Genotypic Regulation of Aflatoxin Accumulation but Not Aspergillus Fungal Growth upon Post-Harvest Infection of Peanut (Arachis hypogaea L.) Seeds
Abstract
1. Introduction
2. Results and Discussion
3. Conclusions
4. Materials and Methods
4.1. Plant Materials
4.2. In Vitro A. flavus Inoculation Using AF-70-GFP
4.3. Visual Tracking of GFP Expression
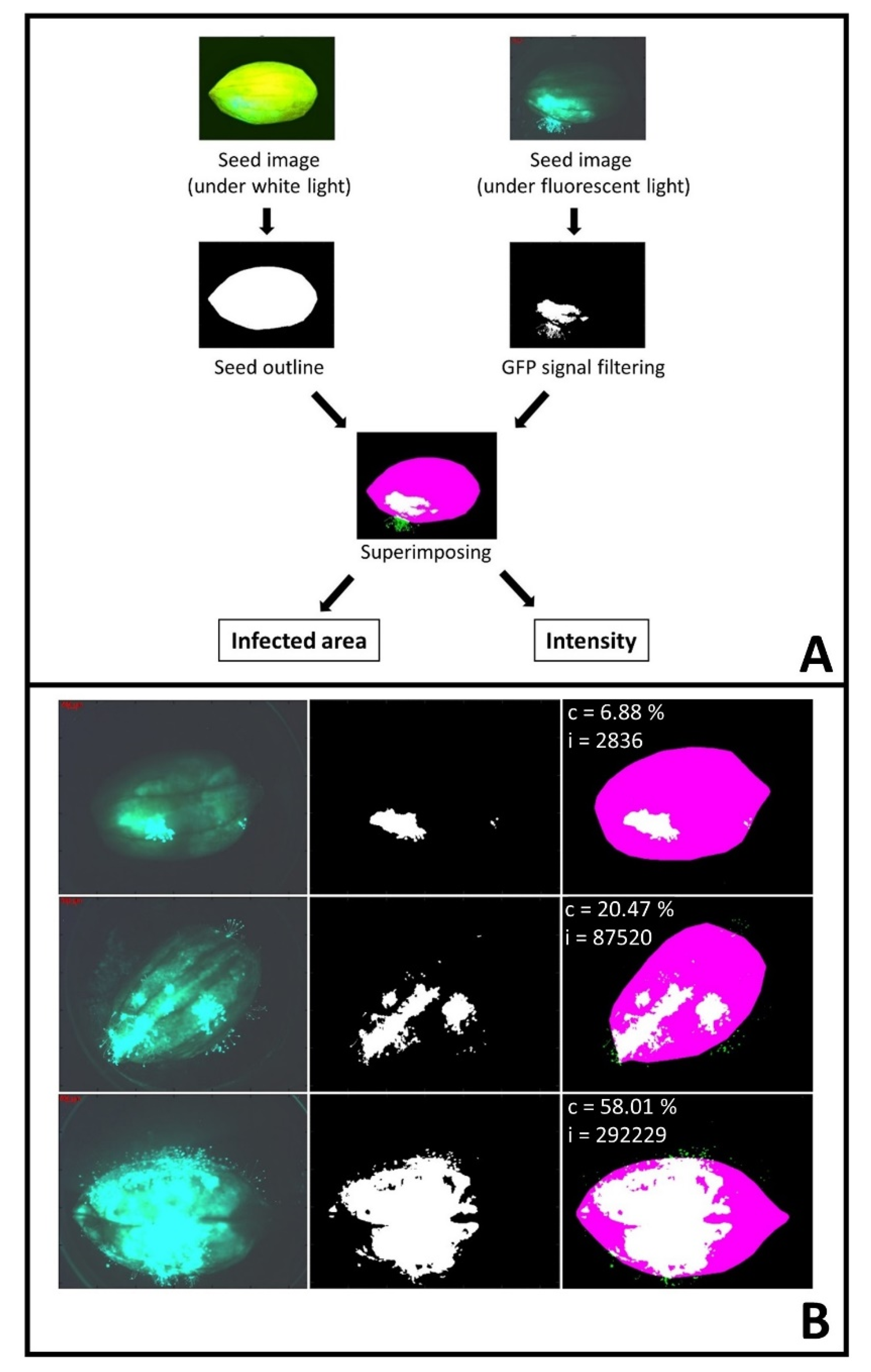
4.4. Image Processing
4.5. GFP and Aflatoxin Analysis
4.6. Rainout Shelter Study
4.7. Statistical Analysis
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| A. flavus | Aspergillus flavus |
| A. parasiticus | Aspergillus parasiticus |
| AUDPC | Area Under the Disease Progress Curve |
| GFP | Green Fluorescent Protein |
| RFU | Relative Fluorescence Unit |
| SICIA | Seed Infection Coverage and Intensity Analyzer |
References
- Chang, H.G.; Markakis, P. Effect of temperature on aflatoxin production in barley by Aspergillus parasiticus. Korean J. Food Sci. Technol. 1982, 14, 162–167. [Google Scholar]
- Rajasekaran, K.; Cary, J.W.; Cotty, P.J.; Cleveland, T.E. Development of a GFP-expressing Aspergillus flavus strain to study fungal invasion, colonization, and resistance in cottonseed. Mycopathologia 2008, 165, 89–97. [Google Scholar] [CrossRef] [PubMed]
- Taheri, N.; Semnani, S.; Roshandel, G.; Namjoo, M.; Keshavarzian, H.; Chogan, A.G.; Kebria, F.G.; Joshaghani, H. Aflatoxin contamination in wheat flour samples from Golestan province, northeast of Iran. Iran. J. Public Health 2012, 41, 42–47. [Google Scholar] [PubMed]
- Elzupir, A.O.; Alamer, A.S.; Dutton, M.F. The occurrence of aflatoxin in rice worldwide: A review. Toxin Rev. 2015, 34, 37–42. [Google Scholar] [CrossRef]
- Ostadrahimi, A.; Ashrafnejad, F.; Kazemi, A.; Sargheini, N.; Mahdavi, R.; Farshchian, M.; Mahluji, S. Aflatoxin in raw and salt-roasted nuts (pistachios, peanuts and walnuts) sold in markets of Tabriz, Iran. Jundishapur J. Microbiol. 2014, 7, e8674. [Google Scholar] [CrossRef] [PubMed]
- Abbas, H.K.; Zablotowicz, R.M.; Shier, W.T.; Johnson, B.J.; Phillips, N.A.; Weaver, M.A.; Abel, C.A.; Bruns, H.A. Aflatoxin and fumonisin in corn (Zea mays) infected by common smut Ustilago maydis. Plant Dis. 2015, 99, 1236–1240. [Google Scholar] [CrossRef]
- Diener, U.L.; Cole, R.J.; Sanders, T.H.; Payne, G.A.; Lee, L.S.; Klich, M.A. Epidemiology of aflatoxin formation by Aspergillus flavus. Annu. Rev. Phytopathol. 1987, 25, 249–270. [Google Scholar] [CrossRef]
- Xue, H.Q.; O’brian, G.; Payne, G.A.; Isleib, T.G. Evaluation of post-harvest aflatoxin production in peanut germplasm with resistance to seed colonization and pre-harvest aflatoxin contamination. Peanut Sci. 2004, 31, 124–134. [Google Scholar] [CrossRef]
- Scheidegger, K.; Payne, G. Unlocking the secrets behind secondary metabolism: A review of Aspergillus flavus from pathogenicity to functional genomics. J. Toxicol. Toxin Rev. 2003, 22, 423–459. [Google Scholar] [CrossRef]
- Verma, R.J. Aflatoxins cause DNA damage. Int. J. Hum. Genet. 2004, 4, 231–236. [Google Scholar]
- Smith, M.C.; Madec, S.; Coton, E.; Hymery, N. Natural co-occurrence of mycotoxins in foods and feeds and their in vitro combined toxicological effects. Toxins 2016, 8, 94. [Google Scholar] [CrossRef] [PubMed]
- Lamb, M.C.; Sternitzke, D.A. Cost of aflatoxin to the farmer, buying point, and sheller segments of the southeast United States peanut industry. Peanut Sci. 2001, 28, 59–63. [Google Scholar] [CrossRef]
- Cole, R.J.; Hill, R.A.; Blankenship, P.D.; Sanders, T.H. Color mutants of Aspergillus flavus and Aspergillus parasiticus in study of preharvest invasion of peanuts. Appl. Environ. Microbiol. 1986, 52, 1128–1131. [Google Scholar] [PubMed]
- Torres, A.M.; Barros, G.G.; Palacios, S.A.; Chulze, S.N.; Battilani, P. Review on pre- and post-harvest management of peanuts to minimize aflatoxin contamination. Food Res. Int. 2014, 62, 11–19. [Google Scholar] [CrossRef]
- Lynch, R.E.; Wilson, D.M. Enhanced infection of peanut, Arachis hypogaea l., seeds with Aspergillus flavus group fungi due to external scarification of peanut pods by the lesser cornstalk borer, Elasmopalpus lignosellus (zeller). Peanut Sci. 1991, 18, 110–116. [Google Scholar] [CrossRef]
- Timper, P.; Wilson, D.M.; Holbrook, C.C.; Maw, B.W. Relationship between Meloidogyne arenaria and aflatoxin contamination in peanut. J. Nematol. 2004, 36, 167–170. [Google Scholar] [PubMed]
- Styer, C.H.; Cloe, R.J.; Hill, R.A. Inoculation and infection of peanut flowers by Aspergillus flavus. Proc. Am. Peanut Res. Educ. Soc. Inc. 1983, 15, 91. [Google Scholar]
- Holbrook, C.C.; Guo, B.Z.; Wilson, D.M.; Timper, P. The U.S. Breeding program to develop peanut with drought tolerance and reduced aflatoxin contamination. Peanut Sci. 2009, 36, 50–53. [Google Scholar] [CrossRef]
- Clevenger, J.; Marasigan, K.; Liakos, V.; Sobolev, V.; Vellidis, G.; Holbrook, C.; Ozias-Akins, P. RNA sequencing of contaminated seeds reveals the state of the seed permissive for pre-harvest aflatoxin contamination and points to a potential susceptibility factor. Toxins 2016, 8, 317. [Google Scholar] [CrossRef] [PubMed]
- Blankenship, P.; Cole, R.; Sanders, T.; Hill, R. Effect of geocarposphere temperature on pre-harvest colonization of drought-stressed peanuts by Aspergillus flavus and subsequent aflatoxin contamination. Mycopathologia 1984, 85, 69–74. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Lei, Y.; Wan, L.; Yan, L.; Lv, J.; Dai, X.; Ren, X.; Guo, W.; Jiang, H.; Liao, B. Comparative transcript profiling of resistant and susceptible peanut post-harvest seeds in response to aflatoxin production by Aspergillus flavus. BMC Plant Biol. 2016, 16, 54. [Google Scholar] [CrossRef] [PubMed]
- Liang, X.Q.; Holbrook, C.C.; Lynch, R.E.; Guo, B.Z. Beta-1,3-glucanase activity in peanut seed (Arachis hypogaea) is induced by inoculation with Aspergillus flavus and copurifies with a conglutin-like protein. Phytopathology 2005, 95, 506–511. [Google Scholar] [CrossRef] [PubMed]
- Sharaf, A.N.; Gaber, A.; Abdelhadi, A.A.; Ragab, A.I.; Korani, W.A. Induction, characterization and genetic analysis of Aspergillus flavus resistant mutants in Arachis hypogaea. Afr. J. Biotechnol. 2011, 10, 17095–17105. [Google Scholar]
- Lorang, J.M.; Tuori, R.P.; Martinez, J.P.; Sawyer, T.L.; Redman, R.S.; Rollins, J.A.; Wolpert, T.J.; Johnson, K.B.; Rodriguez, R.J.; Dickman, M.B.; et al. Green fluorescent protein is lighting up fungal biology. Appl. Environ. Microbiol. 2001, 67, 1987–1994. [Google Scholar] [CrossRef] [PubMed]
- Nigam, S.N.; Waliyar, F.; Aruna, R.; Reddy, S.V.; Lava Kumar, P.; Craufurd, P.Q.; Diallo, A.T.; Ntare, B.R.; Upadhyaya, H.D. Breeding peanut for resistance to aflatoxin contamination at ICRISAT. Peanut Sci. 2009, 36, 42–49. [Google Scholar] [CrossRef]
- Rao, M.J.V.; Upadhyaya, H.D.; Mehan, V.K.; Nigam, S.N.; McDonald, D.; Reddy, N.S. Registration of peanut germplasm ICGV 88145 and ICGV 89104 resistant to seed infection by Aspergillus flavus. Crop Sci. 1995, 35, 1717. [Google Scholar] [CrossRef]
- Dwivedi, S.L.; Varma, T.S. Molecular diversity among groundnut varieties differing in drought tolerance traits. Int. Arachis Newsl. 2002, 22, 34–36. [Google Scholar]
- Waliyar, F.; Hassan, H.; Bosc, J. Sources of resistance to Aspergillus flavus and aflatoxin contamination in groundnut genotypes in West Africa. Plant Dis. 1994, 78, 704–708. [Google Scholar] [CrossRef]
- Asis, R.; Barrionuevo, D.L.; Giorda, L.M.; Nores, M.L.; Aldao, M.A. Aflatoxin production in six peanut (Arachis hypogaea L.) genotypes infected with Aspergillus flavus and Aspergillus parasiticus, isolated from peanut production areas of cordoba, argentina. J. Agric. Food Chem. 2005, 53, 9274–9280. [Google Scholar] [CrossRef] [PubMed]
- Hamidou, F.; Rathore, A.; Waliyar, F.; Vadez, V. Although drought intensity increases aflatoxin contamination, drought tolerance does not lead to less aflatoxin contamination. Field Crops Res. 2014, 156, 103–110. [Google Scholar] [CrossRef]
- Mehan, V.K. Screening groundnuts for resistance to seed invasion by Aspergillus flavus and to aflatoxin production. In Aflatoxin Contamination of Groundnut: Proceedings of the International Workshop, 6–9 Oct. 1987, ICRISAT Center, India; Hall, S.D., Ed.; ICRISAT: Andhar Pradesh, India, 1989; pp. 323–334. [Google Scholar]
- Liang, X.Q.; Luo, M.; Holbrook, C.C.; Guo, B.Z. Storage protein profiles in spanish and runner market type peanuts and potential markers. BMC Plant Biol. 2006, 6, 24. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Scharfenstein, L.L.; Carter-Wientjes, C.; Chang, P.K.; Zhang, D.; Meng, X.; Yu, J. Lack of aflatoxin production by Aspergillus flavus is associated with reduced fungal growth and delayed expression of aflatoxin pathway genes. World Mycotoxin J. 2015, 8, 335–340. [Google Scholar] [CrossRef]
- Holbrook, C., Jr.; Cantonwine, E.; Sullivan, D.; Guo, B.; Wilson, D.; Dong, W. Development of Peanut Germplasm with Improved Drought Tolerance; American Peanut Research and Education Society, Inc.: Birmingham, AL, USA, 2007; p. 21. [Google Scholar]
- Holbrook, C.C.; Timper, P.; Culbreath, A.K.; Kvien, C.K. Registration of ‘Tifguard’ peanut. J. Plant Regist. 2008, 2, 92–94. [Google Scholar] [CrossRef]
- Beute, M.K.; Wynne, J.C.; Emery, D.A. Registration of NC3033 peanut germplasm. Crop Sci. 1976, 16, 887. [Google Scholar] [CrossRef]
- Holbrook, C.C.; Culbreath, A.K. Registration of ‘Tifrunner’ peanut. J. Plant Regist. 2007, 1, 124. [Google Scholar] [CrossRef]
- Gorbet, D.W.; Tillman, B.L. Registration of ‘Florida-07’ peanut. J. Plant Regist. 2009, 3, 14–18. [Google Scholar] [CrossRef]
- Luis, J.M.; Ozias-Akins, P.; Holbrook, C.C.; Kemerait, R.C., Jr.; Snider, J.L.; Liakos, V. Phenotyping peanut genotypes for drought tolerance. Peanut Sci. 2016, 43, 36–48. [Google Scholar] [CrossRef]
- Klich, M.A.; Lee, L.S. Seed viability and aflatoxin production in individual cottonseed naturally contaminated with Aspergillus flavus. J. Am. Oil Chem. Soc. 1982, 59, 545. [Google Scholar] [CrossRef]
- Asis, R.; Muller, V.; Barrionuevo, D.L.; Araujo, S.A.; Aldao, M.A. Analysis of protease activity in Aspergillus flavus and A. parasiticus on peanut seed infection and aflatoxin contamination. Eur. J. Plant Pathol 2009, 124, 391–403. [Google Scholar] [CrossRef]
- Xue, H.Q.; Isleib, T.G.; Payne, G.A.; Novitzky, W.F.; Obrain, G. Aflatoxin production in peanut lines selected to represent a range of linoleic acid concentrations. J. Food Prot. 2005, 68, 126–132. [Google Scholar] [CrossRef] [PubMed]
- Lansden, J.A. Aflatoxin inhibition and fungistasis by peanut tannins. Peanut Sci. 1982, 9, 17–20. [Google Scholar] [CrossRef]
- Huang, S.C.; Yen, G.-C.; Chang, L.-W.; Yen, W.-J.; Duh, P.-D. Identification of an antioxidant, ethyl protocatechuate, in peanut seed testa. J. Agric. Food Chem. 2003, 51, 2380–2383. [Google Scholar] [CrossRef] [PubMed]
- Xue, H.Q.; Isleib, T.G.; Payne, G.A.; Wilson, R.F.; Novitzky, W.P.; O’Brian, G. Comparison of aflatoxin production in normal- and high-oleic backcross-derived peanut lines. Plant Dis. 2003, 87, 1360–1365. [Google Scholar] [CrossRef]
- Qin, H.; Feng, S.; Chen, C.; Guo, Y.; Knapp, S.; Culbreath, A.; He, G.; Wang, M.L.; Zhang, X.; Holbrook, C.C. An integrated genetic linkage map of cultivated peanut (arachis hypogaea L.) constructed from two RIL populations. Theor. Appl. Genet. 2012, 124, 653–664. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Pandey, M.K.; Qiao, L.; Qin, H.; Culbreath, A.K.; He, G.; Varshney, R.K.; Scully, B.T.; Guo, B. Genetic mapping and quantitative trait loci analysis for disease resistance using F2 and F5 generation-based genetic maps derived from ‘Tifrunner’בGT-C20’ in peanut. Plant Genome 2013, 6, 1–10. [Google Scholar] [CrossRef]
- Burow, G.B.; Gardner, H.W.; Keller, N.P. A peanut seed lipoxygenase responsive to Aspergillus colonization. Plant Mol. Biol. 2000, 42, 689–701. [Google Scholar] [CrossRef] [PubMed]
- Tsitsigiannis, D.I.; Kunze, S.; Willis, D.K.; Feussner, I.; Keller, N.P. Aspergillus infection inhibits the expression of peanut 13S-hpode-forming seed lipoxygenases. Mol. Plant-Microbe Interact. 2005, 18, 1081–1089. [Google Scholar] [CrossRef] [PubMed]
- Muller, V.; Ame, M.V.; Carrari, V.; Gieco, J.; Asis, R. Lipoxygenase activation in peanut seed cultivars resistant and susceptible to Aspergillus parasiticus colonization. Phytopathology 2014, 104, 1340–1348. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Dickman, M.B. Bcl-2 family members localize to tobacco chloroplasts and inhibit programmed cell death induced by chloroplast-targeted herbicides. J. Exp. Bot. 2004, 55, 2617–2623. [Google Scholar] [CrossRef] [PubMed]
- Anderson, W.F.; Holbrook, C.C.; Wilson, D.M.; Matheron, M.E. Evaluation of pre-harvest aflatoxin contamination in several potentially resistant peanut genotypes. Peanut Sc. 1995, 22, 29–32. [Google Scholar] [CrossRef]
- Holbrook, C.C.; Kvien, C.K.; Rucker, K.S.; Wilson, D.M.; Hook, J.E.; Matheron, M.E. Pre-harvest aflatoxin contamination in drought-tolerant and drought-intolerant peanut genotypes. Peanut Sci. 2000, 27, 45–48. [Google Scholar] [CrossRef]
- Core Team, R. R: A Language and Environment for Statistical Computing; The R Foundation for Statistical Computing: Vienna, Austria, 2011. [Google Scholar]
| Genotype | Year 2014 | Year 2015 | ||
|---|---|---|---|---|
| Aflatoxin Range (ppb) | Average Aflatoxin B (ppb) | Aflatoxin Range (ppb) | Average Aflatoxin B (ppb) | |
| ICG 1471 | 5 to 35 | 15 | 2 to 7 | 3 |
| Florida-07 | 5 to 18 | 11 | 23 to 641 | 214 |
| Tifrunner | 9 to 48 | 26 | 4 to 1200 | 381 |
| A69 | 3 to 230 | 61 | 3 to 54 | 20 |
| A72 | 3 to 679 | 216 | 3 to 2100 | 425 |
| ICGV88145 | 1 to 1034 | 226 | 1 to 11 | 4 |
| C76-16 | 25 to 535 | 246 | 22 to 599 | 200 |
| Tifguard | 12 to 734 | 272 | n/a | n/a |
| NC 3033 | 220 to 360 | 303 | n/a | n/a |
| GT-C20 | 220 to 39,000 | 4581 | n/a | n/a |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Korani, W.A.; Chu, Y.; Holbrook, C.; Clevenger, J.; Ozias-Akins, P. Genotypic Regulation of Aflatoxin Accumulation but Not Aspergillus Fungal Growth upon Post-Harvest Infection of Peanut (Arachis hypogaea L.) Seeds. Toxins 2017, 9, 218. https://doi.org/10.3390/toxins9070218
Korani WA, Chu Y, Holbrook C, Clevenger J, Ozias-Akins P. Genotypic Regulation of Aflatoxin Accumulation but Not Aspergillus Fungal Growth upon Post-Harvest Infection of Peanut (Arachis hypogaea L.) Seeds. Toxins. 2017; 9(7):218. https://doi.org/10.3390/toxins9070218
Chicago/Turabian StyleKorani, Walid Ahmed, Ye Chu, Corley Holbrook, Josh Clevenger, and Peggy Ozias-Akins. 2017. "Genotypic Regulation of Aflatoxin Accumulation but Not Aspergillus Fungal Growth upon Post-Harvest Infection of Peanut (Arachis hypogaea L.) Seeds" Toxins 9, no. 7: 218. https://doi.org/10.3390/toxins9070218
APA StyleKorani, W. A., Chu, Y., Holbrook, C., Clevenger, J., & Ozias-Akins, P. (2017). Genotypic Regulation of Aflatoxin Accumulation but Not Aspergillus Fungal Growth upon Post-Harvest Infection of Peanut (Arachis hypogaea L.) Seeds. Toxins, 9(7), 218. https://doi.org/10.3390/toxins9070218





