ROS Homeostasis and Plant Salt Tolerance: Plant Nanobiotechnology Updates
Abstract
1. Introduction
2. ROS Generation in Plants under Salt Stress
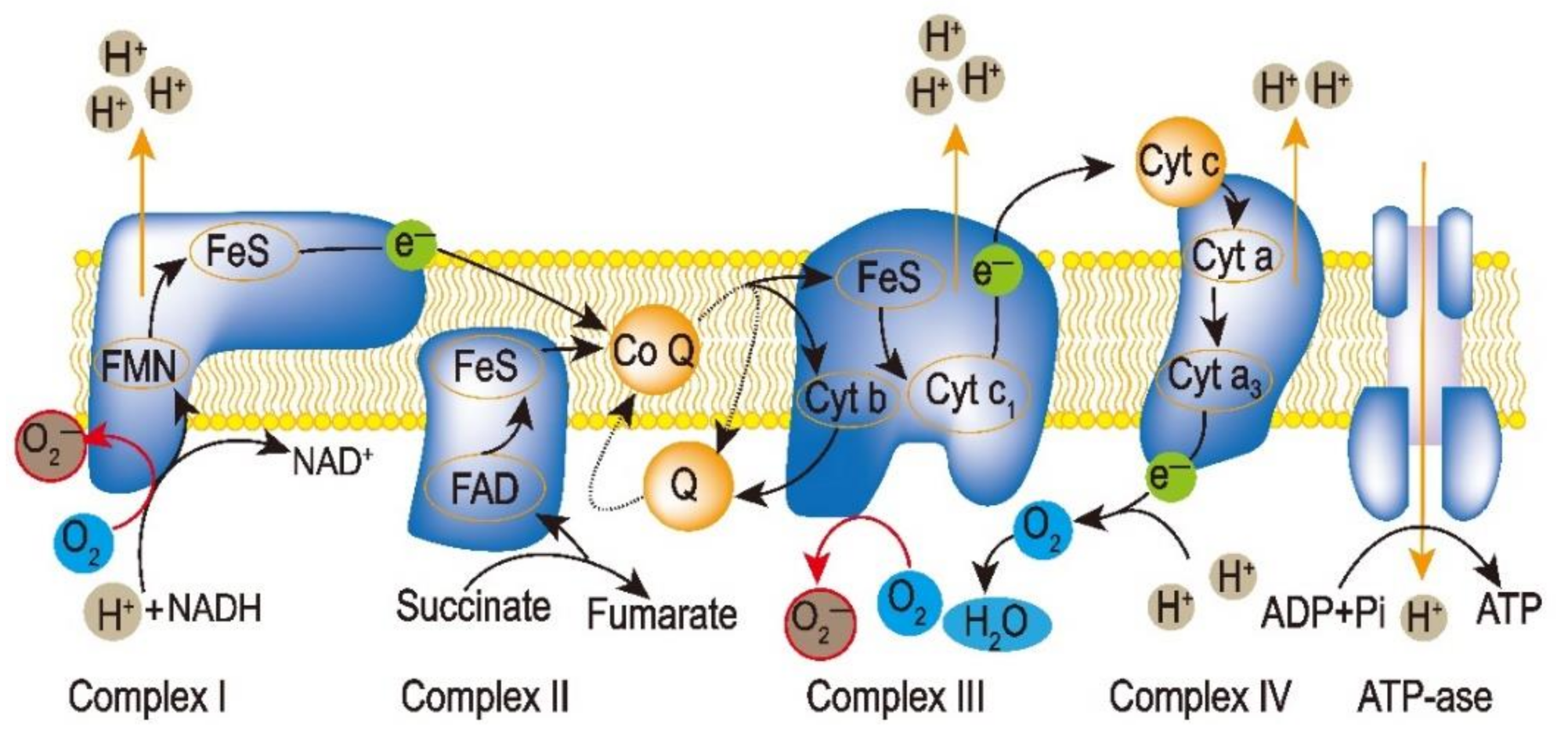
2.1. ROS in Mitochondria
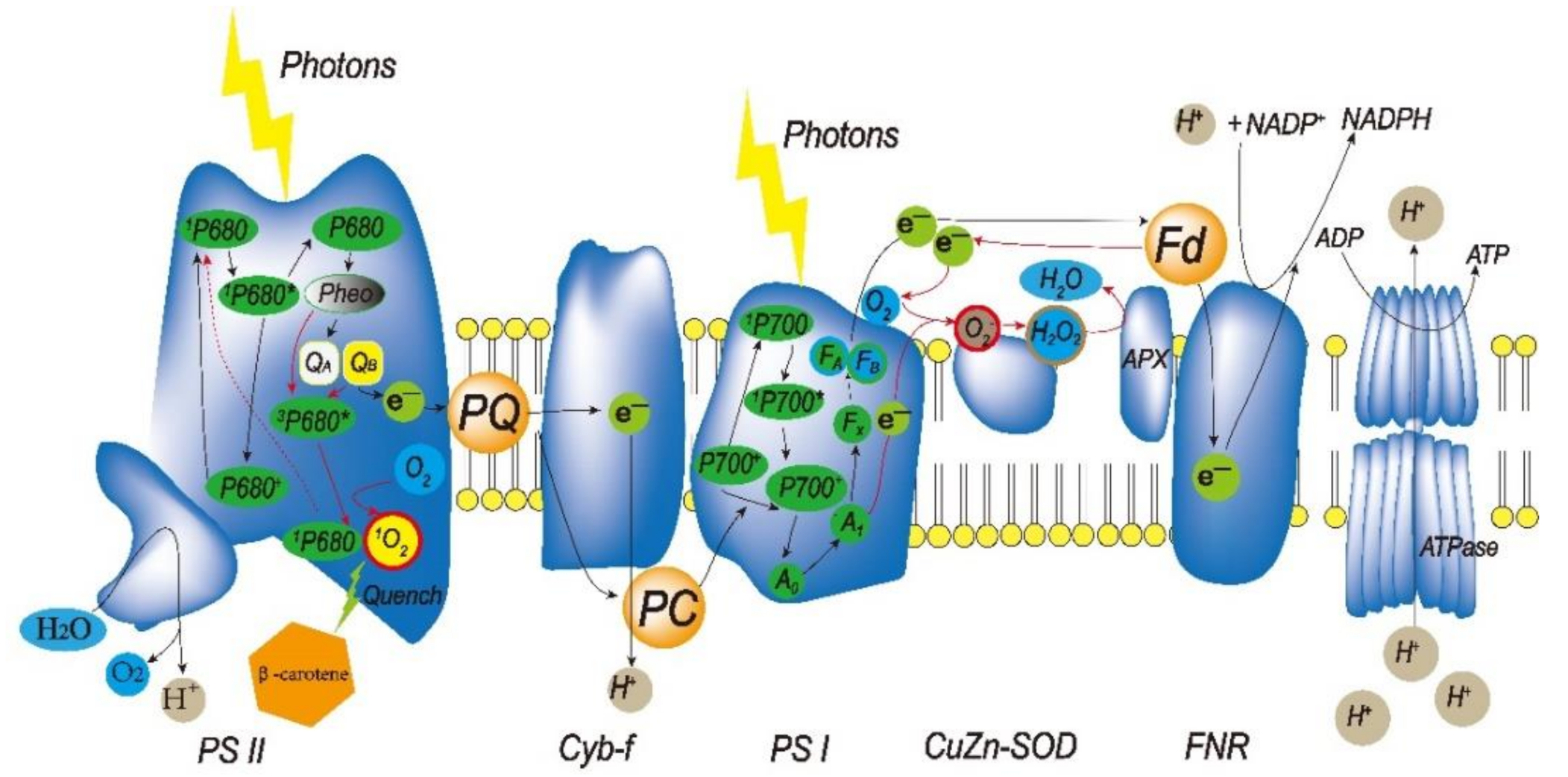
2.2. ROS in Chloroplasts
2.3. ROS in the Apoplast
2.4. ROS in Peroxisomes
3. ROS Scavenging in Plants
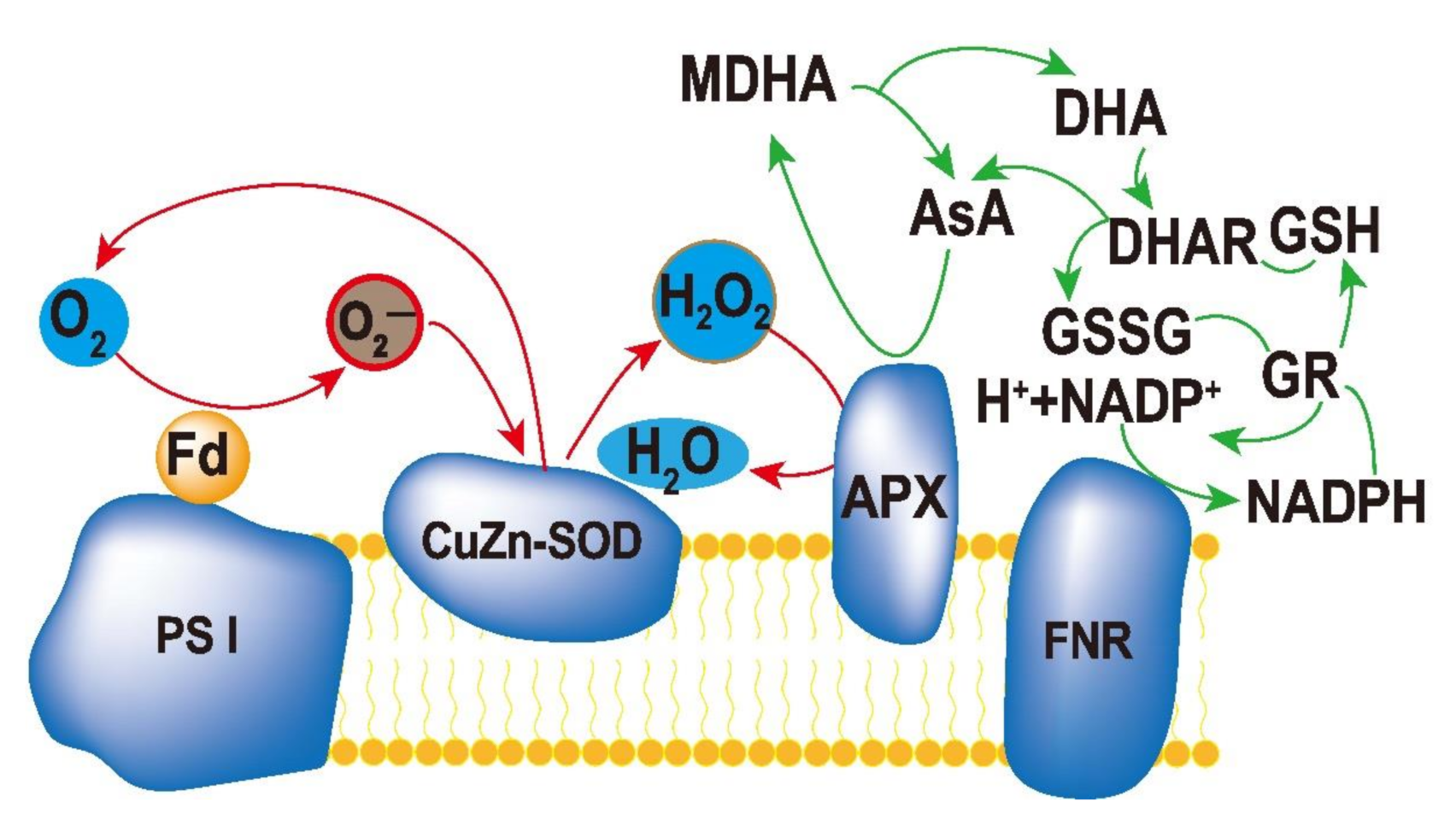
3.1. Enzymatic Scavenging System
- (1) 2H2O → 4[e−] + 4H+ + O2 (The photolysis of H2O in PSII);
- (2) 2O2 + 2[e−] → 2 O2− (Photoreduction of O2 in PSI);
- (3) 2O2− + 2H+ → H2O2 + O2 (SOD catalyzes O2− disproportionation);
- (4) H2O2 + 2AsA → 2H2O + 2MDA (APX catalyzes ASA reduction H2O2);
- (5) 2MDA + 2[e−] + 2H+ → 2AsA.
- Total: 2H2O + O2− → O2 + 2H2O.
3.2. Non-Enzme Scavenging System
4. Plant Nanobiotechnology Approach to Modulating ROS Homeostasis and Plant Salt Tolerance
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- FAO. The Future of Food and Agriculture—Trends and Challenges. Annual Report. FAO: Rome, Italy, 2017. [Google Scholar]
- Shin, W.; Siddikee, A.; Joe, M.M.; Benson, A.; Kim, K.; Selvakumar, G.; Kang, Y.; Jeon, S.; Samaddar, S.; Chatterjee, P. Halotolerant Plant Growth Promoting Bacteria Mediated Salinity Stress Amelioration in Plants. Korean J. Soil Sci. Fertil. 2016, 49, 355–367. [Google Scholar] [CrossRef]
- Dat, J.F.; Vandenabeele, S.; Van Montagu, M.; Van Breusegem, F. Dual Action of The Active Oxygen Species During Plant Stress Responses. Cell. Mol. Life Sci. 2000, 57, 779–795. [Google Scholar] [CrossRef]
- Halliwell, B. Phagocyte-Derived Reactive Species: Salvation or Suicide? Trends Biochem. Sci. 2006, 31, 509–515. [Google Scholar] [CrossRef] [PubMed]
- Gill, S.S.; Tuteja, N. Reactive Oxygen Species and Antioxidant Machinery in Abiotic Stress Tolerance in Crop Plants. Plant Physiol. Biochem. 2010, 48, 909–930. [Google Scholar] [CrossRef] [PubMed]
- Gapper, C.; Dolan, L. Control of Plant Development by Reactive Oxygen Species. Plant Physiol. 2006, 141, 341–345. [Google Scholar] [CrossRef]
- Cai, J.; Chen, T.; Wang, Y.; Qin, G.; Tian, S. Slrem1 Triggers Cell Death by Activating an Oxidative Burst and Other Regulators. Plant Physiol. 2020, 183, 717–732. [Google Scholar] [CrossRef] [PubMed]
- Kora, D.; Bhattacharjee, S. The Interaction of Reactive Oxygen Species and Antioxidants at the Metabolic Interface in Salicylic Acid-Induced Adventitious Root Formation in Mung Bean [Vigna radiata (L.) R. Wilczek]. J. Plant Physiol. 2020, 248, 153152. [Google Scholar] [CrossRef]
- Qi, X.H.; Chen, M.Y.; Liang, D.N.; Xu, Q.; Zhou, F.C.; Chen, X.H. Jasmonic Acid, Ethylene and ROS are Involved in the Response of Cucumber (Cucumis sativus L.) to Aphid Infestation. Sci. Hortic. 2020, 269, 109421. [Google Scholar] [CrossRef]
- Piacentini, D.; Corpas, F.; D’angeli, S.; Altamura, M.; Falasca, G. Cadmium and Arsenic-Induced-Stress Differentially Modulates Arabidopsis Root Architecture, Peroxisome Distribution, Enzymatic Activities and Their Nitric Oxide Content. Plant Physiol. Biochem. 2020, 148, 312–323. [Google Scholar] [CrossRef]
- Zhang, R.; Xue, H.L.; Si, M.; Bi, Y.; Nan, M.N.; Zong, Y.Y.; Long, H.T.; Prusky, D.; Cheng, X.Y. Mechanism of Ca2+-Mediated NOX Modulated in ROS Metabolism Induced by T-2 Toxin in Potato Tuber. Food Chem. 2020, 317, 126416. [Google Scholar] [CrossRef]
- Steffens, B.; Steffens, B. The Role of Ethylene and Ros In Salinity, Heavy Metal, and Flooding Responses in Rice. Front. Plant Sci. 2014, 5, 685. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.J.; Yu, X.Z.; Li, Y.H.; Yang, L. Inhibition of the Mitochondrial Respiratory Components (Complex I and Complex III) as Stimuli to Induce Oxidative Damage in Oryza sativa L. under Thiocyanate Exposure. Chemosphere 2020, 243, 125472. [Google Scholar] [CrossRef]
- Chandrakar, V.; Yadu, B.; Meena, R.K.; Dubey, A.; Keshavkant, S. Arsenic-Induced Genotoxic Responses and Their Amelioration by Diphenylene Iodonium, 24-Epibrassinolide and Proline in Glycine Max L. Plant Physiol. Biochem. 2017, 112, 74–86. [Google Scholar] [CrossRef]
- Tuteja, N.; Singh, M.B.; Misra, M.K.; Bhalla, P.L.; Tuteja, R. Molecular Mechanisms of DNA Damage and Repair: Progress in Plants. Crit. Rev. Biochem. Mol. Biol. 2001, 36, 337–397. [Google Scholar] [CrossRef]
- Racchi, M.L. Antioxidant Defenses in Plants with Attention to Prunus and Citrus spp. Antioxidants 2014, 2, 340–369. [Google Scholar] [CrossRef]
- Mostofa, M.G.; Hossain, M.A.; Fujita, M. Trehalose Pretreatment Induces Salt Tolerance in Rice (Oryza Sativa L.) Seed-Lings: Oxidative Damage and Co-Induction of Antioxidant Defense and Glyoxalase Systems. Protoplasma 2015, 252, 461–475. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Zhou, Y.; Zhai, H.; He, S.; Zhao, N.; Liu, Q. A Novel Sweetpotato Wrky Transcription Factor, Ibwrky2, Positively Regulates Drought and Salt Tolerance in Transgenic Arabidopsis. Biomolecules 2020, 10, 506. [Google Scholar] [CrossRef]
- Khpalwak, W.; Abdel-dayenm, S.M.; Sakugawa, H. Individual and Combined Effects of Fluoranthene, Phenanthrene, Mannitol and Sulfuric Acid on Marigold (Calendula officinalis). Ecotoxicol. Environ. Safe. 2018, 148, 834–841. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.; Chen, Y.E.; Zhao, Y.Q.; Ding, C.B.; Liao, J.Q.; Hu, C.; Zhou, L.J.; Zhang, Z.W.; Yuan, S.; Yuan, M. Exogenous Melatonin Alleviates Oxidative Damages and Protects Photosystem II in Maize Seedlings Under Drought Stress. Front. Plant Sci. 2019, 10, 677. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.N.; Khan, Z.; Luo, T.; Liu, J.; Rizwan, M.; Fahad, S.; Zhang, J.; Xu, Z.; Wu, H.; Hu, L. Seed Priming with Gibberellic Acid and Melatonin in Rapeseed: Consequences for Improving Yield and Seed Quality under Drought and Non-Stress Conditions. Ind. Crop Prod. 2020, 156, 112850. [Google Scholar] [CrossRef]
- Liu, J.; Li, G.; Chen, L.; Gu, J.; Wu, H.; Li, Z. Cerium Oxide Nanoparticles Improves Cotton Salt Tolerance by Enabling Better Ability to Maintain Cytosolic K+/Na+ ratio. Preprint. [CrossRef]
- Wu, H.; Tito, N.; Giraldo, J.P. Anionic Cerium Oxide Nanoparticles Protect Plant Photosynthesis from Abiotic Stress by Scavenging Reactive Oxygen Species. ACS Nano 2017, 11, 11283–11297. [Google Scholar] [CrossRef]
- Djanaguiraman, M.; Nair, R.; Giraldo, J.P.; Prasad, P.V.V. Cerium Oxide Nanoparticles Decrease Drought-Induced Oxidative Damage in Sorghum Leading to Higher Photosynthesis and Grain Yield. Acs Omega 2018, 3, 14406–14416. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharjee, S. Reactive Oxygen Species and Oxidative Burst: Roles in Stress, Senescence and Signal Transduction in Plants. Curr. Sci. 2005, 89, 1113–1121. [Google Scholar]
- Rhoads, D.M.; Umbach, A.L.; Subbaiah, C.C.; Siedow, J.N. Mitochondrial Reactive Oxygen Species. Contribution to Oxidative Stress and Interorganellar Signaling. Plant Physiol. 2006, 141, 357–366. [Google Scholar] [CrossRef] [PubMed]
- Luo, L.; He, Y.; Zhao, Y.; Xu, Q.; Wu, J.; Ma, H.; Guo, H.; Bai, L.; Zuo, J.; Zhou, J.M.; et al. Regulation of Mitochondrial NAD Pool via NAD(+) Transporter 2 is Essential for Matrix NADH Homeostasis and ROS Production in Arabidopsis. Sci. China Life Sci. 2019, 62, 991–1002. [Google Scholar] [CrossRef]
- Crichton, P.G.; Lee, Y.; Kunji, E.R.S. The Molecular Features of Uncoupling Protein 1 Support a Conventional Mitochondrial Carrier-Like Mechanism. Biochimie 2017, 134, 35–50. [Google Scholar] [CrossRef]
- Begcy, K.; Mariano, E.D.; Mattiello, L.; Nunes, A.V.; Mazzafera, P.; Maia, I.G.; Menossi, M. An Arabidopsis Mitochondrial Uncoupling Protein Confers Tolerance to Drought and Salt Stress in Transgenic Tobacco Plants. PLoS ONE 2011, 6, e23776. [Google Scholar] [CrossRef]
- Qin, X.H.; Duan, Z.K.; Zheng, Y.; Liu, W.C.; Guo, S.Y.; Botella, J.R.; Song, C.P. ABC1K10a, an Atypical Kinase, Functions in Plant Salt Stress Tolerance. BMC Plant Biol. 2020, 20, 270. [Google Scholar] [CrossRef]
- Navrot, N.; Rouhier, N.; Gelhaye, E.; Jacquot, J.P. Reactive Oxygen Species Generation and Antioxidant Systems in Plant Mitochondria. Physiol. Plant. 2007, 129, 185–195. [Google Scholar] [CrossRef]
- Filiz, E.; Ozyigit, I.I.; Saracoglu, I.A.; Uras, M.E.; Sen, U.; Yalcin, B. Abiotic Stress-Induced Regulation of Antioxidant Genes in Different Arabidopsis Ecotypes: Microarray Data Evaluation. Biotech. Biotech. Equipment 2019, 33, 128–143. [Google Scholar] [CrossRef]
- Oertli, J.J. Extracellular Salt Accumulation, A Possible Mechanism of Salt Injury in Plants. Agrochim. Pisa 1968, 12, 461–469. [Google Scholar]
- Bouché, N.; Fromm, H. Gaba in Plants: Just A Metabolite? Trends Plant Sci. 2004, 9, 110–115. [Google Scholar] [CrossRef]
- Che-Othman, M.H.; Jacoby, R.P.; Millar, A.H.; Taylor, N.L. Wheat Mitochondrial Respiration Shifts from the Tricarboxylic Acid Cycle to the GABA Shunt under Salt Stress. New Phytol. 2020, 225, 1166–1180. [Google Scholar] [CrossRef] [PubMed]
- Fromm, S.; Senkler, J.; Eubel, H.; Peterhänsel, C.; Braun, H.P. Life Without Complex I: Proteome Analyses of an Arabidopsis Mutant Lacking the Mitochondrial NADH Dehydrogenase Complex. J. Exp. Bot. 2016, 67, 3079–3093. [Google Scholar] [CrossRef] [PubMed]
- Juszczuk, I.M.; Szal, B.; Rychter, A.M. Oxidation-Reduction and Reactive Oxygen Species Homeostasis in Mutant Plants with Respiratory Chain Complex I Dysfunction. Plant Cell Environ. 2012, 35, 296–307. [Google Scholar] [CrossRef]
- Liu, C.Y.; Lee, C.F.; Wei, Y.H. Role of Reactive Oxygen Species-Elicited Apoptosis in the Pathophysiology of Mitochondrial and Neurodegenerative Diseases Associated with Mitochondrial DNA Mutations. J. Formos. Med. Assoc. 2009, 108, 599–611. [Google Scholar] [CrossRef]
- Roach, T.; Krieger-Liszkay, A. Regulation of Photosynthetic Electron Transport and Photoinhibition. Curr. Protein Pept. Sci. 2014, 15, 351–362. [Google Scholar] [CrossRef]
- Alfonso, D.L.; Puppo, A. Reactive Oxygen Species in Plant Signaling; Springer: Berlin/Heidelberg, Germany, 2009. [Google Scholar] [CrossRef]
- Oukarroum, A.; Bussotti, F.; Goltsev, V.; Kalaji, H.M. Correlation Between Reactive Oxygen Species Production and Photochemistry of Photosystems I and Ii in Lemna Gibba L. Plants Under Salt Stress. Environ. Exp. Bot. 2015, 109, 80–88. [Google Scholar] [CrossRef]
- Mendoza, F.; Berry, C.; Prestigiacomo, L.; van Hoewyk, D. Proteasome Inhibition Rapidly Exacerbates Photoinhibition and Impedes Recovery During High Light Stress in Chlamydomonas Reinhardtii. BMC Plant Biol. 2020, 20, 10–22. [Google Scholar] [CrossRef] [PubMed]
- Gal, A.; Zer, H.; Ohad, I. Redox-controlled Thylakoid Protein Phosphorylation. Physiol. Plant. 1997, 100, 863–868. [Google Scholar] [CrossRef]
- Smaili, S.S.; Hsu, Y.T.; Youle, R.J.; Russell, J.T. Mitochondria in Ca2+ Signaling and Apoptosis. J. Bioenerg. Biomembr. 2000, 32, 35–46. [Google Scholar] [CrossRef]
- Gao, Y.; Ma, J.; Zheng, J.; Chen, J.; Chen, M.; Zhou, Y.; Fu, J.; Xu, Z.; Ma, Y. The Elongation Factor GmEF4 is Involved in the Response to Drought and Salt Tolerance in Soybean. Inter. J. Mol. Sci. 2019, 20, 3001. [Google Scholar] [CrossRef]
- Wu, H.; Shabala, L.; Zhou, M.; Shabala, S. Chloroplast-Generated ROS Dominate NaCl-Induced K+Efflux in Wheat Leaf Mesophyll. Plant Signal. Behav. 2015, 10, E1013793. [Google Scholar] [CrossRef] [PubMed]
- Lodeyro, A.F.; Giró, M.; Poli, H.O.; Bettucci, G.; Cortadi, A.; Ferri, A.M.; Carrillo, N. Suppression of Reactive Oxygen Species Accumulation in Chloroplasts Prevents Leaf Damage but Not Growth Arrest in Salt-Stressed Tobacco Plants. PLoS ONE 2016, 11, 7. [Google Scholar] [CrossRef]
- Rahantaniaina, M.S.; Tuzet, A.; Mhamdi, A.; Noctor, G. Missing Links in Understanding Redox Signaling via Thiol/Disulfide Modulation: How is Glutathione Oxidized in Plants? Front. Plant Sci. 2013, 4, 477. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Luo, L.; Xu, J.; Xin, P.; Guo, H.; Wu, J.; Bai, L.; Wang, G.; Chu, J.; Zuo, J.; et al. Malate Transported from Chloroplast to Mitochondrion Triggers Production of ROS and PCD in Arabidopsis Thaliana. Cell Res. 2018, 28, 448–461. [Google Scholar] [CrossRef]
- Zhao, Y.; Yu, H.; Zhou, J.M.; Smith, S.M.; Li, J. Malate Circulation: Linking Chloroplast Metabolism to Mitochondrial ROS. Trends Plant Sci. 2020, 25, 446–454. [Google Scholar] [CrossRef]
- Koch, W.; Kwart, M.; Laubner, M.; Heineke, D.; Stransky, H.; Frommer, W.B.; Tegeder, M. Reduced Amino Acid Content in Transgenic Potato Tubers Due to Antisense Inhibition of the Leaf H+/Amino Acid Symporter StAAP1. Plant J. 2003, 33, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, Y.; Wang, Y. Apoplastic Proteases - Powerful Weapons Against Pathogen Infection in Plants. Plant Commu. 2020, 1, 4. [Google Scholar] [CrossRef]
- Mohanta, T.K.; Bashir, T.; Hashem, A.; Allah, E.F.A.; Khan, A.L.; Al-Harrasi, A.S. Early Events in Plant Abiotic Stress Signaling: Interplay Between Calcium, Reactive Oxygen Species and Phytohormones. J. Plant Growth Regul. 2018, 37, 1033–1049. [Google Scholar] [CrossRef]
- Roychoudhury, A.; Basu, S. Ascorbate-Glutathione and Plant Tolerance to Various Abiotic Stresses; IK International: Delhi, India, 2012. [Google Scholar]
- Podgorska, A.; Burian, M.; Szal, B. Extra-Cellular but Extra-Ordinarily Important for Cells: Apoplastic Reactive Oxygen Species Metabolism. Front. Plant Sci. 2017, 8, 1353. [Google Scholar] [CrossRef]
- Minibayeva, F.; Kolesnikov, O.; Chasov, A.; Beckett, R.P.; Luthje, S.; Vylegzhanina, N.; Buck, F.; Bottger, M. Wound-Induced Apoplastic Peroxidase Activities: Their Roles in the Production and Detoxification of Reactive Oxygen Species. Plant Cell Environ. 2009, 32, 497–508. [Google Scholar] [CrossRef]
- Pang, C.H.; Wang, B.S.; Beyschlag, W.; Murata, J. Oxidative Stress and Salt Tolerance in Plants. Prog. Bot. 2008, 69, 231–245. [Google Scholar]
- Farvardin, A.; González-Hernández, A.I.; Llorens, E.; García-Agustín, P.; Scalschi, L.; Vicedo, B. The Apoplast: A Key Player in Plant Survival. Antioxidants 2020, 9, 604. [Google Scholar] [CrossRef]
- Moschou, P.N.; Paschalidis, K.A.; Delis, I.D.; Andriopoulou, A.H.; Lagiotis, G.D.; Yakoumakis, D.I.; Roubelakis-Angelakis, K.A. Spermidine Exodus and Oxidation in the Apoplast Induced by Abiotic Stress is Responsible for H2O2 Signatures That Direct Tolerance Responses in Tobacco. Plant Cell 2008, 20, 1708–1724. [Google Scholar] [CrossRef]
- Landi, M.; Degl’Innocentie, E.; Pardossi, A.; Guidi, L. Antioxidant and Photosynthetic Responses in Plants Under Boron Toxicity: A Review. Am. J. Agric. Biol. Sci. 2012, 7, 255–270. [Google Scholar] [CrossRef]
- Islinger, M.; Grille, S.; Fahimi, H.D.; Schrader, M. The Peroxisome: An Update on Mysteries. Histochem. Cell Biol. 2012, 137, 547–574. [Google Scholar] [CrossRef] [PubMed]
- del Río, L.A. Peroxisomes as A Cellular Source of Reactive Nitrogen Species Signal Molecules. Arch. Biochem. Biophys. 2011, 506, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Foyer, C.H.; Noctor, G. Redox Sensing and Signalling Associated with Reactive Oxygen in Chloroplasts, Peroxisomes and Mitochondria. Physiol. Plant. 2003, 119, 355–364. [Google Scholar] [CrossRef]
- Sandalio, L.M.; del Río, L.A. Intraorganellar Distribution of Superoxide Dismutase in Plant Peroxisomes (Glyoxysomes and Leaf Peroxisomes). Plant Physiol. 1988, 88, 1215–1218. [Google Scholar] [CrossRef]
- Sandalio, L.M.; Dalurzo, H.C.; Gómez, M.; Romero-Puertas, M.C.; del Río, L.A. Cadmium-Induced Changes in The Growth and Oxidative Metabolism of Pea Plants. J. Exp. Bot. 2001, 52, 2115–2126. [Google Scholar] [CrossRef]
- Jimenez, A.; Hernandez, J.A.; Del Rio, L.A.; Sevilla, F. Evidence for the Presence of The Ascorbate-Glutathione Cycle in Mitochondria and Peroxisomes of Pea Leaves. Plant Physiol. 1997, 114, 275–284. [Google Scholar] [CrossRef]
- Hasanuzzaman, M.; Borhannuddin, B.M.H.M.; Zulfiqar, F.; Raza, A.; Mohsin, S.M.; Mahmud, J.A.; Fujita, M.; Fotopoulos, V. Reactive Oxygen Species and Antioxidant Defense in Plants under Abiotic Stress: Revisiting the Crucial Role of a Universal Defense Regulator. Antioxidants 2020, 9, 681. [Google Scholar] [CrossRef] [PubMed]
- Kritikou, E.A.; Milstein, S.; Vidalain, P.O.; Lettre, G.; Bogan, E.; Doukoumetzidis, K.; Gray, P.; Chappell, T.G.; Vidal, M.; Hengartner, M.O.C. Elegans Gla-3 is a Novel Component of the MAP Kinase MPK-1 Signaling Pathway Required for Germ Cell Survival. Genes Dev. 2006, 20, 2279–2292. [Google Scholar] [CrossRef] [PubMed]
- Aung, K.; Hu, J. The Arabidopsis Tail-Anchored Protein Peroxisomal and Mitochondrial Division Factor1 is Involved in the Morphogenesis and Proliferation of Peroxisomes and Mitochondria. Plant Cell 2011, 23, 4446–4461. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.H.; Ma, F.W.; Wang, Y.H.; Zhang, J.K. The Responses of The Enzymes Related with Ascorbate–Glutathione Cycle During Drought Stress in Apple Leaves. Acta Physiol. Plant. 2010, 33, 173–180. [Google Scholar] [CrossRef]
- Azam, M.; Hameed, L.; Qadri, R.; Ejaz, S.; Aslam, A.; Khan, M.I.; Shen, J.; Zhang, J.; Nafees, M.; Ahmad, I.; et al. Postharvest Ascorbic Acid Application Maintained Physiological and Antioxidant Responses of Guava (Psidium Guajava L.) at Ambient Storage. Food Sci. Technol. 2020, 40, 1–7. [Google Scholar] [CrossRef]
- Opaliński, Ł.; Veenhuis, M.; Van Der Klei, I.J. Peroxisomes: Membrane Events Accompanying Peroxisome Proliferation. Int. J. Biochem. Cell Biol. 2011, 43, 847–851. [Google Scholar] [CrossRef]
- Orth, T.; Reumann, S.; Zhang, X.; Fan, J.; Wenzel, D.; Quan, S.; Hu, J. The Peroxin11 Protein Family Controls Peroxisome Proliferation in Arabidopsis. Plant Cell 2007, 19, 333–350. [Google Scholar] [CrossRef]
- Luhua, S.; Ciftci-Yilmaz, S.; Harper, J.; Cushman, J.; Mittler, R. Enhanced Tolerance to Oxidative Stress in Transgenic Arabidopsis Plants Expressing Proteins of Unknown Function. Plant Physiol. 2008, 148, 280–292. [Google Scholar] [CrossRef]
- Loggini, B.; Scartazza, A.; Brugnoli, E.; Navari-Izzo, F. Antioxidative Defense System, Pigment Composition, and Photosynthetic Efficiency in Two Wheat Cultivars Subjected to Drought. Plant Physiol. 1999, 119, 3–1091. [Google Scholar] [CrossRef]
- Hasanuzzaman, M.; Borhannuddin, B.M.H.M.; Anee, T.I.; Parvin, K.; Nahar, K.; Mahmud, J.A.; Fujita, M. Regulation of Ascorbate-Glutathione Pathway in Mitigating Oxidative Damage in Plants under Abiotic Stress. Antioxidants 2020, 8, 681195–681203. [Google Scholar] [CrossRef] [PubMed]
- Sarker, U.; Oba, S. Catalase, Superoxide Dismutase and Ascorbate-Glutathione Cycle Enzymes Confer Drought Tolerance of Amaranthus tricolor. Sci. Rep. 2018, 8, 16496. [Google Scholar] [CrossRef]
- Mittler, R. Oxidative Stress, Antioxidants and Stress Tolerance. Trends Plant Sci. 2002, 7, 405–410. [Google Scholar] [CrossRef]
- Diaz-Vivancos, P.; Faize, M.; Barba-Espin, G.; Faize, L.; Petri, C.; Hernández, J.A.; Burgos, L. Ectopic Expression of Cytosolic Superoxide Dismutase and Ascorbate Peroxidase Leads to Salt Stress Tolerance in Transgenic Plums. Plant Biotech. J. 2013, 11, 976–985. [Google Scholar] [CrossRef]
- Prihoda, J.; Tanaka, A.; de Paula, W.B.M.; Allen, J.F.; Tirichine, L.; Bowler, C. Chloroplast-Mitochondria Cross-Talk in Diatoms. J. Exp. Bot. 2012, 63, 1543–1557. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.; Zu, C.; Lu, D.; Zheng, Q.; Shen, J.; Wang, H.; Li, D. Effect of Exogenous Selenium Supply on Photosynthesis, Na+ Accumulation and Antioxidative Capacity of Maize (Zea mays L.) under Salinity Stress. Sci. Rep. 2017, 7, 42039. [Google Scholar] [CrossRef]
- Magwanga, R.O.; Lu, P.; Kirungu, J.N.; Lu, H.; Wang, X.; Cai, X.; Zhou, Z.; Zhang, Z.; Salih, H.; Wang, K.; et al. Characterization of The Late Embryogenesis Abundant (Lea) Proteins Family and Their Role in Drought Stress Tolerance in Upland Cotton. BMC Genet. 2018, 19, 1–31. [Google Scholar] [CrossRef]
- Prashanth, S.R.; Sadhasivam, V.; Parida, A. Over Expression of Cytosolic Copper/Zinc Superoxide Dismutase from A Mangrove Plant Avicennia Marina in Indica Rice Var Pusa Basmati-1 Confers Abiotic Stress Tolerance. Transgenic Res. 2008, 17, 281–291. [Google Scholar] [CrossRef]
- Yıldıztugay, E.; Sekmen, A.H.; Turkan, I.; Kucukoduk, M. Elucidation of Physiological and Biochemical Mechanisms of an Endemic Halophyte Centaurea tuzgoluensis under Salt Stress. Plant Physiol. Biochem. 2011, 49, 816–824. [Google Scholar] [CrossRef]
- Srivastava, A.K.; Esrivastava, S.; Lokhande, V.H.; D’souza, S.F.; Esuprasanna, P. Salt Stress Reveals Differential Antioxidant and Energetics Responses in Glycophyte (Brassica Juncea L.) and Halophyte (Sesuvium Portulacastrum L.). Front. Environ. Sci. 2015, 3. [Google Scholar] [CrossRef]
- Shafi, A.; Chauhan, R.; Gill, T.; Swarnkar, M.K.; Sreenivasulu, Y.; Kumar, S.; Kumar, N.; Shankar, R.; Ahuja, P.S.; Singh, A.K. Expression of SOD and APX Genes Positively Regulates Secondary Cell Wall Biosynthesis and Promotes Plant Growth and Yield in Arabidopsis Under Salt Stress. Plant Mol. Biol. 2015, 87, 615–631. [Google Scholar] [CrossRef]
- Paiva, A.L.S.; Passaia, G.; Lobo, A.K.M.; Jardim-Messeder, D.; Silveira, J.A.; Margis-Pinheiro, M. Mitochondrial Glutathione Peroxidase (Osgpx3) Has A Crucial Role in Rice Protection Against Salt Stress. Environ. Exp. Bot. 2019, 158, 12–21. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhao, H.; Zhou, S.; He, Y.; Luo, Q.; Zhang, F.; Qiu, D.; Feng, J.; Wei, Q.; Chen, L.; et al. Expression of Tagf14b, A 14-3-3 Adaptor Protein Gene from Wheat, Enhances Drought and Salt Tolerance in Transgenic Tobacco. Planta 2018, 248, 117–137. [Google Scholar] [CrossRef]
- Borecký, J.; Nogueira, F.T.S.; De Oliveira, K.A.P.; Maia, I.G.; Vercesi, A.E.; Arruda, P. The Plant Energy-Dissipating Mitochondrial Systems: Depicting the Genomic Structure and The Expression Profiles of The Gene Families of Uncoupling Protein and Alternative Oxidase in Monocots and Dicots. J. Exp. Bot. 2006, 57, 849–864. [Google Scholar] [CrossRef]
- Garg, S.K.; Tripathi, M.; Singh, S.K.; Singh, A. Pentachlorophenol Dechlorination and Simultaneous Cr6+ Reduction by Pseudomonas Putida SKG-1 MTCC (10510): Characterization of PCP Dechlorination Products, Bacterial Structure, and Functional Groups. Environ. Sci. Pollut. Res. 2012, 20, 2288–2304. [Google Scholar] [CrossRef] [PubMed]
- Das, K.; Roychoudhury, A. Reactive Oxygen Species (ROS) and Response of Antioxidants as ROS-Scavengers during Environmental Stress in Plants. Front. Environ. Sci. 2014, 2, 53. [Google Scholar] [CrossRef]
- Morales, M.; Munné-Bosch, S. Malondialdehyde: Facts and Artifacts. Plant Physiol. 2019, 180, 1246–1250. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Tan, D.X.; Allan, A.C.; Zuo, B.; Zhao, Y.; Reiter, R.J.; Wang, L.; Wang, Z.; Guo, Y.; Zhou, J. Chloroplastic Biosynthesis of Melatonin and its Involvement in Protection of Plants from Salt Stress. Sci. Rep. 2017, 7, 41236. [Google Scholar] [CrossRef] [PubMed]
- Zurbriggen, M.D.; Tognetti, V.B.; Fillat, M.F.; Hajirezaei, M.-R.; Valle, E.M.; Carrillo, N. Combating Stress with Flavodoxin: A Promising Route for Crop Improvement. Trends Biotechnol. 2008, 26, 531–537. [Google Scholar] [CrossRef]
- Fisher, B.; Yarmolinsky, D.; Abdel-Ghany, S.; Pilon, M.; Pilon-Smits, E.A.; Sagi, M.; van Hoewyk, D. Superoxide Generated from the Glutathione-mediated Reduction of Selenite Damages the Iron-sulfur Cluster of Chloroplastic Ferredoxin. Plant Physiol. Biochem. 2016, 106, 228–235. [Google Scholar] [CrossRef]
- Singh, S. Nanomaterials Exhibiting Enzyme-Like Properties (Nanozymes): Current Advances and Future Perspectives. Front. Chem. 2019, 7, 46. [Google Scholar] [CrossRef] [PubMed]
- Boghossian, A.A.; Sen, F.; Gibbons, B.M.; Sen, S.; Faltermeier, S.M.; Giraldo, J.P.; Zhang, C.T.; Zhang, J.; Heller, D.A.; Strano, M.S. Application of Nanoparticle Antioxidants to Enable Hyperstable Chloroplasts for Solar Energy Harvesting. Adv. Energy Mater. 2013, 3, 881–893. [Google Scholar] [CrossRef]
- Wu, H.; Shabala, L.; Shabala, S.; Giraldo, J.P. Hydroxyl Radical Scavenging by Cerium Oxide Nanoparticles Improves Arabidopsis Salinity Tolerance by Enhancing Leaf Mesophyll Potassium Retention. Environ. Sci. Nano 2018, 5, 1567–1583. [Google Scholar] [CrossRef]
- Sharpe, E.; Frasco, T.; Andreescu, D.; Andreescu, S. Portable Ceria Nanoparticle-Based Assay for Rapid Detection of Foodantioxidants (Nanocerac). Analyst 2012, 138, 249–262. [Google Scholar] [CrossRef] [PubMed]
- Alharby, H.F.; Metwali, E.M.; Fuller, M.P.; Aldhebiani, A.Y. The Alteration of mRNA Expression of SOD and GPX Genes, and Proteins in Tomato (Lycopersicon esculentum Mill) under Stress of NaCl and/or ZnO Nanoparticles. Saudi J. Biol. Sci. 2016, 23, 773–781. [Google Scholar] [CrossRef] [PubMed]
- Shallan, M.A.; Hassan, H.M.M.; Namich, A.A.M.; Ibrahim, A.A. Biochemical and Physiological Effects of TiO2 and SiO2 Nanoparticles on Cotton Plant Under Drought Stress. Res. J. Pharm. Biol. Chem. Sci. 2016, 7, 1540–1551. [Google Scholar]
- Wu, Z.S.; Ren, W.; Wang, D.W.; Li, F.; Liu, B.; Cheng, H.M. High-Energy MnO2 Nanowire/Graphene and Graphene Asymmetric Electrochemical Capacitors. ACS Nano 2010, 4, 5835–5842. [Google Scholar] [CrossRef]
- Zhao, L.; Lu, L.; Wang, A.; Zhang, H.; Huang, M.; Wu, H.; Xing, B.; Wang, Z.; Ji, R. Nano-Biotechnology in Agriculture: Use of Nanomaterials to Promote Plant Growth and Stress Tolerance. J. Agr. Food Chem. 2020, 68, 1935–1947. [Google Scholar] [CrossRef]
- Newkirk, G.; Wu, H.; Santana, I.; Giraldo, J.P. Catalytic Scavenging of Plant Reactive Oxygen Species in vivo by Anionic Cerium Oxide Nanoparticles. JOVE-J. Visual. Exp. 2018, 138, e58373. [Google Scholar] [CrossRef]
- Carles, L.; Rossi, F.; Joly, M.; Besse-Hoggan, P.; Batisson, I.; Artigas, J. Biotransformation of Herbicides by Aquatic Microbial Communities Associated to Submerged Leaves. Environ. Sci. Pollut. Res. 2016, 24, 3664–3674. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Huang, M.; Huang, Y.; Corvini, P.F.-X.; Ji, R.; Zhao, L. Mn3O4 Nanozymes Boost Endogenous Antioxidant Metabolites in Cucumber (Cucumis Sativus) Plant and Enhance Resistance to Salinity Stress. Environ. Sci. Nano 2020, 7, 1692–1703. [Google Scholar] [CrossRef]
- Li, K.; de Mimérand, Y.D.; Jin, X.; Yi, J.; Guo, J. Metal Oxide (ZnO and TiO2) and Fe-Based Metal–Organic-Framework Nanoparticles on 3D-Printed Fractal Polymer Surfaces for Photocatalytic Degradation of Organic Pollutants. ACS Appl. Nano Mater. 2020, 3, 2830–2845. [Google Scholar] [CrossRef]
- Karami, A.; Sepehri, A. Beneficial Role of MWCNTs and SNP on Growth, Physiological and Photosynthesis Performance of Barley Under NaCl Stress. J. Soil Sci. Plant Nutr. 2018, 18. [Google Scholar] [CrossRef]
- Mahmoud, A.W.M.; Abdeldaym, E.A.; Abdelaziz, S.M.; El-Sawy, M.B.I.; Mottaleb, S.A. Synergetic Effects of Zinc, Boron, Silicon, and Zeolite Nanoparticles on Confer Tolerance in Potato Plants Subjected to Salinity. Agronomy 2019, 10, 19. [Google Scholar] [CrossRef]
- Almutairi, Z.M. Influence of Silver Nano-Particles on The Salt Resistance of Tomato (Solanum Lycopersicum) During Germination. Int. J. Agric. Biol. 2016, 18, 449–457. [Google Scholar] [CrossRef]
- Martínez-Ballesta, M.C.; Zapata, L.; Chalbi, N.; Carvajal, M. Multiwalled Carbon Nanotubes Enter Broccoli Cells Enhancing Growth and Water Uptake of Plants Exposed to Salinity. J. Nanobiotechnol. 2016, 14, 1–14. [Google Scholar] [CrossRef]
- Raliya, R.; Saharan, V.; Dimkpa, C.; Biswas, P. Nanofertilizer For Precision and Sustainable Agriculture: Current State and Future Perspectives. J. Agric. Food Chem. 2018, 66, 6487–6503. [Google Scholar] [CrossRef]
- Xu, H.W.; Song, F.B.; Zhu, X.C.; Tong, S.Y. Photosynthesis, Chlorophyll Fluorescence and Non-Structural Carbohydrates Changes in Husk Leaves of Maize in Black Soils Region of Northeast China. Afr. J. Agric. Res. 2010, 5, 785–791. [Google Scholar] [CrossRef]
- Weber, C.R.; Weiss, M.G. Chlorophyll Mutant in Soybeans Provides Teaching Aid. J. Hered. 1959, 50, 53–54. [Google Scholar] [CrossRef]
- Pagliano, C.; Raviolo, M.; Vecchia, F.D.; Gabbrielli, R.; Gonnelli, C.; Rascio, N.; Barbato, R.; La Rocca, N. Evidence for PSII Donor-Side Damage and Photoinhibition Induced by Cadmium Treatment on Rice (Oryza Sativa L.). J. Photochem. Photobiol. B Biol. 2006, 84, 70–78. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Song, Z.; Yang, J.; Wang, Y.; Han, H.Y. Cobalt Ferrite Nanozyme for Efficient Symbiotic Nitrogen Fixation Via Regulating Reactive Oxygen Metabolism. Environ. Sci. Nano 2021, 8, 188–203. [Google Scholar] [CrossRef]
- Chhipa, H. Nanofertilizers and Nanopesticides for Agriculture. Environ. Chem. Lett. 2017, 15, 15–22. [Google Scholar] [CrossRef]
- Wagner, G.; Korenkov, V.; Judy, J.D.; Bertsch, P.M. Nanoparticles Composed of Zn and ZnO Inhibit Peronospora tabacina Spore Germination in vitro and P. tabacina Infectivity on Tobacco Leaves. Nanomaterials 2016, 6, 50. [Google Scholar] [CrossRef]
- Zhang, Y.; Yan, J.; Avellan, A.; Gao, X.; Matyjaszewski, K.; Tilton, R.D.; Lowry, G.V. Temperature- and PH-Responsive Star Polymers as Nanocarriers with Potential for in vivo Agrochemical Delivery. ACS Nano 2020, 14, 10954–10965. [Google Scholar] [CrossRef] [PubMed]
- Zhao, P.; Cao, L.; Ma, D.; Zhou, Z.; Huang, Q.; Pan, C. Translocation, Distribution and Degradation of Prochloraz-Loaded Mesoporous Silica Nanoparticles in Cucumber Plants. Nanoscale 2018, 10, 1798–1806. [Google Scholar] [CrossRef]
- Santana, I.; Wu, H.; Hu, P.; Giraldo, J.P. Targeted Delivery of Nanomaterials with Chemical Cargoes in Plants Enabled by A Biorecognition Motif. Nat. Commun. 2020, 11, 1–12. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, J.; Fu, C.; Li, G.; Khan, M.N.; Wu, H. ROS Homeostasis and Plant Salt Tolerance: Plant Nanobiotechnology Updates. Sustainability 2021, 13, 3552. https://doi.org/10.3390/su13063552
Liu J, Fu C, Li G, Khan MN, Wu H. ROS Homeostasis and Plant Salt Tolerance: Plant Nanobiotechnology Updates. Sustainability. 2021; 13(6):3552. https://doi.org/10.3390/su13063552
Chicago/Turabian StyleLiu, Jiahao, Chengcheng Fu, Guangjing Li, Mohammad Nauman Khan, and Honghong Wu. 2021. "ROS Homeostasis and Plant Salt Tolerance: Plant Nanobiotechnology Updates" Sustainability 13, no. 6: 3552. https://doi.org/10.3390/su13063552
APA StyleLiu, J., Fu, C., Li, G., Khan, M. N., & Wu, H. (2021). ROS Homeostasis and Plant Salt Tolerance: Plant Nanobiotechnology Updates. Sustainability, 13(6), 3552. https://doi.org/10.3390/su13063552






