Menadione-Induced Oxidative Stress Re-Shapes the Oxylipin Profile of Aspergillus flavus and Its Lifestyle
Abstract
:1. Introduction
2. Materials and Methods
2.1. Fungal Strain and Culture Conditions
2.2. Quantification of Mycelial Growth, Conidiogenesis and Aflatoxin B1 Production
2.3. Antioxidant Enzymes and Intracellular ROS Assays
2.4. Identification of Differentially-Expressed Genes via qRT-PCR
2.5. RNA-seq Analysis
2.6. Oxylipins Assay
3. Results
3.1. Physiological Responses to Menadione-Related Oxidative Stress
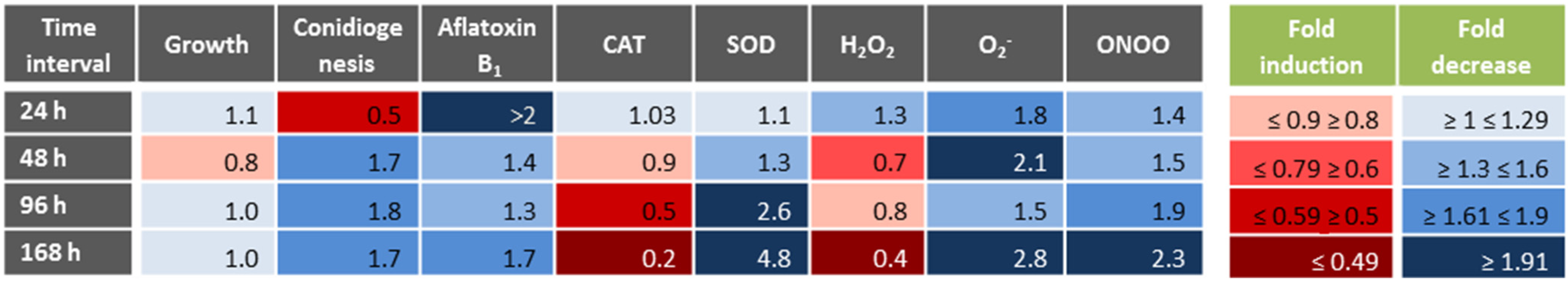
3.2. Correlations
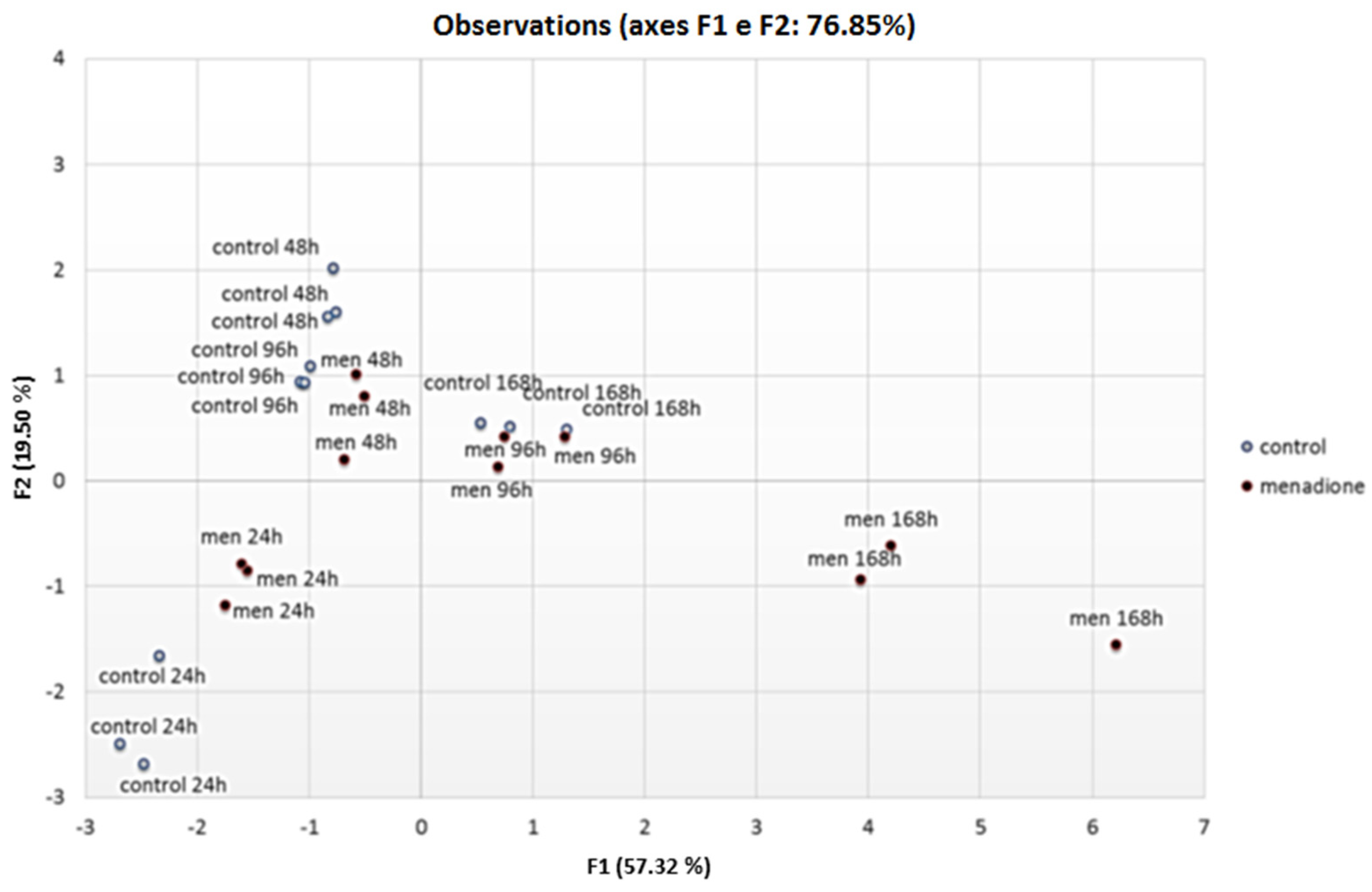
3.3. Transcriptional Analysis of A. flavus Challenged with Menadione
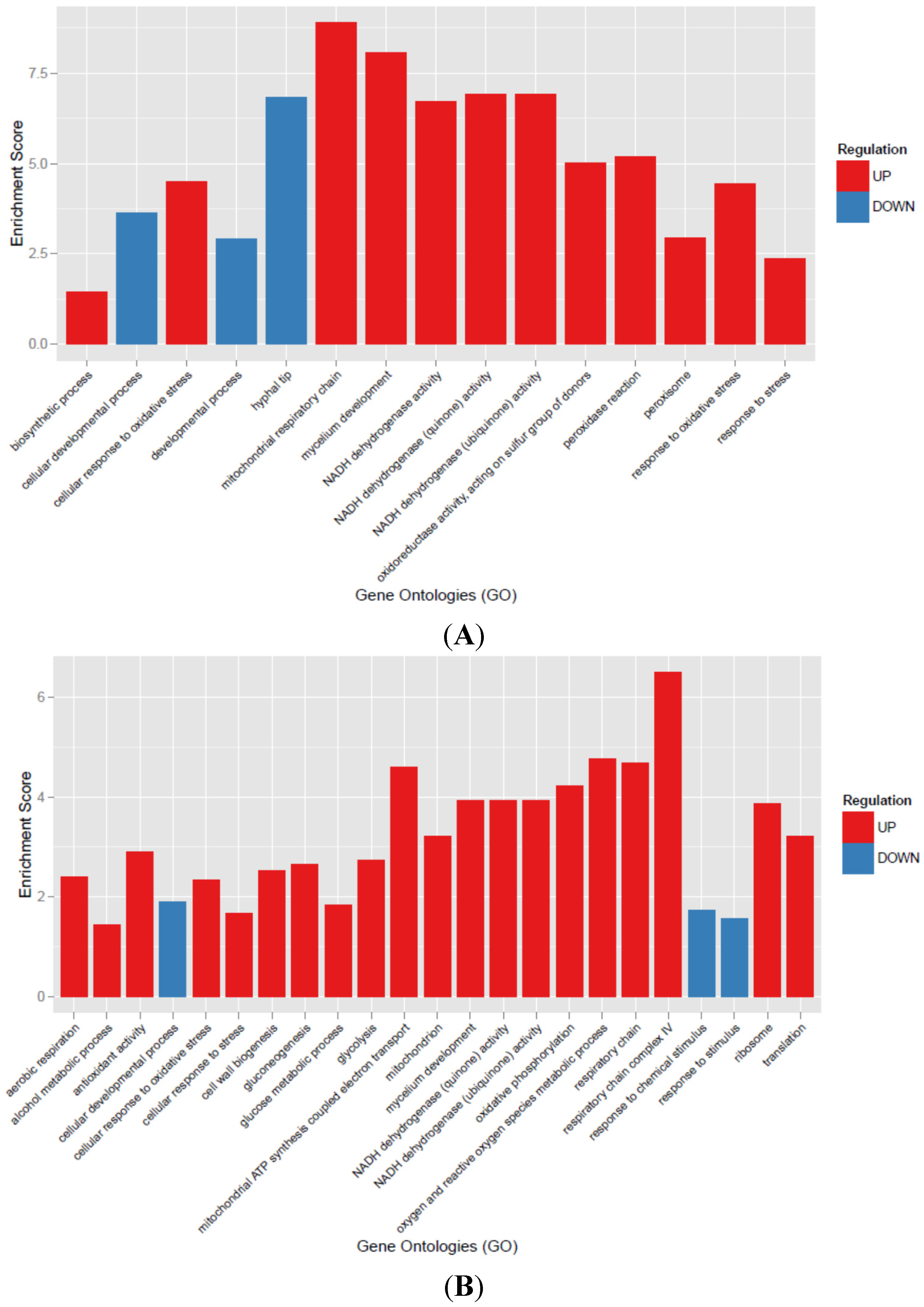
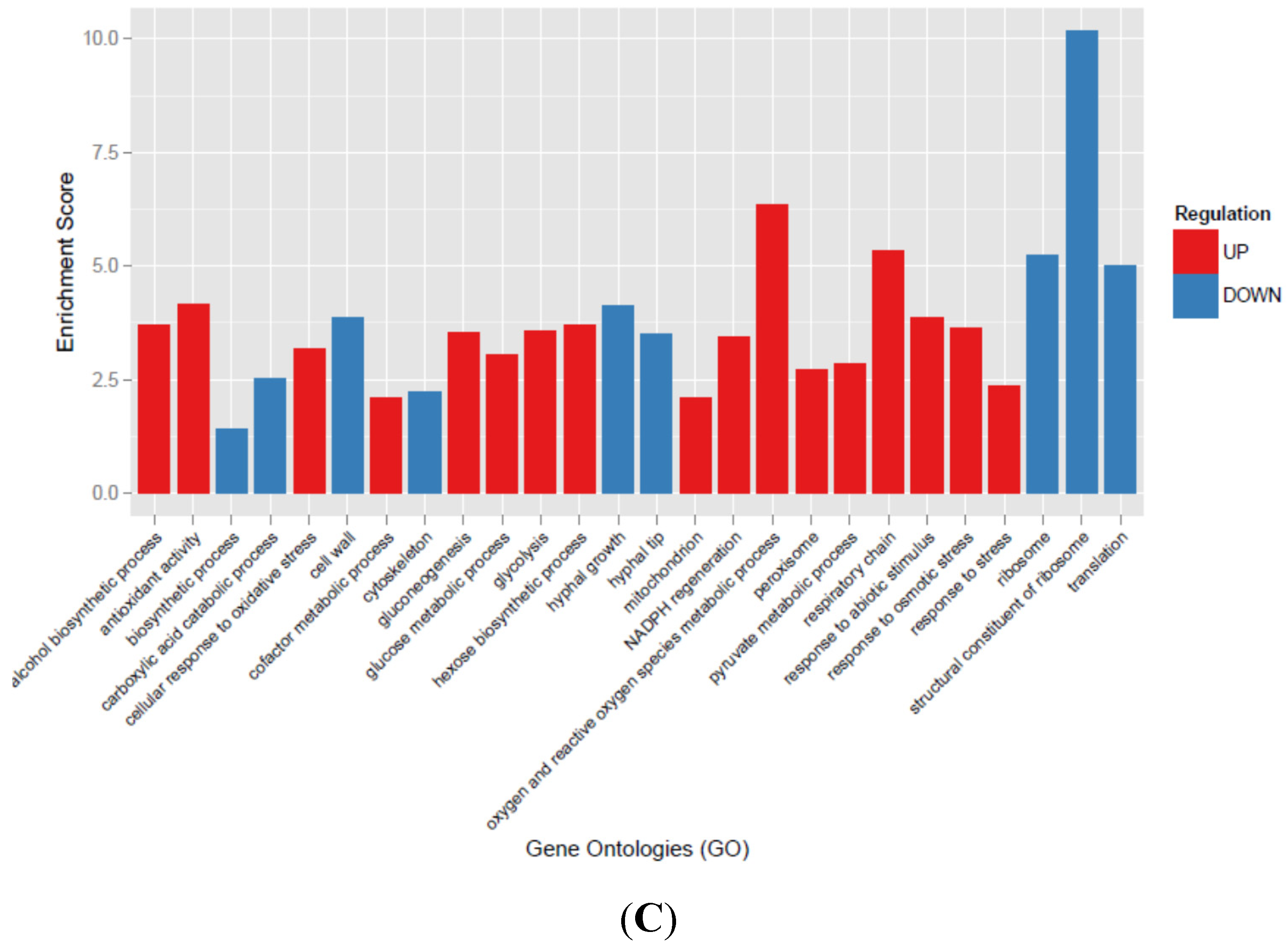
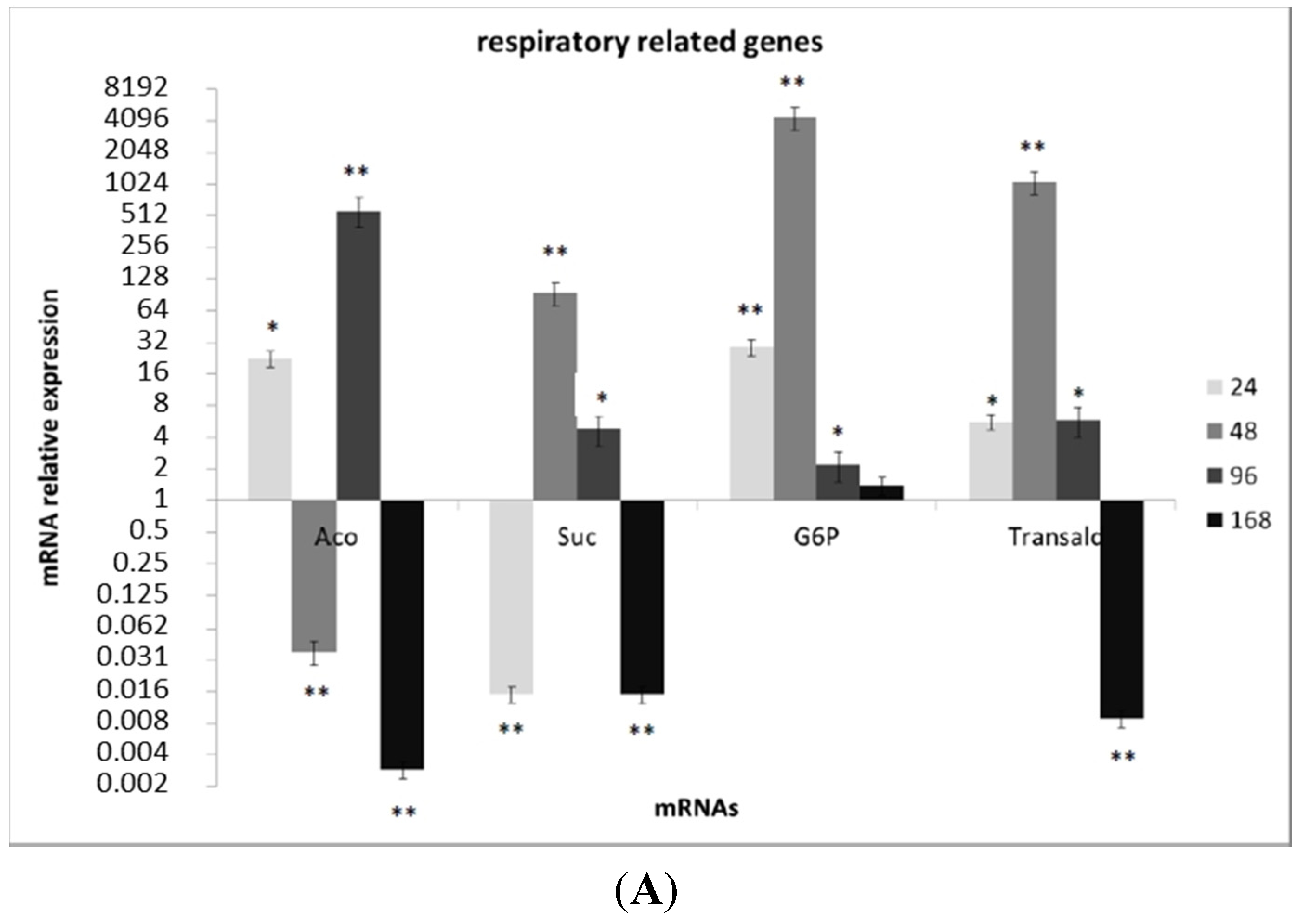
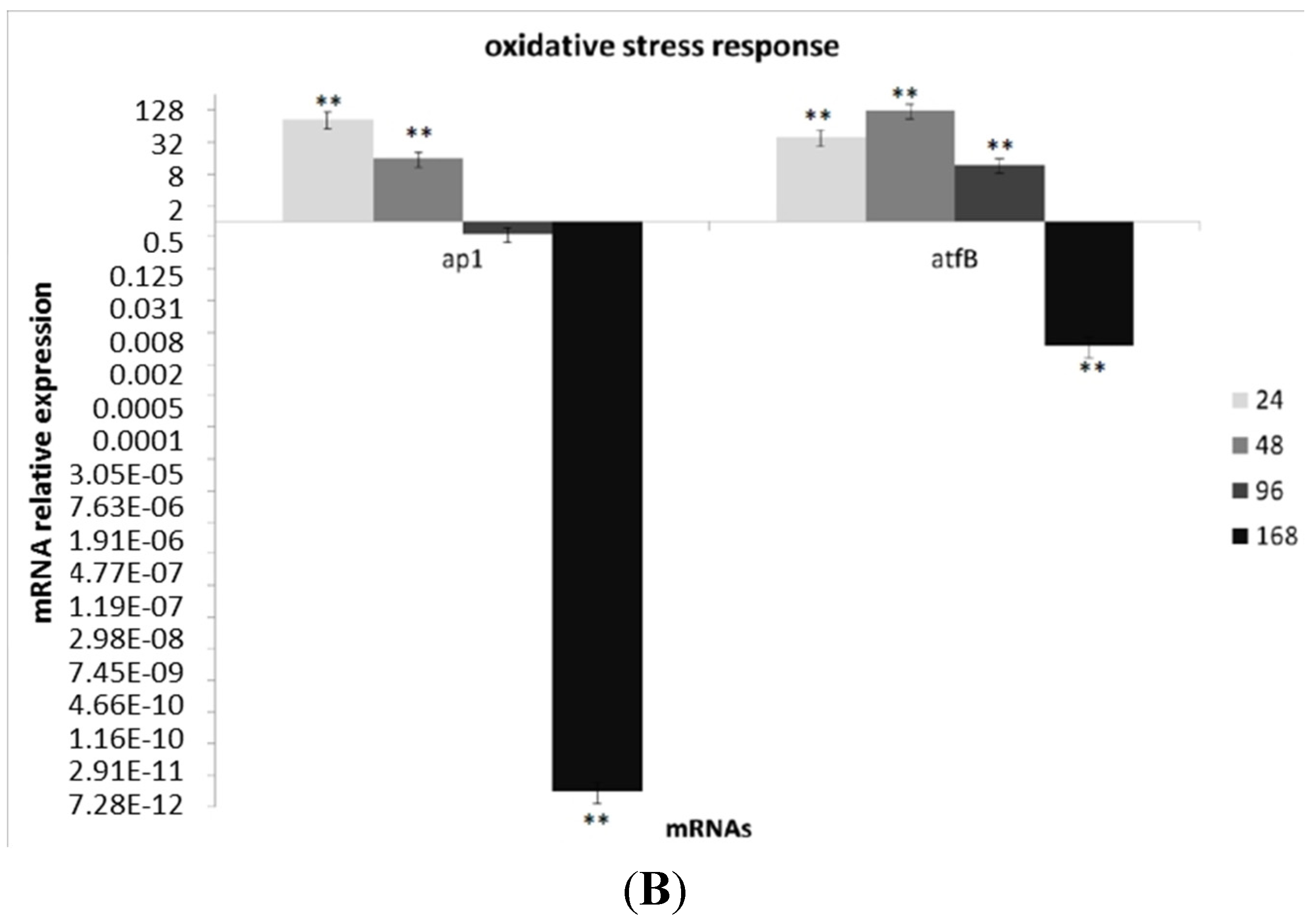
3.4. Oxylipin Analysis of A. flavus Treated with Menadione
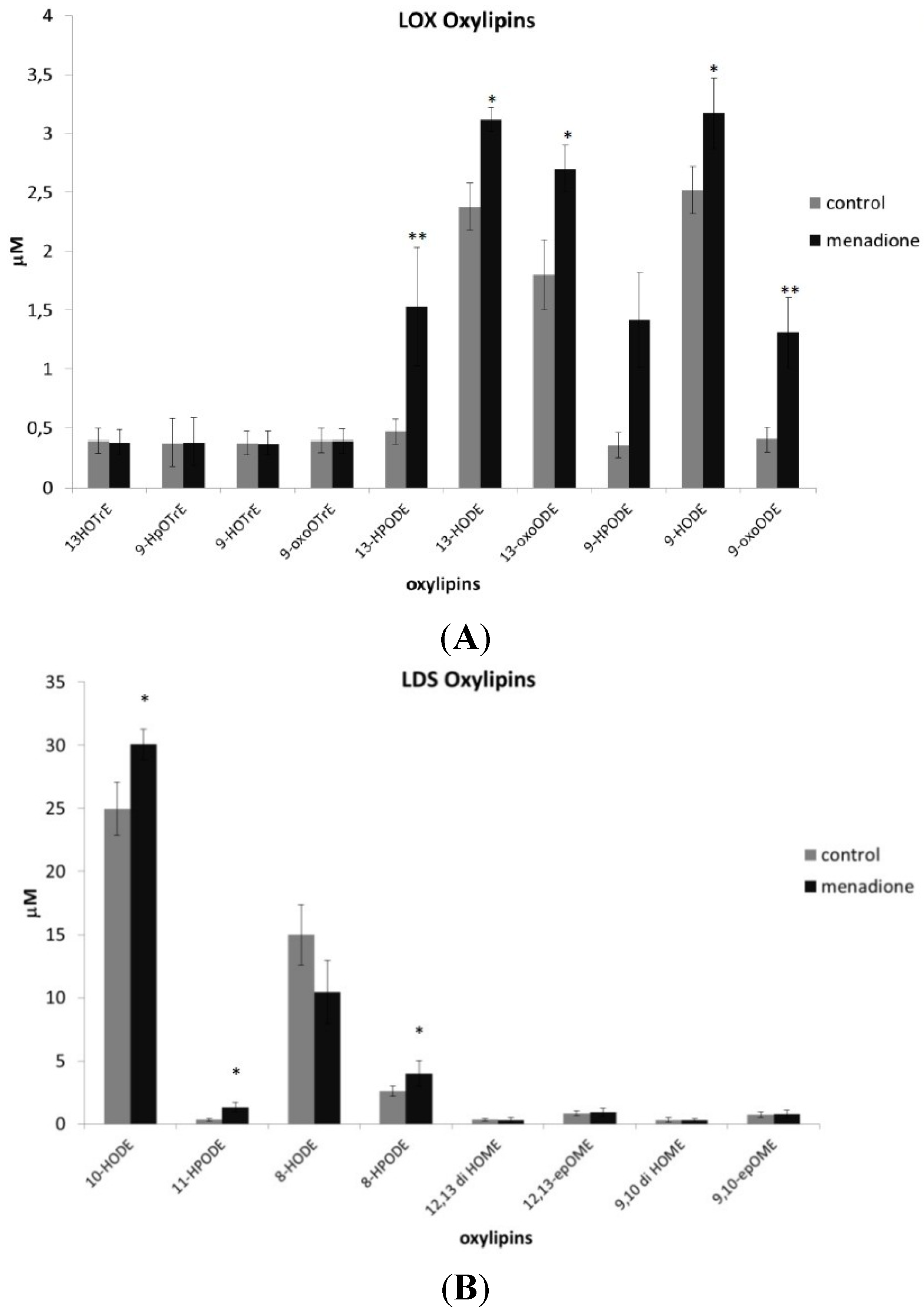
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Jayashree, T.; Subramanyam, C. Oxidative stress as a prerequisite for aflatoxin production by Aspergillus parasiticus. Free Radic. Biol. Med. 2000, 29, 981–985. [Google Scholar] [CrossRef]
- Reverberi, M.; Fabbri, A.A.; Zjalic, S.; Ricelli, A.; Punelli, F.; Fanelli, C. Antioxidant enzymes stimulation in Aspergillus parasiticus by Lentinula edodes inhibits aflatoxin production. Appl. Microbiol. Biotechnol. 2005, 69, 207–215. [Google Scholar] [CrossRef] [PubMed]
- Reverberi, M.; Zjalic, S.; Ricelli, A.; Punelli, F.; Camera, E.; Fabbri, C.; Picardo, M.; Fanelli, C.; Fabbri, A.A. Modulation of antioxidant defence in Aspergillus parasiticus is involved in aflatoxin biosynthesis: A role for the ApyapA gene. Eukaryot. Cell 2008, 7, 988–1000. [Google Scholar] [CrossRef] [PubMed]
- Schrader, M.; Fahimi, H.D. Peroxisomes and oxidative stress. Biochim. Biophys. Acta Mol. Cell Res. 2006, 1763, 1755–1766. [Google Scholar] [CrossRef] [PubMed]
- Halliwell, B. Reactive species and antioxidants. Redox biology is a fundamental theme of aerobic life. Plant Physiol. 2006, 141, 312–322. [Google Scholar] [CrossRef] [PubMed]
- Mittler, R.; Vanderauwera, S.; Suzuki, N.; Miller, G.; Tognetti, V.B.; Vandepoele, K.; Gollery, M.; Shulaev, V.; van Breusegem, F. ROS signalling: The new wave? Trends Plant Sci. 2011, 16, 300–309. [Google Scholar] [CrossRef] [PubMed]
- Apel, K.; Hirt, H. Reactive oxygen species: Metabolism, oxidative stress, and signal transduction. Annu. Rev. Plant Biol. 2004, 55, 373–399. [Google Scholar] [CrossRef] [PubMed]
- Dowling, D.; Simmons, L.W. Reactive oxygen species as universal constraints in life-history evolution. Proc. R. Soc. B 2009, 276, 1737–1745. [Google Scholar] [CrossRef] [PubMed]
- Apostol, I.; Heinstein, P.F.; Low, P.S. Rapid stimulation of an oxidative burst during elicitation of cultured plant cells. Role in defense and signal transduction. Plant Physiol. 1989, 90, 106–116. [Google Scholar] [CrossRef]
- Thorpe, G.W.; Fong, C.S.; Alic, N.; Higgins, V.J.; Dawes, I.W. Cells have distinct mechanisms to maintain protection against different reactive oxygen species: Oxidative-stress-response genes. Proc. Natl. Acad. Sci. USA 2004, 101, 6564–6569. [Google Scholar] [CrossRef] [PubMed]
- Monks, T.J.; Hanzlik, R.P.; Cohen, G.M.; Ross, D.; Graham, D.G. Quinone chemistry and toxicity. Toxicol. Appl. Pharmacol. 1992, 112, 2–16. [Google Scholar] [CrossRef]
- Criddle, D.N.; Gillies, S.; Baumgartner-Wilson, H.K.; Jaffar, M.; Chinje, E.C.; Passmore, S.; Chvanov, M.; Barrow, S.; Gerasimenko, O.V.; Tepikin, A.V.; et al. Menadione-induced reactive oxygen species generation via redox cycling promotes apoptosis of murine pancreatic acinar cells. J. Biol. Chem. 2006, 281, 40485–40492. [Google Scholar] [CrossRef] [PubMed]
- Calvo, A.M.; Wilson, R.A.; Bok, J.W.; Keller, N.P. Relationship between secondary metabolism and fungal development. Microbiol. Mol. Biol. Rev. 2002, 66, 447–459. [Google Scholar] [CrossRef] [PubMed]
- Bonekamp, N.A.; Völkl, A.; Fahimi, H.D.; Schrader, M. Reactive oxygen species and peroxisomes: Struggling for balance. BioFactors 2009, 35, 346–355. [Google Scholar] [CrossRef] [PubMed]
- Reverberi, M.; Zjalic, S.; Punelli, F.; Ricelli, A.; Fabbri, A.A.; Fanelli, C. Apyap1 affects aflatoxin biosynthesis during Aspergillus parasiticus growth in maize seeds. Food Addit. Contam. 2007, 24, 1070–1075. [Google Scholar] [CrossRef] [PubMed]
- Cao, Q.H.; Zhou, Q.X.; Cai, R.X.; Liu, Z.H. Fluorimetric determination of peroxynitrite based on an enzymatic reaction. Anal. Sci. 2005, 21, 445–447. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina Sequence Data. Bioinformatics 2014. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S.L. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013, 14. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Smyth, G.K.; Shi, W. Feature Counts: An efficient general-purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.D.; Davis, J.; McCarthy, D.; Gordon, K.S. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed]
- Scala, V.; Camera, E.; Ludovici, M.; Dall’Asta, C.; Cirlini, M.; Giorni, P.; Battilani, P.; Bello, C.; Fabbri, A.A.; Fanelli, C.; et al. Fusarium verticillioides and maize interaction in vitro: Relation between oxylipin cross-talk and fumonisin synthesis. World Mycotoxin J. 2013, 6, 343–351. [Google Scholar] [CrossRef]
- Ludovici, M.; Ialongo, C.; Reverberi, M.; Beccaccioli, M.; Scarpari, M.; Scala, V. Quantitative profiling of oxylipins through comprehensive LC-MS/MS analysis of Fusarium verticillioides and maize kernels. Food Addit. Contam. A 2014, 31, 2026–2033. [Google Scholar] [CrossRef] [PubMed]
- Strassburg, K.; Huijbrechts, A.M.; Kortekaas, K.A.; Lindeman, J.H.; Pedersen, T.L.; Dane, A. Quantitative profiling of oxylipins through comprehensive LC-MS/MS analysis: Application in cardiac surgery. Anal. Bioanal. Chem. 2012, 404, 1413–1426. [Google Scholar] [CrossRef] [PubMed]
- Narasaiah, K.W.; Sashidhar, R.B.; Subramanyam, C. Biochemical analysis of oxidative stress in the production of aflatoxin and its precursor intermediates. Mycopathologia 2006, 162, 179–189. [Google Scholar] [CrossRef] [PubMed]
- Reverberi, M.; Punelli, M.; Scala, V.; Scarpari, M.; Uva, P.; Mentzen, W.I.; Dolezal, A.L.; Woloshuk, C.; Pinzari, F.; Fabbri, A.A.; et al. Genotypic and phenotypic versatility of Aspergillus flavus during maize exploitation. PLoS ONE 2013, 8, e68735. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.Y.; Roze, L.V.; Linz, J.E. Oxidative stress-related transcription factors in the regulation of secondary metabolism. Toxins 2013, 5, 683–702. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, G.F.; Baltazar Lde, M.; Santos, J.R.; Monteiro, A.S.; Fraga, L.A.; Resende-Stoianoff, M.A.; Santos, D.A. The role of oxidative and nitrosative bursts caused by azoles and amphotericin B against the fungal pathogen Cryptococcus gattii. J. Antimicrob. Chemother. 2013, 68, 1801–1811. [Google Scholar] [CrossRef] [PubMed]
- Roze, L.V.; Chanda, A.; Wee, J.; Awad, D.; Linz, J.E. Stress-related transcription factor AtfB integrates secondary metabolism with oxidative stress response in Aspergilli. J. Biol. Chem. 2011, 286, 35137–35148. [Google Scholar] [CrossRef] [PubMed]
- Szabo, C.; Ischiropoulos, H.; Radi, R. Peroxynitrite: Biochemistry, pathophysiology and development of therapeutics. Nat. Rev. Drug Discov. 2007, 6, 662–680. [Google Scholar] [CrossRef] [PubMed]
- Virag, L.; Szabo, E.; Gergely, P. Peroxynitrite-induced cytotoxicity: Mechanism and opportunities for intervention. Toxicol. Lett. 2003, 11, 113–124. [Google Scholar] [CrossRef]
- Rubbo, H.; Trostchansky, A.; O’Donnell, V.B. Peroxynitrite-mediated lipid oxidation and nitration: Mechanisms and consequences. Arch. Biochem. Biophys. 2009, 484, 167–172. [Google Scholar] [CrossRef] [PubMed]
- Reverberi, M.; Punelli, M.; Smith, C.A.; Zjalic, S.; Scarpari, M.; Scala, V.; Fanelli, C. How peroxisomes affect aflatoxin biosynthesis in Aspergillus flavus. PLoS ONE 2012, 7, e48097. [Google Scholar] [CrossRef] [PubMed]
- Tsitsigiannis, D.I.; Keller, N.P. Oxylipins as developmental and host-fungal communication signals. Trends Microbiol. 2007, 15, 109–118. [Google Scholar] [CrossRef] [PubMed]
- Reverberi, M.; Fabbri, A.A.; Fanelli, C. Oxidative stress and oxylipins in plant-fungus interaction. In Biocommunication of Fungi; Guenther, W., Ed.; Springer: London, UK, 2012. [Google Scholar]
- Reverberi, M.; Punelli, F.; Scarpari, M.; Camera, E.; Zjalic, S.; Ricelli, A.; Fanelli, C.; Fabbri, A.A. Lipoperoxidation affects ochratoxin A biosynthesis in Aspergillus ochraceus and its interaction with wheat seeds. Appl. Microbiol. Biotechnol. 2010, 85, 1935–1946. [Google Scholar] [CrossRef] [PubMed]
- Scala, V.; Giorni, P.; Cirlini, M.; Ludovici, M.; Visentin, I.; Cardinale, F.; Fabbri, A.A.; Fanelli, C.; Reverberi, M.; Battilani, P.; et al. LDS1-produced oxylipins are negative regulators of growth, conidiation and fumonisin synthesis in the fungal maize pathogen Fusarium verticillioides. Front. Microbiol. 2014, 5, 669. [Google Scholar] [CrossRef] [PubMed]
- Tsitsigiannis, D.I.; Kowieski, R.; Zarnowski, R.; Keller, N.P. Three putative oxylipin biosynthetic genes integrate sexual and asexual development in Aspergillus nidulans. Microbiology 2005, 151, 1809–1821. [Google Scholar] [CrossRef] [PubMed]
- Brown, S.; Zarnowski, R.; Sharpee, W.C.; Keller, N.P. Morphological transitions governed by density dependence and lipoxygenase activity in Aspergillus flavus. Appl. Environ. Microbiol. 2008, 74, 5674–5685. [Google Scholar] [CrossRef] [PubMed]
- Scarpari, M.; Punelli, M.; Scala, V.; Zaccaria, M.; Nobili, C.; Ludovici, M.; Camera, E.; Fabbri, A.A.; Reverberi, M.; Fanelli, C. Lipids in Aspergillus flavus-maize interaction. Front. Microbiol. 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Scala, V.; Beccaccioli, M.; Dall’Asta, C.; Giorni, P.; Fanelli, C. Analysis of the expression of genes related to oxylipin biosynthesis in Fusarium verticillioides and maize kernels during their interaction. J. Plant Pathol. 2015, 97, 193–197. [Google Scholar]
- Affeldt, K.J.; Brodhagen, M.; Keller, N.P. Aspergillus oxylipin signalling and quorum sensing pathways depend on G Protein-Coupled Receptors. Toxins 2012, 4, 695–717. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zaccaria, M.; Ludovici, M.; Sanzani, S.M.; Ippolito, A.; Cigliano, R.A.; Sanseverino, W.; Scarpari, M.; Scala, V.; Fanelli, C.; Reverberi, M. Menadione-Induced Oxidative Stress Re-Shapes the Oxylipin Profile of Aspergillus flavus and Its Lifestyle. Toxins 2015, 7, 4315-4329. https://doi.org/10.3390/toxins7104315
Zaccaria M, Ludovici M, Sanzani SM, Ippolito A, Cigliano RA, Sanseverino W, Scarpari M, Scala V, Fanelli C, Reverberi M. Menadione-Induced Oxidative Stress Re-Shapes the Oxylipin Profile of Aspergillus flavus and Its Lifestyle. Toxins. 2015; 7(10):4315-4329. https://doi.org/10.3390/toxins7104315
Chicago/Turabian StyleZaccaria, Marco, Matteo Ludovici, Simona Marianna Sanzani, Antonio Ippolito, Riccardo Aiese Cigliano, Walter Sanseverino, Marzia Scarpari, Valeria Scala, Corrado Fanelli, and Massimo Reverberi. 2015. "Menadione-Induced Oxidative Stress Re-Shapes the Oxylipin Profile of Aspergillus flavus and Its Lifestyle" Toxins 7, no. 10: 4315-4329. https://doi.org/10.3390/toxins7104315
APA StyleZaccaria, M., Ludovici, M., Sanzani, S. M., Ippolito, A., Cigliano, R. A., Sanseverino, W., Scarpari, M., Scala, V., Fanelli, C., & Reverberi, M. (2015). Menadione-Induced Oxidative Stress Re-Shapes the Oxylipin Profile of Aspergillus flavus and Its Lifestyle. Toxins, 7(10), 4315-4329. https://doi.org/10.3390/toxins7104315







