Overlooked Short Toxin-Like Proteins: A Shortcut to Drug Design
Abstract
1. Introduction
2. Results and Discussion
2.1. Thousands of Toxin-Like Secreted Short Proteins in Insects
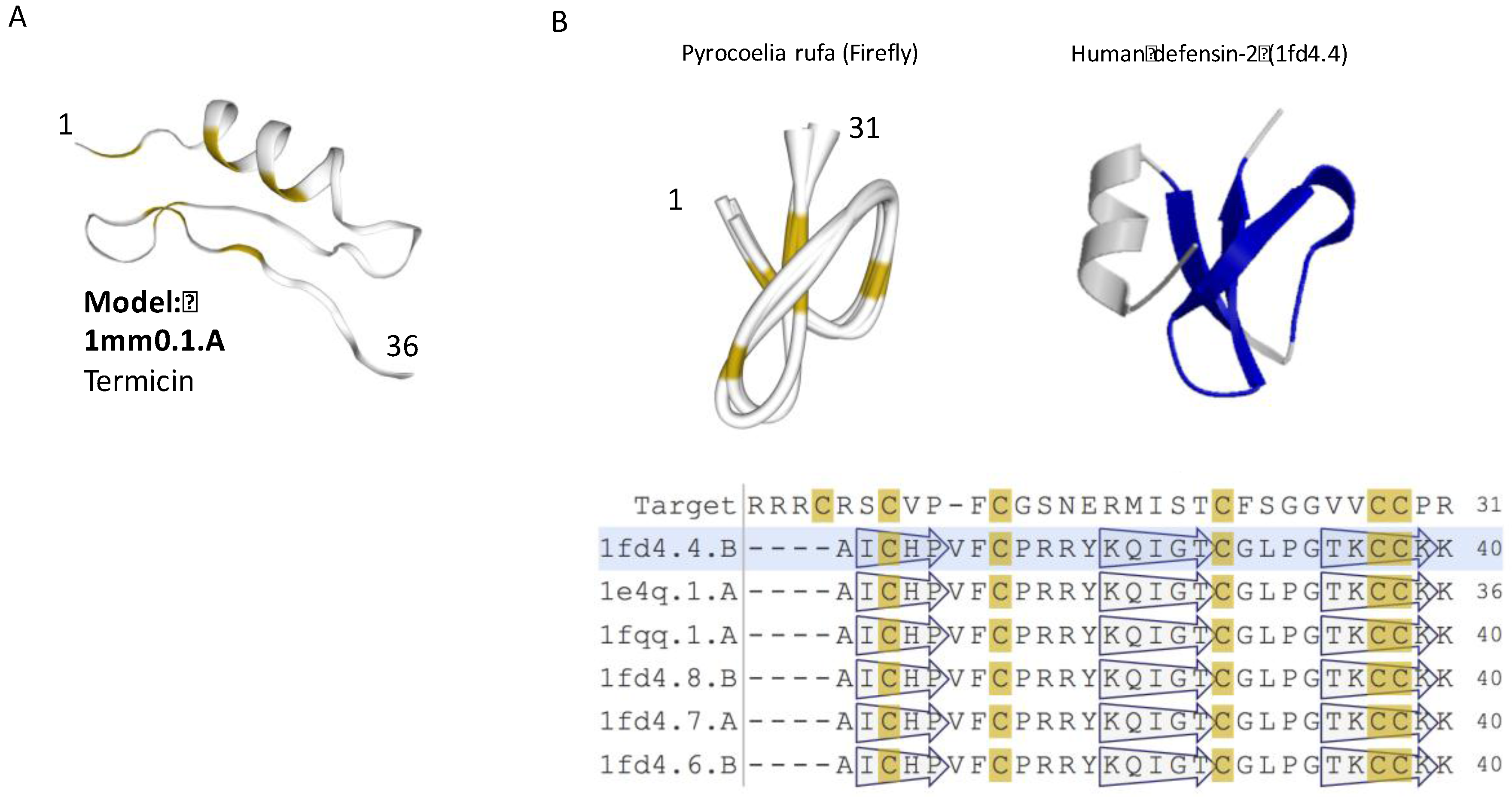
2.2. Most iTOLIP Mini-Proteins Resemble Antibacterial and Antifungal Peptides
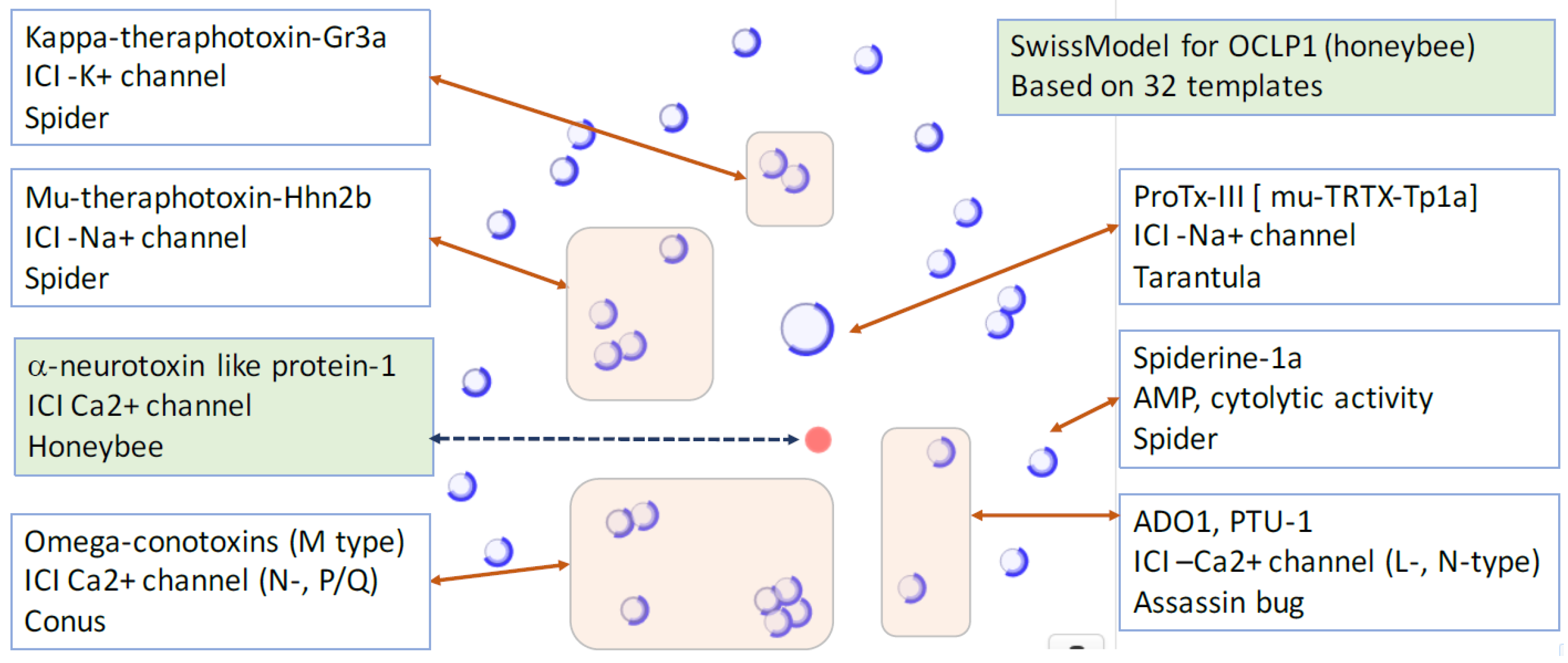
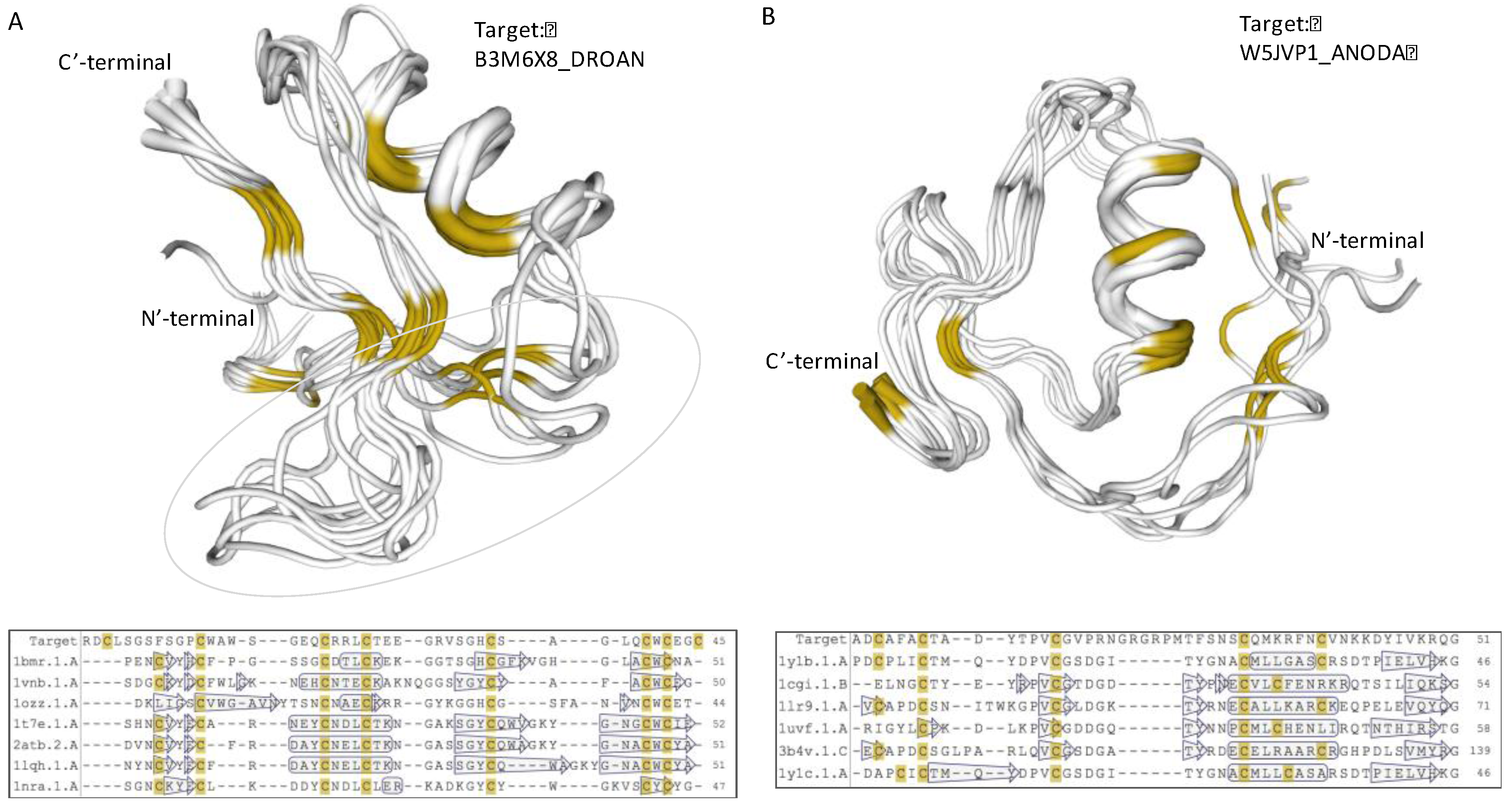
2.3. iTOLIPs as Ion Channel Inhibitors
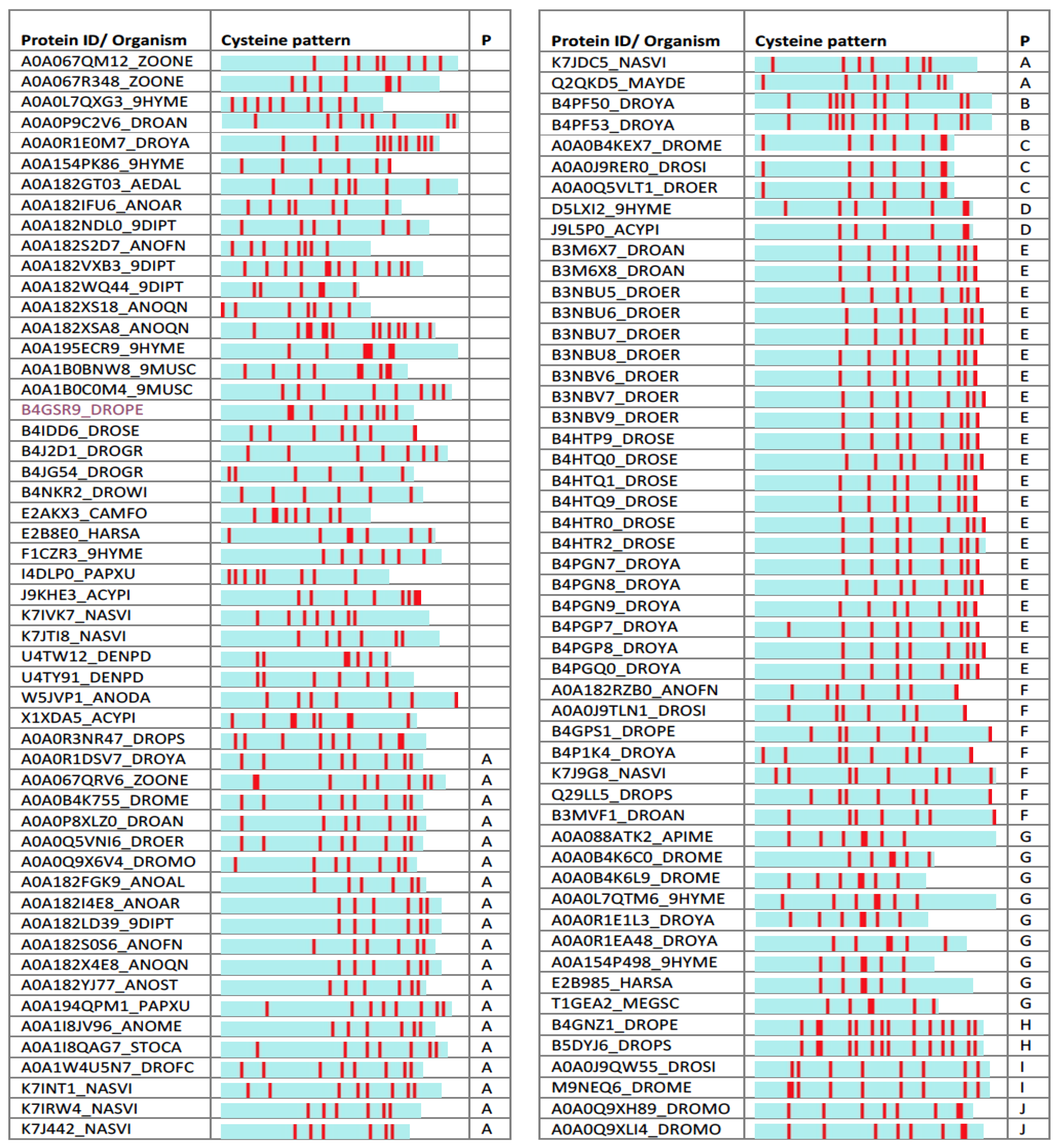
2.4. Uncharacterized iTOLIPs Reveal New Cysteine-Rich Patterns
3. Materials and Methods
3.1. Protein Databases
3.2. Bioinformatics Analysis Tools
3.3. ClanTox Prediction and Scoring
4. Conclusions
Supplementary Materials
Author Contributions
Conflicts of Interest
Abbreviations
| AMP | antimicrobial peptides |
| CSαβ | cysteine-stabilized α-helical and β-sheet |
| ClanTox | classifier of animal toxins |
| CRISP | cysteine rich short proteins |
| ICI | ion channel inhibitor |
| DRS | Drosomycin |
| nAChR | nicotinic acetylcholine receptors |
| OCLP | omega conotoxin-like protein |
| TFP | three-finger proteins |
| iTOLIP | insect toxin-like proteins |
References
- Adermann, K.; John, H.; Standker, L.; Forssmann, W.G. Exploiting natural peptide diversity: Novel research tools and drug leads. Curr. Opin. Biotechnol. 2004, 15, 599–606. [Google Scholar] [CrossRef] [PubMed]
- Alonso, D.; Khalil, Z.; Satkunanthan, N.; Livett, B.G. Drugs from the sea: Conotoxins as drug leads for neuropathic pain and other neurological conditions. Mini Rev. Med. Chem. 2003, 3, 785–787. [Google Scholar] [CrossRef] [PubMed]
- King, G.F. Venoms as a platform for human drugs: Translating toxins into therapeutics. Expert Opin. Biol. Ther. 2011, 11, 1469–1484. [Google Scholar] [CrossRef] [PubMed]
- Proksch, P.; Edrada, R.; Ebel, R. Drugs from the seas-current status and microbiological implications. Appl. Microbiol. Biotechnol. 2002, 59, 125–134. [Google Scholar] [PubMed]
- Bock, J.E.; Gavenonis, J.; Kritzer, J.A. Getting in shape: Controlling peptide bioactivity and bioavailability using conformational constraints. ACS Chem. Biol. 2013, 8, 488–499. [Google Scholar] [CrossRef] [PubMed]
- Vetter, I.; Davis, J.L.; Rash, L.D.; Anangi, R.; Mobli, M.; Alewood, P.F.; Lewis, R.J.; King, G.F. Venomics: A new paradigm for natural products-based drug discovery. Amino Acids 2011, 40, 15–28. [Google Scholar] [CrossRef] [PubMed]
- Bulaj, G. Integrating the discovery pipeline for novel compounds targeting ion channels. Curr. Opin. Chem. Biol. 2008, 12, 441–447. [Google Scholar] [CrossRef] [PubMed]
- Harvey, A.L. Toxins and drug discovery. Toxicon 2014, 92, 193–200. [Google Scholar] [CrossRef] [PubMed]
- Fry, B.G.; Roelants, K.; Champagne, D.E.; Scheib, H.; Tyndall, J.D.; King, G.F.; Nevalainen, T.J.; Norman, J.A.; Lewis, R.J.; Norton, R.S.; et al. The toxicogenomic multiverse: Convergent recruitment of proteins into animal venoms. Annu. Rev. Genom. Hum. Genet. 2009, 10, 483–511. [Google Scholar] [CrossRef] [PubMed]
- Wong, E.S.; Belov, K. Venom evolution through gene duplications. Gene 2012, 496, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, N.; Morpurgo, N.; Linial, M. Novel families of toxin-like peptides in insects and mammals: A computational approach. J. Mol. Biol. 2007, 369, 553–566. [Google Scholar] [CrossRef] [PubMed]
- Fry, B.G.; Wuster, W.; Kini, R.M.; Brusic, V.; Khan, A.; Venkataraman, D.; Rooney, A.P. Molecular evolution and phylogeny of elapid snake venom three-finger toxins. J. Mol. Evol. 2003, 57, 110–129. [Google Scholar] [CrossRef] [PubMed]
- Craik, D.J.; Fairlie, D.P.; Liras, S.; Price, D. The future of peptide-based drugs. Chem. Biol. Drug Des. 2013, 81, 136–147. [Google Scholar] [CrossRef] [PubMed]
- Han, T.S.; Teichert, R.W.; Olivera, B.M.; Bulaj, G. Conus venoms—A rich source of peptide-based therapeutics. Curr. Pharm. Des. 2008, 14, 2462–2479. [Google Scholar] [CrossRef] [PubMed]
- Lavergne, V.; Harliwong, I.; Jones, A.; Miller, D.; Taft, R.J.; Alewood, P.F. Optimized deep-targeted proteotranscriptomic profiling reveals unexplored conus toxin diversity and novel cysteine frameworks. Proc. Natl. Acad. Sci. USA 2015, 112, E3782–E3791. [Google Scholar] [CrossRef] [PubMed]
- Drabeck, D.H.; Dean, A.M.; Jansa, S.A. Why the honey badger don’t care: Convergent evolution of venom-targeted nicotinic acetylcholine receptors in mammals that survive venomous snake bites. Toxicon 2015, 99, 68–72. [Google Scholar] [CrossRef] [PubMed]
- Zambelli, V.; Pasqualoto, K.; Picolo, G.; Chudzinski-Tavassi, A.; Cury, Y. Harnessing the knowledge of animal toxins to generate drugs. Pharmacol. Res. 2016, 112, 30–36. [Google Scholar] [CrossRef] [PubMed]
- Fry, B.G. From genome to “venome” Molecular origin and evolution of the snake venom proteome inferred from phylogenetic analysis of toxin sequences and related body proteins. Genome Res. 2005, 15, 403–420. [Google Scholar] [CrossRef] [PubMed]
- Casewell, N.R.; Wuster, W.; Vonk, F.J.; Harrison, R.A.; Fry, B.G. Complex cocktails: The evolutionary novelty of venoms. Trends Ecol. Evol. 2013, 28, 219–229. [Google Scholar] [CrossRef] [PubMed]
- Sitprija, V.; Sitprija, S. Renal effects and injury induced by animal toxins. Toxicon 2012, 60, 943–953. [Google Scholar] [CrossRef] [PubMed]
- Corzo, G.; Villegas, E.; Gomez-Lagunas, F.; Possani, L.D.; Belokoneva, O.S.; Nakajima, T. Oxyopinins, large amphipathic peptides isolated from the venom of the wolf spider oxyopes kitabensis with cytolytic properties and positive insecticidal cooperativity with spider neurotoxins. J. Biol. Chem. 2002, 277, 23627–23637. [Google Scholar] [CrossRef] [PubMed]
- Edwards, L.P.; Whitter, E.; Hessinger, D.A. Apparent membrane pore-formation by portuguese man-of-war (physalia physalis) venom in intact cultured cells. Toxicon 2002, 40, 1299–1305. [Google Scholar] [CrossRef]
- Slotta, K.H.; Gonzalez, J.; Roth, S. The direct and indirect hemolytic factors from animal venoms. In RUSSELL Animal Toxins; Elsevier: Amsterdam, The Netherlands, 2016; pp. 369–377. [Google Scholar]
- Estrada, G.; Villegas, E.; Corzo, G. Spider venoms: A rich source of acylpolyamines and peptides as new leads for cns drugs. Nat. Prod. Rep. 2007, 24, 145–161. [Google Scholar] [CrossRef] [PubMed]
- Petricevich, V.L. Scorpion venom and the inflammatory response. Mediat. Inflamm. 2010, 2010, 903295. [Google Scholar] [CrossRef] [PubMed]
- Gibbons, A.; Dean, B. The cholinergic system: An emerging drug target for schizophrenia. Curr. Pharm. Des. 2016, 22, 2124–2133. [Google Scholar] [CrossRef] [PubMed]
- Tirosh, Y.; Ofer, D.; Eliyahu, T.; Linial, M. Short toxin-like proteins attack the defense line of innate immunity. Toxins 2013, 5, 1314–1331. [Google Scholar] [CrossRef] [PubMed]
- Tsetlin, V.I. Three-finger snake neurotoxins and ly6 proteins targeting nicotinic acetylcholine receptors: Pharmacological tools and endogenous modulators. Trends Pharmacol. Sci. 2015, 36, 109–123. [Google Scholar] [CrossRef] [PubMed]
- Kini, R.M. Evolution of three-finger toxins—A versatile mini protein scaffold. Acta Chim. Slovenica 2011, 58, 693–701. [Google Scholar]
- Ibanez-Tallon, I.; Miwa, J.M.; Wang, H.L.; Adams, N.C.; Crabtree, G.W.; Sine, S.M.; Heintz, N. Novel modulation of neuronal nicotinic acetylcholine receptors by association with the endogenous prototoxin lynx1. Neuron 2002, 33, 893–903. [Google Scholar] [CrossRef]
- Chimienti, F.; Hogg, R.C.; Plantard, L.; Lehmann, C.; Brakch, N.; Fischer, J.; Huber, M.; Bertrand, D.; Hohl, D. Identification of slurp-1 as an epidermal neuromodulator explains the clinical phenotype of mal de meleda. Hum. Mol. Genet. 2003, 12, 3017–3024. [Google Scholar] [CrossRef] [PubMed]
- Kalia, J.; Milescu, M.; Salvatierra, J.; Wagner, J.; Klint, J.K.; King, G.F.; Olivera, B.M.; Bosmans, F. From foe to friend: Using animal toxins to investigate ion channel function. J. Mol. Biol. 2015, 427, 158–175. [Google Scholar] [CrossRef] [PubMed]
- Mouhat, S.; Andreotti, N.; Jouirou, B.; Sabatier, J.-M. Animal toxins acting on voltage-gated potassium channels. Curr. Pharm. Des. 2008, 14, 2503–2518. [Google Scholar] [CrossRef] [PubMed]
- Norton, R.S. Structure and function of peptide and protein toxins from marine organisms. J. Toxicol. Toxin Rev. 1998, 17, 99–130. [Google Scholar] [CrossRef]
- Terlau, H.; Olivera, B.M. Conus venoms: A rich source of novel ion channel-targeted peptides. Physiol. Rev. 2004, 84, 41–68. [Google Scholar] [CrossRef] [PubMed]
- Quintero-Hernández, V.; Jiménez-Vargas, J.; Gurrola, G.; Valdivia, H.; Possani, L. Scorpion venom components that affect ion-channels function. Toxicon 2013, 76, 328–342. [Google Scholar] [CrossRef] [PubMed]
- Bohlen, C.J.; Chesler, A.T.; Sharif-Naeini, R.; Medzihradszky, K.F.; Zhou, S.; King, D.; Sánchez, E.E.; Burlingame, A.L.; Basbaum, A.I.; Julius, D. A heteromeric texas coral snake toxin targets acid-sensing ion channels to produce pain. Nature 2011, 479, 410. [Google Scholar] [CrossRef] [PubMed]
- Guo, M.; Teng, M.; Niu, L.; Liu, Q.; Huang, Q.; Hao, Q. Crystal structure of the cysteine-rich secretory protein stecrisp reveals that the cysteine-rich domain has a K+ channel inhibitor-like fold. J. Biol. Chem. 2005, 280, 12405–12412. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, G.M.; Orta, G.; Reddy, T.; Koppers, A.J.; Martínez-López, P.; de la Vega-Beltràn, J.L.; Lo, J.C.; Veldhuis, N.; Jamsai, D.; McIntyre, P. Cysteine-rich secretory protein 4 is an inhibitor of transient receptor potential m8 with a role in establishing sperm function. Proc. Natl. Acad. Sci. USA 2011, 108, 7034–7039. [Google Scholar] [CrossRef] [PubMed]
- Diochot, S.; Salinas, M.; Baron, A.; Escoubas, P.; Lazdunski, M. Peptides inhibitors of acid-sensing ion channels. Toxicon 2007, 49, 271–284. [Google Scholar] [CrossRef] [PubMed]
- Mouhat, S.; Jouirou, B.; Mosbah, A.; De Waard, M.; Sabatier, J.-M. Diversity of folds in animal toxins acting on ion channels. Biochem. J. 2004, 378, 717–726. [Google Scholar] [CrossRef] [PubMed]
- Ohno, M.; Menez, R.; Ogawa, T.; Danse, J.M.; Shimohigashi, Y.; Fromen, C.; Ducancel, F.; Zinn-Justin, S.; Le Du, M.H.; Boulain, J.C.; et al. Molecular evolution of snake toxins: Is the functional diversity of snake toxins associated with a mechanism of accelerated evolution? Prog. Nucl. Acid Res. Mol. Biol. 1998, 59, 307–364. [Google Scholar]
- Chang, L.-S. Genetic diversity in snake venom three-finger proteins and phospholipase a2 enzymes. Toxin Rev. 2007, 26, 143–167. [Google Scholar] [CrossRef]
- Casewell, N.R.; Wagstaff, S.C.; Harrison, R.A.; Renjifo, C.; Wüster, W. Domain loss facilitates accelerated evolution and neofunctionalization of duplicate snake venom metalloproteinase toxin genes. Mol. Biol. Evol. 2011, 28, 2637–2649. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, A.; Lee, A.; Campbell, E.; MacKinnon, R. Structure of a pore-blocking toxin in complex with a eukaryotic voltage-dependent K+ channel. Elife 2013, 2, e00594. [Google Scholar] [CrossRef] [PubMed]
- Strix, G. A toxin against pain. Sci. Am. 2005, 292, 88–93. [Google Scholar] [CrossRef]
- Góngora-Benítez, M.; Tulla-Puche, J.; Albericio, F. Multifaceted roles of disulfide bonds. Peptides as therapeutics. Chem. Rev. 2013, 114, 901–926. [Google Scholar] [CrossRef] [PubMed]
- Herzig, V.; King, G.F. The cystine knot is responsible for the exceptional stability of the insecticidal spider toxin ω-hexatoxin-hv1a. Toxins 2015, 7, 4366–4380. [Google Scholar] [CrossRef] [PubMed]
- Kuzmenkov, A.I.; Fedorova, I.M.; Vassilevski, A.A.; Grishin, E.V. Cysteine-rich toxins from lachesana tarabaevi spider venom with amphiphilic c-terminal segments. Biochim. Biophys. Acta 2013, 1828, 724–731. [Google Scholar] [CrossRef] [PubMed]
- Lavergne, V.; Alewood, P.F.; Mobli, M.; King, G.F. The structural universe of disulfide-rich venom peptides. In Venoms to Drugs: Venoms as a Source for the Development of Human Therapeutics; Royal Society of Chemistry: London, UK, 2015. [Google Scholar]
- Avrutina, O. Synthetic cystine-knot miniproteins—Valuable scaffolds for polypeptide engineering. Adv. Exp. Med. Biol. 2016, 917, 121–144. [Google Scholar] [PubMed]
- Rappoport, N.; Karsenty, S.; Stern, A.; Linial, N.; Linial, M. Protonet 6.0: Organizing 10 million protein sequences in a compact hierarchical family tree. Nucl. Acids Res. 2012, 40, D313–D320. [Google Scholar] [CrossRef] [PubMed]
- Ofer, D.; Rappoport, N.; Linial, M. The little known universe of short proteins in insects: A machine learning approach. In Short Views on Insect Genomics and Proteomics; Springer: Berlin, Germany, 2015; pp. 177–202. [Google Scholar]
- Werren, J.H.; Richards, S.; Desjardins, C.A.; Niehuis, O.; Gadau, J.; Colbourne, J.K.; Group, N.G.W. Functional and evolutionary insights from the genomes of three parasitoid nasonia species. Science 2010, 327, 343–348. [Google Scholar] [CrossRef] [PubMed]
- Nygaard, S.; Zhang, G.; Schiøtt, M.; Li, C.; Wurm, Y.; Hu, H.; Zhou, J.; Ji, L.; Qiu, F.; Rasmussen, M. The genome of the leaf-cutting ant acromyrmex echinatior suggests key adaptations to advanced social life and fungus farming. Genome Res. 2011, 21, 1339–1348. [Google Scholar] [CrossRef] [PubMed]
- Rappoport, N.; Linial, M. Trends in genome dynamics among major orders of insects revealed through variations in protein families. BMC Genom. 2015, 16, 583. [Google Scholar] [CrossRef] [PubMed]
- Naamati, G.; Askenazi, M.; Linial, M. Clantox: A classifier of short animal toxins. Nucleic Acids Res. 2009, 37, W363–W368. [Google Scholar] [CrossRef] [PubMed]
- Radivojac, P.; Clark, W.T.; Oron, T.R.; Schnoes, A.M.; Wittkop, T.; Sokolov, A.; Graim, K.; Funk, C.; Verspoor, K.; Ben-Hur, A. A large-scale evaluation of computational protein function prediction. Nat. Methods 2013, 10, 221–227. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, N.; Linial, M. Automatic detection of false annotations via binary property clustering. BMC Bioinform. 2005, 6, 46. [Google Scholar] [CrossRef] [PubMed]
- Ofer, D.; Linial, M. Neuropid: A predictor for identifying neuropeptide precursors from metazoan proteomes. Bioinformatics 2013, 30, 931–940. [Google Scholar] [CrossRef] [PubMed]
- Tirosh, Y.; Linial, I.; Askenazi, M.; Linial, M. Short toxin-like proteins abound in cnidaria genomes. Toxins 2012, 4, 1367–1384. [Google Scholar] [CrossRef] [PubMed]
- Tassanakajon, A.; Somboonwiwat, K.; Amparyup, P. Sequence diversity and evolution of antimicrobial peptides in invertebrates. Dev. Comp. Immunol. 2015, 48, 324–341. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Yuan, K.; Zhang, R.; Ren, X.; Liu, X.; Zhao, S.; Wang, D. Cloning and purification of the first termicin-like peptide from the cockroach eupolyphaga sinensis. J. Venom. Anim. Toxins Incl. Trop. Dis. 2016, 22, 5. [Google Scholar] [CrossRef] [PubMed]
- Fjell, C.D.; Hiss, J.A.; Hancock, R.E.; Schneider, G. Designing antimicrobial peptides: Form follows function. Nat. Rev. Drug Discov. 2011, 11, 37–51. [Google Scholar] [CrossRef] [PubMed]
- Froy, O.; Gurevitz, M. Arthropod defensins illuminate the divergence of scorpion neurotoxins. J. Pept. Sci. 2004, 10, 714–718. [Google Scholar] [CrossRef] [PubMed]
- Froy, O.; Gurevitz, M. New insight on scorpion divergence inferred from comparative analysis of toxin structure, pharmacology and distribution. Toxicon 2003, 42, 549–555. [Google Scholar] [CrossRef]
- Bun Ng, T.; Chi Fai Cheung, R.; Ho Wong, J.; Juan Ye, X. Antimicrobial activity of defensins and defensin-like peptides with special emphasis on those from fungi and invertebrate animals. Curr. Protein Pept. Sci. 2013, 14, 515–531. [Google Scholar]
- Whittington, C.M.; Papenfuss, A.T.; Bansal, P.; Torres, A.M.; Wong, E.S.; Deakin, J.E.; Graves, T.; Alsop, A.; Schatzkamer, K.; Kremitzki, C. Defensins and the convergent evolution of platypus and reptile venom genes. Genome Res. 2008, 18, 986–994. [Google Scholar] [CrossRef] [PubMed]
- Varkey, J.; Singh, S.; Nagaraj, R. Antibacterial activity of linear peptides spanning the carboxy-terminal beta-sheet domain of arthropod defensins. Peptides 2006, 27, 2614–2623. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.; Li, W.; Jiang, D.; Zeng, X. Evidence for the existence of insect defensin-like peptide in scorpion venom. IUBMB Life 2000, 50, 57–61. [Google Scholar] [PubMed]
- Gao, B.; Zhu, S. The drosomycin multigene family: Three-disulfide variants from drosophila takahashii possess antibacterial activity. Sci. Rep. 2016, 6, 32175. [Google Scholar] [CrossRef] [PubMed]
- Deng, X.J.; Yang, W.Y.; Huang, Y.D.; Cao, Y.; Wen, S.Y.; Xia, Q.Y.; Xu, P. Gene expression divergence and evolutionary analysis of the drosomycin gene family in drosophila melanogaster. J. Biomed. Biotechnol. 2009, 2009, 315423. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Su, M.; Hamann, M.T.; Bowling, J.J.; Kim, H.S.; Jung, J.H. Solution structure of a sponge-derived cystine knot peptide and its notable stability. J. Nat. Prod. 2014, 77, 304–310. [Google Scholar] [CrossRef] [PubMed]
- Ovchinnikova, T.V.; Balandin, S.V.; Aleshina, G.M.; Tagaev, A.A.; Leonova, Y.F.; Krasnodembsky, E.D.; Men’shenin, A.V.; Kokryakov, V.N. Aurelin, a novel antimicrobial peptide from jellyfish aurelia aurita with structural features of defensins and channel-blocking toxins. Biochem. Biophys. Res. Commun. 2006, 348, 514–523. [Google Scholar] [CrossRef] [PubMed]
- Cohen, L.; Moran, Y.; Sharon, A.; Segal, D.; Gordon, D.; Gurevitz, M. Drosomycin, an innate immunity peptide of drosophila melanogaster, interacts with the fly voltage-gated sodium channel. J. Biol. Chem. 2009, 284, 23558–23563. [Google Scholar] [CrossRef] [PubMed]
- Stehling, E.G.; Sforca, M.L.; Zanchin, N.I.; Oyama, S., Jr.; Pignatelli, A.; Belluzzi, O.; Polverini, E.; Corsini, R.; Spisni, A.; Pertinhez, T.A. Looking over toxin-k(+) channel interactions. Clues from the structural and functional characterization of alpha-ktx toxin tc32, a kv1.3 channel blocker. Biochemistry 2012, 51, 1885–1894. [Google Scholar] [CrossRef] [PubMed]
- Deuis, J.R.; Dekan, Z.; Wingerd, J.S.; Smith, J.J.; Munasinghe, N.R.; Bhola, R.F.; Imlach, W.L.; Herzig, V.; Armstrong, D.A.; Rosengren, K.J.; et al. Pharmacological characterisation of the highly nav1.7 selective spider venom peptide pn3a. Sci. Rep. 2017, 7, 40883. [Google Scholar] [CrossRef] [PubMed]
- Jablonsky, M.J.; Jackson, P.L.; Krishna, N.R. Solution structure of an insect-specific neurotoxin from the new world scorpion centruroides sculpturatus ewing. Biochemistry 2001, 40, 8273–8282. [Google Scholar] [CrossRef] [PubMed]
- Krimm, I.; Gilles, N.; Sautiere, P.; Stankiewicz, M.; Pelhate, M.; Gordon, D.; Lancelin, J.M. Nmr structures and activity of a novel alpha-like toxin from the scorpion leiurus quinquestriatus hebraeus. J. Mol. Biol. 1999, 285, 1749–1763. [Google Scholar] [CrossRef] [PubMed]
- Mourao, C.B.; Schwartz, E.F. Protease inhibitors from marine venomous animals and their counterparts in terrestrial venomous animals. Mar. Drugs 2013, 11, 2069–2112. [Google Scholar] [CrossRef] [PubMed]
- Boutet, E.; Lieberherr, D.; Tognolli, M.; Schneider, M.; Bansal, P.; Bridge, A.J.; Poux, S.; Bougueleret, L.; Xenarios, I. Uniprotkb/swiss-prot, the manually annotated section of the uniprot knowledgebase: How to use the entry view. In Plant Bioinformatics: Methods and Protocols; Spinger: Berlin, Germany, 2016; pp. 23–54. [Google Scholar]
- Bienert, S.; Waterhouse, A.; de Beer, T.A.; Tauriello, G.; Studer, G.; Bordoli, L.; Schwede, T. The swiss-model repository-new features and functionality. Nucleic Acids Res. 2017, 45, D313–D319. [Google Scholar] [CrossRef] [PubMed]
- Rose, P.W.; Prlić, A.; Altunkaya, A.; Bi, C.; Bradley, A.R.; Christie, C.H.; Costanzo, L.D.; Duarte, J.M.; Dutta, S.; Feng, Z. The rcsb protein data bank: Integrative view of protein, gene and 3d structural information. Nucleic Acids Res. 2016, 45, D271–D281. [Google Scholar] [PubMed]
- Petersen, T.N.; Brunak, S.; von Heijne, G.; Nielsen, H. Signalp 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar] [CrossRef] [PubMed]
- Remmert, M.; Biegert, A.; Hauser, A.; Söding, J. Hhblits: Lightning-fast iterative protein sequence searching by hmm-hmm alignment. Nat. Methods 2012, 9, 173–175. [Google Scholar] [CrossRef] [PubMed]
- Naamati, G.; Askenazi, M.; Linial, M. A predictor for toxin-like proteins exposes cell modulator candidates within viral genomes. Bioinformatics 2010, 26, i482–i488. [Google Scholar] [CrossRef] [PubMed]
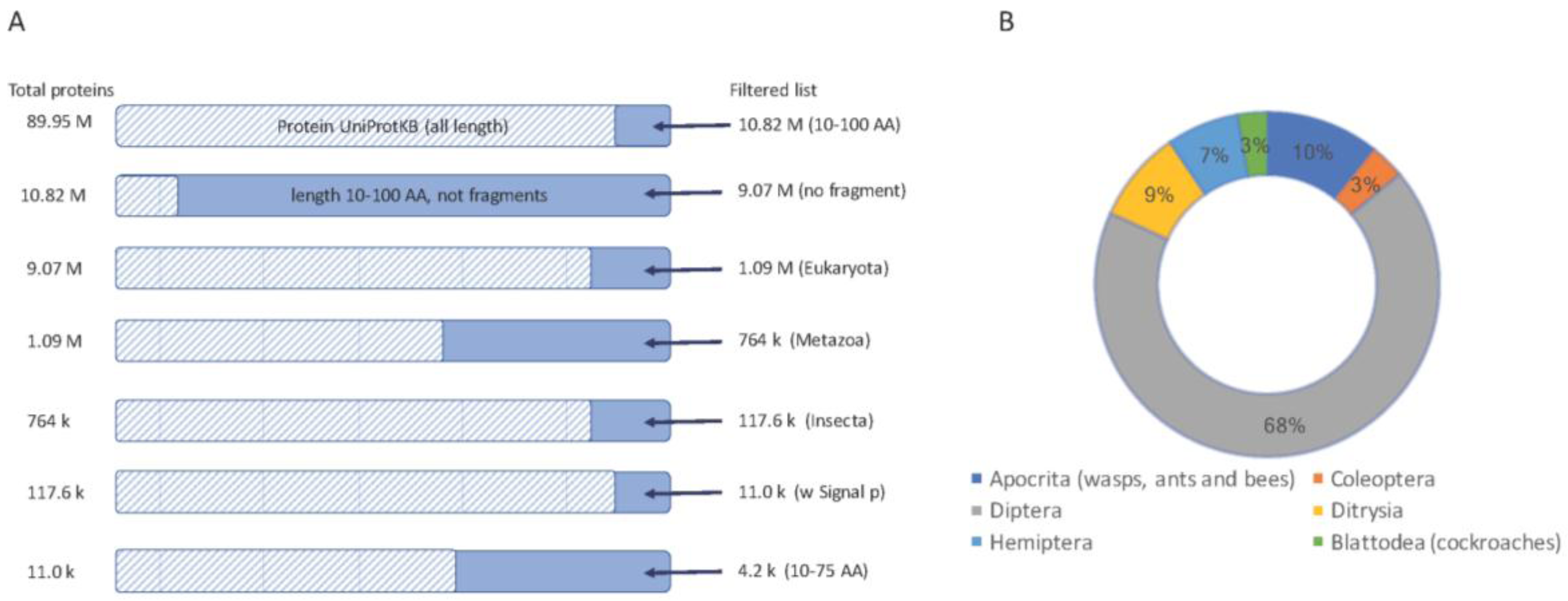
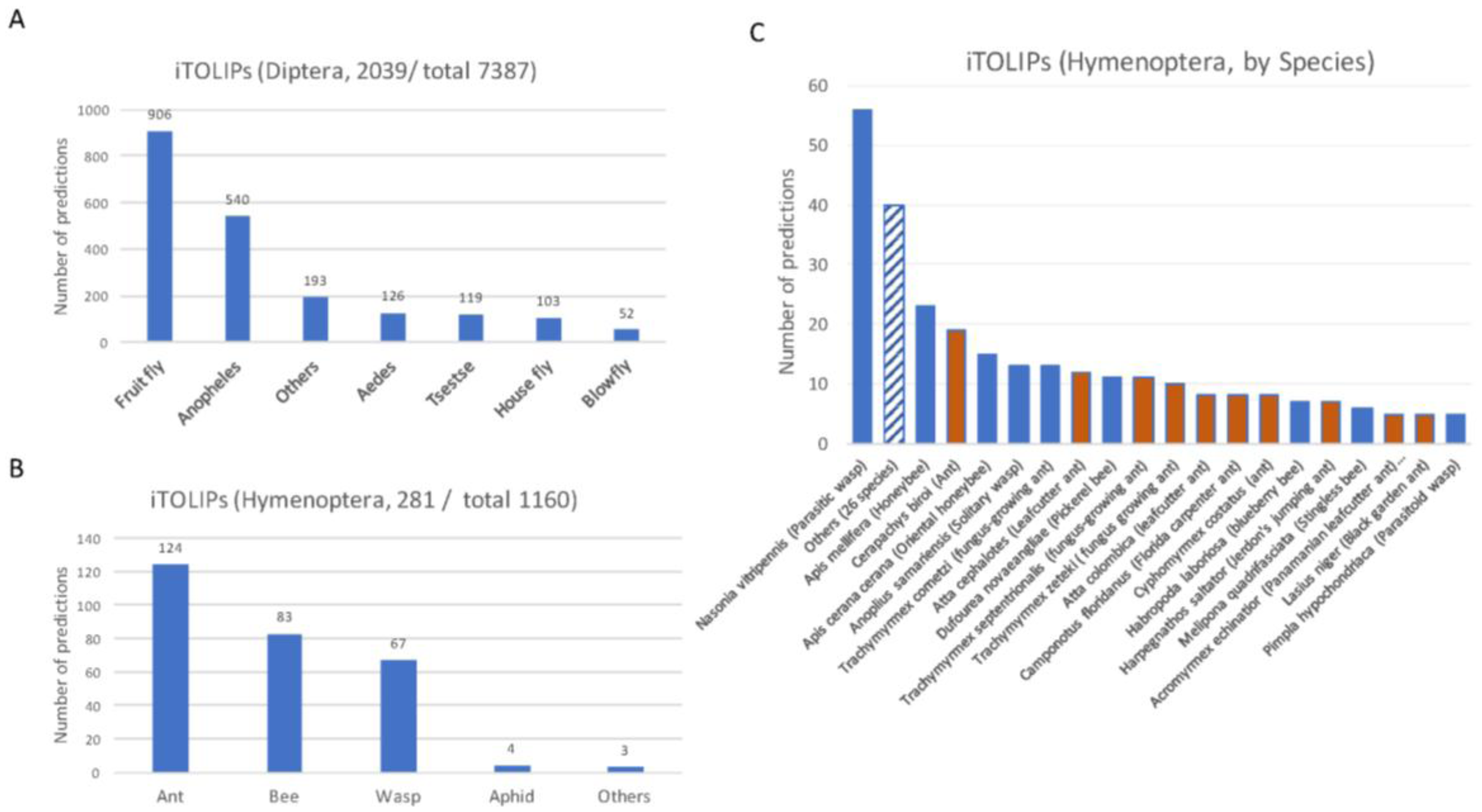
| Insects | Number of Short Proteins | Number of Top Predictions | % Top Predictions from Total | Representative Family |
|---|---|---|---|---|
| Blattoidea | 238 | 124 | 52.1 | Termite |
| Hymenoptera (wasps, ants and bees) | 460 | 35 | 7.6 | Honeybee |
| Ditrysia | 403 | 20 | 5 | Butterfly |
| Polyphaga | 139 | 12 | 8.6 | Beetle |
| Hemiptera | 230 | 16 | 7 | Aphid |
| Pulicidae | 17 | 2 | 11.8 | Flea |
| Acrididae | 9 | 2 | 22.2 | Grasshopper |
| Pseudagrion | 2 | 0 | 0 | Damselfly |
| Psocodea | 24 | 0 | 0 | Lice |
| All insects | 4196 | 379 | 9 |
| UniProtKB | AA (Mature) a | Protein Name | Species | PDB | % Seq. Sim | Description |
|---|---|---|---|---|---|---|
| H9KQJ7 | 74 (54) | ω-conotoxin-like protein 1 | A. mellifera | 2n86.1 | 44.1 | Spiderine-1a |
| A0A084WJA1 | 71 (46) | K-channel toxin α-KTx 18.3 | A. sinensis | 2b68.1 | 24.1 | defensin |
| J7HBU2 | 70 (47) | Salivary toxin-like peptide | N. intermedia | 5t4r.1 | 51.5 | Mu-theraphotoxin-Pn3a |
| J7HIK0 | 70 (47) | Salivary toxin-like peptide | N. intermedia | 5t4r.1 | 51.5 | Mu-theraphotoxin-Pn3a |
| J7HBS6 | 70 (46) | Salivary toxin-like peptide | N. intermedia | 5t4r.1 | 51.5 | Mu-theraphotoxin-Pn3a |
| J7HBT1 | 75 (50) | Salivary toxin-like peptide | N. intermedia | 1d1h.1 | 46.7 | Hanatoxin Type 1 |
| A0A034WXR3 | 60 (36) | Venom toxin-like peptide | A. ervi | 1q3j.1 | 33.3 | ALO3 |
| A0A034WY34 | 61 (37) | Venom toxin-like peptide | A. ervi | 2lqa.1 | 43.8 | Asteropsin A |
| A0A034WWW1 | 51 (37) | Venom toxin-like peptide | A. ervi | 1omn.1 | 48.0 | ω-Conotoxin MVIIC |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Linial, M.; Rappoport, N.; Ofer, D. Overlooked Short Toxin-Like Proteins: A Shortcut to Drug Design. Toxins 2017, 9, 350. https://doi.org/10.3390/toxins9110350
Linial M, Rappoport N, Ofer D. Overlooked Short Toxin-Like Proteins: A Shortcut to Drug Design. Toxins. 2017; 9(11):350. https://doi.org/10.3390/toxins9110350
Chicago/Turabian StyleLinial, Michal, Nadav Rappoport, and Dan Ofer. 2017. "Overlooked Short Toxin-Like Proteins: A Shortcut to Drug Design" Toxins 9, no. 11: 350. https://doi.org/10.3390/toxins9110350
APA StyleLinial, M., Rappoport, N., & Ofer, D. (2017). Overlooked Short Toxin-Like Proteins: A Shortcut to Drug Design. Toxins, 9(11), 350. https://doi.org/10.3390/toxins9110350








