SLUG and Truncated TAL1 Reduce Glioblastoma Stem Cell Growth Downstream of Notch1 and Define Distinct Vascular Subpopulations in Glioblastoma Multiforme
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Lentiviral Transductions
2.3. Human Samples and Histology
2.4. Immunohistochemistry and Immunofluorescence
2.5. Immunofluorescence Followed by Fluorescence In Situ Hybridization (IF-FISH)
2.6. Measures of Cell Growth
2.7. Western Blots and Co-Immunoprecipitation
2.8. Bioinformatics and Statistical Analyses
3. Results
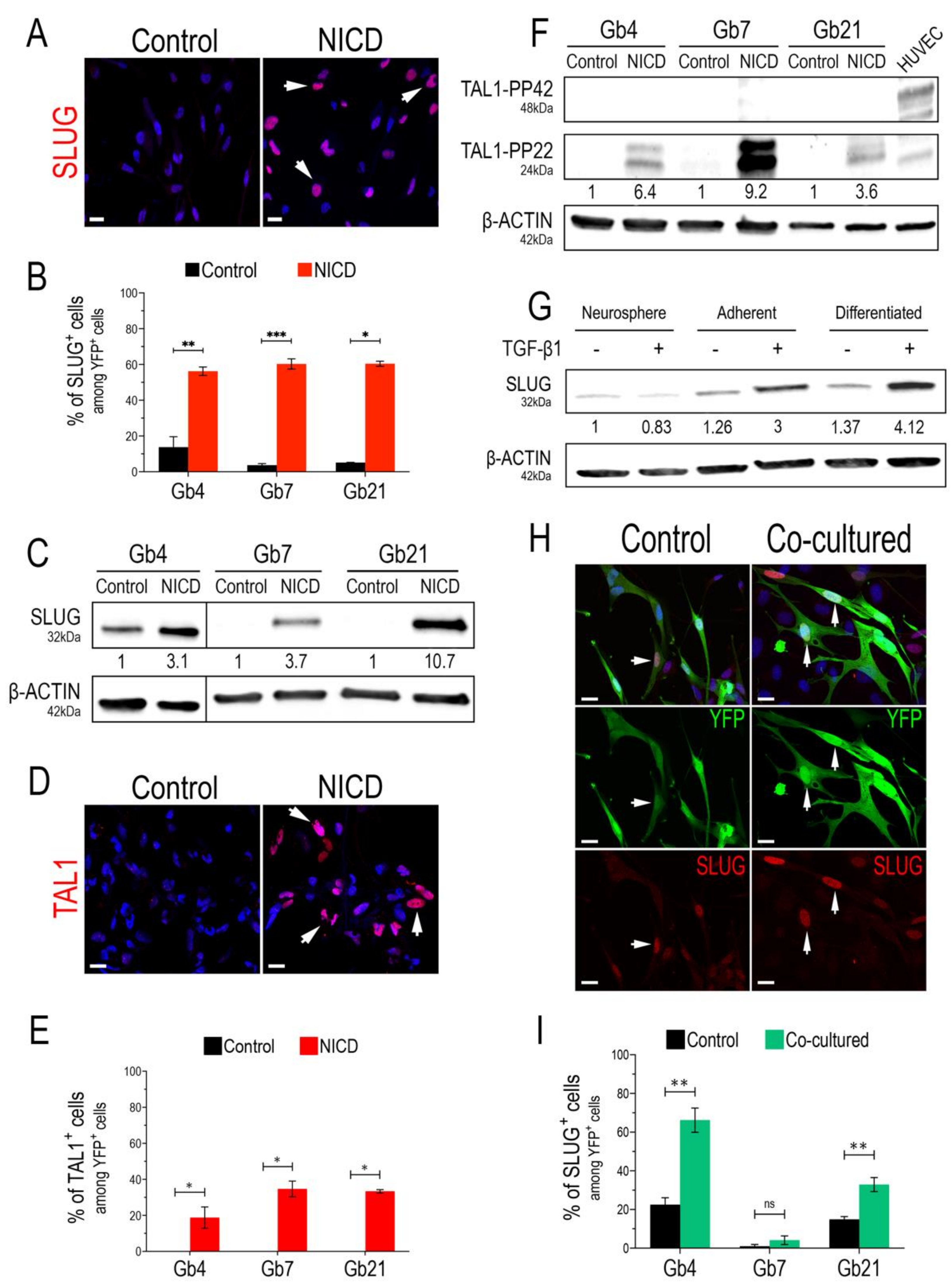
3.1. SLUG and TAL1 Are Inducible in Cultured GSCs
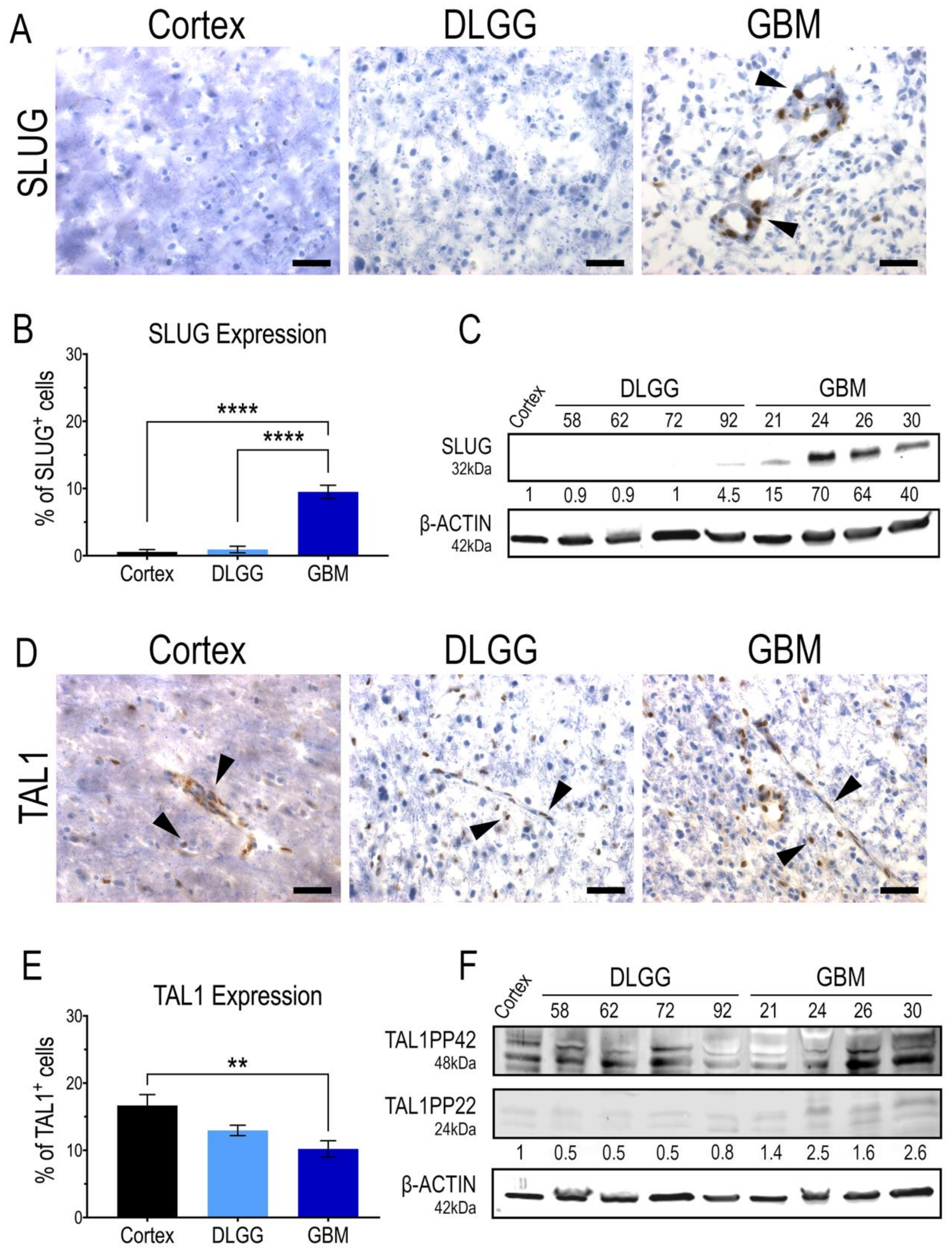
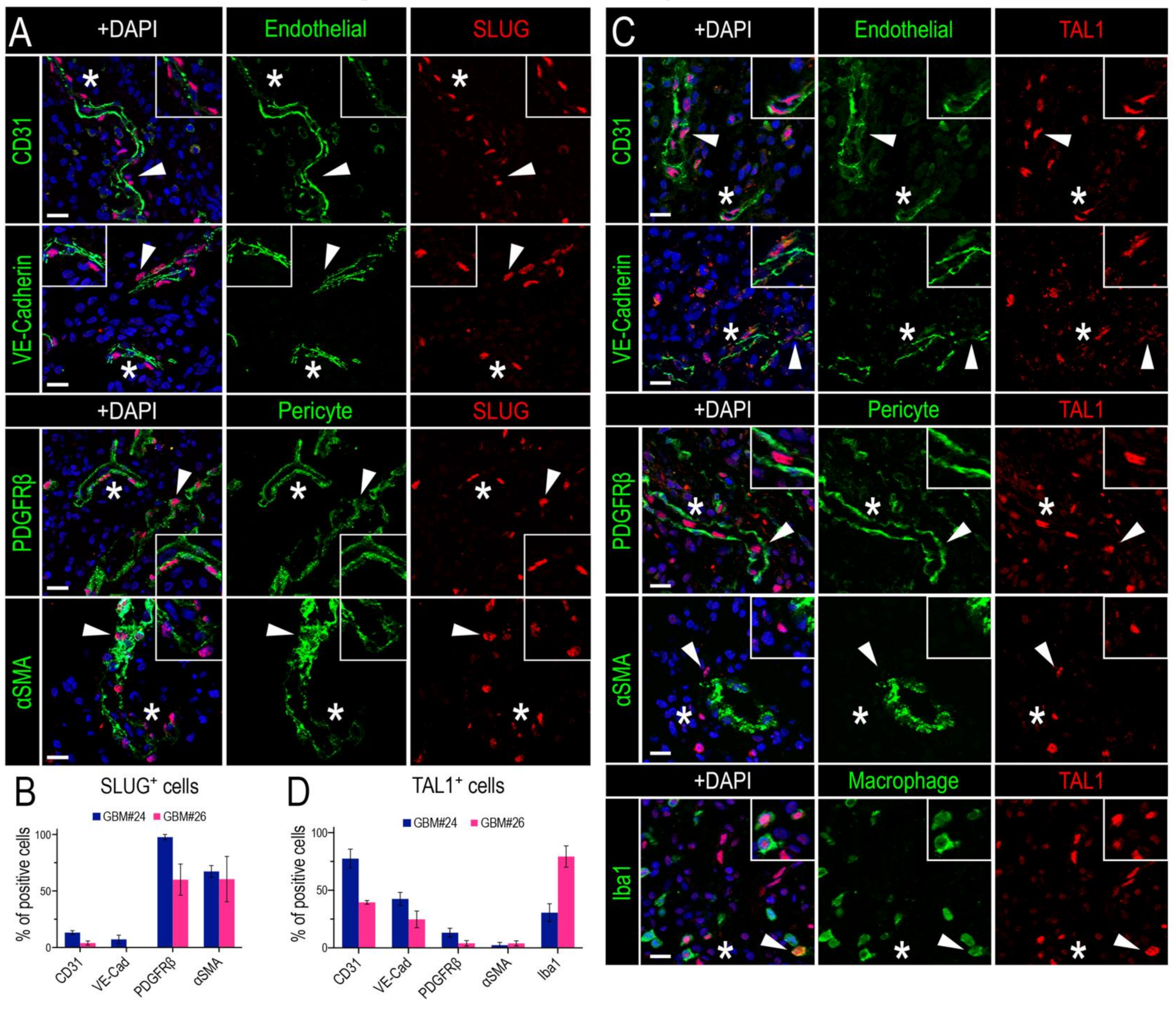
3.2. SLUG and TAL1 Define Mutually Exclusive Subpopulations of Vascular Cells in GBM Resections
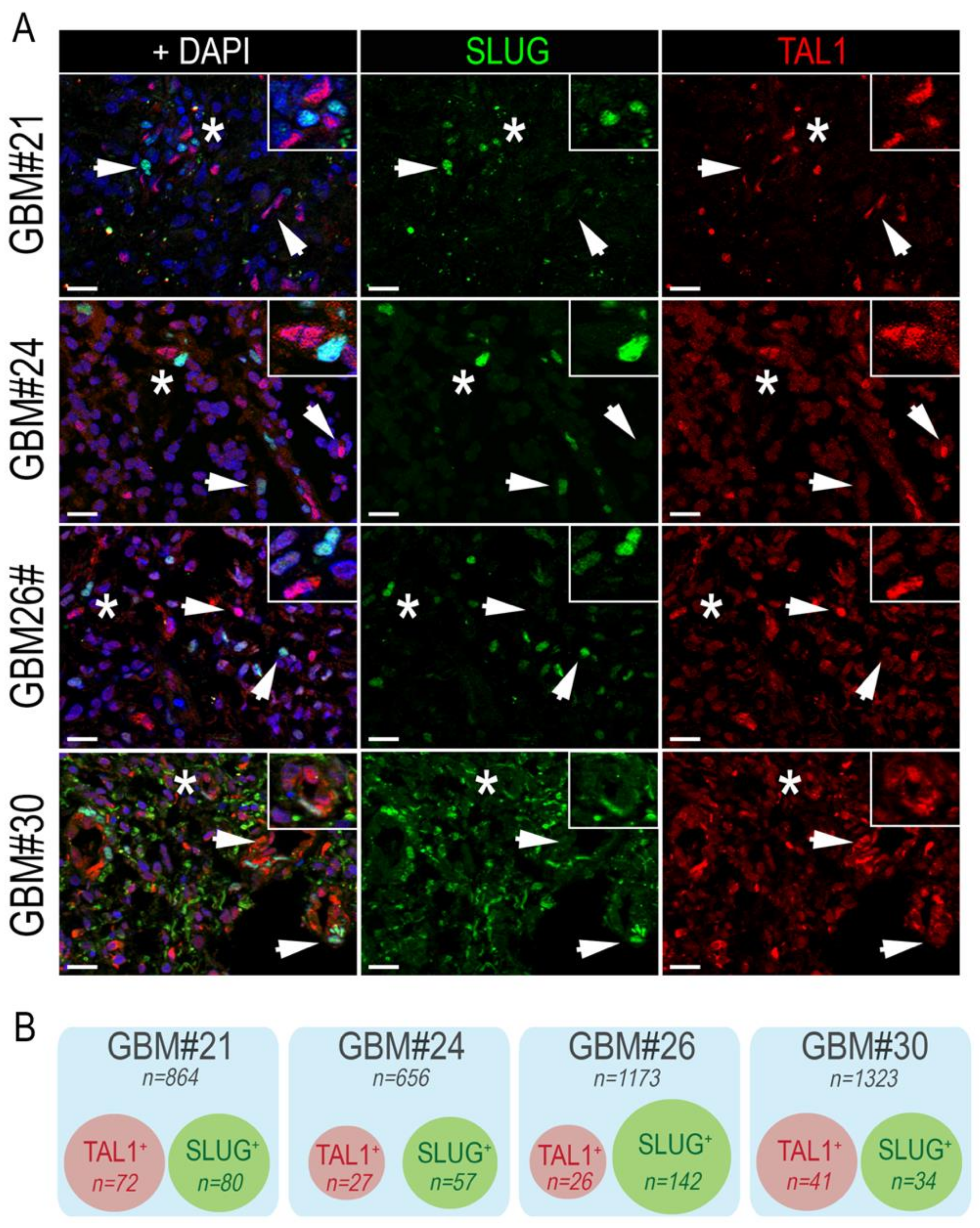
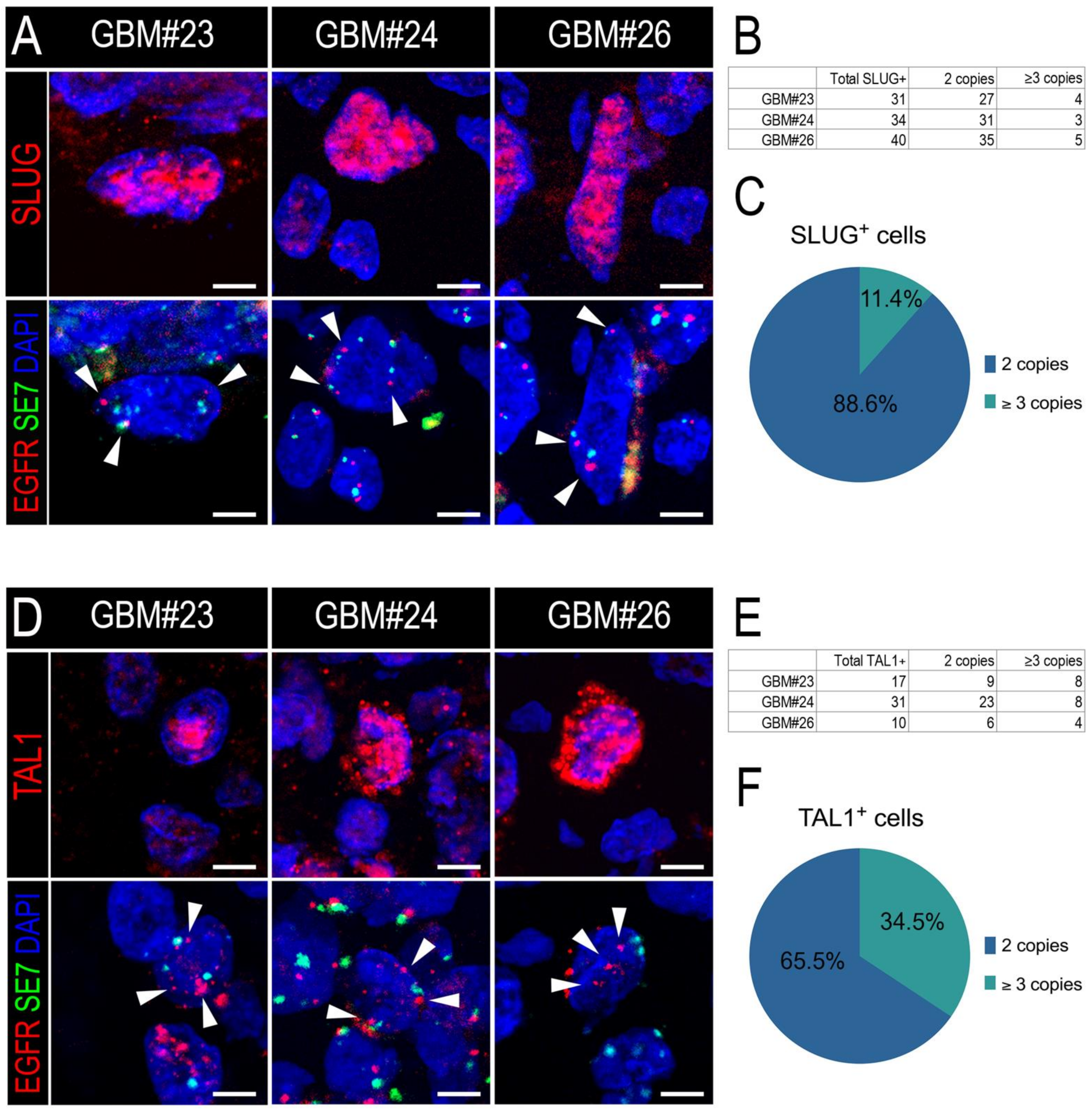
3.3. SLUG and TAL1 Subpopulations Contain Cells of Tumoral Origin in GBM Samples
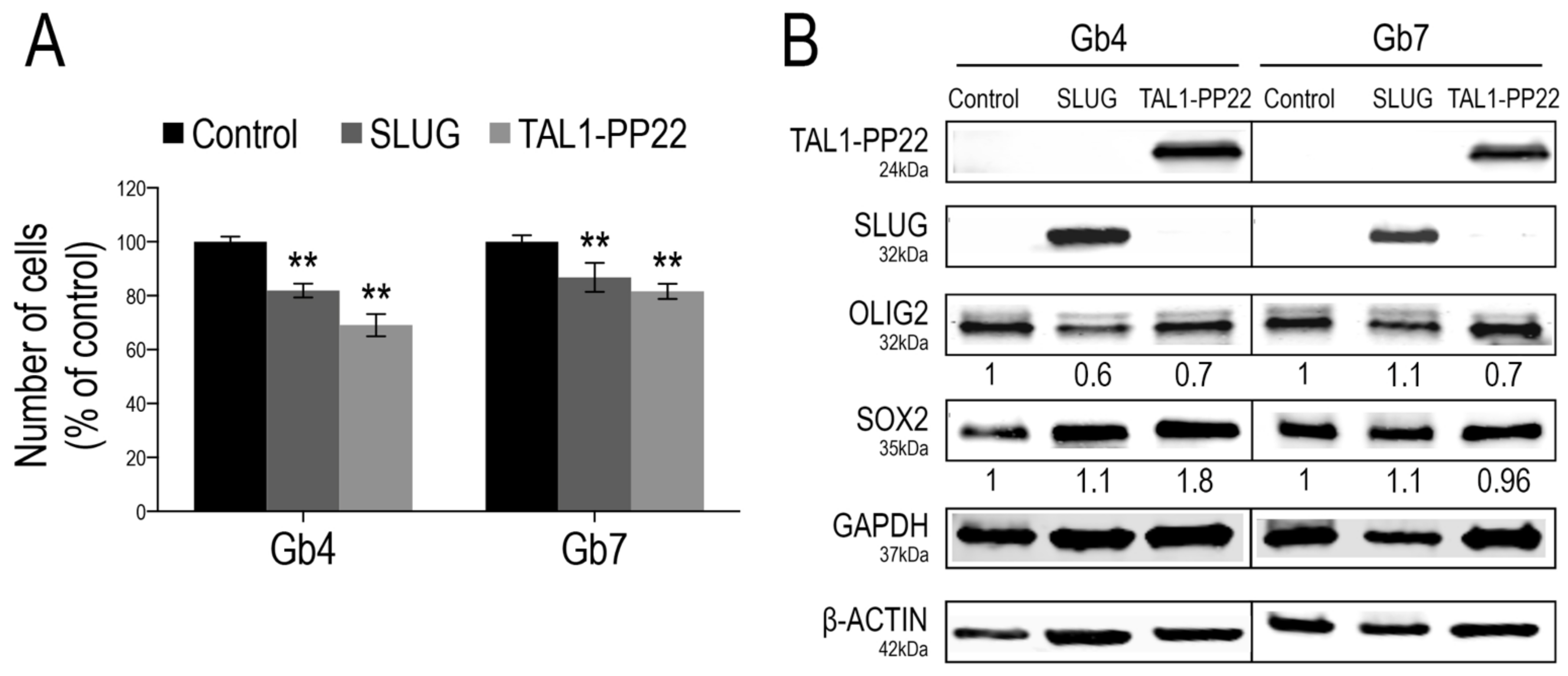
3.4. SLUG and TAL1-PP22 Independently Control the Growth of GSCs In Vitro
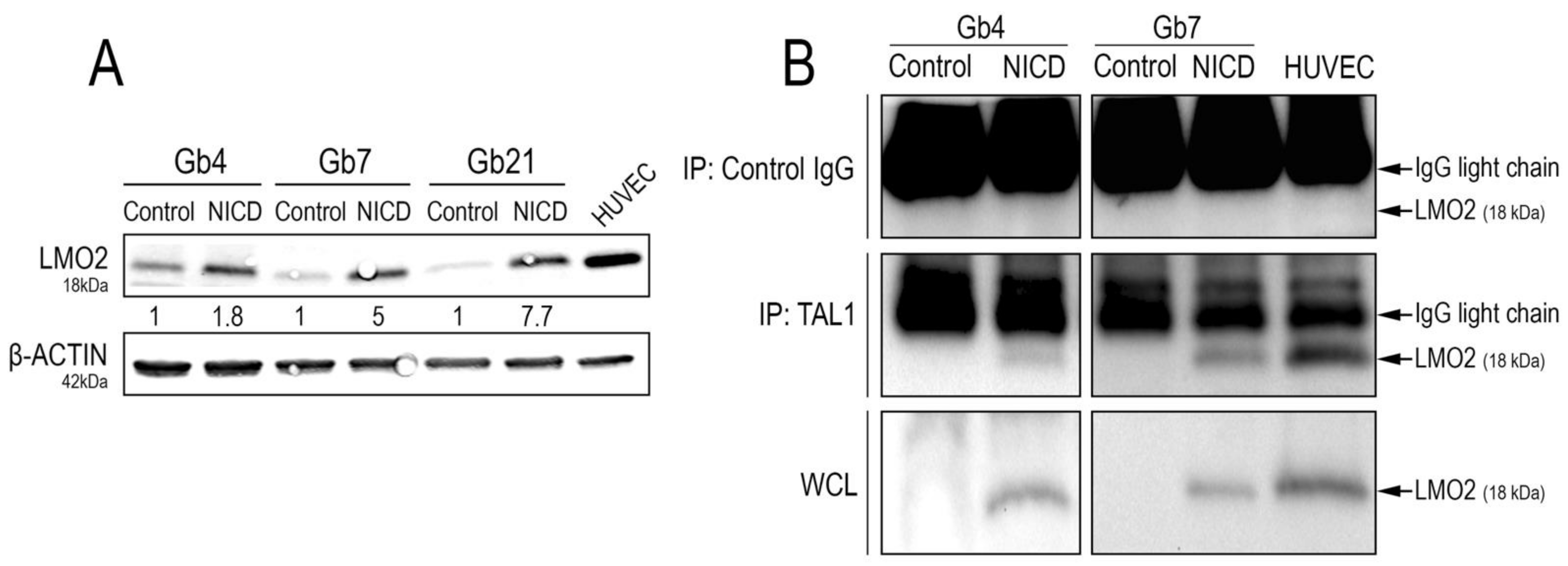
3.5. Truncated TAL1-pp22 Interacts with LMO2 upon Notch1 Activation of Cultured GSCs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Louis, D.N.; Perry, A.; Reifenberger, G.; von Deimling, A.; Figarella-Branger, D.; Cavenee, W.K.; Ohgaki, H.; Wiestler, O.D.; Kleihues, P.; Ellison, D.W. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A summary. Acta Neuropathol. 2016, 131, 803–820. [Google Scholar] [CrossRef]
- Aldape, K.; Brindle, K.M.; Chesler, L.; Chopra, R.; Gajjar, A.; Gilbert, M.R.; Gottardo, N.; Gutmann, D.H.; Hargrave, D.; Holland, E.C.; et al. Challenges to curing primary brain tumours. Nat. Rev. Clin. Oncol. 2019, 16, 509–520. [Google Scholar] [CrossRef] [PubMed]
- Verhaak, R.G.; Hoadley, K.A.; Purdom, E.; Wang, V.; Qi, Y.; Wilkerson, M.D.; Miller, C.R.; Ding, L.; Golub, T.; Mesirov, J.P.; et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 2010, 17, 98–110. [Google Scholar] [CrossRef]
- Patel, A.P.; Tirosh, I.; Trombetta, J.J.; Shalek, A.K.; Gillespie, S.M.; Wakimoto, H.; Cahill, D.P.; Nahed, B.V.; Curry, W.T.; Martuza, R.L.; et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 2014, 344, 1396–1401. [Google Scholar] [CrossRef] [PubMed]
- Brennan, C.W.; Verhaak, R.G.; McKenna, A.; Campos, B.; Noushmehr, H.; Salama, S.R.; Zheng, S.; Chakravarty, D.; Sanborn, J.Z.; Berman, S.H.; et al. The somatic genomic landscape of glioblastoma. Cell 2013, 155, 462–477. [Google Scholar] [CrossRef] [PubMed]
- Broekman, M.L.; Maas, S.L.N.; Abels, E.R.; Mempel, T.R.; Krichevsky, A.M.; Breakefield, X.O. Multidimensional communication in the microenvirons of glioblastoma. Nat. Rev. Neurol. 2018, 14, 482–495. [Google Scholar] [CrossRef] [PubMed]
- Hardee, M.E.; Zagzag, D. Mechanisms of glioma-associated neovascularization. Am. J. Pathol. 2012, 181, 1126–1141. [Google Scholar] [CrossRef] [PubMed]
- Galli, R.; Binda, E.; Orfanelli, U.; Cipelletti, B.; Gritti, A.; De Vitis, S.; Fiocco, R.; Foroni, C.; Dimeco, F.; Vescovi, A. Isolation and characterization of tumorigenic, stem-like neural precursors from human glioblastoma. Cancer Res. 2004, 64, 7011–7021. [Google Scholar] [CrossRef]
- Rennert, R.C.; Achrol, A.S.; Januszyk, M.; Kahn, S.A.; Liu, T.T.; Liu, Y.; Sahoo, D.; Rodrigues, M.; Maan, Z.N.; Wong, V.W.; et al. Multiple subsets of brain tumor initiating cells co-exist in glioblastoma. Stem Cells 2016, 34, 1702–1707. [Google Scholar] [CrossRef] [PubMed]
- Hambardzumyan, D.; Bergers, G. Glioblastoma: Defining Tumor Niches. Trends Cancer 2015, 1, 252–265. [Google Scholar] [CrossRef] [PubMed]
- Filatova, A.; Acker, T.; Garvalov, B.K. The cancer stem cell niche(s): The crosstalk between glioma stem cells and their microenvironment. Biochim. Biophys. Acta 2013, 1830, 2496–2508. [Google Scholar] [CrossRef] [PubMed]
- Zhu, T.S.; Costello, M.A.; Talsma, C.E.; Flack, C.G.; Crowley, J.G.; Hamm, L.L.; He, X.; Hervey-Jumper, S.L.; Heth, J.A.; Muraszko, K.M.; et al. Endothelial cells create a stem cell niche in glioblastoma by providing NOTCH ligands that nurture self-renewal of cancer stem-like cells. Cancer Res. 2011, 71, 6061–6072. [Google Scholar] [CrossRef] [PubMed]
- Calabrese, C.; Poppleton, H.; Kocak, M.; Hogg, T.L.; Fuller, C.; Hamner, B.; Oh, E.Y.; Gaber, M.W.; Finklestein, D.; Allen, M.; et al. A perivascular niche for brain tumor stem cells. Cancer Cell 2007, 11, 69–82. [Google Scholar] [CrossRef] [PubMed]
- Prager, B.C.; Bhargava, S.; Mahadev, V.; Hubert, C.G.; Rich, J.N. Glioblastoma Stem Cells: Driving Resilience through Chaos. Trends Cancer 2020, 6, 223–235. [Google Scholar] [CrossRef]
- Castellan, M.; Guarnieri, A.; Fujimura, A.; Zanconato, F.; Battilana, G.; Panciera, T.; Sladitschek, H.L.; Contessotto, P.; Citron, A.; Grilli, A.; et al. Single-cell analyses reveal YAP/TAZ as regulators of stemness and cell plasticity in Glioblastoma. Nat. Cancer 2021, 2, 174–188. [Google Scholar] [CrossRef]
- Peñuelas, S.; Anido, J.; Prieto-Sánchez, R.M.; Folch, G.; Barba, I.; Cuartas, I.; García-Dorado, D.; Poca, M.A.; Sahuquillo, J.; Baselga, J.; et al. TGF-beta increases glioma-initiating cell self-renewal through the induction of LIF in human glioblastoma. Cancer Cell 2009, 15, 315–327. [Google Scholar] [CrossRef]
- Parmigiani, E.; Taylor, V.; Giachino, C. Oncogenic and Tumor-Suppressive Functions of NOTCH Signaling in Glioma. Cells 2020, 9, 2304. [Google Scholar] [CrossRef]
- Guichet, P.O.; Guelfi, S.; Teigell, M.; Hoppe, L.; Bakalara, N.; Bauchet, L.; Duffau, H.; Lamszus, K.; Rothhut, B.; Hugnot, J.P. Notch1 stimulation induces a vascularization switch with pericyte-like cell differentiation of glioblastoma stem cells. Stem Cells 2015, 33, 21–34. [Google Scholar] [CrossRef] [PubMed]
- Augustus, M.; Pineau, D.; Aimond, F.; Azar, S.; Lecca, D.; Scamps, F.; Muxel, S.; Darlix, A.; Ritchie, W.; Gozé, C.; et al. Identification of CRYAB(+) KCNN3(+) SOX9(+) Astrocyte-Like and EGFR(+) PDGFRA(+) OLIG1(+) Oligodendrocyte-Like Tumoral Cells in Diffuse IDH1-Mutant Gliomas and Implication of NOTCH1 Signalling in Their Genesis. Cancers 2021, 13, 2107. [Google Scholar] [CrossRef] [PubMed]
- Phillips, S.; Kuperwasser, C. SLUG: Critical regulator of epithelial cell identity in breast development and cancer. Cell Adh. Migr. 2014, 8, 578–587. [Google Scholar] [CrossRef]
- Nieto, M.A.; Huang, R.Y.; Jackson, R.A.; Thiery, J.P. Emt: 2016. Cell 2016, 166, 21–45. [Google Scholar] [CrossRef] [PubMed]
- Hajra, K.M.; Chen, D.Y.; Fearon, E.R. The SLUG zinc-finger protein represses E-cadherin in breast cancer. Cancer Res. 2002, 62, 1613–1618. [Google Scholar]
- Mani, S.A.; Guo, W.; Liao, M.J.; Eaton, E.N.; Ayyanan, A.; Zhou, A.Y.; Brooks, M.; Reinhard, F.; Zhang, C.C.; Shipitsin, M.; et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 2008, 133, 704–715. [Google Scholar] [CrossRef]
- Oh, S.J.; Ahn, E.J.; Kim, O.; Kim, D.; Jung, T.Y.; Jung, S.; Lee, J.H.; Kim, K.K.; Kim, H.; Kim, E.H.; et al. The Role Played by SLUG, an Epithelial-Mesenchymal Transition Factor, in Invasion and Therapeutic Resistance of Malignant Glioma. Cell Mol. Neurobiol. 2019, 39, 769–782. [Google Scholar] [CrossRef]
- Cheng, W.Y.; Kandel, J.J.; Yamashiro, D.J.; Canoll, P.; Anastassiou, D. A multi-cancer mesenchymal transition gene expression signature is associated with prolonged time to recurrence in glioblastoma. PLoS ONE 2012, 7, e34705. [Google Scholar] [CrossRef] [PubMed]
- Oh, H.; Hwang, I.; Jang, J.Y.; Wu, L.; Cao, D.; Yao, J.; Ying, H.; Li, J.Y.; Yao, Y.; Hu, B.; et al. Therapy-Induced Transdifferentiation Promotes Glioma Growth Independent of EGFR Signaling. Cancer Res. 2021, 81, 1528–1539. [Google Scholar] [CrossRef]
- Lecuyer, E.; Hoang, T. SCL: From the origin of hematopoiesis to stem cells and leukemia. Exp. Hematol. 2004, 32, 11–24. [Google Scholar] [CrossRef] [PubMed]
- Correia, N.C.; Arcangeli, M.L.; Pflumio, F.; Barata, J.T. Stem Cell Leukemia: How a TALented actor can go awry on the hematopoietic stage. Leukemia 2016, 30, 1968–1978. [Google Scholar] [CrossRef] [PubMed]
- D’Souza, S.L.; Elefanty, A.G.; Keller, G. SCL/Tal-1 is essential for hematopoietic commitment of the hemangioblast but not for its development. Blood 2005, 105, 3862–3870. [Google Scholar] [CrossRef]
- Kim, P.G.; Albacker, C.E.; Lu, Y.F.; Jang, I.H.; Lim, Y.; Heffner, G.C.; Arora, N.; Bowman, T.V.; Lin, M.I.; Lensch, M.W.; et al. Signaling axis involving Hedgehog, Notch, and Scl promotes the embryonic endothelial-to-hematopoietic transition. Proc. Natl. Acad. Sci. USA 2013, 110, E141–E150. [Google Scholar] [CrossRef]
- Visvader, J.E.; Mao, X.; Fujiwara, Y.; Hahm, K.; Orkin, S.H. The LIM-domain binding protein Ldb1 and its partner LMO2 act as negative regulators of erythroid differentiation. Proc. Natl. Acad. Sci. USA 1997, 94, 13707–13712. [Google Scholar] [CrossRef]
- Ema, M.; Faloon, P.; Zhang, W.J.; Hirashima, M.; Reid, T.; Stanford, W.L.; Orkin, S.; Choi, K.; Rossant, J. Combinatorial effects of Flk1 and Tal1 on vascular and hematopoietic development in the mouse. Genes Dev. 2003, 17, 380–393. [Google Scholar] [CrossRef]
- De Val, S.; Black, B.L. Transcriptional control of endothelial cell development. Dev. Cell 2009, 16, 180–195. [Google Scholar] [CrossRef] [PubMed]
- Peng, C.Y.; Yajima, H.; Burns, C.E.; Zon, L.I.; Sisodia, S.S.; Pfaff, S.L.; Sharma, K. Notch and MAML signaling drives Scl-dependent interneuron diversity in the spinal cord. Neuron 2007, 53, 813–827. [Google Scholar] [CrossRef]
- Joshi, K.; Lee, S.; Lee, B.; Lee, J.W.; Lee, S.K. LMO4 controls the balance between excitatory and inhibitory spinal V2 interneurons. Neuron 2009, 61, 839–851. [Google Scholar] [CrossRef] [PubMed]
- Lahti, L.; Haugas, M.; Tikker, L.; Airavaara, M.; Voutilainen, M.H.; Anttila, J.; Kumar, S.; Inkinen, C.; Salminen, M.; Partanen, J. Differentiation and molecular heterogeneity of inhibitory and excitatory neurons associated with midbrain dopaminergic nuclei. Development 2016, 143, 516–529. [Google Scholar] [CrossRef] [PubMed]
- Muroyama, Y.; Fujiwara, Y.; Orkin, S.H.; Rowitch, D.H. Specification of astrocytes by bHLH protein SCL in a restricted region of the neural tube. Nature 2005, 438, 360–363. [Google Scholar] [CrossRef] [PubMed]
- Lazrak, M.; Deleuze, V.; Noel, D.; Haouzi, D.; Chalhoub, E.; Dohet, C.; Robbins, I.; Mathieu, D. The bHLH TAL-1/SCL regulates endothelial cell migration and morphogenesis. J. Cell Sci. 2004, 117, 1161–1171. [Google Scholar] [CrossRef]
- Deleuze, V.; Chalhoub, E.; El-Hajj, R.; Dohet, C.; Le Clech, M.; Couraud, P.O.; Huber, P.; Mathieu, D. TAL-1/SCL and its partners E47 and LMO2 up-regulate VE-cadherin expression in endothelial cells. Mol. Cell. Biol. 2007, 27, 2687–2697. [Google Scholar] [CrossRef]
- Mathieu, D. The bHLH TAL1 protein: A key molecule in the hematopoietic and endothelial systems. J. De La Soc. De Biol. 2009, 203, 143–153. [Google Scholar] [CrossRef]
- Palii, C.G.; Vulesevic, B.; Fraineau, S.; Pranckeviciene, E.; Griffith, A.J.; Chu, A.; Faralli, H.; Li, Y.; McNeill, B.; Sun, J.; et al. Trichostatin A enhances vascular repair by injected human endothelial progenitors through increasing the expression of TAL1-dependent genes. Cell Stem Cell 2014, 14, 644–657. [Google Scholar] [CrossRef] [PubMed]
- Glasker, S.; Li, J.; Xia, J.B.; Okamoto, H.; Zeng, W.; Lonser, R.R.; Zhuang, Z.; Oldfield, E.H.; Vortmeyer, A.O. Hemangioblastomas share protein expression with embryonal hemangioblast progenitor cell. Cancer Res. 2006, 66, 4167–4172. [Google Scholar] [CrossRef]
- Cao, S.; Zheng, J.; Liu, X.; Liu, Y.; Ruan, X.; Ma, J.; Liu, L.; Wang, D.; Yang, C.; Cai, H.; et al. FXR1 promotes the malignant biological behavior of glioma cells via stabilizing MIR17HG. J. Exp. Clin. Cancer Res. 2019, 38, 37. [Google Scholar] [CrossRef] [PubMed]
- Guichet, P.O.; Bieche, I.; Teigell, M.; Serguera, C.; Rothhut, B.; Rigau, V.; Scamps, F.; Ripoll, C.; Vacher, S.; Taviaux, S.; et al. Cell death and neuronal differentiation of glioblastoma stem-like cells induced by neurogenic transcription factors. Glia 2013, 61, 225–239. [Google Scholar] [CrossRef] [PubMed]
- Guichet, P.O.; Guelfi, S.; Ripoll, C.; Teigell, M.; Sabourin, J.C.; Bauchet, L.; Rigau, V.; Rothhut, B.; Hugnot, J.P. Asymmetric Distribution of GFAP in Glioma Multipotent Cells. PLoS ONE 2016, 11, e0151274. [Google Scholar] [CrossRef] [PubMed]
- Azar, S.; Leventoux, N.; Ripoll, C.; Rigau, V.; Goze, C.; Lorcy, F.; Bauchet, L.; Duffau, H.; Guichet, P.O.; Rothhut, B.; et al. Cellular and molecular characterization of IDH1-mutated diffuse low grade gliomas reveals tumor heterogeneity and absence of EGFR/PDGFRalpha activation. Glia 2018, 66, 239–255. [Google Scholar] [CrossRef]
- Bowman, R.L.; Wang, Q.; Carro, A.; Verhaak, R.G.; Squatrito, M. GlioVis data portal for visualization and analysis of brain tumor expression datasets. Neuro-Oncology 2017, 19, 139–141. [Google Scholar] [CrossRef] [PubMed]
- Darmanis, S.; Sloan, S.A.; Croote, D.; Mignardi, M.; Chernikova, S.; Samghababi, P.; Zhang, Y.; Neff, N.; Kowarsky, M.; Caneda, C.; et al. Single-Cell RNA-Seq Analysis of Infiltrating Neoplastic Cells at the Migrating Front of Human Glioblastoma. Cell Rep. 2017, 21, 1399–1410. [Google Scholar] [CrossRef]
- Ren, X.; Gomez, G.A.; Zhang, B.; Lin, S. Scl isoforms act downstream of etsrp to specify angioblasts and definitive hematopoietic stem cells. Blood 2010, 115, 5338–5346. [Google Scholar] [CrossRef] [PubMed]
- Zhen, F.; Lan, Y.; Yan, B.; Zhang, W.; Wen, Z. Hemogenic endothelium specification and hematopoietic stem cell maintenance employ distinct Scl isoforms. Development 2013, 140, 3977–3985. [Google Scholar] [CrossRef]
- Marin, F.; Nieto, M.A. Expression of chicken slug and snail in mesenchymal components of the developing central nervous system. Dev. Dyn. Off. Publ. Am. Assoc. Anat. 2004, 230, 144–148. [Google Scholar] [CrossRef]
- Hoang, T.; Lambert, J.A.; Martin, R. SCL/TAL1 in Hematopoiesis and Cellular Reprogramming. Curr. Top. Dev. Biol. 2016, 118, 163–204. [Google Scholar] [CrossRef]
- Deleuze, V.; El-Hajj, R.; Chalhoub, E.; Dohet, C.; Pinet, V.; Couttet, P.; Mathieu, D. Angiopoietin-2 is a direct transcriptional target of TAL1, LYL1 and LMO2 in endothelial cells. PLoS ONE 2012, 7, e40484. [Google Scholar] [CrossRef]
- Kim, S.H.; Kim, E.J.; Hitomi, M.; Oh, S.Y.; Jin, X.; Jeon, H.M.; Beck, S.; Jin, X.; Kim, J.K.; Park, C.G.; et al. The LIM-only transcription factor LMO2 determines tumorigenic and angiogenic traits in glioma stem cells. Cell Death Differ. 2015, 22, 1517–1525. [Google Scholar] [CrossRef]
- Park, C.G.; Sohn, Y.W.; Kim, E.J.; Kim, S.H.; Kim, S.C.; Kim, H. LIM domain only 2 induces glioma invasion via cytosolic p27(KIP1). Tumour Biol. 2016, 37, 2473–2480. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Bergström, T.; Jiang, Y.; Johansson, P.; Marinescu, V.D.; Lindberg, N.; Segerman, A.; Wicher, G.; Niklasson, M.; Baskaran, S.; et al. The Human Glioblastoma Cell Culture Resource: Validated Cell Models Representing All Molecular Subtypes. EBioMedicine 2015, 2, 1351–1363. [Google Scholar] [CrossRef]
- Shutter, J.R.; Scully, S.; Fan, W.; Richards, W.G.; Kitajewski, J.; Deblandre, G.A.; Kintner, C.R.; Stark, K.L. Dll4, a novel Notch ligand expressed in arterial endothelium. Genes Dev. 2000, 14, 1313–1318. [Google Scholar]
- Merrilees, M.J.; Sodek, J. Synthesis of TGF-beta 1 by vascular endothelial cells is correlated with cell spreading. J. Vasc. Res. 1992, 29, 376–384. [Google Scholar] [CrossRef]
- Yang, H.W.; Menon, L.G.; Black, P.M.; Carroll, R.S.; Johnson, M.D. SNAI2/Slug promotes growth and invasion in human gliomas. BMC Cancer 2010, 10, 301. [Google Scholar] [CrossRef] [PubMed]
- Torsvik, A.; Stieber, D.; Enger, P.; Golebiewska, A.; Molven, A.; Svendsen, A.; Westermark, B.; Niclou, S.P.; Olsen, T.K.; Chekenya Enger, M.; et al. U-251 revisited: Genetic drift and phenotypic consequences of long-term cultures of glioblastoma cells. Cancer Med. 2014, 3, 812–824. [Google Scholar] [CrossRef]
- Zhou, W.; Gross, K.M.; Kuperwasser, C. Molecular regulation of Snai2 in development and disease. J. Cell Sci. 2019, 132, jcs235127. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wakeman, T.P.; Lathia, J.D.; Hjelmeland, A.B.; Wang, X.F.; White, R.R.; Rich, J.N.; Sullenger, B.A. Notch promotes radioresistance of glioma stem cells. Stem Cells 2010, 28, 17–28. [Google Scholar] [CrossRef] [PubMed]
- Thippu Jayaprakash, K.; Michael, A. Notch Inhibition: A Promising Strategy to Improve Radiosensitivity and Curability of Radiotherapy. Clin. Oncol. (R Coll. Radiol.) 2021, 33, e44–e49. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Chen, H.; Zhang, J.; Chen, Y.; Wang, M.; Ma, J.; Hong, L.; Liu, N.; Fan, Q.; Lu, X.; et al. Increased Notch Signaling Enhances Radioresistance of Malignant Stromal Cells Induced by Glioma Stem/Progenitor Cells. PLoS ONE 2015, 10, e0142594. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.C.; Tsai, J.T.; Chao, T.Y.; Ma, H.I.; Liu, W.H. The STAT3/Slug Axis Enhances Radiation-Induced Tumor Invasion and Cancer Stem-like Properties in Radioresistant Glioblastoma. Cancers 2018, 10, 512. [Google Scholar] [CrossRef] [PubMed]
- Mader, L.; Blank, A.E.; Capper, D.; Jansong, J.; Baumgarten, P.; Wirsik, N.M.; Zachskorn, C.; Ehlers, J.; Seifert, M.; Klink, B.; et al. Pericytes/vessel-associated mural cells (VAMCs) are the major source of key epithelial-mesenchymal transition (EMT) factors SLUG and TWIST in human glioma. Oncotarget 2018, 9, 24041–24053. [Google Scholar] [CrossRef] [PubMed]
- Chesnelong, C.; Hao, X.; Cseh, O.; Wang, A.Y.; Luchman, H.A.; Weiss, S. SLUG Directs the Precursor State of Human Brain Tumor Stem Cells. Cancers 2019, 11, 1635. [Google Scholar] [CrossRef]
- Wirsik, N.M.; Ehlers, J.; Mäder, L.; Ilina, E.I.; Blank, A.E.; Grote, A.; Feuerhake, F.; Baumgarten, P.; Devraj, K.; Harter, P.N.; et al. TGF-β activates pericytes via induction of the epithelial-to-mesenchymal transition protein SLUG in glioblastoma. Neuropathol. Appl. Neurobiol. 2021, 47, 768–780. [Google Scholar] [CrossRef]
- Coll-Bonfill, N.; Peinado, V.I.; Pisano, M.V.; Parrizas, M.; Blanco, I.; Evers, M.; Engelmann, J.C.; Garcia-Lucio, J.; Tura-Ceide, O.; Meister, G.; et al. Slug Is Increased in Vascular Remodeling and Induces a Smooth Muscle Cell Proliferative Phenotype. PLoS ONE 2016, 11, e0159460. [Google Scholar] [CrossRef]
- Rosińska, S.; Gavard, J. Tumor Vessels Fuel the Fire in Glioblastoma. Int. J. Mol. Sci. 2021, 22, 6514. [Google Scholar] [CrossRef] [PubMed]
- Guelfi, S.; Duffau, H.; Bauchet, L.; Rothhut, B.; Hugnot, J.P. Vascular Transdifferentiation in the CNS: A Focus on Neural and Glioblastoma Stem-Like Cells. Stem Cells Int. 2016, 2016, 2759403. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.; Huang, Z.; Zhou, W.; Wu, Q.; Donnola, S.; Liu, J.K.; Fang, X.; Sloan, A.E.; Mao, Y.; Lathia, J.D.; et al. Glioblastoma stem cells generate vascular pericytes to support vessel function and tumor growth. Cell 2013, 153, 139–152. [Google Scholar] [CrossRef]
- Calkhoven, C.F.; Muller, C.; Martin, R.; Krosl, G.; Pietsch, H.; Hoang, T.; Leutz, A. Translational control of SCL-isoform expression in hematopoietic lineage choice. Genes Dev. 2003, 17, 959–964. [Google Scholar] [CrossRef]
- Qian, F.; Zhen, F.; Xu, J.; Huang, M.; Li, W.; Wen, Z. Distinct functions for different scl isoforms in zebrafish primitive and definitive hematopoiesis. PLoS Biol. 2007, 5, e132. [Google Scholar] [CrossRef]
- Bernard, O.; Azogui, O.; Lecointe, N.; Mugneret, F.; Berger, R.; Larsen, C.J.; Mathieu-Mahul, D. A third tal-1 promoter is specifically used in human T cell leukemias. J. Exp. Med. 1992, 176, 919–925. [Google Scholar] [CrossRef]
- Bernard, O.; Lecointe, N.; Jonveaux, P.; Souyri, M.; Mauchauffé, M.; Berger, R.; Larsen, C.J.; Mathieu-Mahul, D. Two site-specific deletions and t(1;14) translocation restricted to human T-cell acute leukemias disrupt the 5’ part of the tal-1 gene. Oncogene 1991, 6, 1477–1488. [Google Scholar]
- Elwood, N.J.; Green, A.R.; Melder, A.; Begley, C.G.; Nicola, N. The SCL protein displays cell-specific heterogeneity in size. Leukemia 1994, 8, 106–114. [Google Scholar]
- Lécuyer, E.; Herblot, S.; Saint-Denis, M.; Martin, R.; Begley, C.G.; Porcher, C.; Orkin, S.H.; Hoang, T. The SCL complex regulates c-kit expression in hematopoietic cells through functional interaction with Sp1. Blood 2002, 100, 2430–2440. [Google Scholar] [CrossRef]
- Yamada, Y.; Pannell, R.; Forster, A.; Rabbitts, T.H. The oncogenic LIM-only transcription factor Lmo2 regulates angiogenesis but not vasculogenesis in mice. Proc. Natl. Acad. Sci. USA 2000, 97, 320–324. [Google Scholar] [CrossRef] [PubMed]
- Kolodziej, S.; Kuvardina, O.N.; Oellerich, T.; Herglotz, J.; Backert, I.; Kohrs, N.; Buscato, E.; Wittmann, S.K.; Salinas-Riester, G.; Bonig, H.; et al. PADI4 acts as a coactivator of Tal1 by counteracting repressive histone arginine methylation. Nat. Commun. 2014, 5, 3995. [Google Scholar] [CrossRef]
- Ricci-Vitiani, L.; Pallini, R.; Biffoni, M.; Todaro, M.; Invernici, G.; Cenci, T.; Maira, G.; Parati, E.A.; Stassi, G.; Larocca, L.M.; et al. Tumour vascularization via endothelial differentiation of glioblastoma stem-like cells. Nature 2010, 468, 824–828. [Google Scholar] [CrossRef] [PubMed]
- Soda, Y.; Marumoto, T.; Friedmann-Morvinski, D.; Soda, M.; Liu, F.; Michiue, H.; Pastorino, S.; Yang, M.; Hoffman, R.M.; Kesari, S.; et al. Transdifferentiation of glioblastoma cells into vascular endothelial cells. Proc. Natl. Acad. Sci. USA 2011, 108, 4274–4280. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guelfi, S.; Orsetti, B.; Deleuze, V.; Rigau, V.; Bauchet, L.; Duffau, H.; Rothhut, B.; Hugnot, J.-P. SLUG and Truncated TAL1 Reduce Glioblastoma Stem Cell Growth Downstream of Notch1 and Define Distinct Vascular Subpopulations in Glioblastoma Multiforme. Cancers 2021, 13, 5393. https://doi.org/10.3390/cancers13215393
Guelfi S, Orsetti B, Deleuze V, Rigau V, Bauchet L, Duffau H, Rothhut B, Hugnot J-P. SLUG and Truncated TAL1 Reduce Glioblastoma Stem Cell Growth Downstream of Notch1 and Define Distinct Vascular Subpopulations in Glioblastoma Multiforme. Cancers. 2021; 13(21):5393. https://doi.org/10.3390/cancers13215393
Chicago/Turabian StyleGuelfi, Sophie, Béatrice Orsetti, Virginie Deleuze, Valérie Rigau, Luc Bauchet, Hugues Duffau, Bernard Rothhut, and Jean-Philippe Hugnot. 2021. "SLUG and Truncated TAL1 Reduce Glioblastoma Stem Cell Growth Downstream of Notch1 and Define Distinct Vascular Subpopulations in Glioblastoma Multiforme" Cancers 13, no. 21: 5393. https://doi.org/10.3390/cancers13215393
APA StyleGuelfi, S., Orsetti, B., Deleuze, V., Rigau, V., Bauchet, L., Duffau, H., Rothhut, B., & Hugnot, J.-P. (2021). SLUG and Truncated TAL1 Reduce Glioblastoma Stem Cell Growth Downstream of Notch1 and Define Distinct Vascular Subpopulations in Glioblastoma Multiforme. Cancers, 13(21), 5393. https://doi.org/10.3390/cancers13215393






