Abstract
Probiotics are reported to improve gastrointestinal (GI) function via regulating gut microbiota (GM). However, exactly how probiotics influence GM and GI function in elders is poorly characterized. Therefore, in this study, we assessed the effect of the probiotic Lacticaseibacillus paracasei PS23 (LPPS23) on the GM and GI function of aged mice. There were four groups of senescence-accelerated mouse prone-8 (SAMP8) mice (n = 4): a non-treated control group, a saline control group, a low dose LPPS23 group (1 × 108 colony-forming unit (CFU)/mouse/day), and a high dose LPPS23 group (1 × 109 CFU/mouse/day). Non-treated mice were euthanized at 16 weeks old, and others were euthanized at 28 weeks old. The next-generation sequencing results revealed that LPPS23 enriched Lactobacillus and Candidatus_Saccharimonas, while the abundance of Lachnospiraceae_UCG_001 decreased in aged mice given LPPS23. The abundance of Lactobacillus negatively correlated with the abundance of Erysipelotrichaceae. Moreover, LPPS23 improved the GI function of aged mice due to the longer intestine length, lower intestinal permeability, and higher phagocytosis in LPPS23-treated mice. The ELISA results showed that LPPS23 attenuated the alterations of pro-inflammatory factors and immunoglobulins. The abundance of LPPS23-enriched Lactobacillus was positively correlated with healthy GI function, while Lachnospiraceae_UCG_001, which was repressed by LPPS23, was negatively correlated with a healthy GI function in the aged mice according to Spearman’s correlation analysis. Taken together, LPPS23 can effectively modulate GM composition and improve GI function in aged SAMP8 mice.
1. Introduction
Aging is an important factor in several disorders that influence the quality of life of elderly individuals. The age-related alteration of gut microbiota (GM) composition is linked to age-related diseases, such as cognitive impairment, sarcopenia, and gastrointestinal (GI) dysfunction [1]. Therefore, good GM composition should prevent age-related disorders.
Several studies have addressed the interaction among GM, GI function, and aging. Sovran et al. linked GM alterations to age-associated impairments in GI mucus barrier function and age-related inflammation [2]. Moreover, aging significantly increases GI vulnerability, and 40% of geriatric patients report at least one GI complaint during routine physical examinations [3]. Previous studies have also indicated that age-related GM alterations and GI dysfunctions exist, such as bacterial overgrowth, increased intestinal permeability, and decreased absorption [4,5,6]. Furthermore, GM alterations and GI dysfunctions are suggested to cause age-related inflammation [7,8,9]. Thus, preventing age-related GI dysfunction should cooperate with GM to attenuate the diseases derived by age-related inflammation, such as sarcopenia, cognitive impairment, and metabolic diseases [10].
Probiotic supplementation is widely used to modulate GM composition and enhance GI function [11,12]. In addition, many probiotics have been reported to exert anti-inflammatory properties [13]. Therefore, probiotic supplementation may protect against GI dysfunction and inflammation in the elderly. However, to our knowledge, the precise effects of probiotics on GI function in the elderly remain unclear; only a small number of studies have investigated the effects of probiotic supplements in age-related disorders [14]. Thus, in order to provide a solid foundation for the use of probiotic-based strategies to prevent age-related diseases, it is important to investigate the effects of probiotics on GM and GI dysfunction in the aged subjects.
In our previous work, we demonstrated that probiotics could restructure GM and improve GI function. We also reported that Lacticaseibacillus paracasei PS23 (LPPS23) was beneficial in mitigating age-related diseases, including cognitive impairment and sarcopenia, in SAMP8 (senescence accelerated mouse prone-8) mice [14,15]. Moreover, administration of LPPS23 for 12 weeks significantly reduced age-related inflammation and protein uptake decline in SAMP8 mice [14]. Therefore, we assumed that LPPS23 could regulate the GM composition and GI function to prevent age-related disorders and so further investigated how LPPS23 influenced GM and GI function in the elderly by assessing the interactions between LPPS23 and the host. We administered LPPS23 to SAMP8 mice over the course of their lifespans, from youth into old age, and observed the effects of LPPS23 on GM composition and GI function.
2. Materials and Methods
2.1. Probiotics and Animals
LPPS23 was provided by Dr. Ying-Chieh Tsai (National Yang Ming Chiao Tung University, Taipei, Taiwan). LPPS23 is a strain isolated from healthy human feces and identified by phylogenetic classification of its 16S rRNA gene sequence. LPPS23 can induce anti-inflammatory effects and improve brain and muscle health [14,15,16]. In the present study, LPPS23 was subcultured before used. The doses of live LPPS23 used for administration were 1 × 108 and 1 × 109 colony-forming unit (CFU)/200 μL; SAMP8 mice were bred by Dr. Ming-Fu Wang. To make sure that age was the only factor to induce GI dysfunction, all mice were housed under standard laboratory conditions with a 12/12 h light/dark cycle at 22–24 °C and 40–60% humidity. SAMP8 mice were provided with a commercially available diet (local supplier) and sterile water ad libitum. Well-trained investigators monitored the conditions of the mice every day with no stress or discomfort observed in their behavior. Additionally, during the study, no unhealthy phenomena in the feeding conditions were observed.
The female mice were divided into four groups (n = 4): FC (16-week-old non-aged mice controls), FA (aged mice administered saline), FPS23L (aged mice administered low dose 1 × 108 CFU LPPS23/mouse/day), and FPS23H (aged mice administered high dose 1 × 109 LPPS23 CFU/mouse/day). The dose of LPPS23 was chosen based on our previous studies [14,15], which indicated LPPS23 at 1 × 109 CFU/mouse/day prevented age-related decline in SAMP8 mice. The mice received 200 µL of saline or live LPPS23 at 9:00 a.m. by gavage every day for 12 weeks from 16 weeks of age until 28 weeks of age. The sample size was referred to in the previous studies regarding the composition of GM and gut function in mice [17,18].
The FC group was humanely sacrificed at 16 weeks old, and the other groups were sacrificed after 12 weeks of treatment at 28 weeks old. Intestine, intestinal mucosa, and fecal samples were collected, and intestinal length was measured. The mucosal samples were collected by intestine perfusion with 5 mL phosphate buffered saline (PBS) after sacrifice. Then, the supernatants were isolated by 2000× g centrifuge at 4 °C for 30 min for the analysis of pro-inflammatory factors and immunoglobulins. The animal protocol was approved by the Institutional Animal Care and Use Committee of Shih Chien University (IACUC-10407).
2.2. Intestinal Permeability
On the same day that the saline- or LPPS23-treated mice were to be euthanized, they were deprived of food 6 h prior to oral gavage of 200 μL of 80 mg/mL FITC–dextran (4 kDa; Sigma-Aldrich, St. Louis, MO, USA) followed by 4 h deprivation of both food and water. Blood samples were collected, and fluorescence intensity was measured on fluorescence plates using an excitation wavelength of 490 nm and an emission wavelength of 520 nm.
2.3. Enzyme-Linked Immunosorbent Assays
Pro-inflammatory factors in the intestinal mucosa were determined using sandwich ELISA (enzyme-linked immunosorbent assay) kits produced by Biolegend (San Diego, CA, USA) for tumor necrosis factor (TNF)-α and monocyte chemotactic protein (MCP)-1, according to the manufacturer’s instructions. The ELISA reader (BioTek, Winooski, VT, USA) was used to measure absorbance values of the samples.
2.4. Quantification of IgA, IgE, IgM, and IgG in the Intestinal Mucosa
The levels of IgA, IgE, IgM, and IgG in the intestinal mucosa were analyzed using a mouse IgA ELISA kit (eBioscience, Santa Clara, CA, USA), mouse IgE ELISA kit (Biolegend), mouse IgM ELISA kit (Biolegend), and mouse Total IgG ELISA kit (eBioscience) according to the manufacturers’ instructions. The plates were read at 450 nm.
2.5. Bacterial Genomic DNA Isolation
Total genomic DNA was isolated from 200 mg fecal samples using the QIAamp DNA Stool Mini kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. The concentrations of DNA were measured with a NanoDrop2000 (Thermo Scientific, Waltham, MA, USA), and the samples were stored at −80 °C until use.
2.6. 16S rRNA Gene Sequencing and Data Analysis
The V3-V4 region of the 16S rRNA gene was PCR-amplified using the primers 341F (5′-CCTAYGGGRBGCASCAG-3′) and 806R (5′-GGACTACNNGGGTATCTAAT-3′) to develop the amplicon libraries according to recommended Illumina 16S Metagenomic Sequencing Library Preparation manual protocol. Then, the amplicons were paired-end sequenced (PE 2 × 250) using an Illumina HiSeq 2000 platform according to the manufacturer’s protocol. The paired forward and reverse reads passed quality control and were merged, and then mapped to the Silva database to construct operational taxonomic units (OTUs) at 97% identity through the UPARSE pipeline (drive5, Tiburon, CA, USA) [19].
Quantitative Insights Into Microbial Ecology (QIIME) [20] was used to analyze data. Chimeric sequences were removed using ChimeraSlayer [21]. Sequences with ≥97% similarity were assigned to the same operational taxonomic units (OTUs). The Silva database [22] was used to annotate the genes. Alpha diversity analysis (Shannon index) was assessed using QIIME. Beta diversity was analyzed using weighted principal coordinate analysis (PCoA) and partial least squares discriminant analysis (PLS-DA) using QIIME; PERMANOVA was used for analyzing statistical significance. Linear discriminant analysis (LDA) effect size (LefSe) was performed online using the Galaxy workflow framework [23].
2.7. Statistical Analyses
Data are presented as the mean ± standard error of the mean (SEM). Data of bacteria were analyzed using nonparametric one-way ANOVA with one-way ANOVA post hoc, and the other data were analyzed using one-way ANOVA with a Tukey honestly significant difference (HSD) post hoc test. Rank tests with Spearman’s correlation coefficient were used to assess associations between two bacteria, between bacteria and parameters of intestinal function, between bacteria and pro-inflammatory factors, and between bacteria and Ig. A p-value < 0.05 was considered statistically significant.
3. Results
3.1. Effects of LPPS23 on GM Community Composition and Diversity
The three major bacterial phyla identified in the SAMP8 mice feces were Bacteroidetes, Firmicutes, and Proteobacteria (Figure 1A), which were evaluated by at least 52,597 reads/sample. Moreover, the ratio of Firmicutes/Bacteroidetes, an indicator of gut inflammation [24], was no different among the groups (Figure 1B). Alpha diversity was not significantly different between the FC, FA, FPS23L, and FPS23H groups (Figure 1C). The microbial communities of all groups were separated by the principal co-ordinates analysis (PCoA) (Figure 1D) and partial least squares–discriminant analysis (PLSDA) (p = 0.042) (Figure 1E), though there was no statistical difference in the PCoA analysis (p = 0.153).
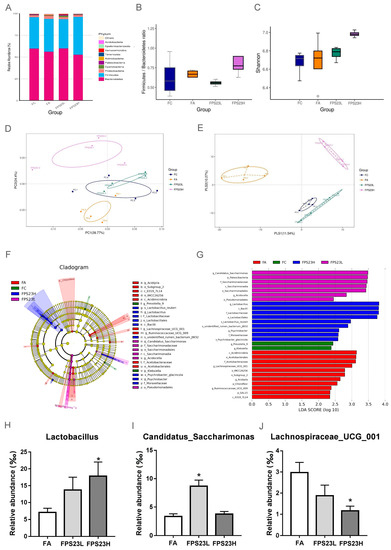
Figure 1.
Gut microbiota composition. (A) Bacterial community distribution; (B) ratio of Firmicutes/Bacteroidetes; (C) alpha diversity indicated by Shannon’s diversity index; (D,E) beta diversity indicated by weighted principal co-ordinates analysis (PCoA) (D) and partial least squares discriminant analysis (PLS-DA) (E) in senescence accelerated mouse prone-8 (SAMP8) mice; (F,G) Linear discriminant analysis effect size (LEfSe) analysis of the gut microbiota of SAMP8 mice; (H–J) relative abundance of Lactobacillus (H), Candidatus_Saccharimonas (I), and Lachnospiraceae UCG 001 (J) in the aged SAMP8 mice. Asterisk (*) indicates significant differences from the female aged mice administered saline (FA) (p < 0.05). n = 4.
3.2. Effects of LPPS23 on Abundant GM
Next, we identified the microbial species that were enriched from the class to species level using LEfSe analysis. Bacteria belonging to the order Lactobacillales increased in the FPS23H mice. Moreover, both the high and low doses of LPPS23 enhanced bacteria belonging to the order Pseudomonadales, and the low dose of LPPS23 promoted bacteria from the class Saccharimonadia. The bacteria enriched in the FA group belonged to the class Acidimicrobiia, genus Lachnospiraceae_UCG_001, genus Ruminococcaceae_ UCG_009, and order Acetobacterales. Bacteria from the genus Prevotella and Klebsiella were enriched in the FC group (Figure 1F,G).
Bacteria with LDA scores > 3.0 at the genus level in the aged mice were also compared between the LPPS23-treated and aged control mice. Lactobacillus (Figure 1H) was significantly enriched in the FPS23H mice, and the enrichment of Candidatus_Saccharimonas (Figure 1I) was observed in the FPS23L mice. Moreover, Lachnospiraceae_UCG_001 (Figure 1J) abundance decreased in FPS23H mice.
3.3. Bacteria Correlated with Lactobacillus
We used Spearman’s correlation analysis to identify bacteria that are correlated with Lactobacillus. Lactobacillus was negatively correlated with bacteria from the family Erysipelotrichaceae in mice (r = −0.62, p = 0.035) (Figure 2).
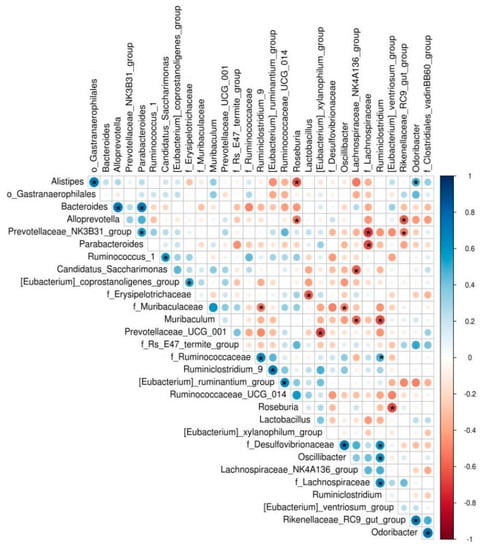
Figure 2.
Spearman’s correlation analysis of the gut microbiota of SAMP8 mice. Asterisk (*) indicates p < 0.05. n = 4.
3.4. Effects of LPPS23 on Intestinal Parameters
Since LPPS23 regulated GM composition in the present study and was reported to enhance the protein uptake in the aged SAMP8 mice [14], LPPS23 might have effects on GI function. As age-related intestinal shortening and barrier dysfunction are associated with GM alterations [25], we measured intestinal length, intestinal permeability, and phagocytotic activity in 16-week-old untreated SAMP8 mice and 28-week-old SAMP8 mice after administration of saline or LPPS23 daily for 12 weeks to assess the effects of LPPS23 on the age-related change in the intestine. Significant age-related declines in intestinal function were observed in the mice including decreased length of the intestine (Figure 3A,B), increased intestinal permeability (Figure 3C), and decreased phagocytotic activity (Figure 3D) in the FA group compared to the FC group. Moreover, LPPS23 treatment prevented these age-related changes in the mice. These results indicated that LPPS23 might maintain intestinal function during the aging process.
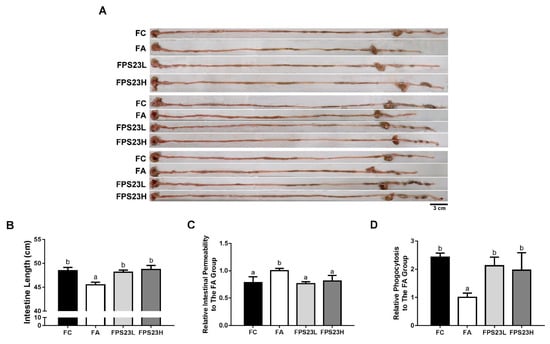
Figure 3.
Intestinal function. (A) Representative experiments of intestine length, (B) intestinal length, (C) intestinal permeability, and (D) phagocytosis in the SAMP8 mice. Different superscript letters (a, b, c) indicate significant differences in one-way ANOVA by Tukey’s honestly significant difference (HSD) post hoc test (p < 0.05). n = 4. Scale bar = 3 cm.
3.5. Effects of LPPS23 on Levels of the Pro-Inflammatory Factors TNF-α and MCP-1
Levels of the pro-inflammatory factors TNF-α and MCP-1 were quantified to assess the effect of LPPS23 on the aging intestine. Both pro-inflammatory factors increased during aging. LPPS23 suppressed these age-related changes in mice (Figure 4A,B). However, there was no dose response effect for LPPS23 (Figure 4A,B).
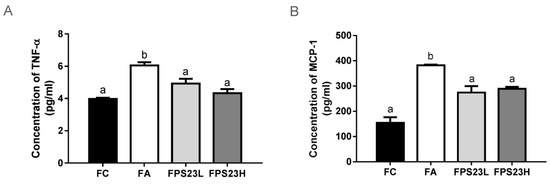
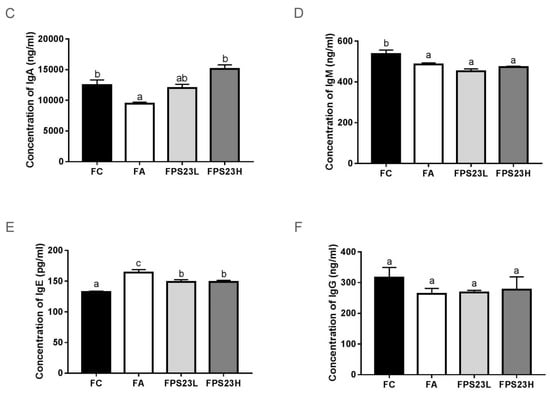
Figure 4.
Pro-inflammatory cytokines and immunoglobulins in the intestinal mucosa. (A) TNF-α; (B) MCP-1; (C) IgA; (D) IgM; (E) IgE; (F) IgG in the SAMP8 mice. Different superscript letters (a, b, c) indicate significant differences in one-way ANOVA by Tukey HSD post-hoc test (p < 0.05). n = 4.
3.6. Effects of LPPS23 on Levels of Immunoglobulins in the Intestinal Mucosa
The levels of immunoglobulins in the intestinal mucosa were also quantified to investigate immune function. Aging reduced the levels of IgA (Figure 4C) and IgM (Figure 4D), induced IgE levels (Figure 4E), and had no impact on IgG (Figure 4F) in mice. Moreover, these age-related changes in IgA (Figure 4C) and IgE (Figure 4E) in mice were significantly reduced by the administration of LPPS23. The higher dose of LPPS23 greatly reduced the age-related decline of IgA more than the lower dose of LPPS23.
3.7. Correlation of LPPS23-Modulated Bacteria and GI Conditions
The correlation of bacteria and GI conditions was analyzed, because LPPS23 significantly altered GI conditions and the abundance of Lactobacillus, Candidatus_Saccharimonas, and Lachnospiraceae_UCG_001 (Figure 5) in aged SAMP8 mice. The abundance of Lactobacillus was positively correlated with the intestine length and phagocytotic activity, while a negative correlation was observed between Lactobacillus and the intestinal permeability and concentration of the pro-inflammatory factors, TNF-α and MCP-1. Candidatus_Saccharimonas (which was significantly increased in the FPS23L group) was negatively correlated with the concentration of mucosal IgE and IgM. Lachnospiraceae_UCG_001 (which was reduced by the high dose of LPPS23) was negatively correlated with the intestine length and concentration of IgA, while it was positively correlated with the concentration of MCP-1 (Figure 5). The trends of correlation with the GI conditions were similar in Lactobacillus and Candidatus_Saccharimonas, but that of Lachnospiraceae_UCG_001 was different from the others (Figure 5).
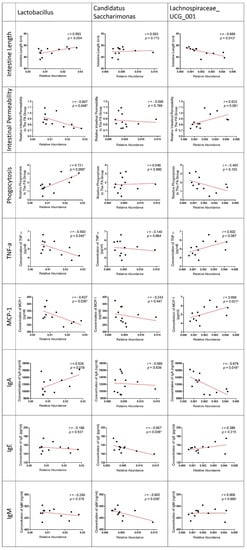
Figure 5.
Correlation of Lacticaseibacillus Paracasei PS23 (LPPS23)-modulated bacteria and gastrointestinal (GI) condition. Rank tests with Spearman’s correlation coefficient were used to assess correlations between bacteria (Lactobacillus, Candidatus_Saccharimonas, and Lachnospiraceae_UCG_ 001) and parameters of intestinal function (intestine length, intestinal permeability, and phagocytosis), pro-inflammatory factors (TNF-α and MCP-1), and immunoglobulin (IgA, IgE, and IgM). Asterisk (*) indicates p < 0.05. n = 12.
4. Discussion
The number of individuals with age-related disorders is increasing due to longer life spans. Our previous studies reported that LPPS23 attenuated the age-related cognitive impairment, muscle loss, and protein uptake decline in SAMP8 mice [14,15], and these age-related problems were associated with the change in GM composition [26,27]. Therefore, we attempted to understand the interaction of LPPS23 and GM in the aged SAMP8 mice. Although the probiotic did not influence the alpha diversity of the GM in the mice, high-dose LPPS23 (109 CFU/mL per day for 12 weeks) altered beta diversity and increased the abundance of Lactobacillus in the GM of the mice. Moreover, a negative correlation was observed between Lactobacillus and Erysipelotrichaceae. Furthermore, we identified three bacterial genera that were significantly regulated by LPPS23 in the aged mice: Lactobacillus, Candidatus_Saccharimonas, and Lachnospiraceae_UCG_001.
Due to the bidirectional effect of GM composition and GI function, the GI function was further investigated in SAMP8 mice. We observed severe intestinal dysfunction associated with inflammation and changes in IgA, IgE, and IgM concentrations in the intestinal mucosa of aged SAMP8 mice. These age-related alterations were alleviated by the administration of LPPS23 and were significantly correlated with the aforementioned LPPS23-modulated bacterial genera. Thus, we have provided the first demonstration of a relationship among probiotics, GM, and aging GI tracts.
Probiotics are thought to influence GI health by regulating the composition of the GM [28]. Several species and strains of Lactobacilli have been reported to change the population of bacteria in the GM, including Lactobacillus casei, Lactobacillus acidophilus, Lactobacillus helveticus, and Lactobacillus rhamnosus [29]. Similarly, we found that LPPS23 altered the beta-diversity of the GM and increased the abundance of Lactobacillus. Enrichment of Lactobacillus has been linked to healthy intestines and reduced inflammation [30,31]. Our results also demonstrated a positive correlation between the abundance of Lactobacillus and factors of GI health, such as a longer intestine, lower intestinal permeability, and higher phagocytosis. Moreover, the FPS23H mice exhibited the highest abundance of Lactocacillus and the best intestinal function among the aged mice groups. Taken together, the results suggest that LPPS23 attenuates the GI dysfunction in aged SAMP8 mice by regulating the GM in ways such as the enrichment of Lactobacillus.
GI dysfunction and inflammation induce each other. Therefore, several probiotics belonging to Lactobacillus have been reported to improve both GI function and inflammation. For example, Oliveira et al. (2018) found that Lactobacillus rhamnosus ST11 reduces intestinal inflammation and GI dysfunction in an adoptive transfer mouse model of experimental colitis [32]. Another study demonstrated that Lactobacillus acidophilus NCFM attenuates TNF-α-induced intestinal dysfunction and inflammation [33]. Our previous study also revealed that LPPS23 improved the intestinal function in protein uptake and prevented the increase in pro-inflammatory factors in the serum, muscle, and brain of aged SAMP8 mice [14,15]. Thus, this raises the question of whether Lactobacillus can also improve GI dysfunction and age-related inflammation in the elderly. The results of this study revealed that LPPS23 increased the abundance of Lactobacillus, which was negatively correlated with the concentration of pro-inflammatory factors TNF-α and MCP-1. Therefore, the results suggest LPPS23 could prevent not only intestinal function but also age-related inflammation by regulating the GM in the aged mice.
Since LPPS23 increased Lactobacillus that benefited intestinal function [34], we examined the correlations between Lactobacillus and other bacterial species. Only Erysipelotrichaceae was correlated negatively with Lactobacillus. Erysipelotrichaceae was enriched in a colorectal cancer mouse model [35] and in mice exhibiting colitis after treatment with dextran sodium sulphate [36]. Moreover, Thevaranjan et al. (2017) demonstrated that Erysipelotrichaceae was increased in aged mice [37]. Therefore, LPPS23 may cause increased Lactobacillus abundance resulting in decreased Erysipelotrichaceae abundance, which in turn may maintain or improve intestinal function and reduce inflammation in aged mice.
In addition to Lactobacillus, the relative abundances of other bacteria were changed in SAMP8 mice treated either with a high or low dose of LPPS23. All of the significantly changed bacteria (LDA score > 3.0 and p < 0.05) at the genus level were related to intestinal inflammation. Lachnospiraceae_UCG_001, a bacterium found to be increased in feces from unhealthy inflamed intestines [38], was reduced in the aged mice treated with the high dose of LPPS23. The relationship between Lachnospiraceae_UCG_001 and unhealthy intestines was also confirmed in the present study by the negative correlation between the abundance of Lachnospiraceae_UCG_001 and health factors, including longer intestines, a higher concentration of IgA, and a lower concentration of MCP-1. Furthermore, Candidatus_Saccharimonas, which was negatively correlated with intestinal inflammation [39,40], was more abundant in the FPS23L than FA mice. Our results also indicated a negative correlation between Candidatus_Saccharimonas and IgE, which is an immunoglobulin that increases because of immunosenescence and GI dysfunction resulting in age-related chronic inflammation [41,42,43]. Therefore, LPPS23 might improve the GI function in the aged SAMP8 mice through establishing an anti-inflammatory microenvironment in the intestine to prevent GI dysfunction caused by age-related inflammation. As such, we were able to determine potential bacterial biomarkers (i.e., Erysipelotrichaceae, Lachnospiraceae_UCG_001, and Candidatus_saccharimonas) of intestinal inflammation and GI function in the aged population.
Because only live LPPS23 was investigated in the present study, it raises the question of whether the effects of LPPS23 were caused when it was passing through or colonizing the intestine. According to the Liao et al. study, both dead and live LPPS23 showed similar abilities in the reduction of inflammation [16]. Therefore, establishing itself in the gut may not be a requirement for LPPS23 to achieve its effect in the present study. However, aged mice were used in the present study, while Liao et al. used young mice. Thus, further study is necessary to understand whether live LPPS23 is required to obtain the results of the present study.
The sample size of this study was referenced from previous studies regarding GM composition and GI function in mice [17,18]. Moreover, our results demonstrate that LPPS23 significantly improved GI function and changed the properties of three bacteria at the genus level, especially Lactobacillus (p < 0.05). The statistical powers were also calculated and were higher than 0.8 in the results of GI function and condition with a significant difference. Thus, the animal count should be reasonable and provide enough statistical significance in the present study. Furthermore, the anti-inflammatory effects of LPPS23 were demonstrated not only in the SAMP8 mice studies [14,15] but also in the C57BL/6J mice studies [16,44]. These studies also suggested that LPPS23 might influence gut microbiota and condition, because the effects of LPPS23 seemed to be implemented either via the gut–brain or gut–muscle axes. Since the present study showed that LPPS23 could prevent inflammation, regulate gut microbiota, and maintain intestine function, the results of the present study should be general and representative. However, it cannot be excluded that performing the study with more samples in each group might reveal more information. Therefore, increasing the sample number will be considered in further research.
SAMP8 mice are used most often for age-related neuropathological research, but more and more studies show that SAMP8 mice are also an optimal model for age-related disorders, such as muscle and mitochondria impairments [14]. The previous studies revealed acceleration in the appearance of senescence and age-related decrease in protein uptake in SAMP8 mice [14,15,45]. In the present study, 28-week-old SAMP8 mice showed several changes in phenomena that were related to aging intestines in rodents and humans. For example, increases in intestinal permeability and pro-inflammatory cytokines are observed in both aged rodents and humans [4,5,46,47]. Therefore, accelerated senescence seems to occur systemically in SAMP8 mice, which should be an appropriate model for intestinal age-related studies. It is unsurprising that SAMP8 mice were utilized for studying GI function by different scientists [48,49,50]. However, there were still some phenomena related to aging intestines in humans not assessed in the previous and present studies. Therefore, more investigations are necessary to evaluate how well SAMP8 mice can represent humans in aged-related GI function studies.
Although the results of the present study strongly supported the LPPS23 effectively modulating gut microbiota composition and improving gastrointestinal function in aged SAMP8 mice, there were limitations. First, the sample number was relatively small in the present study. The study was performed with well-trained investigators, inbred strain mice from the same batch, and in a well-controlled environment to lower the variation. Therefore, significant differences were revealed between the groups. However, to increase the samples, we may need more information to understand the effect of LPPS23 on microbiota and GI condition. Second, the effect of LPPS23 was observed in mice, not humans. Thus, a clinical trial will be necessary to confirm the function of LPPS23 in the future.
In conclusion, we have provided the first piece of evidence that LPPS23 can prevent age-related intestinal dysfunction and inflammation by modulating the GM. LPPS23 supplementation merits further research as a potential strategy to maintain GI health in elderly individuals.
Author Contributions
L.-H.C. designed the study, performed the experiments, and drafted the article. M.-F.W., C.-C.C., S.-Y.H. and C.-H.P. drafted the article. Y.-T.Y. and C.-H.H. performed the data analysis and edited the article. C.-H.C. performed the experiments and contributed to interpretation of the data. H.-Y.H. designed the study, supervised the project, and drafted the article. All authors discussed the results and contributed to the manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported in part by Ministry of Science and Technology, Taiwan (108-2321-B-038-007 and 108-2410-H-030-017).
Institutional Review Board Statement
The animal protocol was approved by the Institutional Animal Care and Use Committee of Shih Chien University (IACUC-10407).
Data Availability Statement
The data in this study are available on request from the corresponding author.
Acknowledgments
The authors would like to thank Lin-Ai Li and Hwai-Hsyan Ku for technical assistance.
Conflicts of Interest
The authors have no competing interests to declare.
References
- Santoro, A.; Zhao, J.C.; Wu, L.; Carru, C.; Biagi, E.; Franceschi, C. Microbiomes other than the gut: Inflammaging and age-related diseases. Semin. Immunopathol. 2020, 42, 589–605. [Google Scholar] [CrossRef]
- Sovran, B.; Hugenholtz, F.; Elderman, M.; Van Beek, A.A.; Graversen, K.; Huijskes, M.; Boekschoten, M.V.; Savelkoul, H.F.J.; De Vos, P.; Dekker, J.; et al. Age-associated impairment of the mucus barrier function is associated with profound changes in microbiota and immunity. Sci. Rep. 2019, 9, 1437. [Google Scholar] [CrossRef] [PubMed]
- Hall, K.E.; Proctor, D.D.; Fisher, L.; Rose, S. American gastroenterological association future trends committee report: Effects of aging of the population on gastroenterology practice, education, and research. Gastroenterology 2005, 129, 1305–1338. [Google Scholar] [CrossRef]
- Hollander, D.; Tarnawski, H. Aging-associated increase in intestinal absorption of macromolecules. Gerontology 1985, 31, 133–137. [Google Scholar] [CrossRef] [PubMed]
- Annaert, P.; Brouwers, J.; Bijnens, A.; Lammert, F.; Tack, J.; Augustijns, P. Ex vivo permeability experiments in excised rat intestinal tissue and in vitro solubility measurements in aspirated human intestinal fluids support age-dependent oral drug absorption. Eur. J. Pharm Sci. 2010, 39, 15–22. [Google Scholar] [CrossRef]
- Drozdowski, L.; Thomson, A.B. Aging and the intestine. World J. Gastroenterol. 2006, 12, 7578–7584. [Google Scholar] [CrossRef] [PubMed]
- Biagi, E.; Nylund, L.; Candela, M.; Ostan, R.; Bucci, L.; Pini, E.; Nikkila, J.; Monti, D.; Satokari, R.; Franceschi, C.; et al. Through ageing, and beyond: Gut microbiota and inflammatory status in seniors and centenarians. PLoS ONE 2010, 5, e10667. [Google Scholar] [CrossRef]
- Walrath, T.; Dyamenahalli, K.U.; Hulsebus, H.J.; McCullough, R.L.; Idrovo, J.P.; Boe, D.M.; McMahan, R.H.; Kovacs, E.J. Age-related changes in intestinal immunity and the microbiome. J. Leukocyte Biol. 2020, 1–17. [Google Scholar] [CrossRef]
- An, R.; Wilms, E.; Masclee, A.A.M.; Smidt, H.; Zoetendal, E.G.; Jonkers, D. Age-dependent changes in gi physiology and microbiota: Time to reconsider? Gut 2018, 67, 2213–2222. [Google Scholar] [CrossRef] [PubMed]
- Franceschi, C.; Garagnani, P.; Parini, P.; Giuliani, C.; Santoro, A. Inflammaging: A new immune-metabolic viewpoint for age-related diseases. Nat. Rev. Endocrinol. 2018, 14, 576–590. [Google Scholar] [CrossRef]
- Holmes, E.; Li, J.V.; Marchesi, J.R.; Nicholson, J.K. Gut microbiota composition and activity in relation to host metabolic phenotype and disease risk. Cell Metab. 2012, 16, 559–564. [Google Scholar] [CrossRef]
- Malaguarnera, G.; Leggio, F.; Vacante, M.; Motta, M.; Giordano, M.; Biondi, A.; Basile, F.; Mastrojeni, S.; Mistretta, A.; Malaguarnera, M.; et al. Probiotics in the gastrointestinal diseases of the elderly. J. Nutr. Health Aging 2012, 16, 402–410. [Google Scholar] [CrossRef]
- Lomax, A.R.; Calder, P.C. Prebiotics, immune function, infection and inflammation: A review of the evidence. Br. J. Nutr. 2009, 101, 633–658. [Google Scholar] [CrossRef]
- Chen, L.H.; Huang, S.Y.; Huang, K.C.; Hsu, C.C.; Yang, K.C.; Li, L.A.; Chan, C.H.; Huang, H.Y. Lactobacillus paracasei ps23 decelerated age-related muscle loss by ensuring mitochondrial function in samp8 mice. Aging 2019, 11, 756–770. [Google Scholar] [CrossRef]
- Huang, S.Y.; Chen, L.H.; Wang, M.F.; Hsu, C.C.; Chan, C.H.; Li, J.X.; Huang, H.Y. Lactobacillus paracasei ps23 delays progression of age-related cognitive decline in senescence accelerated mouse prone 8 (samp8) mice. Nutrients 2018, 10, 894. [Google Scholar] [CrossRef]
- Liao, J.F.; Hsu, C.C.; Chou, G.T.; Hsu, J.S.; Liong, M.T.; Tsai, Y.C. Lactobacillus paracasei ps23 reduced early-life stress abnormalities in maternal separation mouse model. Benef. Microbes 2019, 10, 425–436. [Google Scholar] [CrossRef]
- Desai, M.S.; Seekatz, A.M.; Koropatkin, N.M.; Kamada, N.; Hickey, C.A.; Wolter, M.; Pudlo, N.A.; Kitamoto, S.; Terrapon, N.; Muller, A.; et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 2016, 167, 1339–1353.e1321. [Google Scholar] [CrossRef] [PubMed]
- Lam, Y.Y.; Ha, C.W.Y.; Hoffmann, J.M.A.; Oscarsson, J.; Dinudom, A.; Mather, T.J.; Cook, D.I.; Hunt, N.H.; Caterson, I.D.; Holmes, A.J.; et al. Effects of dietary fat profile on gut permeability and microbiota and their relationships with metabolic changes in mice. Obesity 2015, 23, 1429–1439. [Google Scholar] [CrossRef]
- Edgar, R.C. Uparse: Highly accurate otu sequences from microbial amplicon reads. Nat. Methods 2013, 10, 996–998. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. Qiime allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef]
- Haas, B.J.; Gevers, D.; Earl, A.M.; Feldgarden, M.; Ward, D.V.; Giannoukos, G.; Ciulla, D.; Tabbaa, D.; Highlander, S.K.; Sodergren, E.; et al. Chimeric 16s rrna sequence formation and detection in sanger and 454-pyrosequenced pcr amplicons. Genome Res. 2011, 21, 494–504. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glockner, F.O. The silva ribosomal rna gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef]
- Afgan, E.; Baker, D.; Batut, B.; van den Beek, M.; Bouvier, D.; Cech, M.; Chilton, J.; Clements, D.; Coraor, N.; Gruning, B.A.; et al. The galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 2018, 46, W537–W544. [Google Scholar] [CrossRef]
- Ma, X.W.; Hu, Y.C.; Li, X.; Zheng, X.T.; Wang, Y.T.; Zhang, J.M.; Fu, C.M.; Geng, F.N. Periplaneta americana ameliorates dextran sulfate sodium-induced ulcerative colitis in rats by keap1/nrf-2 activation, intestinal barrier function, and gut microbiota regulation. Front. Pharmacol. 2018, 9, 944. [Google Scholar] [CrossRef]
- Nagpal, R.; Mainali, R.; Ahmadi, S.; Wang, S.; Singh, R.; Kavanagh, K.; Kitzman, D.W.; Kushugulova, A.; Marotta, F.; Yadav, H. Gut microbiome and aging: Physiological and mechanistic insights. Nutr. Healthy Aging 2018, 4, 267–285. [Google Scholar] [CrossRef]
- Prokopidis, K.; Cervo, M.M.; Gandham, A.; Scott, D. Impact of protein intake in older adults with sarcopenia and obesity: A gut microbiota perspective. Nutrients 2020, 12, 2285. [Google Scholar] [CrossRef]
- Ticinesi, A.; Tana, C.; Nouvenne, A.; Prati, B.; Lauretani, F.; Meschi, T. Gut microbiota, cognitive frailty and dementia in older individuals: A systematic review. Clin. Interv. Aging 2018, 13, 1497–1511. [Google Scholar] [CrossRef]
- Verdu, E.F.; Bercik, P.; Collins, S.M. Effect of probiotics on gastrointestinal function: Evidence from animal models. Therap. Adv. Gastroenterol. 2009, 2, 31–35. [Google Scholar] [CrossRef]
- Azad, M.A.; Sarker, M.; Li, T.J.; Yin, J. Probiotic species in the modulation of gut microbiota: An overview. Biomed. Res. Int. 2018, 2018, 9478630. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.N.; Wu, H.; Chen, Y.Z.; Chen, Y.J.; Shen, X.Z.; Liu, T.T. Altered molecular signature of intestinal microbiota in irritable bowel syndrome patients compared with healthy controls: A systematic review and meta-analysis. Dig. Liver Dis. 2017, 49, 331–337. [Google Scholar] [CrossRef] [PubMed]
- Ganji-Arjenaki, M.; Rafieian-Kopaei, M. Probiotics are a good choice in remission of inflammatory bowel diseases: A meta analysis and systematic review. J. Cell Physiol. 2018, 233, 2091–2103. [Google Scholar] [CrossRef]
- Oliveira, M.; Bosco, N.; Perruisseau, G.; Nicolas, J.; Segura-Roggero, I.; Duboux, S.; Briand, M.; Blum, S.; Benyacoub, J. Lactobacillus paracasei reduces intestinal inflammation in adoptive transfer mouse model of experimental colitis. Clin. Dev. Immunol. 2011, 2011, 807483. [Google Scholar] [CrossRef][Green Version]
- Wang, H.; Zhang, Q.; Niu, Y.; Zhang, X.; Lu, R. Surface-layer protein from lactobacillus acidophilus ncfm attenuates tumor necrosis factor-alpha-induced intestinal barrier dysfunction and inflammation. Int. J. Biol. Macromol. 2019, 136, 27–34. [Google Scholar] [CrossRef]
- Vaughan, E.E.; Mollet, B.; deVos, W.M. Functionality of probiotics and intestinal lactobacilli: Light in the intestinal tract tunnel. Curr. Opin. Biotechnol. 1999, 10, 505–510. [Google Scholar] [CrossRef]
- Zhu, Q.C.; Jin, Z.M.; Wu, W.; Gao, R.Y.; Guo, B.M.; Gao, Z.G.; Yang, Y.Z.; Qin, H.L. Analysis of the intestinal lumen microbiota in an animal model of colorectal cancer. PLoS ONE 2014, 9, e90849. [Google Scholar] [CrossRef]
- Palm, N.W.; de Zoete, M.R.; Cullen, T.W.; Barry, N.A.; Stefanowski, J.; Hao, L.M.; Degnan, P.H.; Hu, J.Z.; Peter, I.; Zhang, W.; et al. Immunoglobulin a coating identifies colitogenic bacteria in inflammatory bowel disease. Cell 2014, 158, 1000–1010. [Google Scholar] [CrossRef] [PubMed]
- Thevaranjan, N.; Puchta, A.; Schulz, C.; Naidoo, A.; Szamosi, J.C.; Verschoor, C.P.; Loukov, D.; Schenck, L.P.; Jury, J.; Foley, K.P.; et al. Age-associated microbial dysbiosis promotes intestinal permeability, systemic inflammation, and macrophage dysfunction. Cell Host Microbe 2017, 21, 455–456. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, X.L.; Zhou, M.; Kang, C.; Lang, H.D.; Chen, M.T.; Hui, S.C.; Wang, B.; Mi, M.T. Crosstalk between gut microbiota and sirtuin-3 in colonic inflammation and tumorigenesis. Exp. Mol. Med. 2018, 50, 21. [Google Scholar] [CrossRef]
- Li, Q.Q.; Chen, C.; Liu, C.P.; Sun, W.N.; Liu, X.H.; Ci, Y.F.; Song, Y. The effects of cellulose on aom/dss-treated c57bl/6 colorectal cancer mice by changing intestinal flora composition and inflammatory factors. Nutr. Cancer 2020, 73, 502–513. [Google Scholar] [CrossRef]
- Li, H.L.; Lu, L.; Wang, X.S.; Qin, L.Y.; Wang, P.; Qiu, S.P.; Wu, H.; Huang, F.; Zhang, B.B.; Shi, H.L.; et al. Alteration of gut microbiota and inflammatory cytokine/chemokine profiles in 5-fluorouracil induced intestinal mucositis. Front. Cell. Infect. Microbiol. 2017, 7, 455. [Google Scholar] [CrossRef]
- Yu, L.C. Intestinal epithelial barrier dysfunction in food hypersensitivity. J. Allergy 2012, 2012, 596081. [Google Scholar] [CrossRef] [PubMed]
- De Martinis, M.; Sirufo, M.M.; Viscido, A.; Ginaldi, L. Food Allergies and Ageing. Int. J. Mol. Sci. 2019, 20, 5580. [Google Scholar] [CrossRef] [PubMed]
- Navines-Ferrer, A.; Serrano-Candelas, E.; Molina-Molina, G.J.; Martin, M. Ige-related chronic diseases and anti-ige-based treatments. J. Immunol. Res. 2016, 2016, 8163803. [Google Scholar] [CrossRef]
- Wei, C.L.; Wang, S.; Yen, J.T.; Cheng, Y.F.; Liao, C.L.; Hsu, C.C.; Wu, C.C.; Tsai, Y.C. Antidepressant-like activities of live and heat-killed lactobacillus paracasei ps23 in chronic corticosterone-treated mice and possible mechanisms. Brain Res. 2019, 1711, 202–213. [Google Scholar] [CrossRef] [PubMed]
- Takeda, T.; Hosokawa, M.; Takeshita, S.; Irino, M.; Higuchi, K.; Matsushita, T.; Tomita, Y.; Yashuhira, K.; Hamamoto, H.; Shimizu, K.; et al. A new murine model of accelerated senescence. Mech. Ageing Dev. 1981, 17, 183–194. [Google Scholar] [CrossRef]
- Steegenga, W.T.; de Wit, N.J.; Boekschoten, M.V.; Ijssennagger, N.; Lute, C.; Keshtkar, S.; Bromhaar, M.M.; Kampman, E.; de Groot, L.C.; Muller, M. Structural, functional and molecular analysis of the effects of aging in the small intestine and colon of c57bl/6j mice. BMC Med. Genom. 2012, 5, 38. [Google Scholar] [CrossRef]
- Man, A.L.; Bertelli, E.; Rentini, S.; Regoli, M.; Briars, G.; Marini, M.; Watson, A.J.; Nicoletti, C. Age-associated modifications of intestinal permeability and innate immunity in human small intestine. Clin. Sci. 2015, 129, 515–527. [Google Scholar] [CrossRef]
- Yamamoto, K.; Kushida, M.; Tsuduki, T. The effect of dietary lipid on gut microbiota in a senescence-accelerated prone mouse model (samp8). Biogerontology 2018, 19, 367–383. [Google Scholar] [CrossRef] [PubMed]
- Ben Othman, S.; Ido, K.; Masuda, R.; Gotoh, S.; Hosoda-Yabe, R.; Kitaguchi, K.; Yabe, T. Senescence-accelerated mouse prone 8 mice exhibit specific morphological changes in the small intestine during senescence and after pectin supplemented diet. Exp. Gerontol. 2020, 142, 111099. [Google Scholar] [CrossRef]
- Xie, Z.; Lu, H.; Yang, S.; Zeng, Y.; Li, W.; Wang, L.; Luo, G.; Fang, F.; Zeng, T.; Cheng, W. Salidroside attenuates cognitive dysfunction in senescence-accelerated mouse prone 8 (samp8) mice and modulates inflammation of the gut-brain axis. Front. Pharmacol. 2020, 11, 568423. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).