Deep-Learning-Based Digitization of Protein-Self-Assembly to Print Biodegradable Physically Unclonable Labels for Device Security
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Fractal Pattern Acquisition via Multiscale Imaging
2.3. Fabrication of Flexible and Biodegradable PUF Labels
3. Results
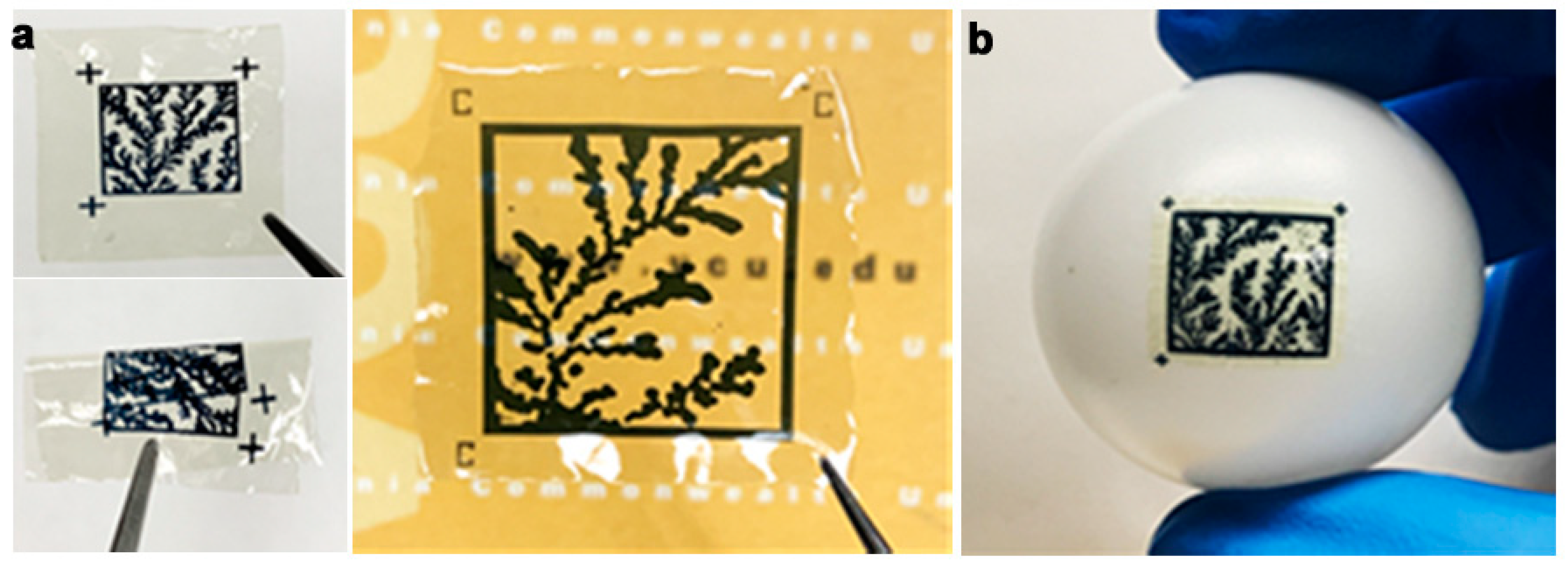
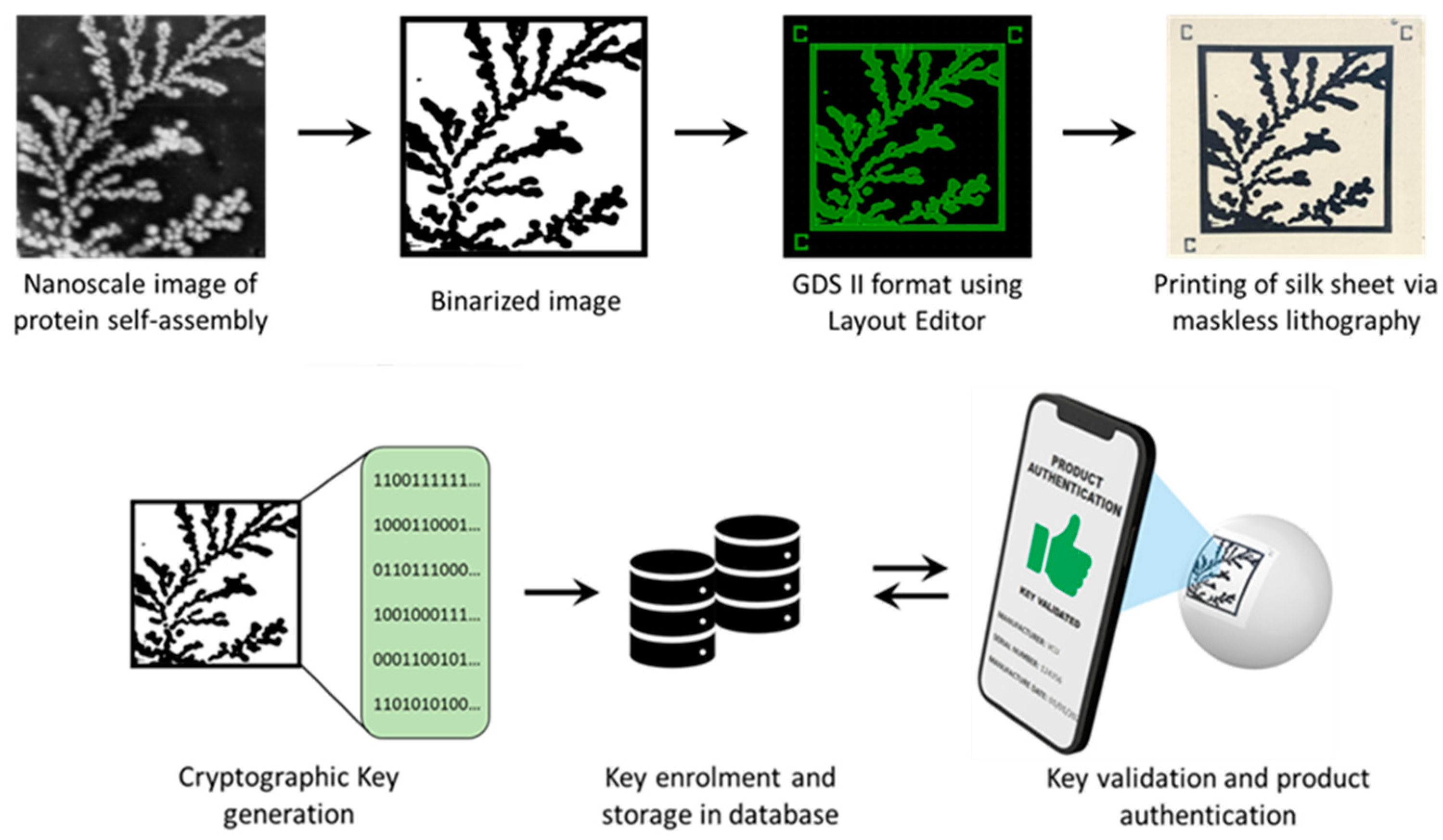
3.1. Printed Image PUFs
3.2. Cryptographic Keys
3.2.1. Image Processing
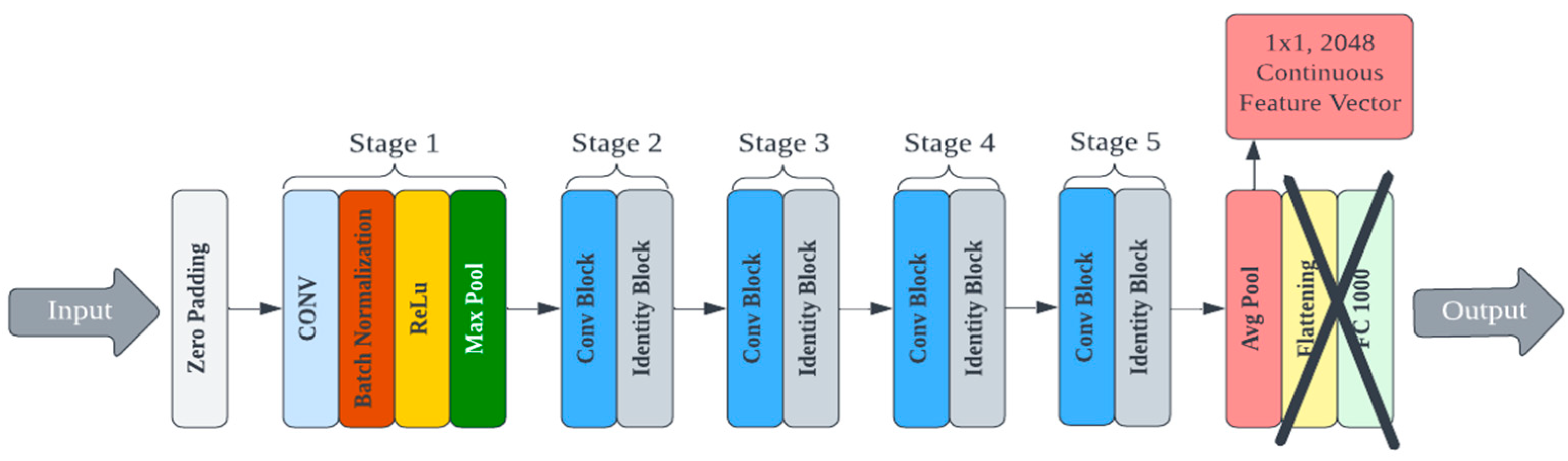
3.2.2. Deep Learning
3.3. Performance of the PUF Devices
3.4. Key Authentication
3.5. Validation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Mackey, T.K.; Nayyar, G. A review of existing and emerging digital technologies to combat the global trade in fake medicines. Expert Opin. Drug Saf. 2017, 16, 587–602. [Google Scholar] [CrossRef]
- Zafar, S.; Nazir, M.; Bakhshi, T.; Khattak, H.A.; Khan, S.; Bilal, M.; Choo, K.-K.R.; Kwak, K.-S.; Sabah, A. A systematic review of bio-cyber interface technologies and security issues for internet of bio-nano things. IEEE Access 2021, 9, 93529–93566. [Google Scholar] [CrossRef]
- Lewis, T.G. Critical Infrastructure Protection in Homeland Security: Defending a Networked Nation; John Wiley & Sons: Hoboken, NJ, USA, 2019. [Google Scholar]
- Soon, J.M.; Manning, L. Developing anti-counterfeiting measures: The role of smart packaging. Food Res. Int. 2019, 123, 135–143. [Google Scholar] [CrossRef]
- Picard, J.; Landry, P.; Bolay, M. Counterfeit Detection with QR Codes. In Proceedings of the 21st ACM Symposium on Document Engineering, Limerick, Ireland, 24–27 August 2021; Association for Computing Machinery: New York, NY, USA, 2021. [Google Scholar]
- Mukhopadhyay, D. PUFs as promising tools for security in internet of things. IEEE Des. Test 2016, 33, 103–115. [Google Scholar] [CrossRef]
- Halak, B.; Zwolinski, M.; Mispan, M.S. Overview of PUF-Based Hardware Security Solutions for the Internet of Things. In Proceedings of the 2016 IEEE 59th International Midwest Symposium on Circuits and Systems (MWSCAS), Abu Dhabi, United Arab, 16–19 October 2016; IEEE: Piscataway, NJ, USA, 2016. [Google Scholar]
- Pappu, R.; Recht, B.; Taylor, J.; Gershenfeld, N. Physical one-way functions. Science 2002, 297, 2026–2030. [Google Scholar] [CrossRef]
- Arppe, R.; Sørensen, T.J. Physical unclonable functions generated through chemical methods for anti-counterfeiting. Nat. Rev. Chem. 2017, 1, 0031. [Google Scholar] [CrossRef]
- Erozan, A.T.; Hefenbrock, M.; Beigl, M.; Aghassi-Hagmann, J.; Tahoori, M.B. Image PUF: A Physical Unclonable Function for Printed Electronics based on Optical Variation of Printed Inks. IACR Cryptol. Eprint Arch. 2019, 2019, 1419. [Google Scholar]
- Wigger, B.; Meissner, T.; Förste, A.; Jetter, V.; Zimmermann, A. Using unique surface patterns of injection moulded plastic components as an image based Physical Unclonable Function for secure component identification. Sci. Rep. 2018, 8, 4738. [Google Scholar] [CrossRef]
- Zerrouki, F.; Ouchani, S.; Bouarfa, H. A survey on silicon PUFs. J. Syst. Archit. 2022, 127, 102514. [Google Scholar] [CrossRef]
- Wali, A.; Dodda, A.; Wu, Y.; Pannone, A.; Reddy Usthili, L.K.; Ozdemir, S.K.; Ozbolat, I.T.; Das, S. Biological physically unclonable function. Commun. Phys. 2019, 2, 39. [Google Scholar] [CrossRef]
- Leem, J.W.; Kim, M.S.; Choi, S.H.; Kim, S.-R.; Kim, S.-W.; Song, Y.M.; Young, R.J.; Kim, Y.L. Edible unclonable functions. Nat. Commun. 2020, 11, 328. [Google Scholar] [CrossRef]
- Bae, H.J.; Bae, S.; Park, C.; Han, S.; Kim, J.; Kim, L.N.; Kim, K.; Song, S.H.; Park, W.; Kwon, S. Biomimetic microfingerprints for anti-counterfeiting strategies. Adv. Mater. 2015, 27, 2083–2089. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Shu, Z.; Zhang, L.; Chen, Y.; Feng, Z.; Hu, Y.; Huang, F.; Wang, P.; Li, D.; Yao, Y. Random Nanofracture-Enabled Physical Unclonable Function. Adv. Mater. Technol. 2021, 6, 2001073. [Google Scholar] [CrossRef]
- Smith, A.F.; Patton, P.; Skrabalak, S.E. Plasmonic nanoparticles as a physically unclonable function for responsive anti-counterfeit nanofingerprints. Adv. Funct. Mater. 2016, 26, 1315–1321. [Google Scholar] [CrossRef]
- Kim, M.S.; Lee, G.J.; Leem, J.W.; Choi, S.; Kim, Y.L.; Song, Y.M. Revisiting silk: A lens-free optical physical unclonable function. Nat. Commun. 2022, 13, 247. [Google Scholar] [CrossRef]
- Hu, Y.W.; Zhang, T.P.; Wang, C.F.; Liu, K.K.; Sun, Y.; Li, L.; Lv, C.F.; Liang, Y.C.; Jiao, F.H.; Zhao, W.B. Flexible and biocompatible physical unclonable function anti-counterfeiting label. Adv. Funct. Mater. 2021, 31, 2102108. [Google Scholar] [CrossRef]
- Whitesides, G.M.; Grzybowski, B. Self-assembly at all scales. Science 2002, 295, 2418–2421. [Google Scholar] [CrossRef]
- Philp, D.; Stoddart, J.F. Self-assembly in natural and unnatural systems. Angew. Chem.-Int. Ed. Engl. 1996, 35, 1155–1196. [Google Scholar] [CrossRef]
- Bohr, H.; Kühle, A.; Sørensen, A.H.; Bohr, J. Hierarchical organization in aggregates of protein molecules. Z. Für Phys. D At. Mol. Clust. 1997, 40, 513–515. [Google Scholar] [CrossRef]
- Meakin, P.; Jullien, R. The Effects of Restructuring on the Geometry of Clusters Formed by Diffusion-Limited, Ballistic, and Reaction-Limited Cluster Cluster Aggregation. J. Chem. Phys. 1988, 89, 246–250. [Google Scholar] [CrossRef]
- Khire, T.S.; Kundu, J.; Kundu, S.C.; Yadavalli, V.K. The fractal self-assembly of the silk protein sericin. Soft Matter 2010, 6, 2066–2071. [Google Scholar] [CrossRef]
- Pal, R.K.; Farghaly, A.A.; Collinson, M.M.; Kundu, S.C.; Yadavalli, V.K. Photolithographic Micropatterning of Conducting Polymers on Flexible Silk Matrices. Adv. Mater 2016, 28, 1406–1412. [Google Scholar] [CrossRef]
- Tokuyama, M.; Kawasaki, K. Fractal Dimensions For Diffusion-Limited Aggregation. Phys. Lett. A 1984, 100, 337–340. [Google Scholar] [CrossRef]
- Choi, J.; Crowdy, D.; Bazant, M.Z. Diffusion-limited aggregation on curved surfaces. EPL (Europhys. Lett.) 2010, 91, 46005. [Google Scholar] [CrossRef]
- Hurd, A.J.; Schaefer, D.W. Diffusion-limited aggregation in two dimensions. Phys. Rev. Lett. 1985, 54, 1043. [Google Scholar] [CrossRef] [PubMed]
- Kato, N.; Sato, S.; Yamanaka, A.; Yamada, H.; Fuwa, N.; Nomura, M. Silk protein, sericin, inhibits lipid peroxidation and tyrosinase activity. Biosci. Biotechnol. Biochem. 1998, 62, 145–147. [Google Scholar] [CrossRef]
- Witten Jr, T.A.; Sander, L.M. Diffusion-limited aggregation, a kinetic critical phenomenon. Phys. Rev. Lett. 1981, 47, 1400. [Google Scholar] [CrossRef]
- Sander, L.M. Diffusion-limited aggregation: A kinetic critical phenomenon? Contemp. Phys. 2000, 41, 203–218. [Google Scholar] [CrossRef]
- Goh, T.Y.; Basah, S.N.; Yazid, H.; Safar, M.J.A.; Saad, F.S.A. Performance analysis of image thresholding: Otsu technique. Measurement 2018, 114, 298–307. [Google Scholar] [CrossRef]
- Kurland, N.E.; Dey, T.; Kundu, S.C.; Yadavalli, V.K. Precise Patterning of Silk Microstructures Using Photolithography. Adv. Mater. 2013, 25, 6207–6212. [Google Scholar] [CrossRef]
- Kurland, N.E.; Dey, T.; Wang, C.; Kundu, S.C.; Yadavalli, V.K. Silk protein lithography as a route to fabricate sericin microarchitectures. Adv. Mater 2014, 26, 4431–4437. [Google Scholar] [CrossRef]
- Xu, M.; Yadavalli, V.K. Flexible Biosensors for the Impedimetric Detection of Protein Targets Using Silk-Conductive Polymer Biocomposites. ACS Sens. 2019, 4, 1040–1047. [Google Scholar] [CrossRef]
- Pal, R.K.; Farghaly, A.A.; Wang, C.; Collinson, M.M.; Kundu, S.C.; Yadavalli, V.K. Conducting polymer-silk biocomposites for flexible and biodegradable electrochemical sensors. Biosens. Bioelectron. 2016, 81, 294–302. [Google Scholar] [CrossRef] [PubMed]
- Pal, R.K.; Kundu, S.C.; Yadavalli, V.K. Biosensing using photolithographically micropatterned electrodes of PEDOT:PSS on ITO substrates. Sens. Actuators B Chem. 2017, 242, 140–147. [Google Scholar] [CrossRef]
- Alani, M.M. Applications of machine learning in cryptography: A survey. In Proceedings of the 3rd International Conference on Cryptography, Security and Privacy, Kuala Lumpur, Malaysia, 19–21 January 2019; Association for Computing Machinery: New York, NY, USA, 2019. [Google Scholar]
- Shen, F.; Xu, Y.; Liu, L.; Yang, Y.; Huang, Z.; Shen, H.T. Unsupervised deep hashing with similarity-adaptive and discrete optimization. IEEE Trans. Pattern Anal. Mach. Intell. 2018, 40, 3034–3044. [Google Scholar] [CrossRef]
- Liu, N.; Mou, H.; Tang, J.; Wan, L.; Li, Q.; Yuan, Y. Fully Connected Hashing Neural Networks for Indexing Large-Scale Remote Sensing Images. Mathematics 2022, 10, 4716. [Google Scholar] [CrossRef]
- Lin, K.; Yang, H.-F.; Hsiao, J.-H.; Chen, C.-S. Deep learning of binary hash codes for fast image retrieval. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition Workshops, Boston, MA, USA, 7–12 June 2015. [Google Scholar]
- Team, Keras. Keras Documentation: Keras Applications. 2020. Available online: https://keras.io/api/applications/ (accessed on 1 March 2023).
- Seepers, R.M.; Strydis, C.; Sourdis, I.; De Zeeuw, C.I. On using a von neumann extractor in heart-beat-based security. In Proceedings of the 2015 IEEE Trustcom/BigDataSE/ISPA, Helsinki, Finland, 20–22 August 2015; IEEE: Piscataway, NJ, USA, 2015. [Google Scholar]
- Rukhin, A.; Soto, J.; Nechvatal, J.; Smid, M.; Barker, E. A Statistical Test Suite for Random and Pseudorandom Number Generators for Cryptographic Applications; Technical Report; Booz-Allen and Hamilton Inc.: Mclean, VA, USA, 2001. [Google Scholar]
- Lu, C.-S.; Hsu, C.-Y. Geometric distortion-resilient image hashing scheme and its applications on copy detection and authentication. Multimed. Syst. 2005, 11, 159–173. [Google Scholar] [CrossRef]
- Ketkar, N. Introduction to keras. In Deep Learning with Python: A Hands-On Introduction; Apress: New York, NY, USA, 2017; pp. 97–111. [Google Scholar]



| NIST Statistical Test | p-Value | Proportion | Result |
|---|---|---|---|
| Frequency | 0.040108 | 54/54 | Pass |
| Block Frequency | 0.574903 | 54/54 | Pass |
| Cumulative Sums | 0.000274 | 54/54 | Pass |
| 0.023545 | 54/54 | Pass | |
| Runs | 0.137282 | 54/54 | Pass |
| Longest Run of Ones | 0.883171 | 54/54 | Pass |
| Approximate Entropy | 0.574903 | 53/54 | Pass |
| Sample # | AFM Image | Printed Label | % Match with AFM Image | |
|---|---|---|---|---|
| Image #1 | Image #2 | |||
| 1 |  |  | 90.7% | 89.8% |
| 2 | 89.4% | 89.5% | ||
| 3 | 93.3% | 93.7% | ||
| 4 | 89.2% | 89.3% | ||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pradhan, S.; Rajagopala, A.D.; Meno, E.; Adams, S.; Elks, C.R.; Beling, P.A.; Yadavalli, V.K. Deep-Learning-Based Digitization of Protein-Self-Assembly to Print Biodegradable Physically Unclonable Labels for Device Security. Micromachines 2023, 14, 1678. https://doi.org/10.3390/mi14091678
Pradhan S, Rajagopala AD, Meno E, Adams S, Elks CR, Beling PA, Yadavalli VK. Deep-Learning-Based Digitization of Protein-Self-Assembly to Print Biodegradable Physically Unclonable Labels for Device Security. Micromachines. 2023; 14(9):1678. https://doi.org/10.3390/mi14091678
Chicago/Turabian StylePradhan, Sayantan, Abhi D. Rajagopala, Emma Meno, Stephen Adams, Carl R. Elks, Peter A. Beling, and Vamsi K. Yadavalli. 2023. "Deep-Learning-Based Digitization of Protein-Self-Assembly to Print Biodegradable Physically Unclonable Labels for Device Security" Micromachines 14, no. 9: 1678. https://doi.org/10.3390/mi14091678
APA StylePradhan, S., Rajagopala, A. D., Meno, E., Adams, S., Elks, C. R., Beling, P. A., & Yadavalli, V. K. (2023). Deep-Learning-Based Digitization of Protein-Self-Assembly to Print Biodegradable Physically Unclonable Labels for Device Security. Micromachines, 14(9), 1678. https://doi.org/10.3390/mi14091678








