Vitamin E Deficiency Disrupts Gene Expression Networks during Zebrafish Development
Abstract
1. Introduction
2. Methods
2.1. Zebrafish Husbandry
2.2. RNA Extraction and Sequencing
2.3. Data Deposition
2.4. Data Processing and Statistical Analysis
2.5. Gene Ontology and Network Enrichment
2.6. Western Blotting
3. Results
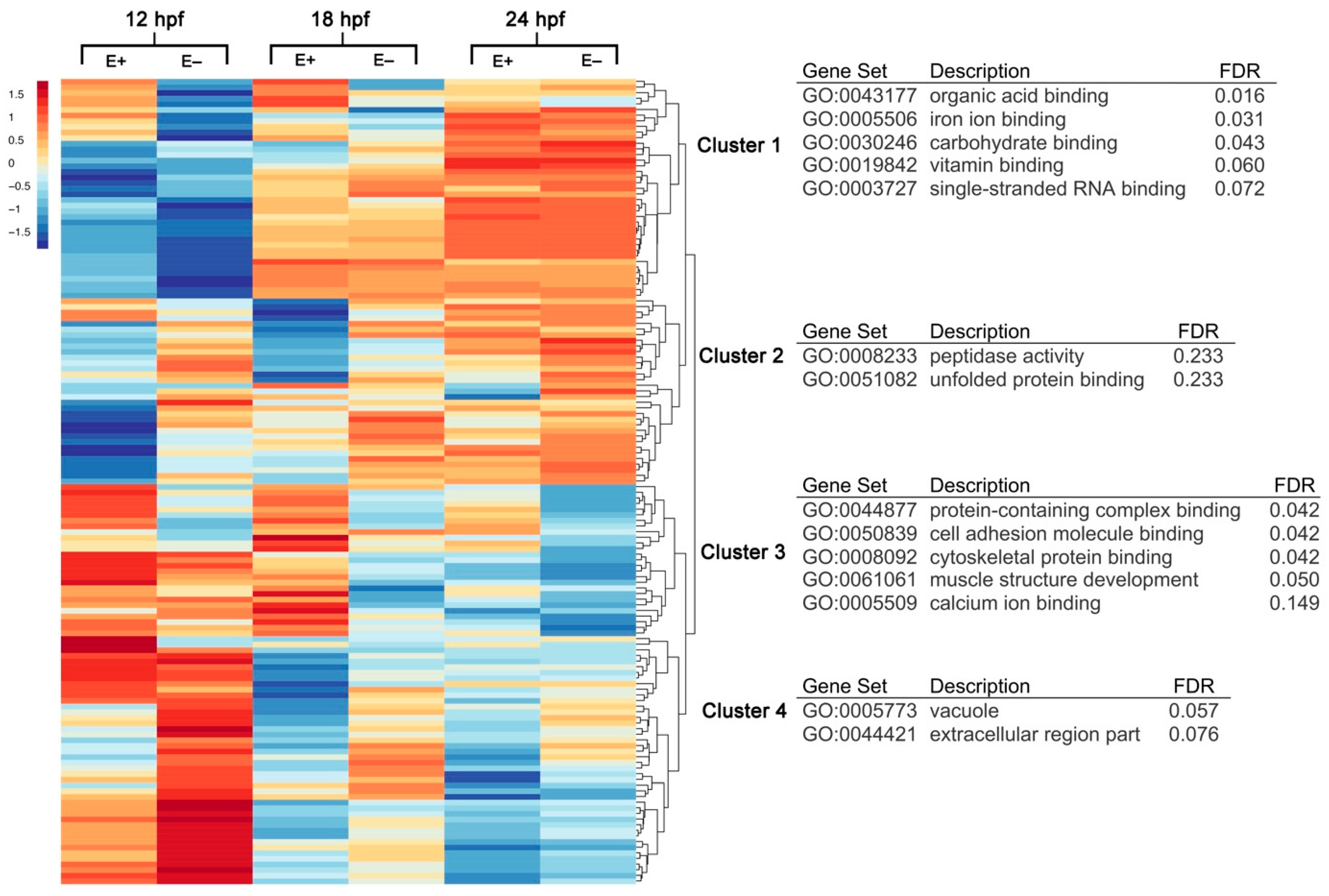
3.1. Hierarchical Clustering and Gene Annotation
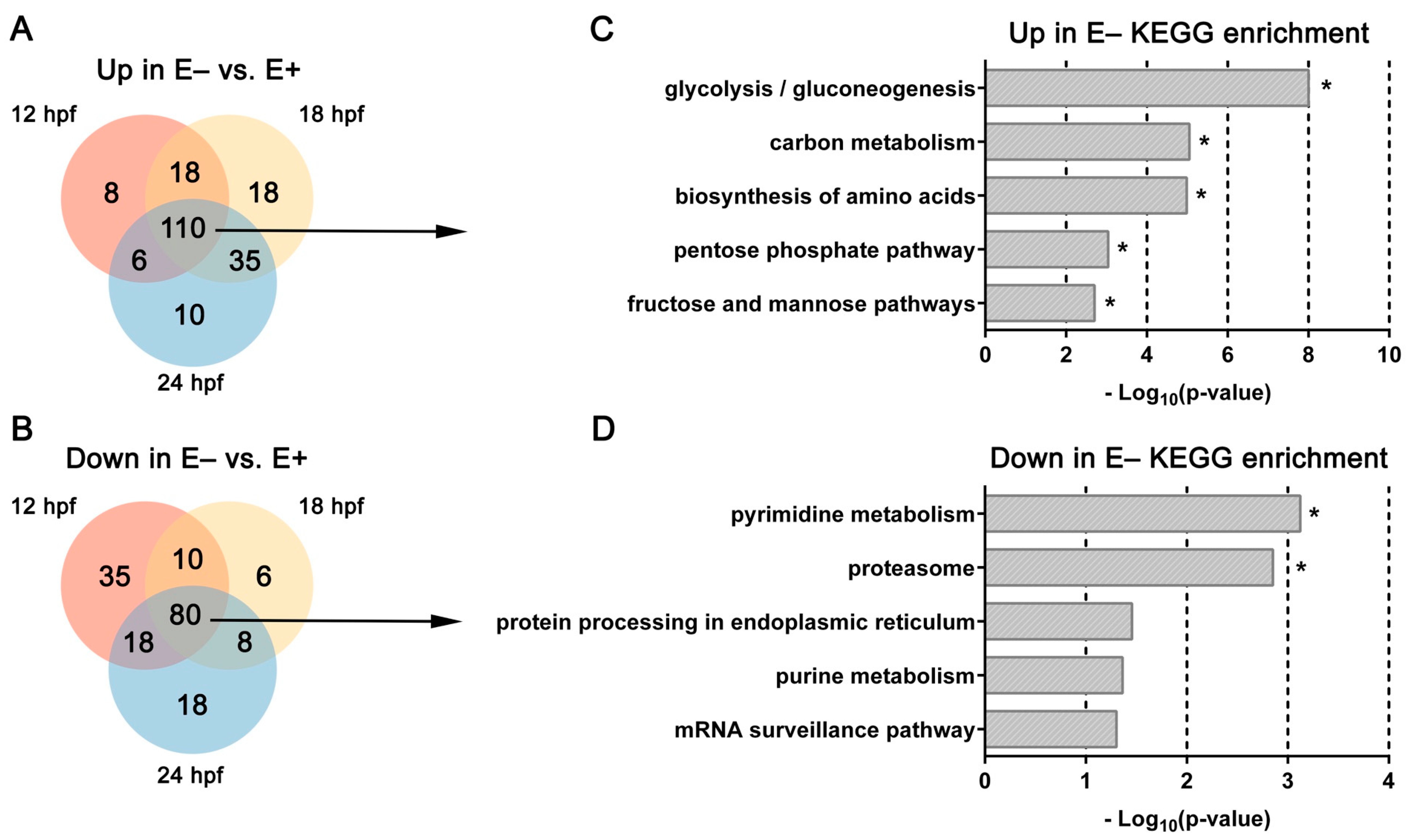
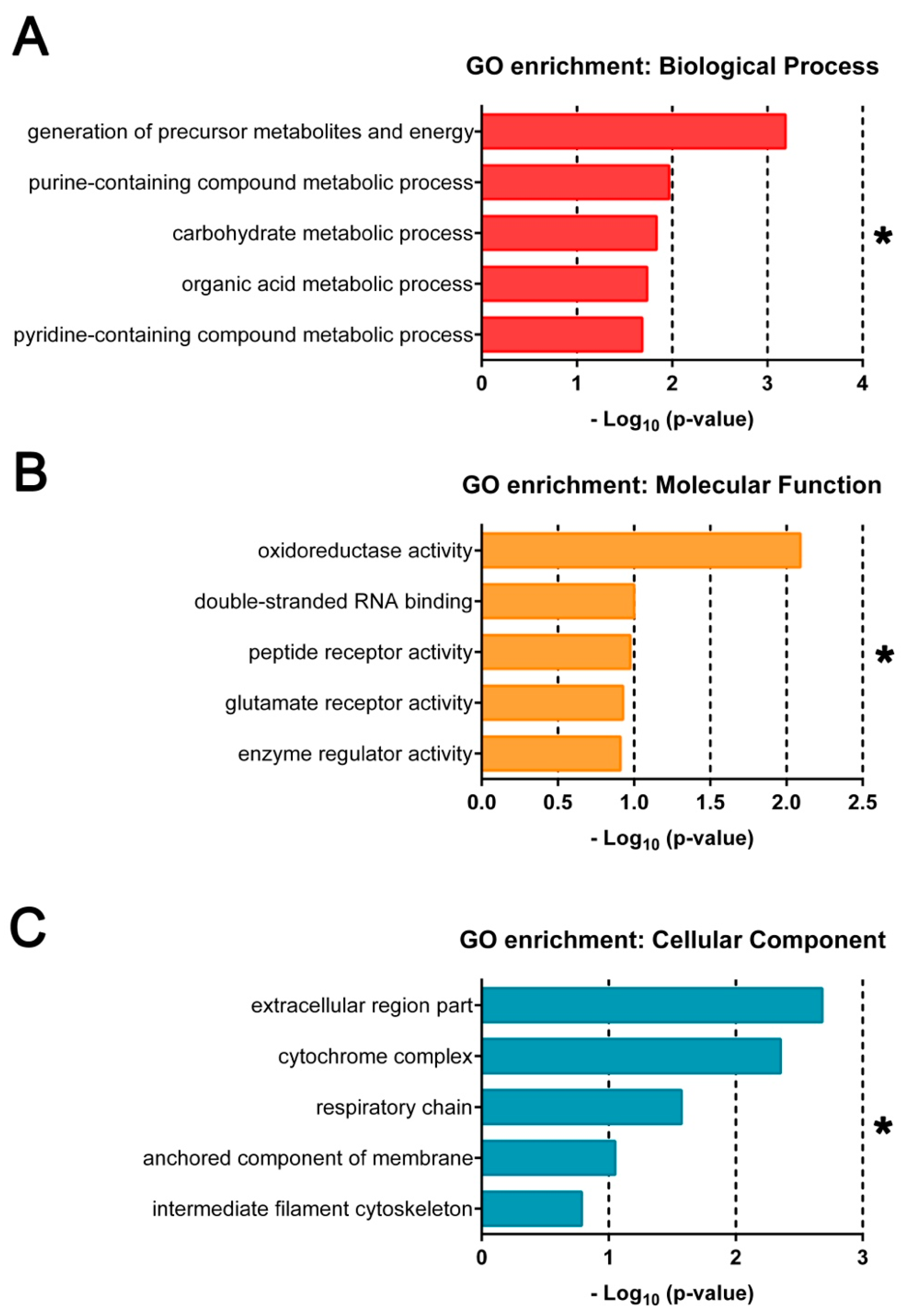
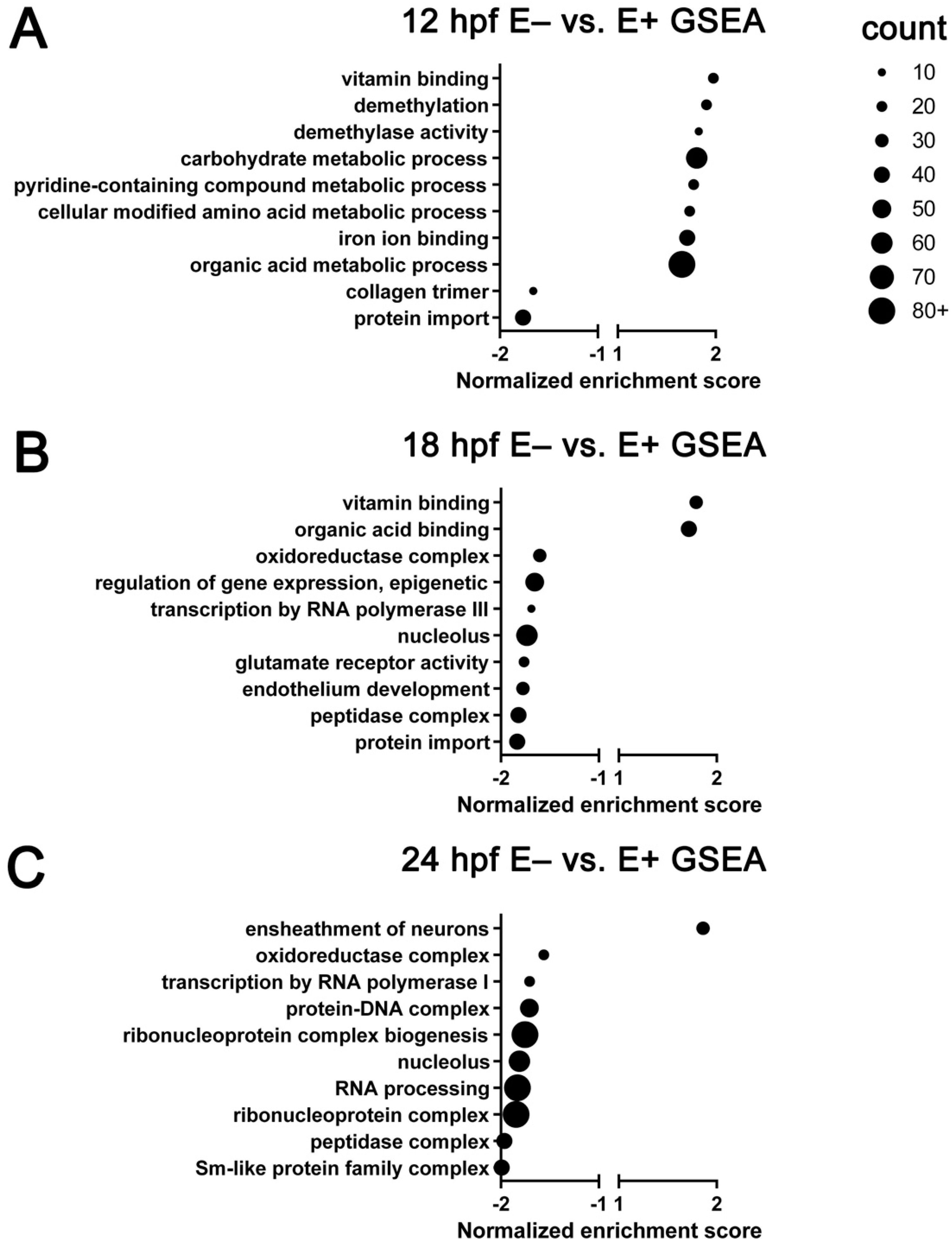
3.2. Top Differentially Expressed Genes in E– Embryos
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Traber, M.G.; Atkinson, J. Vitamin E, antioxidant and nothing more. Free Radic. Biol. Med. 2007, 43, 4–15. [Google Scholar] [CrossRef]
- Zingg, J.M. Vitamin E: Regulatory Role on Signal Transduction. IUBMB Life 2019, 71, 456–478. [Google Scholar] [CrossRef]
- Mene-Saffrane, L.; DellaPenna, D. Biosynthesis, regulation and functions of tocochromanols in plants. Plant Physiol. Biochem. 2010, 48, 301–309. [Google Scholar] [CrossRef]
- Evans, H.M.; Bishop, K.S. On the Existence of a Hitherto Unrecognized Dietary Factor Essential for Reproduction. Science 1922, 56, 650–651. [Google Scholar] [CrossRef] [PubMed]
- Steele, C.E.; Jeffery, E.H.; Diplock, A.T. The effect of vitamin E and synthetic antioxidants on the growth in vitro of explanted rat embryos. J. Reprod. Fertil. 1974, 38, 115–123. [Google Scholar] [CrossRef] [PubMed]
- Jishage, K.; Tachibe, T.; Ito, T.; Shibata, N.; Suzuki, S.; Mori, T.; Hani, T.; Arai, H.; Suzuki, H. Vitamin E is essential for mouse placentation but not for embryonic development itself. Biol. Reprod. 2005, 73, 983–987. [Google Scholar] [CrossRef]
- Altman, P.L.; Dittmer, D.S. (Eds.) Growth, including reproduction and morphological development; Federation of American Societies for Experimental Biology: Washington, DC, USA, 1962; Volume xiv, p. 608. [Google Scholar]
- Kimmel, C.B.; Ballard, W.W.; Kimmel, S.R.; Ullmann, B.; Schilling, T.F. Stages of embryonic development of the zebrafish. Dev. Dyn. 1995, 203, 253–310. [Google Scholar] [CrossRef] [PubMed]
- Miller, G.W.; Ulatowski, L.; Labut, E.M.; Lebold, K.M.; Manor, D.; Atkinson, J.; Barton, C.L.; Tanguay, R.L.; Traber, M.G. The alpha-tocopherol transfer protein is essential for vertebrate embryogenesis. PLoS ONE 2012, 7, e47402. [Google Scholar] [CrossRef]
- Miller, G.W.; Labut, E.M.; Lebold, K.M.; Floeter, A.; Tanguay, R.L.; Traber, M.G. Zebrafish (Danio rerio) fed vitamin E-deficient diets produce embryos with increased morphologic abnormalities and mortality. J. Nutr. Biochem. 2012, 23, 478–486. [Google Scholar] [CrossRef]
- McDougall, M.; Choi, J.; Kim, H.K.; Bobe, G.; Stevens, J.F.; Cadenas, E.; Tanguay, R.; Traber, M.G. Lethal dysregulation of energy metabolism during embryonic vitamin E deficiency. Free Radic. Biol. Med. 2017, 104, 324–332. [Google Scholar] [CrossRef]
- Head, B.; La Du, J.; Tanguay, R.L.; Kioussi, C.; Traber, M.G. Vitamin E is necessary for zebrafish nervous system development. Sci. Rep. 2020, 10, 15028. [Google Scholar] [CrossRef] [PubMed]
- Santander, N.; Lizama, C.; Parga, M.J.; Quiroz, A.; Perez, D.; Echeverria, G.; Ulloa, L.; Palma, V.; Rigotti, A.; Busso, D. Deficient Vitamin E Uptake During Development Impairs Neural Tube Closure in Mice Lacking Lipoprotein Receptor SR-BI. Sci. Rep.-Uk 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Homanics, G.E.; Maeda, N.; Traber, M.G.; Kayden, H.J.; Dehart, D.B.; Sulik, K.K. Exencephaly and hydrocephaly in mice with targeted modification of the apolipoprotein B (Apob) gene. Teratology 1995, 51, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Ulatowski, L.; Parker, R.; Warrier, G.; Sultana, R.; Butterfield, D.A.; Manor, D. Vitamin E is essential for Purkinje neuron integrity. Neuroscience 2014, 260, 120–129. [Google Scholar] [CrossRef] [PubMed]
- Yokota, T.; Igarashi, K.; Uchihara, T.; Jishage, K.; Tomita, H.; Inaba, A.; Li, Y.; Arita, M.; Suzuki, H.; Mizusawa, H.; et al. Delayed-onset ataxia in mice lacking alpha -tocopherol transfer protein: Model for neuronal degeneration caused by chronic oxidative stress. Proc. Natl. Acad. Sci. USA 2001, 98, 15185–15190. [Google Scholar] [CrossRef]
- Tixier, V.; Bataille, L.; Etard, C.; Jagla, T.; Weger, M.; Daponte, J.P.; Strahle, U.; Dickmeis, T.; Jagla, K. Glycolysis supports embryonic muscle growth by promoting myoblast fusion. Proc. Natl. Acad. Sci. USA 2013, 110, 18982–18987. [Google Scholar] [CrossRef]
- McDougall, M.; Choi, J.; Kim, H.K.; Bobe, G.; Stevens, J.F.; Cadenas, E.; Tanguay, R.; Traber, M.G. Lipid quantitation and metabolomics data from vitamin E-deficient and -sufficient zebrafish embryos from 0 to 120 hours-post-fertilization. Data Brief. 2017, 11, 432–441. [Google Scholar] [CrossRef]
- Zhang, J.; Head, B.; Leonard, S.W.; Choi, J.; Tanguay, R.L.; Traber, M.G. Vitamin E deficiency dysregulates thiols, amino acids and related molecules during zebrafish embryogenesis. Redox. Biol. 2020, 38, 101784. [Google Scholar] [CrossRef]
- Lee, M.S.; Bonner, J.R.; Bernard, D.J.; Sanchez, E.L.; Sause, E.T.; Prentice, R.R.; Burgess, S.M.; Brody, L.C. Disruption of the folate pathway in zebrafish causes developmental defects. BMC Dev. Biol. 2012, 12, 12. [Google Scholar] [CrossRef]
- White, R.J.; Collins, J.E.; Sealy, I.M.; Wali, N.; Dooley, C.M.; Digby, Z.; Stemple, D.L.; Murphy, D.N.; Billis, K.; Hourlier, T.; et al. A high-resolution mRNA expression time course of embryonic development in zebrafish. Elife 2017, 6. [Google Scholar] [CrossRef]
- Shimobayashi, M.; Hall, M.N. Multiple amino acid sensing inputs to mTORC1. Cell Res. 2016, 26, 7–20. [Google Scholar] [CrossRef]
- Garza-Lombo, C.; Petrosyan, P.; Tapia-Rodriguez, M.; Valdovinos-Flores, C.; Gonsebatt, M.E. Systemic L-buthionine-S-R-sulfoximine administration modulates glutathione homeostasis via NGF/TrkA and mTOR signaling in the cerebellum. Neurochem. Int. 2018, 121, 8–18. [Google Scholar] [CrossRef] [PubMed]
- Kearns, C.A.; Ravanelli, A.M.; Cooper, K.; Appel, B. Fbxw7 Limits Myelination by Inhibiting mTOR Signaling. J. Neurosci. 2015, 35, 14861–14871. [Google Scholar] [CrossRef] [PubMed]
- Milanese, C.; Bombardieri, C.R.; Sepe, S.; Barnhoorn, S.; Payan-Gomez, C.; Caruso, D.; Audano, M.; Pedretti, S.; Vermeij, W.P.; Brandt, R.M.C.; et al. DNA damage and transcription stress cause ATP-mediated redesign of metabolism and potentiation of anti-oxidant buffering. Nat. Commun. 2019, 10, 4887. [Google Scholar] [CrossRef] [PubMed]
- Podda, M.; Weber, C.; Traber, M.G.; Packer, L. Simultaneous determination of tissue tocopherols, tocotrienols, ubiquinols, and ubiquinones. J. Lipid Res. 1996, 37, 893–901. [Google Scholar] [CrossRef]
- Liao, Y.; Wang, J.; Jaehnig, E.J.; Shi, Z.; Zhang, B. WebGestalt 2019: Gene set analysis toolkit with revamped UIs and APIs. Nucleic. Acids Res. 2019, 47, W199–W205. [Google Scholar] [CrossRef] [PubMed]
- Ruzicka, L.; Howe, D.G.; Ramachandran, S.; Toro, S.; Van Slyke, C.E.; Bradford, Y.M.; Eagle, A.; Fashena, D.; Frazer, K.; Kalita, P.; et al. The Zebrafish Information Network: New support for non-coding genes, richer Gene Ontology annotations and the Alliance of Genome Resources. Nucleic. Acids Res. 2019, 47, D867–D873. [Google Scholar] [CrossRef] [PubMed]
- Pang, Z.; Chong, J.; Li, S.; Xia, J. MetaboAnalystR 3.0: Toward an Optimized Workflow for Global Metabolomics. Metabolites 2020, 10, 186. [Google Scholar] [CrossRef] [PubMed]
- Scheldeman, C.; Mills, J.D.; Siekierska, A.; Serra, I.; Copmans, D.; Iyer, A.M.; Whalley, B.J.; Maes, J.; Jansen, A.C.; Lagae, L.; et al. mTOR-related neuropathology in mutant tsc2 zebrafish: Phenotypic, transcriptomic and pharmacological analysis. Neurobiol. Dis. 2017, 108, 225–237. [Google Scholar] [CrossRef]
- Love, C.E.; Prince, V.E. Expression and retinoic acid regulation of the zebrafish nr2f orphan nuclear receptor genes. Dev. Dyn. 2012, 241, 1603–1615. [Google Scholar] [CrossRef]
- Thisse, C.; Thisse, B. High-resolution in situ hybridization to whole-mount zebrafish embryos. Nat. Protoc. 2008, 3, 59–69. [Google Scholar] [CrossRef] [PubMed]
- Tanimura, T.; Shepard, T.H. Glucose metabolism by rat embryos in vitro. Proc. Soc. Exp. Biol. Med. 1970, 135, 51–54. [Google Scholar] [CrossRef] [PubMed]
- Miller, G.W.; Truong, L.; Barton, C.L.; Labut, E.M.; Lebold, K.M.; Traber, M.G.; Tanguay, R.L. The influences of parental diet and vitamin E intake on the embryonic zebrafish transcriptome. Comp. Biochem. Physiol. Part D Genomics. Proteomics. 2014, 10, 22–29. [Google Scholar] [CrossRef] [PubMed]
- Motorykin, I.; Traber, M.G.; Tanguay, R.L.; Maier, C.S. Proteome-driven elucidation of adaptive responses to combined vitamin E and C deficiency in zebrafish. J. Proteome. Res. 2014, 13, 1647–1656. [Google Scholar] [CrossRef] [PubMed]
- Moazzami, A.A.; Andersson, R.; Kamal-Eldin, A. Changes in the metabolic profile of rat liver after alpha-tocopherol deficiency as revealed by metabolomics analysis. NMR Biomed. 2011, 24, 499–505. [Google Scholar] [CrossRef] [PubMed]
- Lebold, K.M.; Jump, D.B.; Miller, G.W.; Wright, C.L.; Labut, E.M.; Barton, C.L.; Tanguay, R.L.; Traber, M.G. Vitamin E deficiency decreases long-chain PUFA in zebrafish (Danio rerio). J. Nutr. 2011, 141, 2113–2118. [Google Scholar] [CrossRef]
- Lebold, K.M.; Lohr, C.V.; Barton, C.L.; Miller, G.W.; Labut, E.M.; Tanguay, R.L.; Traber, M.G. Chronic vitamin E deficiency promotes vitamin C deficiency in zebrafish leading to degenerative myopathy and impaired swimming behavior. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 2013, 157, 382–389. [Google Scholar] [CrossRef]
- Zhang, C.; Skamagki, M.; Liu, Z.; Ananthanarayanan, A.; Zhao, R.; Li, H.; Kim, K. Biological Significance of the Suppression of Oxidative Phosphorylation in Induced Pluripotent Stem Cells. Cell Rep 2017, 21, 2058–2065. [Google Scholar] [CrossRef]
- Kraus, P.; V, S.; Yu, H.B.; Xing, X.; Lim, S.L.; Adler, T.; Pimentel, J.A.; Becker, L.; Bohla, A.; Garrett, L.; et al. Pleiotropic functions for transcription factor zscan10. PLoS ONE 2014, 9, e104568. [Google Scholar] [CrossRef]
- Fishwick, K.J.; Li, R.A.; Halley, P.; Deng, P.; Storey, K.G. Initiation of neuronal differentiation requires PI3-kinase/TOR signalling in the vertebrate neural tube. Dev. Biol. 2010, 338, 215–225. [Google Scholar] [CrossRef]
- Kadoya, M.; Sasai, N. Negative Regulation of mTOR Signaling Restricts Cell Proliferation in the Floor Plate. Front Neurosci. 2019, 13, 1022. [Google Scholar] [CrossRef] [PubMed]
- Makky, K.; Tekiela, J.; Mayer, A.N. Target of rapamycin (TOR) signaling controls epithelial morphogenesis in the vertebrate intestine. Dev. Biol. 2007, 303, 501–513. [Google Scholar] [CrossRef] [PubMed]
- Mathews, E.S.; Appel, B. Cholesterol Biosynthesis Supports Myelin Gene Expression and Axon Ensheathment through Modulation of P13K/Akt/mTor Signaling. J. Neurosci. 2016, 36, 7628–7639. [Google Scholar] [CrossRef] [PubMed]
- Blauth, K.; Banerjee, S.; Bhat, M.A. Axonal ensheathment and intercellular barrier formation in Drosophila. Int. Rev. Cell Mol. Biol. 2010, 283, 93–128. [Google Scholar] [CrossRef]
- Woodhoo, A.; Sommer, L. Development of the Schwann cell lineage: From the neural crest to the myelinated nerve. Glia 2008, 56, 1481–1490. [Google Scholar] [CrossRef]
- Barraud, P.; Seferiadis, A.A.; Tyson, L.D.; Zwart, M.F.; Szabo-Rogers, H.L.; Ruhrberg, C.; Liu, K.J.; Baker, C.V. Neural crest origin of olfactory ensheathing glia. Proc. Natl. Acad. Sci. USA 2010, 107, 21040–21045. [Google Scholar] [CrossRef]
- McGraw, H.F.; Snelson, C.D.; Prendergast, A.; Suli, A.; Raible, D.W. Postembryonic neuronal addition in zebrafish dorsal root ganglia is regulated by Notch signaling. Neural. Dev. 2012, 7, 23. [Google Scholar] [CrossRef]
- Marchi, D.; Santhakumar, K.; Markham, E.; Li, N.; Storbeck, K.H.; Krone, N.; Cunliffe, V.T.; van Eeden, F.J.M. Bidirectional crosstalk between Hypoxia-Inducible Factor and glucocorticoid signalling in zebrafish larvae. PLoS Genet. 2020, 16, e1008757. [Google Scholar] [CrossRef]
- Pescador, N.; Cuevas, Y.; Naranjo, S.; Alcaide, M.; Villar, D.; Landazuri, M.O.; Del Peso, L. Identification of a functional hypoxia-responsive element that regulates the expression of the egl nine homologue 3 (egln3/phd3) gene. Biochem. J. 2005, 390, 189–197. [Google Scholar] [CrossRef]
- Packer, J.E.; Slater, T.F.; Willson, R.L. Direct observation of a free radical interaction between vitamin E and vitamin C. Nature 1979, 278, 737–738. [Google Scholar] [CrossRef]
- Drouin, G.; Godin, J.-R.; Pagé, B. The genetics of vitamin C loss in vertebrates. Curr. Genom. 2011, 12, 371–378. [Google Scholar] [CrossRef] [PubMed]
- Nishikimi, M.; Yagi, K. Molecular basis for the deficiency in humans of gulonolactone oxidase, a key enzyme for ascorbic acid biosynthesis. Am. J. Clin. Nutr. 1991, 54, 1203S–1208S. [Google Scholar] [CrossRef] [PubMed]
- Hasselholt, S.; Tveden-Nyborg, P.; Lykkesfeldt, J. Distribution of vitamin C is tissue specific with early saturation of the brain and adrenal glands following differential oral dose regimens in guinea pigs. Br. J. Nutr. 2015, 113, 1539–1549. [Google Scholar] [CrossRef] [PubMed]
- Tveden-Nyborg, P.; Vogt, L.; Schjoldager, J.G.; Jeannet, N.; Hasselholt, S.; Paidi, M.D.; Christen, S.; Lykkesfeldt, J. Maternal vitamin C deficiency during pregnancy persistently impairs hippocampal neurogenesis in offspring of guinea pigs. PLoS ONE 2012, 7, e48488. [Google Scholar] [CrossRef]
- Meredith, M.E.; Harrison, F.E.; May, J.M. Differential regulation of the ascorbic acid transporter SVCT2 during development and in response to ascorbic acid depletion. Biochem. Biophys Res. Commun. 2011, 414, 737–742. [Google Scholar] [CrossRef]
- D’Aniello, C.; Cermola, F.; Palamidessi, A.; Wanderlingh, L.G.; Gagliardi, M.; Migliaccio, A.; Varrone, F.; Casalino, L.; Matarazzo, M.R.; De Cesare, D.; et al. Collagen Prolyl Hydroxylation-Dependent Metabolic Perturbation Governs Epigenetic Remodeling and Mesenchymal Transition in Pluripotent and Cancer Cells. Cancer Res. 2019, 79, 3235–3250. [Google Scholar] [CrossRef]
- Bretaud, S.; Nauroy, P.; Malbouyres, M.; Ruggiero, F. Fishing for collagen function: About development, regeneration and disease. Sem. Cell Dev. Biol. 2019, 89, 100–108. [Google Scholar] [CrossRef]
- Henry, C.A.; Crawford, B.D.; Yan, Y.L.; Postlethwait, J.; Cooper, M.S.; Hille, M.B. Roles for zebrafish focal adhesion kinase in notochord and somite morphogenesis. Dev. Biol. 2001, 240, 474–487. [Google Scholar] [CrossRef]
- Li, R.F.; Wu, T.Y.; Mou, Y.Z.; Wang, Y.S.; Chen, C.L.; Wu, C.Y. Nr2f1b control venous specification and angiogenic patterning during zebrafish vascular development. J. Biomed. Sci. 2015, 22, 104. [Google Scholar] [CrossRef]
- Meyer, D.N.; Baker, B.B.; Baker, T.R. Ancestral TCDD Exposure Induces Multigenerational Histologic and Transcriptomic Alterations in Gonads of Male Zebrafish. Toxicol. Sci. 2018, 164, 603–612. [Google Scholar] [CrossRef]
- Iribarne, M.; Masai, I. Neurotoxicity of cGMP in the vertebrate retina: From the initial research on rd mutant mice to zebrafish genetic approaches. J. Neurogenet 2017, 31, 88–101. [Google Scholar] [CrossRef] [PubMed]
- Iwasa, K.; Shima, K.; Komai, K.; Nishida, Y.; Yokota, T.; Yamada, M. Retinitis pigmentosa and macular degeneration in a patient with ataxia with isolated vitamin E deficiency with a novel c.717 del C mutation in the TTPA gene. J. Neurol. Sci. 2014, 345, 228–230. [Google Scholar] [CrossRef] [PubMed]
- Finno, C.J.; Bordbari, M.H.; Gianino, G.; Ming-Whitfield, B.; Burns, E.; Merkel, J.; Britton, M.; Durbin-Johnson, B.; Sloma, E.A.; McMackin, M.; et al. An innate immune response and altered nuclear receptor activation defines the spinal cord transcriptome during alpha-tocopherol deficiency in Ttpa-null mice. Free Radic. Biol. Med. 2018, 120, 289–302. [Google Scholar] [CrossRef] [PubMed]
- Finno, C.J.; Peterson, J.; Kang, M.; Park, S.; Bordbari, M.H.; Durbin-Johnson, B.; Settles, M.; Perez-Flores, M.C.; Lee, J.H.; Yamoah, E.N. Single-Cell RNA-seq Reveals Profound Alterations in Mechanosensitive Dorsal Root Ganglion Neurons with Vitamin E Deficiency. iScience 2019, 21, 720–735. [Google Scholar] [CrossRef]
- Fischer, A.; Pallauf, J.; Gohil, K.; Weber, S.U.; Packer, L.; Rimbach, G. Effect of selenium and vitamin E deficiency on differential gene expression in rat liver. Biochem. Biophys Res. Commun. 2001, 285, 470–475. [Google Scholar] [CrossRef]
- Gohil, K.; Schock, B.C.; Chakraborty, A.A.; Terasawa, Y.; Raber, J.; Farese, R.V., Jr.; Packer, L.; Cross, C.E.; Traber, M.G. Gene expression profile of oxidant stress and neurodegeneration in transgenic mice deficient in alpha-tocopherol transfer protein. Free Radic. Biol. Med. 2003, 35, 1343–1354. [Google Scholar] [CrossRef]
| 12 hpf | 18 hpf | 24 hpf | |||
|---|---|---|---|---|---|
| Gene Symbol | Log2FC in E– | Gene Symbol | Log2FC in E– | Gene Symbol | Log2FC in E– |
| nitr3r.1l ↑↑↓ | 6.129 | cngb3.2 ↑↑ | 2.944 | olfce2 ↑↑↑ | 5.822 |
| si:dkey-90m5.4 | 3.135 | olfce2 ↑↑↑ | 2.472 | si:dkey-23k10.3 ↓↑ | 4.389 |
| tnnc1a | 3.048 | ighd ↑↑ | 2.326 | si:dkey-238d18.15 ↑↑ | 3.404 |
| olfce2 ↑↑↑ | 3.023 | anpepa ↑↑ | 2.129 | polr3c | 2.768 |
| cd40lg | 2.924 | nitr3r.1l ↑↑↓ | 2.068 | topaz1 | 2.242 |
| slc25a55b | 2.496 | si:dkey-238d18.15 ↑↑ | 2.003 | anpepa ↑↑ | 1.815 |
| ribc1 | 2.386 | rnps1 | 1.801 | si:dkey-238d18.5 | 1.813 |
| ighd ↑↑ | 2.327 | pvalb4 | 1.714 | cngb3.2 ↑↑ | 1.722 |
| si:dkey-159f12.2 | 2.225 | snx21 | 1.662 | dntt | 1.516 |
| scdb | 2.145 | si:dkey-90m5.4 | 1.635 | f2rl1.2 | 1.505 |
| b3gat2 | −4.178 | serpina7 ↓↓↓ | −4.396 | nitr3r.1l ↑↑↓ | −2.692 |
| serpina7 ↓↓↓ | −2.993 | si:dkey-15j16.6 ↓↓ | −2.998 | si:ch211-191i18.2 | −2.657 |
| loxhd1b ↓↓ | −2.787 | si:dkey-73p2.2 ↓↓↓ | −2.901 | si:dkey-15j16.6 ↓↓ | −2.648 |
| si:dkey-73p2.2 ↓↓↓ | −2.673 | zgc:158445 | −2.839 | serpina7 ↓↓↓ | −2.629 |
| asb13a.1 | −2.666 | si:ch1073-13h15.3 | −2.680 | si:dkey-73p2.2 ↓↓↓ | −2.231 |
| rfesd | −2.532 | si:dkey-23k10.3 ↓↑ | −2.460 | zgc:173585 | −1.617 |
| aqp8b | −2.518 | prkra | −2.174 | loxhd1b ↓↓ | −1.575 |
| nr2f1b | −2.205 | grin1b | −1.573 | zgc:101562 | −1.453 |
| proca | −2.069 | kiaa1549lb | −1.570 | insl5a | −1.206 |
| si:ch1073-268j14.1 | −2.041 | ormdl3 | −1.417 | dock11 | −1.125 |
| Symbol | Name | Log2FC in E– vs. E+ | FDR | ||
|---|---|---|---|---|---|
| 12 hpf | 18 hpf | 24 hpf | |||
| nr2f1b | Nuclear receptor subfamily 2, group F, member 1b | −2.21 | 0.17 | 0.12 | 0.007 |
| mafaa | v-maf avian musculoaponeurotic fibrosarcoma oncogene homolog Aa | −1.83 | −0.22 | 0.13 | 0.008 |
| hsf1 | Heat shock transcription factor 1 | 0.68 | 0.55 | −0.08 | 0.013 |
| zgc:101562 | zgc:101562 | −0.94 | −1.11 | −1.45 | 0.020 |
| tead1b | TEA domain family member 1b | 0.46 | 0.26 | 0.15 | 0.024 |
| mbd3b | Methyl-CpG binding domain protein 3b | −0.32 | −0.35 | −0.23 | 0.024 |
| tox2 | TOX high mobility group box family member 2 | −0.48 | −0.41 | −0.08 | 0.027 |
| hmgb2b | High mobility group box 2b | −0.14 | 0.49 | 0.04 | 0.047 |
| bhlhe40 | Basic helix-loop-helix family, member e40 | 0.86 | 0.17 | 0.12 | 0.061 |
| tbx18 | T-box transcription factor | −0.81 | −0.33 | −0.42 | 0.066 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Head, B.; Ramsey, S.A.; Kioussi, C.; Tanguay, R.L.; Traber, M.G. Vitamin E Deficiency Disrupts Gene Expression Networks during Zebrafish Development. Nutrients 2021, 13, 468. https://doi.org/10.3390/nu13020468
Head B, Ramsey SA, Kioussi C, Tanguay RL, Traber MG. Vitamin E Deficiency Disrupts Gene Expression Networks during Zebrafish Development. Nutrients. 2021; 13(2):468. https://doi.org/10.3390/nu13020468
Chicago/Turabian StyleHead, Brian, Stephen A. Ramsey, Chrissa Kioussi, Robyn L. Tanguay, and Maret G. Traber. 2021. "Vitamin E Deficiency Disrupts Gene Expression Networks during Zebrafish Development" Nutrients 13, no. 2: 468. https://doi.org/10.3390/nu13020468
APA StyleHead, B., Ramsey, S. A., Kioussi, C., Tanguay, R. L., & Traber, M. G. (2021). Vitamin E Deficiency Disrupts Gene Expression Networks during Zebrafish Development. Nutrients, 13(2), 468. https://doi.org/10.3390/nu13020468







