Comparative Sequence Analysis of TRI1 of Fusarium
Abstract
1. Introduction
2. Results and Discussion
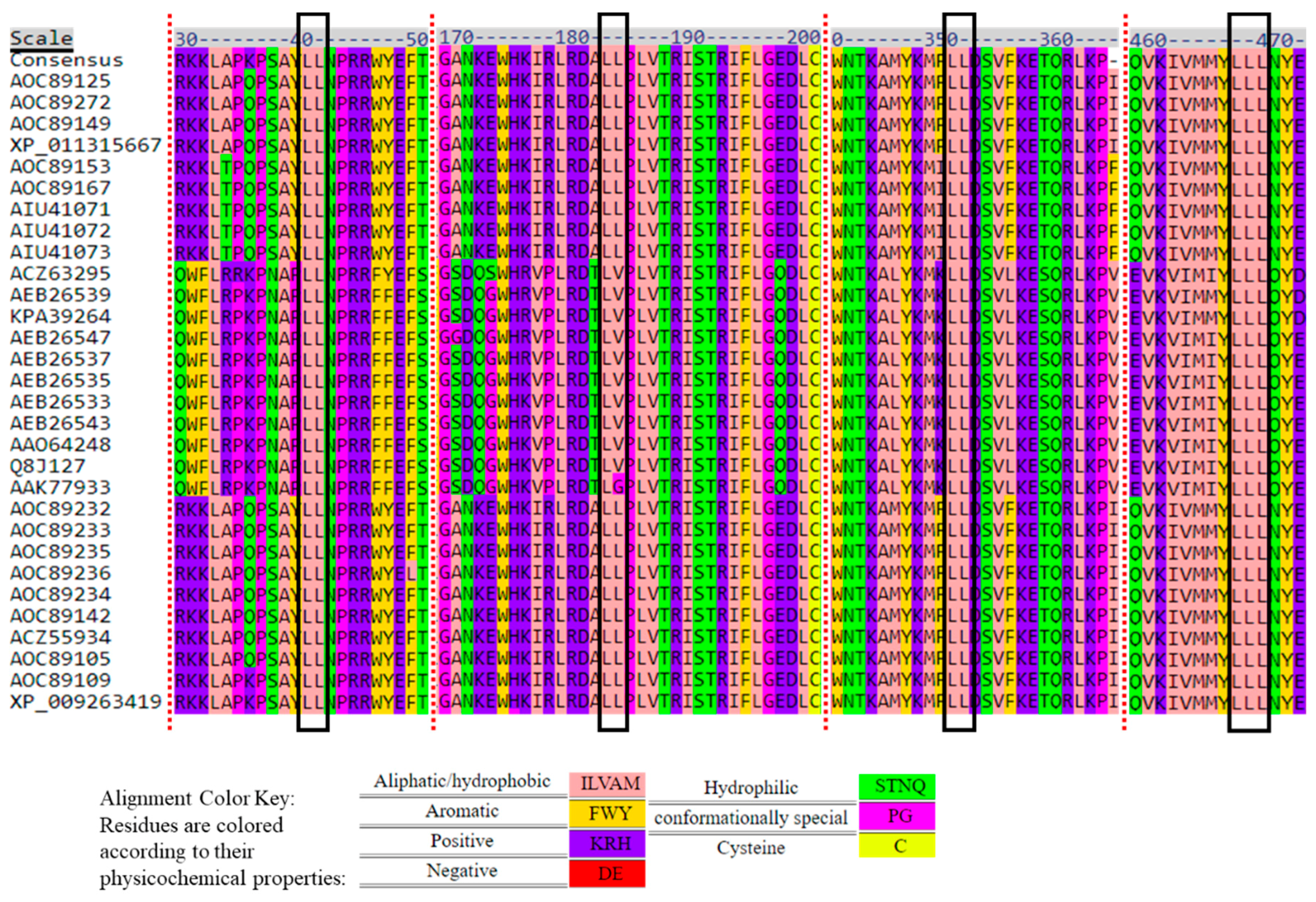
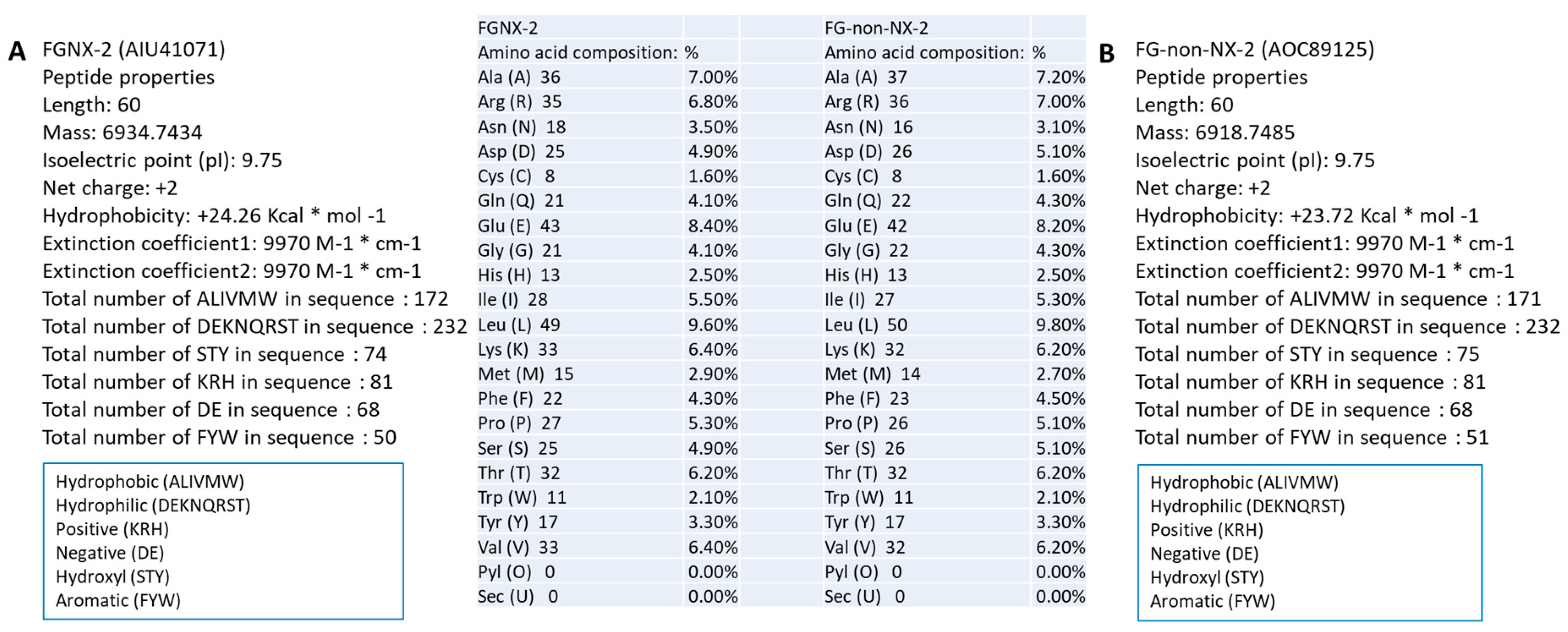
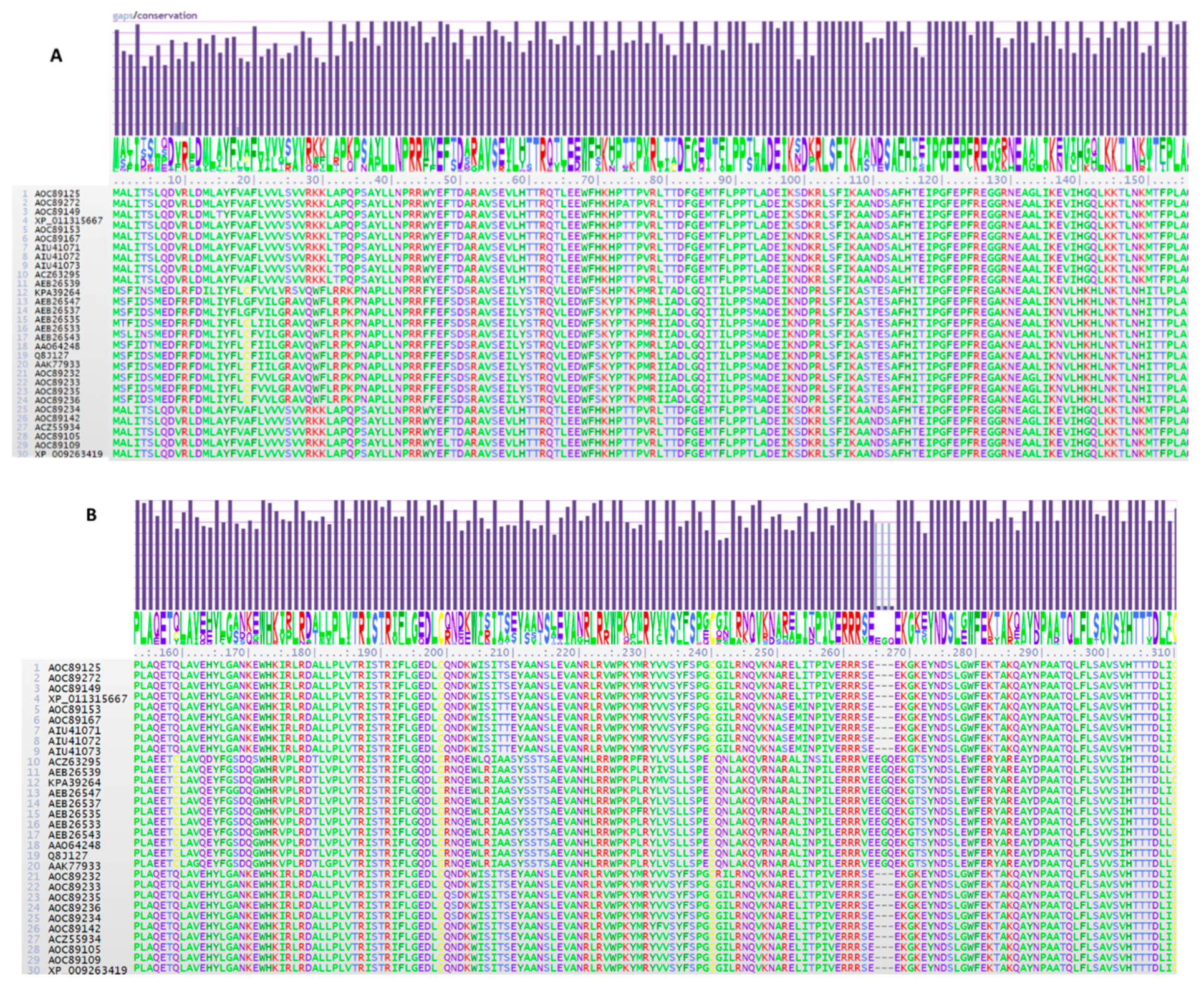
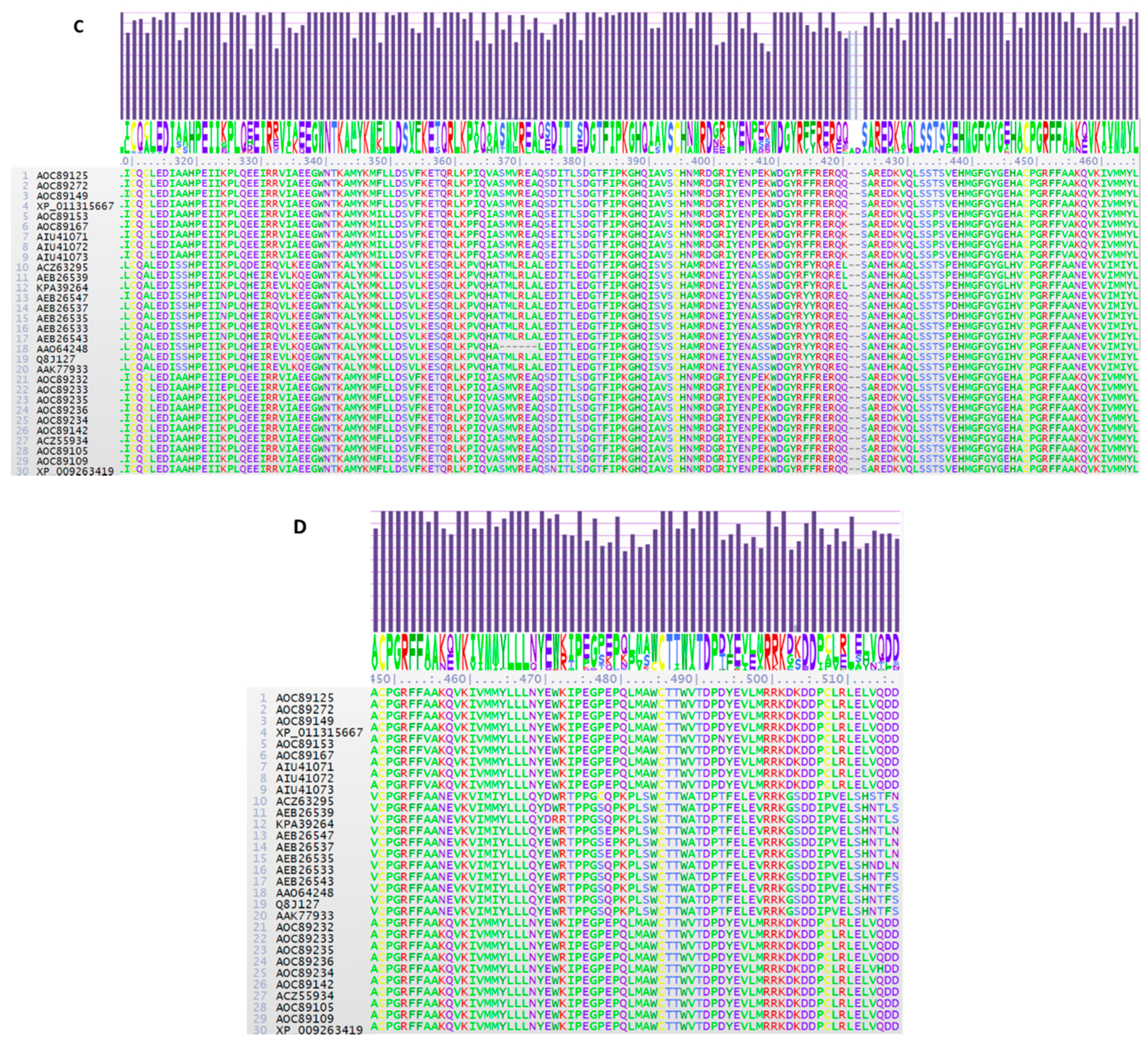
2.1. TRI1 Peptide Primary Sequence Comparisons
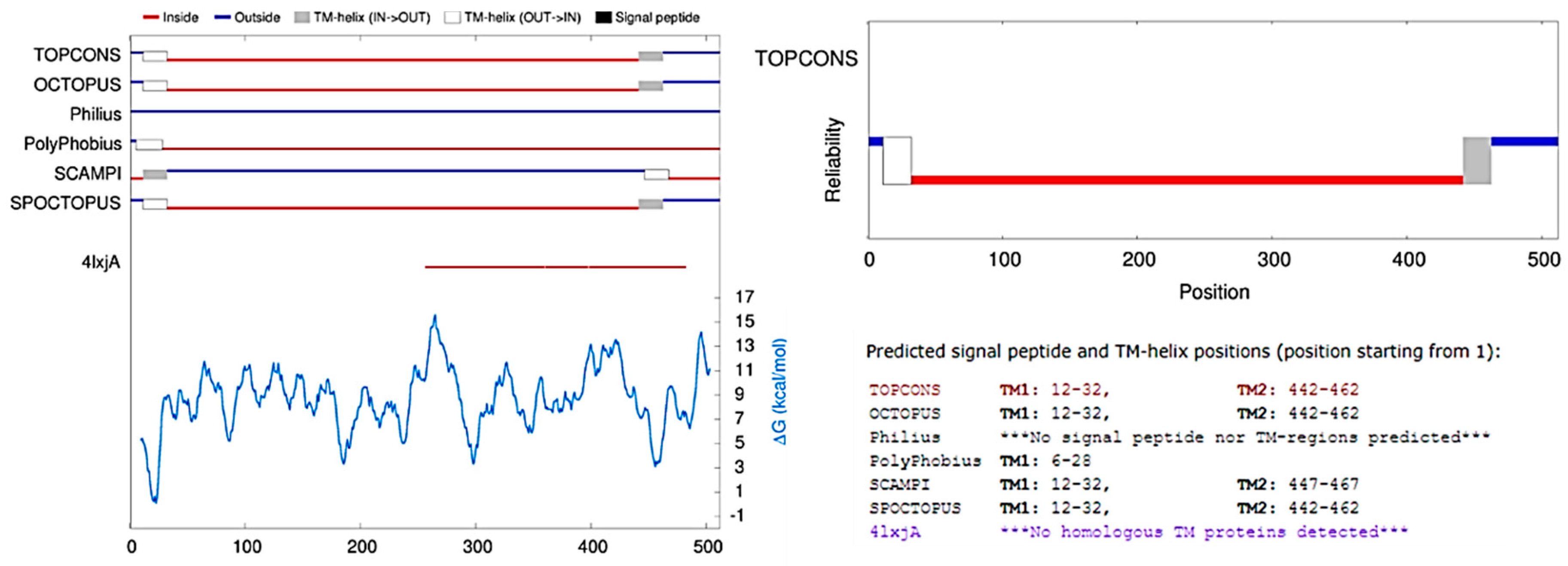
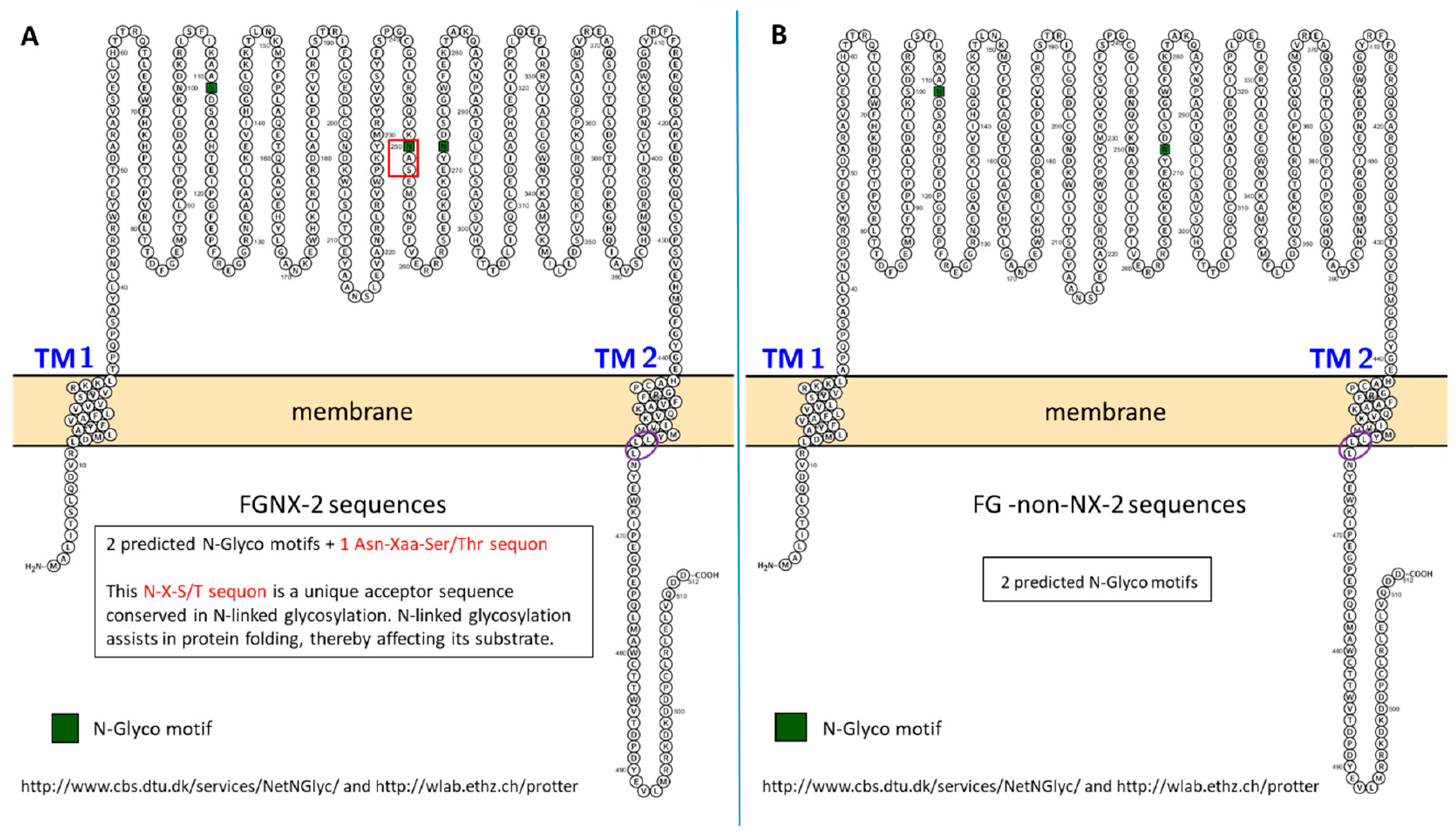
2.2. Topology
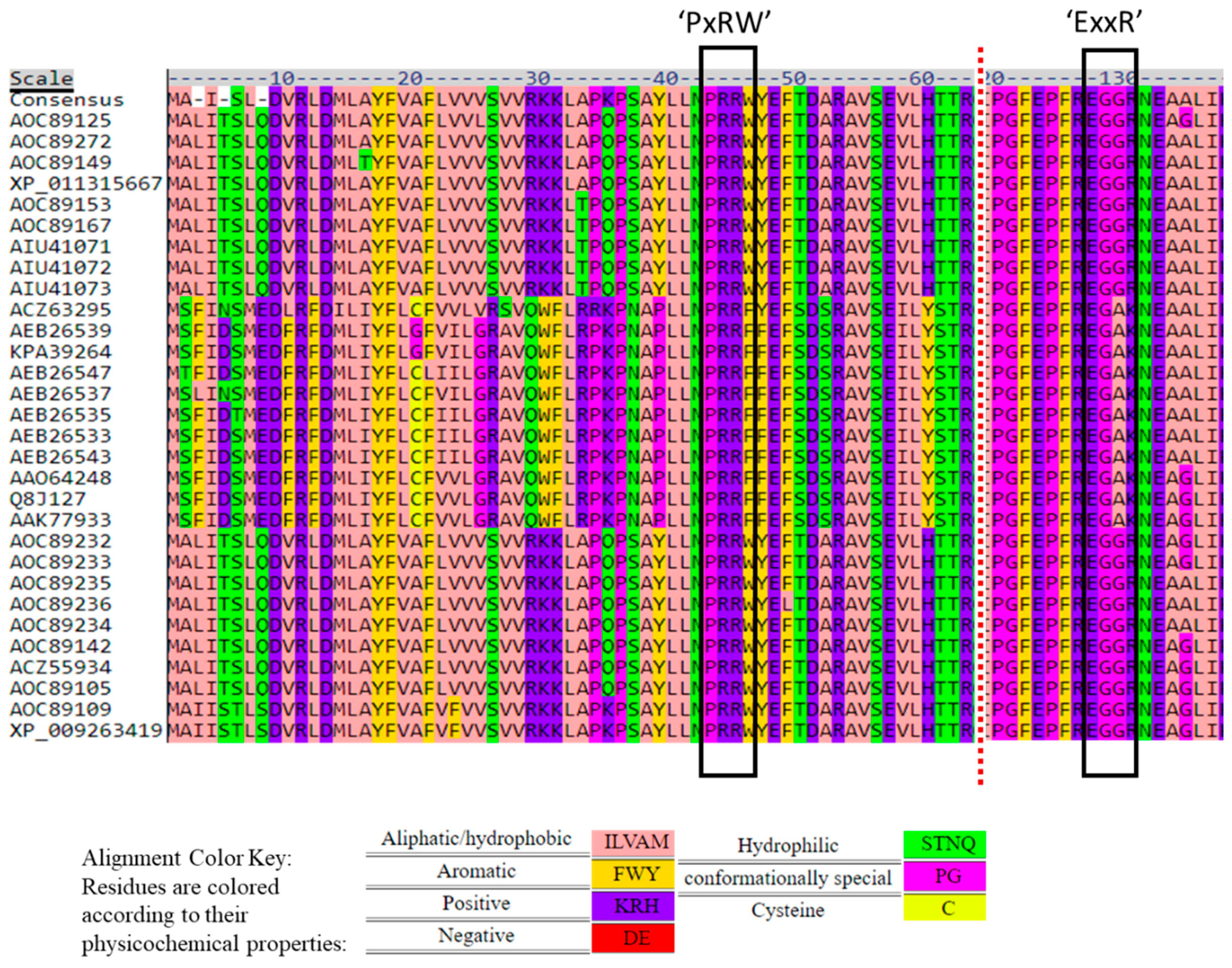
2.3. PER/PxRW and ExxR Motifs
2.4. Dileucine (LL) Motifs
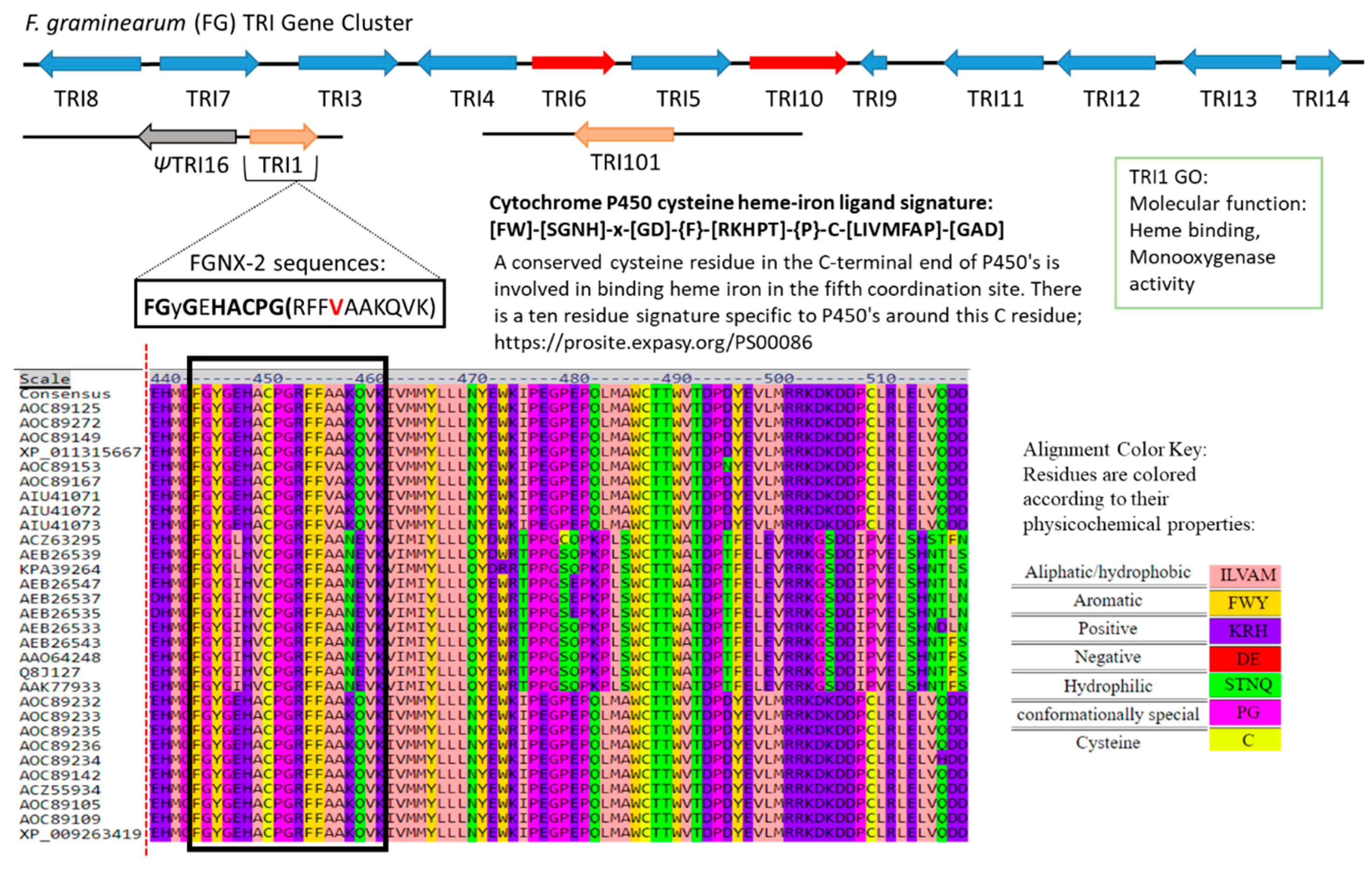
2.5. Heme Motif
2.6. TRI1 DNA Polymorphism Profile and Evidence of Selection
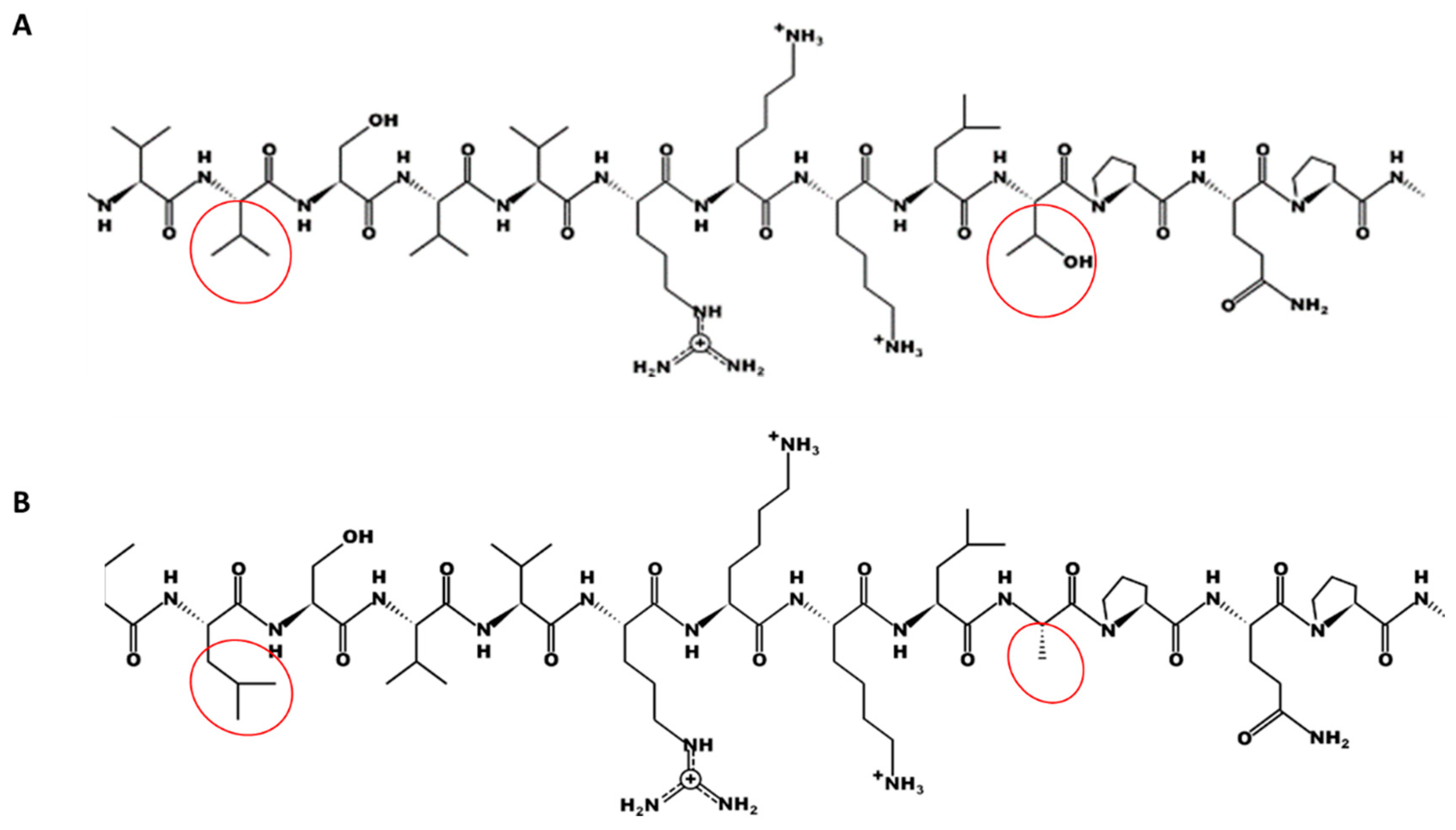
2.7. TRI1p Residue Substitutions
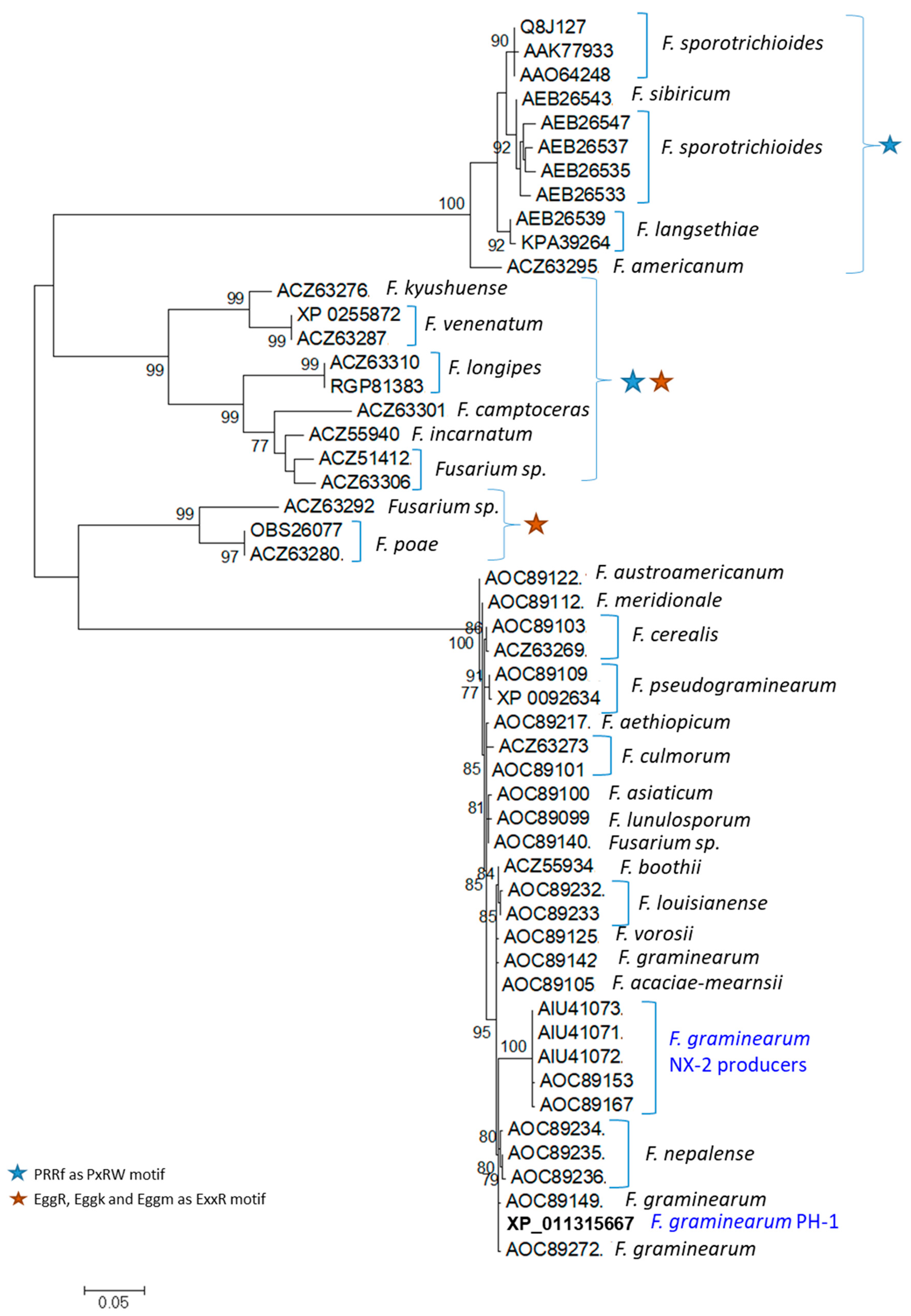
2.8. Phylogeny of TRI1p Sequences of Fusarium
2.9. N-Linked Glycosylation
3. Conclusions
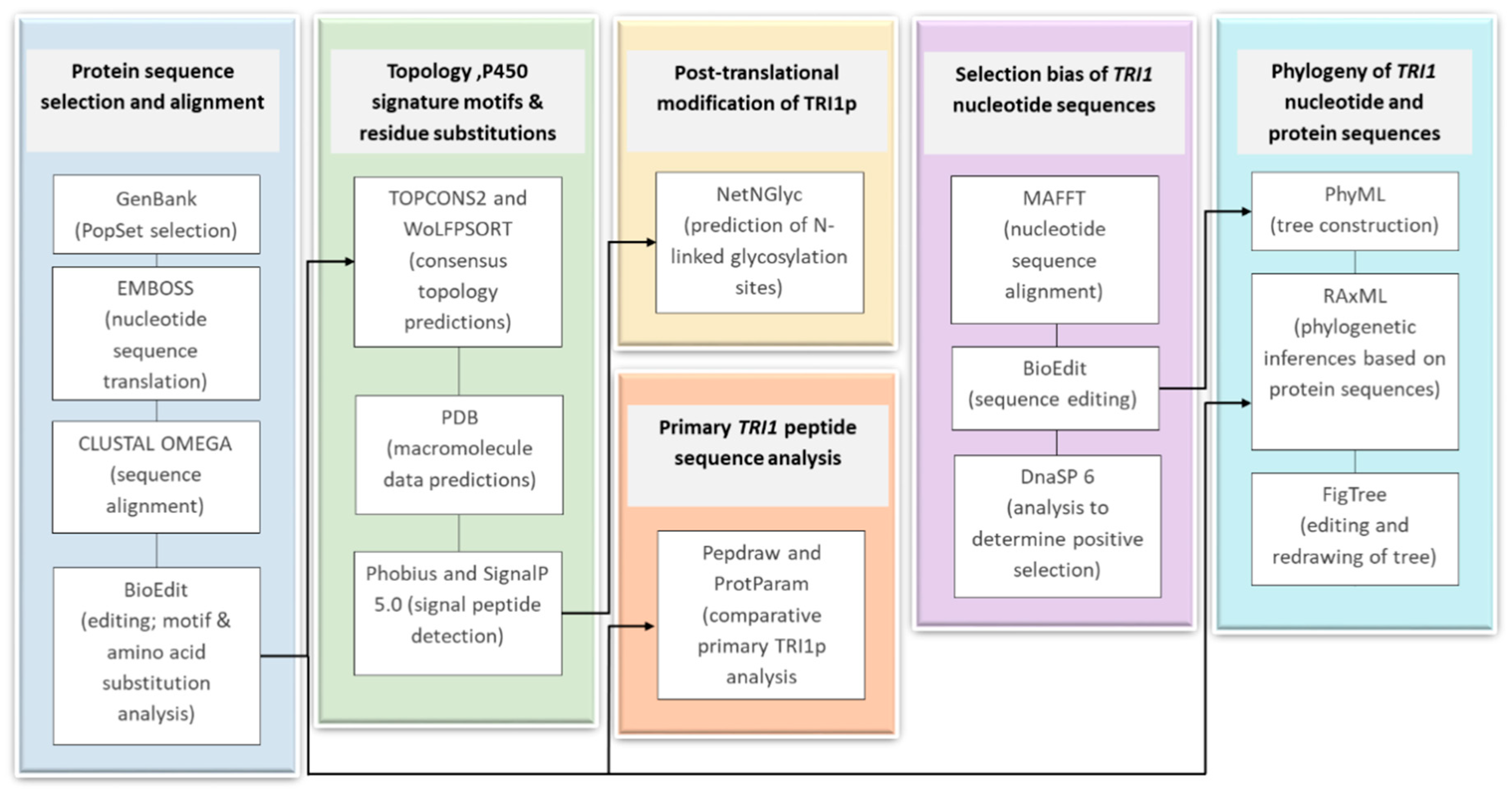
4. Materials and Methods
4.1. Protein Sequence Selection and Alignment
4.2. Comparative Primary Peptide Sequence Analysis
4.3. Topology Analysis
4.4. Signature Motifs of P450s
4.5. Selection Bias
4.6. Phylogeny of TRI1 Nucleotide and Protein Sequences
4.7. Post-Translational Modification of TRI1p
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Desjardins, A.E.; Hohn, T.M.; McCormick, S.P. Effect of gene disruption of trichodiene synthase on the virulence of Gibberella pulicaris. Mol. Plant Microbe Interact. 1992, 5, 214–222. [Google Scholar] [CrossRef]
- Desjardins, A.E.; Proctor, R.H.; McCormick, S.P.; Hohn, T.M. Reduced Virulence of Trichothecene Antibiotic-Nonproducing Mutants of Gibberella zeae in Wheat Field Tests. In Proceedings of the Fusarium Head Scab: Global Status and Future Prospects, El Batan, Mexico, 13–17 October 1996; Dubin, H.J., Gilchrist, L., McNab, A., Eds.; BCIMMYT: El Batan, Mexico, 1996. [Google Scholar]
- Cuzick, A.; Urban, M.; Hammond-Kosack, K. Fusarium graminearum gene deletion mutants map1 and tri5 reveal similarities and differences in the pathogenicity requirements to cause disease on Arabidopsis and wheat floral tissue. New Phytol. 2008, 177, 990–1000. [Google Scholar] [CrossRef] [PubMed]
- Proctor, R.H.; McCormick, S.P.; Kim, H.S.; Cardoza, R.E.; Stanley, A.M.; Lindo, L.; Kelly, A.; Brown, D.W.; Lee, T.; Vaughan, M.M.; et al. Evolution of structural diversity of trichothecenes, a family of toxins produced by plant pathogenic and entomopathogenic fungi. PLoS Pathog. 2018, 14, e1006946. [Google Scholar] [CrossRef] [PubMed]
- McCormick, S.P.; Stanley, A.M.; Stover, N.A.; Alexander, N.J. Trichothecenes: From simple to complex mycotoxins. Toxins 2011, 3, 802–814. [Google Scholar] [CrossRef]
- Cundliffe, E.; Davies, J.E. Inhibition of initiation, elongation, and termination of eukaryotic protein synthesis by trichothecene fungal toxins. Antimicrob. Agents Chemother. 1977, 11, 491–499. [Google Scholar] [CrossRef]
- Sudakin, D.L. Trichothecenes in the environment: Relevance to human health. Toxicol. Lett. 2003, 143, 97–107. [Google Scholar] [CrossRef]
- Pestka, J.J. Deoxynivalenol: Mechanisms of action, human exposure, and toxicological relevance. Arch. Toxicol. 2010, 84, 663–679. [Google Scholar] [CrossRef]
- Alexander, N.J.; Proctor, R.H.; McCormick, S.P. Genes, gene clusters, and biosynthesis of trichothecenes and fumonisins in Fusarium. Toxin Rev. 2009, 28, 198–215. [Google Scholar] [CrossRef]
- McCormick, S.P.; Harris, L.J.; Alexander, N.J.; Ouellet, T.; Saparno, A.; Allard, S.; Desjardins, A.E. Tri1 in Fusarium graminearum encodes a P450 oxygenase. Appl. Environ. Microbiol. 2004, 70, 2044–2051. [Google Scholar] [CrossRef]
- McCormick, S.P.; Alexander, N.J.; Proctor, R.H. Fusarium TRI4 encodes a multifunctional oxygenase required for trichothecene biosynthesis. Can. J. Microbiol. 2006, 52, 636–642. [Google Scholar] [CrossRef]
- Alexander, N.J.; McCormick, S.P.; Waalwijk, C.; van der Lee, T.; Proctor, R.H. The genetic basis for 3-ADON and 15-ADON trichothecene chemotypes in Fusarium. Fungal Genet. Biol. 2011, 48, 485–495. [Google Scholar] [CrossRef] [PubMed]
- Caspi, R.; Billington, R.; Fulcher, C.A.; Keseler, I.M.; Kothari, A.; Krummenacker, M.; Latendresse, M.; Midford, P.E.; Ong, Q.; Ong, W.K.; et al. The MetaCyc database of metabolic pathways and enzymes. Nucleic Acids Res. 2017, 46, D633–D639. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M.; Tokai, T.; Takahashi-Ando, N.; Ohsato, S.; Fujimura, M. Molecular and genetic studies of Fusarium trichothecene biosynthesis: Pathways, genes, and evolution. Biosci. Biotechnol. Biochem. 2007, 71, 2105–2123. [Google Scholar] [CrossRef]
- Van der Lee, T.; Zhang, H.; van Diepeningen, A.; Waalwijk, C. Biogeography of Fusarium graminearum species complex and chemotypes: A review. Food Addit. Contam. Part A 2015, 32, 453–460. [Google Scholar] [CrossRef]
- Ward, T.J.; Clear, R.M.; Rooney, A.P.; O’Donnell, K.; Gaba, D.; Patrick, S.; Starkey, D.E.; Gilbert, J.; Geiser, D.M.; Nowicki, T.W. An adaptive evolutionary shift in Fusarium head blight pathogen populations is driving the rapid spread of more toxigenic Fusarium graminearum in North America. Fungal Genet. Biol. 2008, 45, 473–484. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhang, Z.; Van der Lee, T.; Chen, W.Q.; Xu, J.; Xu, J.S.; Yang, L.; Yu, D.; Waalwijk, C.; Feng, J. Population genetic analyses of Fusarium asiaticum populations from barley suggest a recent shift favoring 3ADON producers in southern China. Phytopathology 2010, 100, 328–336. [Google Scholar] [CrossRef] [PubMed]
- Gale, L.R.; Harrison, S.A.; Ward, T.J.; O’Donnell, K.; Milus, E.A.; Gale, S.W.; Kistler, H.C. Nivalenol-type populations of Fusarium graminearum and F. asiaticum are prevalent on wheat in southern Louisiana. Phytopathology 2011, 101, 124–134. [Google Scholar] [CrossRef] [PubMed]
- Gomes, L.B.; Ward, T.J.; Badiale-Furlong, E.; Del Ponte, E.M. Species composition, toxigenic potential and pathogenicity of Fusarium graminearum species complex isolates from southern Brazilian rice. Plant Pathol. 2015, 64, 980–987. [Google Scholar] [CrossRef]
- Liang, J.M.; Xayamongkhon, H.; Broz, K.; Dong, Y.; McCormick, S.P.; Abramova, S.; Ward, T.J.; Ma, Z.H.; Kistler, H.C. Temporal dynamics and population genetic structure of Fusarium graminearum in the upper midwestern United States. Fungal Genet. Biol. 2014, 73, 83–92. [Google Scholar] [CrossRef]
- Kelly, A.C.; Clear, R.M.; O’Donnell, K.; McCormick, S.; Turkington, T.K.; Tekauz, A.; Gilbert, J.; Kistler, H.C.; Busman, M.; Ward, T.J. Diversity of Fusarium head blight populations and trichothecene toxin types reveals regional differences in pathogen composition and temporal dynamics. Fungal Genet. Biol. 2015, 82, 22–31. [Google Scholar] [CrossRef]
- Varga, E.; Wiesenberger, G.; Hametner, C.; Ward, T.J.; Dong, Y.; Schöfbeck, D.; McCormick, S.; Broz, K.; Stückler, R.; Schuhmacher, R.; et al. New tricks of an old enemy: Isolates of Fusarium graminearum produce a type A trichothecene mycotoxin. Environ. Microbiol. 2015, 17, 2588–2600. [Google Scholar] [CrossRef] [PubMed]
- Kimura, Y.; Hisano, Y.; Kawahara, A.; Higashijima, S.I. Efficient generation of knock-in transgenic zebrafish carrying reporter/driver genes by CRISPR/Cas9-mediated genome engineering. Sci. Rep. 2014, 4, 6545. [Google Scholar] [CrossRef] [PubMed]
- Lofgren, L.; Riddle, J.; Dong, Y.; Kuhnem, P.R.; Cummings, J.A.; Del Ponte, E.M.; Bergstrom, G.C.; Kistler, H.C. A high proportion of NX-2 genotype strains are found among Fusarium graminearum isolates from north-eastern New York State. Eur. J. Plant Pathol. 2018, 150, 791–796. [Google Scholar] [CrossRef]
- Menke, J.; Weber, J.; Broz, K.; Kistler, H.C. Cellular development associated with induced mycotoxin synthesis in the filamentous fungus Fusarium graminearum. PLoS ONE 2013, 8, e63077. [Google Scholar] [CrossRef]
- Kistler, H.C.; Broz, K. Cellular compartmentalization of secondary metabolism. Front. Microbiol. 2015, 6, 68. [Google Scholar] [CrossRef]
- Boenisch, M.J.; Broz, K.L.; Purvine, S.O.; Chrisler, W.B.; Nicora, C.D.; Connolly, L.R.; Freitag, M.; Baker, S.E.; Kistler, H.C. Structural reorganization of the fungal endoplasmic reticulum upon induction of mycotoxin biosynthesis. Sci. Rep. 2017, 7, 44296. [Google Scholar] [CrossRef]
- Hernández-Martínez, F.; Briones-Roblero, C.I.; Nelson, D.R.; Rivera-Orduña, F.N.; Zúñiga, G. Cytochrome P450 complement (CYPome) of Candida oregonensis, a gut-associated yeast of bark beetle, Dendroctonus rhizophagus. Fungal Biol. 2016, 120, 1077–1089. [Google Scholar] [CrossRef]
- Córdova, P.; Gonzalez, A.M.; Nelson, D.R.; Gutiérrez, M.S.; Baeza, M.; Cifuentes, V.; Alcaíno, J. Characterization of the cytochrome P450 monooxygenase genes (P450ome) from the carotenogenic yeast Xanthophyllomyces dendrorhous. BMC Genom. 2017, 18, 540. [Google Scholar] [CrossRef]
- Chen, W.; Lee, M.K.; Jefcoate, C.; Kim, S.C.; Chen, F.; Yu, J.H. Fungal cytochrome p450 monooxygenases: Their distribution, structure, functions, family expansion, and evolutionary origin. Genome Biol. Evol. 2014, 6, 1620–1634. [Google Scholar] [CrossRef]
- Sello, M.M.; Jafta, N.; Nelson, D.R.; Chen, W.; Yu, J.H.; Parvez, M.; Kgosiemang, I.K.R.; Monyaki, R.; Raselemane, S.C.; Qhanya, L.B.; et al. Diversity and evolution of cytochrome P450 monooxygenases in Oomycetes. Sci. Rep. 2015, 5, 11572. [Google Scholar] [CrossRef]
- Sezutsu, H.; Le Goff, G.; Feyereisen, R. Origins of P450 diversity. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2013, 368, 20120428. [Google Scholar] [CrossRef] [PubMed]
- Pandey, K.N. Functional roles of short sequence motifs in the endocytosis of membrane receptors. Front. Biosci. 2009, 14, 5339–5360. [Google Scholar] [CrossRef] [PubMed]
- Bonifacino, J.S.; Traub, L.M. Signals for sorting of transmembrane proteins to endosomes and lysosomes. Annu. Rev. Biochem. 2003, 72, 395–447. [Google Scholar] [CrossRef] [PubMed]
- Kelly, D.E.; Kraševec, N.; Mullins, J.; Nelson, D.R. The CYPome (cytochrome P450 complement) of Aspergillus nidulans. Fungal Genet. Biol. 2009, 46, S53–S61. [Google Scholar] [CrossRef]
- Guengerich, F.P.; Waterman, M.R.; Egli, M. Recent structural insights into cytochrome P450 function. Trends Pharmacol. Sci. 2016, 37, 625–640. [Google Scholar] [CrossRef]
- EMBL-EBI. Available online: https://www.ebi.ac.uk/ (accessed on 18 October 2019).
- Danielson, P.B. The cytochrome P450 superfamily: Biochemistry, evolution and drug metabolism in humans. Curr. Drug Metab. 2002, 3, 561–597. [Google Scholar] [CrossRef]
- Yasui, H.; Hayashi, S.; Sakurai, H. Possible involvement of singlet oxygen species as multiple oxidants in P450 catalytic reactions. Drug Metab. Pharmacokinet. 2005, 20, 1–13. [Google Scholar] [CrossRef]
- Ortiz de Montellano, P.R. Hydrocarbon hydroxylation by cytochrome P450 enzymes. Chem. Rev. 2010, 110, 932–948. [Google Scholar] [CrossRef]
- Munro, A.W.; Girvan, H.M.; McLean, K.J. Variations on a (t) heme-novel mechanisms, redox partners and catalytic functions in the cytochrome P450 superfamily. Nat. Prod. Rep. 2007, 24, 585–609. [Google Scholar] [CrossRef]
- Syed, K.; Mashele, S.S. Comparative analysis of P450 signature motifs EXXR and CXG in the large and diverse kingdom of fungi: Identification of evolutionarily conserved amino acid patterns characteristic of P450 family. PLoS ONE 2014, 9, e95616. [Google Scholar] [CrossRef]
- Hargrove, T.Y.; Wawrzak, Z.; Liu, J.; Waterman, M.R.; Nes, W.D.; Lepesheva, G.I. Structural complex of sterol 14α-demethylase (CYP51) with 14α-methylenecyclopropyl-Δ7-24, 25-dihydrolanosterol. J. Lipid Res. 2012, 53, 311–320. [Google Scholar] [CrossRef] [PubMed]
- Johnston, J.B.; Ouellet, H.; Podust, L.M.; Ortiz de Montellano, P.R. Structural control of cytochrome P450-catalyzed ω-hydroxylation. Arch. Biochem. Biophys. 2011, 507, 86–94. [Google Scholar] [CrossRef] [PubMed]
- Shoji, O.; Kunimatsu, T.; Kawakami, N.; Watanabe, Y. Highly selective hydroxylation of benzene to phenol by wild-type cytochrome P450BM3 assisted by decoy molecules. Angew. Chem. 2013, 52, 6606–6610. [Google Scholar] [CrossRef] [PubMed]
- Neuman, R.C., Jr. Substituent effects. In Organic Chemistry; University of California: Riverside, CA, USA, 2013; pp. 1–35. [Google Scholar]
- Maleczka, R.E.; Shi, F.; Holmes, D.; Smith, M.R. C-H activation/borylation/oxidation: A one-pot unified route to meta-substituted phenols bearing ortho-/para-directing groups. J. Am. Chem. Soc. 2003, 125, 7792–7793. [Google Scholar] [CrossRef] [PubMed]
- Chotana, G.A.; Rak, M.A.; Smith, M.R. Sterically directed functionalization of aromatic C-H bonds: Selective borylation ortho to cyano groups in arenes and heterocycles. J. Am. Chem. Soc. 2005, 127, 10539–10544. [Google Scholar] [CrossRef] [PubMed]
- Kelly, A.; Proctor, R.H.; Belzile, F.; Chulze, S.N.; Clear, R.M.; Cowger, C.; Elmer, W.; Lee, T.; Obanor, F.; Waalwijk, C.; et al. The geographic distribution and complex evolutionary history of the NX-2 trichothecene chemotype from Fusarium graminearum. Fungal Genet. Biol. 2016, 95, 39–48. [Google Scholar] [CrossRef]
- Kelly, A.C.; Ward, T.J. Population genomics of Fusarium graminearum reveals signatures of divergent evolution within a major cereal pathogen. PLoS ONE 2018, 13, e0194616. [Google Scholar] [CrossRef]
- Jung, S.T.; Lauchli, R.; Arnold, F.H. Cytochrome P450: Taming a wild type enzyme. Curr. Opin. Biotechnol. 2011, 22, 809–817. [Google Scholar] [CrossRef]
- Aebi, M. N-linked protein glycosylation in the ER. Biochim. Biophys. Acta Mol. Cell Res. 2013, 1833, 2430–2437. [Google Scholar] [CrossRef]
- Ortiz de Montellano, P.R. Cytochrome P450: Structure, Mechanism, and Biochemistry, 3rd ed.; Springer: New York, NY, USA, 2005; p. 463. [Google Scholar]
- Schalk, M.; Croteau, R. A single amino acid substitution (F363I) converts the regiochemistry of the spearmint (−)-limonene hydroxylase from a C6-to a C3-hydroxylase. Proc. Natl. Acad. Sci. USA 2000, 97, 11948–11953. [Google Scholar] [CrossRef]
- Meek, I.B.; Peplow, A.W.; Ake, C., Jr.; Phillips, T.D.; Beremand, M.N. TRI1 encodes the cytochrome P450 monooxygenase for C-8 hydroxylation during trichothecene biosynthesis in Fusarium sporotrichioides and resides upstream of another new TRI gene. Appl. Environ. Microbiol. 2003, 69, 1607–1613. [Google Scholar] [CrossRef] [PubMed]
- Zamir, L.O.; Devor, K.A.; Sauriol, F. Biosynthesis of the trichothecene 3-acetyldeoxynivalenol. Identification of the oxygenation steps after isotrichodermin. J. Biol. Chem. 1991, 266, 14992–15000. [Google Scholar]
- Romero, P.A.; Arnold, F.H. Exploring protein fitness landscapes by directed evolution. Nat. Rev. Mol. Cell Biol. 2009, 10, 866. [Google Scholar] [CrossRef] [PubMed]
- Karlovsky, P. Biological detoxification of the mycotoxin deoxynivalenol and its use in genetically engineered crops and feed additives. Appl. Microbiol. Biotechnol. 2011, 91, 491–504. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Tan, Y.; Liu, N.; Yan, Z.; Liao, Y.; Chen, J.; De Saeger, S.; Yang, H.; Zhang, Q.; Wu, A. Detoxification of deoxynivalenol via glycosylation represents novel insights on antagonistic activities of Trichoderma when confronted with Fusarium graminearum. Toxins 2016, 8, 335. [Google Scholar] [CrossRef]
- Varga, E.; Wiesenberger, G.; Woelflingseder, L.; Twaruschek, K.; Hametner, C.; Vaclaviková, M.; Malachová, A.; Marko, D.; Berthiller, F.; Adam, G. Less-toxic rearrangement products of NX-toxins are formed during storage and food processing. Toxicol. Lett. 2018, 284, 205–212. [Google Scholar] [CrossRef]
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein identification and analysis tools on the ExPASy server. In The Proteomics Protocols Handbook, 1st ed.; Walker, J.M., Ed.; Humana Press: Totowa, NJ, USA, 2005; pp. 571–607. [Google Scholar]
- Tsirigos, K.D.; Peters, C.; Shu, N.; Käll, L.; Elofsson, A. The TOPCONS web server for combined membrane protein topology and signal peptide prediction. Nucleic Acids Res. 2015, 43, W401–W407. [Google Scholar] [CrossRef]
- Käll, L.; Krogh, A.; Sonnhammer, E.L. An HMM posterior decoder for sequence feature prediction that includes homology information. Bioinformatics 2005, 21, i251–i257. [Google Scholar] [CrossRef]
- Nielsen, H.; Tsirigos, K.D.; Brunak, S.; von Heijne, G. A brief history of protein sorting prediction. Protein J. 2019, 38, 1–17. [Google Scholar] [CrossRef]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Gracia, A. DnaSP 6: DNA sequence polymorphism analysis of large datasets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef]
- Kryazhimskiy, S.; Plotkin, J.B. The population genetics of dN/dS. PLoS Genet. 2008, 4, e1000304. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.X.; Li, W.H. Statistical tests of neutrality of mutations. Genetics 1993, 133, 693–709. [Google Scholar] [PubMed]
- Ramírez-Soriano, A.; Ramos-Onsins, S.E.; Rozas, J.; Calafell, F.; Navarro, A. Statistical power analysis of neutrality tests under demographic expansions, contractions and bottlenecks with recombination. Genetics 2008, 179, 555–567. [Google Scholar] [CrossRef] [PubMed]
- Rokas, A. Phylogenetic analysis of protein sequence data using the randomized axelerated maximum likelihood (RAXML) program. Curr. Protoc. Mol. Biol. 2011, 96, 1–14. [Google Scholar] [CrossRef] [PubMed]

| DNA Polymorphism Parameter | FGNX-2 | FG-Non-NX-2 | 52 Fusarium Sequences |
|---|---|---|---|
| N | 5 | 4 | 52 |
| h | 3 | 2 | 46 |
| Hd | 0.70000 | 1.00000 | 0.99500 |
| Pi | 0.00052 | 0.00065 | 0.30227 |
| k | 0.80000 | 1.00000 | 463.382 |
| Ct | 0.14 | 0.14 | 0.14 |
| C | 0.049 | 0.049 | 0.049 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramdass, A.C.; Villafana, R.T.; Rampersad, S.N. Comparative Sequence Analysis of TRI1 of Fusarium. Toxins 2019, 11, 689. https://doi.org/10.3390/toxins11120689
Ramdass AC, Villafana RT, Rampersad SN. Comparative Sequence Analysis of TRI1 of Fusarium. Toxins. 2019; 11(12):689. https://doi.org/10.3390/toxins11120689
Chicago/Turabian StyleRamdass, Amanda C., Ria T. Villafana, and Sephra N. Rampersad. 2019. "Comparative Sequence Analysis of TRI1 of Fusarium" Toxins 11, no. 12: 689. https://doi.org/10.3390/toxins11120689
APA StyleRamdass, A. C., Villafana, R. T., & Rampersad, S. N. (2019). Comparative Sequence Analysis of TRI1 of Fusarium. Toxins, 11(12), 689. https://doi.org/10.3390/toxins11120689





