Distinguishing Early Successional Plant Communities Using Ground-Level Hyperspectral Data
Abstract
:1. Introduction
2. Experimental Section
2.1. Study Site
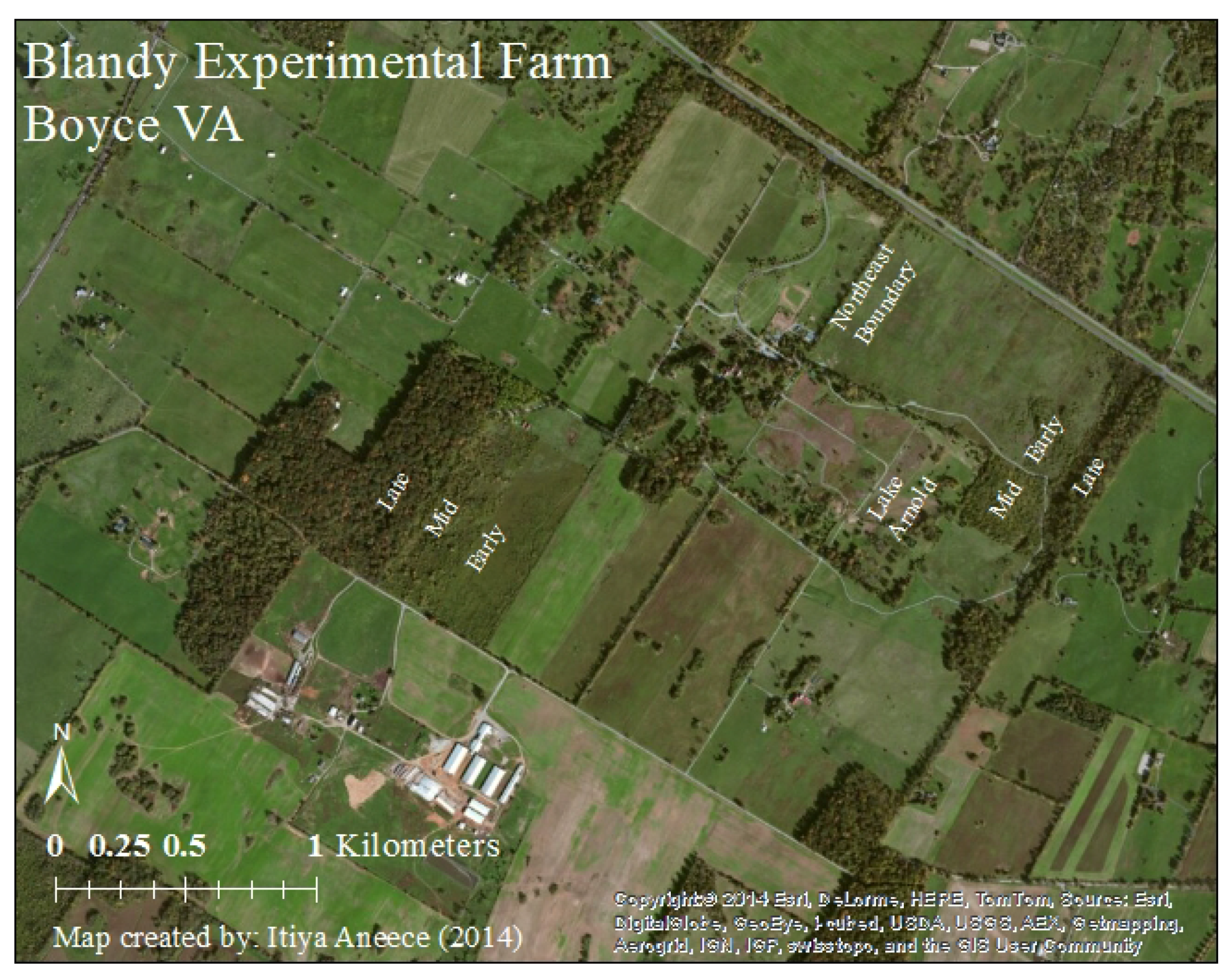

2.2. Field Methods

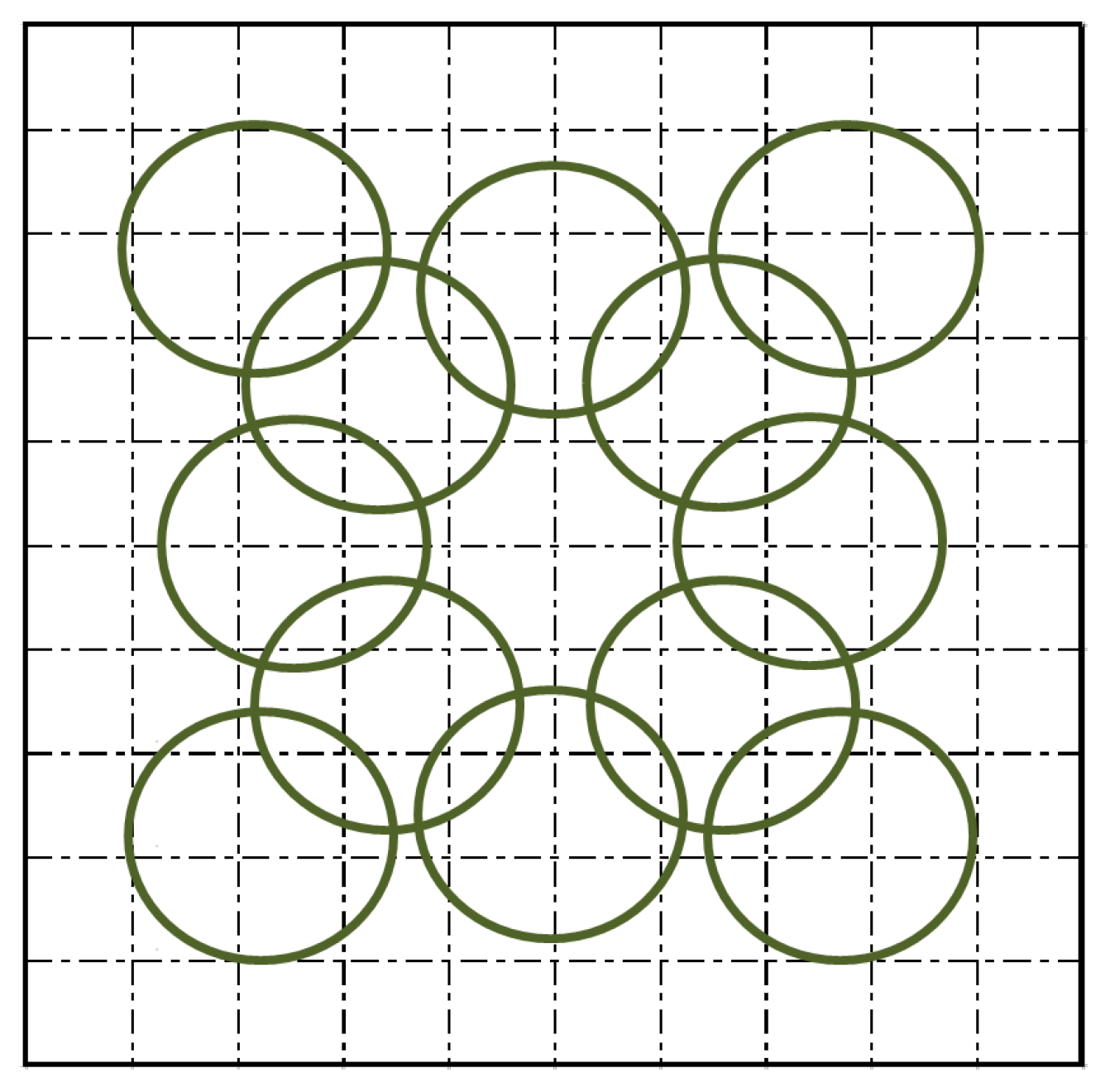
| Plot | Number of Spectra | Date of Collection |
|---|---|---|
| lacp1 | 37 | 5 July 2014 |
| lacp2 | 36 | 5 July 2014 |
| lacp3 | 36 | 5 July 2014 |
| nebcp1 | 36 | 4 July 2014 |
| nebcp2 | 36 | 4 July 2014 |
| nebcp3 | 36 | 4 July 2014 |
| neecp1 | 37 | 4 July 2014 |
| neecp2 | 37 | 4 July 2014 |
| neecp3 | 36 | 4 July 2014 |
| swecp1 | 35 | 5 July 2014 |
| swecp2 | 36 | 5 July 2014 |
| swecp3 | 38 | 5 July 2014 |
2.3. Data Analysis and Statistical Methods
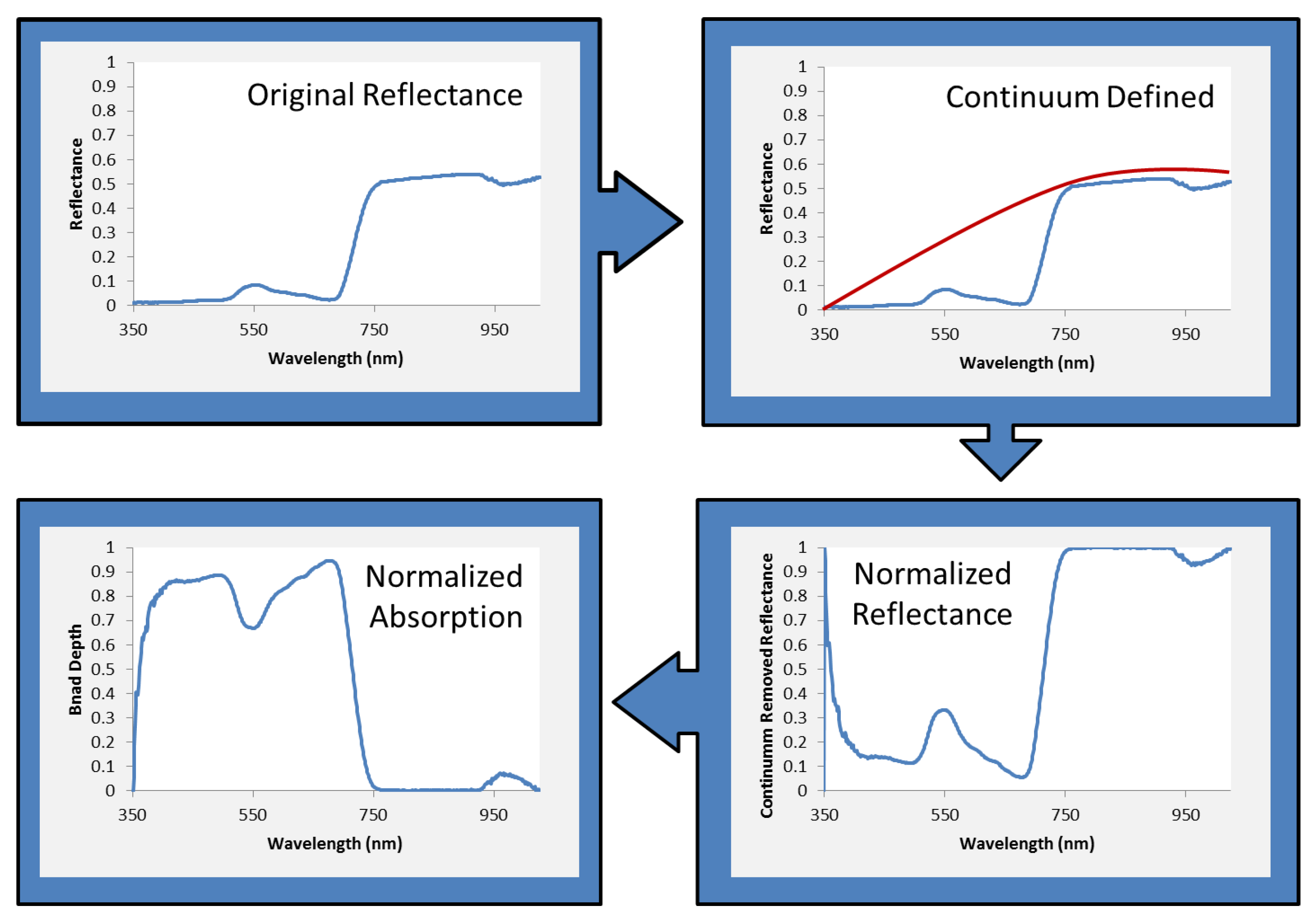
3. Results and Discussion
3.1. Species Ordinations

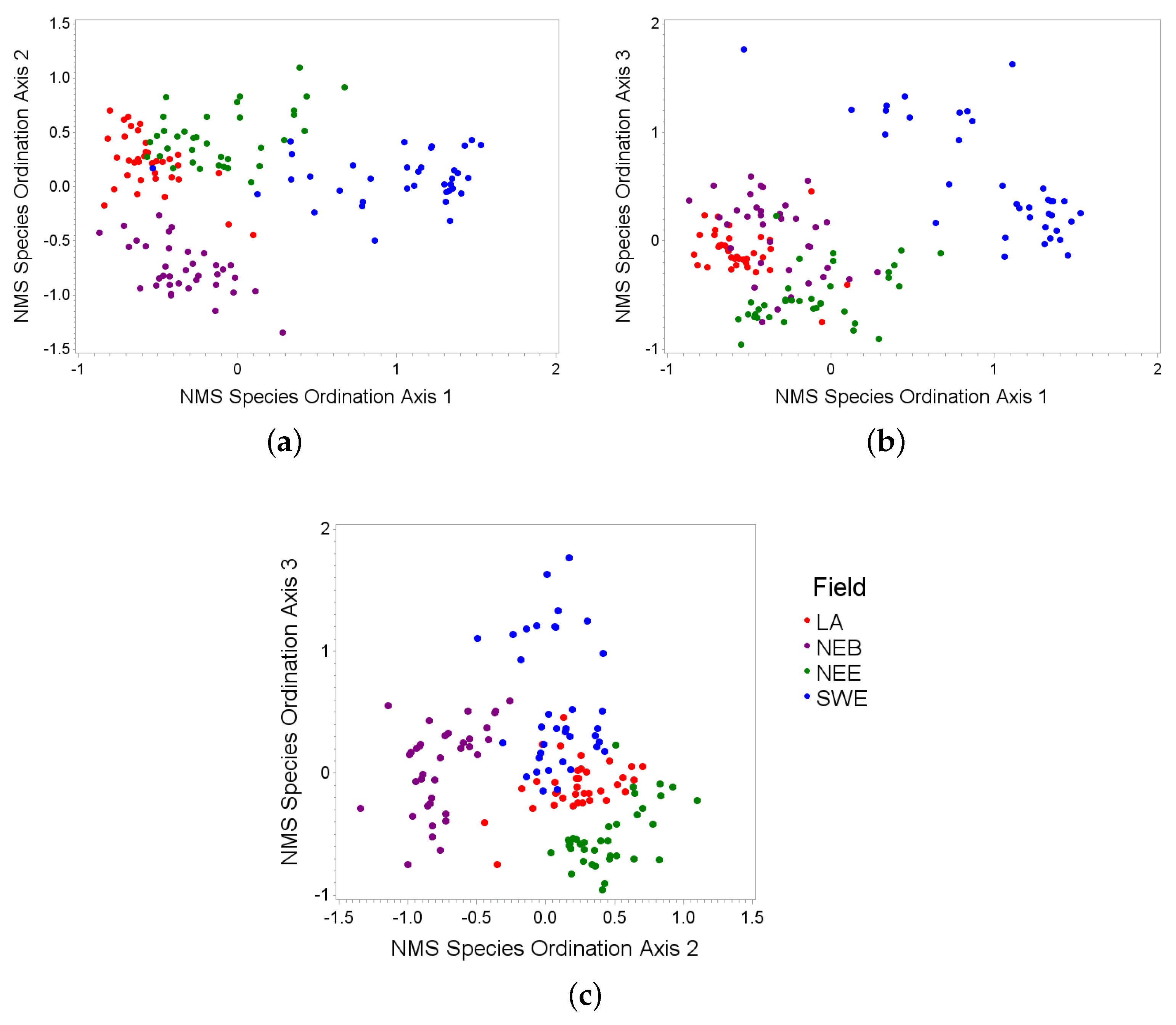
| Axis | Species | Partial R-Square |
|---|---|---|
| Axis 1 | Solidago altissima | 0.6962 |
| Axis 1 | Bromus japonicus | 0.1117 |
| Axis 2 | Festuca rubra | 0.5538 |
| Axis 2 | Galium verum | 0.1251 |
| Axis 2 | Muhlenbergia schreberi | 0.0737 |
| Axis 2 | Achillea millefolium | 0.0433 |
| Axis 2 | Bromus commutatus | 0.0363 |
| Axis 3 | Galium verum | 0.3817 |
| Axis 3 | Solidago gigantea | 0.2266 |
| Axis 3 | Dactylis glomerata | 0.0813 |
| Axis 3 | Bromus japonicus | 0.0573 |
| Axis 3 | Ambrosia artemisiifolia | 0.0476 |
| Axis 3 | Lonicera japonica | 0.0336 |
3.2. Spectral Ordinations
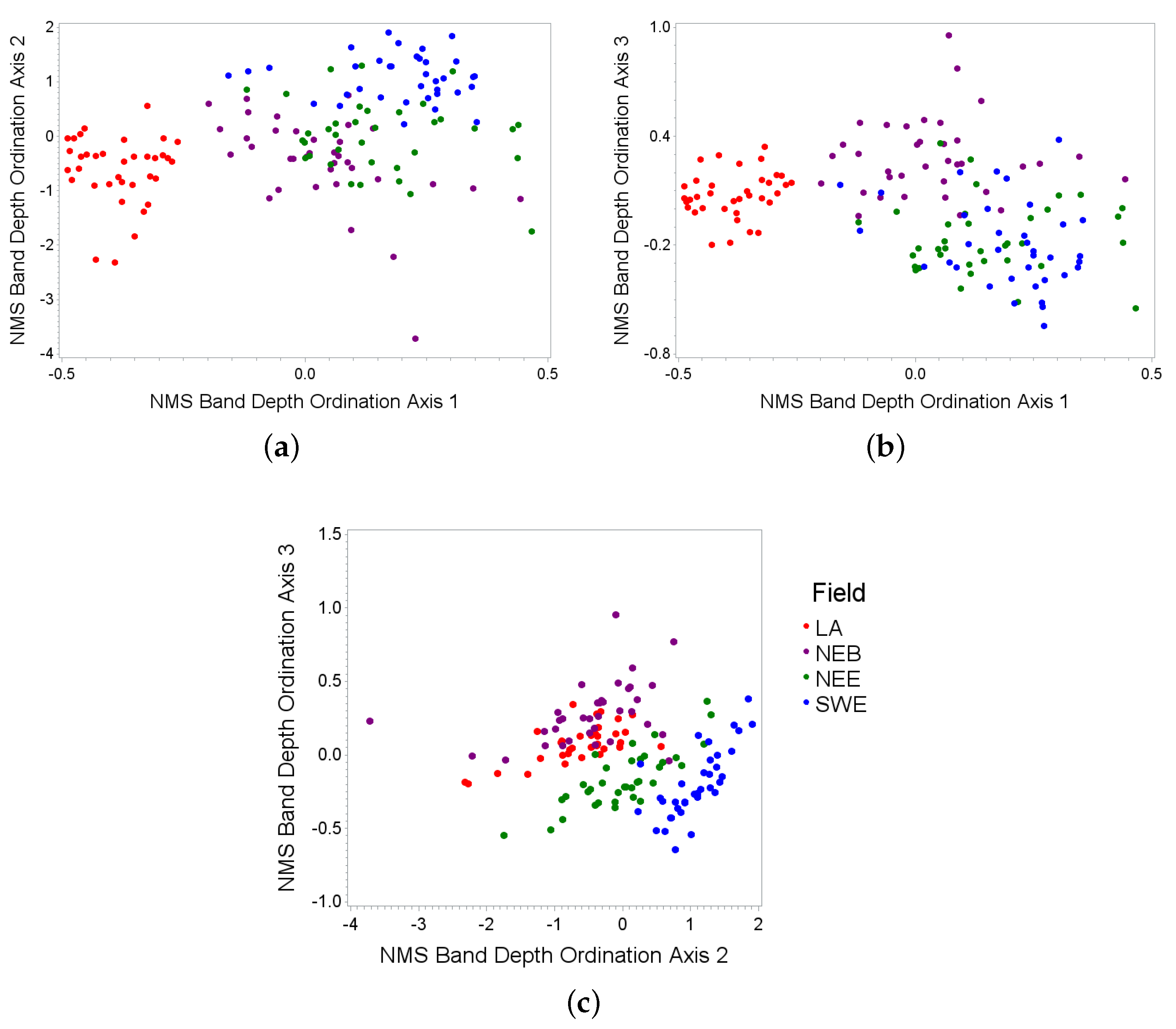
| Axis | 435 | 525 | 575 | 635 | 680 | 710 | 750 | 835 | 970 |
|---|---|---|---|---|---|---|---|---|---|
| Axis 1 | 3.60 | . | . | 7.40 | −7.66 | −4.78 | 1.78 | . | 8.48 |
| Axis 2 | 6.31 | −1.65 | 3.44 | 7.28 | . | . | 2.86 | . | 2.22 |
| Axis 3 | . | −4.11 | 6.96 | −6.41 | . | 3.79 | 5.31 | 5.59 | . |
3.3. Discriminant Analyses
| Number of Observations Classified into Plot | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| lacp1 | lacp2 | lacp3 | nebcp1 | nebcp2 | nebcp3 | neecp1 | neecp2 | neecp3 | swecp1 | swecp2 | swecp3 | Total | Producer’s Accuracy | |
| lacp1 | 12 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 12 | 85.7% |
| lacp2 | 0 | 12 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 12 | 100% |
| lacp3 | 2 | 0 | 8 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 12 | 88.9% |
| nebcp1 | 0 | 0 | 0 | 4 | 5 | 2 | 0 | 1 | 0 | 0 | 0 | 0 | 12 | 44.4% |
| nebcp2 | 0 | 0 | 0 | 4 | 6 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 12 | 46.2% |
| nebcp3 | 0 | 0 | 0 | 1 | 2 | 6 | 0 | 0 | 2 | 1 | 0 | 0 | 12 | 54.5% |
| neecp1 | 0 | 0 | 0 | 0 | 0 | 0 | 4 | 3 | 2 | 2 | 1 | 0 | 12 | 44.4% |
| neecp2 | 0 | 0 | 1 | 0 | 0 | 0 | 3 | 6 | 1 | 0 | 0 | 1 | 12 | 40.0% |
| neecp3 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 3 | 7 | 0 | 0 | 0 | 12 | 50.0% |
| swecp1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 5 | 1 | 4 | 12 | 45.5% |
| swecp2 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 8 | 2 | 12 | 72.7% |
| swecp3 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 1 | 9 | 12 | 56.3% |
| Total | 14 | 12 | 9 | 9 | 13 | 11 | 9 | 15 | 14 | 11 | 11 | 16 | 144 | |
| User’s Accuracy | 100% | 100% | 66.7% | 33.3% | 50.0% | 50.0% | 33.3% | 50.0% | 58.3% | 41.7% | 66.7% | 75.0% | ||
| Overall Accuracy | 60.4% | |||||||||||||
| Number of Observations Classified into Plot | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| lacp1 | lacp2 | lacp3 | nebcp1 | nebcp2 | nebcp3 | neecp1 | neecp2 | neecp3 | swecp1 | swecp2 | swecp3 | Total | Producer’s Accuracy | |
| lacp1 | 11 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 12 | 78.6% |
| lacp2 | 1 | 10 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 12 | 83.3% |
| lacp3 | 2 | 1 | 7 | 0 | 0 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 12 | 77.8% |
| nebcp1 | 0 | 0 | 0 | 8 | 3 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 12 | 88.9% |
| nebcp2 | 0 | 0 | 0 | 1 | 10 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 12 | 71.4% |
| nebcp3 | 0 | 0 | 1 | 0 | 1 | 9 | 0 | 0 | 1 | 0 | 0 | 0 | 12 | 75.0% |
| neecp1 | 0 | 0 | 0 | 0 | 0 | 0 | 8 | 0 | 1 | 2 | 0 | 1 | 12 | 80.0% |
| neecp2 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 7 | 0 | 0 | 0 | 4 | 12 | 77.8% |
| neecp3 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 10 | 0 | 0 | 0 | 12 | 77.0% |
| swecp1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 11 | 1 | 0 | 12 | 73.3% |
| swecp2 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 9 | 1 | 12 | 90.0% |
| swecp3 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 11 | 12 | 64.7% |
| Total | 14 | 12 | 9 | 9 | 14 | 12 | 10 | 9 | 13 | 15 | 10 | 17 | 144 | |
| User’s Accuracy | 91.7% | 83.3% | 58.3% | 66.7% | 83.3% | 75% | 66.7% | 58.3% | 83.3% | 91.7% | 75.0% | 91.7% | ||
| Overall Accuracy | 77.1% | |||||||||||||
| Simulated Broad Bands | Narrow Bands | |
|---|---|---|
| Matthew’s Correlation Coefficient | 0.548 | 0.745 |
| Overall Accuracy | 60.4% | 77.1% |
| Average Producer’s Accuracy | 60.7% | 78.1% |
| Average User’s Accuracy | 60.4% | 77.1% |
3.4. Other Considerations
3.5. Context of Key Findings
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Appendix. Persistence of Invasive Plants
References
- Wang, J.; Epstein, H.; Wang, L. Soil CO2 flux and its controls during secondary succession. J. Geophys. Res. 2010, 115. [Google Scholar] [CrossRef]
- Mosher, E.; Silander, J.; Latimer, A. The role of land-use history in major invasions by woody plant species in the northeastern North American landscape. Biol. Invasions 2009, 11, 2317–2328. [Google Scholar] [CrossRef]
- Chapin, S., III; Walker, B.; Hobbs, R.; Hooper, D.; Lawton, J.; Sala, O.; Tilman, D. Biotic control over the functioning of ecosystems. Science 1997, 277, 500–504. [Google Scholar] [CrossRef]
- Kuhman, T.; Pearson, S.; Turner, M. Agricultural land-use history increases non-native plant invasion in a southern Appalachian forest a century after abandonment. Can. J. For. Res. 2011, 41, 920–929. [Google Scholar] [CrossRef]
- DeMeester, J.; DeB Richter, D. Differences in wetland nitrogen cycling between the invasive grass Microstegium vimineum and a diverse plant community. Ecol. Appl. 2010, 20, 609–619. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, J.; Williams, P.; Timmins, S. Secondary forest succession differs through naturalised gorse and native kanuka near Wellington and Nelson. N. Z. J. Ecol. 2007, 31, 22–38. [Google Scholar]
- Yoshida, K.; Oka, S. Invasion of Leucaena leucocephala and its effects on the native plant community in the Ogasawara (Bonin) Islands. Weed Technol. 2004, 18, 1371–1375. [Google Scholar] [CrossRef]
- Feldpausch, T.; Rondon, M.; Fernandes, E.; Riha, S.; Wandelli, E. Carbon and nutrient accumulation in secondary forests regenerating on pastures in central Amazonia. Ecol. Appl. 2004, 14, 164–176. [Google Scholar] [CrossRef]
- Kassi N’Dja, J.; Decocq, G. Successional patterns of plant species and community diversity in a semi-deciduous tropical forest under shifting cultivation. J. Veg. Sci. 2008, 19, 809–820. [Google Scholar] [CrossRef]
- Otto, R.; Krusi, B.; Burga, C.; Fernandez-Palacios, J. Old-field succession along a precipitation gradient in the semi-arid coastal region of Tenerife. J. Arid Environ. 2006, 65, 156–178. [Google Scholar] [CrossRef]
- Cunard, C.; Lee, T. Is patience a virtue? Succession, light, and the death of invasive glossy buckthorn (Frangula alnus). Biol. Invasions 2009, 11, 577–586. [Google Scholar] [CrossRef]
- Simberloff, D. Invasions of plant communities—More of the same, something very different, or both? Am. Midl. Nat. 2010, 163, 220–233. [Google Scholar] [CrossRef]
- Grau, H.R.; Arturi, M.F.; Brown, A.D.; Acenolaza, P.G. Floristic and structural patterns along a chronosequence of secondary forest succession in Argentinean subtropical montane forests. For. Ecol. Manag. 1997, 95, 161–171. [Google Scholar] [CrossRef]
- Leicht-Young, S.A.; O’Donnell, H.; Latimer, A.M.; Silander, J.A. Effects of an invasive plant species, Celastrus orbiculatus, on soil composition and processes. Am. Midl. Nat. 2009, 161, 219–231. [Google Scholar] [CrossRef]
- Wilfong, B.; Gorchov, D.; Henry, M. Detecting an invasive shrub in deciduous forest understories using remote sensing. Weed Sci. 2009, 57, 512–520. [Google Scholar] [CrossRef]
- Zhang, J.; Rivard, B.; Sanchez-Azofeifa, A.; Castro-Esau, K. Intra- and inter-class spectral variability of tropical tree species at La Selva, Costa Rica: Implications for species identification using HYDICE imagery. Remote Sens. Environ. 2006, 105, 129–141. [Google Scholar] [CrossRef]
- Schmidt, K.S.; Skidmore, A.K. Exploring spectral discrimination of grass species in African rangelands. Int. J. Remote Sens. 2001, 22, 3421–3434. [Google Scholar] [CrossRef]
- Bradley, B.; Mustard, J. Characterizing the landscape dynamics of an invasive plant and risk of invasion using remote sensing. Ecol. Appl. 2006, 16, 1132–1147. [Google Scholar] [CrossRef]
- Smith, A.; Blackshaw, R. Weed: Crop discrimination using remote sensing: A detached leaf experiment. Weed Technol. 2003, 17, 811–820. [Google Scholar] [CrossRef]
- Jafari, R.; Lewis, M. Arid land characterisation with EO-1 Hyperion hyperspectral data. Int. J. Appl. Earth Obs. Geoinf. 2012, 19, 298–307. [Google Scholar] [CrossRef]
- Burkholder, A. Seasonal Trends in Separability of Leaf Reflectance Spectra for Ailanthus altissima and Four Other Tree Species. Master’s Thesis, Eberly College of Arts and Sciences at West Virginia University, Morgantown, WV, USA, 2010. [Google Scholar]
- Narumalani, S.; Mishra, D.; Wilson, R.; Reece, P.; Kohler, A. Detecting and mapping four invasive species along the floodplain of North Platte River, Nebraska. Weed Technol. 2009, 23, 99–107. [Google Scholar] [CrossRef]
- Pinard, V.; Bannari, A. Spectroradiometric analysis in a hyperspectral use perspective to discriminate between forest species. In Proceedings of the 2003 IEEE International Geoscience and Remote Sensing Symposium, IGARSS’03, Toulouse, France, 21–25 July 2003; Volume 7, pp. 4301–4303.
- Rud, R.; Shoshany, M.; Alchanatis, V.; Cohen, Y. Application of spectral features’ ratios for improving classification in partially calibrated hyperspectral imagery: A case study of separating Mediterranean vegetation species. J. Real-Time Image Process. 2006, 1, 143–152. [Google Scholar] [CrossRef]
- Gai, Y.Y.; Fan, W.J.; Xu, X.R.; Zhang, Y.Z. Flower species identification and coverage estimation based on hyperspectral remote sensing data. In Proceedings of the 2011 IEEE International Geoscience and Remote Sensing Symposium (IGARSS), Vancouver, BC, Canada, 24–29 July 2011; pp. 1243–1246.
- Xiao, Y.; Zhao, W.; Zhou, D.; Gong, H. Sensitivity analysis of vegetation reflectance to biochemical and biophysical variables at leaf, canopy, and regional scales. IEEE Trans. Geosci. Remote Sens. 2014, 52, 4014–4024. [Google Scholar] [CrossRef]
- Mahlein, A. Detection, Identification, and Quantification of Fungal Diseases of Sugar Beet Leaves Using Imaging and Non-Imaging Hyperspectral Techniques. Ph.D. Thesis, Universitats- und Landesbibliothek Bonn, Bonn, Germany, 2011. [Google Scholar]
- Merzlyak, M.N.; Gitelson, A.A.; Chivkunova, O.B.; Solovchenko, A.E.; Pogosyan, S.I. Application of reflectance spectroscopy for analysis of higher plant pigments. Russ. J. Plant Physiol. 2003, 50, 704–710. [Google Scholar] [CrossRef]
- Buschmann, C.; Nagel, E.; Rang, S.; Stober, F. Interpretation of reflectance spectra of terrestrial vegetation based on specifical plant test systems. In Proceedings of the 10th Annual International Geoscience and Remote Sensing Symposium, IGARSS ’90, College Park, MD, USA, 20–24 May 1990; pp. 1927–1930.
- Papes, M.; Tupayachi, R.; Martinez, P.; Peterson, A.; Powell, G. Using hyperspectral satellite imagery for regional inventories: A test with tropical emergent trees in the Amazon Basin. J. Veg. Sci. 2010, 21, 342–354. [Google Scholar] [CrossRef]
- Gamon, J.; Berry, J. Facultative and constitutive pigment effects on the Photochemical Reflectance Index (PRI) in sun and shade conifer needles. Isr. J. Plant Sci. 2012, 60, 85–95. [Google Scholar] [CrossRef]
- Kokaly, R.; Clark, R. Spectroscopic determination of leaf biochemistry using band-depth analysis of absorption features and stepwise multiple linear regression. Remote Sens. Environ. 1999, 67, 267–287. [Google Scholar] [CrossRef]
- Roberts, D.; Ustin, S.L.; Ogunjemio, S.; Greenberg, J.; Dobrowski, S.; Chen, J.; Hinckley, T. Spectral and structural measures of northwest forest vegetation at leaf to landscape scales. Ecosystems 2004, 7, 545–562. [Google Scholar] [CrossRef]
- Wang, L.; Shaner, P.; Macko, S. Foliar delta 15N patterns along successional gradients at plant community and species levels. Geophys. Res. Lett. 2007, 34, L16403. [Google Scholar] [CrossRef]
- Bowers, M.A. University of Virginia’s Blandy Experimental Farm. Bull. Ecol. Soc. Am. 1997, 78, 220–225. [Google Scholar]
- Mitchell, J.; Glenn, N.; Sankey, T.; Derryberry, D.; Germino, M. Remote sensing of sagebrush canopy nitrogen. Remote Sens. Environ. 2012, 124, 217–223. [Google Scholar] [CrossRef]
- Sanches, I.; Filho, C.; Kokaly, R. Spectroscopic remote sensing of plant stress at leaf and canopy levels using the chlorophyll 680 nm absorption feature with continuum removal. ISPRS J. Photogramm. Remote Sens. 2014, 97, 111–122. [Google Scholar] [CrossRef]
- Schmidt, K.; Skidmore, A. Spectral discrimination of vegetation types in a coastal wetland. Remote Sens. Environ. 2003, 85, 92–108. [Google Scholar] [CrossRef]
- Clark, R.; Roush, T. Reflectance spectroscopy: Quantitative analysis techniques for remote sensing applications. J. Geophys. Res. 1984, 89, 6329–6340. [Google Scholar] [CrossRef]
- Schmidtlein, S.; Zimmermann, P.; Schupferling, R.; Weiß, C. Mapping the floristic continuum: Ordination space position estimated from imaging spectroscopy. J. Veg. Sci. 2007, 18, 131–140. [Google Scholar] [CrossRef]
- EDDMapS. EDDMapS: Early Detection and Distribution Mapping System. Available online: http://www.eddmaps.org/ (accessed on 5 March 2014).
- USDA Plants Database. USDA: United States Department of Agriculture, Natural Resources Conservation Services, Plants Database. Available online: http://www.plants.usda.gov/ (accessed on 3 March 2015).
- Grime, J. Benefits of plant diversity to ecosystems: Immediate, filter and founder effects. J. Ecol. 1998, 86, 902–910. [Google Scholar] [CrossRef]
- Blackburn, G.A. Hyperspectral remote sensing of plant pigments. J. Exp. Bot. 2006, 58, 855–867. [Google Scholar] [CrossRef] [PubMed]
- Koning, R. Light Reactions. Plant Physiology Information Website. 1994. Available online: http://plantphys.info/plant_physiology/lightrxn.shtml (accessed on 22 May 2015).
- Sims, D.; Gamon, J. Relationships between leaf pigment content and spectral reflectance across a wide range of species, leaf structures and developmental stages. Remote Sens. Environ. 2002, 81, 337–354. [Google Scholar] [CrossRef]
- Gamon, J.; Field, C.; Roberts, D.; Ustin, S.; Valentini, R. Functional patterns in an annual grassland during an AVIRIS overflight. Remote Sens. Environ. 1993, 44, 239–253. [Google Scholar] [CrossRef]
- Wang, J.; Xu, R.; Yang, S. Estimation of plant water content by spectral absorption features centered at 1450 nm and 1940 nm regions. Environ. Monit. Assess. 2009, 157, 459–469. [Google Scholar] [CrossRef] [PubMed]
- Laidler, G.; Treitz, P.; Atkinson, D. Remote sensing of Arctic vegetation: Relations between the NDVI, spatial resolution and vegetation cover on Boothia Peninsula, Nunavut. Arctic 2008, 61, 1–13. [Google Scholar] [CrossRef]
- Cochrane, M.A. Using vegetation reflectance variability for species level classification of hyperspectral data. Int. J. Remote Sens. 2000, 21, 2075–2087. [Google Scholar] [CrossRef]
- Baron, J.; Barber, M.; Adams, M.; Agboola, J.; Allen, E.; Bealey, W.; Bobbink, R.; Bobrovsky, M.V.; Bowman, W.; Branquinho, C.; et al. The effects of atmospheric nitrogen deposition on terrestrial and freshwater biodiversity. In Nitrogen Deposition, Critical Loads and Biodiversity; Sutton, M., Mason, K., Sheppard, L., Sverdrup, H., Haeuber, R., Hicks, W., Eds.; Springer: Dordrecht, The Netherlands, 2014; pp. 465–480. [Google Scholar]
- Castro, H.; Fortunel, C.; Freitas, H. Effects of land abandonment on plant litter decomposition in a Montado system: Relation to litter chemistry and community functional parameters. Plant Soil 2010, 333, 181–190. [Google Scholar] [CrossRef]
- Aragon, R.; Morales, J. Species composition and invasion in NW Argentinian secondary forests: Effects of land use history, environment and landscape. J. Veg. Sci. 2003, 14, 195–204. [Google Scholar] [CrossRef]
- Riedel, S.; Epstein, H. Edge effects on vegetation and soils in a Virginia old-field. Plant Soil 2005, 270, 13–22. [Google Scholar] [CrossRef]
- Rooney, T.; Rogers, D. Colonization and effects of garlic mustard (Alliaria petiolata), European buckthorn (Rhamnus cathartica), and Bell’s honeysuckle (Lonicera x bella) on understory plants after five decades in southern Wisconsin forests. Invasive Plant Sci. Manag. 2011, 4, 317–325. [Google Scholar] [CrossRef]
- Seabloom, E.; Borer, E.; Buckley, Y.; Cleland, E.; Davies, K.; Firn, J.; Harpole, W.; Hautier, Y.; Lind, E.; MacDougall, A.; et al. Predicting invasion in grassland ecosystems: Is exotic dominance the real embarrassment of richness? Glob. Chang. Biol. 2013, 19, 3677–3687. [Google Scholar] [CrossRef] [PubMed]
- Arroyo-Mora, J.; Sanchez-Azofeifa, A.; Kalacska, M.; Rivard, B.; Calvo-Alvarado, J.; Janzen, D. Secondary forest detection in a neotropical dry forest landscape using Landsat 7 ETM+ and IKONOS imagery. Biotropica 2005, 37, 497–507. [Google Scholar] [CrossRef]
- Weidenhamer, J.; Callaway, R. Direct and indirect effects of invasive plants on soil chemistry and ecosystem function. J. Chem. Ecol. 2010, 36, 59–69. [Google Scholar] [CrossRef] [PubMed]
- Crowley, J.; Brickey, D.; Rowan, L. Airborne imaging spectrometer data of the Ruby Mountains, Montana: Mineral discrimination using relative absorption band-depth images. Remote Sens. Environ. 1989, 29, 121–134. [Google Scholar] [CrossRef]
- Rice, M.; Cloutis, E.; Bell, J.; Bish, D.; Horgan, B.; Mertzman, S.; Craig, M.; Renaut, R.; Gautason, B.; Mountain, B. Reflectance spectra diversity of silica-rich materials: Sensitivity to environment and implications for detections on Mars. Icarus 2013, 223, 499–533. [Google Scholar] [CrossRef]
- Castro-Esau, K.; Sanchez-Azofeifa, G.; Caelli, T. Discrimination of lianas and trees with leaf-level hyperspectral data. Remote Sens. Environ. 2004, 90, 353–372. [Google Scholar] [CrossRef]
- Carlson, K.; Asner, G.; Hughes, R.; Ostertag, R.; Martin, R. Hyperspectral remote sensing of canopy biodiversity in Hawaiian lowland rainforests. Ecosystems 2007, 10, 536–549. [Google Scholar] [CrossRef]
- Broge, N.; Leblanc, E. Comparing prediction power and stability of broadband and hyperspectral vegetation indices for estimation of green leaf area index and canopy chlorophyll density. Remote Sens. Environ. 2001, 76, 156–172. [Google Scholar] [CrossRef]
- Clevers, J.; Kooistra, L. Using hyperspectral remote sensing data for retrieving canopy chlorophyll and nitrogen content. IEEE J. Sel. Top. Appl. Earth Obs. 2012, 5, 574–583. [Google Scholar] [CrossRef]
- Gao, Z.; Zhang, L. Multi-seasonal spectral characteristics analysis of coastal salt marsh vegetation in Shanghai, China. Estuar. Coast. Shelf Sci. 2006, 69, 217–224. [Google Scholar] [CrossRef]
- EnMAP. Mission|Enmap. Available online: http://www.enmap.org/?q=mission (accessed on 20 October 2015).
- Hook, S. Welcome to HyspIRI Mission Study Website: Hyperspectral Infrared Imager. Available online: https://hyspiri.jpl.nasa.gov/ (accessed on 20 October 2015).
- Feldman, S. Biological control of plumeless thistle (Carduus acanthoides L.) in Argentina. Weed Sci. 1997, 45, 534–537. [Google Scholar]
- Zhang, R.; Heberling, J.; Haner, E.; Shea, K. Tolerance of two invasive thistles to repeated disturbance. Ecol. Res. 2011, 26, 575–581. [Google Scholar] [CrossRef]
- Allen, M.; Shea, K. Spatial segregation of congeneric invaders in central Pennsylvania, USA. Biol. Invasions 2006, 8, 509–521. [Google Scholar] [CrossRef]
- Knight, K.; Kurylo, J.; Endress, A.; Stewart, J.; Reich, P. Ecology and ecosystem impacts of common buckthorn (Rhamnus cathartica): A review. Biol. Invasions 2007, 9, 925–937. [Google Scholar] [CrossRef]
- Fryer, J. Celastrus orbiculatus . Fire Effects Information System, (Online); U.S. Department of Agriculture, Forest Service, Rocky Mountain Research Station, Fire Sciences Laboratory (Producer), 2011. Available online: http://www.fs.fed.us/database/feis/ (accessed on 30 October 2014).
- Ladwig, L.; Meiners, S. Liana host preference and implications for deciduous forest regeneration. J. Torrey Bot. Soc. 2010, 137, 103–112. [Google Scholar] [CrossRef]
- Pavlovic, N.; Leicht-Young, S. Are temperate mature forests buffered from invasive lianas? J. Torrey Bot. Soc. 2011, 138, 85–92. [Google Scholar] [CrossRef]
- Pooler, M.; Dix, R.; Feely, J. Interspecific hybridizations between the native bittersweet, Celastrus scandens, and the introduced invasive species, C. orbiculatus. Southeast. Nat. 2002, 1, 69–76. [Google Scholar] [CrossRef]
- Kim, H.; Hoelmer, K.; Lee, W.; Kwon, Y.; Lee, S. Molecular and morphological identification of the soybean aphid and other Aphis species on the primary host Rhamnus davurica in Asia. Ann. Entomol. Soc. Am. 2010, 103, 532–543. [Google Scholar] [CrossRef]
- Heimpel, G.; Frelich, L.; Landis, D.; Hopper, K.; Hoelmer, K.; Sezen, Z.; Asplen, M.; Wu, K. European buckthorn and Asian soybean aphid as components of an extensive invasional meltdown in North America. Biol. Invasions 2010, 12, 2913–2931. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aneece, I.; Epstein, H. Distinguishing Early Successional Plant Communities Using Ground-Level Hyperspectral Data. Remote Sens. 2015, 7, 16588-16606. https://doi.org/10.3390/rs71215850
Aneece I, Epstein H. Distinguishing Early Successional Plant Communities Using Ground-Level Hyperspectral Data. Remote Sensing. 2015; 7(12):16588-16606. https://doi.org/10.3390/rs71215850
Chicago/Turabian StyleAneece, Itiya, and Howard Epstein. 2015. "Distinguishing Early Successional Plant Communities Using Ground-Level Hyperspectral Data" Remote Sensing 7, no. 12: 16588-16606. https://doi.org/10.3390/rs71215850
APA StyleAneece, I., & Epstein, H. (2015). Distinguishing Early Successional Plant Communities Using Ground-Level Hyperspectral Data. Remote Sensing, 7(12), 16588-16606. https://doi.org/10.3390/rs71215850







