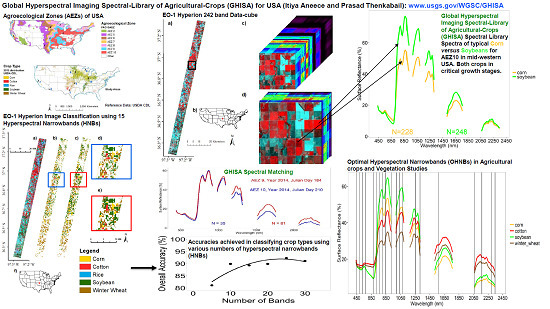
Accuracies Achieved in Classifying Five Leading World Crop Types and their Growth Stages Using Optimal Earth Observing-1 Hyperion Hyperspectral Narrowbands on Google Earth Engine
Abstract
1. Introduction
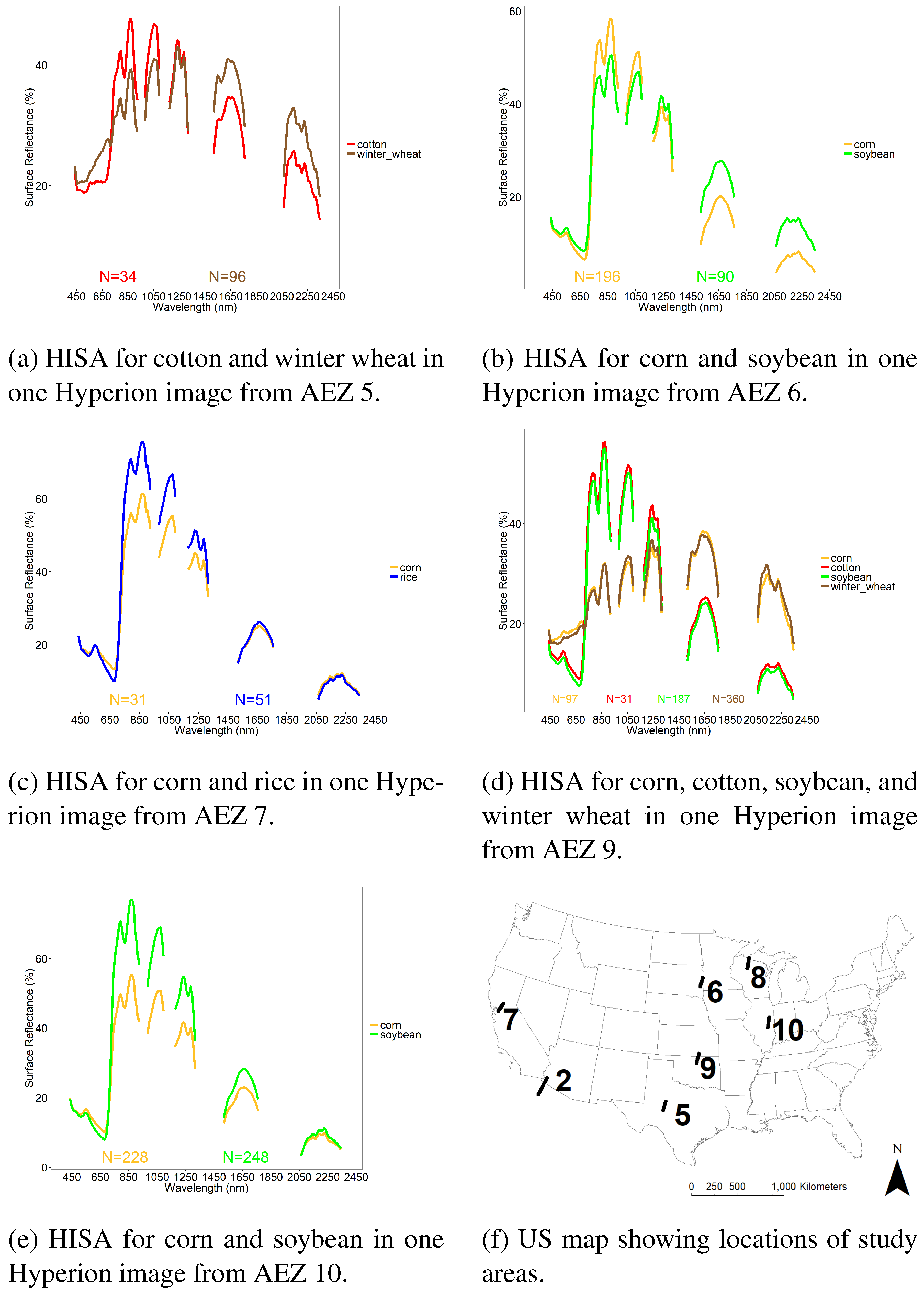
- Develop a Hyperspectral Imaging Spectral library of Agricultural crops (HISA), of five principal crops of the US, using Hyperion satellite data.
- Establish optimal hyperspectral narrowbands (OHNBs) of Hyperion to study agricultural crops in the US and to overcome data redundancy.
- Classify the five leading principal crops in the US using multiple Hyperion images from distinct AEZs to determine the strengths and limitations of OHNBs in classifying crop types and crop growth stages.
- Demonstrate the power of computing large volumes of Hyperion data on the GEE cloud computing platform. Although much research has been done on determining best bands for studying crops, this study contributes unique knowledge due to its use of 99 Hyperion images throughout different AEZs in the US to study five globally dominant crops. The results from this research will help us prepare for processing large datasets that will be generated by upcoming hyperspectral satellites like EnMAP and the Surface Biology and Geology mission (formerly HyspIRI mission) [57], as well as the DLR Earth Sensing Imaging Spectrometer (DESIS) which is already on the ISS-MUSES platform [58], and automating that processing in a cloud-computing platform. This will enable us to study and characterize agricultural crops and advance their modeling and mapping, which in turn will help in advancing food security analysis.
2. Materials and Methodology
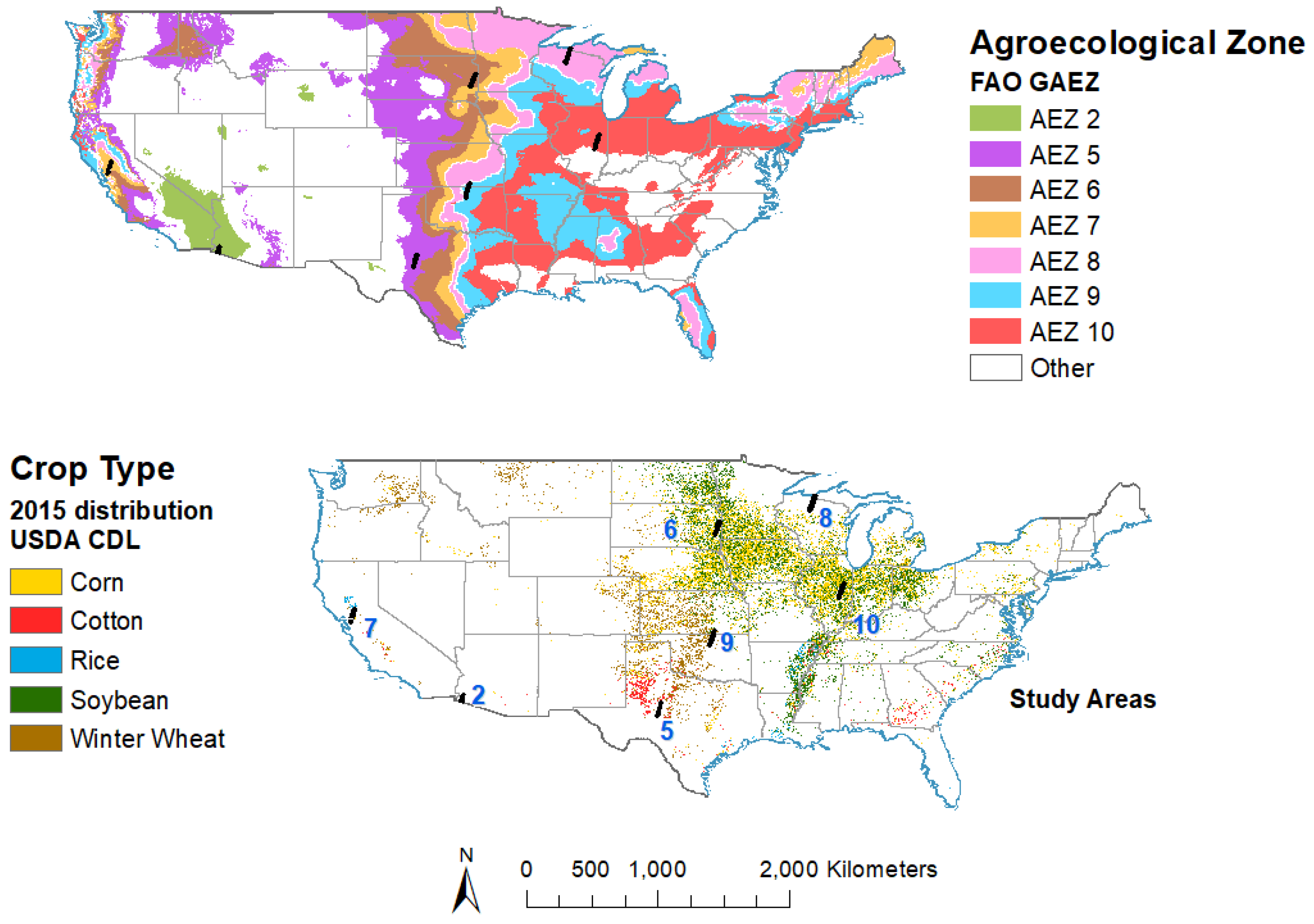
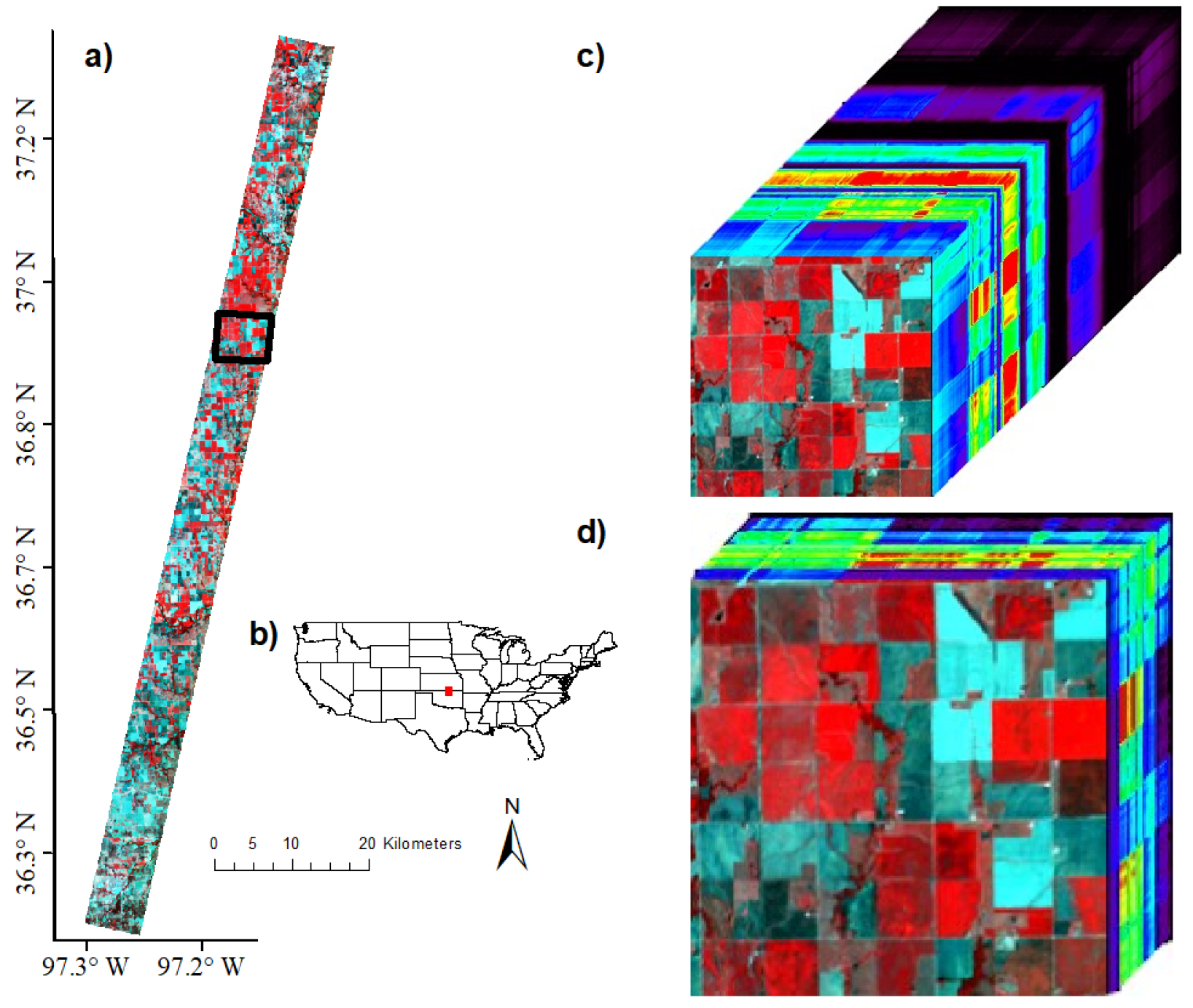
2.1. Study Area
2.2. Datasets
2.2.1. Reference Data of Agricultural Crops in the US
2.2.2. Preprocessing Hyperion Images: Steps Used in This Study
2.2.3. Atmospheric Correction of Hyperion Images
2.2.4. Distribution of Data into Training and Validation Datasets
Reference Training and Validation Datasets from Hyperion Images for Crop Type Linear Discriminant Analysis (LDA)
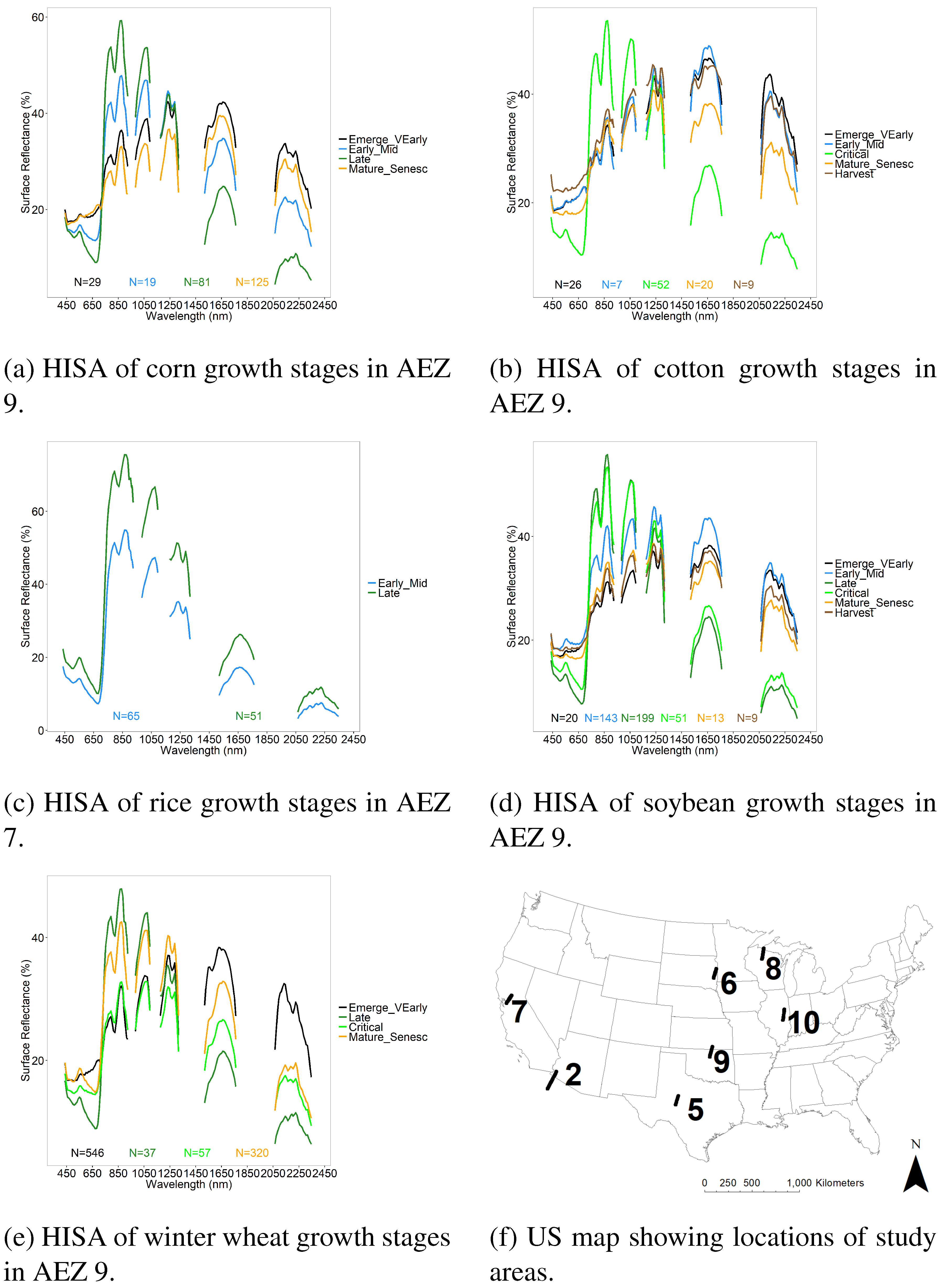
Reference Training and Validation Datasets from Hyperion Images for Crop Growth Stage Differentiation Using LDA
Reference Training and Validation Datasets from Hyperion Images for Crop Type Image Classification and Mapping Using Support Vector Machines (SVM) in GEE
2.3. Selection of OHNBs by Data Mining and Overcoming Data Redundancy
2.4. LDA for Classifying Crop Types
2.5. LDA for Classifying Crop Growth Stages
2.6. Hyperion Image Classification for Establishing Crop Types
3. Results
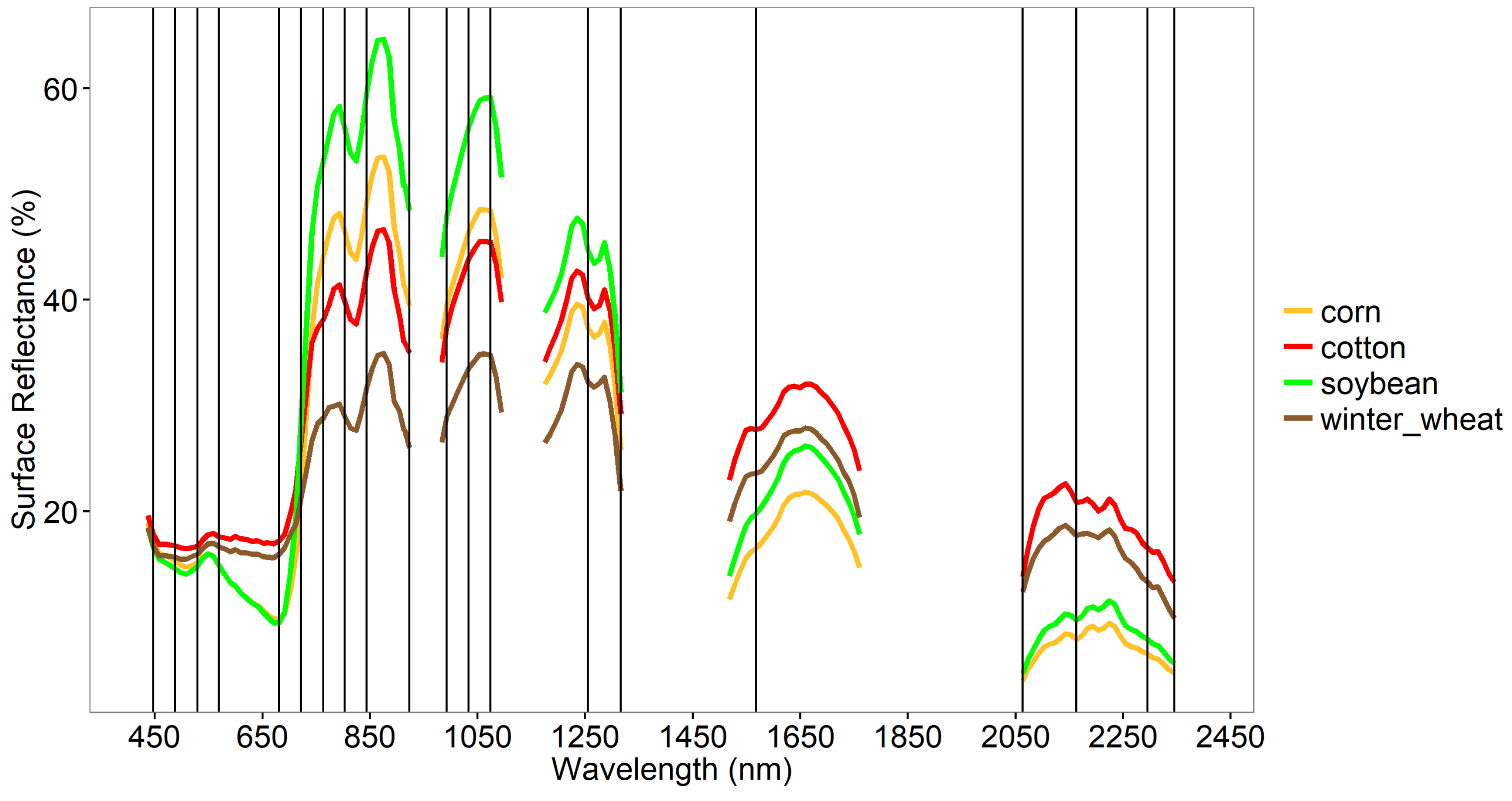
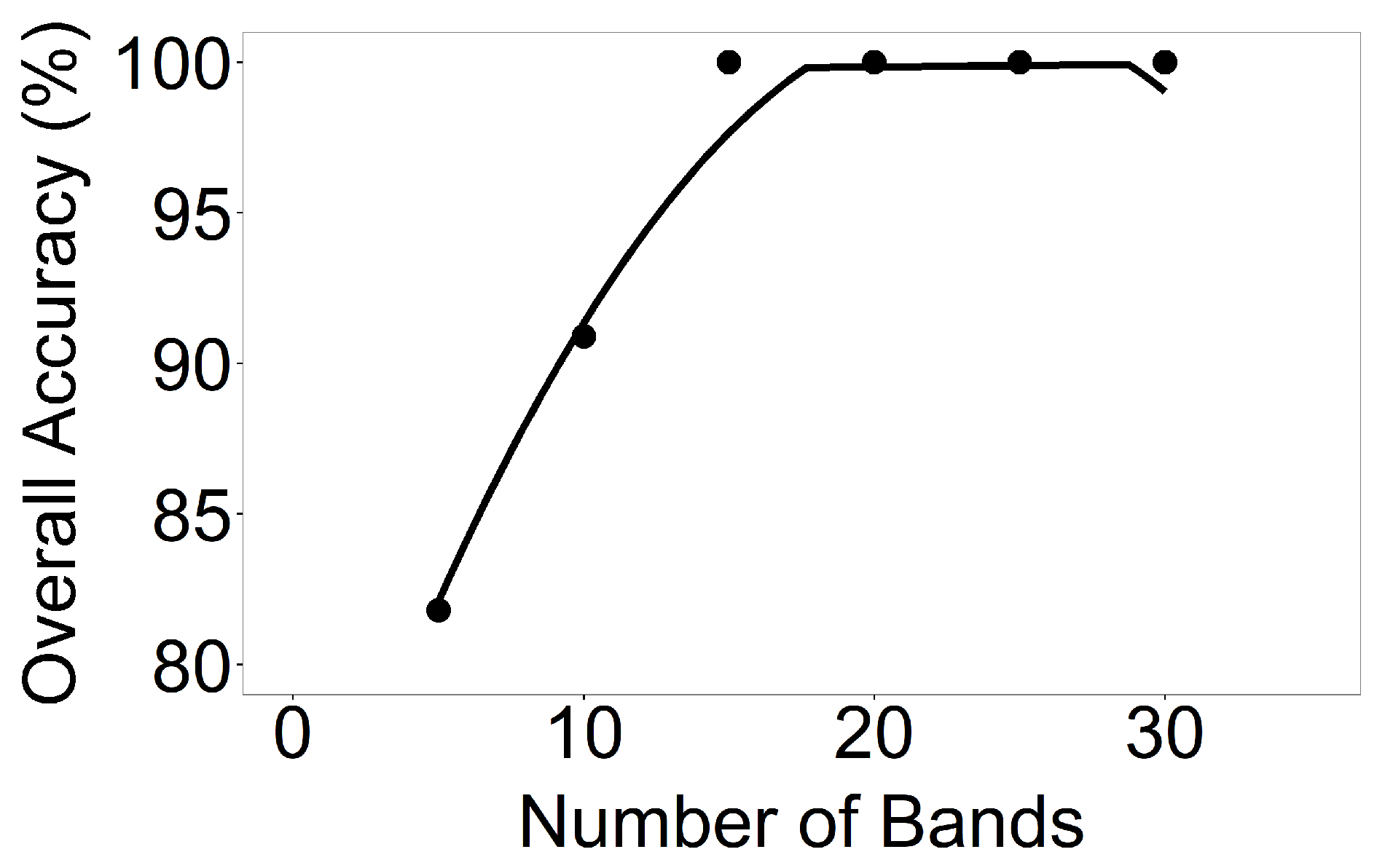
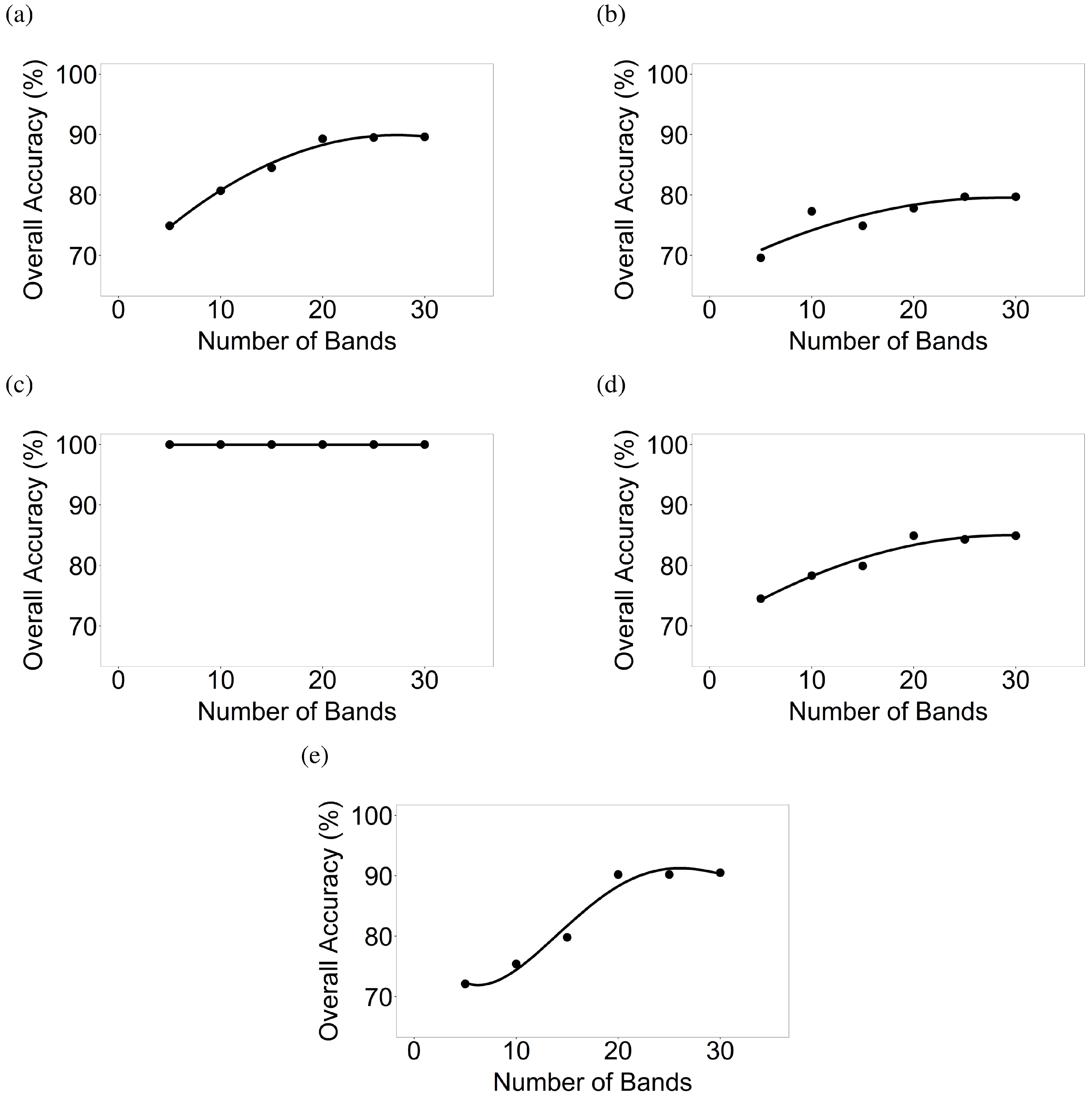
3.1. Establishing OHNBs
3.2. LDA Across Crop Types
3.3. LDA Across Growth Stages
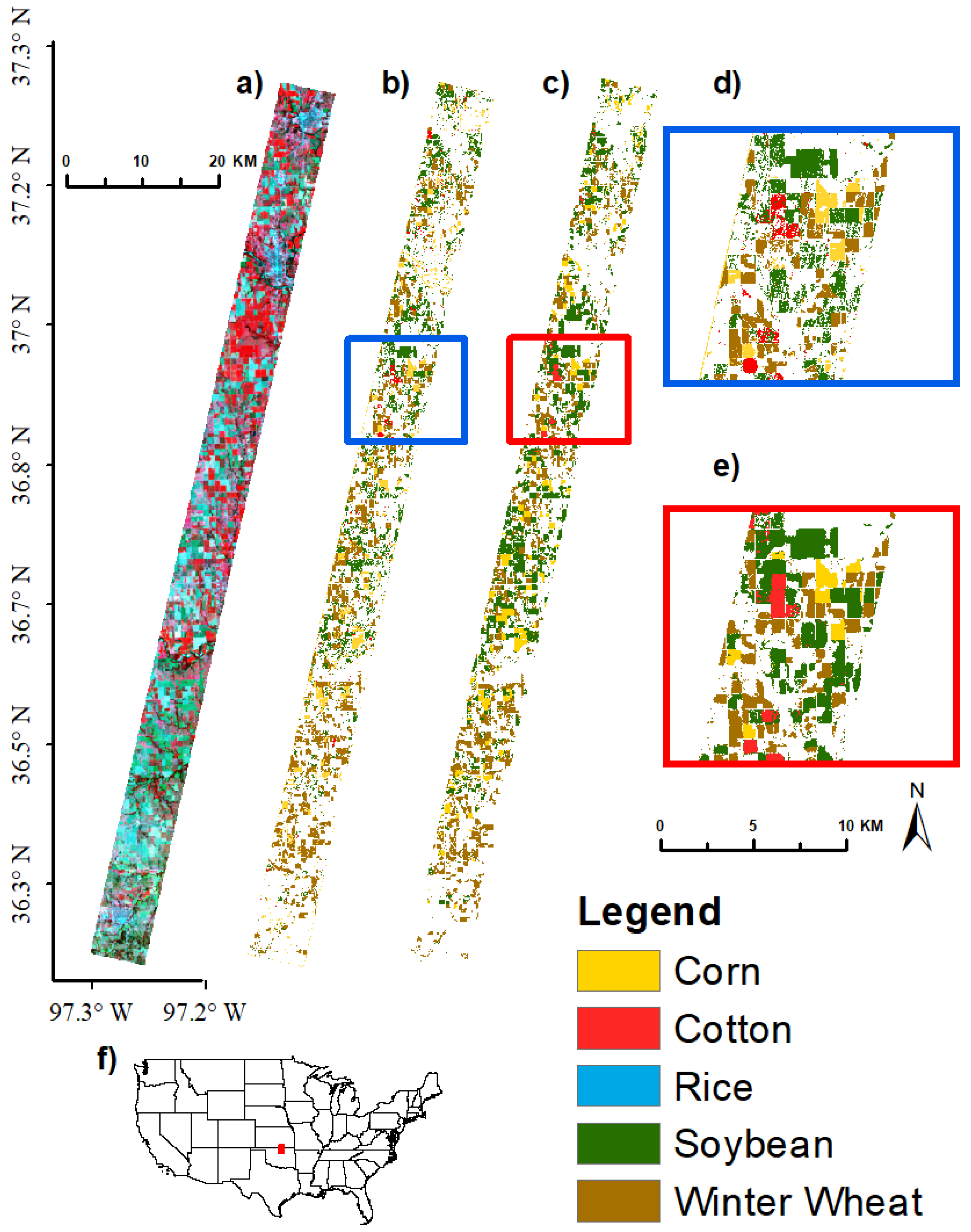
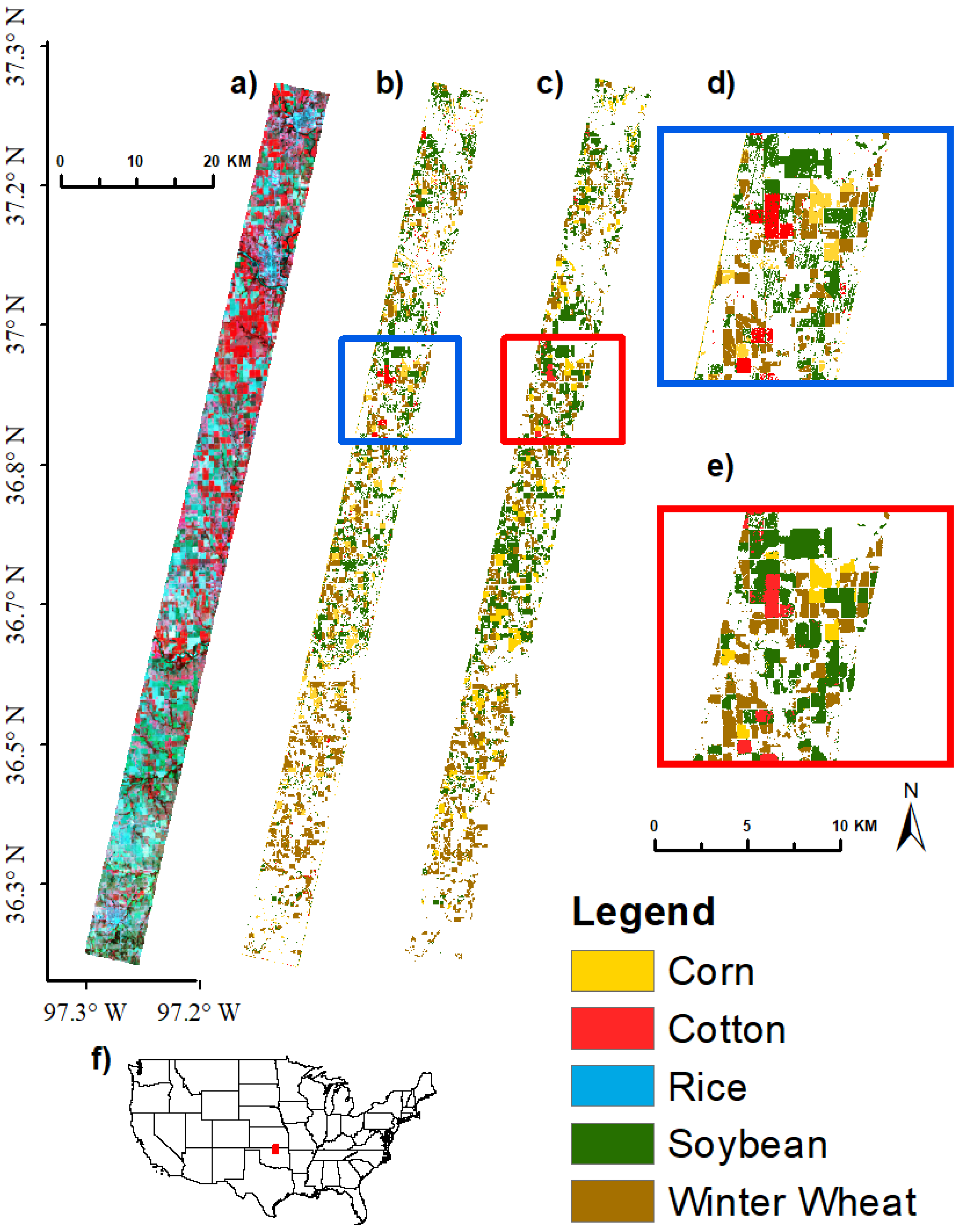
3.4. Image Classification Using Support Vector Machine
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Odegard, I.; van der Voet, E. The future of food—Scenarios and the effect on natureal resource use in agriculture in 2050. Ecol. Econ. 2014, 97, 51–59. [Google Scholar] [CrossRef]
- Kruse, F.; Baugh, W.; Perry, S. Validation of DigitalGlobe WorldView-3 Earth imaging satellite shortwave infrared bands for mineral mapping. J. Appl. Remote Sens. 2015, 9, 096044-1–096044-17. [Google Scholar] [CrossRef]
- Lambert, M.; Traore, P.; Blaes, X.; Baret, P.; Defourny, P. Estimating smallholder crops production at village level from Sentinel-2 time series in Mali’s cotton belt. Remote Sens. Environ. 2018, 216, 647–657. [Google Scholar] [CrossRef]
- Jarolmasjed, S.; Sankaran, S.; Kalcsits, L.; Khot, L. Proximal hyperspectral sensing of stomatal conductance to monitor the efficacy of exogenous abscisic acid applications in apple trees. Crop Protect. 2018, 109, 42–50. [Google Scholar] [CrossRef]
- McDowell, M.; Kruse, F. Enhanced compositional mapping through integrated full-range spectral analysis. Remote Sens. 2016, 8, 757. [Google Scholar] [CrossRef]
- Sun, J.; Shi, S.; Gong, W.; Yang, J.; Du, L.; Song, S.; Chen, B.; Zhang, Z. Evaluation of hyperspectral LiDAR for monitoring rice leaf nitrogen by comparison with multispectral LiDAR and passive spectrometer. Nat. Sci. Rep. 2017, 7, 9. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Huang, W.; Niu, Z.; Wang, Y.; Wang, C.; Li, W.; Hao, P.; Yu, B. Fine crop mapping by combining high spectral and high spatial resolution remote sensing data in complex heterogeneous areas. Comput. Electron. Agric. 2017, 139, 1–9. [Google Scholar] [CrossRef]
- Marshall, M.; Thenkabail, P. Advantage of hyperspectral EO-1 Hyperion over multispectral IKONOS, GeoEye-1, WorldView-2, Landsat ETM+, and MODIS vegetation indices in crop biomass estimation. ISPRS J. Photogramm. Remote Sens. 2015, 108, 205–218. [Google Scholar] [CrossRef]
- He, K.; Rocchini, D.; Neteler, M.; Nagendra, H. Benefits of hyperspectral remote sensing for tracking plant invasions. Divers. Distrib. 2011, 17, 381–392. [Google Scholar] [CrossRef]
- Underwood, E.; Ustin, S.; Ramirez, C. A comparison of spatial and spectral image resolution for mapping invasive plants in coastal California. J. Environ. Manag. 2007, 39, 63–83. [Google Scholar] [CrossRef]
- Underwood, E.; Ustin, S.; DiPietro, D. Mapping nonnative plants using hyperspectral imagery. Remote Sens. Environ. 2003, 86, 150–161. [Google Scholar] [CrossRef]
- Carlson, K.; Asner, G.; Hughes, R.; Ostertag, R.; Martin, R. Hyperspectral remote sensing of canopy biodiversity in Hawaiian lowland rainforests. Ecosystems 2007, 10, 536–549. [Google Scholar] [CrossRef]
- Banskota, A.; Wynne, R.; Kayastha, N. Improving within-genus tree species discrimination using the discrete wavelet transform applied to airborne hyperspectral data. Int. J. Remote Sens. 2011, 32, 3551–3563. [Google Scholar] [CrossRef]
- Cho, M.; Mathieu, R.; Asner, G.; Naidoo, L.; van Aardt, J.; Ramoelo, A.; Debba, P.; Wessels, K.; Main, R.; Smit, I.; et al. Mapping tree species composition in South African savannas using an integrated airborne spectral and LiDAR system. Remote Sens. Environ. 2012, 125, 214–226. [Google Scholar] [CrossRef]
- Naidoo, L.; Cho, M.; Mathieu, R.; Asner, G. Classification of savanna tree species, in the Greater Kruger National Park region, by integrating hyperspectral and LiDAR data in a Random Forest data mining environment. ISPRS J. Photogramm. Remote Sens. 2012, 69, 167–179. [Google Scholar] [CrossRef]
- Zhang, J.; Rivard, B.; Sanchez-Azofeifa, A.; Castro-Esau, K. Intra- and inter-class spectral variability of tropical tree species at La Selva, Costa Rica: Implications for species identification using HYDICE imagery. Remote Sens. Environ. 2006, 105, 129–141. [Google Scholar] [CrossRef]
- Shafri, H.; Suhaili, A.; Mansor, S. The performance of maximum likelihood, spectral angle mapper, neural network and decision tree classifiers in hyperspectral image analysis. J. Comput. Sci. 2007, 3, 419–423. [Google Scholar] [CrossRef]
- Andrew, M.; Ustin, S. Spectral and physiological uniqueness of perennial pepperweed (Lepidium latifolium). Weed Sci. 2006, 54, 1051–1062. [Google Scholar] [CrossRef]
- Zomer, R.; Trabucco, A.; Ustin, S. Building spectral libraries for wetlands land cover classification and hyperspectral remote sensing. J. Environ. Manag. 2009, 90, 2170–2177. [Google Scholar] [CrossRef]
- Daughtry, C.; Hunt, E., Jr.; Doraiswamy, P.; McMurtrey, J., III. Remote sensing the spatial distribution of crop residues. Agron. J. 2005, 97, 864–871. [Google Scholar] [CrossRef]
- He, Y.; Mui, A. Scaling up semi-arid grassland biochemical content from the leaf to the canopy level: Challenges and opportunities. Sensors 2010, 10, 11072–11087. [Google Scholar] [CrossRef] [PubMed]
- Beland, M.; Roberts, D.; Peterson, S.; Biggs, T.; Kokaly, R.; Piazza, S.; Roth, K.; Khanna, S.; Ustin, S. Mapping changing distributions of dominant species in oil-contaminated salt marshes of Louisiana using imaging spectroscopy. Remote Sens. Environ. 2016, 182, 192–207. [Google Scholar] [CrossRef]
- Peterson, S.; Roberts, D.; Beland, M.; Kokaly, R.; Ustin, S. Oil detection in the coastal marshes of Louisiana using MESMA applied to band subsets of AVIRIS data. Remote Sens. Environ. 2015, 159, 222–231. [Google Scholar] [CrossRef]
- Kefauver, S.; Penuelas, J.; Ustin, S. Applications of hyeprspectral remote sensing and GIS for assessing forest health and air pollution. In Proceedings of the 2012 IEEE International Geoscience and Remote Sensing Symposium (IGARSS), Munich, Germany, 22–27 July 2012. [Google Scholar]
- Smith, A.; Blackshaw, R. Weed-crop discrimination using remote sensing: A detached leaf experiment. Weed Technol. 2003, 17, 811–820. [Google Scholar] [CrossRef]
- Sahoo, R.; Ray, S.; Manjunath, K. Hyperspectral remote sensing of agriculture. Curr. Sci. 2015, 108, 848–859. [Google Scholar]
- Yang, C.; Everitt, J.; Bradford, J. Yield estimation from hyperspectral imagery using spectral angle mapper (SAM). Trans. ASABE 2008, 51, 729–737. [Google Scholar] [CrossRef]
- Jafari, R.; Lewis, M. Arid land characterization with EO-1 Hyperion hyperspectral data. Int. J. Appl. Earth Obs. Geoinf. 2012, 19, 298–307. [Google Scholar] [CrossRef]
- Thenkabail, P.; Enclona, E.; Ashton, M.; Legg, C.; De Dieu, M. Hyperion, IKONOS, ALI, and ETM+ sensors in the study of African rainforests. Remote Sens. Environ. 2004, 90, 23–43. [Google Scholar] [CrossRef]
- Asner, G.; Heidebrecht, K. Imaging spectroscopy for desertification studies: Comparing AVIRIS and EO-1 Hyperion in Argentina drylands. IEEE Trans. Geosci. Remote Sens. 2003, 41, 1283–1296. [Google Scholar] [CrossRef]
- Oskouei, M.; Babakan, S. Detection of alteration minerals using Hyperion data analysis in Lahroud. J. Indian Soc. Remote Sens. 2016, 44, 713–721. [Google Scholar] [CrossRef]
- Dadon, A.; Ben-Dor, E.; Beyth, M.; Karnieli, A. Examination of spaceborne imaging spectroscopy data utility for stratigraphic and lithologic mapping. J. Appl. Remote Sens. 2011, 5, 053507-1–053507-14. [Google Scholar] [CrossRef]
- Papes, M.; Tupayachi, R.; Martinez, P.; Peterson, A.; Powell, G. Using hyperspectral satellite imagery for regional inventories: A test with tropical emergent trees in the Amazon Basin. J. Veg. Sci. 2010, 21, 342–354. [Google Scholar] [CrossRef]
- Somers, B.; Asner, G. Tree species mapping in tropical forests using multi-temporal imaging spectroscopy: Wavelength adaptive spectral mixture analysis. Int. J. Appl. Earth Obs. Geoinf. 2014, 31, 57–66. [Google Scholar] [CrossRef]
- Somers, B.; Asner, G. Mapping tropical rainforest canopies using multi-temporal spaceborne imaging spectroscopy. Proc. SPIE 2013, 8887, 888704-1–888704-8. [Google Scholar]
- Christian, B.; Krishnayya, N. Classification of tropical trees growing in a sanctuary using Hyperion (EO-1) and SAM algorithm. Curr. Sci. 2009, 96, 1601–1607. [Google Scholar]
- George, R.; Padalia, H.; Kushwaha, S. Forest tree species discrimination in western Himalaya using EO-1 Hyperion. Int. J. Appl. Earth Obs. Geoinf. 2014, 28, 140–149. [Google Scholar] [CrossRef]
- Walsh, S.; McCleary, A.; Mena, C.; Shao, Y.; Tuttle, J.; Gonzalez, A.; Atkinson, R. QuickBird and Hyperion data analysis of an invasive plant species in the Galapagos Islands of Ecuador: Implications for control and land use management. Remote Sens. Environ. 2008, 112, 1927–1941. [Google Scholar] [CrossRef]
- Somers, B.; Asner, G. Multi-temporal hyperspectral mixture analysis and feature selection for invasive species mapping in rainforests. Remote Sens. Environ. 2013, 136, 14–27. [Google Scholar] [CrossRef]
- Huemmrich, K.; Gamon, J.; Tweedie, C.; Campbell, P.; Landis, D.; Middleton, E. Arctic tundra vegetation functional types based on photosynthetic physiology and optical properties. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2013, 6, 265–275. [Google Scholar] [CrossRef]
- Mielke, C.; Boesche, N.; Rogass, C.; Kaufmann, H.; Gauert, C.; de Wit, M. Spaceborne mine waste minerology monitoring in South Africa, applications for modern push-broom missions: Hyperion/ OLI and EnMAP/ Sentinel-2. Remote Sens. 2014, 6, 6790–6816. [Google Scholar] [CrossRef]
- Sims, N.; Culvenor, D.; Newnham, G.; Coops, N.; Hopmans, P. Towards the operational use of satellite hyperspectral image data for mapping nutrient status and fertilizer requirements in Australian plantation forests. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2013, 6, 320–328. [Google Scholar] [CrossRef]
- Zhang, Q.; Middleton, E.; Cheng, Y.; Landis, D. Variations of foliage chlorophyll fAPAR and foliage non-chlorophyll fAPAR (fAPARchl, fAPARnon-chl) at the Harvard Forest. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2013, 6, 2254–2264. [Google Scholar] [CrossRef]
- Roberts, D.; Dennison, P.; Gardner, M.; Hetzel, Y.; Ustin, S.; Lee, C. Evaluation of the potential of Hyperion for fire danger assessment by comparison to the airborne visible/ infrared imaging spectrometer. IEEE Trans. Geosci. Remote Sens. 2003, 41, 1297–1310. [Google Scholar] [CrossRef]
- Mitri, G.; Gitas, I. Mapping postfire vegetation recovery using EO-1 Hyperion imagery. IEEE Trans. Geosci. Remote Sens. 2010, 48, 1613–1618. [Google Scholar] [CrossRef]
- Bannari, A.; Staenz, K.; Champagne, C.; Khurshid, K. Spatial variability mapping of crop residue using Hyperion (EO-1) hyperspectral data. Remote Sens. 2015, 7, 8107–8127. [Google Scholar] [CrossRef]
- Sonmez, N.; Slater, B. Measuring intensity of tillage and plant residue cover using remote sensing. Eur. J. Remote Sens. 2016, 49, 121–135. [Google Scholar] [CrossRef]
- Bhojaraja, B.E.; Shetty, A.; Nagaraj, M.K.; Manju, P. Age-based classification of arecanut crops: A case study of Channagiri, Karnataka, India. Geocarto Int. 2015, 1–11. [Google Scholar] [CrossRef]
- Breunig, F.; Galvao, L.; Formaggio, A.; Epiphanio, J. Classification of soybean varieties using different techniques: Case study with Hyperion and sensor spectral resolution simulations. J. Appl. Remote Sens. 2011, 5, 053533-1–053533-15. [Google Scholar] [CrossRef]
- Pan, Z.; Huang, J.; Wang, F. Multi range spectral feature fitting for hyperspectral imagery in extracting oilseed rape planting area. Int. J. Appl. Earth Obs. Geoinf. 2013, 25, 21–29. [Google Scholar] [CrossRef]
- Lamparelli, R.; Johann, J.; Dos Santos, E.; Esquerdo, J.; Rocha, J. Use of data mining and spectral profiles to differentiate condition after harvest of coffee plants. Eng. Agric. Jaboticabal 2012, 32, 184–196. [Google Scholar] [CrossRef]
- Moharana, S.; Dutta, S. Spatial variability of chlorophyll and nitrogen content of rice from hyperspectral imagery. ISPRS J. Photogramm. Remote Sens. 2016, 122, 17–29. [Google Scholar] [CrossRef]
- Houborg, R.; McCabe, M.; Angel, Y.; Middleton, E. Detection of chlorophyll and leaf area index dynamics from sub-weekly hyperspectral imagery. Proc. SPIE 2016, 9998. [Google Scholar] [CrossRef]
- Thenkabail, P.; Mariotto, I.; Gumma, M.; Middleton, E.; Landis, D.; Huemmrich, K. Selection of hyperspectral narrowbands (HNBs) and composition of hyperspectral two band vegetation indices (HVIs) for biophysical characterization and discrimination of crop types using field reflectance and Hyperion/ EO-1 data. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2013, 6, 427–439. [Google Scholar] [CrossRef]
- Mariotto, I.; Thenkabail, P.; Huete, A.; Slonecker, T.; Platonov, A. Hyperspectral versus multispectral crop-productivity modeling and type discrimination for the HyspIRI mission. Remote Sens. Environ. 2013, 139, 291–305. [Google Scholar] [CrossRef]
- Gorelick, N.; Hancher, M.; Dixon, M.; Ilyushchenko, S.; Thau, D.; Moore, R. Google Earth Engine: Planetary-scale geospatial analysis for everyone. Remote Sens. Environ. 2017, 202, 18–27. [Google Scholar] [CrossRef]
- National Academies of Sciences, Engineering and Medicine. Thriving on Our Changing Planet: A Decadal Strategy for Earth Observation from Space; The National Academies Press: Washington, DC, USA, 2018. [Google Scholar] [CrossRef]
- Eckardt, A.; Horack, J.; Lehmann, F.; Krutz, D.; Drescher, J.; Whorton, M.; Soutullo, M. DESIS (DLR Earth Sensing Imaging Spectrometer for the ISS-MUSES platform). In Proceedings of the 2015 IEEE International Geoscience and Remote Sensing Symposium (IGARSS), Milan, Italy, 26–31 July 2015; pp. 1457–1459. [Google Scholar] [CrossRef]
- FAO—Food and Agriculture Organization of the United Nations. GAEZ-–Global Agro-Ecological Zones; FAO: Rome, Italy, 2018. [Google Scholar]
- NASS. USDA National Agricultural Statistics Service Cropland Data Layer. 2014. Available online: https://nassgeodata.gmu.edu/CropScape/ (accessed on 28 September 2018).
- NASS. USDA CropScape and Cropland Data Layer-Metadata; Technical Report; United States Department of Agriculture, National Agricultural Statistics Service, NASS Marketing and Information Services Office: Washington, DC, USA, 2018.
- NASS. United States Department of Agriculture: National Agricultural Statistics Service. 2017. Available online: https://www.nass.usda.gov/ (accessed on 28 September 2018).
- NASS. USDA Crop Production 2017 Summary: January 2018; Technical Report; United States Department of Agriculture, National Agricultural Statistics Service, NASS Marketing and Information Services Office: Washington, DC, USA, 2018.
- Aneece, I.; Thenkabail, P. Spaceborne hyperspectral EO-1 Hyperion data pre-processing: Methods, approaches, and algorithms. In Hyperspectral Remote Sensing of Vegetation; Taylor and Francis Inc./CRC Press: Boca Raton, FL, USA, 2018. [Google Scholar]
- Pervez, W.; Uddin, V.; Khan, S.; Khan, J. Satellite-based land use mapping: Comparative analysis of Landsat-8, Advanced Land Imager, and big data Hyperion imagery. J. Appl. Remote Sens. 2016, 10, 026004-1–026004-20. [Google Scholar] [CrossRef]
- Das, P.; Seshasai, M. Multispectral sensor spectral resolution simulations for generation of hyperspectral vegetation indices for Hyperion data. Geocarto Int. 2015, 30, 686–700. [Google Scholar] [CrossRef]
- Farifteh, J.; Nieuwenhuis, W.; Garcia-Melendez, E. Mapping spatial variations of iron oxide by-product minerals from EO-1 Hyperion. Int. J. Remote Sens. 2013, 34, 682–699. [Google Scholar] [CrossRef]
- San, B.; Suzen, M. Evaluation of cross-track illumination in EO-1 Hyperion imagery for lithological mapping. Int. J. Remote Sens. 2011, 32, 7873–7889. [Google Scholar] [CrossRef]
- Datt, B.; McVicar, T.; Van Niel, T.; Jupp, D.; Pearlman, J. Preprocessing EO-1 Hyperion hyperspectral data to support the application of agricultural indexes. IEEE Trans. Geosci. Remote Sens. 2003, 41, 1246–1259. [Google Scholar] [CrossRef]
- Gueymard, C. SMARTS2, a Simple Model of the Atmospheric Radiative Transfer of Sunshine: Algorithms and Performance Assessment; Technical Report FSEC-PF-270-95; Florida Solar Energy Center, University of Central Florida: Cocoa, FL, USA, 1995. [Google Scholar]
- Gueymard, C. Parameterized transmittance model for direct beam and circumsolar spectral irradiance. Sol. Energy 2001, 71, 325–346. [Google Scholar] [CrossRef]
- Seidel, F.; Kokhanovsky, A.; Schaepman, M. Fast and simple model for atmospheric radiative transfer. Atmos. Meas. Tech. 2010, 3, 1129–1141. [Google Scholar] [CrossRef]
- Chavez, P., Jr. Image-based atmospheric corrections- revisited and improved. Photogramm. Eng. Remote Sens. 1996, 62, 1025–1036. [Google Scholar]
- Gopinathan, K.; Polokoana, P. Estimation of hourly beam and diffuse solar radiation. Sol. Wind Technol. 1986, 3, 223–229. [Google Scholar] [CrossRef]
- NOAA. NOAA Integrated Surface Database. 2017. Available online: https://www.ncdc.noaa.gov/isd (accessed on 8 December 2016).
- Abatzoglou, J. Development of gridded surface meteorological data for ecological applications and modelling. Int. J. Climatol. 2013, 33, 121–131. [Google Scholar] [CrossRef]
- NASS. United States Department of Agriculture National Agricultural Statistics Service, Research and Science: Cropscape and Cropland Data Layers- FAQs. 2018. Available online: https://www.nass.usda.gov/Research_and_Science/Cropland/sarsfaqs2.php (accessed on 28 September 2018).
- Stevens, A.; Ramirez-Lopez, L. Package ‘Prospectr’. Technical Report. 2015. Available online: https://cran.r-project.org/web/packages/prospectr/prospectr.pdf (accessed on 8 June 2018).
- Thenkabail, P.; Gumma, M.; Telaguntla, P.; Mohammed, I. Hyperspectral remote sensing of vegetation and agricultural crops. Photogramm. Eng. Remote Sens. 2014, 80, 697–709. [Google Scholar]
- Mulla, D. Twenty five years of remote sensing in precision agriculture: Key advances and remaining knowledge gaps. Biosyst. Eng. 2013, 114, 358–371. [Google Scholar] [CrossRef]
- Bajwa, S.; Kulkarni, S.; Lyon, J.; Huete, A. Hyperspectral data mining. In Hyperspectral Remote Sensing of Vegetation; Thenkabail, P., Ed.; CRC Press, Taylor and Francis Group: Boca Raton, FL, USA, 2011; pp. 93–120. [Google Scholar]
- Laparra, V.; Malo, J.; Camps-Valls, G. Dimensionality reduction via regression in hyperspectral imagery. IEEE J. Sel. Top. Signal Process. 2015, 9, 1026–1036. [Google Scholar] [CrossRef]
- Ding, C.; He, X.; Zha, H.; Simon, H. Adaptive dimension reduction for clustering high dimensional data. In Proceedings of the International Conference on Data Mining, Maebashi City, Japan, 9–12 December 2002; pp. 1–8. [Google Scholar]
- Binol, H.; Ochilov, S.; Alam, M.; Bai, A. Target oriented dimensionality reduction of hyperspectral data by Kernel Fukunaga-Koontz Transform. Opt. Lasers Eng. 2017, 89, 123–130. [Google Scholar] [CrossRef]
- Chang, P.; Yang, J.; Den, W.; Wu, C. Characterizing and locating air pollution sources in a complex industrial district using optical remote sensing technology and multivariate statistical modeling. Environ. Sci. Pollut. Res. 2014, 21, 10852–10866. [Google Scholar] [CrossRef]
- Shahdoosti, H.; Mirzapour, F. Spectral-spatial feature extraction using orthogonal linear discriminant analysis for classification of hyperspectral data. Eur. J. Remote Sens. 2017, 50, 111–124. [Google Scholar] [CrossRef]
- Maxwell, A.; Warner, T.; Fang, F. Implementation of machine-learning classification in remote sensing: An applied review. Int. J. Remote Sens. 2018, 39, 2784–2817. [Google Scholar] [CrossRef]
- Maxwell, A.; Warner, T.; Strager, M.; Conley, F.; Sharp, A. Assessing machine-learning algorithms and image- and LiDAR-derived variables for GEOBIA classification of mining and mine reclamation. Int. J. Remote Sens. 2015, 36, 954–978. [Google Scholar] [CrossRef]
- Xiong, J.; Thenkabail, P.; Tilton, J.; Gumma, M.; Teluguntla, P.; Oliphant, A.; Congalton, R.; Yadav, K.; Gorelick, N. Nominal 30-m cropland extent map of continental Africa by integration pixel-based and object-based algorithms using Sentinel-2 and Landsat-8 data on Google Earth Engine. Remote Sens. 2017, 9, 1065. [Google Scholar] [CrossRef]
- Hsu, C.; Chang, C.; Lin, C. A Practical Guide to Support Vector Classification. 2010, pp. 1–10. Available online: https://www.csie.ntu.edu.tw/~cjlin/papers/guide/guide.pdf (accessed on 24 January 2013).
- USGS. USGS EarthExplorer—Home. 2018. Available online: https://earthexplorer.usgs.gov/ (accessed on 28 September 2018).
- Suarez, L.; Apan, A.; Werth, J. Detection of phenoxy herbicide dosage in cotton crops through the analysis of hyperspectral data. Int. J. Remote Sens. 2017, 38, 6528–6553. [Google Scholar] [CrossRef]
- Marshall, M.; Thenkabail, P. Biomass modeling of four leading world crops using hyperspectral narrowbands in support of HyspIRI mission. Photogramm. Eng. Remote Sens. 2014, 80, 757–772. [Google Scholar] [CrossRef]
- Nugent, P.; Shaw, J.; Jha, P.; Scherrer, B.; Donelick, A.; Kumar, V. Discrimination of herbicide-resistant kochia with hyperspectral imaging. J. Appl. Remote Sens. 2018, 12, 016037-1–016037-10. [Google Scholar] [CrossRef]
- Feng, H.; Jiang, N.; Huang, C.; Fang, W.; Yang, W.; Chen, G.; Xiong, L.; Liu, Q. A hyperspectral imaging system for an accurate prediction of the above-ground biomass of individual rice plants. Rev. Sci. Instrum. 2013, 84, 095107-1–095107-10. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Miller, J.; Pattey, E.; Haboudane, D.; Strachan, I.; Hinther, M. Monitoring crop biomass accumulation using multi-temporal hyperspectral remote sensing data. In Proceedings of the IEEE International Geosicence and Remote Sensing Symposium, Anchorage, AK, USA, 20–24 September 2004; Volume 3, pp. 1637–1640. [Google Scholar]
- Mutanga, O.; Skidmore, A. Narrow band vegetation indices overcome the saturation problem in biomass estimation. Int. J. Remote Sens. 2004, 25, 3999–4014. [Google Scholar] [CrossRef]
- Cutter, M.; Kellar-Bland, H. CHRIS Data Format; Technical Report SmarTeam 0114848; Surrey Satellite Technology LTD- Chris Operations: Guildford, UK, 2008. [Google Scholar]
- Thenkabail, P.; Aneece, I.; Teluguntla, P.; Oliphant, A.; Foley, D.; Williamson, D. Global Hyperspectral Imaging Spectroscopy of Agricultural-Crops & Vegetation (GHISA). 2019. Available online: www.usgs.gov/WGSC/GHISA (accessed on 30 May 2018).
| Crop | US Area Acres (Hectares) | Portion of US Crops % | World Area Acres (Hectares) | Portion of World Crops % |
|---|---|---|---|---|
| Corn | 90,886,000 | 28.6 | 561,176,321 | 12.7 |
| (36,780,259) | (227,099,999) | |||
| Cotton | 12,055,000 | 3.8 | 131,954,274 | 3.0 |
| (4,878,485) | (53,400,000) | |||
| Rice | 2,562,000 | 0.8 | 483,338,126 | 10.9 |
| (1,036,804) | (195,599,999) | |||
| Soybean | 89,513,000 | 28.1 | 229,066,689 | 5.2 |
| (36,224,625) | (92,700,000) | |||
| Wheat * | 46,012,000 | 14.5 | 995,340,477 | 22.5 |
| (18,620,395) | (402,800,000) | |||
| Principal Crops ** | 318,184,000 | 75.8 | 3,061,635,676 | 54.3 |
| (128,764,496) | (1,238,999,999) |
| AEZ | Years | Crop | Accuracies | ||
|---|---|---|---|---|---|
| Overall (%) | Producer’s (%) | User’s (%) | |||
| 2 | 2011–2012 | All | 86.0 | ||
| Cotton | 96.1 | 94.9 | |||
| 5 | 2013–2015 | All | 80.5 | ||
| Cotton | 93.2 | 88.3 | |||
| Winter Wheat | 92.7 | 88.2 | |||
| 6 | 2011–2014 | All | 79 | ||
| Corn | 95.4 | 94.4 | |||
| Soybean | 95.2 | 95.6 | |||
| 7 | 2012 | All | 83.7 | ||
| Corn | 89.3 | 89.0 | |||
| Rice | 99.5 | 98.6 | |||
| 8 | 2008–2014 | All | 87.2 | ||
| Corn | 95.7 | 94.0 | |||
| 9 | 2008–2015 | All | 83.4 | ||
| Corn | 88.5 | 89.1 | |||
| Cotton | 85.5 | 81.1 | |||
| Soybean | 80.3 | 78.8 | |||
| Winter Wheat | 93.5 | 93.1 | |||
| 10 | 2009–2015 | All | 94.6 | ||
| Corn | 97.6 | 97.7 | |||
| Soybean | 96.3 | 96.4 | |||
| AEZ ** | Number of Hyperion Images * | Years | Crops | |
|---|---|---|---|---|
| Crop Type-Discrimination | Crop Growth Stage-Discrimination | |||
| 2 | 0 | 4 | 2011–2012 | Cotton |
| 5 | 1 | 12 | 2013–2015 | Cotton, Winter Wheat |
| 6 | 3 | 14 | 2011–2014 | Corn, Soybean |
| 7 | 2 | 2 | 2012 | Corn, Rice |
| 8 | 0 | 25 | 2008–2014 | Corn |
| 9 | 2 | 19 | 2008–2015 | Corn, Cotton, Soybean, Winter Wheat |
| 10 | 3 | 23 | 2009–2015 | Corn, Soybean |
| Total | 11 | 99 | 2008–2015 | Corn, Cotton, Rice, Soybean, Winter Wheat |
| AEZ | Hyperion Image | Crop Type | Total Sample Size | Training Sample Size | Validation Sample Size |
|---|---|---|---|---|---|
| 5 | Image 1 | Cotton | 34 | 25 | 9 |
| Winter Wheat | 96 | 72 | 24 | ||
| Other | 260 | 195 | 65 | ||
| 6 | Image 1 | Corn | 103 | 77 | 26 |
| Soybean | 183 | 137 | 46 | ||
| Other | 377 | 283 | 94 | ||
| Image 2 | Corn | 196 | 147 | 49 | |
| Soybean | 90 | 67 | 23 | ||
| Other | 310 | 232 | 78 | ||
| Image 3 | Corn | 184 | 138 | 46 | |
| Soybean | 95 | 71 | 24 | ||
| Other | 320 | 240 | 80 | ||
| 7 | Image 1 | Corn | 34 | 25 | 9 |
| Rice | 65 | 48 | 17 | ||
| Other | 200 | 150 | 50 | ||
| Image 2 | Corn | 31 | 23 | 8 | |
| Rice | 51 | 38 | 13 | ||
| Other | 200 | 150 | 50 | ||
| 9 | Image 1 | Corn | 97 | 72 | 25 |
| Cotton | 31 | 23 | 8 | ||
| Soybean | 187 | 140 | 47 | ||
| Winter Wheat | 360 | 270 | 90 | ||
| Other | 1100 | 825 | 275 | ||
| Image 2 | Corn | 81 | 60 | 21 | |
| Soybean | 90 | 67 | 23 | ||
| Winter Wheat | 279 | 209 | 70 | ||
| Other | 936 | 702 | 234 | ||
| 10 | Image 1 | Corn | 270 | 202 | 68 |
| Soybean | 282 | 211 | 71 | ||
| Other | 350 | 262 | 88 | ||
| Image 2 | Corn | 252 | 189 | 63 | |
| Soybean | 278 | 208 | 70 | ||
| Other | 350 | 262 | 88 | ||
| Image 3 | Corn | 228 | 171 | 57 | |
| Soybean | 248 | 186 | 62 | ||
| Other | 350 | 262 | 88 |
| Crop Type | AEZ | Number of Hyperion Images | Growth Stage | Sample Size (N) | ||
|---|---|---|---|---|---|---|
| Total | Training | Validation | ||||
| Corn | 6, 8, 9, and 10 | 48 | Emerge_VEarly | 190 | 142 | 48 |
| Early_Mid | 265 | 198 | 67 | |||
| Late | 696 | 522 | 174 | |||
| Critical | 779 | 584 | 195 | |||
| Mature_Senesc | 499 | 374 | 125 | |||
| Harvest | 128 | 96 | 32 | |||
| Cotton | 2, 5, and 9 | 25 | Emerge_VEarly | 215 | 161 | 54 |
| Early_Mid | 316 | 237 | 79 | |||
| Critical | 197 | 147 | 50 | |||
| Mature_Senesc | 81 | 60 | 21 | |||
| Harvest | 14 | 10 | 4 | |||
| Rice | 7 | 2 | Early_Mid | 65 | 48 | 17 |
| Late | 51 | 38 | 13 | |||
| Soybean | 6, 9, and 10 | 36 | Emerge_VEarly | 132 | 99 | 33 |
| Early_Mid | 581 | 435 | 146 | |||
| Late | 291 | 218 | 73 | |||
| Critical | 815 | 611 | 204 | |||
| Mature_Senesc | 183 | 137 | 46 | |||
| Harvest | 84 | 63 | 21 | |||
| Winter Wheat | 5 and 9 | 24 | Emerge_VEarly | 760 | 570 | 190 |
| Late | 37 | 27 | 10 | |||
| Critical | 70 | 52 | 18 | |||
| Mature_Senesc | 474 | 355 | 119 | |||
| Number | This Study | Thenkabail et al. (2014) | Thenkabail et al. (2013) | Feature |
|---|---|---|---|---|
| 1 | 447 | None | 450 | Nitrogen, Senescing: sensitivity to changes in leaf nitrogen. reflectance changes due to pigments is moderate to low. Sensitive to senescing (yellow and yellow green leaves). |
| 2 | 488 * | 490 | 490 | Carotenoid, Light use efficiency (LUE), Stress in vegetation: Sensitive to senescing and loss of chlorophyll\browning, ripening, crop yield, and soil background effects. |
| 3 | 529 | 531 | 531 | Light use efficiency (LUE), Xanthophyll cycle, Stress in vegetation, pest and disease: Senescing and loss of chlorophyll\browning, ripening, crop yield, and soil background effects. |
| 4 | 569 | 570 | 570 | Pigments (Anthrocyanins, Chlorophyll), Nitrogen: negative change in reflectance per unit change in wavelength is maximum as a result of sensitivity to vegetation vigor, pigment, and N. |
| 5 | 681 * | 682 | 687 | Biophysical quantities and yield: leaf area index, wet and dry biomass, plant height, grain yield, crop type, crop discrimination. |
| 6 | 722 * | 720 | 720 | Stress and chlorophyll: Nitrogen stress, crop stress, crop growth stage studies. |
| 7 | 763 * | None | 760 | Biophysical quantities and yield: leaf area index, wet and dry biomass, plant height, grain yield, crop type, crop discrimination, total chlorophyll. |
| 8 | 803 | None | None | Water absorption band. |
| 9 | 844 | 855 | 855 | Biophysical quantities and yield: leaf area index, wet and dry biomass, plant height, grain yield, crop type, crop discrimination, total chlorophyll. |
| 10 | 923 | 910 | None | Biophysical quantities and yield: leaf area index, wet and dry biomass, plant height, grain yield, crop type, crop discrimination, total chlorophyll. |
| 11 | 993 | 970 | 970 | Moisture, biomass, and protein: peak NIR reflectance. Useful for computing crop moisture sensitivity index. |
| 12 | 1033 | None | 1045 | Biophysical and biochemical quantities: leaf area index, wet and dry biomass, plant height, grain yield, crop type, crop discrimination, total chlorophyll, anthocyanin, carotenoids. |
| 13 | 1074 | 1075 | None | Biophysical and biochemical quantities: leaf area index, wet and dry biomass, plant height, grain yield, crop type, crop discrimination, total chlorophyll, anthocyanin, carotenoids. |
| 14 | 1175 | 1180 | 1180 | Water absorption band. |
| 15 | 1215 | None | None | Water sensitivity: water band index, leaf water, biomass. Reflectance peak in 1050–1300 nm. |
| 16 | 1255 | 1245 | 1245 | Water sensitivity: water band index, leaf water, biomass. Reflectance peak in 1050–1300 nm. |
| 17 | 1316 | None | None | Water sensitivity: water band index, leaf water, biomass. Reflectance peak in 1050–1300 nm. |
| 18 | 1528 | 1518 | None | Moisture and biomass: A point of most rapid rise in spectra with unit change in wavelength in SWIR. Sensitive to plant moisture. |
| 19 | 1568 | None | 1548 | Moisture and biomass: A point of most rapid rise in spectra with unit change in wavelength in SWIR. Sensitive to plant moisture. |
| 20 | 1609 | None | 1620 | Heavy metal stress, Moisture sensitivity: Heavy metal stress due to reduction in Chlorophyll. Sensitivity to plant moisture fluctuations in ESWIR. Use as an index with 1548 or 1620 or 1690 nm. |
| 21 | 1649 | 1650 | 1650 | Heavy metal stress, Moisture sensitivity: Heavy metal stress due to reduction in Chlorophyll. Sensitivity to plant moisture fluctuations in ESWIR. Use as an index with 1548 or 1620 or 1690 nm. |
| 22 | 1699 | None | 1690 | Lignin, biomass, starch, moisture: sensitive to lignin, biomass, starch. Discriminating crops and vegetation. |
| 23 | 1760 | None | 1760 | Water absorption band: highest moisture absorption trough in FSWIR. Use as an index with any one of 2025 nm, 2133 nm, and 2213 am. Affected by noise at times. |
| 24 | 2063 | None | 2050 | Water absorption band: highest moisture absorption trough in FSWIR. Use as an index with any one of 2025 nm, 2133 nm, and 2213 am. Affected by noise at times. |
| 25 | 2103 | 2133 | 2133 | Litter (plant litter), lignin, cellulose: typically highest reflectivity in FSWIR for vegetation. Litter-soil differentiation. |
| 26 | 2163 | None | 2173 | Litter (plant litter), lignin, cellulose: typically highest reflectivity in FSWIR for vegetation. Litter-soil differentiation. |
| 27 | 2204 | 2205 | 2205 | Litter, lignin, cellulose, sugar, starch, protein; Heavy metal stress: typically, second highest reflectivity in FSWIR for vegetation. Heavy metal stress due to reduction in Chlorophyll. |
| 28 | 2254 | 2260 | None | Moisture and biomass: moisture absorption trough in far shortwave infrared. A point of most rapid change in slope of spectra based on land cover, vegetation type, and vigor. |
| 29 | 2295 | 2295 | 2295 | Stress: sensitive to soil background and plant stress. |
| 30 | 2345 | 2359 | None | Cellulose, protein, nitrogen: sensitive to crop stress, lignin, and starch. |
| Band | Frequency of Occurrence | Region |
|---|---|---|
| 447 | 14 | VIS |
| 722 | 11 | RE |
| 803 | 4 | NIR |
| 923 | 7 | H20 |
| 2345 | 14 | SWIR |
| 488 | 9 | VIS |
| 844 | 4 | NIR |
| 993 | 3 | H20 |
| 2063 | 13 | SWIR |
| 529 | 6 | VIS |
| 1033 | 2 | NIR |
| 2295 | 10 | SWIR |
| 681 | 4 | VIS |
| 1074 | 2 | NIR |
| 1316 | 9 | SWIR |
| 569 | 3 | VIS |
| 763 | 1 | NIR |
| 2163 | 7 | SWIR |
| 1255 | 5 | SWIR |
| 1568 | 5 | SWIR |
| 2254 | 5 | SWIR |
| 1175 | 4 | SWIR |
| 1528 | 4 | SWIR |
| 1649 | 4 | SWIR |
| 1699 | 4 | SWIR |
| 1609 | 3 | SWIR |
| 1760 | 2 | SWIR |
| 2103 | 2 | SWIR |
| 1215 | 1 | SWIR |
| 2204 | 1 | SWIR |
| Corn | Cotton | Rice | Soybean | Winter Wheat | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AEZ | Hyperion Image & Year | Producer’s Accuracy (%) | User’s Accuracy (%) | Producer’s Accuracy (%) | User’s Accuracy (%) | Producer’s Accuracy (%) | User’s Accuracy (%) | Producer’s Accuracy (%) | User’s Accuracy (%) | Producer’s Accuracy (%) | User’s Accuracy (%) | Overall Accuracy (%) |
| AEZ 5 | Image 1 (2015) | - | - | 100 | 100 | - | - | - | - | 87.5 | 84.0 | 92.7 |
| AEZ 6 | Image 1 (2011) | 92.3 | 88.9 | - | - | - | - | 91.1 | 97.6 | - | - | 95.4 |
| AEZ 6 | Image 2 (2012) | 98.0 | 96.0 | - | - | - | - | 86.4 | 90.5 | - | - | 95.1 |
| AEZ 6 | Image 3 (2012) | 95.7 | 89.8 | - | - | - | - | 95.8 | 95.8 | - | - | 95.2 |
| AEZ 7 | Image 1 (2012) | 75.0 | 85.7 | - | - | 68.8 | 100 | - | - | - | - | 90.4 |
| AEZ 7 | Image 2 (2012) | 87.5 | 53.9 | - | - | 92.3 | 100 | - | - | - | - | 89.9 |
| AEZ 9 | Image 1 (2010) | 75.0 | 81.8 | 25.0 (75.0) * | 50.0 (85.7) * | - | - | 70.2 | 67.4 | 82.2 | 92.5 | 86.3 (88.8) |
| AEZ 9 | Image 2 (2014) | 90.0 | 90.0 | - | - | - | - | 59.1 | 56.5 | 37.1 (54.3) * | 48.2 (55.9) * | 73.5 (75.5) * |
| AEZ 10 | Image 1 (2013) | 95.6 | 98.5 | - | - | - | - | 92.8 | 91.4 | - | - | 93.1 |
| AEZ 10 | Image 2 (2013) | 92.1 | 90.6 | - | - | - | - | 95.7 | 93.1 | - | - | 93.0 |
| AEZ 10 | Image 3 (2015) | 94.6 | 89.8 | - | - | - | - | 93.6 | 89.2 | - | - | 90.1 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aneece, I.; Thenkabail, P. Accuracies Achieved in Classifying Five Leading World Crop Types and their Growth Stages Using Optimal Earth Observing-1 Hyperion Hyperspectral Narrowbands on Google Earth Engine. Remote Sens. 2018, 10, 2027. https://doi.org/10.3390/rs10122027
Aneece I, Thenkabail P. Accuracies Achieved in Classifying Five Leading World Crop Types and their Growth Stages Using Optimal Earth Observing-1 Hyperion Hyperspectral Narrowbands on Google Earth Engine. Remote Sensing. 2018; 10(12):2027. https://doi.org/10.3390/rs10122027
Chicago/Turabian StyleAneece, Itiya, and Prasad Thenkabail. 2018. "Accuracies Achieved in Classifying Five Leading World Crop Types and their Growth Stages Using Optimal Earth Observing-1 Hyperion Hyperspectral Narrowbands on Google Earth Engine" Remote Sensing 10, no. 12: 2027. https://doi.org/10.3390/rs10122027
APA StyleAneece, I., & Thenkabail, P. (2018). Accuracies Achieved in Classifying Five Leading World Crop Types and their Growth Stages Using Optimal Earth Observing-1 Hyperion Hyperspectral Narrowbands on Google Earth Engine. Remote Sensing, 10(12), 2027. https://doi.org/10.3390/rs10122027