1. Introduction
Phytoplankton plays a critical role in ocean ecosystems and the global carbon cycle via carbon fixation during photosynthesis, and they account for up to 50% of the total primary production on Earth [
1]. More importantly, phytoplankton can serve as a “biological pump” by moving fixed carbon into deep ocean [
2]. One of the factors affecting carbon fixation and sinking rate is the size of the phytoplankton cell [
3]. Phytoplankton size structures, expressed as phytoplankton size classes (PSCs), are divided into three size classes: microplankton (>20 μm), nanoplankton (2–20 μm), and picoplankton (<2 μm) [
4]. PSCs are found to closely relate to phytoplankton functional types (PFTs) [
5]. Therefore, synoptic mapping of PSCs has been recently targeted by ocean color remote sensing community [
6].
The mechanism of PSCs retrieval from ocean color data lies in the fact that PSCs are closely associated with the phytoplankton abundance as well as inherent and apparent optical properties in waters [
6,
7,
8]. Hitherto, several PSCs retrieval approaches have been developed [
8], including abundance-based, spectral-based, and statistical-based ones [
6]. Abundance-based methods employ chlorophyll
a concentrations to infer phytoplankton size structure, because large phytoplankton cells are generally associated with high biomass and small cells with low biomass [
7]. Spectral-based methods depend on the spectral shape of either phytoplankton absorption or particulate backscattering [
9,
10,
11,
12].
Machine learning techniques have also successfully applied to estimate PSCs and PFTs. For examples, Raitsos et al. [
13] and Brewin et al. [
8] applied artificial neural networks to retrieve PFTs and PSCs from bio-optical, spatial, temporal, and physical features; Organelli et al. [
14] calibrated a partial least squares (PLS) models with in situ particulate absorption coefficients for PSCs retrieval in the Mediterranean Sea. In addition to the original spectral features of waters, their spectral derivatives and indices were also applied in model development, and feature selection techniques were applied to identify sensitive features for PSCs and PFTs estimations. For examples, Torrecilla et al. [
15] found, through the sensitivity test in cluster analysis, that the second derivatives of original spectral features of waters worked better than band ratios and original features in discriminating phytoplankton pigment assemblages; and Li et al. [
16] used support vector machine recursive feature elimination (SVM-RFE) to select sensitive spectral features and then applied them to develop PSCs estimation models with SVM regression.
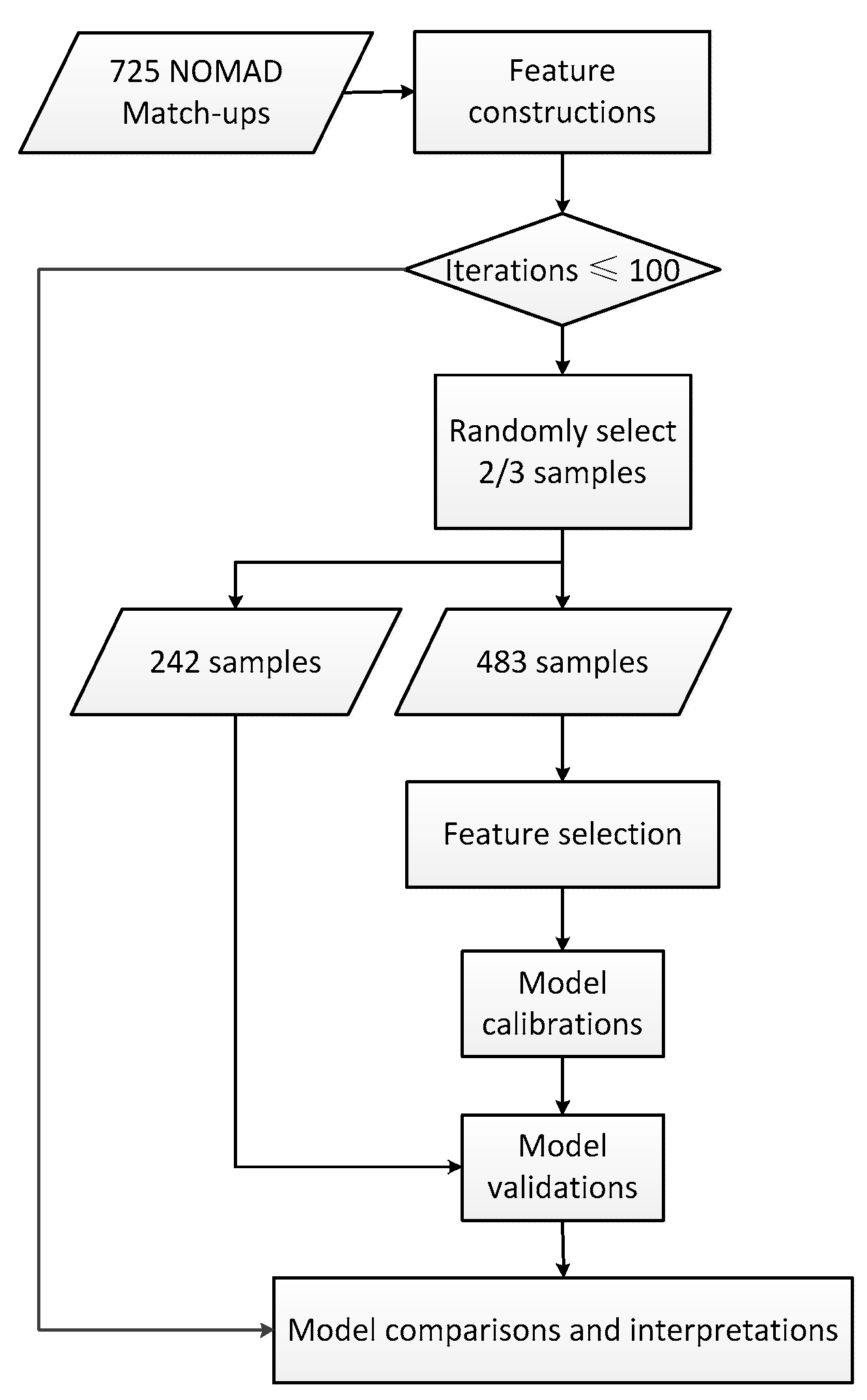
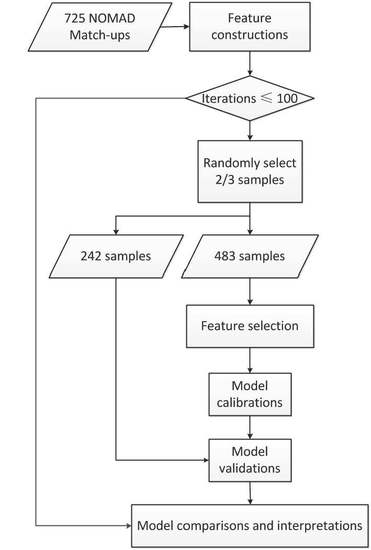
Considering the successful applications of abovementioned methods, machine learning techniques may be promising for PSCs mapping over global oceans, while statistical PSCs modeling mainly involves sensitive feature selection and model calibration. However, few comprehensive comparisons of machine learning techniques for inferring PSCs with remote sensing data were carried out in the literature. Therefore, this study, using the NOMAD HPLC database and satellite-derived products, aimed to compare the effectiveness of machine learning techniques for selecting useful spectral features and developing PSCs retrieval models. Three feature selection algorithms, including genetic algorithm, successive projection algorithm and SVM-RFE, and four modeling techniques, including PLS, ANN, SVM and random forests, were tested. The results from this study would be a good reference for statistical estimations of PSCs with remote sensing techniques.
4. Discussion
This study demonstrated the effectiveness of machine learning techniques in inferring phytoplankton size classes from satellite-sensed data. A stable rank order was found among four modeling techniques (RF > SVM > ANN > PLS) in terms of their prediction performance for inferring PSCs. RF performed the best, and was slightly better than SVM, followed by ANN. The better performance achieved by RF could be attributed to the combination of multiple diverse individual decision trees. Additionally, the out-of-bag error estimation used in RF modeling guarantees its generalization and resistance to overfitting [
37,
41]. The good results obtained by SVM should lie in its complex fitting properties, even for non-linear data, through RBF kernel mapping [
42]. Some studies indicated that RF performed better than SVM, while contrary results could also be found in the literature [
41,
43,
44]. This might be explained by the fact that SVM worked better for small sample size, while RF was thought to be more stable and reliable for large and high dimensional datasets. Considering the growing numbers of PSCs databases [
8], RF may be a better choice for PSCs estimations globally. However, PLS was not recommended due to its obvious lower accuracy.
This study emphasized the usefulness of feature selection algorithms in selecting sensitive features for PSC inferring. Although the models calibrated with all variables produced equivalent prediction performance, feature selection techniques could dramatically reduce model complexity by selecting a few sensitive features. The importance of ecological variables for PSCs estimations were highlighted, and SST, wind stress and temporal variables were among the features frequently selected, because these factors could directly affect phytoplankton growth and reproduction [
13]. Additionally, the less accurate predictions obtained by the models calibrated with only ocean color data also justified their importance. The geographic information was not incorporated in modeling following the suggestion by Raitsos and Lavender [
13], since a global PSCs database covering oceans worldwide is still not available.
The SVM-RFE used in this study showed its advantage over GA and SPA in selecting sensitive PSCs features. This might be explained by: (1) as an embedded method, the feature ranking in SVM-RFE was more effective, because it directly evaluates feature importance according to its contribution to the model [
30]; and (2) SVM was more effective than PLS regression [
42,
45], which was used as the learning machine in GA and SPA. Theoretically, SVM can also be used as a learning machine in GA; however, its efficiency would be reduced further, since it demands constant iterations to optimize initial random chromosomes to identify sensitive features [
26,
27]. Comparatively, SPA produced the worst performance, which might be explained by the fewer features it selected for modeling. Moreover SPA was designed to eliminate collinearities among features, which may exclude some useful and include some non-correlated but uninformative variables [
27]. For example, CV(443, 490, 510), selected by SPA both for microplankton and picoplankton, produced very low correlation to other features as well as PSCs.
This study indicated that the three-component method based on Chl
a alone could not achieve accurate estimations of PSCs. Similar result was also found by Alvain et al. [
46] for PFTs. However, Chl-a still plays important roles in PSCs retrieval [
13]. Even though Chl
a was not included in modeling, the features closely related to it might be selected instead. For examples, spectral curvatures, like CV(443, 490, 555) and CV(490, 510, 555), could be used to estimate Chl
a [
47]; and
aph_443, instead of Chl
a, was also used in some abundance-based models [
8]. Moreover, the results shown in
Table 7 also supported this statement, since many prominent features were closely related to Chl
a. The mean parameters for the three-component model in our study was different from those of Brewin et al. [
7], which could be explained by the different dataset used to fit models [
48]. Considering the parameters obtained from different studies (refers to Table 4.2 in [
6]), the parameters obtained in this study (
Table 2) should be valid.
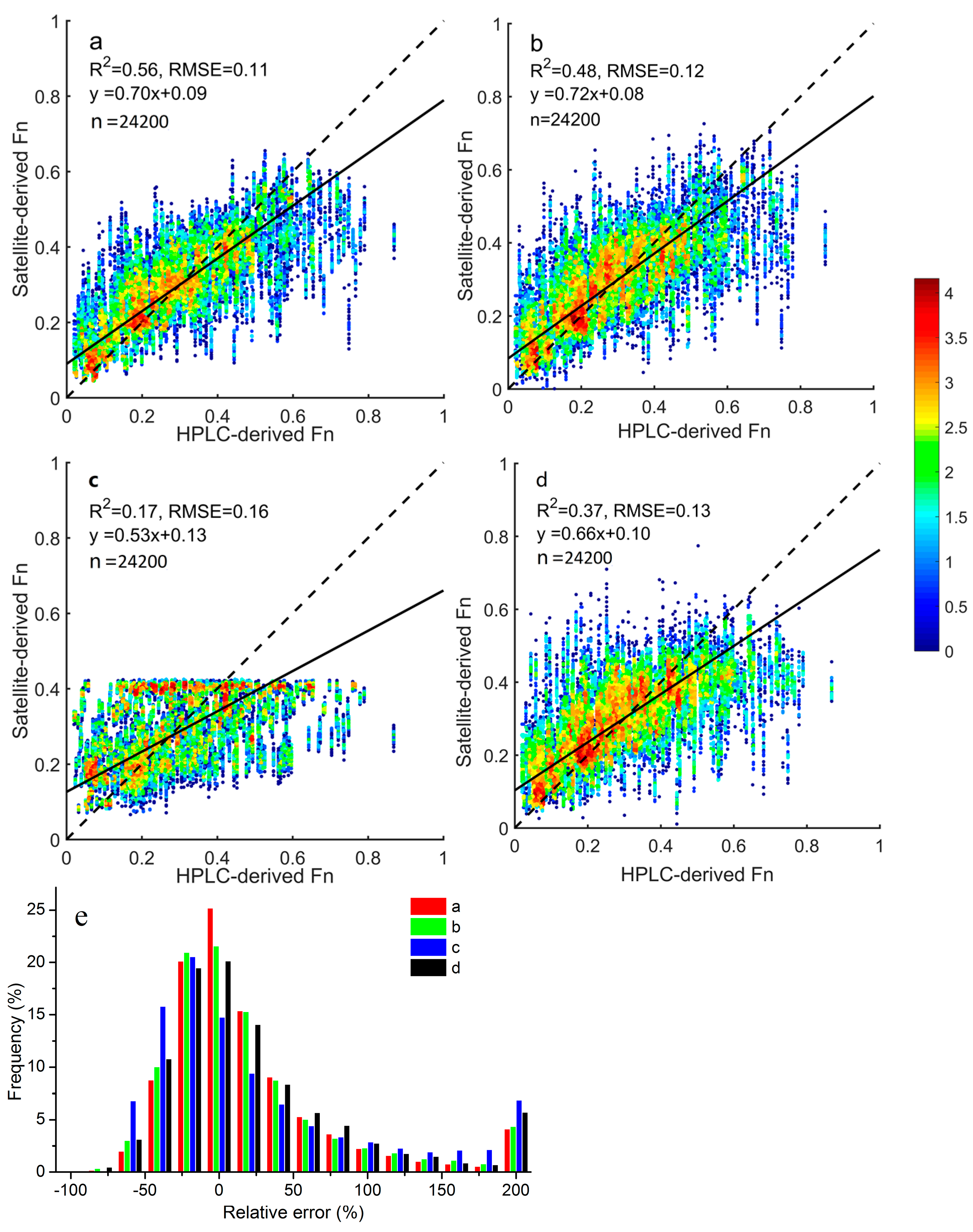
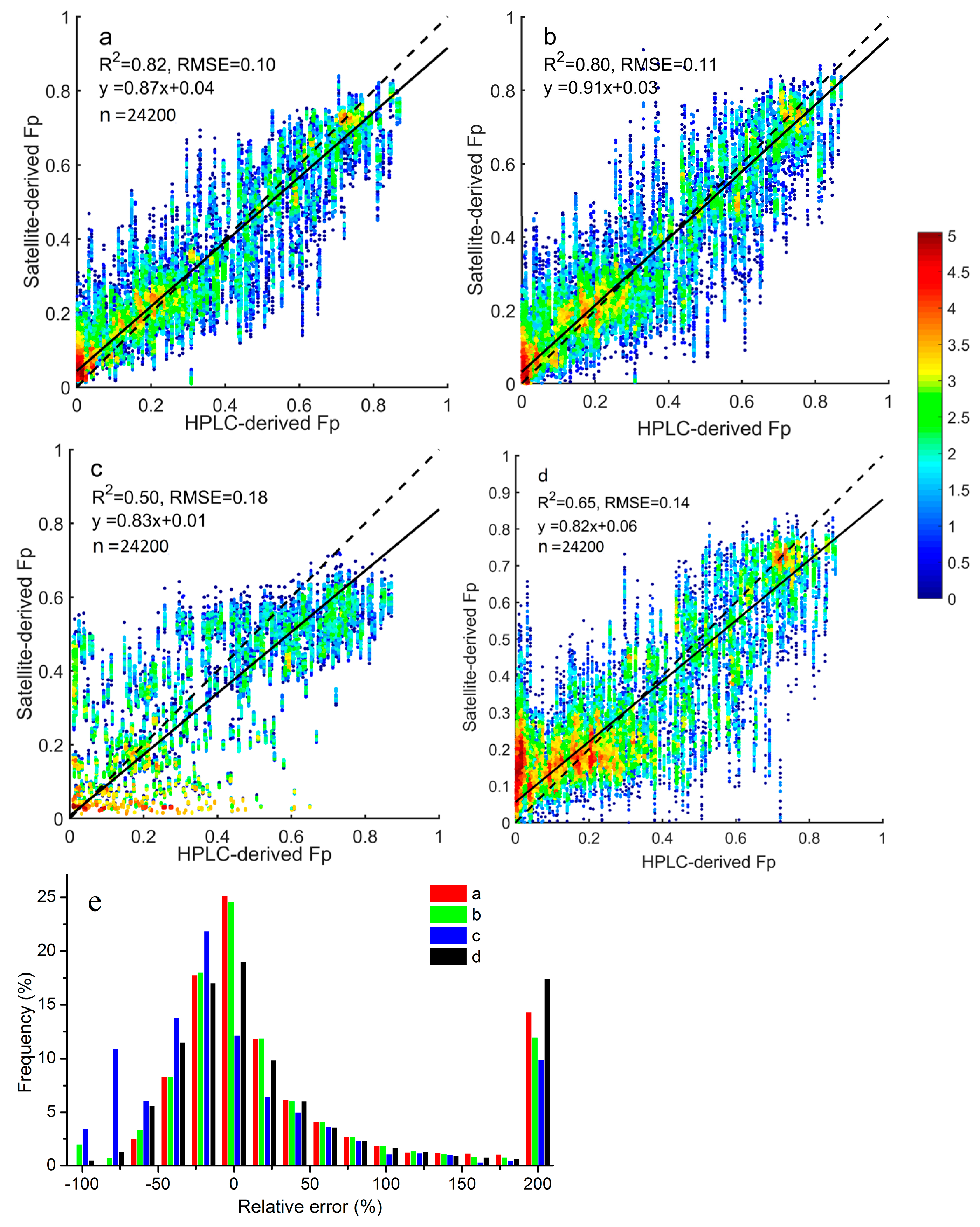
We found that microplankton and picoplankton fractions could be estimated accurately from space, and inferring nanoplankton fractions still appeared to be challenging. This was consistent with the findings by Li et al. [
16]. Such results could be explained by several reasons: (1) the microplankton and picoplankton abundances tend to increase and decrease monotonically, respectively, as a function of total Chl
a, while the Fn appears to first increase and then decrease with Chl
a [
6,
49,
50]; (2) several features have been found moderately correlated to Fm and Fp, while only one with Fn; (3) Fn shows narrow range of variations than Fm and Fp, which might partially account for its low determination of coefficients; and (4) the determination deviations introduced by HPLC should also be considered, since some types phytoplankton could be found both in picoplankton and nanoplankton [
6,
51]. The samples with relative errors >200% for Fp were largely accounted for by those with low picoplankton fractions (<2%), since a tiny absolute deviation would also tend to result in a large relative error.
5. Conclusions
Three feature selection algorithms (GA, SPA, and SVM-RFE) were applied to select the features sensitive to phytoplankton size classes from a total of 39 ocean color, biological, physical, and temporal variables, and four modeling techniques (PLS, ANN, SVM, and RF) were used to calibrate PSCs retrieval models. The embedded feature selection method, SVM-RFE, worked better than the other two wrapper methods (i.e., GA and SPA), and random forests produced the highest prediction performance. Therefore, the combination of SVM-RFE and RF in further applications was recommended. Moreover, besides popular ocean color data, the satellite-sensed ecological factors were found useful for PSCs inferring.