Simple Summary
The exposure of cancer cells to cadmium compounds may be associated with the acceleration of tumor progression. It is known that cadmium is a transcriptional regulator, and the study of differentially expressed genes has enabled the identification and classification of cadmium-associated molecular signatures as useful biomarkers that are potentially transferable to clinical research. This review recapitulates the studies that report the detection of such signatures in breast, gastric, colon, liver, lung, and nasopharyngeal tumor cell models, as specifically demonstrated by individual gene or whole genome expression profiling.
Abstract
The exposure of cancer cells to cadmium and its compounds is often associated with the development of more malignant phenotypes, thereby contributing to the acceleration of tumor progression. It is known that cadmium is a transcriptional regulator that induces molecular reprogramming, and therefore the study of differentially expressed genes has enabled the identification and classification of molecular signatures inherent in human neoplastic cells upon cadmium exposure as useful biomarkers that are potentially transferable to clinical research. This review recapitulates selected studies that report the detection of cadmium-associated signatures in breast, gastric, colon, liver, lung, and nasopharyngeal tumor cell models, as specifically demonstrated by individual gene or whole genome expression profiling. Where available, the molecular, biochemical, and/or physiological aspects associated with the targeted gene activation or silencing in the discussed cell models are also outlined.
1. Introduction: A Short Excursus on Cadmium and Eukaryotic Cell Systems
Cadmium (Cd) is an underground mineral extracted as part of zinc deposits which, along with its compounds, exhibits a broad range of applications in the industry spanning from battery components to stabilizers for plastics, electroplated coatings for non-ferrous metals and dye pigments. Volcanic eruption and the erosion of Cd-containing rocks represent the major natural sources of the metal; on the other hand, humans are mainly exposed to Cd via cigarette smoking and, to a lesser extent, food ingestion, inhalation of polluted air, and the ingestion of contaminated drinking water [1].
Cd is not essential for the human body and does not exert any useful cellular metabolic effect. The cellular uptake of free Cd ions and complexes of Cd with small organic molecules involves various ion channels, carriers, and ATPase pumps, whereas Cd-protein complexes are internalized via receptor-mediated endocytosis, as extensively reviewed by the authors in [2]. Once accumulated in the intracellular milieu, the metal may interact with the thiol groups of cysteines present in cellular proteins, thus inducing an impairment of the functions of the enzymes located in different cellular compartments. The numerous biological targets and complex mechanisms of the action of Cd ions have been explored in depth in previously-published reviews [3,4,5]. It is known that Cd can cause mitochondrial dysfunction, exert genotoxic and epigenotoxic effects, and can also interfere with a number of cell proliferation-linked signalization pathways. Reports in the literature have shown that mitochondria are one of the main intracellular targets for Cd toxicity due to the blocking of the electron transfer chain at complex III, which is considered the principal site for the production of reactive oxygen species (ROS), thereby favoring the over-production of the latter at the expense of ATP. This leads to a dissipation of mitochondrial transmembrane potential with a consequent alteration of not only the cell’s redox status but also mitochondrial and nuclear gene expression and the genome integrity [6,7]. In fact, it is widely acknowledged that the genome-related effects of Cd are based upon its ability to damage the antioxidant defense and DNA repair systems, and also due to the replacement of zinc by Cd in p53 with a consequent impairment of protein activity, thereby triggering the occurrence of DNA strand breaks and chromosomal aberrations [8,9]. The inhibition of p53, coupled with the direct stimulation of mitogenic signals via calcium and inositol triphosphate second messengers, is also responsible for Cd-dependent enhanced cell proliferation. On the other hand, in specific cell systems the metal was conversely found to induce endoplasmic reticulum stress due to severe ROS and calcium signaling, which ultimately leads to apoptotic and/or autophagic cell death [10,11,12,13]. As a further DNA-directed activity, evidence has been produced that Cd may be regarded as an epigenetic modulator, given its ability to alter the global DNA methylation pattern via the down-regulation of DNA methyltransferase activity and demethylation which, upon prolonged exposure, turns into enhanced enzyme activity and genome hypermethylation. These opposite alterations may result in the up-regulation of cellular protooncogenes and the silencing of oncosuppressor genes, respectively [14,15].
Evidence collected in the literature has highlighted that the exposure of cancer cells to Cd compounds may be associated with the development of more malignant phenotypes, thereby contributing not only to the onset of cell transformation in the tumor initiation stage, which has been widely reviewed in several articles e.g., in [16,17], but also to the acceleration of tumor progression. An example of this aspect is the growth-promoting “metalloestrogen” role played by Cd on estrogen receptor α-positive breast cancer cells. On the other hand, based upon the intrinsic cytospecificity, Cd has been proven to act as a two-edged sword, conversely down-regulating viability and causing the death of a number of tumor cell lines, such as those derived from triple-negative breast cancer and hepatocarcinoma [18,19,20]. Cd is a transcriptional regulator that induces molecular reprogramming [21]; therefore, the study of differentially expressed genes has enabled the identification and classification of the molecular signatures inherent in human neoplastic cells upon Cd exposure as useful biomarkers that are potentially transferable to clinical research. The goal of this review is to recapitulate selected studies that report the detection of such signatures, as specifically demonstrated by individual gene or whole genome expression profiling, in particular focusing on breast, gastric, colon, liver, lung and nasopharyngeal tumor cell models. Where available, the molecular, biochemical and/or physiological aspects associated with the targeted gene activation or silencing in the discussed cell models will be also briefly outlined.
2. Molecular Signatures in Breast Cancer Cells
The search for Cd exposure-associated molecular signatures has been conducted in two cell lines, i.e., MDA-MB231 and MCF-7. The former was established from a pleural effusion of a triple-negative breast cancer (TNBC) of basal morphology, which was negative for estrogen receptor (ER) α and progesterone receptor (PR) expression and its p53 was inactivated by a mutation in codon 280 of exon 8. The ERα-positive and PR-negative MCF-7 cell line was established from a pleural effusion of a highly hormone-responsive malignant adenocarcinoma [22,23,24].
2.1. Molecular Signatures in MDA-MB231 Cells
Concerning TNBC cells, about a decade ago papers published by my group of researchers first contributed to this issue by investigating the effect of administering different concentrations of CdCl2 to MDA-MB231 cells and also comparing the tumor cell behavior with that of HB2 cells, a nonneoplastic immortalized line from the human breast epithelium [25]. The obtained results demonstrated the cytotoxic effect of the molecule with a 50% inhibitory concentration (IC50) at 96 h of 5 μM, which conversely was ineffective in modifying the survival and growth of HB2 cells [18]. Among the biological aspects of MDA-MB231 cells studied under this experimental condition, molecular signatures were searched that demonstrated an up- or down-regulation of the expression levels of genes coding for heat shock proteins (hsp), metallothioneins (MT), cytochrome oxidase subunits, and other factors related to apoptosis, signal transduction, and growth control, as summarized sinoptically in Table 1 (see refs. [18,26,27,28,29,30]).

Table 1.
Gene expression changes in MDA-MB231 breast cancer cells exposed to 5mM CdCl2 for 96 h.
In light of the literature on the subject, some of the produced data allow for the following comments to be made.
2.1.1. Expression Levels of MTs
MTs are a group of cysteine-rich, low molecular weight proteins able to bind metals and act as controllers of cellular homeostasis through protection against oxidative stress, heavy metal toxicity, and DNA damages. The exposure of MDA-MB231 cells to CdCl2 resulted in the up-regulation of MTIA and MTIF whilst MTIG was down-regulated. It is noteworthy that MTIA mRNA and proteins were proven to be induced by the exposure of human proximal tubule cells to Cd concentrations, eliciting a downfall of cell viability [31]. In addition, MTIA overexpression provided HEK293 embryonic kidney cells with more resistance to the Cd administration than other MT isoforms [32]. Taking the data from the literature into account, it is conceivable that the up-regulation of this MT isoform in the MDA-MB231 cells may represent a cellular defense reaction specifically set up against Cd intoxication. Interestingly, Chang et al. [33] validated MTIA mRNA expression as a molecular marker for renal dysfunction in occupational Cd exposure. Concerning the other two MTs, Albrecht et al. [34] reported a significant increase in the MT1F transcripts in RWPE-1 human prostate epithelial cells at the time points immediately preceding cell death due to exposure to 3, 6 and 12 mM Cd2+ over a 13-day period, which may be consistent with the cytotoxic effect exerted on MDA-MB231 cells. Moreover, the down-regulation of MTIG, whose reduction was found to induce glutathione depletion and lipid peroxidation in hepatocellular carcinoma cells [35], may be a contributing factor to MDA-MB231 cell death.
2.1.2. Expression Levels of Heat Shock Proteins
The up-regulation of HSPB1 and the down-regulation of HSPA5, HSPD1, HSP90AB1 and TRAP1 in CdCl2-treated MDA-MB231 cells has been demonstrated. It is known that oxidative damage and ROS generation may selectively trigger a significant decrease in the HSPA5 levels [36] and therefore these data are consistent with the findings of a study into the increase in mitochondrial respiratory activity and the accumulation of ROS in treated MDA-MB231 cells, as shown by Cannino et al. [27]. Moreover, the down-regulation of HSPD1 and TRAP1, coding for two mitochondrial chaperonins, could be responsible for the accumulation of ROS as reported by the authors in [37,38], whereas the down-regulation of HSPA8 may contribute to the decrease in cell protection against ROS-mediated lipid peroxidation during the oxidative challenge. Interestingly, Masuda et al. [39] suggested that Bcl-2 may be one of the regulators of TRAP1 expression, which is consistent with the data demonstrating the concomitant down-regulation of BCL2 and TRAP1 in exposed MDA-MB231 cells. Exposure to 10 mM CdCl2 for 24 h was found to be responsible for the decrease in the expression level of HSP90AB1, contributing to the metal-promoted lethal effect on cholinergic neurons [40]. Kindas-Mugge et al. [41] demonstrated the decrease in the proliferation rate of HSBP1-overexpressing MDA-MB231 cells. It is conceivable that the restraining effect of hsp27, the product of HSBP1, on cell growth behavior could be jointly responsible with the induction of cell death for the halving of the MDA-MB231 cell numbers observed after 96 h-treatment with 5 µM CdCl2.
2.1.3. Expression Levels of Cytochrome Oxidase Components
The exposure of MDA-MB231 cells to CdCl2 resulted in the down-regulation of COX2 and COX4, thereby suggesting a relationship between the reduction in the amount of COX and the production of ROS, given that mitochondrial proteins are some of the earliest cellular targets because of their immediate proximity to the ROS-generating sources [27].
2.1.4. Expression Levels of Proliferation- and Apoptosis-Related Genes
The exposure of MDA-MB231 cells to CdCl2 determined the down-regulation of BCL2 and AEG1 and the up-regulation of DAPK, WAF1, RIPK, PLP2, FOS, JUN, and several genes coding for caspases, the enzymatic components of the apoptotic machinery. This result suggests that the metal directs the tumor cells towards a type of death which shares several features with programmed cell death. In fact, the Bcl-2 protein is an apoptosis-suppressor factor, whereas death-associated protein kinase-1 is involved in the onset of the death pathways of apoptosis, as well as autophagy and programmed necrosis. In MDA-MB231 cells, cyclin-dependent kinase inhibitor 1 was proven to negatively modulate the cell cycle progression due to binding to both the cyclin–CDK complex and proliferating the cell nuclear antigen, and the RIPK1 product, a kinase forming part of the tumor necrosis factor receptor-1 (TNFR1) complex I, which is known to be one of the most important upstream mediators of NF-κB signaling as well as an important regulator of cell death [42,43,44]. In breast cancer, the astrocyte elevated gene-1 protein was found to be an agonist of the Wnt pathway which regulates cell proliferation and is involved in the control of the NF-kB pathway and the expression of proliferation-promoting FOS/JUN genes [29,45]. The knockdown of endogenous AEG1 was proven to sensitize MDA-MB-231 cells to TRAIL-induced apoptosis both in vitro and in vivo. Therefore, in line with the data of Zhang et al. [46], the possibility that CdCl2-mediated AEG1 down-regulation may facilitate the intrinsic and extrinsic apoptosis pathways via the decrease in BCL2 expression levels and CASP8 up-regulation could be considered. In addition, PLP2 over-expression was also found to be correlated with the increase in CASP8 expression levels [30].
2.1.5. Expression Levels of MAPKs
CdCl2 treatment on MDA-MB231 cells was efficient in decreasing the expression levels of MAPK11 and −14 and increasing that of MAPK12, although an opposite variation in the accumulation of the protein product of the latter, i.e., mitogen-activated protein kinase p38γ was recorded. Zhong et al. [47] demonstrated that in case of oxidative stress, the activation of the MAPK11/12/13/14/NF-kB pathway may induce protective autophagy via the transcriptional activation of the autophagy-related genes MAP1LC3B, BAG3, and HSPA1A in HeLa cells treated with an anticancer copper complex. Therefore, it might be assumed that MAPK down-regulation following the exposure of breast cancer cells to CdCl2 may contribute to the cytotoxic effect through the inhibition of such a protective cell response.
More recently, additional Cd-related molecular signatures in MDA-MB231 cells have been brought to light. Wei et al. [48,49] reported that non-cytotoxic concentrations of CdCl2 (0.5–1 µM) trigger cell migration, an ability which in vivo can result in an enhanced metastatic attitude during TNBC progression. Using luciferase reporter assays, after prolonged Cd treatment (3 µM for 4 weeks), they demonstrated the up-regulation of the transcriptional activity of the T-cell factor/lymphoid enhancer-binding factor (TCF/LEF), whose contribution to the Wnt signaling machinery is well-known, and Snail, a target of the Notch1 pathway primarily involved in epithelial-mesenchymal transition (EMT) processes as demonstrated by knockdown experiments [50,51]. The effect on the TCF/LEF appeared to be due to Cd-induced signaling via the interaction with the integrin/focal adhesion kinase (FAK) pathway leading to the inhibition of GSK3β and consequent β-catenin accumulation and nuclear translocation. On the other hand, Cd induced the elevation of Snail transcription by up-regulating the activity of its promoter. Moreover, their results also suggested that Cd may inhibit miR-200 that post-transcriptionally restrains the translation of ZEB-1 which, in turn, is a direct suppressor of the transcription of the gene encoding the epithelial marker E-cadherin, thus further contributing to the regulation of EMT [52].
2.2. Molecular Signatures in MCF-7 Cells
Concerning the ERα-positive MCF-7 cell line, in 2013 Lubovac-Pilav et al. [53] investigated the effect of chronic (up to 6 months) 10−7 M CdCl2 exposure on the gene expression pattern by high throughput microarray technology followed by hierarchical clustering analysis and functional annotations. More recently, Liang et al. [54] complemented the analysis by producing data on the epigenomic as well as the transcriptomic profiles induced by short-term (72 h) 60 µM CdCl2 treatment, further identifying also in this case the critical pathways and genes via bioinformatic studies. Their cumulative results provide a broad picture of the impact of the exposure of breast cancer cells to the metal compound at the gene signature level, with 795 differentially expressed genes identified by the authors in [53] and 997 differentially expressed genes identified by the authors in [54]. In particular, taking their most significant findings on the highest changes in gene expression levels and the data from the literature into account, the following comments can be made.
2.2.1. Expression Levels of Breast Cancer-Associated Factors
Ninety-seven genes identified by the authors in [53] were known to be breast cancer-associated, thereby confirming that Cd is active in accelerating breast carcinogenesis by modulating the expression of genes involved in tumor progression. Table 2 shows those breast carcinogenesis-associated genes positioned among the top 30 up- and down-regulated ones upon chronic metal exposure, all of which were also linked to endocrine disruption effects, while some of them were also linked to the regulation of cell proliferation and differentiation (e.g., CRABP1 and CCNE1) and ROS production (e.g., UCP2). The identification of gene signatures that were significantly regulated by short-term CdCl2 exposure and strongly associated to breast carcinogenesis through the protein-protein network analysis led Liang et al. [54] to extract TXNRD1 and CCT3 from the list of 400 validated genes as those endowed with the highest degree of expression change, hazard ratio difference, and degree of connectivity. The first gene codes for thioredoxin reductase 1, an intracellular redox sensor and antioxidant enzyme whose deregulation may be involved in breast cancer initiation [55]. CCT3 protein product is the chaperonin containing TCP1 subunit 3, a component of a complex which catalyses the correct folding of proteins involved in cytoskeletal assembly and the cell cycle and whose involvement in the promotion of breast cancer cell proliferation and metastasis via Wnt has been recently acknowledged [56,57]. It is also of note that one of the most significant Gene Ontology (GO) terms in which a large amount of the genes validated by Liang et al. [54] were enriched was the “Wnt signaling pathway”, which is known to be highly dysregulated in breast tumor [58].

Table 2.
Breast carcinogenesis-associated gene signatures in MCF-7 cells submitted to chronic CdCl2 exposure [53].
2.2.2. Expression Levels of Metal Ion- and Xenobiotic-Binding Factors
The chronic exposure of MCF-7 cells to CdCl2 was proven to affect the expression level of the genes coding for not only Cd- but also calcium-, zinc- and iron-binding proteins due to the mimesis and interference played by Cd towards the other metals. Table 3 shows those genes positioned among the top 30 up- and down-regulated ones in the “GO molecular functions: metal ions” category. More specifically, one of them was subcategorized as “cadmium ion binding” (MT1F), five were subcategorized as “calcium ion binding” (ANXA3, ANXA2P2, ATP2A3, FKBP9 and OCM), one was subcategorized as both (MT1X), and four were subcategorized as “zinc ion binding” (CBFA2T3, MT2A, PDLIM1 and ZNHIT2). This was consistent with the ability of Cd to mimic other divalent cations thus stimulating the expression of metal transporters. The up-regulation of MT1 expression after the short-term exposure of MCF-7 cells to Cd was also reported by Darwish et al. [59] who also demonstrated the concurrent over-expression of MDR1 and MRP2 genes, coding for the xenobiotic transporters multidrug resistance-associated protein 1 and 2, respectively, which are known to also be involved in Cd efflux as a cytoprotective reaction to cell intoxication [60].

Table 3.
Metal ion binding-associated gene signatures in the MCF-7 cells submitted to chronic CdCl2 exposure [53].
2.2.3. Expression Levels of Cell Growth-Associated Factors
Another GO category which was analyzed in chronically CdCl2-intoxicated MCF-7 cells was “GO biological process: cell growth” [54]. Two were the genes found among the top 30 up-regulated ones, that is, cyclin E1-coding CCNE1 which was subcategorized under the GO terms “cell division” and “cell cycle”, and the zinc ion transporter-coding CBFA2T3 included in the “cell proliferation” subcategory. It is known that cyclin E1 is an oncogene which controls the G1/S phase transition and its expression is directly related to the aggressive potential of breast cancer [61]; on the other hand, the association of metal treatment on breast tumor cells and CBFA2T3 over-expression is still undetermined. In addition, consistent with the Cd-induced positive effect on breast tumor cell growth, Siewit et al. [62] demonstrated the up-regulation of CCND1 and MYC, coding for cyclin D1 and c-myc proteins, and the down-regulation of CDKN1A, coding for the p21/WAF1 protein, after 24 h of MCF-7 cell exposure to micromolar CdCl2 concentrations.
2.2.4. Expression Levels of Methyltransferases
Following MCF-7 cell treatment with 0.1, 0.5 and 1 mM CdCl2 for 24 h and 48 h, Ghosh et al. [63] reported the Cd dose-dependent increase in the expression of the PRMT5 and EZH2 genes coding for the proteins arginine methyltransferase 5 and histone-lysine N-methyltransferase enhancer of zeste homolog 2, respectively, which are two critical epigenetic modulators responsible for the development of multiple cancer histotypes. The experimental data collected suggested that such expressional modulations could be due to CpG island demethylation in the gene promoter sites as a positive feedback loop of the Cd-induced down-regulation of methyltransferase activity, and/or the Cd-promoted increase in the levels of NFYA and E2F1 transcription factors and their subsequent enriched recruitment to the demethylated gene promoters.
2.2.5. Expression Levels of Heat Shock Proteins
The dose-dependent up-regulation of the HSPA2 gene, coding for hsp70 protein, was reported by Darwish et al. [59] in short-term (24 h) Cd-treated MCF-7 cells (10 and 100 µM concentrations) and was associated with the role of this molecular chaperone in preventing protein degradation under stress conditions.
2.2.6. Expression Levels of Antioxidant System and Inflammatory Markers
In the same paper, Darwish et al. [59] demonstrated the drastic dose-dependent down-regulation of some genes whose products are involved in the protection against oxidative damage, i.e., glutathione S-transferase omega 1, NAD(P)H quinone dehydrogenase 1, superoxide dismutase 1 and 2, and catalase, and the up-regulation of HO-1, coding for the antioxidant heme oxigenase 1 enzyme, which is known to be triggered by Cd in MCF-7 cells via the p38 kinase pathway and via Nrf2 [64]. Moreover, they found that Cd determined a significant dose-dependent accumulation of the mRNAs of cyclooxygenase-2, the tumor necrosis factor-α, and interleukin 8 and 10, whose induction is conceivably linked to inflammation and cell damage in various organs. Interestingly, a reduction in the expression of these genes and a concurrent increase in the expression of the genes for metallothionein-1 and the xenobiotic transporters was obtained by the co-exposure of MCF-7 cells with both Cd and fat-soluble vitamins (mainly D, whereas A and E to a lesser degree). This confirmed the cytoprotective effects of the latter which make them a dietary supplement worth investigating for people at high risk of exposure to Cd.
3. Molecular Signatures in Cancer Cells of the Gastrointestinal Tract
3.1. Molecular Signatures in Gastric Cancer Cells
The search for Cd exposure-associated molecular signatures has been conducted in cell lines isolated from gastric adenocarcinoma, i.e., MKN28 (well-differentiated), SNU638 (poorly differentiated with the mutated p53 gene) and AGS, the latter characterized by gene expressions typical of tumors that likely process from intestinal metaplasia [65,66]. In these cell lines Khoi et al. [67] examined the effects of a 4 h-treatment with 20 µM Cd on the expression levels of the PLAUR gene, coding for urokinase-plasminogen activator receptor (uPAR), demonstrating its time-dependent up-regulation. A more detailed investigation on the sole Cd-treated AGS cells brought evidence that PLAUR over-expression was to be ascribed to the activation of the ERK1/2-NFkB-AP1 pathway and that uPAR accumulation was a likely mediator of Cd-induced stimulation of cell invasiveness.
3.2. Molecular Signatures in Colon Cancer Cells
The search for Cd exposure-associated molecular signatures has been conducted in the poorly differentiated RKO cell line, which bears mutations in BRAF and PIK3CA oncogenes, and the HT-29 cell line, isolated from a primary colon adenocarcinoma [68,69].
3.2.1. Molecular Signatures in RKO Cells
The subchronic low-dose exposure of RKO cells to Cd (50 µM for 24 h) [70] resulted in the differential expression of twenty genes, most of which belong to the hsp-coding family and are listed in Table 4, that can be considered as potential carcinogenesis-linked genes under the experimental conditions used. Among the other genes shown to undergo expression alterations, some were related to the detoxication of chemical carcinogens (AKR1C2), cell survival (DCD, IDH1), migration-promoting EMT transition (P4HB), and colon cancer metastasis (PGK1) [71,72,73].

Table 4.
Differentially expressed hsp-coding genes in RKO colon carcinoma cells submitted to subchronic low-dose CdCl2 exposure [70].
3.2.2. Molecular Signatures in HT-29 Cells
More recently, dealing with the cyclooxygenase inflammatory pathway, a study by Naji et al. [74] using the luciferase reporter assay demonstrated the early (6 h) and time-dependent 100 nM CdCl2-driven increase in COX2 expression at the transcriptional level in HT-29 colon adenocarcinoma cells. It is widely acknowledged that the cyclooxygenase pathway is involved in colorectal malignant progression, and also promotes the migration of tumor cells via the activation of the ROS-p38-COX-2-PGE2 and the ROS-Akt pathways. Therefore, the up-regulation of the inducible COX-2 isoform may be considered as one of the mechanisms by which exposure to environmental Cd pollutants maximizes colorectal malignancy in exposed individuals. In addition, in cells treated with 0.05–10 µM CdCl2 for 24 h, Iftode et al. [75] reported the suppression of the expression of DNMT1 and DNMT3B genes, coding for DNA methyltransferase-1 and -3β and which act as hypomethylating agents likely responsible for the carcinogenetic properties of Cd.
4. Molecular Signatures in Liver Cancer Cells
The search for Cd exposure-associated molecular signatures has been conducted in HepG2 cells, isolated from a differentiated hepatocellular carcinoma.
Fabbri et al. [76] and Urani et al. [77] performed a whole-genome analysis by cDNA microarray after 24 h exposure of this cell line to 2 and 10 µM CdCl2, considered to be low human-relevant metal concentrations. In particular, the work of Urani and coworkers has also focused on intracellular zinc displacement by Cd and its molecular consequences, as zinc is an acknowledged second messenger and transcriptional regulator. Their investigation into gene expression profiling was complemented by a miRNA expression analysis, due to the considerable involvement of their altered regulation in the carcinogenetic process. A group of eleven genes, all belonging to the metallothionein-coding family (i.e., MT1A, -B, -E, -F, -G, -H, -JP, -L, -M, -X and -2A) were found up-regulated after exposure to CdCl2 concentrations whereas the down-regulation of 12 miRNAs and the differential-expression of 949 genes was proven in response to the treatment with the sole higher metal concentration. In particular, the down-regulation affected the genes involved in the liver function pathways (e.g., fatty acid/cholesterol metabolism and hemostasis) while the up-regulation concerned the genes involved in inflammation and cancer progression (e.g., cytokine-cytokine receptor interaction-, focal adhesion- and MAPK signaling). A subset of relevant genes was further submitted to validate their differential expression through real time-PCR (Table 5).

Table 5.
PCR-validated up-regulated genes in HepG2 hepatocarcinoma cells exposed to 10 µM CdCl2 exposure [76,77].
An analysis of the KEGG database indicated that most validated genes (CAPN2, COL1A1, ITGA2, ITGA3, ITGB1, JUN and LAMB3) were associated with focal adhesions whose regulation is involved in liver cancer cell invasion and metastasis, in association with the dysregulation of the cytoskeletal component [78]. Interestingly, Urani and coworkers also found the up-regulation of other adherent junction pathway-related genes, i.e., SNAI1, MET, TGFBR2, RAC and CDC42, thus confirming the facilitating role of Cd in cell motility and metastatization processes. The other validated genes, i.e., FOS, HSPA6 and GADD45B, were associated to the MAP kinase pathway, which is known to play a role in hepatocarcinoma cell survival and tumor growth [79]. It is also of note that the two top pathways of the Cd-dependent dysregulated miRNAs were related to focal adhesions and the MAP kinase cascade.
Distinct sets of 330 and 181 genes were found impaired by the authors in [80] after acute and chronic low concentration-CdCl2 treatment of HepG2 cells, respectively. The majority of them were involved in detoxification and metabolic processes in the exposure conditions in the former, and in the regulation of various signaling pathways including the inflammatory and insulin responses in the latter. A subset of relevant genes was further submitted to validate their differential expression through real time-PCR (Table 6).

Table 6.
PCR-validated differentially expressed genes in HepG2 hepatocarcinoma cells exposed to 0.5 µM acute or 0.1 µM chronic CdCl2 treatment [80].
Under acute exposure to the metal compound, as expected for cells undergoing detoxification, the up-regulation of some members of the metallothionein gene family were found, conceivably driven, at least in part, by the down-regulation of SPINK1 as reported for colon cancer cells [81]. Down-regulation was also observed for ecto-5′-nucleotidase (a.k.a. CD73), the major enzyme responsible for the enzymatic dephosphorylation of the inflammation-promoting extracellular adenosine 5′-monophosphate nucleotide to the immunosuppressive adenosine. Conceivably, such down-regulation might be an indicator of a pro-inflammatory activity [82]. Interestingly, CYP3A7, whose product is involved in drug metabolism, was either down- or up-regulated following acute or chronic exposure, respectively, and this might be associated to the Cd cytotoxic insult in the former and the onset of a detoxification reaction by stressed cells in the latter [83,84]. Following chronic treatment, the up-regulation of the extracellular matrix protein nephronectin may complement the down-regulation of NT5E and be related to the stimulation of an inflammatory reaction, according to the evidence of protein localization in inflammatory foci in animal models of hepatitis [85]. Other validated differentially-expressed genes were: (i) DNAJB9, whose up-regulation may be associated with the inhibition of p53-induced senescence leading to cell mitogenic signalization and transformation; (ii) ADH4, whose lowered expression is linked to the stimulation of several cancer related pathways, including ATR, FOXM1, FOXO, MTOR, NOTCH, and the p53 downstream pathway; (iii) EGR1 coding for an anti-tumorigenic zinc-finger transcription factor; and (iv) ID1 whose down-regulation may impair the cell redox state by overproduction of ROS [86,87,88,89].
By comparison with MCF-7 breast tumor cells, 48 h-exposure to Cd also up-regulated the expression of PRMT5 and EZH2 methyltransferase genes in HepG2 cells, albeit to a lesser extent, as evidenced by the luciferase reporter assays, thereby confirming the impact of the metal on the expression of the two oncogenic epigenetic regulators also in this neoplastic cytotype [63].
5. Molecular Signatures in Lung Cancer Cells
The search for Cd exposure-associated molecular signatures has been conducted in the following cell lines isolated from non-small cell lung carcinoma tissues: (i) H460 characterized by mutant K-ras and wild-type p53 [90,91] and its Cd-resistant derivative RH460 established after selection via the exposure of the parental line to increasing Cd concentrations [92]; (ii) H1299 established from a lymph node metastasis of the lung from a patient who had received prior radiation therapy and endowed with a homozygous partial p53 deletion determining the lack of protein expression; and (iii) A549 characterized by mutant K-ras, wild-type EGFR and the properties of type II alveolar epithelial cells [93].
5.1. Molecular Signatures in H460 and RH460 Cells
Kim et al. [94] studied the induction of the multidrug resistance-associated protein 1 (MRP1), coded by the ABCC1 gene, by Cd treatment in responsive H460 vs. resistant RH460 cells. This protein is a multitasking transporter broadly involved in many aspects of cell biology and pathology spanning from cell survival and differentiation to inflammation and cancer [95]. The acquired Cd resistance of RH460 cells determines the absence of Cd-induced apoptosis and autophagy, occurring in the parental line, due to the lack of glycogen synthase kinase (GSK)-3β phosphorylation at serine residue and the consequent intracellular relocalization of the molecule [96]. Using inhibitors and siRNAs against MRP1 and GSK-3β and overexpressing GSK-3β-HA, Kim et al. [94] revealed the role played by the kinase in the modulation of the expression of the MRP1 molecular signature through both transcriptional regulation and direct interaction with p-Ser GSK3αβ which intervenes in MRP1 stabilization and intracellular redistribution. The obtained data represent a promising tool for the formulation of GSK-3β serine phosphorylation-inducing chemotherapeutics aimed to treat multidrug resistant lung tumors.
5.2. Molecular Signatures in H1299 Cells
It is known that various metals, including Cd, interfere with the localization, folding and function of members of the p53 protein family [97]. Within this context, Adámik et al. [98] examined whether CdCl2 impaired the function of p63 and p73 as transcription factors. To this purpose, the different p53 family isoforms were co-transfected with p53 family-dependent luciferase reporter vectors (pGL3-MDM2-APP, pGL3-PGM1 and pGL3-BAX) into p53-deficient H1299 cells. The obtained data demonstrated that the p63 and p73 transactivation of some of the p53-dependent promoters was inhibited by exposure to 20–50 mM CdCl2. This was also demonstrated in light of the data confirming the impairment of the binding of the factors’ core domains to p53 consensus sequences, as revealed by electrophoretic mobility shift assays. Thus, the sensitivity of p53 family proteins to Cd appears to be conserved and also active in cell-based assays and, conceivably, the resulting modulation of gene signature expression may play a central role in metal carcinogenesis.
5.3. Molecular Signatures in A549 Cells
Fujiki et al. [99] submitted A549 lung tumor cells to prolonged 20 mM CdCl2 exposure and observed the onset of a high proliferative rate, EMT, stress fiber formation, cell locomotion, and resistance to antitumor drugs. From a molecular point of view, the involvement of Notch1 signalization in the Cd-promoted malignant progression was demonstrated, and which was maintained for a considerable time after the removal of CdCl2 from the culture medium. In particular, the cell treatment was associated with the up-regulation of NOTCH1, JAG2, coding for Notch-ligand Jagged-2 protein, and MMP2, coding for matrix metalloprotease 2, an invasion-facilitating collagenase which is a prognostic factor for non-small cell lung cancer [100]. On the other hand, the down-regulation of NOTCH3 was a further gene signature of cell exposure to CdCl2. The cumulative results obtained indicated that the transcriptional activity of Notch1 was stimulated by hypoxia-inducible factor 1 (HIF-1) and that the insulin-like growth factor 1 receptor (IGF-1R)/Akt/ERK/p70 S6 kinase 1 (S6K1) cascade could cooperate with Notch1 signaling and HIF-1 following the CdCl2 treatment of A549 cells.
6. Molecular Signatures in Nasal Septum and Nasopharyngeal Cancer Cells
Since inhalation is the primary route of Cd intake, mainly by cigarette smoking and also by occupational exposure during working activities related to nickel–cadmium batteries, electroplating, and paint pigments, RPMI-2650, CNE-1 and CNE-2 cell lines were used as an in vitro model system to examine the molecular targets of Cd-induced cancer progression in nasal and nasopharyngeal epithelia.
6.1. Molecular Signatures in RPMI-2650 Cells
The RPMI-2650 line was isolated from the pleural effusion of a patient with an anaplastic squamous carcinoma of the nasal septum [101]. In the light of the observed Cd-dependent up-regulation of the intracellular ROS level and the down-regulation of cell proliferation, Lee et al. [102] compared the mRNA expression patterns of RPMI-2650 cells grown in control conditions or exposed to 0.75 µM Cd acetate for 72 h via differential display analysis. Following a preliminary analysis of gene expression, AKR1C3, coding for the aldo-keto reductase family 1 member C3 protein, was proven to undergo an increase in expression, regulated by Cd at the transcription/translation level; this up-regulation was also confirmed by Western blot analysis. This molecular signature is a hormone activity regulator and prostaglandin F synthase is responsible for monitoring the occupancy of hormone receptors and controlling cell proliferation and differentiation in a hormone-independent way [103]. Based on the reported Cd-induced accumulation of the Nrf2 transcription factor in the nucleoplasm and the restraining of the Cd-dependent increase in the AKR1C3 protein levels by a phosphoinositide 3-kinase (PI3K) inhibitor, it was suggested that the up-regulation of AKR1C3 may result from the augmentation of intracellular ROS, at least in part through the activation of Nrf2 and the onset of PI3K-related signalization, thereby contributing to an adaptive intracellular response to Cd cytotoxicity.
6.2. Molecular Signatures in CNE-1 and CNE-2 Cells
CNE-1 and CNE-2 cell lines were established from a well-differentiated and a poorly differentiated nasopharyngeal squamous carcinoma, respectively, a highly invasive and metastatic malignant tumor with unique ethnic and geographic distribution and prominent incidences in South China and some African areas only [104,105]. With the aim to mimic chronic low-level Cd exposure, Peng et al. [106] exposed both cell lines to a non-toxic CdCl2 concentration (1 µM) for up to two weeks and observed the acquisition of more proliferative and aggressive characteristics by the cells both in vitro and in vivo. Molecular analyses revealed that chronic Cd treatment induced a remarkable up-regulation of CCND1, CCNE1, MYC and JUN, thereby demonstrating the activation of Wnt/β-catenin signaling, which is also in the parallel confirmatory data on increased β-catenin protein immunostaining in Western blots. Further studies highlighted that chronic Cd exposure induced the down-regulation of CSNK1A1, coding for the α isoform of casein kinase I, via the hypermethylation of the promoter CpG islands. On the other hand, given that this enzyme is a negative regulator of the Wnt/β-catenin pathway [107], this molecular event might be involved, at least in part, in the exacerbation of the malignant phenotype by neoplastic nasopharyngeal cells.
7. Conclusions
Malignant growth is a multistep process driven by accumulating genetic alterations that progressively lead the cells to acquire novel abilities such as self-sufficiency in growth signals, insensitivity to growth-inhibitory signals, evasion of apoptosis, limitless replicative potential, sustained angiogenesis, tissue-invading and metastasizing attitude, oncopromoting inflammatory ability, energy metabolism reprogramming and immunoevasion (the “hallmarks of cancer “ [108,109]).
Since the establishment of the HeLa cell line in 1951, human tumor cells in culture have represented the most extensively used model system to study cancer biology providing the wealth of information that we currently possess about the molecular, biochemical, and genetic aspects of oncodevelopment and the modes of action of potentially anti-cancer compounds. On the other hand, the clinical relevance and usefulness of these in vitro models is still controversial, mostly due to the lack of interactions with other cell types and the signaling and structural molecule-containing extracellular microenvironments which influence the action of drugs in vivo. Nevertheless, the growing body of data on the molecular profiles and biological characterizations arising from established and novel in vitro preclinical models cannot be ignored, thereby representing a highly useful resource for the testing of scientifical hypotheses focused on the elucidation of carcinogenetic mechanisms and the initial assessment of cancer treatments [110,111].
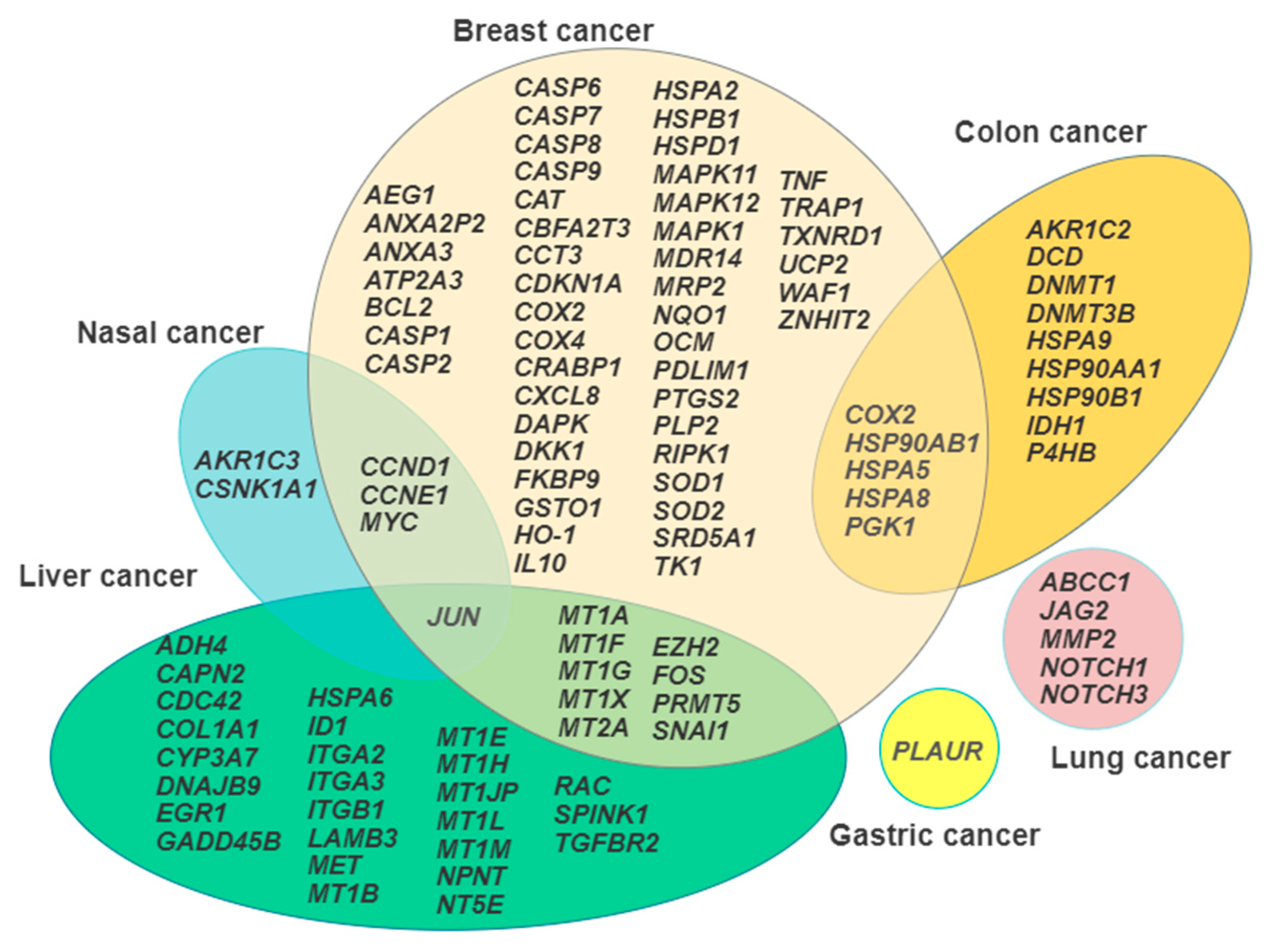
It is generally accepted that the knowledge of the repertoire of genes transcribed under a given circumstance, such as the exposure to environmental pollutants, may aid in the dissection of the mode of action of the toxicants under investigation [112]. This review has pointed out that different neoplastic cytotypes undergo a wide range of Cd-induced changes in their transcriptional profiles which can consequently influence protein expression, signal transduction, and cell metabolism, and ultimately tie in with the previously listed traits of the carcinogenetic process. Figure 1 summarizes in a Venn diagram the representation the gene signatures discussed, showing the areas of overlap between the different cancer cell models, which is a useful procedure to select the most promising candidates for further translation in in vivo studies.
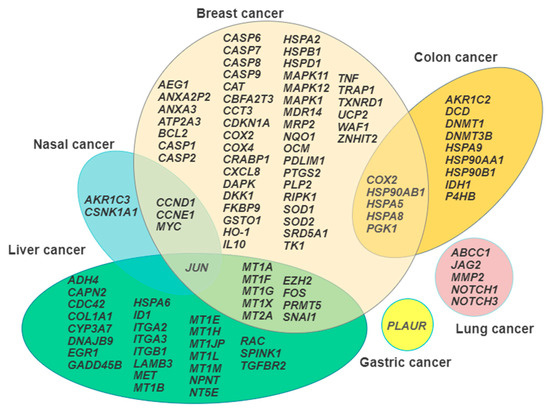
Figure 1.
Venn diagram illustrating the gene signature arrays in each cancer cell model examined and the overlapping areas among the model systems.
The individuation of single or panels of novel expression markers can provide a broader picture of the numerous and complex effects exerted by the metal and, in addition, has potential as alternatives to traditional biomarkers for the efficient assessment of the tumor progression stages in diverse cancers. On the other hand, descriptive epidemiological studies can certainly take advantage from detailed studies at the molecular level, which provide new avenues for analytic research. The Cd levels in blood, urine, feces, liver, kidney, hair, and other tissues can be easily measured and can provide an indication of recent or total exposure to Cd, urine being the first choice medium for biomonitoring [113] and inductively coupled plasma-mass spectrometry of hair is a powerful tool to recapitulate the history of Cd exposure [114]. Thus, a combination of a chemical evaluation of Cd with the analysis of the arrays of the so-called “biomarkers of effects”, such as the molecular signatures, can provide a useful tool for the identification of marker genes in risk assessment [115,116], serving in both prognostic and predictive applications for the screening of exposed populations, as already reported in the cases of Cd-induced neuro- and nefrotoxicity [117,118,119].
Funding
This research was funded by the University of Palermo (Italy), grants FFR 2018 and FFR 2020.
Conflicts of Interest
The author declares no conflict of interest.
References
- World Health Organization. Exposure to Cadmium: A Major Public Health Concern. 2019. Available online: https://apps.who.int/iris/bitstream/handle/10665/329480/WHO-CED-PHE-EPE-19.4.3-eng.pdf?ua=1 (accessed on 18 January 2021).
- Thévenod, F.; Fels, J.; Lee, W.K.; Zarbock, R. Channels, transporters and receptors for cadmium and cadmium complexes in eukaryotic cells: Myths and facts. Biometals 2019, 32, 469–489. [Google Scholar] [CrossRef] [PubMed]
- Maret, W.; Moulis, J.M. The bioinorganic chemistry of cadmium in the context of its toxicity. Met. Ions Life Sci. 2013, 11, 1–29. [Google Scholar] [CrossRef] [PubMed]
- Choong, G.; Liu, Y.; Templeton, D.M. Interplay of calcium and cadmium in mediating cadmium toxicity. Chem. Biol. Interact. 2014, 211, 54–65. [Google Scholar] [CrossRef] [PubMed]
- Namdarghanbari, M.A.; Bertling, J.; Krezoski, S.; Petering, D.H. Toxic metal proteomics: Reaction of the mammalian zinc proteome with Cd2+. J. Inorg. BioChem. 2014, 136, 115–121. [Google Scholar] [CrossRef]
- Branca, J.J.V.; Pacini, A.; Gulisano, M.; Taddei, N.; Fiorillo, C.; Becatti, M. Cadmium-Induced cytotoxicity: Effects on mitochondrial electron transport chain. Front. Cell Dev. Biol. 2020, 8, 604377. [Google Scholar] [CrossRef]
- Genchi, G.; Sinicropi, M.S.; Lauria, G.; Carocci, A.; Catalano, A. The effects of cadmium toxicity. Int. J. Environ. Res. Public Health 2020, 17, 3782. [Google Scholar] [CrossRef]
- Méplan, C.; Mann, K.; Hainaut, P. Cadmium induces conformational modifications of wild-type p53 and suppresses p53 response to DNA damage in cultured cells. J. Biol. Chem. 1999, 274, 31663–31670. [Google Scholar] [CrossRef]
- Anetor, J.I. Rising environmental cadmium levels in developing countries: Threat to genome stability and health. Niger. J. Physiol. Sci. 2012, 27, 103–115. [Google Scholar] [CrossRef]
- Fujiwara, Y.; Lee, J.Y.; Tokumoto, M.; Satoh, M. Cadmium renal toxicity via apoptotic pathways. Biol. Pharm. Bull. 2012, 35, 1892–1897. [Google Scholar] [CrossRef]
- Thévenod, F.; Lee, W.K. Cadmium and cellular signaling cascades: Interactions between cell death and survival pathways. Arch. Toxicol. 2013, 87, 1743–1786. [Google Scholar] [CrossRef]
- Luevano, J.; Damodaran, C. A review of molecular events of cadmium-induced carcinogenesis. J. Environ. Pathol. Toxicol. Oncol. 2014, 33, 183–194. [Google Scholar] [CrossRef] [PubMed]
- Tokumoto, M.; Lee, J.Y.; Satoh, M. Transcription factors and downstream genes in cadmium toxicity. Biol. Pharm. Bull. 2019, 42, 1083–1088. [Google Scholar] [CrossRef]
- Koedrith, P.; Kim, H.; Weon, J.I.; Seo, Y.R. Toxicogenomic approaches for understanding molecular mechanisms of heavy metal mutagenicity and carcinogenicity. Int. J. Hyg. Environ. Health 2013, 216, 587–598. [Google Scholar] [CrossRef]
- Ijomone, O.M.; Ijomone, O.K.; Iroegbu, J.D.; Ifenatuoha, C.W.; Olung, N.F.; Aschner, M. Epigenetic influence of environmentally neurotoxic metals. Neurotoxicology 2020, 81, 51–65. [Google Scholar] [CrossRef] [PubMed]
- Hartwig, A. Cadmium and cancer. Met. Ions Life Sci. 2013, 11, 491–507. [Google Scholar] [CrossRef]
- Chen, Q.Y.; Costa, M. A comprehensive review of metal-induced cellular transformation studies. Toxicol. Appl. Pharm. 2017, 331, 33–40. [Google Scholar] [CrossRef]
- Sirchia, R.; Longo, A.; Luparello, C. Cadmium regulation of apoptotic and stress response genes in tumoral and immortalized epithelial cells of the human breast. Biochimie 2008, 90, 1578–1590. [Google Scholar] [CrossRef] [PubMed]
- Byrne, C.; Divekar, S.D.; Storchan, G.B.; Parodi, D.A.; Martin, M.B. Metals and breast cancer. J. Mammary Gland Biol. Neoplasia 2013, 18, 63–73. [Google Scholar] [CrossRef]
- Skipper, A.; Sims, J.N.; Yedjou, C.G.; Tchounwou, P.B. Cadmium chloride induces DNA damage and apoptosis of human liver carcinoma cells via oxidative stress. Int. J. Environ. Res. Public Health 2016, 13, 88. [Google Scholar] [CrossRef]
- Luparello, C.; Sirchia, R.; Longo, A. Cadmium as a transcriptional modulator in human cells. Crit. Rev. Toxicol. 2011, 1, 75–82. [Google Scholar] [CrossRef]
- Levenson, A.S.; Jordan, V.C. MCF-7: The first hormone-responsive breast cancer cell line. Cancer Res. 1997, 57, 3071–3078. [Google Scholar]
- Gartel, A.L.; Feliciano, C.; Tyner, A.L. A new method for determining the status of p53 in tumor cell lines of different origin. Oncol. Res. 2003, 13, 405–408. [Google Scholar] [CrossRef]
- Huovinen, M.; Loikkanen, J.; Myllynen, P.; Vähäkangas, K.H. Characterization of human breast cancer cell lines for the studies on p53 in chemical carcinogenesis. Toxicol. Vitr. 2011, 25, 1007–1017. [Google Scholar] [CrossRef]
- Caradonna, F.; Luparello, C. Cytogenetic characterization of HB2 epithelial cells from the human breast. Vitr. Cell Dev. Biol. Anim. 2014, 50, 48–55. [Google Scholar] [CrossRef] [PubMed]
- Luparello, C.; Sirchia, R.; Paci, L.; Miceli, V.; Vella, R.; Scudiero, R.; Trinchella, F. Response to cadmium stress by neoplastic and immortalized human breast cells: Evidence for different modulation of gene expression. In Trends in Signal Transduction Research; Meyer, J.N., Ed.; Nova Science Publ.: Happauge, NY, USA, 2007; pp. 87–113. [Google Scholar]
- Cannino, G.; Ferruggia, E.; Luparello, C.; Rinaldi, A.M. Effects of cadmium chloride on some mitochondria-related activity and gene expression of human MDA-MB231 breast tumor cells. J. Inorg. BioChem. 2008, 102, 1668–1676. [Google Scholar] [CrossRef] [PubMed]
- Casano, C.; Agnello, M.; Sirchia, R.; Luparello, C. Cadmium effects on p38/MAPK isoforms in MDA-MB231 breast cancer cells. Biometals 2010, 23, 83–92. [Google Scholar] [CrossRef]
- Luparello, C.; Longo, A.; Vetrano, M. Exposure to cadmium chloride influences astrocyte-elevated gene-1 (AEG-1) expression in MDA-MB231 human breast cancer cells. Biochimie 2012, 94, 207–213. [Google Scholar] [CrossRef] [PubMed]
- Longo, A.; Librizzi, M.; Luparello, C. Effect of transfection with PLP2 antisense oligonucleotides on gene expression of cadmium-treated MDA-MB231 breast cancer cells. Anal. Bioanal. Chem. 2013, 405, 1893–1901. [Google Scholar] [CrossRef]
- Bylander, J.E.; Li, S.L.; Sens, M.A.; Hazen-Martin, D.; Re, G.G.; Sens, D.A. Induction of metallothionein mRNA and protein following exposure of cultured human proximal tubule cells to cadmium. Toxicol. Lett. 1994, 71, 111–122. [Google Scholar] [CrossRef]
- Li, J.; Liu, Y.; Ru, B. Effect of metallothionein on cell viability and its interactions with cadmium and zinc in HEK293 cells. Cell Biol. Int. 2005, 29, 843–848. [Google Scholar] [CrossRef]
- Chang, X.; Jin, T.; Chen, L.; Nordberg, M.; Lei, L. Metallothionein I isoform mRNA expression in peripheral lymphocytes as a biomarker for occupational cadmium exposure. Exp. Biol. Med. 2009, 234, 666–672. [Google Scholar] [CrossRef] [PubMed]
- Albrecht, A.L.; Singh, R.K.; Somji, S.; Sens, M.A.; Sens, D.A.; Garrett, S.H. Basal and metal-induced expression of metallothionein isoform 1 and 2 genes in the RWPE-1 human prostate epithelial cell line. J. Appl. Toxicol. 2008, 28, 283–293. [Google Scholar] [CrossRef]
- Sun, X.; Niu, X.; Chen, R.; He, W.; Chen, D.; Kang, R.; Tang, D. Metallothionein-1G facilitates sorafenib resistance through inhibition of ferroptosis. Hepatology 2016, 64, 488–500. [Google Scholar] [CrossRef]
- Paschen, W.; Mengesdorf, T.; Althausen, S.; Hotop, S. Peroxidative stress selectively down-regulates the neuronal stress response activated under conditions of endoplasmic reticulum dysfunction. J. Neurochem. 2001, 76, 1916–1924. [Google Scholar] [CrossRef]
- Hua, G.; Zhang, Q.; Fan, Z. Heat shock protein 75 (TRAP1) antagonizes reactive oxygen species generation and protects cells from granzyme M-mediated apoptosis. J. Biol. Chem. 2007, 282, 20553–20560. [Google Scholar] [CrossRef]
- Tang, H.; Chen, Y.; Liu, X.; Wang, S.; Lv, Y.; Wu, D.; Wang, Q.; Luo, M.; Deng, H. Downregulation of HSP60 disrupts mitochondrial proteostasis to promote tumorigenesis and progression in clear cell renal cell carcinoma. Oncotarget 2016, 7, 38822–38834. [Google Scholar] [CrossRef] [PubMed]
- Masuda, Y.; Shima, G.; Aiuchi, T.; Horie, M.; Hori, K.; Nakajo, S.; Kajimoto, S.; Shibayama-Imazu, T.; Nakaya, K. Involvement of tumor necrosis factor receptor-associated protein 1 (TRAP1) in apoptosis induced by beta-hydroxyisovalerylshikonin. J. Biol. Chem. 2004, 279, 42503–42515. [Google Scholar] [CrossRef] [PubMed]
- Moyano, P.; García, J.M.; Lobo, M.; Anadón, M.J.; Sola, E.; Pelayo, A.; García, J.; Frejo, M.T.; Pino, J.D. Cadmium alters heat shock protein pathways in SN56 cholinergic neurons, leading to Aβ and phosphorylated Tau protein generation and cell death. Food Chem. Toxicol. 2018, 121, 297–308. [Google Scholar] [CrossRef] [PubMed]
- Kindas-Mügge, I.; Micksche, M.; Trautinger, F. Modification of growth in small heat shock (hsp27) gene transfected breast carcinoma. Anticancer Res. 1998, 18, 413–417. [Google Scholar] [PubMed]
- Bialik, S.; Kimchi, A. The DAP-kinase interactome. Apoptosis 2014, 19, 316–328. [Google Scholar] [CrossRef] [PubMed]
- Tseng, T.H.; Chien, M.H.; Lin, W.L.; Wen, Y.C.; Chow, J.M.; Chen, C.K.; Kuo, T.C.; Lee, W.J. Inhibition of MDA-MB-231 breast cancer cell proliferation and tumor growth by apigenin through induction of G2/M arrest and histone H3 acetylation-mediated p21WAF1/CIP1 expression. Environ. Toxicol. 2017, 32, 434–444. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, N.; Kowald, L.; van Wijk, S.J.L.; Fulda, S. Differential involvement of TAK1, RIPK1 and NF-κB signaling in Smac mimetic-induced cell death in breast cancer cells. Biol. Chem. 2019, 400, 171–180. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Dai, Y.; Wang, L.; Li, L. Astrocyte elevated gene-1 promotes the proliferation and invasion of breast cancer cells by activating the Wnt/β-catenin signaling pathway. Oncol. Lett. 2017, 13, 2385–2390. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhang, N.; Wang, X.; Huo, Q.; Li, X.; Wang, H.; Schneider, P.; Hu, G.; Yang, Q. The oncogene metadherin modulates the apoptotic pathway based on the tumor necrosis factor superfamily member TRAIL (Tumor Necrosis Factor-related Apoptosis-inducing Ligand) in breast cancer. J. Biol. Chem. 2013, 288, 9396–9407. [Google Scholar] [CrossRef]
- Zhong, W.; Zhu, H.; Sheng, F.; Tian, Y.; Zhou, J.; Chen, Y.; Li, S.; Lin, J. Activation of the MAPK11/12/13/14 (p38 MAPK) pathway regulates the transcription of autophagy genes in response to oxidative stress induced by a novel copper complex in HeLa cells. Autophagy 2014, 10, 1285–1300. [Google Scholar] [CrossRef]
- Wei, Z.; Shaikh, Z.A. Cadmium stimulates metastasis-associated phenotype in triple-negative breast cancer cells through integrin and β-catenin signaling. Toxicol. Appl. Pharm. 2017, 328, 70–80. [Google Scholar] [CrossRef]
- Wei, Z.; Shan, Z.; Shaikh, Z.A. Epithelial-mesenchymal transition in breast epithelial cells treated with cadmium and the role of Snail. Toxicol. Appl. Pharm. 2018, 344, 46–55. [Google Scholar] [CrossRef]
- Hrckulak, D.; Kolar, M.; Strnad, H.; Korinek, V. TCF/LEF transcription factors: An update from the internet resources. Cancers 2016, 8, 70. [Google Scholar] [CrossRef]
- Cano, A.; Neto, M.A. Snail Transcription Factors. In Encyclopedia of Cancer; Schwab, M., Ed.; Springer: Berlin/Heidelberg, Germany, 2011. [Google Scholar] [CrossRef]
- Korpal, M.; Kang, Y. The emerging role of miR-200 family of microRNAs in epithelial-mesenchymal transition and cancer metastasis. RNA Biol. 2008, 5, 115–119. [Google Scholar] [CrossRef]
- Lubovac-Pilav, Z.; Borràs, D.M.; Ponce, E.; Louie, M.C. Using expression profiling to understand the effects of chronic cadmium exposure on MCF-7 breast cancer cells. PLoS ONE 2013, 8, e84646. [Google Scholar] [CrossRef]
- Liang, Z.Z.; Zhu, R.M.; Li, Y.L.; Jiang, H.M.; Li, R.B.; Tang, L.Y.; Wang, Q.; Ren, Z.F. Differential epigenetic and transcriptional profile in MCF-7 breast cancer cells exposed to cadmium. Chemosphere 2020, 261, 128148. [Google Scholar] [CrossRef]
- Dong, C.; Zhang, L.; Sun, R.; Liu, J.; Yin, H.; Li, X.; Zheng, X.; Zeng, H. Role of thioredoxin reductase 1 in dysplastic transformation of human breast epithelial cells triggered by chronic oxidative stress. Sci. Rep. 2016, 6, 36860. [Google Scholar] [CrossRef]
- Qu, H.; Zhu, F.; Dong, H.; Hu, X.; Han, M. Upregulation of CCT-3 Induces breast cancer cell proliferation through miR-223 competition and Wnt/β-catenin signaling pathway activation. Front. Oncol. 2020, 10, 533176. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.; Bu, S.; Wang, X.; Zhang, H.; Ge, H. Suppression of CCT3 inhibits the proliferation and migration in breast cancer cells. Cancer Cell Int. 2020, 20, 218. [Google Scholar] [CrossRef] [PubMed]
- Castagnoli, L.; Tagliabue, E.; Pupa, S.M. Inhibition of the Wnt signalling pathway: An Avenue to control breast cancer aggressiveness. Int. J. Mol. Sci. 2020, 21, 9069. [Google Scholar] [CrossRef] [PubMed]
- Darwish, W.S.; Chen, Z.; Li, Y.; Wu, Y.; Chiba, H.; Hui, S.P. Identification of cadmium-produced lipid hydroperoxides, transcriptomic changes in antioxidant enzymes, xenobiotic transporters, and pro-inflammatory markers in human breast cancer cells (MCF7) and protection with fat-soluble vitamins. Environ. Sci. Pollut. Res. Int. 2020, 27, 1978–1990. [Google Scholar] [CrossRef]
- Carrière, P.; Mantha, M.; Champagne-Paradis, S.; Jumarie, C. Characterization of basolateral-to-apical transepithelial transport of cadmium in intestinal TC7 cell monolayers. Biometals 2011, 24, 857–874. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, Y.; Liu, B.; Song, A. Identifying breast cancer subtypes associated modules and biomarkers by integrated bioinformatics analysis. BioSci. Rep. 2021, 41, BSR20203200. [Google Scholar] [CrossRef]
- Siewit, C.L.; Gengler, B.; Vegas, E.; Puckett, R.; Louie, M.C. Cadmium promotes breast cancer cell proliferation by potentiating the interaction between ERalpha and c-Jun. Mol. Endocrinol. 2010, 24, 981–992. [Google Scholar] [CrossRef]
- Ghosh, K.; Chatterjee, B.; Behera, P.; Kanade, S.R. The carcinogen cadmium elevates CpG-demethylation and enrichment of NFYA and E2F1 in the promoter of oncogenic PRMT5 and EZH2 methyltransferases resulting in their elevated expression in vitro. Chemosphere 2020, 242, 125186. [Google Scholar] [CrossRef] [PubMed]
- Alam, J.; Wicks, C.; Stewart, D.; Gong, P.; Touchard, C.; Otterbein, S.; Choi, A.M.; Burow, M.E.; Tou, J. Mechanism of heme oxygenase-1 gene activation by cadmium in MCF-7 mammary epithelial cells. Role of p38 kinase and Nrf2 transcription factor. J. Biol. Chem. 2000, 275, 27694–27702. [Google Scholar] [CrossRef]
- Bae, S.I.; Park, J.G.; Kim, Y.I.; Kim, W.H. Genetic alterations in gastric cancer cell lines and their original tissues. Int. J. Cancer 2000, 87, 512–516. [Google Scholar] [CrossRef]
- Ji, J.; Chen, X.; Leung, S.Y.; Chi, J.T.; Chu, K.M.; Yuen, S.T.; Li, R.; Chan, A.S.; Li, J.; Dunphy, N.; et al. Comprehensive analysis of the gene expression profiles in human gastric cancer cell lines. Oncogene 2002, 21, 6549–6556. [Google Scholar] [CrossRef]
- Khoi, P.N.; Xia, Y.; Lian, S.; Kim, H.D.; Kim, D.H.; Joo, Y.E.; Chay, K.O.; Kim, K.K.; Jung, Y.D. Cadmium induces urokinase-type plasminogen activator receptor expression and the cell invasiveness of human gastric cancer cells via the ERK-1/2, NF-κB, and AP-1 signaling pathways. Int. J. Oncol. 2014, 45, 1760–1768. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, D.; Eide, P.W.; Eilertsen, I.A.; Danielsen, S.A.; Eknæs, M.; Hektoen, M.; Lind, G.E.; Lothe, R.A. Epigenetic and genetic features of 24 colon cancer cell lines. Oncogenesis 2013, 2, e71. [Google Scholar] [CrossRef]
- Martínez-Maqueda, D.; Miralles, B.; Recio, I. HT29 Cell Line. In The Impact of Food Bioactives on Health; Verhoeckx, K., Ed.; Springer: Cham, Switzerland, 2015; pp. 113–124. [Google Scholar] [CrossRef]
- Kwon, J.Y.; Weon, J.I.; Koedrith, P.; Park, K.S.; Kim, I.S.; Seo, Y.R. Identification of molecular candidates and interaction networks via integrative toxicogenomic analysis in a human cell line following low-dose exposure to the carcinogenic metals cadmium and nickel. Oncol. Rep. 2013, 30, 1185–1194. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, S.S.; Glatzle, J.; Bajaeifer, K.; Bühler, S.; Lehmann, T.; Königsrainer, I.; Vollmer, J.P.; Sipos, B.; Ahmad, S.S.; Northoff, H.; et al. Phosphoglycerate kinase 1 as a promoter of metastasis in colon cancer. Int. J. Oncol. 2013, 43, 586–590. [Google Scholar] [CrossRef]
- Chen, W.D.; Zhang, Y. Regulation of aldo-keto reductases in human diseases. Front. Pharm. 2012, 3, 35. [Google Scholar] [CrossRef] [PubMed]
- Xia, W.; Zhuang, J.; Wang, G.; Ni, J.; Wang, J.; Ye, Y. P4HB promotes HCC tumorigenesis through downregulation of GRP78 and subsequent upregulation of epithelial-to-mesenchymal transition. Oncotarget 2017, 8, 8512–8521. [Google Scholar] [CrossRef]
- Naji, S.; Issa, K.; Eid, A.; Iratni, R.; Eid, A.H. Cadmium Induces migration of colon cancer cells: Roles of reactive oxygen species, P38 and cyclooxygenase-2. Cell Physiol. BioChem. 2019, 52, 1517–1534. [Google Scholar] [CrossRef] [PubMed]
- Iftode, A.; Drăghici, G.A.; Macașoi, I.; Marcovici, I.; Coricovac, D.E.; Dragoi, R.; Tischer, A.; Kovatsi, L.; Tsatsakis, A.M.; Cretu, O.; et al. Exposure to cadmium and copper triggers cytotoxic effects and epigenetic changes in human colorectal carcinoma HT-29 cells. Exp. Ther. Med. 2021, 21, 100. [Google Scholar] [CrossRef]
- Fabbri, M.; Urani, C.; Sacco, M.G.; Procaccianti, C.; Gribaldo, L. Whole genome analysis and microRNAs regulation in HepG2 cells exposed to cadmium. ALTEX Altern. Anim. Exp. 2012, 29, 173–182. [Google Scholar] [CrossRef]
- Urani, C.; Melchioretto, P.; Bruschi, M.; Fabbri, M.; Sacco, M.G.; Gribaldo, L. Impact of cadmium on intracellular zinc levels in HepG2 cells: Quantitative evaluations and molecular effects. Biomed. Res. Int. 2015, 2015, 949514. [Google Scholar] [CrossRef] [PubMed]
- Panera, N.; Crudele, A.; Romito, I.; Gnani, D.; Alisi, A. Focal adhesion kinase: Insight into molecular roles and functions in hepatocellular carcinoma. Int. J. Mol. Sci. 2017, 18, 99. [Google Scholar] [CrossRef]
- Delire, B.; Stärkel, P. The Ras/MAPK pathway and hepatocarcinoma: Pathogenesis and therapeutic implications. Eur. J. Clin. Investig. 2015, 45, 609–623. [Google Scholar] [CrossRef]
- Cartularo, L.; Laulicht, F.; Sun, H.; Kluz, T.; Freedman, J.H.; Costa, M. Gene expression and pathway analysis of human hepatocellular carcinoma cells treated with cadmium. Toxicol. Appl. Pharm. 2015, 288, 399–408. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, R.; Pandey, S.K.; Goel, S.; Bhatia, V.; Shukla, S.; Jing, X.; Dhanasekaran, S.M.; Ateeq, B. SPINK1 promotes colorectal cancer progression by downregulating Metallothioneins expression. Oncogenesis 2015, 4, e162. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Jia, J.; Zhang, D. Purinergic signalling in liver diseases: Pathological functions and therapeutic opportunities. JHEP Rep. 2020, 2, 100165. [Google Scholar] [CrossRef] [PubMed]
- Nibourg, G.A.; Huisman, M.T.; van der Hoeven, T.V.; van Gulik, T.M.; Chamuleau, R.A.; Hoekstra, R. Stable overexpression of pregnane X receptor in HepG2 cells increases its potential for bioartificial liver application. Liver Transpl. 2010, 16, 1075–1085. [Google Scholar] [CrossRef] [PubMed]
- Gerets, H.H.; Tilmant, K.; Gerin, B.; Chanteux, H.; Depelchin, B.O.; Dhalluin, S.; Atienzar, F.A. Characterization of primary human hepatocytes, HepG2 cells, and HepaRG cells at the mRNA level and CYP activity in response to inducers and their predictivity for the detection of human hepatotoxins. Cell Biol. Toxicol. 2012, 28, 69–87. [Google Scholar] [CrossRef] [PubMed]
- Inagaki, F.F.; Tanaka, M.; Inagaki, N.F.; Yagai, T.; Sato, Y.; Sekiguchi, K.; Oyaizu, N.; Kokudo, N.; Miyajima, A. Nephronectin is upregulated in acute and chronic hepatitis and aggravates liver injury by recruiting CD4 positive cells. BioChem. Biophys. Res. Commun. 2013, 430, 751–756. [Google Scholar] [CrossRef] [PubMed]
- Magee, N.; Zhang, Y. Role of early growth response 1 in liver metabolism and liver cancer. Hepatoma Res. 2017, 3, 268–277. [Google Scholar] [CrossRef] [PubMed]
- Yin, X.; Tang, B.; Li, J.H.; Wang, Y.; Zhang, L.; Xie, X.Y.; Zhang, B.H.; Qiu, S.J.; Wu, W.Z.; Ren, Z.G. ID1 promotes hepatocellular carcinoma proliferation and confers chemoresistance to oxaliplatin by activating pentose phosphate pathway. J. Exp. Clin. Cancer Res. 2017, 36, 166. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.J.; Jung, Y.J.; Lee, S.; Kim, J.I.; Han, J.A. DNAJB9 Inhibits p53-dependent oncogene-induced senescence and induces cell transformation. Mol. Cells 2020, 43, 397–407. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Li, T.; Kong, D.; You, H.; Kong, F.; Tang, R. Prognostic implications of alcohol dehydrogenases in hepatocellular carcinoma. BMC Cancer 2020, 20, 1204. [Google Scholar] [CrossRef]
- Mitsudomi, T.; Viallet, J.; Mulshine, J.L.; Linnoila, R.I.; Minna, J.D.; Gazdar, A.F. Mutations of ras genes distinguish a subset of non-small-cell lung cancer cell lines from small-cell lung cancer cell lines. Oncogene 1991, 6, 1353–1362. [Google Scholar]
- Mitsudomi, T.; Steinberg, S.M.; Nau, M.M.; Carbone, D.; D’Amico, D.; Bodner, S.; Oie, H.K.; Linnoila, R.I.; Mulshine, J.L.; Minna, J.D.; et al. p53 gene mutations in non-small-cell lung cancer cell lines and their correlation with the presence of ras mutations and clinical features. Oncogene 1992, 7, 171–180. [Google Scholar]
- Park, C.H.; Lee, B.H.; Ahn, S.G.; Yoon, J.H.; Oh, S.H. Serine 9 and tyrosine 216 phosphorylation of GSK-3β differentially regulates autophagy in acquired cadmium resistance. Toxicol. Sci. 2013, 135, 380–389. [Google Scholar] [CrossRef] [PubMed]
- A549—A model for Non-Small Cell Lung Cancer. Available online: https://www.covance.com/industry-solutions/oncology/preclinical/tumor-spotlights/model-spotlight-a549-a-model-for-non-small-cell-lung-cancer.html (accessed on 6 April 2021).
- Kim, H.R.; Lee, K.Y.; Ahn, S.G.; Lee, B.H.; Jung, K.T.; Yoon, J.H.; Yoon, H.E.; Oh, S.H. Transcriptional regulation, stabilization, and subcellular redistribution of multidrug resistance-associated protein 1 (MRP1) by glycogen synthase kinase 3αβ: Novel insights on modes of cadmium-induced cell death stimulated by MRP1. Arch. Toxicol. 2015, 89, 1271–1284. [Google Scholar] [CrossRef]
- Cole, S.P. Multidrug resistance protein 1 (MRP1, ABCC1), a “multitasking” ATP-binding cassette (ABC) transporter. J. Biol. Chem. 2014, 289, 30880–30888. [Google Scholar] [CrossRef] [PubMed]
- Mancinelli, R.; Carpino, G.; Petrungaro, S.; Mammola, C.L.; Tomaipitinca, L.; Filippini, A.; Facchiano, A.; Ziparo, E.; Giampietri, C. Multifaceted roles of GSK-3 in cancer and autophagy-related diseases. Oxid. Med. Cell Longev. 2017, 2017, 4629495. [Google Scholar] [CrossRef]
- Phatak, V.M.; Muller, P.A.J. Metal toxicity and the p53 protein: An intimate relationship. Toxicol. Res. 2015, 4, 576–591. [Google Scholar] [CrossRef]
- Adámik, M.; Bažantová, P.; Navrátilová, L.; Polášková, A.; Pečinka, P.; Holaňová, L.; Tichý, V.; Brázdová, M. Impact of cadmium, cobalt and nickel on sequence-specific DNA binding of p63 and p73 in vitro and in cells. BioChem. Biophys. Res. Commun. 2015, 456, 29–34. [Google Scholar] [CrossRef] [PubMed]
- Fujiki, K.; Inamura, H.; Miyayama, T.; Matsuoka, M. Involvement of Notch1 signaling in malignant progression of A549 cells subjected to prolonged cadmium exposure. J. Biol. Chem. 2017, 292, 7942–7953. [Google Scholar] [CrossRef]
- Qian, Q.; Wang, Q.; Zhan, P.; Peng, L.; Wei, S.Z.; Shi, Y.; Song, Y. The role of matrix metalloproteinase 2 on the survival of patients with non-small cell lung cancer: A systematic review with meta-analysis. Cancer Invest. 2010, 28, 661–669. [Google Scholar] [CrossRef] [PubMed]
- Moore, G.E.; Sandberg, A.A. Studies of a human tumor cell line with a diploid karyotype. Cancer 1964, 17, 170–175. [Google Scholar] [CrossRef]
- Lee, Y.J.; Lee, G.J.; Baek, B.J.; Heo, S.H.; Won, S.Y.; Im, J.H.; Cho, M.K.; Nam, H.S.; Lee, S.H. Cadmium-induced up-regulation of aldo-keto reductase 1C3 expression in human nasal septum carcinoma RPMI-2650 cells: Involvement of reactive oxygen species and phosphatidylinositol 3-kinase/Akt. Environ. Toxicol. Pharm. 2011, 31, 469–478. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; He, S.; Chen, Y.; Liu, Y.; Feng, F.; Liu, W.; Guo, Q.; Zhao, L.; Sun, H. Overview of AKR1C3: Inhibitor achievements and disease insights. J. Med. Chem. 2020, 63, 11305–11329. [Google Scholar] [CrossRef]
- Zeng, Y. Establishment of an epitheloid cell line and a fusiform cell line from a patient with nasopharyngeal carcinoma. Sci. Sin. 1978, 21, 127–134. [Google Scholar]
- Zhang, S.H.; Gao, X.K.; Zeng, Y. Cytogenetic studies on an epithelial cell line derived from poorly differentiated nasopharyngeal carcinoma. Yi Chuan Xue Bao 1983, 10, 498–503. [Google Scholar]
- Peng, L.; Huang, Y.T.; Zhang, F.; Chen, J.Y.; Huo, X. Chronic cadmium exposure aggravates malignant phenotypes of nasopharyngeal carcinoma by activating the wnt/β-catenin signaling pathway via hypermethylation of the casein kinase 1α promoter. Cancer Manag. Res. 2019, 11, 81–93. [Google Scholar] [CrossRef]
- Cruciat, C.M. Casein kinase 1 and Wnt/β-catenin signaling. Curr. Opin. Cell Biol. 2014, 31, 46–55. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Weinstein, J.N. Drug discovery: Cell lines battle cancer. Nature 2012, 483, 544–545. [Google Scholar] [CrossRef]
- Gillet, J.P.; Varma, S.; Gottesman, M.M. The clinical relevance of cancer cell lines. J. Natl. Cancer Inst. 2013, 105, 7452–7458. [Google Scholar] [CrossRef] [PubMed]
- Poblete-Naredo, I.; Albores, A. Molecular biomarkers to assess health risks due to environmental contaminants exposure. Biomedica 2016, 36, 309–335. [Google Scholar] [CrossRef]
- Tiesjema, B.; Mengelers, M. Biomonitoring of Lead and Cadmium: Preliminary Study on the Added Value for Human Exposure and Effect Assessment. Available online: https://www.rivm.nl/bibliotheek/rapporten/2016-0215.pdf (accessed on 10 April 2021).
- Pozebon, D.; Scheffler, G.L.; Dressler, V.L. Elemental hair analysis: A review of procedures and applications. Anal. Chim. Acta 2017, 992, 1–23. [Google Scholar] [CrossRef]
- Dakeshita, S.; Kawai, T.; Uemura, H.; Hiyoshi, M.; Oguma, E.; Horiguchi, H.; Kayama, F.; Aoshima, K.; Shirahama, S.; Rokutan, K.; et al. Gene expression signatures in peripheral blood cells from Japanese women exposed to environmental cadmium. Toxicology 2009, 257, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Bollati, V.; Marinelli, B.; Apostoli, P.; Bonzini, M.; Nordio, F.; Hoxha, M.; Pegoraro, V.; Motta, V.; Tarantini, L.; Cantone, L.; et al. Exposure to metal-rich particulate matter modifies the expression of candidate microRNAs in peripheral blood leukocytes. Environ. Health Perspect. 2010, 118, 763–768. [Google Scholar] [CrossRef]
- Krauskopf, J.; Bergdahl, I.A.; Johansson, A.; Palli, D.; Lundh, T.; Kyrtopoulos, S.A.; de Kok, T.M.; Kleinjans, J.C. Blood transcriptome response to environmental metal exposure reveals potential biological processes related to Alzheimer’s disease. Front. Public Health 2020, 8, 557587. [Google Scholar] [CrossRef] [PubMed]
- Yuan, W.; Liu, L.; Liang, L.; Huang, K.; Deng, Y.; Dong, M.; Chen, J.; Wang, G.; Zou, F. MiR-122-5p and miR-326-3p: Potential novel biomarkers for early detection of cadmium exposure. Gene 2020, 724, 144156. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Huang, Z.; Chen, B.; Lu, Q.; Cao, L.; Chen, W. LncRNA-ENST00000446135 is a novel biomarker of cadmium toxicity in 16HBE cells, rats, and Cd-exposed workers and regulates DNA damage and repair. Toxicol. Res. 2020, 9, 823–834. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).