Temporal Transferability of Pine Forest Attributes Modeling Using Low-Density Airborne Laser Scanning Data
Abstract
1. Introduction
2. Materials and Methods
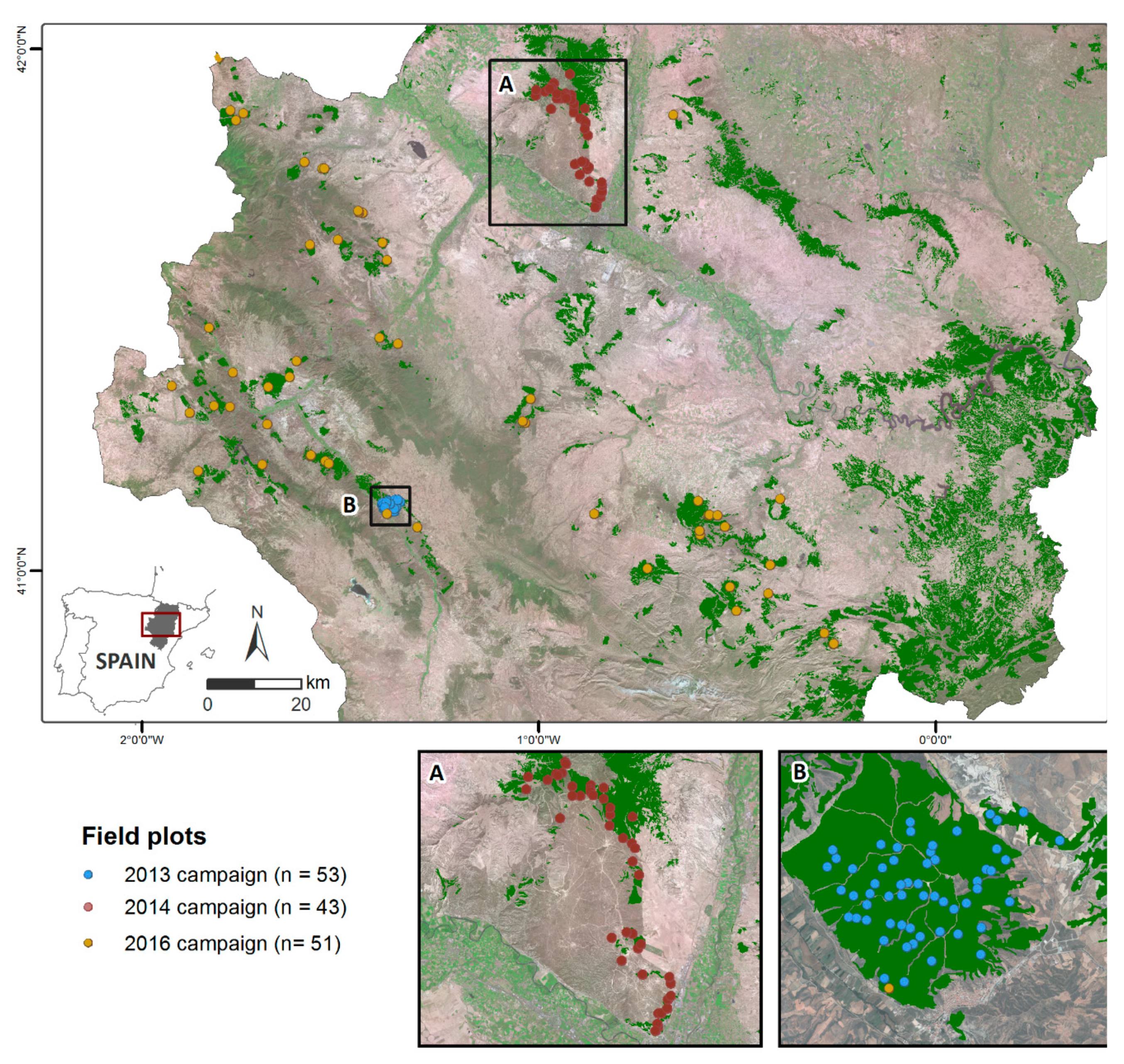
2.1. Study Area
2.2. Forest Inventory Data
2.3. Inventories Updating and Stand Variable Computation
2.4. ALS Data and Processing
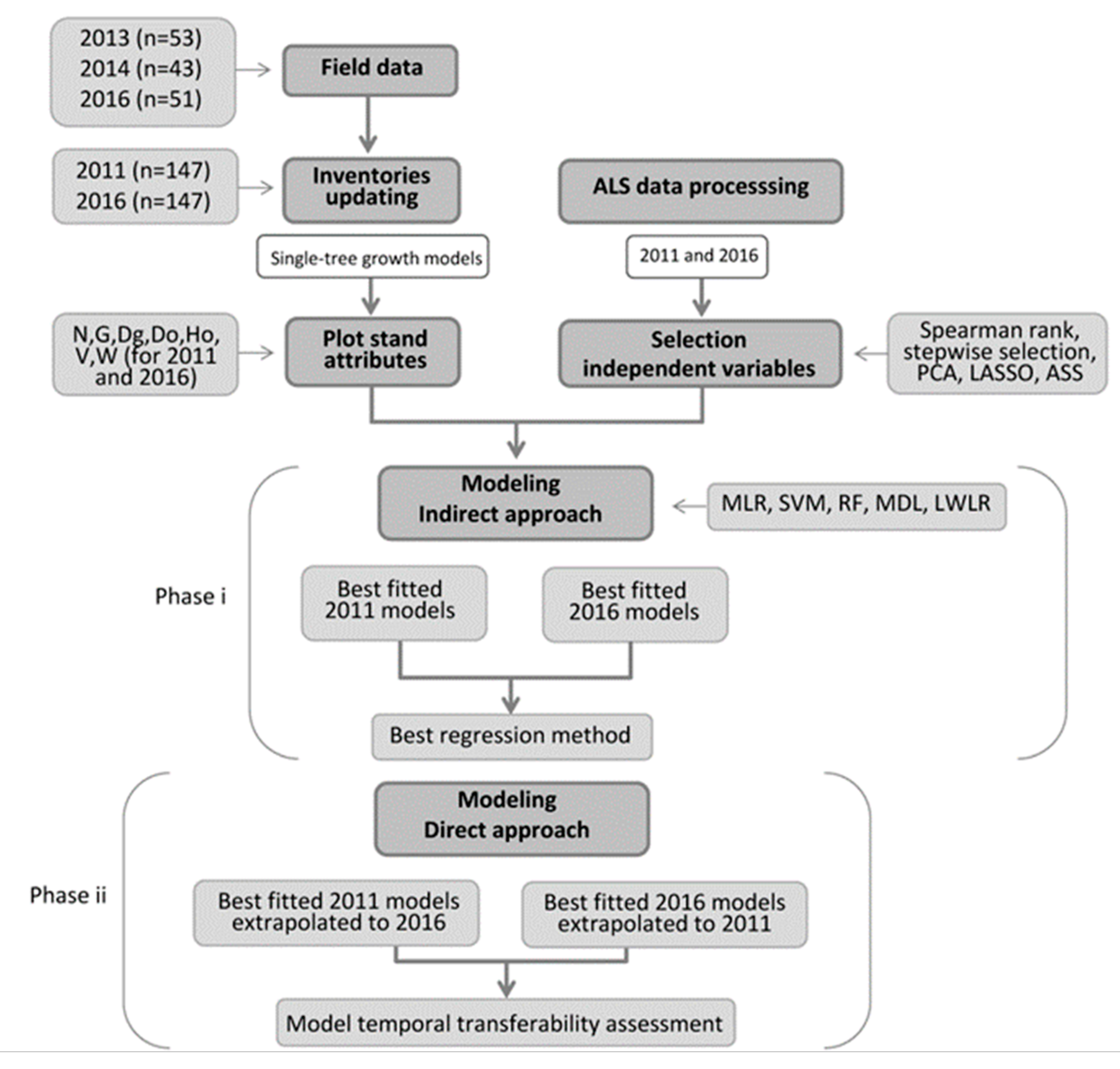
2.5. Modeling of Forest Stand Attributes and Temporal Tranferability Assessment
2.5.1. Variable Selection and Attributes Modeling Using the Indirect Approach
2.5.2. Assessment of Temporal Transferability by Applying a Direct Approach
3. Results
3.1. Field Plot Computation
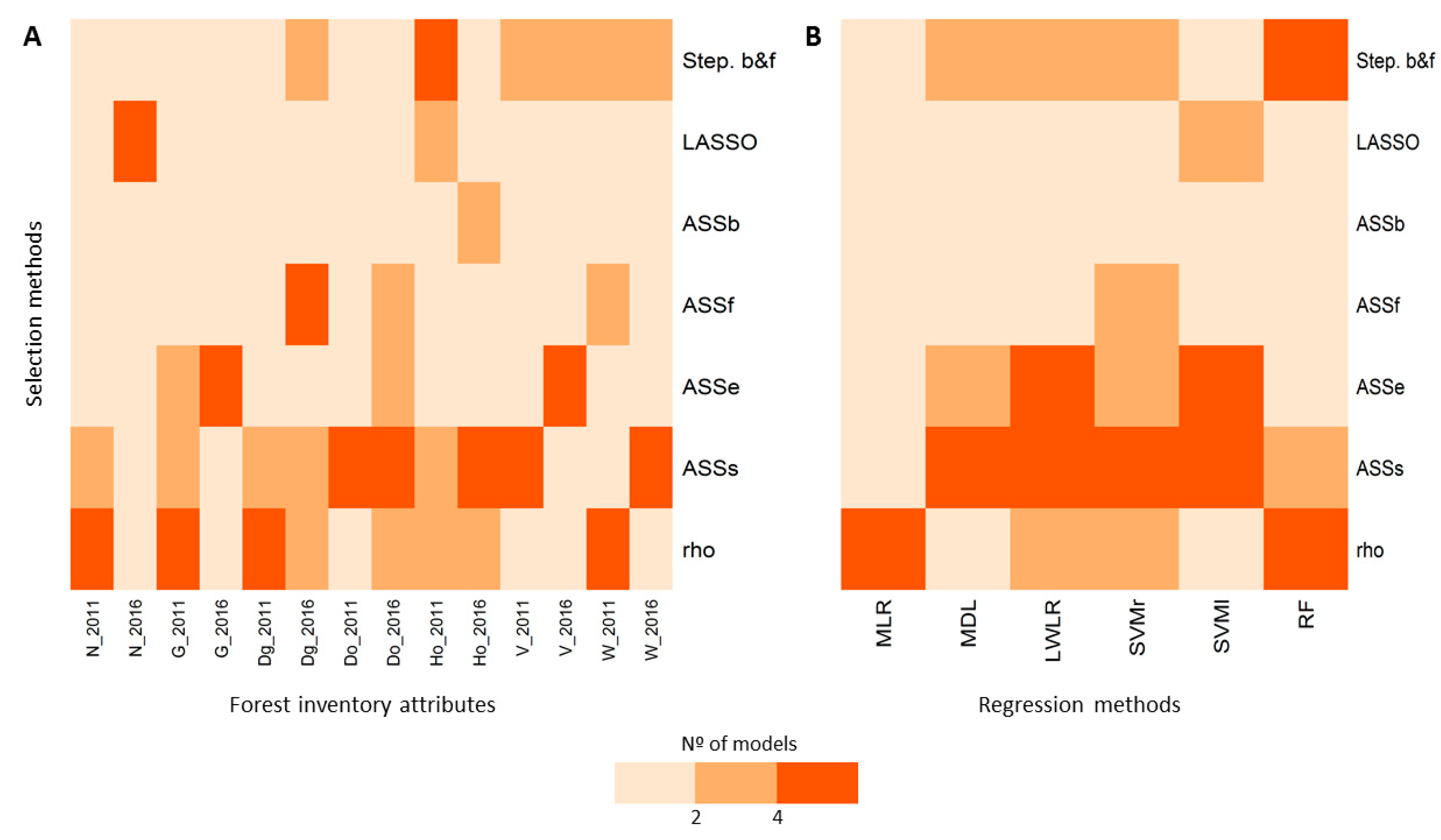
3.2. Variable Selection
3.3. Indirect Approach
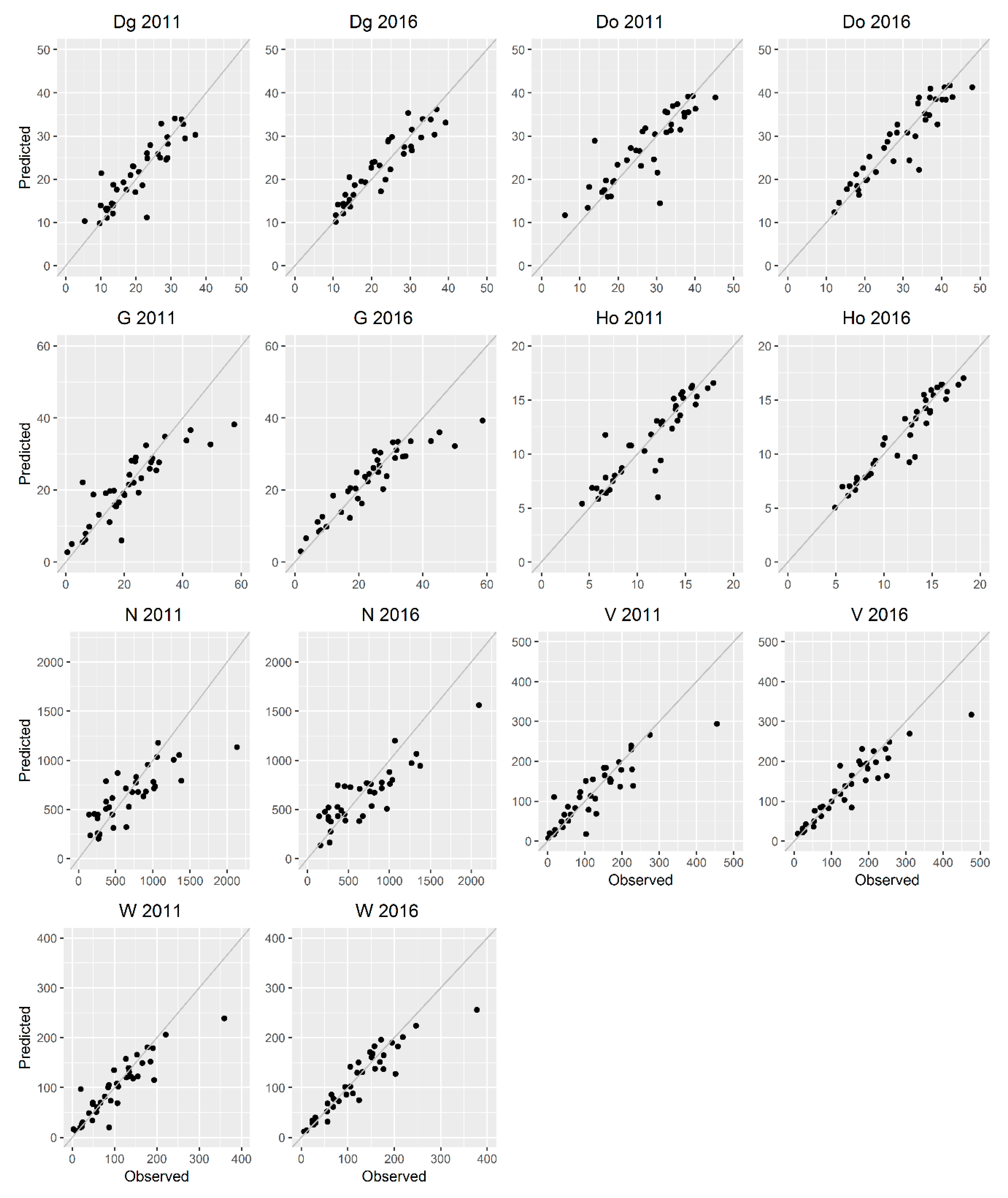
3.4. Direct Approach
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
| Macro-Classes | Classes | ALS Computed Metrics | Abbreviations |
|---|---|---|---|
| Canopy height metrics (CHM) | Lower height variables | Minimum elevation | Elev. minimum |
| 01th percentile of the return heights | P01 | ||
| 05th percentile of the return heights | P05 | ||
| 10th percentile of the return heights | P10 | ||
| 20th percentile of the return heights | P20 | ||
| 25th percentile of the return heights | P25 | ||
| L moment 1 elevation | Elev. L1 | ||
| L moment 2 elevation | Elev. L2 | ||
| Mean height variables | Mean elevation | Elev. Mean | |
| Mode elevation | Elev. Mode | ||
| 30th percentile of the return heights | P30 | ||
| 40th percentile of the return heights | P40 | ||
| 50th percentile of the return heights | P50 | ||
| 60th percentile of the return heights | P60 | ||
| 70th percentile of the return heights | P70 | ||
| L moment 3 elevation | Elev. L3 | ||
| Elevation quadratic mean | Elev. SQRT mean SQ | ||
| Elevation cubic mean | Elev. CUR mean CUBE | ||
| Higher height variables | 75th percentile of the return heights | P75 | |
| 80th percentile of the return heights | P80 | ||
| 90th percentile of the return heights | P90 | ||
| 95th percentile of the return heights | P95 | ||
| 99th percentile of the return heights | P99 | ||
| Maximum elevation | Elev. maximum | ||
| L moment 4 elevation | Elev. L4 | ||
| Canopy height variability metrics (CHVM) | Variability | Standard deviation of point heights distribution | Elev. SD |
| Variance of point heights distribution | Elev. Variance | ||
| Coefficient of variation of point heights distribution | Elev. CV | ||
| Skewness of point heights distribution | Elev. Skewness | ||
| kurtosis of point heights distribution | Elev. Kurtosis | ||
| Interquartile distance of point heights distribution | Elev. IQ | ||
| Average Absolute Deviation of point heights distribution | Elev. AAD | ||
| Variability L moment | L moment coefficient of variation of point heights distribution | Elev. LCV | |
| L moment skewness of point heights distribution | Elev. Lskewness | ||
| L moment kurtosis of point heights distribution | Elev. Lkurtosis | ||
| Canopy density metrics (CDM) | % first, % all returns, canopy relief ratio | percentage of first returns above the 2.00 | % first ret. above 2.00 |
| percentage of all returns above the 2.00 | % all ret. above 2.00 | ||
| percentage of first returns above the mean | % first ret. above mean | ||
| percentage of first returns above the mode | % first ret. above mode | ||
| percentage of all returns above the mean | % all ret. above mean | ||
| percentage of all returns above the mode | % all ret. above mode | ||
| Canopy relief ratio | CRR | ||
| All returns Total returns-1 | All returns above 2.00 divided by the total first returns × 100 | (All ret. above 2.00)/(total first ret.) × 100 | |
| All returns above mean divided by the total first returns × 100 | (All ret. above mean)/(total first ret.) × 100 | ||
| All returns above mode divided by the total first returns × 100 | (All ret. above mode)/(total first ret.) × 100 |
| Fitting Phase | Validation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ALS Metrics | Model | SM | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| P90 + (All ret. above mean)/(total first ret.) × 100 | MLR | Step. | 347.22 | 49.08 | 0.00 | 350.67 | 49.57 | 8.76 | 0.53 |
| Elev. L2 + Elev. Variance + (All ret. above 2.00)/(total first ret.) × 100 | MDL | ASSs | 235.89 | 33.34 | −0.83 | 292.37 | 41.33 | −1.48 | 0.68 |
| P99 + Elev. IQ + % first ret. above 2.00 | LWLR | Rho | 205.80 | 29.09 | −9.79 | 310.97 | 43.96 | −11.39 | 0.65 |
| P99 + Elev. IQ + % first ret. above 2.00 | SVMr | rho | 257.09 | 36.34 | 28.81 | 272.76 | 38.55 | 26.99 | 0.72 |
| Elev. L2 + Elev. Variance + (All ret. above 2.00)/(total first ret.) × 100 | SVMl | ASSs | 319.34 | 45.14 | 60.68 | 309.56 | 43.76 | 64.83 | 0.65 |
| P99 + Elev. SD + % first ret. above 2.00 | RF | rho | 151.86 | 21.46 | 1.91 | 303.56 | 42.91 | 6.91 | 0.66 |
| Fitting Phase | Validation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ALS Metrics | Model | SM | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| Elev. mean + Elev. L kurtosis + Canopy relief ratio | MLR | Step. | 358.84 | 51.47 | 0.00 | 363.62 | 52.15 | 11.57 | 0.45 |
| Elev. maximum + Elev. L kurtosis + % first ret. above 2.00 | MDL | LASSO | 243.13 | 34.87 | 1.14 | 322.89 | 46.31 | 8.15 | 0.61 |
| Elev. maximum + Elev. L kurtosis + % first ret. above 2.00 | LWLR | LASSO | 204.63 | 29.35 | 4.55 | 333.20 | 47.79 | 11.06 | 0.57 |
| Elev. maximum + Elev. L kurtosis + % first ret. above 2.00 | SVMr | LASSO | 250.87 | 35.98 | 13.95 | 278.58 | 39.96 | 11.83 | 0.67 |
| Elev. maximum + Elev. L kurtosis + % first ret. above 2.00 | SVMl | LASSO | 322.11 | 46.20 | 29.31 | 313.41 | 44.95 | 36.04 | 0.59 |
| Elev. maximum + Elev. L kurtosis + % first ret. above 2.00 | RF | LASSO | 159.15 | 22.83 | −1.71 | 302.57 | 43.40 | −10.81 | 0.60 |
| Fitting Phase | Validation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ALS Metrics | Model | SM | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| Elev. minimum + Elev. Kurtosis + (All ret. above mode)/(total first ret.) × 100 | MLR | rho | 5.80 | 29.80 | 0.00 | 6.01 | 30.89 | 0.19 | 0.64 |
| P10 + % first ret. above 2.00 | MDL | rho | 4.61 | 23.69 | 0.21 | 5.23 | 26.85 | 0.38 | 0.74 |
| P05 + % first ret. above mean | LWLR | ASSe | 4.07 | 20.92 | 0.01 | 5.53 | 28.42 | 0.12 | 0.70 |
| Elev. minimum + Elev. Kurtosis + (All ret. above mode)/(total first ret.) × 100 | SVMr | ASSs | 4.43 | 22.77 | −0.10 | 4.77 | 24.51 | −0.10 | 0.77 |
| Elev. minimum + Elev. Kurtosis + (All ret. above mode)/(total first ret.) × 100 | SVMl | ASSs | 4.85 | 24.92 | 0.10 | 4.87 | 25.05 | 0.05 | 0.75 |
| P10 + % first ret. above 2.00 | RF | rho | 2.61 | 13.41 | 0.02 | 5.19 | 26.69 | 0.06 | 0.73 |
| Fitting Phase | Validation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ALS Metrics | Model | SM | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| Elev. minimum +% all ret. above mode | MLR | rho | 9.27 | 42.11 | 0.00 | 9.19 | 41.76 | 0.21 | 0.15 |
| P75 + Elev. CUR mean CUBE + (All ret. above 2.00)/(total first ret.) × 100 | MDL | ASSe | 3.65 | 16.57 | 0.19 | 4.43 | 20.11 | 0.12 | 0.82 |
| P75 + Elev. CUR mean CUBE + (All ret. above 2.00)/(total first ret.) × 100 | LWLR | ASSe | 2.84 | 12.93 | 0.01 | 5.05 | 22.94 | −0.10 | 0.77 |
| P75 + Elev. CUR mean CUBE + (All ret. above 2.00)/(total first ret.) × 100 | SVMr | ASSe | 3.88 | 17.61 | 0.41 | 4.14 | 18.80 | 0.30 | 0.84 |
| P75 + Elev. CUR mean CUBE + (All ret. above 2.00)/(total first ret.) × 100 | SVMl | ASSe | 4.38 | 19.89 | 0.44 | 4.43 | 20.12 | 0.35 | 0.81 |
| P10 + Elev. minimum + % first ret. above mean | RF | ASSf | 2.32 | 10.56 | 0.08 | 4.64 | 21.06 | 0.37 | 0.81 |
| Fitting Phase | Validation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ALS Metrics | Model | SM | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| P90 + % first ret. above mean | MLR | rho | 3.80 | 18.44 | 0.00 | 3.84 | 18.60 | −0.03 | 0.77 |
| P90 + (All ret. above 2.00)/(total first ret.) × 100 | MDL | ASSs | 3.30 | 15.97 | 0.16 | 3.78 | 18.34 | 0.00 | 0.78 |
| P90 + (All ret. above 2.00)/(total first ret.) × 100 | LWLR | ASSs | 2.98 | 14.46 | −0.02 | 3.75 | 18.19 | −0.22 | 0.78 |
| P90 + Elev. Std.dev + % first ret. above mean | SVMr | rho | 3.38 | 16.38 | 0.19 | 3.56 | 17.25 | 0.06 | 0.81 |
| P90 + Elev. Std.dev + % first ret. above mean | SVMl | rho | 3.76 | 18.24 | 0.08 | 3.88 | 18.80 | −0.06 | 0.78 |
| P90 + Elev. Std.dev + (All ret. above mean)/(total first ret.) × 100 | RF | rho | 1.89 | 9.17 | −0.02 | 3.75 | 18.18 | −0.04 | 0.79 |
| Fitting Phase | Validation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ALS Metrics | Model | SM | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| P90 + Elev. LCV + (All ret. above mean)/(total first ret.) × 100 | MLR | Step. | 3.48 | 15.53 | 0.00 | 3.63 | 16.22 | −0.07 | 0.82 |
| Elev. maximum + Elev. IQ + (All ret. above 2.00)/(total first ret.) × 100 | MDL | ASSf | 3.12 | 13.93 | 0.23 | 3.71 | 16.58 | 0.21 | 0.82 |
| P90 + Elev. LCV + (All ret. above mean)/(total first ret.) × 100 | LWLR | Step. | 2.59 | 11.58 | −0.03 | 3.91 | 17.48 | −0.04 | 0.80 |
| Elev. maximum + Elev. IQ + (All ret. above 2.00)/(total first ret.) × 100 | SVMr | ASSf | 3.03 | 13.53 | 0.21 | 3.42 | 15.28 | 0.11 | 0.85 |
| P90 + Elev. mode + % first ret. above mode | SVMl | ASSs | 3.45 | 15.40 | 0.20 | 3.57 | 15.95 | 0.11 | 0.83 |
| P90 + Elev. LCV + (All ret. above mean)/(total first ret.) × 100 | RF | rho | 1.65 | 7.39 | 0.01 | 3.59 | 16.05 | 0.05 | 0.82 |
| Fitting Phase | Validation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ALS Metrics | Model | SM | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| Elev. skewness + Elev. Lkurtosis + P25 | MLR | rho | 5.34 | 20.17 | 0.00 | 5.42 | 20.47 | 0.07 | 0.62 |
| P90+ (All ret. above 2.00)/(total first ret.) × 100 | MDL | ASSf | 3.99 | 15.07 | 0.10 | 4.27 | 16.12 | 0.11 | 0.77 |
| P90+ (All ret. above 2.00)/(total first ret.) × 100 | LWLR | ASSf | 3.49 | 13.17 | −0.03 | 4.29 | 16.19 | −0.05 | 0.77 |
| P90+ (All ret. above 2.00)/(total first ret.) x 100 | SVMr | ASSf | 4.11 | 15.53 | 0.19 | 4.07 | 15.36 | 0.11 | 0.79 |
| P90+ (All ret. above 2.00)/(total first ret.) × 100 | SVMl | ASSf | 4.24 | 16.01 | 0.22 | 4.20 | 15.85 | 0.11 | 0.78 |
| P90 + % first ret. above mean | RF | rho | 2.13 | 8.04 | −0.11 | 4.38 | 16.55 | −0.36 | 0.76 |
| Fitting Phase | Validation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ALS Metrics | Model | SM | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| P90 + Elev. CV + (All ret. above mean)/(total first ret.) × 100 | MLR | Step. | 3.53 | 12.33 | 0.00 | 3.63 | 12.68 | −0.08 | 0.85 |
| P90 + Elev. variance + Elev. L2 | MDL | ASSs | 3.26 | 11.40 | 0.18 | 3.47 | 12.13 | 0.23 | 0.86 |
| P95 + Elev. CV | LWLR | ASSs | 2.88 | 10.07 | −0.03 | 3.63 | 12.70 | −0.09 | 0.84 |
| P95 + Elev. CV | SVMr | ASSs | 3.25 | 11.35 | 0.40 | 3.40 | 11.89 | 0.33 | 0.87 |
| Elev. Std.dev + Elev. Variance + P05 | SVMl | ASSe | 3.26 | 11.40 | 0.18 | 3.36 | 11.75 | 0.11 | 0.87 |
| P95 + Elev. CV | RF | ASSs | 1.61 | 5.64 | −0.02 | 3.62 | 12.64 | 0.00 | 0.85 |
| Fitting Phase | Validation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ALS Metrics | Model | SM | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| Elev. LCV + Elev. Lkurtosis + P01 | MLR | rho | 2.21 | 20.24 | 0.00 | 2.26 | 20.71 | 0.05 | 0.63 |
| P90 + Elev. kurtosis | MDL | Step. | 1.24 | 11.36 | 0.10 | 1.44 | 13.18 | 0.08 | 0.85 |
| P90 + Elev. skewness | LWLR | ASSs | 1.16 | 10.69 | −0.02 | 1.40 | 12.83 | 0.01 | 0.86 |
| P90 + Elev. variance + % All ret. above mean | SVMr | ASSf | 1.32 | 12.11 | 0.11 | 1.34 | 12.30 | 0.09 | 0.87 |
| Elev. L1 + Elev. maximum | SVMl | LASSO | 1.42 | 12.99 | 0.09 | 1.40 | 12.82 | 0.05 | 0.86 |
| P90 + Canopy relief ratio | RF | Step. | 0.72 | 6.65 | −0.01 | 1.46 | 13.41 | −0.06 | 0.84 |
| Fitting Phase | Validation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ALS Metrics | Model | SM | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| Elev. minimum + Elev. CV + Canopy relief ratio | MLR | rho | 2.41 | 20.90 | 0.00 | 2.52 | 21.91 | 0.01 | 0.51 |
| P95 + Elev. Std.dev | MDL | ASSs | 0.92 | 7.99 | 0.07 | 0.95 | 8.27 | 0.07 | 0.93 |
| P95 + Elev. variance | LWLR | ASSs | 0.79 | 6.87 | 0.02 | 0.98 | 8.48 | 0.00 | 0.93 |
| P95 + Elev. Std.dev | SVMr | ASSs | 0.86 | 7.48 | 0.03 | 1.02 | 8.83 | 0.03 | 0.92 |
| P90 + Elev. variance + Elev. SQRT mean SQ | SVMl | ASSb | 0.96 | 8.30 | 0.12 | 1.00 | 8.69 | 0.08 | 0.93 |
| P95 + Elev. variance + (All ret. above mean)/(total first ret.) × 100 | RF | rho | 0.43 | 3.76 | −0.01 | 1.00 | 8.72 | −0.03 | 0.92 |
| Fitting Phase | Validation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ALS Metrics | Model | SM | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| P20 + (All ret. above 2.00)/(total first ret.) × 100 | MDL | ASSs | 30.15 | 28.63 | 1.69 | 33.39 | 31.71 | 2.13 | 0.81 |
| P20 + (All ret. above 2.00)/(total first ret.) × 100 | LWLR | ASSs | 25.89 | 24.58 | 0.05 | 34.09 | 32.37 | 0.24 | 0.80 |
| Elev. L2 + Elev. CUR mean CUBE + % first ret. above mean | SVMr | Step. | 28.87 | 27.42 | 2.59 | 29.71 | 28.22 | 1.79 | 0.84 |
| P20 + Elev. L skewness + (All ret. above mean)/(total first ret.) × 100 | SVMl | ASSs | 34.25 | 32.52 | 0.88 | 34.30 | 32.58 | 0.09 | 0.79 |
| P20 + Elev. L skewness + % first ret. above 2.00 | RF | ASSs | 16.80 | 15.96 | 0.17 | 34.28 | 32.55 | −0.56 | 0.78 |
| Fitting Phase | Validation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ALS Metrics | Model | SM | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| P75 + Elev. CUR mean CUBE + (All ret. above 2.00)/(total first ret.) × 100 | MDL | ASSe | 24.87 | 20.02 | −0.34 | 29.63 | 23.85 | −0.19 | 0.88 |
| P75 + Elev. CUR mean CUBE + (All ret. above 2.00)/(total first ret.) × 100 | LWLR | ASSe | 20.26 | 16.30 | −0.08 | 31.80 | 25.59 | −0.06 | 0.85 |
| P75 + Elev. CUR mean CUBE + (All ret. above 2.00)/(total first ret.) × 100 | SVMr | ASSe | 24.69 | 19.87 | 2.65 | 26.35 | 21.20 | 1.92 | 0.90 |
| P75 + Elev. CUR mean CUBE + (All ret. above 2.00)/(total first ret.) × 100 | SVMl | ASSe | 30.49 | 24.54 | 2.60 | 31.14 | 25.06 | 1.48 | 0.86 |
| Elev. L2 + Elev. CUR mean CUBE + % first ret. above 2.00 | RF | Step. | 15.25 | 12.27 | −0.38 | 31.73 | 25.53 | 0.32 | 0.86 |
| Fitting Phase | Validation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ALS Metrics | Model | SM | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| Elev. L2 + Elev. CUR mean CUBE + % first ret. above 2.00 | MDL | Step. | 22.02 | 24.21 | 0.68 | 28.30 | 31.12 | 1.82 | 0.76 |
| P10 + Elev. CUR mean CUBE + % first ret. above mean | LWLR | rho | 18.34 | 20.17 | 0.23 | 29.12 | 32.02 | 0.04 | 0.74 |
| P10 + Elev. SQRT mean SQ + (All ret. above mean)/(total first ret.) × 100 | SVMr | rho | 23.00 | 25.29 | 0.75 | 24.29 | 26.71 | −0.03 | 0.82 |
| P10 + Canopy relief ratio + (All ret. above mean)/(total first ret.) × 100 | SVMl | ASSf | 26.60 | 29.25 | 0.50 | 26.82 | 29.49 | 0.11 | 0.79 |
| P10 + Elev. CUR mean CUBE + (All ret. above mean)/(total first ret.) × 100 | RF | rho | 14.39 | 15.83 | −0.05 | 29.43 | 32.36 | 0.24 | 0.75 |
| Fitting Phase | Validation | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ALS Metrics | Model | SM | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| P75 + Elev. CUR mean CUBE + (All ret. above 2.00)/(total first ret.) × 100 | MDL | ASSe | 19.66 | 18.44 | −0.64 | 23.44 | 21.98 | −0.43 | 0.88 |
| P75 + Elev. CUR mean CUBE + (All ret. above 2.00)/(total first ret.) × 100 | LWLR | ASSe | 16.11 | 15.11 | −0.11 | 25.75 | 24.15 | 0.18 | 0.85 |
| Elev. L2 + Elev. CUR mean CUBE + % first ret. above 2.00 | SVMr | Step. | 18.82 | 17.65 | 1.23 | 20.06 | 18.81 | 0.56 | 0.90 |
| P75 + Elev. CUR mean CUBE + (All ret. above 2.00)/(total first ret.) × 100 | SVMl | ASSe | 22.78 | 21.37 | 1.65 | 23.43 | 21.98 | 0.80 | 0.87 |
| P20 + Elev. CUR mean CUBE + All ret. above 2.00)/(total first ret.) × 100 | RF | Step. | 12.58 | 11.80 | 0.13 | 22.38 | 20.99 | 0.01 | 0.87 |

References
- Lal, R. Sequestration of atmospheric CO2 in global carbon pools. Energy Environ. Sci. 2008, 1, 86–100. [Google Scholar] [CrossRef]
- Pan, Y.; Birdsey, R.A.; Fang, J.; Houghton, R.; Kauppi, P.E.; Kurz, W.A.; Phillips, O.L.; Shvidenko, A.; Lewis, S.L.; Canadell, J.G.; et al. A large and persistent carbon sink in the world’s forests. Science 2011, 333, 988–993. [Google Scholar] [CrossRef] [PubMed]
- Zhao, K.; Suarez, J.C.; Garcia, M.; Hu, T.; Wang, C.; Londo, A. Utility of multitemporal lidar for forest and carbon monitoring: Tree growth, biomass dynamics, and carbon flux. Remote Sens. Environ. 2018, 204, 883–897. [Google Scholar] [CrossRef]
- Zald, H.S.J.; Wulder, M.A.; White, J.C.; Hilker, T.; Hermosilla, T.; Hobart, G.W.; Coops, N.C. Integrating Landsat pixel composites and change metrics with lidar plots to predictively map forest structure and aboveground biomass in Saskatchewan, Canada. Remote Sens. Environ. 2016, 176, 188–201. [Google Scholar] [CrossRef]
- Pflugmacher, D.; Cohen, W.B.; Kennedy, R.E.; Yang, Z. Using Landsat-derived disturbance and recovery history and lidar to map forest biomass dynamics. Remote Sens. Environ. 2014, 151, 124–137. [Google Scholar] [CrossRef]
- Ferraz, A.; Saatchi, S.; Bormann, K.J.; Painter, T.H. Fusion of NASA Airborne Snow Observatory (ASO) Lidar Time Series over Mountain Forest Landscapes. Remote Sens. 2018, 10, 164. [Google Scholar] [CrossRef]
- Hummel, S.; Hudak, A.T.; Uebler, E.H.; Falkowski, M.J.; Megown, K.A. A Comparison of Accuracy and Cost of LiDAR versus Stand Exam Data for Landscape Management on the Malheur National Forest. J. For. 2011, 109, 267–273. [Google Scholar]
- Hudak, A.T.; Evans, J.S.; Stuart Smith, A.M. LiDAR Utility for Natural Resource Managers. Remote Sens. 2009, 1, 934–951. [Google Scholar] [CrossRef]
- Cao, L.; Coops, N.C.; Innes, J.L.; Sheppard, S.R.J.; Fu, L.; Ruan, H.; She, G. Estimation of forest biomass dynamics in subtropical forests using multi-temporal airborne LiDAR data. Remote Sens. Environ. 2016, 178, 158–171. [Google Scholar] [CrossRef]
- Dubayah, R.O.; Sheldon, S.L.; Clark, D.B.; Hofton, M.A.; Blair, J.B.; Hurtt, G.C.; Chazdon, R.L. Estimation of tropical forest height and biomass dynamics using lidar remote sensing at la Selva, Costa Rica. J. Geophys. Res. Biogeosci. 2010, 115, 1–17. [Google Scholar] [CrossRef]
- Stoker, J.M.; Harding, D.; Parrish, J. The need for a national LIDAR dataset. Photogramm. Eng. Remote Sensing 2008, 74, 1066–1068. [Google Scholar]
- Marinelli, D.; Paris, C.; Bruzzone, L. A Novel Approach to 3-D Change Detection in Multitemporal LiDAR Data Acquired in Forest Areas. IEEE Trans. Geosci. Remote Sens. 2018, 56, 3030–3046. [Google Scholar] [CrossRef]
- Fekety, P.A.; Falkowski, M.J.; Hudak, A.T. Temporal transferability of LiDAR-based imputation of forest inventory attributes. Can. J. Forest Res. 2015, 45, 422–435. [Google Scholar] [CrossRef]
- Socha, J.; Pierzchalski, M.; Bałazy, R.; Ciesielski, M. Modelling top height growth and site index using repeated laser scanning data. For. Ecol. Manag. 2017, 406, 307–317. [Google Scholar] [CrossRef]
- Gatziolis, D.; Fried, J.S.; Monleon, V.S. Challenges to Estimating Tree Height via LiDAR in Closed-Canopy Forests: A Parable from Western Oregon. For. Sci. 2010, 56, 139–155. [Google Scholar]
- Hopkinson, C.; Chasmer, L.; Hall, R.J. The uncertainty in conifer plantation growth prediction from multi-temporal lidar datasets. Remote Sens. Environ. 2008, 112, 1168–1180. [Google Scholar] [CrossRef]
- Næsset, E.; Gobakken, T. Estimating forest growth using canopy metrics derived from airborne laser scanner data. Remote Sens. Environ. 2005, 96, 453–465. [Google Scholar] [CrossRef]
- Yu, X.; Hyyppä, J.; Kaartinen, H.; Maltamo, M.; Hyyppä, H. Obtaining plotwise mean height and volume growth in boreal forests using multi-temporal laser surveys and various change detection techniques. Int. J. Remote Sens. 2008, 29, 1367–1386. [Google Scholar] [CrossRef]
- Yu, X.; Hyyppä, J.; Hyyppä, H.; Maltamo, M. Effects of flight altitude on tree height estimation using airborne laser scanning. ISPRS Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2004, XXXVI, 96–101. [Google Scholar]
- St-Onge, B.; Vepakomma, U. Assessing forest gap dynamics and growth using multi-temporal laser-scanner data. ISPRS Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2004, XXXVI, Par, 173–178. [Google Scholar]
- Poudel, K.P.; Flewelling, J.W.; Temesgen, H. Predicting Volume and Biomass Change from Multi-Temporal Lidar Sampling and Remeasured Field Inventory Data in Panther Creek Watershed, Oregon, USA. Forests 2018, 9, 28. [Google Scholar] [CrossRef]
- Temesgen, H.; Strunk, J.; Andersen, H.E.; Flewelling, J. Evaluating different models to predict biomass increment from multi-temporal lidar sampling and remeasured field inventory data in south-central Alaska. Math. Comput. For. Nat. Res. Sci. 2015, 7, 66–80. [Google Scholar]
- Réjou-Méchain, M.; Tymen, B.; Blanc, L.; Fauset, S.; Feldpausch, T.R.; Monteagudo, A.; Phillips, O.L.; Richard, H.; Chave, J. Using repeated small-footprint LiDAR acquisitions to infer spatial and temporal variations of a high-biomass Neotropical forest. Remote Sens. Environ. 2015, 169, 93–101. [Google Scholar] [CrossRef]
- Skowronski, N.S.; Clark, K.L.; Gallagher, M.; Birdsey, R.A.; Hom, J.L. Airborne laser scanner-assisted estimation of aboveground biomass change in a temperate oak–pine forest. Remote Sens. Environ. 2014, 151, 166–174. [Google Scholar] [CrossRef]
- Meyer, V.; Saatchi, S.S.; Chave, J.; Dalling, J.W.; Bohlman, S.; Fricker, G.A.; Robinson, C.; Neumann, M.; Hubbell, S. Detecting tropical forest biomass dynamics from repeated airborne lidar measurements. Biogeosciences 2013, 10, 5421–5438. [Google Scholar] [CrossRef]
- Bollandsås, O.M.; Gregoire, T.G.; Næsset, E.; Øyen, B.H. Detection of biomass change in a Norwegian mountain forest area using small footprint airborne laser scanner data. Stat. Methods Appl. 2013, 22, 113–129. [Google Scholar] [CrossRef]
- Hudak, A.T.; Strand, E.K.; Vierling, L.A.; Byrne, J.C.; Eitel, J.U.H.; Martinuzzi, S.; Falkowski, M.J. Quantifying aboveground forest carbon pools and fluxes from repeat LiDAR surveys. Remote Sens. Environ. 2012, 123, 25–40. [Google Scholar] [CrossRef]
- Noordermeer, L.; Bollandsås, O.M.; Gobakken, T.; Næsset, E. Direct and indirect site index determination for Norway spruce and Scots pine using bitemporal airborne laser scanner data. For. Ecol. Manag. 2018, 428, 104–114. [Google Scholar] [CrossRef]
- Mccarley, T.R.; Kolden, C.A.; Vaillant, N.M.; Hudak, A.T.; Smith, A.M.S.; Wing, B.M.; Kellogg, B.S.; Kreitler, J. Multi-temporal LiDAR and Landsat quantification of fire-induced changes to forest structure. Remote Sens. Environ. 2017, 419–432. [Google Scholar] [CrossRef]
- Vepakomma, U.; Kneeshaw, D.; St-Onge, B. Interactions of multiple disturbances in shaping boreal forest dynamics: A spatially explicit analysis using multi-temporal lidar data and high-resolution imagery. J. Ecol. 2010, 98, 526–539. [Google Scholar] [CrossRef]
- Vepakomma, U.; St-Onge, B.; Kneeshaw, D. Spatially explicit characterization of boreal forest gap dynamics using multi-temporal lidar data. Remote Sens. Environ. 2008, 112, 2326–2340. [Google Scholar] [CrossRef]
- Solberg, S.; Næsset, E.; Hanssen, K.H.; Christiansen, E. Mapping defoliation during a severe insect attack on Scots pine using airborne laser scanning. Remote Sens. Environ. 2006, 102, 364–376. [Google Scholar] [CrossRef]
- Guerra-Hernández, J.; Görgens, E.B.; García-Gutiérrez, J.; Rodriguez, L.C.E.; Tomé, M.; González-Ferreiro, E. Comparison of ALS based models for estimating aboveground biomass in three types of Mediterranean forest. Eur. J. Remote Sens. 2016, 49, 185. [Google Scholar] [CrossRef]
- Mehtatalo, L.; Virolainen, A.; Tuomela, J.; Packalen, P. Estimating Tree Height Distribution Using Low-Density ALS Data With and Without Training Data. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2015, 8, 1432–1441. [Google Scholar] [CrossRef]
- Shendryk, I.; Hellström, M.; Klemedtsson, L.; Kljun, N. Low-Density LiDAR and Optical Imagery for Biomass Estimation over Boreal Forest in Sweden. Forests 2014, 5, 992–1010. [Google Scholar] [CrossRef]
- Domingo, D.; Lamelas, M.; Montealegre, A.; García-Martín, A.; de la Riva, J.; Domingo, D.; Lamelas, M.T.; Montealegre, A.L.; García-Martín, A.; de la Riva, J. Estimation of Total Biomass in Aleppo Pine Forest Stands Applying Parametric and Nonparametric Methods to Low-Density Airborne Laser Scanning Data. Forests 2018, 9, 158. [Google Scholar] [CrossRef]
- Montealegre, A.L.; Lamelas, M.T.; de la Riva, J.; García-Martín, A.; Escribano, F. Use of low point density ALS data to estimate stand-level structural variables in Mediterranean Aleppo pine forest. Forestry 2016, 89, 373–382. [Google Scholar] [CrossRef]
- Garcia-Gutierrez, J.; Gonzalez-Ferreiro, E.; Riquelme-Santos, J.C.; Miranda, D.; Dieguez-Aranda, U.; Navarro-Cerrillo, R.M. Evolutionary feature selection to estimate forest stand variables using LiDAR. Int. J. Appl. Earth Obs. Geoinf. 2014, 26, 119–131. [Google Scholar] [CrossRef]
- Jakubowski, M.K.; Guo, Q.; Kelly, M. Tradeoffs between lidar pulse density and forest measurement accuracy. Remote Sens. Environ. 2013, 130, 245–253. [Google Scholar] [CrossRef]
- Gobakken, T.; Næsset, E. Assessing effects of laser point density, ground sampling intensity, and field sample plot size on biophysical stand properties derived from airborne laser scanner data. Can. J. For. Res. 2008, 38, 1095–1109. [Google Scholar] [CrossRef]
- Lovell, J.L.; Jupp, D.L.B.; Newnham, G.J.; Coops, N.C.; Culvenor, D.S. Simulation study for finding optimal lidar acquisition parameters for forest height retrieval. For. Ecol. Manag. 2005, 214, 398–412. [Google Scholar] [CrossRef]
- Roussel, J.-R.; Caspersen, J.; Béland, M.; Thomas, S.; Achim, A. Removing bias from LiDAR-based estimates of canopy height: Accounting for the effects of pulse density and footprint size. Remote Sens. Environ. 2017, 198, 1–16. [Google Scholar] [CrossRef]
- Garcia, M.; Saatchi, S.; Casas, A.; Koltunov, A.; Ustin, S.; Ramirez, C.; Garcia-Gutierrez, J.; Balzter, H. Quantifying biomass consumption and carbon release from the California Rim fire by integrating airborne LiDAR and Landsat OLI data. J. Geophys. Res. Biogeosci. 2017. [Google Scholar] [CrossRef]
- Singh, K.K.; Chen, G.; McCarter, J.B.; Meentemeyer, R.K. Effects of LiDAR point density and landscape context on estimates of urban forest biomass. ISPRS J. Photogramm. Remote Sens. 2015, 101, 310–322. [Google Scholar] [CrossRef]
- Ruiz, L.; Hermosilla, T.; Mauro, F.; Godino, M.; Ruiz, L.A.; Hermosilla, T.; Mauro, F.; Godino, M. Analysis of the Influence of Plot Size and LiDAR Density on Forest Structure Attribute Estimates. Forests 2014, 5, 936–951. [Google Scholar] [CrossRef]
- García, O. Dimensionality reduction in growth models: An example. FBMIS 2003, 1, 1–15. [Google Scholar]
- Thornley, J. Modelling forest ecosystems: The Edinburgh Forest Model. In Forest Sustainability: Theory and Practice; CAB International: Wallingford, UK, 2006. [Google Scholar]
- Montero, G.; Serrada, R. La situación de los bosques y el sector forestal en España - ISFE 2013; Sociedad Española de Ciencias Forestales: Lourizán (Pontevedra), Spain, 2013. [Google Scholar]
- Cuadrat, J.M.; Saz, M.A.; Vicente-Serrano, S.M. Atlas Climático de Aragón; Gobierno de Aragón: Aragón, Zaragoza, Spain, 2007. [Google Scholar]
- Granados, M.E.; Vilagrosa, A.; Chirino, E.; Vallejo, V.R. Reforestation with resprouter species to increase diversity and resilience in Mediterranean pine forests. For. Ecol. Manag. 2016, 362, 231–240. [Google Scholar] [CrossRef]
- Gasque, M.; García-Fayos, P. Interaction between Stipa tenacissima and Pinus halepensis: Consequences for reforestation and the dynamics of grass steppes in semi-arid Mediterranean areas. For. Ecol. Manag. 2004, 189, 251–261. [Google Scholar] [CrossRef]
- Hernandez-Tecles, E.; Osem, Y.; Alfaro-Sanchez, R.; de las Heras, J. Vegetation structure of planted versus natural Aleppo pine stands along a climatic gradient in Spain. Ann. For. Sci. 2015, 72, 641–650. [Google Scholar] [CrossRef]
- Hair, J.F.; Prentice, E.; Cano, D. Anaálisis Multivariante; Prentice-Hall: Madrid, Spain, 1999; pp. 1–795. ISBN 9788483220351. [Google Scholar]
- Næsset, E. Predicting forest stand characteristics with airborne scanning laser using a practical two-stage procedure and field data. Remote Sens. Environ. 2002, 80, 88–99. [Google Scholar] [CrossRef]
- Alonso Ponce SIMANFOR - Sistema de Simulación de Manejo Forestal Sostenible. Gestión Forestal Sostenible 2018. Available online: http://sostenible.palencia.uva.es/content/simanfor-sistema-de-simulacion-de-manejo-forestal-sostenible (accessed on 28 September 2018).
- Bravo, F.; Rodriguez, F.; Ordóñez, C. A web-based application to simulate alternatives for sustainable forest management: SIMANFOR. For. Syst. 2012, 21, 4. [Google Scholar] [CrossRef]
- Schröder, J.; Gadow, K. von Testing a new competition index for Maritime pine in northwestern Spain. Can. J. For. Res. 1999, 29, 280–283. [Google Scholar] [CrossRef]
- Rojo-Alboreca, A.; Cabanillas-Saldaña, A.M.; Barrio-Anta, M.; Notivol-Paíno, E.; Gorgoso-Varela, J.J. Site index curves for natural Aleppo pine forests in the central Ebro valley (Spain). Madera y Bosques 2017, 23, 143. [Google Scholar] [CrossRef]
- Ruiz-Peinado, R.; Del Rio, M.; Montero, G. New models for estimating the carbon sink capacity of Spanish softwood species. For. Syst. 2011, 20, 176. [Google Scholar] [CrossRef]
- PNOA Plan Nacional de Ortofotografía Aérea. Available online: http://pnoa.ign.es/presentacion (accessed on 12 April 2016).
- Evans, J.S.; Hudak, A.T. A Multiscale Curvature Algorithm for Classifying Discrete Return LiDAR in Forested Environments. IEEE Trans. Geosci. Remote Sens. 2007, 45, 1029–1038. [Google Scholar] [CrossRef]
- Montealegre, A.; Lamelas, M.; Riva, J. Interpolation Routines Assessment in ALS-Derived Digital Elevation Models for Forestry Applications. Remote Sens. 2015, 7, 8631–8654. [Google Scholar] [CrossRef]
- Evans, J.S.; Hudak, A.T.; Faux, R.; Smith, A.M.S. Discrete Return Lidar in Natural Resources: Recommendations for Project Planning, Data Processing, and Deliverables. Remote Sens. 2009, 1, 776–794. [Google Scholar] [CrossRef]
- McGaughey, R. FUSION/LDV: Software for LIDAR data analysis and visualization - V3.10. USDA For Serv. 2014, 1–212. [Google Scholar]
- Nilsson, M. Estimation of tree heights and stand volume using an airborne lidar system. Remote Sens. Environ. 1996, 56, 1–7. [Google Scholar] [CrossRef]
- Næsset, E.; Økland, T. Estimating tree height and tree crown properties using airborne scanning laser in a boreal nature reserve. Remote Sens. Environ. 2002, 79, 105–115. [Google Scholar] [CrossRef]
- Darlington, R.B.; Horst, P. Factor Analysis of Data Matrices. Am. J. Psychol. 1966, 79, 344–346. [Google Scholar] [CrossRef]
- Kaiser, H.F. The varimax criterion for analytic rotation in factor analysis. Psychometrika 1958, 23, 187–200. [Google Scholar] [CrossRef]
- Tibshirani, R. Regression shrinkage and selection via the lasso: A retrospective. J. R. Stat. Soc. Stat. Methodol. Ser. B 2011, 73, 273–282. [Google Scholar] [CrossRef]
- Miller, A.J. Subset Selection in Regression. In Monographs on Statistics and Applied Probability 95; Isham, V., Keiding, T., Louis, N., Tibshirani, R.R., Tong, H., Eds.; Chapman & Hall/CRC: New York, NY, USA, 2002; p. 256. Available online: https://ncss-wpengine.netdna-ssl.com/wp-content/themes/ncss/pdf/Procedures/NCSS/Subset_Selection_in_Multiple_Regression.pdf (accessed on 23 January 2019).
- Hermosilla, T.; Ruiz, L.A.; Kazakova, A.N.; Coops, N.C.; Moskal, L.M. Estimation of forest structure and canopy fuel parameters from small-footprint full-waveform LiDAR data. Int. J. Wildand Fire 2014, 23, 224. [Google Scholar] [CrossRef]
- Kumar, L.; Sinha, P.; Taylor, S.; Alqurashi, A.F. Review of the use of remote sensing for biomass estimation to support renewable energy generation. J. Appl. Remote Sens. 2015, 9, 097696. [Google Scholar] [CrossRef]
- Means, J.; Acker, S.; Harding, D.; Blair, J.; Lefsky, M.; Cohen, W.; Harmon, M.; McKee, W. Use of large-footprint scanning airborne lidar to estimate forest stand characteristics in the western Cascades of Oregon. Remote Sens. Environ. 1999, 67, 298–308. [Google Scholar] [CrossRef]
- García, D.; Godino, M.; Mauro, F. Lidar: Aplicación Práctica Al Inventario Forestal; Editorial Academica Española: Madrid, Spain, 2012; p. 196. ISBN 3659009555. [Google Scholar]
- Mountrakis, G.; Im, J.; Ogole, C. Support vector machines in remote sensing: A review. ISPRS J. Photogramm. Remote Sens. 2011, 66, 247–259. [Google Scholar] [CrossRef]
- Liaw, A.; Wiener, M. Classification and Regression by randomForest. R News 2002, 2, 18–22. [Google Scholar]
- Van Essen, D.C.; Drury, H.A.; Dickson, J.; Harwell, J.; Hanlon, D.; Anderson, C.H. An Integrated Software Suite for Surface-based Analyses of Cerebral Cortex. J. Am. Med. Inf. Assoc. 2001, 8, 443–459. [Google Scholar] [CrossRef]
- García-Gutiérrez, J.; Martínez-Álvarez, F.; Troncoso, A.; Riquelme, J.C. A comparison of machine learning regression techniques for LiDAR-derived estimation of forest variables. Neurocomputing 2015, 167, 24–31. [Google Scholar] [CrossRef]
- Görgens, E.B.; Montaghi, A.; Rodriguez, L.C.E. A performance comparison of machine learning methods to estimate the fast-growing forest plantation yield based on laser scanning metrics. Comput. Electron. Agric. 2015, 116, 221–227. [Google Scholar] [CrossRef]
- Stojanova, D.; Panov, P.; Gjorgjioski, V.; Kobler, A.; Džeroski, S. Estimating vegetation height and canopy cover from remotely sensed data with machine learning. Ecol. Inf. 2010, 5, 256–266. [Google Scholar] [CrossRef]
- Nemenyi, P. Distribution-Free Multiple Comparisons. Ph.D. Thesis, Princeton University, Princenton, NY, USA, 1963. [Google Scholar]
- Watt, M.S.; Meredith, A.; Watt, P.; Gunn, A.; Andersen, H.; Reutebuch, S.; Nelson, R. Use of LiDAR to estimate stand characteristics for thinning operations in young Douglas-fir plantations. N. Z. J. For. Sci. 2013, 43, 1–10. [Google Scholar] [CrossRef]
- Amaro, A.; Reed, D.; Soares, P. Modelling Forest Systems; CABI publishing: Lisbon, Portugal, 2003; Available online: https://bit.ly/2FNPtVj (accessed on 23 January 2019).
- Hansen, E.; Gobakken, T.; Bollandsås, O.; Zahabu, E.; Næsset, E.; Hansen, E.H.; Gobakken, T.; Bollandsås, O.M.; Zahabu, E.; Næsset, E. Modeling Aboveground Biomass in Dense Tropical Submontane Rainforest Using Airborne Laser Scanner Data. Remote Sens. 2015, 7, 788–807. [Google Scholar] [CrossRef]
- Kristensen, T.; Næsset, E.; Ohlson, M.; Bolstad, P.V.; Kolka, R. Mapping Above- and Below-Ground Carbon Pools in Boreal Forests: The Case for Airborne Lidar. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Silva, C.A.; Klauberg, C.; Hudak, A.T.; Vierling, L.A.; Liesenberg, V.; Carvalho, S.P.C.E.; Rodriguez, L.C.E. A principal component approach for predicting the stem volume in Eucalyptus plantations in Brazil using airborne LiDAR data. Forestry 2016, 89. [Google Scholar] [CrossRef]
- Hopkinson, C.; Chasmer, L.; Barr, A.G.; Kljun, N.; Black, T.A.; McCaughey, J.H. Monitoring boreal forest biomass and carbon storage change by integrating airborne laser scanning, biometry and eddy covariance data. Remote Sens. Environ. 2016, 181, 82–95. [Google Scholar] [CrossRef]
- Guerra-Hernandez, J.; Gonzalez-Ferreiro, E.; Sarmento, A.; Silva, J.; Nunes, A.; Correia, A.C.; Fontes, L.; Tomé, M.; Diaz-Varela, R. Short Communication. Using high resolution UAV imagery to estimate tree variables in Pinus pinea plantation in Portugal. For. Syst. 2016, 25. [Google Scholar] [CrossRef]
- Domingo, D.; Lamelas-Gracia, M.T.; Montealegre-Gracia, A.L.; de la Riva-Fernández, J. Comparison of regression models to estimate biomass losses and CO2 emissions using low-density airborne laser scanning data in a burnt Aleppo pine forest. Eur. J. Remote Sens. 2017, 50, 384–396. [Google Scholar] [CrossRef]
- Gagliasso, D.; Hummel, S.; Temesgen, H. A comparison of selected parametric and non-parametric imputation methods for estimating forest biomass and basal area. Open J. For. 2014, 4, 42–48. [Google Scholar] [CrossRef]
| Date of the Campaign | Field Data | Variables | Units |
|---|---|---|---|
| First: June to July 2013 | Green crown height Total height Dbh | Stand density (N) Basal area (G) Squared mean diameter (Dg) Dominant diameter (Do) Dominant height (Ho) Volume over bark (V) Total tree biomass (W) | stems ha−1 m2 ha−1 cm cm m m3 ha−1 tons ha−1 |
| Second: July to September 2014 | |||
| Third: April 2016 |
| Characteristics | Year 2011 | Year 2016 |
|---|---|---|
| Time period | January to February | September to November |
| Laser scanning system | Leica ALS60 | Leica ALS80 |
| Wavelength | 1,064 nm | 1064 nm |
| Average flying altitude over sea level | 3,000 m | 3150 m |
| Pulse repetition frequency | ~70 kHz | 176–286 kHz |
| Scanning frequency | ~45 kHz | 28–59 Hz |
| Maximum scan angle | 29° | 25° |
| Nominal point density | 0.5 points m−2 | 1 points m−2 |
| Average point density | 0.64 points m−2 | 1.25 points m−2 |
| Accuracy of the point cloud (RMSEz) | ≤0.2 m | 0.09 m |
| Forest Inventory Attribute | Min. | Max. | Range | Mean | Standard Deviation |
|---|---|---|---|---|---|
| N (stems ha−1) | 99.03 | 3200.00 | 3100.97 | 715.61 | 486.54 |
| G (m2 ha−1) | 0.82 | 58.89 | 58.07 | 21.47 | 10.04 |
| Dg (cm) | 9.04 | 43.52 | 34.48 | 21.67 | 8.01 |
| Do (cm) | 9.21 | 47.96 | 38.76 | 27.79 | 8.73 |
| Ho (m) | 4.69 | 18.90 | 14.21 | 11.32 | 3.54 |
| V (m3 ha−1) | 2.21 | 467.62 | 465.41 | 118.71 | 77.79 |
| W (tons ha−1) | 2.89 | 373.02 | 370.14 | 101.91 | 60.69 |
| Inventory Attribute | Min. 2011 | Min. 2016 | Max. 2011 | Max. 2016 | Range 2011 | Range 2016 | Mean 2011 | Mean 2016 | SD 2011 | SD 2016 |
|---|---|---|---|---|---|---|---|---|---|---|
| N (stems ha−1) | 99.03 | 99.03 | 3405.67 | 3161.81 | 3306.64 | 3062.79 | 709.64 | 699.20 | 500.86 | 481.00 |
| G (m2 ha−1) | 0.11 | 0.91 | 57.56 | 58.69 | 57.45 | 57.77 | 19.71 | 22.26 | 9.97 | 10.14 |
| Dg (cm) | 3.29 | 9.55 | 41.41 | 45.05 | 38.12 | 35.50 | 20.72 | 22.45 | 7.99 | 8.40 |
| Do (cm) | 3.35 | 9.72 | 45.85 | 49.19 | 42.50 | 39.47 | 26.59 | 28.72 | 8.84 | 9.09 |
| Ho (m) | 4.24 | 4.90 | 18.46 | 19.08 | 14.22 | 14.17 | 10.97 | 11.58 | 3.70 | 3.60 |
| V (m3 ha−1) | 0.35 | 2.51 | 454.77 | 476.02 | 454.42 | 473.51 | 107.31 | 126.45 | 74.83 | 81.48 |
| W (tons ha−1) | 1.34 | 3.14 | 359.22 | 377.82 | 357.88 | 374.68 | 92.46 | 108.26 | 58.10 | 63.63 |
| Fitting Phase | Validation | |||||||
|---|---|---|---|---|---|---|---|---|
| Attribute | ALS Metrics | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| N 2011 N 2016e | P99 + ElevIQ + % first ret. Above 2.00 | 257.09 | 36.34 | 28.81 | 272.76 | 38.55 | 26.99 | 0.72 |
| 265.62 | 38.10 | 17.99 | 295.83 | 42.43 | 20.49 | 0.64 | ||
| G 2011 G 2016e | Elev. minimum + Elev. kurtosis + % first ret. above mean | 4.43 | 22.77 | −0.10 | 4.77 | 24.51 | −0.10 | 0.77 |
| 4.18 | 19.01 | 0.20 | 5.51 | 25.05 | 0.57 | 0.71 | ||
| Dg 2011 Dg 2016e | P90 + Elev. SD + % first ret. above mean | 3.38 | 16.38 | 0.19 | 3.56 | 17.25 | 0.06 | 0.81 |
| 3.02 | 13.48 | 0.19 | 3.43 | 15.35 | 0.06 | 0.85 | ||
| Do 2011 Do 2016e | P90 + (All ret. Above 2)/(total first ret) × 100 | 4.11 | 15.53 | 0.19 | 4.07 | 15.36 | 0.11 | 0.79 |
| 3.43 | 11.99 | 0.41 | 3.53 | 12.33 | 0.31 | 0.86 | ||
| Ho 2011 Ho 2016e | P90 + Elev. variance + % all ret. above mean | 1.32 | 12.11 | 0.11 | 1.34 | 12.30 | 0.09 | 0.87 |
| 0.86 | 7.47 | 0.10 | 0.98 | 8.54 | 0.10 | 0.93 | ||
| V 2011 V 2016e | Elev. L2 + Elev. cubic mean + % first ret. above mean | 28.87 | 27.42 | 2.59 | 29.71 | 28.22 | 1.79 | 0.84 |
| 25.03 | 20.15 | 3.14 | 26.00 | 20.92 | 2.64 | 0.90 | ||
| W 2011 | P10 + Elev. Quadratic mean + (All ret. Above mean)/(total first ret) × 100 | 23.00 | 25.29 | 0.75 | 24.29 | 26.71 | −0.03 | 0.82 |
| W 2016e | 19.63 | 18.41 | 1.80 | 21.39 | 20.06 | 1.08 | 0.89 | |
| Fitting Phase | Validation | |||||||
|---|---|---|---|---|---|---|---|---|
| Attribute | ALS Metrics | RMSE | % RMSE | Bias | RMSE | % RMSE | Bias | R2 |
| N 2011e | Elev. maximum + Elev. L kurtosis + % first ret. Above 2.00 | 256.69 | 36.28 | 33.73 | 340.20 | 48.09 | 49.31 | 0.55 |
| N 2016 | 250.87 | 35.98 | 13.95 | 278.58 | 39.96 | 11.83 | 0.67 | |
| G 2011e | P75 + Elev. CUR mean CUBE + (All ret. Above 2)/(total first ret) × 100 | 4.97 | 25.54 | 0.26 | 5.04 | 25.88 | 0.13 | 0.74 |
| G 2016 | 3.88 | 17.61 | 0.41 | 4.14 | 18.80 | 0.30 | 0.84 | |
| Dg 2011e | Elev. maximum + Elev. IQ + (All ret. Above 2)/(total first ret) × 100 | 3.54 | 17.14 | 0.14 | 3.77 | 18.25 | 0.00 | 0.79 |
| Dg 2016 | 3.03 | 13.53 | 0.21 | 3.42 | 15.28 | 0.11 | 0.85 | |
| Do 2011e | P99 + Elev. CV | 4.20 | 15.85 | 0.25 | 4.18 | 15.79 | 0.16 | 0.78 |
| Do 2016 | 3.25 | 11.35 | 0.40 | 3.40 | 11.89 | 0.33 | 0.87 | |
| Ho 2011e | P95 + Elev. SD | 1.32 | 12.12 | 0.03 | 1.38 | 12.64 | 0.03 | 0.86 |
| Ho 2016 | 0.86 | 7.48 | 0.03 | 1.02 | 8.83 | 0.03 | 0.92 | |
| V 2011e | P75 + Elev. CUR mean CUBE + (All ret. Above 2)/(total first ret) × 100 | 29.97 | 28.46 | 1.51 | 30.96 | 29.40 | 0.84 | 0.83 |
| V 2016 | 24.69 | 19.87 | 2.65 | 26.35 | 21.20 | 1.92 | 0.90 | |
| W 2011e | Elev. L2 + Elev. CUR mean CUBE + % first ret. Above 2.00 | 23.11 | 25.42 | 0.96 | 23.36 | 25.69 | 0.27 | 0.83 |
| W 2016 | 18.82 | 17.65 | 1.23 | 20.06 | 18.81 | 0.56 | 0.90 | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Domingo, D.; Alonso, R.; Lamelas, M.T.; Montealegre, A.L.; Rodríguez, F.; de la Riva, J. Temporal Transferability of Pine Forest Attributes Modeling Using Low-Density Airborne Laser Scanning Data. Remote Sens. 2019, 11, 261. https://doi.org/10.3390/rs11030261
Domingo D, Alonso R, Lamelas MT, Montealegre AL, Rodríguez F, de la Riva J. Temporal Transferability of Pine Forest Attributes Modeling Using Low-Density Airborne Laser Scanning Data. Remote Sensing. 2019; 11(3):261. https://doi.org/10.3390/rs11030261
Chicago/Turabian StyleDomingo, Darío, Rafael Alonso, María Teresa Lamelas, Antonio Luis Montealegre, Francisco Rodríguez, and Juan de la Riva. 2019. "Temporal Transferability of Pine Forest Attributes Modeling Using Low-Density Airborne Laser Scanning Data" Remote Sensing 11, no. 3: 261. https://doi.org/10.3390/rs11030261
APA StyleDomingo, D., Alonso, R., Lamelas, M. T., Montealegre, A. L., Rodríguez, F., & de la Riva, J. (2019). Temporal Transferability of Pine Forest Attributes Modeling Using Low-Density Airborne Laser Scanning Data. Remote Sensing, 11(3), 261. https://doi.org/10.3390/rs11030261










