Alcohol Consumption Is Associated with Poor Prognosis in Obese Patients with COVID-19: A Mendelian Randomization Study Using UK Biobank
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population from UK Biobank
2.2. Exposure of Interest
2.3. Genetic Data
2.4. Other Potential Confounding Risk Factors for COVID-19
2.5. Ascertainment of Outcomes
2.6. Statistical Analysis
2.7. Sensitivity Analysis
2.8. Mediation Analysis
3. Results
3.1. Characteristics of Participants
3.2. Instrumental Variable Associations
3.3. Observational Associations and Instrumental Variable Associations
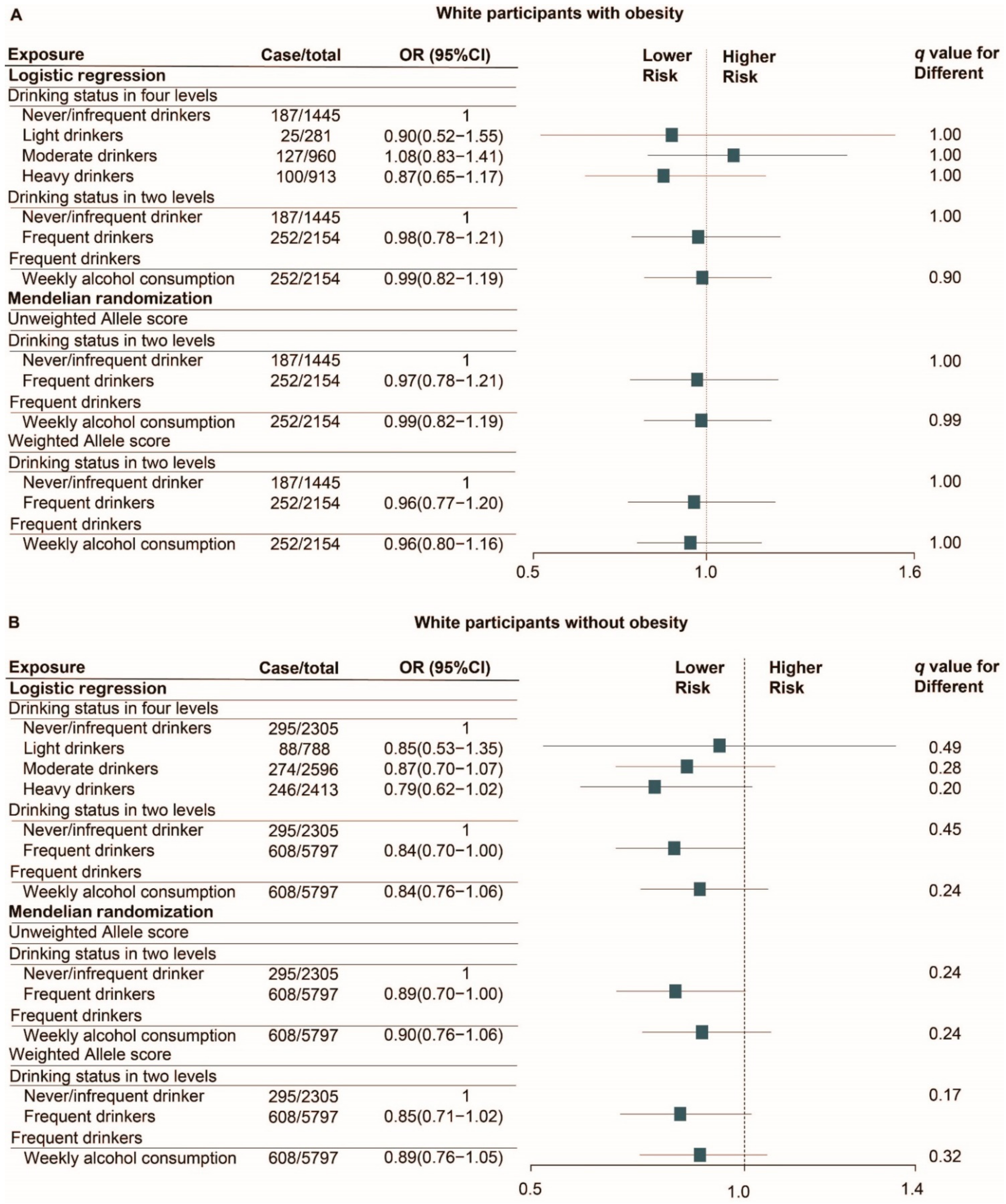
3.4. Alcohol and Risk of SARS-CoV-2 Infection
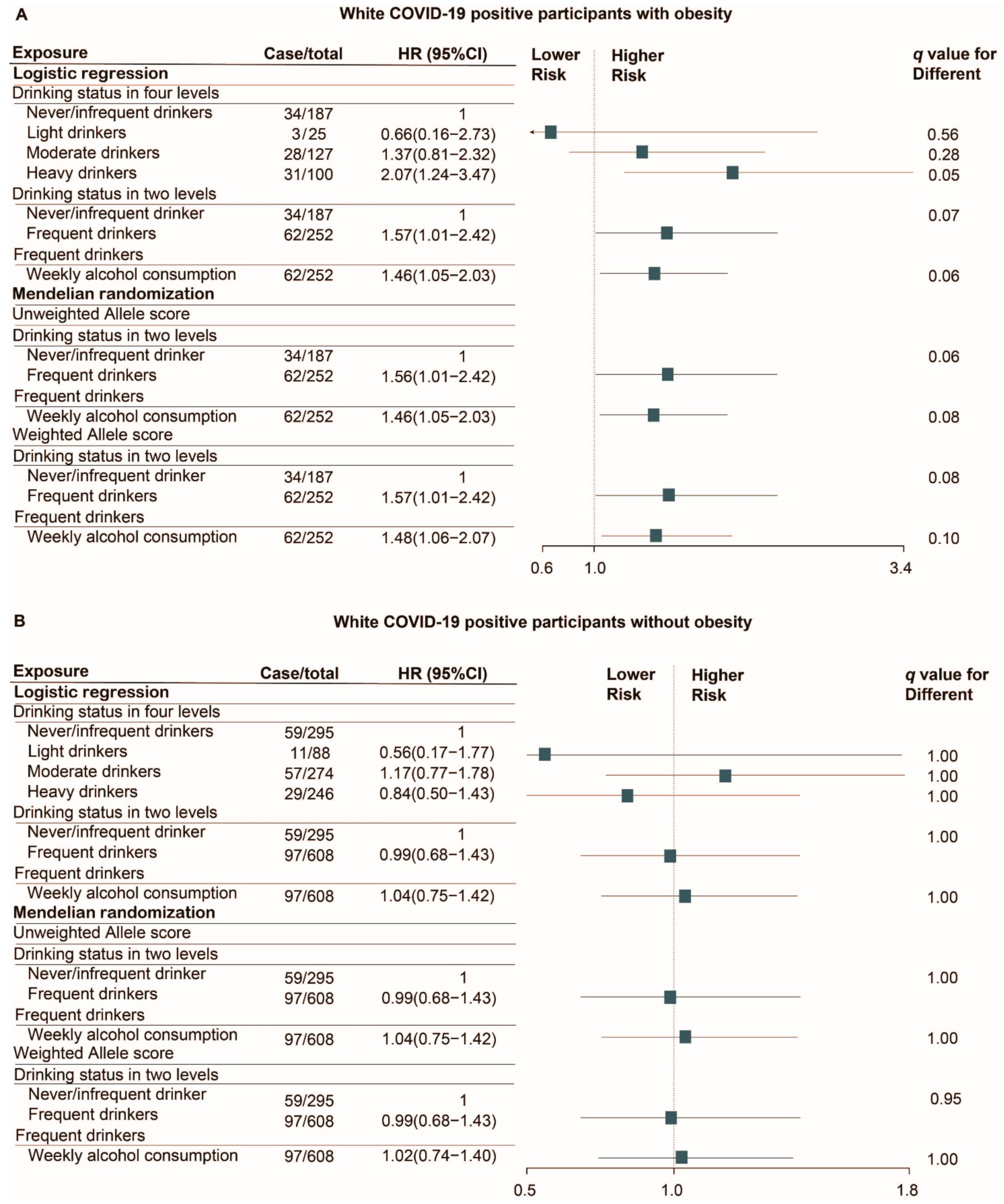
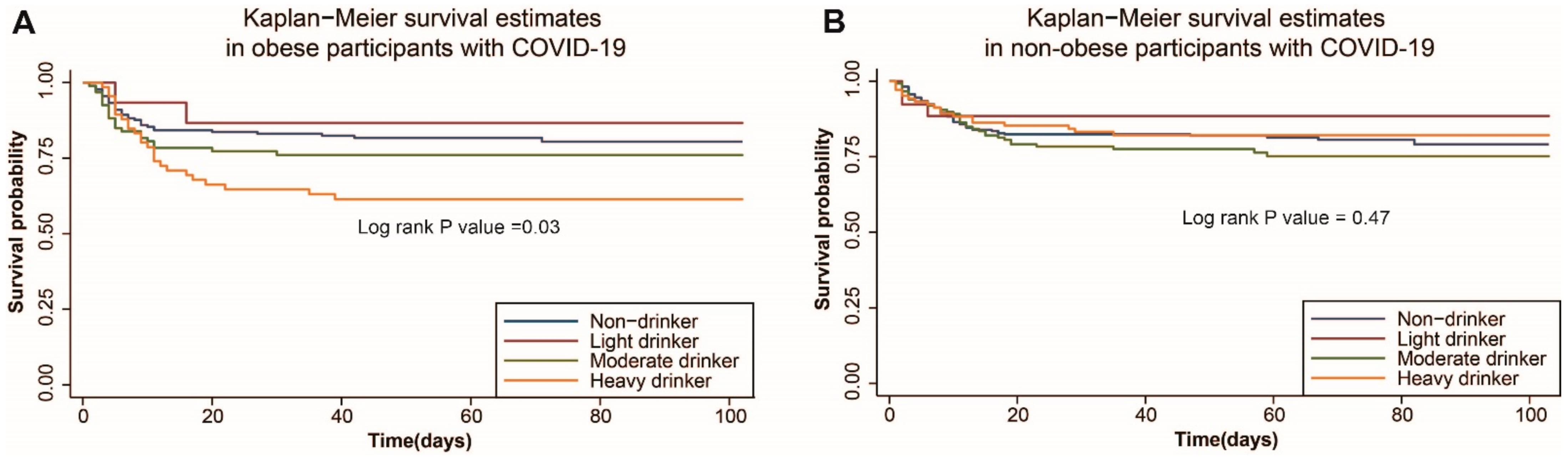
3.5. Alcohol and Risk of Death in White COVID-19 Positive Patients
3.6. Sensitivity Analysis
3.6.1. Association between Alcohol Consumption and Outcomes in Overweight but Not Obese Patients
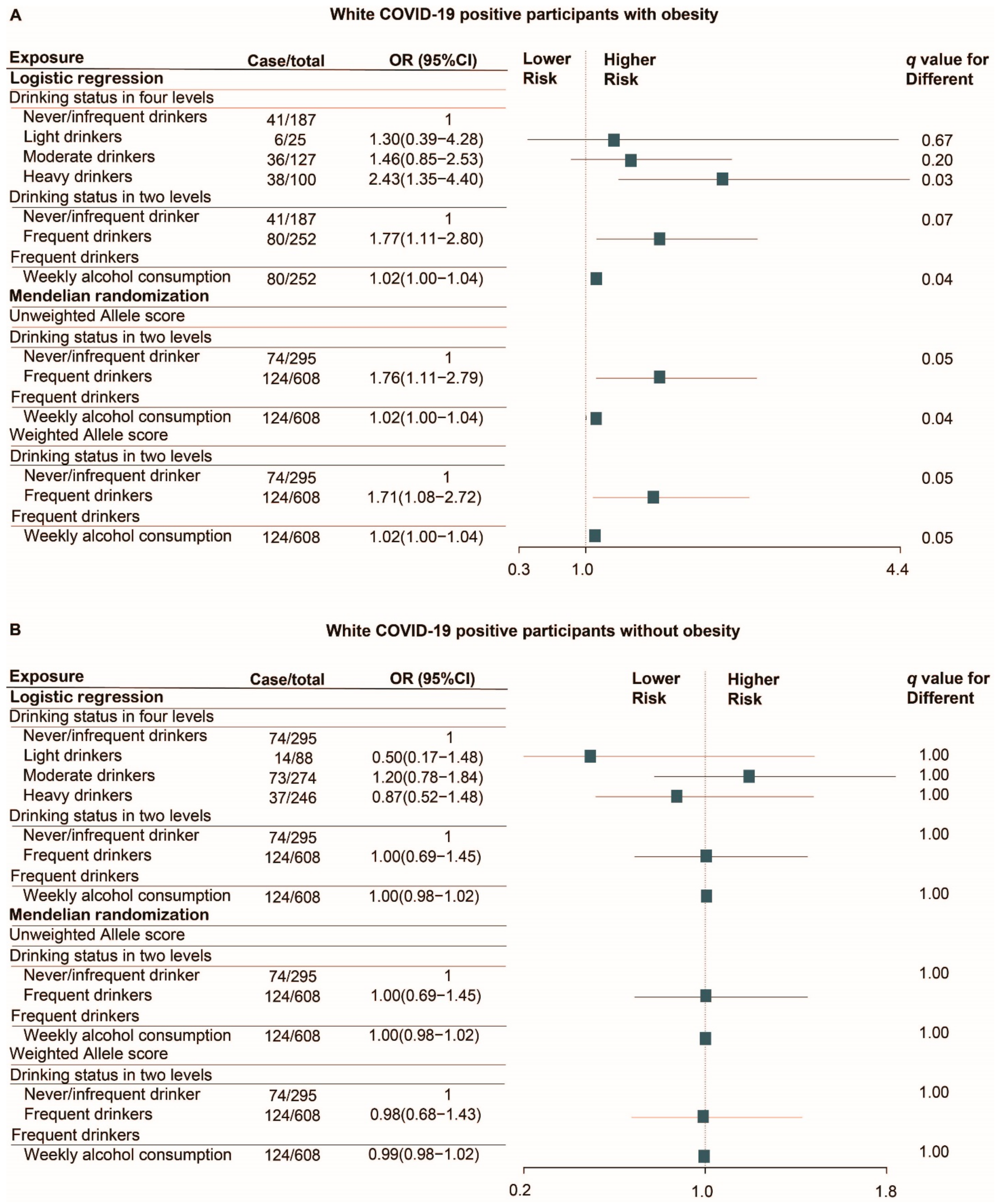
3.6.2. Risk of Severe Clinical Outcomes in White COVID-19 Positive Patients
3.6.3. Differences in the Association between Alcohol Consumption and Outcomes between Frequent and Non-Drinkers
3.7. Mediation Analysis
4. Discussion
4.1. Possible Explanations for an Interaction between Alcohol Consumption and Worse Outcomes of COVID-19 in Patients with Obesity
4.2. Strengths and Limitations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bergmann, C.C.; Silverman, R.H. COVID-19: Coronavirus replication, pathogenesis, and therapeutic strategies. Clevel. Clin. J. Med. 2020, 87, 321–327. [Google Scholar] [CrossRef] [PubMed]
- WHO. WHO Coronavirus Disease (COVID-19) Dashboard. Available online: https://covid19.who.int/ (accessed on 12 September 2020).
- Zhou, F.; Yu, T.; Du, R.; Fan, G.; Liu, Y.; Liu, Z.; Xiang, J.; Wang, Y.; Song, B.; Gu, X.; et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: A retrospective cohort study. Lancet 2020, 395, 1054–1062. [Google Scholar] [CrossRef]
- Wu, Z.; McGoogan, J.M. Characteristics of and Important Lessons From the Coronavirus Disease 2019 (COVID-19) Outbreak in China: Summary of a Report of 72314 Cases From the Chinese Center for Disease Control and Prevention. JAMA 2020, 323, 1239–1242. [Google Scholar] [CrossRef]
- Bednarski, P.J. Nielsen Data Says Alcoholic Beverage Sales through the Roof. Available online: https://www.mediapost.com/publications/article/349635/nielsen-data-says-alcoholic-beverage-sales-through.html (accessed on 12 September 2020).
- Pollard, M.S.; Tucker, J.S.; Green, H.D., Jr. Changes in Adult Alcohol Use and Consequences During the COVID-19 Pandemic in the US. JAMA Netw. Open 2020, 3, e2022942. [Google Scholar] [CrossRef]
- Happel, K.I.; Nelson, S. Alcohol, immunosuppression, and the lung. Proc. Am. Thorac. Soc. 2005, 2, 428–432. [Google Scholar] [CrossRef]
- Simet, S.M.; Sisson, J.H. Alcohol’s Effects on Lung Health and Immunity. Alcohol. Res. 2015, 37, 199–208. [Google Scholar] [PubMed]
- Hemann, E.A.; McGill, J.L.; Legge, K.L. Chronic ethanol exposure selectively inhibits the influenza-specific CD8 T cell response during influenza a virus infection. Alcohol. Clin. Exp. Res. 2014, 38, 2403–2413. [Google Scholar] [CrossRef]
- Samokhvalov, A.V.; Irving, H.M.; Rehm, J. Alcohol consumption as a risk factor for pneumonia: A systematic review and meta-analysis. Epidemiol. Infect. 2010, 138, 1789–1795. [Google Scholar] [CrossRef]
- Simou, E.; Leonardi-Bee, J.; Britton, J. The Effect of Alcohol Consumption on the Risk of ARDS. Chest 2018, 154, 58–68. [Google Scholar] [CrossRef]
- Romano, M.; Ruggiero, A.; Squeglia, F.; Maga, G.; Berisio, R. A Structural View of SARS-CoV-2 RNA Replication Machinery: RNA Synthesis, Proofreading and Final Capping. Cells 2020, 9, 1267. [Google Scholar] [CrossRef]
- Sanz-Garcia, C.; Poulsen, K.L.; Bellos, D.; Wang, H.; McMullen, M.R.; Li, X.; Chattopadhyay, S.; Sen, G.; Nagy, L.E. The non-transcriptional activity of IRF3 modulates hepatic immune cell populations in acute-on-chronic ethanol administration in mice. J. Hepatol. 2019, 70, 974–984. [Google Scholar] [CrossRef]
- Lee, J.H.; Shim, Y.R.; Seo, W.; Kim, M.H.; Choi, W.M.; Kim, H.H.; Kim, Y.E.; Yang, K.; Ryu, T.; Jeong, J.M.; et al. Mitochondrial Double-Stranded RNA in Exosome Promotes Interleukin-17 Production Through Toll-Like Receptor 3 in Alcohol-associated Liver Injury. Hepatology 2020, 72, 609–625. [Google Scholar] [CrossRef]
- Luther, J.; Khan, S.; Gala, M.K.; Kedrin, D.; Sridharan, G.; Goodman, R.P.; Garber, J.J.; Masia, R.; Diagacomo, E.; Adams, D.; et al. Hepatic gap junctions amplify alcohol liver injury by propagating cGAS-mediated IRF3 activation. Proc. Natl. Acad. Sci. USA 2020, 117, 11667–11673. [Google Scholar] [CrossRef]
- Gao, B.; Ahmad, M.F.; Nagy, L.E.; Tsukamoto, H. Inflammatory pathways in alcoholic steatohepatitis. J. Hepatol. 2019, 70, 249–259. [Google Scholar] [CrossRef]
- Neufeld, M.; Lachenmeier, D.W.; Ferreira-Borges, C.; Rehm, J. Is Alcohol an “Essential Good” During COVID-19? Yes, but Only as a Disinfectant! Alcohol. Clin. Exp. Res. 2020, 44, 1906–1909. [Google Scholar] [CrossRef]
- Luk, T.T.; Zhao, S.; Weng, X.; Wong, J.Y.; Wu, Y.S.; Ho, S.Y.; Lam, T.H.; Wang, M.P. Exposure to health misinformation about COVID-19 and increased tobacco and alcohol use: A population-based survey in Hong Kong. Tob. Control 2020. [Google Scholar] [CrossRef]
- WHO. Alcohol and COVID-19: What You Need to Know. 2020. Available online: https://www.euro.who.int/en/health-topics/disease-prevention/alcohol-use/data-and-statistics/fact-sheet-alcohol-and-covid-19-what-you-need-to-know-2020 (accessed on 12 September 2020).
- Sanchez-Roige, S.; Palmer, A.A.; Fontanillas, P.; Elson, S.L.; Adams, M.J.; Howard, D.M.; Edenberg, H.J.; Davies, G.; The 23andMe Research Team; The Substance Use Disorder Working Group of the Psychiatric Genomics Consortium; et al. Genome-Wide Association Study Meta-Analysis of the Alcohol Use Disorders Identification Test (AUDIT) in Two Population-Based Cohorts. Am. J. Psychiatry 2019, 176, 107–118. [Google Scholar] [CrossRef]
- Bennett, D.A.; Holmes, M.V. Mendelian randomisation in cardiovascular research: An introduction for clinicians. Heart 2017, 103, 1400–1407. [Google Scholar] [CrossRef]
- Noyce, A.J.; Nalls, M.A. Mendelian Randomization—The Key to Understanding Aspects of Parkinson’s Disease Causation? Mov. Disord. 2016, 31, 478–483. [Google Scholar] [CrossRef]
- Thompson, A.; Cook, J.; Choquet, H.; Jorgenson, E.; Yin, J.; Kinnunen, T.; Barclay, J.; Morris, A.P.; Pirmohamed, M. Functional validity, role, and implications of heavy alcohol consumption genetic loci. Sci. Adv. 2020, 6, eaay5034. [Google Scholar] [CrossRef]
- Liu, M.; Jiang, Y.; Wedow, R.; Li, Y.; Brazel, D.M.; Chen, F.; Datta, G.; Davila-Velderrain, J.; McGuire, D.; Tian, C.; et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat. Genet. 2019, 51, 237–244. [Google Scholar] [CrossRef]
- Canela-Xandri, O.; Rawlik, K.; Tenesa, A. An atlas of genetic associations in UK Biobank. Nat. Genet. 2018, 50, 1593–1599. [Google Scholar] [CrossRef]
- Burgess, S.; Small, D.S.; Thompson, S.G. A review of instrumental variable estimators for Mendelian randomization. Stat. Methods Med. Res. 2017, 26, 2333–2355. [Google Scholar] [CrossRef] [PubMed]
- NIAAA. Drinking Levels Defined. Available online: https://www.niaaa.nih.gov/alcohol-health/overview-alcohol-consumption/moderate-binge-drinking (accessed on 12 September 2020).
- Howe, L.J.; Lawson, D.J.; Davies, N.M.; St Pourcain, B.; Lewis, S.J.; Davey Smith, G.; Hemani, G. Genetic evidence for assortative mating on alcohol consumption in the UK Biobank. Nat. Commun. 2019, 10, 5039. [Google Scholar] [CrossRef]
- Beasley, M.; Freidin, M.B.; Basu, N.; Williams, F.M.K.; Macfarlane, G.J. What is the effect of alcohol consumption on the risk of chronic widespread pain? A Mendelian randomisation study using UK Biobank. Pain 2019, 160, 501–507. [Google Scholar] [CrossRef]
- Teumer, A. Common Methods for Performing Mendelian Randomization. Front. Cardiovasc. Med. 2018, 5, 51. [Google Scholar] [CrossRef] [PubMed]
- Flegal, K.M.; Kit, B.K.; Graubard, B.I. Body mass index categories in observational studies of weight and risk of death. Am. J. Epidemiol. 2014, 180, 288–296. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B Methodol. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Zhang, Z. Propensity score method: A non-parametric technique to reduce model dependence. Ann. Transl. Med. 2017, 5, 7. [Google Scholar] [CrossRef]
- Wolff, D.; Nee, S.; Hickey, N.S.; Marschollek, M. Risk factors for Covid-19 severity and fatality: A structured literature review. Infection 2020, 49, 15–28. [Google Scholar] [CrossRef]
- Banerjee, M.; Gupta, S.; Sharma, P.; Shekhawat, J.; Gauba, K. Obesity and COVID-19: A Fatal Alliance. Indian J. Clin. Biochem. 2020, 35, 410–417. [Google Scholar] [CrossRef] [PubMed]
- Nelson, S.; Kolls, J.K. Alcohol, host defence and society. Nat. Rev. Immunol. 2002, 2, 205–209. [Google Scholar] [CrossRef]
- Tleyjeh, I.M.; Ghomrawi, H.M.; Steckelberg, J.M.; Montori, V.M.; Hoskin, T.L.; Enders, F.; Huskins, W.C.; Mookadam, F.; Wilson, W.R.; Zimmerman, V.; et al. Conclusion about the association between valve surgery and mortality in an infective endocarditis cohort changed after adjusting for survivor bias. J. Clin. Epidemiol. 2010, 63, 130–135. [Google Scholar] [CrossRef]
- Kruglikov, I.L.; Scherer, P.E. The Role of Adipocytes and Adipocyte-Like Cells in the Severity of COVID-19 Infections. Obesity 2020, 28, 1187–1190. [Google Scholar] [CrossRef]
- Kang, L.; Sebastian, B.M.; Pritchard, M.T.; Pratt, B.T.; Previs, S.F.; Nagy, L.E. Chronic ethanol-induced insulin resistance is associated with macrophage infiltration into adipose tissue and altered expression of adipocytokines. Alcohol. Clin. Exp. Res. 2007, 31, 1581–1588. [Google Scholar] [CrossRef]
- Chang, B.; Xu, M.J.; Zhou, Z.; Cai, Y.; Li, M.; Wang, W.; Feng, D.; Bertola, A.; Wang, H.; Kunos, G.; et al. Short- or long-term high-fat diet feeding plus acute ethanol binge synergistically induce acute liver injury in mice: An important role for CXCL1. Hepatology 2015, 62, 1070–1085. [Google Scholar] [CrossRef] [PubMed]
- Hwang, S.; Ren, T.; Gao, B. Obesity and binge alcohol intake are deadly combination to induce steatohepatitis: A model of high-fat diet and binge ethanol intake. Clin. Mol. Hepatol. 2020, 26, 586. [Google Scholar] [CrossRef]
- Okuno, F.; Arai, M.; Ishii, H.; Shigeta, Y.; Ebihara, Y.; Takagi, S.; Tsuchiya, M. Mild but prolonged elevation of serum angiotensin converting enzyme (ACE) activity in alcoholics. Alcohol 1986, 3, 357–359. [Google Scholar] [CrossRef]
- Traversy, G.; Chaput, J.P. Alcohol Consumption and Obesity: An Update. Curr. Obes. Rep. 2015, 4, 122–130. [Google Scholar] [CrossRef]
- Griffith, G.J.; Morris, T.T.; Tudball, M.J.; Herbert, A.; Mancano, G.; Pike, L.; Sharp, G.C.; Sterne, J.; Palmer, T.M.; Davey Smith, G.; et al. Collider bias undermines our understanding of COVID-19 disease risk and severity. Nat. Commun. 2020, 11, 5749. [Google Scholar] [CrossRef]
| ADH1B rs1229984 | SLC39A8 rs13107325 | GCKR rs1260326 | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Variables | 1/1 (Slow) n = 12,172 | 1/2 or 2/2 (Fast) n = 765 | q-Value | 1/1 (Slow) n = 11,160 | 1/2 or 2/2 (Fast) n = 1756 | q-Value | 1/1 (sLow) n = 4947 | 1/2 or 2/2 (Fast) n = 7920 | q-Value |
| Age (years), n (%) | 0.88 | 0.79 | 1.00 | ||||||
| <65 | 3500 (28.8) | 241 (31.5) | 3274 (29.3) | 463 (26.4) | 1461 (29.5) | 2252 (28.4) | |||
| ≥65 | 8672 (71.2) | 524 (68.5) | 7886 (70.7) | 1293 (73.6) | 3486 (70.5) | 5668 (71.6) | |||
| Male, n (%) | 5977 (49.1) | 358 (46.8) | 0.81 | 5443 (48.4) | 881 (50.2) | 0.80 | 2452 (49.6) | 3856 (48.7) | 0.22 |
| Race, n (%) | <0.001 | <0.001 | <0.001 | ||||||
| No white | 767 (6.3) | 141 (18.5) | 879 (7.9) | 28 (1.6) | 512 (10.4) | 380 (4.8) | |||
| White | 11,359 (93.7) | 623 (81.5) | 10,252 (92.1) | 1720 (98.4) | 4416 (89.6) | 7514 (95.2) | |||
| BMI categories, n (%) | |||||||||
| Normal weight (18.5–24.9) | 3201 (27.0) | 221 (29.4) | 3019 (27.7) | 396 (23.0) | 1263 (26.3) | 2137 (27.6) | |||
| Underweight (<18.5) | 59 (0.5) | 5 (0.7) | 1.00 | 54 (0.5) | 10 (0.6) | 0.76 | 23 (0.5) | 40 (0.5) | 0.99 |
| Overweight (25–29.9) | 4891 (41.2) | 341 (45.4) | 1.00 | 4460 (41.0) | 762 (44.3) | 0.002 | 1946 (40.5) | 3267 (42.2) | 0.95 |
| Obesity (≥30) | 3719 (31.3) | 184 (24.5) | 0.06 | 3350 (30.8) | 551 (32.1) | 0.01 | 1577 (32.8) | 2299 (29.7) | 0.07 |
| Blood type, n (%) | |||||||||
| OO | 5128 (42.2) | 330 (43.1) | 4706 (42.2) | 742 (42.4) | 2098 (42.5) | 3331 (42.1) | |||
| AA + AO | 416 (3.4) | 31 (4.1) | 0.91 | 388 (3.5) | 58 (3.3) | 0.97 | 169 (3.4) | 274 (3.5) | 0.91 |
| BB + BO | 1221 (10.0) | 108 (14.1) | 0.77 | 1155 (10.4) | 172 (9.8) | 0.85 | 520 (10.5) | 800 (10.1) | 0.53 |
| AB | 5390 (44.3) | 296 (38.7) | 0.93 | 4900 (44.0) | 780 (44.5) | 0.90 | 2154 (43.6) | 3504 (44.3) | 1.00 |
| Current smoking, n (%) | |||||||||
| No | 10,656 (87.6) | 673 (88.0) | 9775 (87.6) | 1536 (87.6) | 4318 (87.3) | 6947 (87.8) | |||
| Only occasionally | 369 (3.0) | 27 (3.5) | 1.00 | 351 (3.1) | 44 (2.5) | 0.79 | 157 (3.2) | 236 (3.0) | 0.97 |
| Most or all days | 1137 (9.3) | 65 (8.5) | 0.87 | 1027 (9.2) | 173 (9.9) | 0.81 | 470 (9.5) | 729 (9.2) | 0.50 |
| Comorbidities, n (%) | |||||||||
| Upper gastrointestinal diseases | |||||||||
| Oesophagitis | 579 (4.8) | 21 (2.7) | 0.81 | 518 (4.6) | 81 (4.6) | 0.86 | 188 (3.8) | 409 (5.2) | 0.02 |
| GERD | 1363 (11.2) | 74 (9.7) | 0.99 | 1226 (11.0) | 207 (11.8) | 0.85 | 559 (11.3) | 871 (11.0) | 0.56 |
| Peptic ulcer | 428 (3.5) | 25 (3.3) | 1.00 | 396 (3.5) | 57 (3.2) | 0.56 | 175 (3.5) | 277 (3.5) | 1.00 |
| Gastritis/duodenitis | 1458 (12.0) | 74 (9.7) | 0.81 | 1311 (11.7) | 218 (12.4) | 0.58 | 570 (11.5) | 953 (12.0) | 0.99 |
| Chronic lower respiratory diseases | |||||||||
| COPD | 753 (6.2) | 33 (4.3) | 0.84 | 677 (6.1) | 109 (6.2) | 0.60 | 295 (6.0) | 489 (6.2) | 1.00 |
| Emphysema | 141 (1.2) | 6 (0.8) | 0.97 | 118 (1.1) | 28 (1.6) | 0.41 | 50 (1.0) | 97 (1.2) | 0.49 |
| Bronchitis/Bronchiectasis | 196 (1,6) | 8 (1.0) | 0.92 | 165 (1.5) | 39 (2.2) | 0.23 | 71 (1.4) | 131 (1.7) | 1.00 |
| Asthma | 1383 (11.4) | 78 (10.2) | 0.92 | 1259 (11.3) | 199 (11.3) | 0.97 | 538 (10.9) | 914 (11.5) | 0.46 |
| Chronic heart diseases | |||||||||
| Heart failure | 471 (3.9) | 35 (4.6) | 0.67 | 440 (3.9) | 64 (3.6) | 0.81 | 184 (3.7) | 318 (4.0) | 0.51 |
| Hypertensive | 4444 (36.5) | 252 (32.9) | 0.75 | 4065 (36.4) | 625 (35.6) | 0.53 | 1815 (36.7) | 2857 (36.1) | 1.00 |
| Chronic ischaemic heart disease | 1640 (13.5) | 97 (12.7) | 0.94 | 1496 (13.4) | 239 (13.6) | 0.93 | 628 (12.7) | 1101 (13.9) | 0.09 |
| Diabetes mellitus | 1475 (12.1) | 91 (11.9) | 0.94 | 1349 (12.1) | 215 (12.2) | 0.86 | 662 (13.4) | 893 (11.3) | 0.15 |
| Serious liver diseases | 88 (0.7) | 3 (0.4) | 0.89 | 79 (0.7) | 12 (0.7) | 0.90 | 29 (0.6) | 62 (0.8) | 0.47 |
| Renal failure | 897 (7.4) | 52 (6.8) | 0.93 | 807 (7.2) | 139 (7.9) | 0.72 | 388 (7.8) | 556 (7.0) | 0.52 |
| Insomnia | 9463 (77.8) | 601 (78.6) | 0.82 | 8675 (77.8) | 1373 (78.2) | 0.89 | 3887 (78.7) | 6128 (77.4) | 0.20 |
| Dementia | 91 (0.7) | 2 (0.3) | 0.88 | 80 (0.7) | 13 (0.7) | 0.93 | 44 (0.9) | 49 (0.6) | 0.16 |
| Tumor | 1246 (10.2) | 77 (10.1) | 0.90 | 1143 (10.2) | 177 (10.1) | 0.82 | 505 (10.2) | 815 (10.3) | 0.64 |
| AIDS | 11 (0.1) | 2 (0.3) | 0.72 | 12 (0.1) | 1 (0.1) | 0.83 | 4 (0.1) | 8 (0.1) | 1.00 |
| Alcohol Consumption (Standard Drink/Weekly) in Frequent Drinkers (95% CI) (n = 8131) | OR of Being a Heavy Drinker in the Whole Cohort (95% CI) (n = 11982) | |
|---|---|---|
| ADH1B one or two fast alleles vs. none | −2.78 (−4.08–−1.47) | 0.525 (0.43–0.65) |
| F-test | 17.42 | — |
| p-value | <0.001 | <0.001 |
| SLC39A8 one or two fast alleles vs. none | −0.92 (−1.71–−0.13) | 0.87 (0.78–0.98) |
| F-test | 5.1 | — |
| p-value | <0.001 | 0.022 |
| GCKR one or two fast alleles vs. none | −0.79 (−1.36–−0.22) | 0.92(0.84–0.99) |
| F-test | 7.33 | — |
| p-value | <0.001 | 0.039 |
| Unweighted allele score | −0.73 (−1.07–−0.39) | 0.89 (0.89–0.94) |
| F-test | 17.47 | — |
| p-value | <0.001 | <0.001 |
| Weighted allele score | −15.07 (−20.83–9.31) | 0.05 (0.02–0.12) |
| F-test | 26.29 | — |
| p-value | <0.001 | <0.001 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fan, X.; Liu, Z.; Poulsen, K.L.; Wu, X.; Miyata, T.; Dasarathy, S.; Rotroff, D.M.; Nagy, L.E. Alcohol Consumption Is Associated with Poor Prognosis in Obese Patients with COVID-19: A Mendelian Randomization Study Using UK Biobank. Nutrients 2021, 13, 1592. https://doi.org/10.3390/nu13051592
Fan X, Liu Z, Poulsen KL, Wu X, Miyata T, Dasarathy S, Rotroff DM, Nagy LE. Alcohol Consumption Is Associated with Poor Prognosis in Obese Patients with COVID-19: A Mendelian Randomization Study Using UK Biobank. Nutrients. 2021; 13(5):1592. https://doi.org/10.3390/nu13051592
Chicago/Turabian StyleFan, Xiude, Zhengwen Liu, Kyle L. Poulsen, Xiaoqin Wu, Tatsunori Miyata, Srinivasan Dasarathy, Daniel M. Rotroff, and Laura E. Nagy. 2021. "Alcohol Consumption Is Associated with Poor Prognosis in Obese Patients with COVID-19: A Mendelian Randomization Study Using UK Biobank" Nutrients 13, no. 5: 1592. https://doi.org/10.3390/nu13051592
APA StyleFan, X., Liu, Z., Poulsen, K. L., Wu, X., Miyata, T., Dasarathy, S., Rotroff, D. M., & Nagy, L. E. (2021). Alcohol Consumption Is Associated with Poor Prognosis in Obese Patients with COVID-19: A Mendelian Randomization Study Using UK Biobank. Nutrients, 13(5), 1592. https://doi.org/10.3390/nu13051592







