Evaluating the Potential for Cross-Interactions of Antitoxins in Type II TA Systems
Abstract
1. Introduction
2. Paradigms for Type II TA Systems
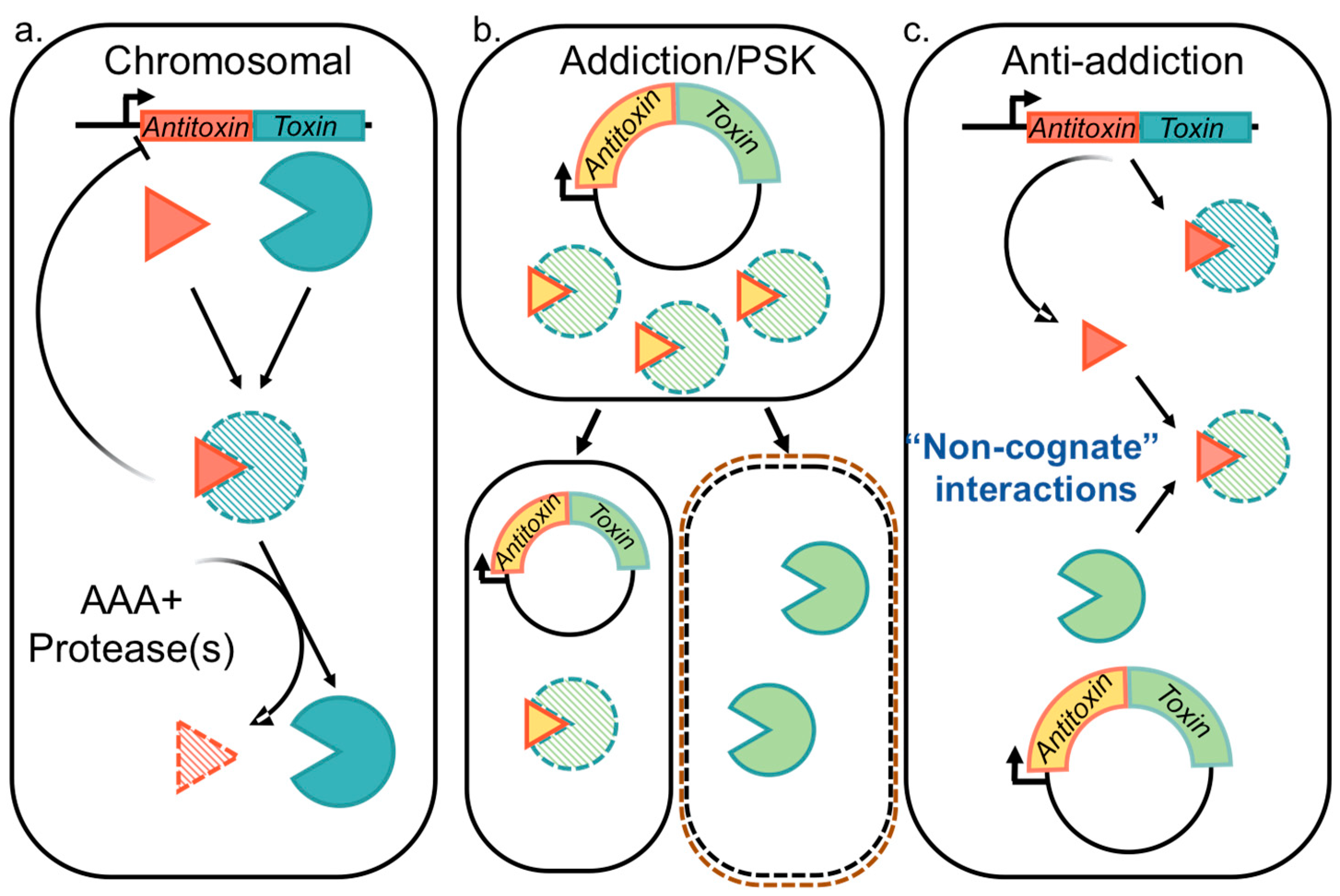
2.1. Functions Attributed to Chromosomal Type II TA Systems
2.2. Plasmid Selection and/or Addiction via Type II TA Systems
2.3. Type II TA Systems Mediating Anti-Addition through Antitoxin Cross-Interactions
3. Conservation of Type II TA System Folds and Cognate Antitoxin Interactions
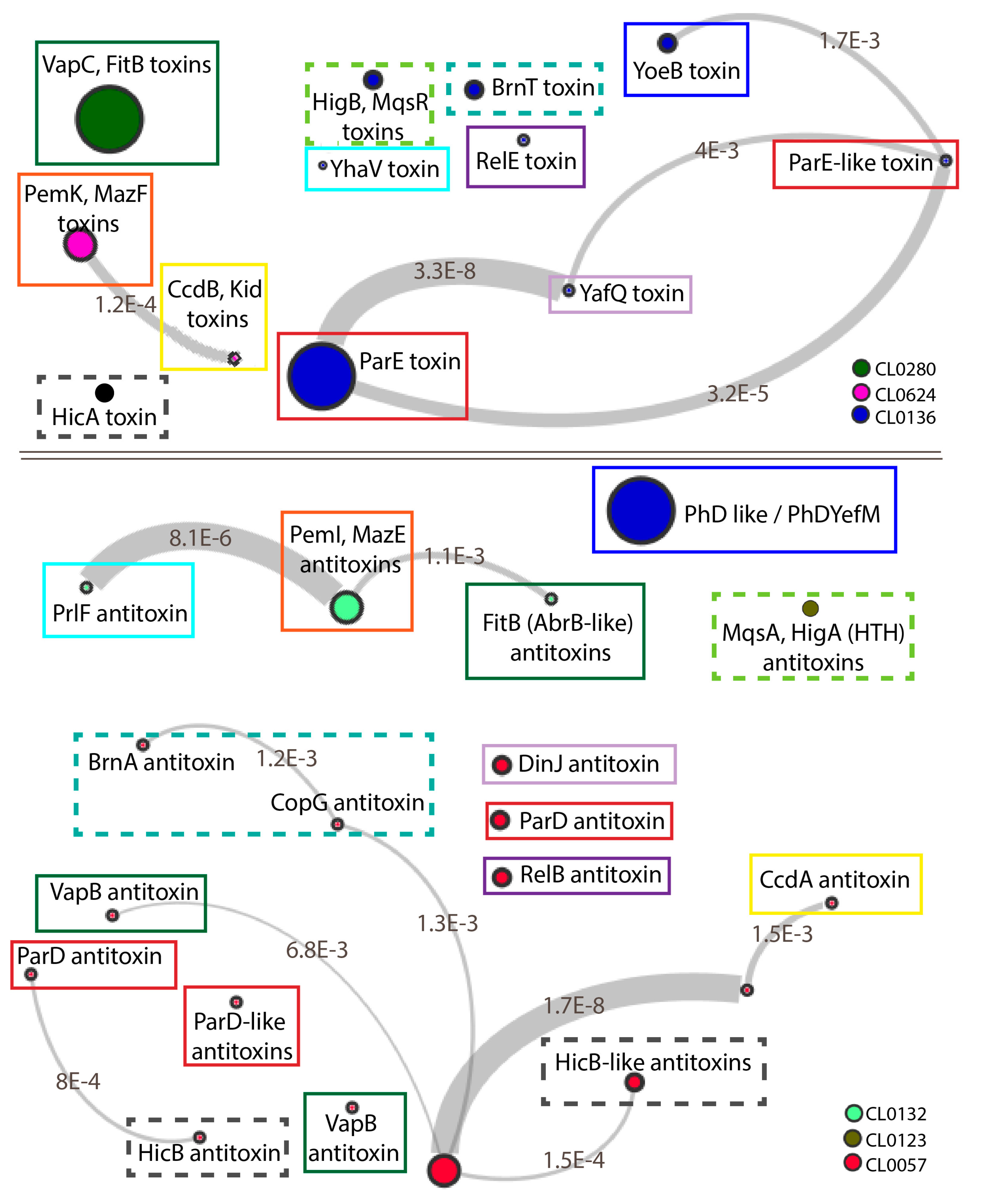
3.1. Toxin Families Share Conserved Folds but Interact with Different Families of Antitoxins
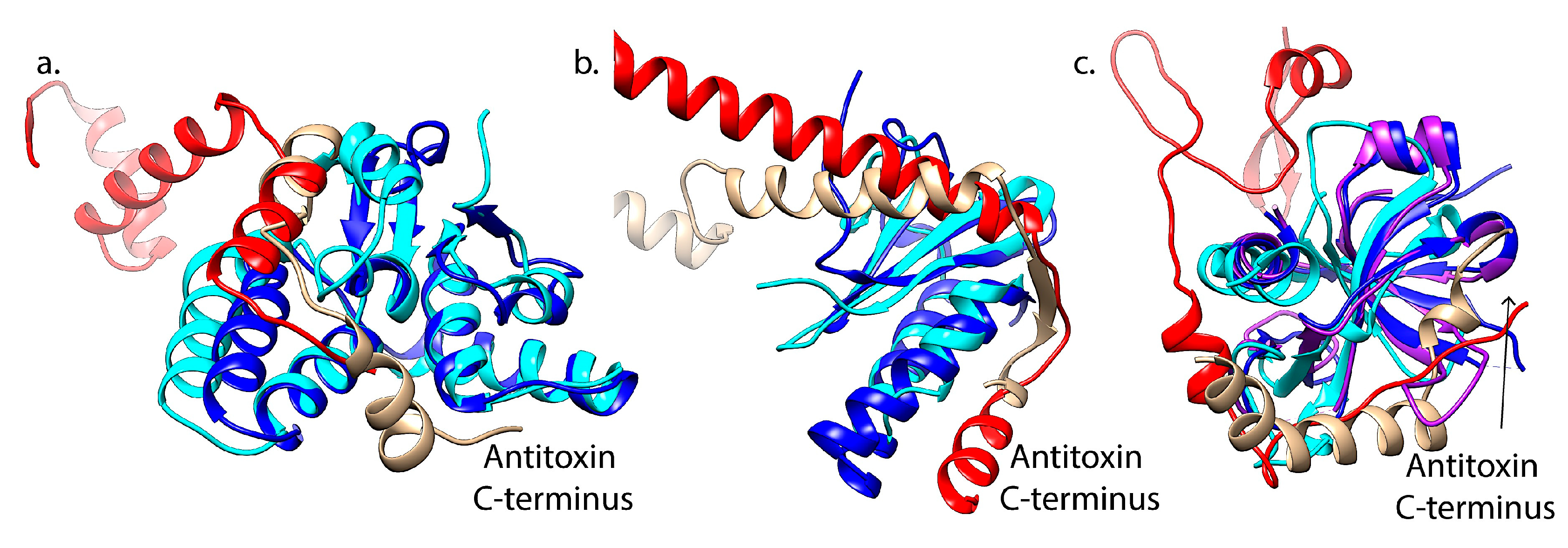
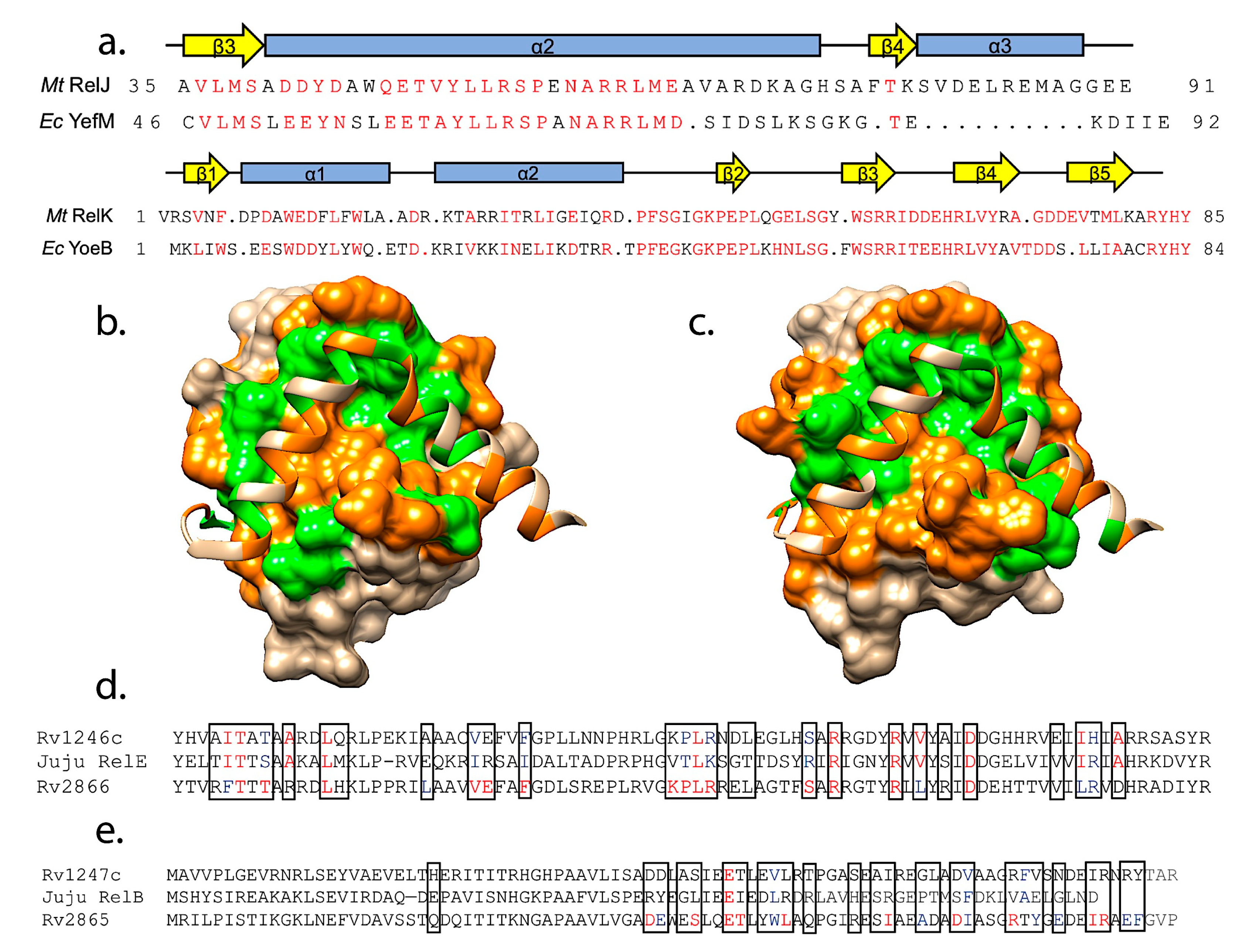
3.2. Interactions with Cognate Antitoxins
4. Feasibility for Cross-Interactions of Type II TA Systems
4.1. Examples of Antitoxin Cross-Reactivity
4.2. Orphan Antitoxins
4.3. Predictions of Antitoxin Cross-Reactivity
5. Discussion
Funding
Acknowledgments
Conflicts of Interest
References
- Anantharaman, V.; Aravind, L. New connections in the prokaryotic toxin-antitoxin network: Relationship with the eukaryotic nonsense-mediated RNA decay system. Genome Biol. 2003, 4, R81. [Google Scholar] [CrossRef] [PubMed]
- Pandey, P.D.; Gerde, K.S. Toxin-antitoxin loci are highly abundant in free-living but lost from host-associated prokaryotes. Nucleic Acids Res. 2005, 33, 966–976. [Google Scholar] [CrossRef]
- Melderen, V.L.; Jurenas, D.; Garcia-Pino, A. Messing up translation from the start: How AtaT inhibits translation initiation in E. coli. RNA Biol. 2018, 15, 303–307. [Google Scholar] [CrossRef]
- Lima-Mendez, G.; Alvarenga, D.O.; Ross, K.; Hallet, B.; Van Melderen, L.; Varani, A.M.; Chandler, M. Toxin-Antitoxin Gene Pairs Found in Tn3 Family Transposons Appear To Be an Integral Part of the Transposition Module. mBio 2020, 11, e00452-20. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.A.; Kamruzzaman, M.; Iredell, J.R. Specialised functions of two common plasmid mediated toxin-antitoxin systems, ccdAB and pemIK, in Enterobacteriaceae. bioRxiv 2020. [Google Scholar] [CrossRef]
- Huguet, K.T.; Gonnet, M.; Doublet, B.; Cloeckaert, A. A toxin antitoxin system promotes the maintenance of the IncA/C-mobilizable Salmonella Genomic Island 1. Sci. Rep. 2016, 6, 32285. [Google Scholar] [CrossRef] [PubMed]
- Makarova, S.K.; Wolf, Y.I.; Koonin, E.V. Comprehensive comparative-genomic analysis of type 2 toxin-antitoxin systems and related mobile stress response systems in prokaryotes. Biol. Direct 2009, 4, 19–56. [Google Scholar] [CrossRef] [PubMed]
- Fernández-García, L.; Blasco, L.; Lopez, M.; Bou, G.; García-Contreras, R.; Wood, T.; Tomas, M. Toxin-antitoxin systems in clinical pathogens. Toxins (Basel) 2016, 8, 227–249. [Google Scholar] [CrossRef]
- Kedzierska, B.; Hayes, F. Emerging Roles of Toxin-Antitoxin Modules in Bacterial Pathogenesis. Molecules 2016, 21, 790. [Google Scholar] [CrossRef]
- Ramisetty, M.B.C.; Santhosh, R.S. Endoribonuclease type II toxin-antitoxin systems: Functional or selfish? Microbiology 2017, 163, 931–939. [Google Scholar] [CrossRef]
- Yang, E.Q.; Walsh, T.R. Toxin-antitoxin systems and their role in disseminating and maintaining antimicrobial resistance. FEMS Microbiol. Rev. 2017, 41, 343–353. [Google Scholar] [CrossRef] [PubMed]
- Fraikin, N.; Goormaghtigh, F.; van Melderen, L. Type II Toxin-Antitoxin Systems: Evolution and Revolutions. J. Bacteriol. 2020, 202, e00763-19. [Google Scholar] [CrossRef] [PubMed]
- Jurėnas, D.; van Melderen, L. The Variety in the Common Theme of Translation Inhibition by Type II Toxin–Antitoxin Systems. Front. Genet. 2020, 11, 262. [Google Scholar] [CrossRef]
- Hall, M.A.; Gollan, B.; Helaine, S. Toxin-antitoxin systems: Reversible toxicity. Curr. Opin. Microbiol. 2017, 36, 102–110. [Google Scholar] [CrossRef]
- Page, R.; Peti, W. Toxin-antitoxin systems in bacterial growth arrest and persistence. Nat. Chem. Biol. 2016, 12, 208–214. [Google Scholar] [CrossRef]
- Ronneau, S.; Helaine, S. Clarifying the Link between Toxin-Antitoxin Modules and Bacterial Persistence. J. Mol. Biol 2019, 431, 3462–3471. [Google Scholar] [CrossRef]
- Unterholzner, J.S.; Poppenberger, B.; Rozhon, W. Toxin-antitoxin systems: Biology, identification, and application. Mob. Genet. Elem. 2013, 3, e26219. [Google Scholar] [CrossRef]
- Schuster, F.C.; Bertram, R. Toxin-antitoxin systems are ubiquitous and versatile modulators of prokaryotic cell fate. FEMS Microbiol. Lett. 2013, 340, 73–85. [Google Scholar] [CrossRef]
- Beyer, M.H.; Iwai, H. Off-Pathway-Sensitive Protein-Splicing Screening Based on a Toxin/Antitoxin System. ChemBioChem 2019, 20, 1933–1938. [Google Scholar] [CrossRef] [PubMed]
- Klimina, K.M.; Kasianov, A.S.; Poluektova, E.U.; Emelyanov, K.V.; Voroshilova, V.N.; Zakharevich, N.V.; Kudryavtseva, A.V.; Makeev, V.J.; Danilenko, V.N. Employing toxin-antitoxin genome markers for identification of Bifidobacterium and Lactobacillus strains in human metagenomes. PeerJ 2019, 7, e6554. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.M.; Kim, D.H.; Jin, C.; Lee, B.J. A Systematic Overview of Type II and III Toxin-Antitoxin Systems with a Focus on Druggability. Toxins (Basel) 2018, 10, 515. [Google Scholar] [CrossRef] [PubMed]
- Równicki, M.; Pieńko, T.; Czarnecki, J.; Kolanowska, M.; Bartosik, D.; Trylska, J. Artificial Activation of Escherichia coli mazEF and hipBA Toxin-Antitoxin Systems by Antisense Peptide Nucleic Acids as an Antibacterial Strategy. Front. Microbiol. 2018, 9, 2870. [Google Scholar] [CrossRef]
- Lioy, V.S.; Rey, O.; Balsa, D.; Pellicer, T.; Alonso, J.C. A toxin-antitoxin module as a target for antimicrobial development. Plasmid 2010, 63, 31–39. [Google Scholar] [CrossRef]
- Fedorec, A.J.; Ozdemir, T.; Doshi, A.; Ho, Y.K.; Rosa, L.; Rutter, J.; Velazquez, O.; Pinheiro, V.B.; Danino, T.; Barnes, C.P. Two New Plasmid Post-segregational Killing Mechanisms for the Implementation of Synthetic Gene Networks in Escherichia coli. iScience 2019, 14, 323–334. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.K.; Lee, B.J. Structure, Biology, and Therapeutic Application of Toxin-Antitoxin Systems in Pathogenic Bacteria. Toxins (Basel) 2016, 8, 305. [Google Scholar] [CrossRef] [PubMed]
- Magnuson, R.D. Hypothetical functions of toxin-antitoxin systems. J. Bacteriol. 2007, 189, 6089–6092. [Google Scholar] [CrossRef]
- Harms, A.; Brodersen, D.E.; Mitarai, N.; Gerdes, K. Toxins, Targets, and Triggers: An Overview of Toxin-Antitoxin Biology. Mol. Cell 2018, 70, 768–784. [Google Scholar] [CrossRef]
- Goeders, N.; van Melderen, L. Toxin-antitoxin systems as multilevel interaction systems. Toxins (Basel) 2014, 6, 304–324. [Google Scholar] [CrossRef]
- Fozo, M.E.; Hemm, M.R.; Storz, G. Small Toxic Proteins and the Antisense RNAs That Repress Them. Microbiol. Mol. Biol. Rev. 2008, 72, 579–589. [Google Scholar] [CrossRef]
- Blower, R.T.; Salmond, G.P.; Luisi, B.F. Balancing at survival’s edge: The structure and adaptive benefits of prokaryotic toxin-antitoxin partners. Curr. Opin. Struct. Biol. 2011, 21, 109–118. [Google Scholar] [CrossRef]
- Fineran, P.C.; Blower, T.R.; Foulds, I.J.; Humphreys, D.P.; Lilley, K.S.; Salmond, G.P. The phage abortive infection system, ToxIN, functions as a protein–RNA toxin–antitoxin pair. Proc. Natl. Acad. Sci. USA 2009, 106, 894. [Google Scholar] [CrossRef] [PubMed]
- Samson, J.E.; Spinelli, S.; Cambillau, C.; Moineau, S. Structure and activity of AbiQ, a lactococcal endoribonuclease belonging to the type III toxin-antitoxin system. Mol. Microbiol. 2013, 87, 756–768. [Google Scholar] [CrossRef] [PubMed]
- Blower, T.R.; Short, F.L.; Rao, F.; Mizuguchi, K.; Pei, X.Y.; Fineran, P.C.; Luisi, B.F.; Salmond, G.P. Identification and classification of bacterial Type III toxin-antitoxin systems encoded in chromosomal and plasmid genomes. Nucleic Acids Res. 2012, 40, 6158–6173. [Google Scholar] [CrossRef] [PubMed]
- Goeders, N.; Chai, R.; Chen, B.; Day, A.; Salmond, G.P. Structure, Evolution, and Functions of Bacterial Type III Toxin-Antitoxin Systems. Toxins (Basel) 2016, 8, 282. [Google Scholar] [CrossRef] [PubMed]
- Dy, R.L.; Przybilski, R.; Semeijn, K.; Salmond, G.P.; Fineran, P.C. A widespread bacteriophage abortive infection system functions through a Type IV toxin-antitoxin mechanism. Nucleic Acids Res. 2014, 42, 4590–4605. [Google Scholar] [CrossRef]
- Masuda, H.; Tan, Q.; Awano, N.; Wu, K.P.; Inouye, M. YeeU enhances the bundling of cytoskeletal polymers of MreB and FtsZ, antagonizing the CbtA (YeeV) toxicity in Escherichia coli. Mol. Microbiol. 2012, 84, 979–989. [Google Scholar] [CrossRef]
- Masuda, H.; Tan, Q.; Awano, N.; Yamaguchi, Y.; Inouye, M. A novel membrane-bound toxin for cell division, CptA (YgfX), inhibits polymerization of cytoskeleton proteins, FtsZ and MreB, in Escherichia coli. FEMS Microbiol. Lett. 2012, 328, 174–181. [Google Scholar] [CrossRef]
- Wang, X.; Lord, D.M.; Cheng, H.Y.; Osbourne, D.O.; Hong, S.H.; Sanchez-Torres, V.; Quiroga, C.; Zheng, K.; Herrmann, T.; Peti, W.; et al. A new type V toxin-antitoxin system where mRNA for toxin GhoT is cleaved by antitoxin GhoS. Nat. Chem. Biol. 2012, 8, 855–861. [Google Scholar] [CrossRef]
- Barbosa, L.C.B.; dos Santos Carrijo, R.; da Conceição, M.B.; Campanella, J.E.M.; Júnior, E.C.; Secches, T.O.; Bertolini, M.C.; Marchetto, R. Characterization of an OrtT-like toxin of Salmonella enterica serovar Houten. Braz. J. Microbiol. 2019, 50, 839–848. [Google Scholar] [CrossRef]
- Islam, S.; Benedik, M.J.; Wood, T.K. Orphan toxin OrtT (YdcX) of Escherichia coli reduces growth during the stringent response. Toxins (Basel) 2015, 7, 299–321. [Google Scholar] [CrossRef]
- Aakre, C.D.; Phung, T.N.; Huang, D.; Laub, M.T. A bacterial toxin inhibits DNA replication elongation through a direct interaction with the beta sliding clamp. Mol. Cell 2013, 52, 617–628. [Google Scholar] [CrossRef]
- Gabrisko, M.; Barak, I. Evolution of the SpoIISABC Toxin-Antitoxin-Antitoxin System in Bacilli. Toxins (Basel) 2016, 8, 180. [Google Scholar] [CrossRef]
- Jimmy, S.; Saha, C.K.; Kurata, T.; Stavropoulos, C.; Oliveira, S.R.A.; Koh, A.; Cepauskas, A.; Takada, H.; Rejman, D.; Tenson, T.; et al. A widespread toxin-antitoxin system exploiting growth control via alarmone signaling. Proc. Natl. Acad. Sci. USA 2020, 117, 10500–10510. [Google Scholar] [CrossRef] [PubMed]
- Sala, A.; Calderon, V.; Bordes, P.; Genevaux, P. TAC from Mycobacterium tuberculosis: A paradigm for stress-responsive toxin-antitoxin systems controlled by SecB-like chaperones. Cell Stress Chaperones 2013, 18, 129–135. [Google Scholar] [CrossRef] [PubMed]
- Wilcox, B.; Osterman, I.; Serebryakova, M.; Lukyanov, D.; Komarova, E.; Gollan, B.; Morozova, N.; Wolf, Y.I.; Makarova, K.S.; Helaine, S.; et al. Escherichia coli ItaT is a type II toxin that inhibits translation by acetylating isoleucyl-tRNAIle. Nucleic Acids Res. 2018, 46, 7873–7885. [Google Scholar] [CrossRef] [PubMed]
- VanDrisse, M.C.; Parks, A.R.; Escalante-Semerena, J.C. A Toxin Involved in Salmonella Persistence Regulates Its Activity by Acetylating Its Cognate Antitoxin, a Modification Reversed by CobB Sirtuin Deacetylase. mBio 2017, 8, e00708-17. [Google Scholar] [CrossRef] [PubMed]
- Horesh, G.; Fino, C.; Harms, A.; Dorman, M.J.; Parts, L.; Gerdes, K.; Heinz, E.; Thomson, N.R. Type II and type IV toxin-antitoxin systems show different evolutionary patterns in the global Klebsiella pneumoniae population. Nucleic Acids Res. 2020, 48, 4357–4370. [Google Scholar] [CrossRef]
- Yamaguchi, Y.; Park, J.H.; Inouye, M. Toxin-antitoxin systems in bacteria and archaea. Ann. Rev. Genet. 2011, 45, 61–79. [Google Scholar] [CrossRef]
- Xie, Y.; Wei, Y.; Shen, Y.; Li, X.; Zhou, H.; Tai, C.; Deng, Z.; Ou, H.Y. TADB 2.0: An updated database of bacterial type II toxin-antitoxin loci. Nucleic Acids Res. 2018, 46, D749–D753. [Google Scholar] [CrossRef]
- Akarsu, H.; Bordes, P.; Mansour, M.; Bigot, D.J.; Genevaux, P.; Falquet, L. TASmania: A bacterial Toxin-Antitoxin Systems database. PLoS Comput. Biol. 2019, 15, e1006946. [Google Scholar] [CrossRef]
- Leplae, R.; Geeraerts, D.; Hallez, R.; Guglielmini, J.; Dreze, P.; Van Melderen, L. Diversity of bacterial type II toxin-antitoxin systems: A comprehensive search and functional analysis of novel families. Nucleic Acids Res. 2011, 39, 5513–5525. [Google Scholar] [CrossRef] [PubMed]
- Chan, T.W.; Balsa, D.; Espinosa, M. One cannot rule them all: Are bacterial toxins-antitoxins druggable? FEMS Microbiol. Rev. 2015, 39, 522–540. [Google Scholar] [CrossRef] [PubMed]
- Bernard, P.; Couturier, M. Cell killing by the F plasmid CcdB protein involves poisoning of DNA-topoisomerase II complexes. J. Mol. Biol. 1992, 226, 735–745. [Google Scholar] [CrossRef]
- Melderen, V.L.; Bernard, P.; Couturier, M. Lon-dependent proteolysis of CcdA is the key control for activation of CcdB in plasmid-free segregant bacteria. Mol. Microbiol. 1994, 11, 1151–1157. [Google Scholar] [CrossRef] [PubMed]
- Roberts, C.R.; Strom, A.R.; Helinski, D.R. The parDE operon of the broad-host-range plasmid RK2 specifies growth inhibition associated with plasmid loss. J. Mol. Biol. 1994, 237, 35–51. [Google Scholar] [CrossRef]
- Sobecky, P.A.; Easter, C.L.; Bear, P.D.; Helinski, D.R. Characterization of the stable maintenance properties of the par region of broad-host-range plasmid RK2. J. Bacteriol. 1996, 178, 2086–2093. [Google Scholar] [CrossRef][Green Version]
- Yarmolinsky, M.B. Programmed cell death in bacterial populations. Science 1995, 267, 836. [Google Scholar] [CrossRef]
- Koonin, E.V.; Makarova, K.S.; Aravind, L. Horizontal Gene Transfer in Prokaryotes: Quantification and Classification. Annu. Rev. Microbiol. 2001, 55, 709–742. [Google Scholar] [CrossRef]
- de Bast, S.M.; Mine, N.; van Melderen, L. Chromosomal toxin-antitoxin systems may act as antiaddiction modules. J. Bacteriol. 2008, 190, 4603–4609. [Google Scholar] [CrossRef]
- Makarova, S.K.; Wolf, Y.I.; Koonin, E.V. Comparative genomics of defense systems in archaea and bacteria. Nucleic Acids Res. 2013, 41, 4360–4377. [Google Scholar] [CrossRef]
- Hernández-Arriaga, A.M.; Chan, W.T.; Espinosa, M.; Díaz-Orejas, R. Conditional Activation of Toxin-Antitoxin Systems: Postsegregational Killing and Beyond. Microbiol. Spectr. 2014, 2, 175–192. [Google Scholar] [CrossRef] [PubMed]
- Mine, N.; Guglielmini, J.; Wilbaux, M.; Van Melderen, L. The decay of the chromosomally encoded ccdO157 toxin-antitoxin system in the Escherichia coli species. Genetics 2009, 181, 1557–1566. [Google Scholar] [CrossRef]
- Ramisetty, M.B.C.; Santhosh, R.S. Horizontal gene transfer of chromosomal Type II toxin–antitoxin systems of Escherichia coli. FEMS Microbiol. Lett. 2016, 363. [Google Scholar] [CrossRef] [PubMed]
- Ramisetty, B.C.M.; Sudhakari, P.A. Bacterial ‘Grounded’ Prophages: Hotspots for Genetic Renovation and Innovation. Front. Genet. 2019, 10, 65. [Google Scholar] [CrossRef] [PubMed]
- Koonin, V.E.; Wolf, Y.I. Genomics of bacteria and archaea: The emerging dynamic view of the prokaryotic world. Nucleic Acids Res. 2008, 36, 6688–6719. [Google Scholar] [CrossRef]
- Andersen, S.B.; Ghoul, M.; Griffin, A.S.; Petersen, B.; Johansen, H.K.; Molin, S. Diversity, Prevalence, and Longitudinal Occurrence of Type II Toxin-Antitoxin Systems of Pseudomonas aeruginosa Infecting Cystic Fibrosis Lungs. Front. Microbiol. 2017, 8, 1180. [Google Scholar] [CrossRef] [PubMed]
- Diaz-Orejas, R.; Espinosa, M.; Yeo, C.C. The Importance of the Expendable: Toxin-Antitoxin Genes in Plasmids and Chromosomes. Front. Microbiol. 2017, 8, 1479. [Google Scholar] [CrossRef]
- Gupta, K.; Tripathi, A.; Sahu, A.; Varadarajan, R. Contribution of the chromosomal ccdAB operon to bacterial drug tolerance. J. Bacteriol. 2017, 199, e00397-17. [Google Scholar] [CrossRef]
- Ames, J.R.; Muthuramalingam, M.; Murphy, T.; Najar, F.Z.; Bourne, C.R. Expression of different ParE toxins results in conserved phenotypes with distinguishable classes of toxicity. MicrobiologyOpen 2019, 8, e902. [Google Scholar] [CrossRef]
- Fiebig, A.; Castro Rojas, C.M.; Siegal-Gaskins, D.; Crosson, S. Interaction specificity, toxicity and regulation of a paralogous set of ParE/RelE-family toxin-antitoxin systems. Mol. Microbiol. 2010, 77, 236–251. [Google Scholar] [CrossRef]
- Muthuramalingam, M.; White, J.C.; Murphy, T.; Ames, J.R.; Bourne, C.R. The toxin from a ParDE toxin-antitoxin system found in Pseudomonas aeruginosa offers protection to cells challenged with anti-gyrase antibiotics. Mol. Microbiol. 2018, 111, 441–454. [Google Scholar] [CrossRef] [PubMed]
- Tamman, H.; Ainelo, A.; Ainsaar, K.; Hõrak, R. A moderate toxin, GraT, modulates growth rate and stress tolerance of Pseudomonas putida. J. Bacteriol. 2014, 196, 157–169. [Google Scholar] [CrossRef] [PubMed]
- Wilbaux, M.; Mine, N.; Guérout, A.M.; Mazel, D.; Van Melderen, L. Functional interactions between coexisting toxin-antitoxin systems of the ccd family in Escherichia coli O157:H7. J. Bacteriol. 2007, 189, 2712–2719. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Garcia, L.; Kim, J.S.; Tomas, M.; Wood, T.K. Toxins of toxin/antitoxin systems are inactivated primarily through promoter mutations. J. Appl. Microbiol. 2019, 127, 1859–1868. [Google Scholar] [CrossRef] [PubMed]
- Deter, H.S.; Jensen, R.V.; Mather, W.H.; Butzin, N.C. Mechanisms for Differential Protein Production in Toxin-Antitoxin Systems. Toxins (Basel) 2017, 9, 211. [Google Scholar] [CrossRef]
- Singh, R.; Barry, C.E., 3rd; Boshoff, H.I. The three RelE homologs of Mycobacterium tuberculosis have individual, drug-specific effects on bacterial antibiotic tolerance. J. Bacteriol. 2010, 192, 1279–1291. [Google Scholar] [CrossRef]
- Pontes, H.M.; Groisman, E.A. Slow growth determines nonheritable antibiotic resistance in Salmonella enterica. Sci. Signal. 2019, 12, eaax3938. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, A.; Dewan, P.C.; Barua, B.; Varadarajan, R. Additional role for the ccd operon of F-plasmid as a transmissible persistence factor. Proc. Natl Acad. Sci. USA 2012, 109, 12497–12502. [Google Scholar] [CrossRef] [PubMed]
- Kamruzzaman, M.; Iredell, J. A ParDE-family toxin antitoxin system in major resistance plasmids of Enterobacteriaceae confers antibiotic and heat tolerance. Sci. Rep. 2019, 9, 9872. [Google Scholar] [CrossRef]
- Gupta, M.; Nayyar, N.; Chawla, M.; Sitaraman, R.; Bhatnagar, R.; Banerjee, N. The chromosomal parDE2 toxin-antitoxin system of Mycobacterium tuberculosis H37Rv: Genetic and functional characterization. Front. Microbiol. 2016, 7, 886. [Google Scholar] [CrossRef]
- Harms, A.; Stanger, F.V.; Scheu, P.D.; de Jong, I.G.; Goepfert, A.; Glatter, T.; Gerdes, K.; Schirmer, T.; Dehio, C. Adenylylation of Gyrase and Topo IV by FicT Toxins Disrupts Bacterial DNA Topology. Cell Rep. 2015, 12, 1497–1507. [Google Scholar] [CrossRef] [PubMed]
- Janssen, D.B.; Garza-Sanchez, F.; Hayes, C.S. YoeB toxin is activated during thermal stress. MicrobiologyOpen 2015, 4, 682–697. [Google Scholar] [CrossRef]
- Chan, W.T.; Domenech, M.; Moreno-Córdoba, I.; Navarro-Martínez, V.; Nieto, C.; Moscoso, M.; García, E.; Espinosa, M. The Streptococcus pneumoniae yefM-yoeB and relBE Toxin-Antitoxin Operons Participate in Oxidative Stress and Biofilm Formation. Toxins (Basel) 2018, 10, 378. [Google Scholar] [CrossRef] [PubMed]
- Ainelo, A.; Porosk, R.; Kilk, K.; Rosendahl, S.; Remme, J.; Hõrak, R. Pseudomonas putida Responds to the Toxin GraT by Inducing Ribosome Biogenesis Factors and Repressing TCA Cycle Enzymes. Toxins (Basel) 2019, 11, 103. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; McAnulty, M.J.; Wood, T.K. Toxin YafQ Reduces Escherichia coli Growth at Low Temperatures. PLoS ONE 2016, 11, e0161577. [Google Scholar] [CrossRef] [PubMed]
- Korch, S.B.; Malhotra, V.; Contreras, H.; Clark-Curtiss, J.E. The Mycobacterium tuberculosis relBE toxin:antitoxin genes are stress-responsive modules that regulate growth through translation inhibition. J. Microbiol. 2015, 53, 783–795. [Google Scholar] [CrossRef]
- Christensen, K.S.; Gerdes, K. Delayed-relaxed response explained by hyperactivation of RelE. Mol. Microbiol. 2004, 53, 587–597. [Google Scholar] [CrossRef]
- Tashiro, Y.; Kawata, K.; Taniuchi, A.; Kakinuma, K.; May, T.; Okabe, S. RelE-mediated dormancy is enhanced at high cell density in Escherichia coli. J. Bacteriol. 2012, 194, 1169–1176. [Google Scholar] [CrossRef]
- Armalytė, J.; Jurėnas, D.; Krasauskas, R.; Čepauskas, A.; Sužiedėlienė, E. The higBA Toxin-Antitoxin Module From the Opportunistic Pathogen Acinetobacter baumannii—Regulation, Activity, and Evolution. Front. Microbiol. 2018, 9, 732. [Google Scholar] [CrossRef]
- Wood, L.T.; Wood, T.K. The HigB/HigA toxin/antitoxin system of Pseudomonas aeruginosa influences the virulence factors pyochelin, pyocyanin, and biofilm formation. MicrobiolOpen 2016, 5, 499–511. [Google Scholar] [CrossRef]
- Ramisetty, B.; Ghosh, D.; Roy Chowdhury, M.; Santhosh, R.S. What Is the Link between Stringent Response, Endoribonuclease Encoding Type II Toxin-Antitoxin Systems and Persistence? Front. Microbiol. 2016, 7, 1882. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Inouye, M. RatA (YfjG), an Escherichia coli toxin, inhibits 70S ribosome association to block translation initiation. Mol. Microbiol. 2011, 79, 1418–1429. [Google Scholar] [CrossRef] [PubMed]
- Norton, P.J.; Mulvey, M.A. Toxin-antitoxin systems are important for niche-specific colonization and stress resistance of uropathogenic Escherichia coli. PLoS Pathog. 2012, 8, e1002954. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, P.; Arora, G.; Singh, M.; Kidwai, S.; Narayan, O.P.; Singh, R. MazF ribonucleases promote Mycobacterium tuberculosis drug tolerance and virulence in guinea pigs. Nat. Commun. 2015, 6, 6059. [Google Scholar] [CrossRef]
- Ren, D.; Walker, A.N.; Daines, D.A. Toxin-antitoxin loci vapBC-1 and vapXD contribute to survival and virulence in nontypeable Haemophilus influenzae. BMC Microbiol. 2012, 12, 263. [Google Scholar] [CrossRef]
- Jaiswal, S.; Paul, P.; Padhi, C.; Ray, S.; Ryan, D.; Dash, S.; Suar, M. The Hha-TomB Toxin-Antitoxin System Shows Conditional Toxicity and Promotes Persister Cell Formation by Inhibiting Apoptosis-Like Death in S. Typhimurium. Sci. Rep. 2016, 6, 38204. [Google Scholar] [CrossRef]
- Helaine, S.; Cheverton, A.M.; Watson, K.G.; Faure, L.M.; Matthews, S.A.; Holden, D.W. Internalization of Salmonella by macrophages induces formation of nonreplicating persisters. Science 2014, 343, 204–208. [Google Scholar] [CrossRef]
- Fraikin, N.; Rousseau, C.J.; Goeders, N.; Van Melderen, L. Reassessing the Role of the Type II MqsRA Toxin-Antitoxin System in Stress Response and Biofilm Formation: mqsA Is Transcriptionally Uncoupled from mqsR. mBio 2019, 10, e02678-19. [Google Scholar] [CrossRef]
- Gerdes, K.; Christensen, S.K.; Lobner-Olesen, A. Prokaryotic toxin-antitoxin stress response loci. Nat. Rev. Microbiol. 2005, 3, 371–382. [Google Scholar] [CrossRef]
- Kim, Y.; Wang, X.; Zhang, X.S.; Grigoriu, S.; Page, R.; Peti, W.; Wood, T.K. Escherichia coli toxin/antitoxin pair MqsR/MqsA regulate toxin CspD. Environ. Microbiol. 2010, 12, 1105–1121. [Google Scholar] [CrossRef]
- Wade, T.J.; Laub, M.T. Concerns about “Stress-Induced MazF-Mediated Proteins in Escherichia coli”. mBio 2019, 10, e00825-19. [Google Scholar] [CrossRef] [PubMed]
- Culviner, H.P.; Laub, M.T. Global Analysis of the E. coli Toxin MazF Reveals Widespread Cleavage of mRNA and the Inhibition of rRNA Maturation and Ribosome Biogenesis. Mol. Cell 2018, 70, 868–880. [Google Scholar] [CrossRef] [PubMed]
- Nigam, A.; Ziv, T.; Oron-Gottesman, A. Engelberg-Kulka Stress-Induced MazF-Mediated Proteins in Escherichia coli. mBio 2019, 10, e00340-19. [Google Scholar] [PubMed]
- Harms, A.; Liesch, M.; Körner, J.; Québatte, M.; Engel, P.; Dehio, C. A bacterial toxin-antitoxin module is the origin of inter-bacterial and inter-kingdom effectors of Bartonella. PLoS Genet. 2017, 13, e1007077. [Google Scholar] [CrossRef] [PubMed]
- Harms, A.; Maisonneuve, E.; Gerdes, K. Mechanisms of bacterial persistence during stress and antibiotic exposure. Science 2016, 354, aaf4268. [Google Scholar] [CrossRef] [PubMed]
- Goormaghtigh, F.; Fraikin, N.; Putrinš, M.; Hallaert, T.; Hauryliuk, V.; Garcia-Pino, A.; Sjödin, A.; Kasvandik, S.; Udekwu, K.; Tenson, T.; et al. Reassessing the Role of Type II Toxin-Antitoxin Systems in Formation of Escherichia coli Type II Persister Cells. mBio 2018, 9, e00640-18. [Google Scholar] [CrossRef] [PubMed]
- Goormaghtigh, F.; Fraikin, N.; Putrinš, M.; Hauryliuk, V.; Garcia-Pino, A.; Udekwu, K.; Tenson, T.; Kaldalu, N.; Van Melderen, L. Reply to Holden and Errington, “Type II Toxin-Antitoxin Systems and Persister Cells”. mBio 2018, 9, e01838-18. [Google Scholar] [CrossRef] [PubMed]
- Zielenkiewicz, U.; Ceglowski, P. The toxin-antitoxin system of the streptococcal plasmid pSM19035. J. Bacteriol. 2005, 187, 6094–6105. [Google Scholar] [CrossRef]
- Chan, T.W.; Espinosa, M. The Streptococcus pneumoniae pezAT Toxin-Antitoxin System Reduces beta-Lactam Resistance and Genetic Competence. Front. Microbiol. 2016, 7, 1322. [Google Scholar] [CrossRef]
- Piscotta, J.F.; Jeffrey, P.D.; Link, A.J. ParST is a widespread toxin-antitoxin module that targets nucleotide metabolism. Proc. Natl. Acad. Sci. USA 2019, 116, 826–834. [Google Scholar] [CrossRef]
- Yuan, J.; Yamaichi, Y.; Waldor, M.K. The three Vibrio cholerae chromosome II-encoded ParE toxins degrade chromosome I following loss of chromosome II. J. Bacteriol. 2011, 193, 611–619. [Google Scholar] [CrossRef] [PubMed]
- Wozniak, A.R.; Waldor, M.K. A toxin-antitoxin system promotes the maintenance of an integrative conjugative element. PLoS Genet. 2009, 5, e1000439. [Google Scholar] [CrossRef] [PubMed]
- Hughes-Games, A.; Roberts, A.P.; Davis, S.A.; Hill, D.J. Identification of integrative and conjugative elements in pathogenic and commensal Neisseriaceae species via genomic distributions of DNA uptake sequence dialects. Microb. Genom. 2020, 6, e000372. [Google Scholar]
- Berni, B.; Soscia, C.; Djermoun, S.; Ize, B.; Bleves, S. A Type VI Secretion System Trans-Kingdom Effector Is Required for the Delivery of a Novel Antibacterial Toxin in Pseudomonas aeruginosa. Front. Microbiol. 2019, 10, 1218. [Google Scholar] [CrossRef]
- Cao, Z.; Casabona, M.G.; Kneuper, H.; Chalmers, J.D.; Palmer, T. The type VII secretion system of Staphylococcus aureus secretes a nuclease toxin that targets competitor bacteria. Nat. Microbiol. 2016, 2, 16183. [Google Scholar] [CrossRef]
- Whitney, J.C.; Peterson, S.B.; Kim, J.; Pazos, M.; Verster, A.J.; Radey, M.C.; Kulasekara, H.D.; Ching, M.Q.; Bullen, N.P.; Bryant, D.; et al. A broadly distributed toxin family mediates contact-dependent antagonism between gram-positive bacteria. Elife 2017, 6, e26938. [Google Scholar] [CrossRef]
- Gerdes, K.; Bech, F.W.; Jørgensen, S.T.; Løbner-Olesen, A.; Rasmussen, P.B.; Atlung, T.; Boe, L.; Karlstrom, O.; Molin, S.; Von Meyenburg, K. Mechanism of postsegregational killing by the hok gene product of the parB system of plasmid R1 and its homology with the relF gene product of the E. coli relB operon. EMBO J. 1986, 5, 2023–2029. [Google Scholar] [CrossRef]
- Ogura, T.; Hiraga, S. Mini-F plasmid genes that couple host cell division to plasmid proliferation. Proc. Natl. Acad. Sci. USA 1983, 80, 4784–4788. [Google Scholar] [CrossRef]
- Aizenman, E.; Engelberg-Kulka, H.; Glaser, G. An Escherichia coli chromosomal “addiction module” regulated by guanosine 3′,5′-bispyrophosphate: A model for programmed bacterial cell death. Proc. Natl. Acad. Sci. USA 1996, 93, 6059–6063. [Google Scholar] [CrossRef]
- Jensen, B.R.; Gerdes, K. Programmed cell death in bacteria: Proteic plasmid stabilization systems. Mol. Microbiol. 1995, 17, 205–210. [Google Scholar] [CrossRef]
- Zander, I.; Shmidov, E.; Roth, S.; Ben-David, Y.; Shoval, I.; Shoshani, S.; Danielli, A.; Banin, E. Characterization of PfiT/PfiA toxin-antitoxin system of Pseudomonas aeruginosa that affects cell elongation and prophage induction. Environ. Microbiol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Liu, X.; Tang, K.; Wang, W.; Guo, Y.; Wang, X. Prophage encoding toxin/antitoxin system PfiT/PfiA inhibits Pf4 production in Pseudomonas aeruginosa. Microb. Biotechnol. 2020, 13, 1132–1144. [Google Scholar] [CrossRef] [PubMed]
- Bustamante, P.; Iredell, J.R. Carriage of type II toxin-antitoxin systems by the growing group of IncX plasmids. Plasmid 2017, 91, 19–27. [Google Scholar] [CrossRef] [PubMed]
- Cooper, F.T.; Paixao, T.; Heinemann, J.A. Within-host competition selects for plasmid-encoded toxin-antitoxin systems. Proc. Biol. Sci. 2010, 277, 3149–3155. [Google Scholar] [CrossRef] [PubMed]
- Fico, S.; Mahillon, J. TasA-tasB, a new putative toxin-antitoxin (TA) system from Bacillus thuringiensis pGI1 plasmid is a widely distributed composite mazE-doc TA system. BMC Genom. 2006, 7, 259. [Google Scholar] [CrossRef]
- Hernández-Ramírez, K.C.; Chávez-Jacobo, V.M.; Valle-Maldonado, M.I.; Patiño-Medina, J.A.; Díaz-Pérez, S.P.; Jácome-Galarza, I.E.; Ortiz-Alvarado, R.; Meza-Carmen, V.; Ramírez-Díaz, M.I. Plasmid pUM505 encodes a Toxin-Antitoxin system conferring plasmid stability and increased Pseudomonas aeruginosa virulence. Microb. Pathog. 2017, 112, 259–268. [Google Scholar] [CrossRef] [PubMed]
- Poirel, L.; Bonnin, R.A.; Nordmann, P. Genetic features of the widespread plasmid coding for the carbapenemase OXA-48. Antimicrob. Agents Chemother. 2012, 56, 559–562. [Google Scholar] [CrossRef]
- Mnif, B.; Vimont, S.; Boyd, A.; Bourit, E.; Picard, B.; Branger, C.; Denamur, E.; Arlet, G. Molecular characterization of addiction systems of plasmids encoding extended-spectrum beta-lactamases in Escherichia coli. J. Antimicrob. Chemother. 2010, 65, 1599–1603. [Google Scholar] [CrossRef]
- Davis, L.T.; Helinski, D.R.; Roberts, R.C. Transcription and autoregulation of the stabilizing functions of broad-host-range plasmid RK2 in Escherichia coli, Agrobacterium tumefaciens and Pseudomonas aeruginosa. Mol. Microbiol. 1992, 6, 1981–1994. [Google Scholar] [CrossRef]
- Gronlund, H.; Gerdes, K. Toxin-antitoxin systems homologous with relBE of Escherichia coli plasmid P307 are ubiquitous in prokaryotes. J. Mol. Biol. 1999, 285, 1401–1415. [Google Scholar] [CrossRef]
- Wang, J.; Stephan, R.; Zurfluh, K.; Hächler, H.; Fanning, S. Characterization of the genetic environment of bla ESBL genes, integrons and toxin-antitoxin systems identified on large transferrable plasmids in multi-drug resistant Escherichia coli. Front. Microbiol. 2014, 5, 716. [Google Scholar] [PubMed]
- Hayes, F. A family of stability determinants in pathogenic bacteria. J. Bacteriol. 1998, 180, 6415–6418. [Google Scholar] [CrossRef]
- Bravo, A.; de Torrontegui, G.; Diaz, R. Identification of components of a new stability system of plasmid R1, ParD, that is close to the origin of replication of this plasmid. Mol. Gen. Genet. 1987, 210, 101–110. [Google Scholar] [CrossRef] [PubMed]
- Diago-Navarro, E.; Hernandez-Arriaga, A.M.; López-Villarejo, J.; Muñoz-Gómez, A.J.; Kamphuis, M.B.; Boelens, R.; Lemonnier, M.; Díaz-Orejas, R. parD toxin-antitoxin system of plasmid R1--basic contributions, biotechnological applications and relationships with closely-related toxin-antitoxin systems. FEBS J. 2010, 277, 3097–3117. [Google Scholar] [CrossRef] [PubMed]
- Tsuchimoto, S.; Ohtsubo, H.; Ohtsubo, E. Two genes, pemK and pemI, responsible for stable maintenance of resistance plasmid R100. J. Bacteriol. 1988, 170, 1461–1466. [Google Scholar] [CrossRef]
- Bukowski, M.; Hyz, K.; Janczak, M.; Hydzik, M.; Dubin, G.; Wladyka, B. Identification of novel mazEF/pemIK family toxin-antitoxin loci and their distribution in the Staphylococcus genus. Sci. Rep. 2017, 7, 13462. [Google Scholar] [CrossRef]
- Lehnherr, H.; Maguin, E.; Jafri, S.; Yarmolinsky, M.B. Plasmid addiction genes of bacteriophage P1: Doc, which causes cell death on curing of prophage, and phd, which prevents host death when prophage is retained. J. Mol. Biol. 1993, 233, 414–428. [Google Scholar] [CrossRef]
- Tian, Q.B.; Hayashi, T.; Murata, T.; Terawaki, Y. Gene product identification and promoter analysis of hig locus of plasmid Rts1. Biochem. Biophys Res. Commun. 1996, 225, 679–684. [Google Scholar] [CrossRef]
- Dziewit, L.; Jazurek, M.; Drewniak, L.; Baj, J.; Bartosik, D. The SXT conjugative element and linear prophage N15 encode toxin-antitoxin-stabilizing systems homologous to the tad-ata module of the Paracoccus aminophilus plasmid pAMI2. J. Bacteriol. 2007, 189, 1983–1997. [Google Scholar] [CrossRef]
- Grady, R.; Hayes, F. Axe-Txe, a broad-spectrum proteic toxin-antitoxin system specified by a multidrug-resistant, clinical isolate of Enterococcus faecium. Mol. Microbiol. 2003, 47, 1419–1432. [Google Scholar] [CrossRef]
- Rosvoll, T.C.; Pedersen, T.; Sletvold, H.; Johnsen, P.J.; Sollid, J.E.; Simonsen, G.S.; Jensen, L.B.; Nielsen, K.M.; Sundsfjord, A. PCR-based plasmid typing in Enterococcus faecium strains reveals widely distributed pRE25-, pRUM-, pIP501- and pHTbeta-related replicons associated with glycopeptide resistance and stabilizing toxin-antitoxin systems. FEMS Immunol. Med. Microbiol. 2010, 58, 254–268. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.X.; Guo, X.K.; Chuan, W.U.; Bo, B.I.; Ren, S.X.; Wu, C.F.; Zhao, G.P. Characterization of a novel toxin-antitoxin module, VapBC, encoded by Leptospira interrogans chromosome. Cell Res. 2004, 14, 208–216. [Google Scholar] [CrossRef] [PubMed]
- Jurėnas, D.; Chatterjee, S.; Konijnenberg, A.; Sobott, F.; Droogmans, L.; Garcia-Pino, A.; Van Melderen, L. AtaT blocks translation initiation by N-acetylation of the initiator tRNAfMet. Nat. Chem. Biol. 2017, 13, 640. [Google Scholar] [CrossRef] [PubMed]
- Wen, Y.; Behiels, E.; Devreese, B. Toxin-Antitoxin systems: Their role in persistence, biofilm formation, and pathogenicity. Pathog. Dis. 2014, 70, 240–249. [Google Scholar] [CrossRef] [PubMed]
- Jurenaite, M.; Markuckas, A.; Suziedeliene, E. Identification and characterization of type II toxin-antitoxin systems in the opportunistic pathogen Acinetobacter baumannii. J. Bacteriol. 2013, 195, 3165–3172. [Google Scholar] [CrossRef] [PubMed]
- McVicker, G.; Tang, C.M. Deletion of toxin-antitoxin systems in the evolution of Shigella sonnei as a host-adapted pathogen. Nat. Microbiol. 2016, 2, 16204. [Google Scholar] [CrossRef]
- Cooper, F.T.; Heinemann, J.A. Postsegregational killing does not increase plasmid stability but acts to mediate the exclusion of competing plasmids. Proc. Natl. Acad. Sci. USA 2000, 97, 12643–12648. [Google Scholar] [CrossRef]
- Song, S.; Wood, T.K. Post-segregational Killing and Phage Inhibition Are Not Mediated by Cell Death Through Toxin/Antitoxin Systems. Front. Microbiol. 2018, 9, 814. [Google Scholar] [CrossRef]
- Otsuka, Y.; Yonesaki, T. Dmd of bacteriophage T4 functions as an antitoxin against Escherichia coli LsoA and RnlA toxins. Mol. Microbiol. 2012, 83, 669–681. [Google Scholar] [CrossRef]
- Blower, T.R.; Fineran, P.C.; Johnson, M.J.; Toth, I.K.; Humphreys, D.P.; Salmond, G.P.C. Mutagenesis and functional characterization of the RNA and protein components of the toxIN abortive infection and toxin-antitoxin locus of Erwinia. J. Bacteriol. 2009, 191, 6029–6039. [Google Scholar] [CrossRef]
- Dy, R.L.; Richter, C.; Salmond, G.P.; Fineran, P.C. Remarkable Mechanisms in Microbes to Resist Phage Infections. Annu. Rev. Virol. 2014, 1, 307–331. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; Akusobi, C.; Fang, X.; Salmond, G.P. Environmental T4-Family Bacteriophages Evolve to Escape Abortive Infection via Multiple Routes in a Bacterial Host Employing “Altruistic Suicide” through Type III Toxin-Antitoxin Systems. Front. Microbiol. 2017, 8, 1006. [Google Scholar] [CrossRef] [PubMed]
- Park, J.S.; Son, W.S.; Lee, B.J. Structural overview of toxin-antitoxin systems in infectious bacteria: A target for developing antimicrobial agents. Biochim. Biophys. Acta 2013, 1843, 1155–1167. [Google Scholar] [CrossRef]
- Arbing, M.A.; Handelman, S.K.; Kuzin, A.P.; Verdon, G.; Wang, C.; Su, M.; Rothenbacher, F.P.; Abashidze, M.; Liu, M.; Hurley, J.M.; et al. Crystal structures of Phd-Doc, HigA, and YeeU establish multiple evolutionary links between microbial growth-regulating toxin-antitoxin systems. Structure (London) 2010, 18, 996–1010. [Google Scholar] [CrossRef] [PubMed]
- Meinhart, A.; Alonso, J.C.; Sträter, N.; Saenger, W. Crystal structure of the plasmid maintenance system epsilon/zeta: Functional mechanism of toxin zeta and inactivation by epsilon 2 zeta 2 complex formation. Proc. Natl. Acad. Sci. USA 2003, 100, 1661–1666. [Google Scholar] [CrossRef]
- Jaen-Luchoro, D.; Aliaga-Lozano, F.; Gomila, R.M.; Gomila, M.; Salva-Serra, F.; Lalucat, J.; Bennasar-Figueras, A. First insights into a type II toxin-antitoxin system from the clinical isolate Mycobacterium sp. MHSD3, similar to epsilon/zeta systems. PLoS ONE 2017, 12, e0189459. [Google Scholar] [CrossRef]
- Fernández-Bachiller, M.I.; Brzozowska, I.; Odolczyk, N.; Zielenkiewicz, U.; Zielenkiewicz, P.; Rademann, J. Mapping Protein-Protein Interactions of the Resistance-Related Bacterial Zeta Toxin-Epsilon Antitoxin Complex (epsilon(2)zeta(2)) with High Affinity Peptide Ligands Using Fluorescence Polarization. Toxins (Basel) 2016, 8, 222. [Google Scholar] [CrossRef]
- Budde, P.P.; Davis, B.M.; Yuan, J.; Waldor, M.K. Characterization of a higBA toxin-antitoxin locus in Vibrio cholerae. J. Bacteriol. 2007, 189, 491–500. [Google Scholar] [CrossRef]
- Yang, J.; Zhou, K.; Liu, P.; Dong, Y.; Gao, Z.; Zhang, J.; Liu, Q. Structural insight into the E. coli HigBA complex. Biochem. Biophys. Res. Commun. 2016, 478, 1521–1527. [Google Scholar] [CrossRef]
- Sterckx, Y.G.J.; Jové, T.; Shkumatov, A.V.; Garcia-Pino, A.; Geerts, L.; De Kerpel, M.; Lah, J.; De Greve, H.; Van Melderen, L.; Loris, R. A unique hetero-hexadecameric architecture displayed by the Escherichia coli O157 PaaA2-ParE2 antitoxin-toxin complex. J. Mol. Biol. 2016, 428, 1589–1603. [Google Scholar] [CrossRef]
- Aakre, C.D.; Herrou, J.; Phung, T.N.; Perchuk, B.S.; Crosson, S.; Laub, M.T. Evolving new protein-protein interaction specificity through promiscuous intermediates. Cell 2015, 163, 594–606. [Google Scholar] [CrossRef] [PubMed]
- El-Gebali, S.; Mistry, J.; Bateman, A.; Eddy, S.R.; Luciani, A.; Potter, S.C.; Qureshi, M.; Richardson, L.J.; Salazar, G.A.; Smart, A.; et al. The Pfam protein families database in 2019. Nucleic Acids Res. 2019, 47, D427–D432. [Google Scholar] [CrossRef]
- Chan, T.W.; Espinosa, M.; Yeo, C.C. Keeping the Wolves at Bay: Antitoxins of Prokaryotic Type II Toxin-Antitoxin Systems. Front. Mol. Biosci. 2016, 3, 9. [Google Scholar] [CrossRef] [PubMed]
- Senissar, M.; Manav, M.C.; Brodersen, D.E. Structural conservation of the PIN domain active site across all domains of life. Protein Sci. 2017, 26, 1474–1492. [Google Scholar] [CrossRef] [PubMed]
- Matelska, D.; Steczkiewicz, K.; Ginalski, K. Comprehensive classification of the PIN domain-like superfamily. Nucleic Acids Res. 2017, 45, 6995–7020. [Google Scholar] [CrossRef]
- Arcus, V.L.; McKenzie, J.L.; Robson, J.; Cook, G.M. The PIN-domain ribonucleases and the prokaryotic VapBC toxin-antitoxin array. Protein Eng. Des. Sel. 2011, 24, 33–40. [Google Scholar] [CrossRef]
- Miallau, L.; Faller, M.; Chiang, J.; Arbing, M.; Guo, F.; Cascio, D.; Eisenberg, D. Structure and proposed activity of a member of the VapBC family of toxin-antitoxin systems. VapBC-5 from Mycobacterium tuberculosis. J. Biol. Chem. 2009, 284, 276–283. [Google Scholar] [CrossRef]
- Maté, M.J.; Vincentelli, R.; Foos, N.; Raoult, D.; Cambillau, C.; Ortiz-Lombardía, M. Crystal structure of the DNA-bound VapBC2 antitoxin/toxin pair from Rickettsia felis. Nucleic Acids Res. 2012, 40, 3245–3258. [Google Scholar]
- Mattison, K.; Wilbur, J.S.; So, M.; Brennan, R.G. Structure of FitAB from Neisseria gonorrhoeae bound to DNA reveals a tetramer of toxin-antitoxin heterodimers containing pin domains and ribbon-helix-helix motifs. J. Biol. Chem. 2006, 281, 37942–37951. [Google Scholar] [CrossRef]
- Min, A.B.; Miallau, L.; Sawaya, M.R.; Habel, J.; Cascio, D.; Eisenberg, D. The crystal structure of the Rv0301-Rv0300 VapBC-3 toxin-antitoxin complex from M. tuberculosis reveals a Mg(2)(+) ion in the active site and a putative RNA-binding site. Protein Sci. 2012, 21, 1754–1767. [Google Scholar]
- Schureck, M.A.; Dunkle, J.A.; Maehigashi, T.; Miles, S.J.; Dunham, C.M. Defining the mRNA recognition signature of a bacterial toxin protein. Proc. Natl. Acad. Sci. USA 2015, 112, 13862–13867. [Google Scholar] [CrossRef] [PubMed]
- Schureck, M.A.; Repack, A.; Miles, S.J.; Marquez, J.; Dunham, C.M. Mechanism of endonuclease cleavage by the HigB toxin. Nucleic Acids Res. 2016, 44, 7944–7953. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.; Sterckx, Y.; Mitchenall, L.A.; Maxwell, A.; Loris, R.; Waldor, M.K. Vibrio cholerae ParE2 poisons DNA gyrase via a mechanism distinct from other gyrase inhibitors. J. Biol. Chem. 2010, 285, 40397–40408. [Google Scholar] [CrossRef] [PubMed]
- Brown, B.L.; Grigoriu, S.; Kim, Y.; Arruda, J.M.; Davenport, A.; Wood, T.K.; Peti, W.; Page, R. Three dimensional structure of the MqsR:MqsA complex: A novel TA pair comprised of a toxin homologous to RelE and an antitoxin with unique properties. PLoS Pathog. 2009, 5, e1000706. [Google Scholar] [CrossRef] [PubMed]
- Hallez, R.; Geeraerts, D.; Sterckx, Y.; Mine, N.; Loris, R.; Van Melderen, L. New toxins homologous to ParE belonging to three-component toxin-antitoxin systems in Escherichia coli O157:H7. Mol. Microbiol. 2010, 76, 719–732. [Google Scholar] [CrossRef]
- Bøggild, A.; Sofos, N.; Andersen, K.R.; Feddersen, A.; Easter, A.D.; Passmore, L.A.; Brodersen, D.E. The crystal structure of the intact E. coli RelBE toxin-antitoxin complex provides the structural basis for conditional cooperativity. Structure (London) 2012, 20, 1641–1658. [Google Scholar]
- Dalton, M.K.; Crosson, S. A conserved mode of protein recognition and binding in a ParD-ParE toxin-antitoxin complex. Biochemistry 2010, 49, 2205–2215. [Google Scholar] [CrossRef]
- Hargreaves, D.; Santos-Sierra, S.; Giraldo, R.; Sabariegos-Jareño, R.; de la Cueva-Méndez, G.; Boelens, R.; Díaz-Orejas, R.; Rafferty, J.B. Structural and functional analysis of the kid toxin protein from E. coli plasmid R1. Structure 2002, 10, 1425–1433. [Google Scholar]
- Kamada, K.; Hanaoka, F.; Burley, S.K. Crystal structure of the MazE/MazF complex: Molecular bases of antidote-toxin recognition. Mol. Cell 2003, 11, 875–884. [Google Scholar] [CrossRef]
- De Jonge, N.; Garcia-Pino, A.; Buts, L.; Haesaerts, S.; Charlier, D.; Zangger, K.; Wyns, L.; De Greve, H.; Loris, R. Rejuvenation of CcdB-poisoned gyrase by an intrinsically disordered protein domain. Mol. Cell 2009, 35, 154–163. [Google Scholar] [CrossRef]
- Bahassi, E.M.; O’Dea, M.H.; Allali, N.; Messens, J.; Gellert, M.; Couturier, M. Interactions of CcdB with DNA gyrase. Inactivation of Gyra, poisoning of the gyrase-DNA complex, and the antidote action of CcdA. J. Biol. Chem. 1999, 274, 10936–10944. [Google Scholar] [CrossRef] [PubMed]
- Kampranis, C.S.; Howells, A.J.; Maxwell, A. The interaction of DNA gyrase with the bacterial toxin CcdB: Evidence for the existence of two gyrase-CcdB complexes. J. Mol. Biol. 1999, 293, 733–744. [Google Scholar] [CrossRef] [PubMed]
- Simic, M.; De Jonge, N.; Loris, R.; Vesnaver, G.; Lah, J. Driving forces of gyrase recognition by the addiction toxin CcdB. J. Biol. Chem. 2009, 284, 20002–20010. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.B.; López-Villarejo, J.; Diago-Navarro, E.; Mitchenall, L.A.; Barendregt, A.; Heck, A.J.; Lemonnier, M.; Maxwell, A.; Díaz-Orejas, R. A common origin for the bacterial toxin-antitoxin systems parD and ccd, suggested by analyses of toxin/target and toxin/antitoxin interactions. PLoS ONE 2012, 7, e46499. [Google Scholar] [CrossRef]
- Ruiz-Echevarría, M.J.; Giménez-Gallego, G.; Sabariegos-Jareño, R.; Díaz-Orejas, R. Kid, a small protein of the parD stability system of plasmid R1, is an inhibitor of DNA replication acting at the initiation of DNA synthesis. J. Mol. Biol. 1995, 247, 568–577. [Google Scholar] [CrossRef]
- Agarwal, S.; Mishra, N.K.; Bhatnagar, S.; Bhatnagar, R. PemK toxin of Bacillus anthracis is a ribonuclease: An insight into its active site, structure, and function. J. Biol Chem. 2010, 285, 7254–7270. [Google Scholar] [CrossRef]
- Masuda, Y.; Miyakawa, K.; Nishimura, Y.; Ohtsubo, E. chpA and chpB, Escherichia coli chromosomal homologs of the pem locus responsible for stable maintenance of plasmid R100. J. Bacteriol. 1993, 175, 6850–6856. [Google Scholar] [CrossRef]
- Kamphuis, M.B.; Monti, M.C.; van den Heuvel, R.H.; Santos-Sierra, S.; Folkers, G.E.; Lemonnier, M.; Díaz-Orejas, R.; Heck, A.J.; Boelens, R. Interactions between the toxin Kid of the bacterial parD system and the antitoxins Kis and MazE. Proteins 2007, 67, 219–231. [Google Scholar] [CrossRef]
- Madl, T.; Van Melderen, L.; Mine, N.; Respondek, M.; Oberer, M.; Keller, W.; Khatai, L.; Zangger, K. Structural basis for nucleic acid and toxin recognition of the bacterial antitoxin CcdA. J. Mol. Biol. 2006, 364, 170–185. [Google Scholar] [CrossRef]
- Walling, R.L.; Butler, J.S. Structural Determinants for Antitoxin Identity and Insulation of Cross Talk between Homologous Toxin-Antitoxin Systems. J. Bacteriol. 2016, 198, 3287–3295. [Google Scholar] [CrossRef]
- Yang, M.; Gao, C.; Wang, Y.; Zhang, H.; He, Z.G. Characterization of the interaction and cross-regulation of three Mycobacterium tuberculosis RelBE modules. PLoS ONE 2010, 5, e10672. [Google Scholar] [CrossRef]
- Połom, D.; Boss, L.; Węgrzyn, G.; Hayes, F.; Kędzierska, B. Amino acid residues crucial for specificity of toxin-antitoxin interactions in the homologous Axe-Txe and YefM-YoeB complexes. FEBS J. 2013, 280, 5906–5918. [Google Scholar] [CrossRef] [PubMed]
- Nolle, N.; Schuster, C.F.; Bertram, R. Two paralogous yefM-yoeB loci from Staphylococcus equorum encode functional toxin-antitoxin systems. Microbiology 2013, 159, 1575–1585. [Google Scholar] [CrossRef]
- Ahidjo, B.A.; Kuhnert, D.; McKenzie, J.L.; Machowski, E.E.; Gordhan, B.G.; Arcus, V.; Abrahams, G.L.; Mizrahi, V. VapC toxins from Mycobacterium tuberculosis are ribonucleases that differentially inhibit growth and are neutralized by cognate VapB antitoxins. PLoS ONE 2011, 6, e21738. [Google Scholar] [CrossRef] [PubMed]
- Iqbal, N.; Guérout, A.M.; Krin, E.; Le Roux, F.; Mazel, D. Comprehensive Functional Analysis of the 18 Vibrio cholerae N16961 Toxin-Antitoxin Systems Substantiates Their Role in Stabilizing the Superintegron. J. Bacteriol. 2015, 197, 2150–2159. [Google Scholar] [CrossRef] [PubMed]
- Tandon, H.; Sharma, A.; Wadhwa, S.; Varadarajan, R.; Singh, R.; Srinivasan, N.; Sandhya, S. Bioinformatic and mutational studies of related toxin-antitoxin pairs in Mycobacterium tuberculosis predict and identify key functional residues. J. Biol. Chem. 2019, 294, 9048–9063. [Google Scholar] [CrossRef]
- Ramage, R.H.; Connolly, L.E.; Cox, J.S. Comprehensive functional analysis of Mycobacterium tuberculosis toxin-antitoxin systems: Implications for pathogenesis, stress responses, and evolution. PLoS Genet. 2009, 5, e1000767. [Google Scholar] [CrossRef]
- Jin, G.; Pavelka, M.S., Jr.; Butler, J.S. Structure-function analysis of VapB4 antitoxin identifies critical features of a minimal VapC4 toxin-binding module. J. Bacteriol. 2015, 197, 1197–1207. [Google Scholar] [CrossRef][Green Version]
- Gucinski, G.C.; Michalska, K.; Garza-Sánchez, F.; Eschenfeldt, W.H.; Stols, L.; Nguyen, J.Y.; Goulding, C.W.; Joachimiak, A.; Hayes, C.S. Convergent Evolution of the Barnase/EndoU/Colicin/RelE (BECR) Fold in Antibacterial tRNase Toxins. Structure 2019, 27, 1660–1674. [Google Scholar] [CrossRef]
- Goulard, C.; Langrand, S.; Carniel, E.; Chauvaux, S. The Yersinia pestis chromosome encodes active addiction toxins. J. Bacteriol. 2010, 192, 3669–3677. [Google Scholar] [CrossRef]
- Sierra, S.S.; Giraldo, R.; Orejas, R.D. Functional interactions between chpB and parD, two homologous conditional killer systems found in the Escherichia coli chromosome and in plasmid R1. FEMS Microbiol. Lett. 1998, 168, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Santos-Sierra, S.; Giraldo, R.; Diaz-Orejas, R. Functional interactions between homologous conditional killer systems of plasmid and chromosomal origin. FEMS Microbiol. Lett. 1997, 152, 51–56. [Google Scholar] [CrossRef]
- Zhu, L.; Sharp, J.D.; Kobayashi, H.; Woychik, N.A.; Inouye, M. Noncognate Mycobacterium tuberculosis toxin-antitoxins can physically and functionally interact. J. Biol. Chem. 2010, 285, 39732–39738. [Google Scholar] [CrossRef] [PubMed]
- Pinaud, L.; Ferrari, M.L.; Friedman, R.; Jehmlich, N.; von Bergen, M.; Phalipon, A.; Sansonetti, P.J.; Campbell-Valois, F.X. Identification of novel substrates of Shigella T3SA through analysis of its virulence plasmid-encoded secretome. PLoS ONE 2017, 12, e0186920. [Google Scholar] [CrossRef] [PubMed]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Burley, S.K.; Berman, H.M.; Bhikadiya, C.; Bi, C.; Chen, L.; Di Costanzo, L.; Christie, C.; Dalenberg, K.; Duarte, J.M.; Dutta, S.; et al. RCSB Protein Data Bank: Biological macromolecular structures enabling research and education in fundamental biology, biomedicine, biotechnology and energy. Nucleic Acids Res. 2019, 47, D464–D474. [Google Scholar] [CrossRef]
- Li, G.Y.; Zhang, Y.; Inouye, M.; Ikura, M. Inhibitory mechanism of Escherichia coli RelE-RelB toxin-antitoxin module involves a helix displacement near an mRNA interferase active site. J. Biol. Chem. 2009, 284, 14628–14636. [Google Scholar] [CrossRef]
- Takagi, H.; Kakuta, Y.; Okada, T.; Yao, M.; Tanaka, I.; Kimura, M. Crystal structure of archaeal toxin-antitoxin RelE-RelB complex with implications for toxin activity and antitoxin effects. Nat. Struct. Mol. Biol. 2005, 12, 327–331. [Google Scholar] [CrossRef]
- Francuski, D.; Saenger, W. Crystal structure of the antitoxin-toxin protein complex RelB-RelE from Methanococcus jannaschii. J. Mol. Biol. 2009, 393, 898–908. [Google Scholar] [CrossRef]
- Garcia-Pino, A.; Christensen-Dalsgaard, M.; Wyns, L.; Yarmolinsky, M.; Magnuson, R.D.; Gerdes, K.; Loris, R. Doc of prophage P1 is inhibited by its antitoxin partner Phd through fold complementation. J. Biol. Chem. 2008, 283, 30821–30827. [Google Scholar] [CrossRef]
- Kamada, K.; Hanaoka, F. Conformational change in the catalytic site of the ribonuclease YoeB toxin by YefM antitoxin. Mol. Cell 2005, 19, 497–509. [Google Scholar] [CrossRef] [PubMed]
- Miallau, L.; Jain, P.; Arbing, M.A.; Cascio, D.; Phan, T.; Ahn, C.J.; Chan, S.; Chernishof, I.; Maxson, M.; Chiang, J.; et al. Comparative proteomics identifies the cell-associated lethality of M. tuberculosis RelBE-like toxin-antitoxin complexes. Structure 2013, 21, 627–637. [Google Scholar] [PubMed]
- Schureck, M.A.; Maehigashi, T.; Miles, S.J.; Marquez, J.; Cho, S.E.; Erdman, R.; Dunham, C.M. Structure of the Proteus vulgaris HigB-(HigA)2-HigB toxin-antitoxin complex. J. Biol. Chem. 2014, 289, 1060–1070. [Google Scholar] [CrossRef]
- Hadži, S.; Garcia-Pino, A.; Haesaerts, S.; Jurėnas, D.; Gerdes, K.; Lah, J.; Loris, R. Ribosome-dependent Vibrio cholerae mRNAse HigB2 is regulated by a beta-strand sliding mechanism. Nucleic Acids Res. 2017, 45, 4972–4983. [Google Scholar] [CrossRef]
- Talavera, A.; Tamman, H.; Ainelo, A.; Konijnenberg, A.; Hadži, S.; Sobott, F.; Garcia-Pino, A.; Hõrak, R.; Loris, R. A dual role in regulation and toxicity for the disordered N-terminus of the toxin GraT. Nat. Commun. 2019, 10, 972. [Google Scholar] [CrossRef] [PubMed]
- Yoon, W.S.; Seok, S.H.; Won, H.S.; Cho, T.; Lee, S.J.; Seo, M.D. Structural changes of antitoxin HigA from Shigella flexneri by binding of its cognate toxin HigB. Int. J. Biol. Macromol. 2019, 130, 99–108. [Google Scholar] [CrossRef]
- Krissinel, E.; Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 2007, 372, 774–797. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera—A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- Russell, A.D.; Hatfull, G.F. PhagesDB: The actinobacteriophage database. Bioinformatics 2017, 33, 784–786. [Google Scholar] [CrossRef]
- Dienemann, C.; Bøggild, A.; Winther, K.S.; Gerdes, K.; Brodersen, D.E. Crystal structure of the VapBC toxin-antitoxin complex from Shigella flexneri reveals a hetero-octameric DNA-binding assembly. J. Mol. Biol. 2011, 414, 713–722. [Google Scholar] [CrossRef]
- Lee, I.G.; Lee, S.J.; Chae, S.; Lee, K.Y.; Kim, J.H.; Lee, B.J. Structural and functional studies of the Mycobacterium tuberculosis VapBC30 toxin-antitoxin system: Implications for the design of novel antimicrobial peptides. Nucleic Acids Res. 2015, 43, 7624–7637. [Google Scholar] [CrossRef] [PubMed]
- Korch, B.S.; Contreras, H.; Clark-Curtiss, J.E. Three Mycobacterium tuberculosis Rel toxin-antitoxin modules inhibit mycobacterial growth and are expressed in infected human macrophages. J. Bacteriol. 2009, 191, 1618–1630. [Google Scholar] [CrossRef] [PubMed]
| TA System | Plasmid | Host Bacteria | Chromosomal Homolog/Xenolog | Citations |
|---|---|---|---|---|
| ParDE | RK2 */RP4 * | (Enterobacteriaceae) Escherichia coli | ParDE | [55,79,129] |
| RelBE | P307, IncB/C | E. coli | RelBE | [130,131] |
| VapBC | IncF | E. coli | [131] | |
| StbDE | R485 | Morganella morganii | [132] | |
| CcdAB, Kid/Kis (PemKI) | F, R1/R100, pCHP91 | E. coli, Erwinia chrysanthemi, Staphylococcus | MazEF | [118,133,134,135,136] |
| PhD-Doc | Bacteriophage P1 | E. coli | [137] | |
| HigBA | pRTS1 | Proteus vulgaris | [138] | |
| Tad-ata | pAMI2 | Paracoccus aminophilus | [139] | |
| Axe-txe | pRUM | Enterococcus faecalis | YoeB-YefM | [140,141] |
| TasA-TasB | pGI1 | Bacillus thuringiensis | [125] | |
| VapBC | pMYSH6000 | Shigella flexneri | [142] | |
| AtaRT | pB171-like plasmid | E.coli | AtaRT | [143] |
| Omega-epsilon-zeta | pSM19035 *, pVER1/2 | Streptococcus pyogenes. Enterococcus faecium | PezAT | [141,144] |
| RelBE, HigBA, HTH/GNAT, SplTA | p3ABAYE | Acinetobacter baumannii | [145] | |
| YacAB | PWR100 | E. coli | [146] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tu, C.-H.; Holt, M.; Ruan, S.; Bourne, C. Evaluating the Potential for Cross-Interactions of Antitoxins in Type II TA Systems. Toxins 2020, 12, 422. https://doi.org/10.3390/toxins12060422
Tu C-H, Holt M, Ruan S, Bourne C. Evaluating the Potential for Cross-Interactions of Antitoxins in Type II TA Systems. Toxins. 2020; 12(6):422. https://doi.org/10.3390/toxins12060422
Chicago/Turabian StyleTu, Chih-Han, Michelle Holt, Shengfeng Ruan, and Christina Bourne. 2020. "Evaluating the Potential for Cross-Interactions of Antitoxins in Type II TA Systems" Toxins 12, no. 6: 422. https://doi.org/10.3390/toxins12060422
APA StyleTu, C.-H., Holt, M., Ruan, S., & Bourne, C. (2020). Evaluating the Potential for Cross-Interactions of Antitoxins in Type II TA Systems. Toxins, 12(6), 422. https://doi.org/10.3390/toxins12060422








