP2. J. Virol. 1993, 67, 7428–7435. [Google Scholar] [CrossRef] [Green Version]
- Jones, K.; Wockner, L.; Brennan, R.M.; Keane, C.; Chattopadhyay, P.K.; Roederer, M.; Price, D.A.; Cole, D.K.; Hassan, B.; Beck, K.; et al. The impact of HLA class I and EBV latency-II antigen-specific CD8(+) T cells on the pathogenesis of EBV(+) Hodgkin lymphoma. Clin. Exp. Immunol. 2016, 183, 206–220. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yao, Y.; Liu, N.; Zhou, Z.; Shi, L. Influence of ERAP1 and ERAP2 gene polymorphisms on disease susceptibility in different populations. Hum. Immunol. 2019, 80, 325–334. [Google Scholar] [CrossRef] [PubMed]
- Jiang, P.; Veenstra, R.N.; Seitz, A.; Nolte, I.M.; Hepkema, B.G.; Visser, L.; van den Berg, A.; Diepstra, A. Interaction between ERAP alleles and HLA class I types support a role of antigen presentation in Hodgkin lymphoma development. Cancers 2021, 13, 414. [Google Scholar] [CrossRef] [PubMed]
- Praest, P.; Luteijn, R.D.; Brak-Boer, I.G.J.; Lanfermeijer, J.; Hoelen, H.; Ijgosse, L.; Costa, A.I.; Gorham, R.D.; Lebbink, R.J.; Wiertz, E.J.H.J. The influence of TAP1 and TAP2 gene polymorphisms on TAP function and its inhibition by viral immune evasion proteins. Mol. Immunol. 2018, 101, 55–64. [Google Scholar] [CrossRef] [PubMed]
- Diepstra, A.; Poppema, S.; Boot, M.; Visser, L.; Nolte, I.M.; Niens, M.; Te Meerman, G.J.; van den Berg, A. HLA-G protein expression as a potential immune escape mechanism in classical Hodgkin’s lymphoma. Tissue Antigens 2008, 71, 219–226. [Google Scholar] [CrossRef]
- Rodgers, J.R.; Cook, R.G. MHC class Ib molecules bridge innate and acquired immunity. Nat. Rev. Immunol. 2005, 5, 459–471. [Google Scholar] [CrossRef]
| cHL Patient | Total | HLA I+ | HLA I− | p-Value | HLA II+ | HLA II− | p-Value |
|---|---|---|---|---|---|---|---|
| (n = 338) | (n = 85) | (n = 219) | (n = 184) | (n = 127) | |||
| Gender, n (%) | |||||||
| Male | 177 (52.4) | 55 (35.0) | 102 (65.0) | 0.0045 a | 96 (58.9) | 67 (41.1) | ns a |
| Female | 161 (47.6) | 30 (20.4) | 117 (79.6) | 88 (59.5) | 60 (40.5) | ||
| Age | |||||||
| Median (range) | 32 (12–78) | 37 (16–72) | 29 (12–78) | 0.0032 b | 30 (14–78) | 36 (15–70) | 0.0067 b |
| EBV status, n (%) | |||||||
| EBV positive | 86 (25.4) | 44 (57.9) | 32 (42.1) | <0.00001 a | 43 (58.1) | 31 (41.9) | ns a |
| EBV negative | 250 (74.0) | 41 (18.1) | 185 (81.9) | 140 (59.3) | 96 (40.7) | ||
| NA | 2 (0.6) | 0 (0) | 2 (100) | 1 (100) | 0 (0) | ||
| Subtype, n (%) | |||||||
| NS | 261 (77.2) | 53 (22.4) | 184 (77.6) | 0.00016 b | 147 (62.3) | 89 (37.7) | ns a |
| MC | 43 (12.7) | 22 (56.4) | 17 (43.6) | 25 (59.5) | 17 (40.5) | ||
| LR | 7 (2.1) | 1 (16.7) | 5 (83.3) | 2 (28.6) | 5 (71.4) | ||
| LD | 1 (0.3) | 0 (0) | 1 (100) | 0 (0) | 1 (100) | ||
| NOS | 26 (7.7) | 9 (42.9) | 12 (57.1) | 10 (40.0) | 15 (60.0) | ||
| cHL | HLA Type | OR | p-Value |
|---|---|---|---|
| Overall | B*44/45 | 0.68 | 1.4 × 10−4 |
| DRB1*04 | 0.59 | 1.8 × 10−6 | |
| DRB1*07 | 0.50 | 1.3 × 10−7 | |
| EBV− | B*07 | 1.49 | 1.9 × 10−4 |
| DRB1*15/16 | 1.62 | 5.0 × 10−6 | |
| DRB1*11/12 | 1.38 | 8.7 × 10−3 | |
| EBV+ | A*01 | 2.88 | 5.1 × 10−12 |
| A*02 | 0.52 | 2.7 × 10−5 | |
| B*08 | 2.12 | 2.0 × 10−6 | |
| B*37 | 3.87 | 6.4 × 10−8 | |
| DRB1*03 | 1.70 | 9.7 × 10−4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tan, G.W.; Jiang, P.; Nolte, I.M.; Kushekhar, K.; Veenstra, R.N.; Hepkema, B.G.; Jarrett, R.F.; van den Berg, A.; Diepstra, A. HLA Expression in Relation to HLA Type in Classic Hodgkin Lymphoma Patients. Cancers 2021, 13, 5833. https://doi.org/10.3390/cancers13225833
Tan GW, Jiang P, Nolte IM, Kushekhar K, Veenstra RN, Hepkema BG, Jarrett RF, van den Berg A, Diepstra A. HLA Expression in Relation to HLA Type in Classic Hodgkin Lymphoma Patients. Cancers. 2021; 13(22):5833. https://doi.org/10.3390/cancers13225833
Chicago/Turabian StyleTan, Geok Wee, Peijia Jiang, Ilja M. Nolte, Kushi Kushekhar, Rianne N. Veenstra, Bouke G. Hepkema, Ruth F. Jarrett, Anke van den Berg, and Arjan Diepstra. 2021. "HLA Expression in Relation to HLA Type in Classic Hodgkin Lymphoma Patients" Cancers 13, no. 22: 5833. https://doi.org/10.3390/cancers13225833
APA StyleTan, G. W., Jiang, P., Nolte, I. M., Kushekhar, K., Veenstra, R. N., Hepkema, B. G., Jarrett, R. F., van den Berg, A., & Diepstra, A. (2021). HLA Expression in Relation to HLA Type in Classic Hodgkin Lymphoma Patients. Cancers, 13(22), 5833. https://doi.org/10.3390/cancers13225833








