GMDS Intragenic Deletions Associate with Congenital Heart Disease including Ebstein Anomaly
Abstract
1. Introduction
2. Materials and Methods
2.1. Clinical Data
2.2. Array-CGH
2.3. Next-Generation Sequencing Targeted Panel
2.4. CNV Detection Using Droplet Digital PCR (ddPCR)
3. Results
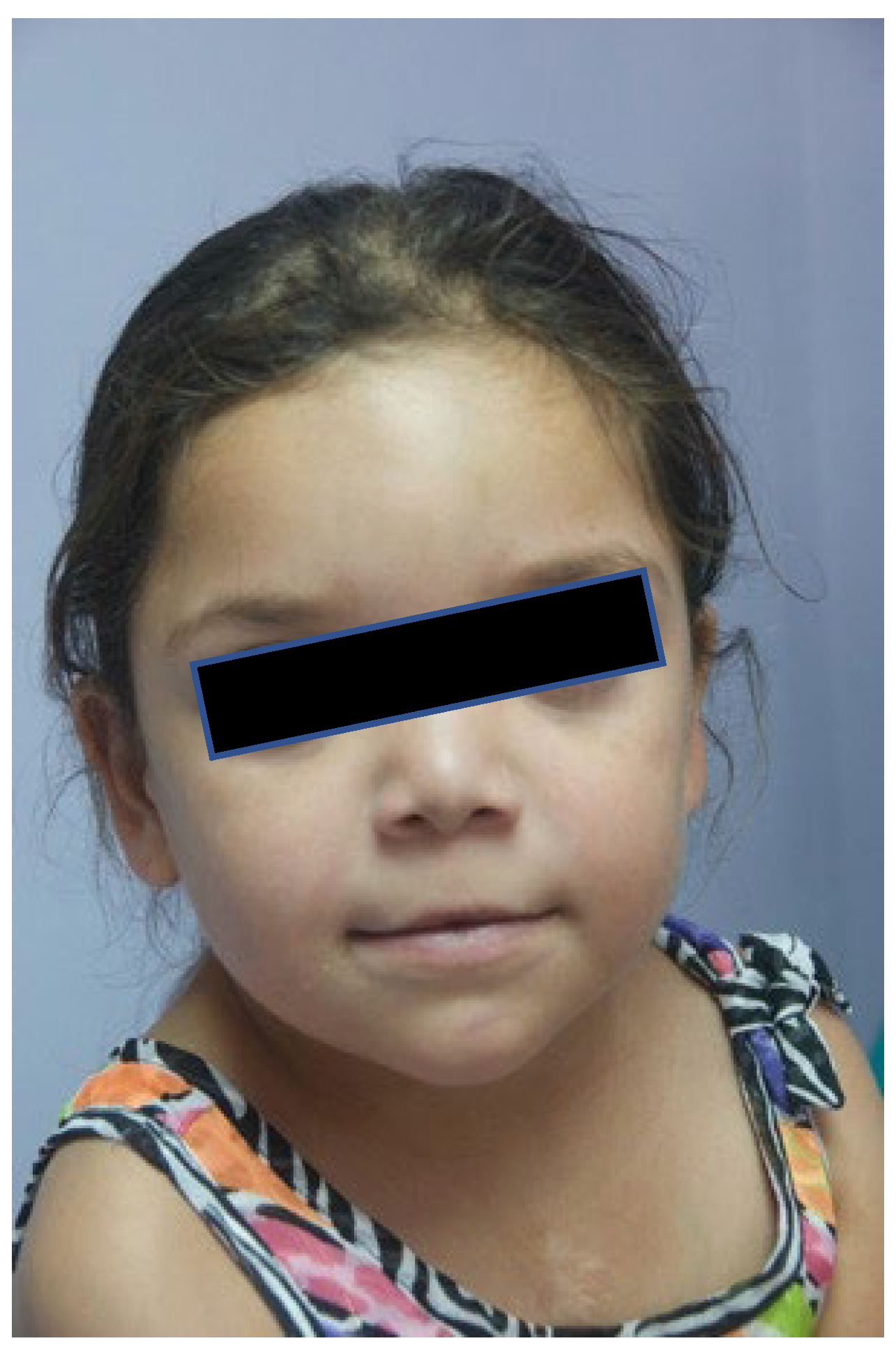
3.1. The Clinical Description of the Affected Patient Is Presented in This Section
3.2. Molecular and Cytogenetic Studies
3.3. Genetic Investigations in Large Ebstein Anomaly Cohort
3.4. Genetic Investigations in Caribbean Ebstein Cohort
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Attenhofer Jost, C.; Connolly, H.; Dearani, J.; Edwards, W.; Danielson, G. Ebstein’s Anomaly. Circulation 2007, 115, 227–285. [Google Scholar] [CrossRef]
- Correa-Villasenor, A.; Ferencz, C.; Neill, C.A.; Wilson, P.D.; Boughman, J.A. Ebstein’s malformation of the tricuspid valve: Genetic and environmental factors. The Baltimore-Washington Infant Study Group. Teratology 1994, 50, 137–147. [Google Scholar] [CrossRef]
- Lupo, P.J.; Langlois, P.H.; Mitchell, L.E. Epidemiology of Ebstein anomaly: Prevalence and patterns in Texas, 1999–2005. Am. J. Med. Genet. Part A 2011, 155, 1007–1014. [Google Scholar] [CrossRef]
- Pradat, P.; Francannet, C.; Harris, J.; Robert, E. The epidemiology of cardiovascular defects, part I: A study based on data from three large registries of congenital malformations. Pediatr. Cardiol. 2003, 24, 195–221. [Google Scholar] [CrossRef] [PubMed]
- Boyle, B.; Garne, E.; Loane, M.; Addor, M.-C.; Arriola, L.; Cavero-Carbonell, C.; Gatt, M.; Lelong, N.; Lynch, C.; Nelen, V.; et al. The changing epidemiology of Ebstein’s anomaly and its relationship with maternal mental health conditions: A European registry-based study. Cardiol. Young 2017, 27, 677–685. [Google Scholar] [CrossRef]
- Sicko, R.J.; Browne, M.L.; Rigler, S.L.; Druschel, C.M.; Liu, G.; Fan, R.; Romitti, P.A.; Caggana, M.; Kay, D.M.; Brody, L.C.; et al. Genetic Variants in Isolated Ebstein Anomaly Implicated in Myocardial Development Pathways. PLoS ONE 2016, 11, e0165174. [Google Scholar] [CrossRef] [PubMed]
- Yuan, S.M. Ebstein’s Anomaly: Genetics, Clinical Manifestations, and Management. Pediatr. Neurol. 2017, 58, 211–215. [Google Scholar] [CrossRef]
- Fesslova, V.; Brankovic, J.; Lalatta, F.; Villa, L.; Meli, V.; Piazza, L.; Ricci, C. Recurrence of congenital heart disease in cases with familial risk screened prenatally by echocardiography. J. Pregnancy 2011, 2011, 368067. [Google Scholar] [CrossRef] [PubMed]
- Hirono, K.; Hata, Y.; Ibuki, K.; Yoshimura, N. Familial Ebstein’s anomaly, left ventricular noncompaction, and ventricular septal defect associated with an MYH7 mutation. J. Thorac. Cardiovasc. 2014, 148, e223–e226. [Google Scholar] [CrossRef]
- Siehr, S.; Punn, R.; Priest, J.; Lowenthal, A. Ebstein anomaly and Trisomy 21: A rare association. Ann. Pediatr. Cardiol. 2014, 7, 67–69. [Google Scholar] [CrossRef]
- Pierpont, M.E.; Basson, C.T.; Benson, D.W., Jr.; Gelb, B.D.; Giglia, T.M.; Goldmuntz, E.; McGee, G.; Sable, D.A.; Srivastava, D.; Webb, C.L. Genetic Basis for Congenital Heart Defects: A Scientific Statement from the American Heart Association Congenital Cardiac Defects Committee, Council on Cardiovascular Disease in the Young. Circulation 2007, 115, 3015–3038. [Google Scholar] [CrossRef] [PubMed]
- Vanderhauwaert, L.G.; Fryns, J.P.; Dumoulin, M.; Logghe, N. Cardiovascular malformations in Turner’s and Noonan’s syn-drome. BHJ 1978, 40, 500–509. [Google Scholar] [CrossRef] [PubMed]
- Digilio, M.C.; Bernardini, L.; Lepri, F.R.; Giuffrida, M.G.; Guida, V.; Baban, A.; Versacci, P.; Capolino, R.; Torres, B.; De Luca, A.; et al. Ebstein anomaly: Genetic heterogeneity and association with microdeletions 1p36 and 8p23.1. Am. J. Med. Genet. Part A 2011, 155, 2196–2202. [Google Scholar] [CrossRef]
- Giannakou, A.; Sicko, R.J.; Zhang, W.; Romitti, P.; Browne, M.L.; Caggana, M.; Brody, L.C.; Jelliffe-Pawlowski, L.; Shaw, G.M.; Kay, D.M.; et al. Copy number variants in Ebstein anomaly. PLoS ONE 2017, 12, e0188168. [Google Scholar] [CrossRef] [PubMed]
- Postma, A.V.; van Engelen, K.; van de Meerakker, J.; Rahman, T.; Probst, S.; Baars, M.J.; Bauer, U.; Pickardt, T.; Sperling, S.R.; Berger, F.; et al. Mutations in the Sarcomere Gene MYH7 in Ebstein Anomaly. Circ. Cardiovasc. Genet. 2011, 4, 43–50. [Google Scholar] [CrossRef] [PubMed]
- Vander Velde, E.; Vriend, J.W.J.; Mannens, M.; Uiterwaal, C.; Brand, R.; Mulder, B.J.M. NEW STUDY CONCOR, an initiative towards a national registry and DNA-bank of patients with congenital heart disease in the Netherlands- Rationale, design, and first results. Eur. J. Epidemiol. 2005, 20, 549–557. [Google Scholar] [CrossRef]
- Available online: https://concor.net/en/aboutconcor/methodology.html (accessed on 27 December 2022).
- Chen, G.; Yu, X.; Zhang, M.; Zheng, A.; Wang, Z.; Zuo, Y.; Liang, Q.; Jiang, D.; Chen, Y.; Zhao, L.; et al. Inhibition of Euchromatic Histone Lysine Methyltransferase 2 (EHMT2) Suppresses the Proliferation and Invasion of Cervical Cancer Cells. Cytogenet. Genome Res. 2019, 158, 205–212. [Google Scholar] [CrossRef]
- Silversides, C.K.; Lionel, A.C.; Costain, G.; Merico, D.; Migita, O.; Liu, B.; Yuen, T.; Rickaby, J.; Thiruvahindrapuram, B.; Marshall, C.R.; et al. Rare Copy Number Variations in Adults with Tetralogy of Fallot Implicate Novel Risk Gene Pathways. PLOS Genet. 2012, 8, e1002843. [Google Scholar] [CrossRef]
- Becker, D.J.; Lowe, J.B. Fucose: Biosynthesis and biological function in mammals. Glycobiology 2003, 13, 41R–53R. [Google Scholar] [CrossRef]
- Available online: http://www.informatics.jax.org/allele/MGI:4363597 (accessed on 1 June 2022).
- Song, Y.; Willer, J.R.; Scherer, P.C.; Panzer, J.A.; Kugath, A.; Skordalakes, E.; Gregg, R.G.; Willer, G.B.; Balice-Gordon, R.J. Neural and synaptic defects in slytherin, a zebrafish model for human congenital disorders of glycosylation. PLoS ONE 2010, 5, e13743. [Google Scholar] [CrossRef]
- Moloney, D.J.; Panin, V.M.; Johnston, S.H.; Chen, J.; Shao, L.; Wilson, R.; Wang, Y.; Stanley, P.; Irvine, K.D.; Haltiwanger, R.S.; et al. Fringe is a glycosyltransferase that modifies Notch. Nature 2000, 406, 369–375. [Google Scholar] [CrossRef] [PubMed]
- MacGrogan, D.; Luna-Zurita, L.; de la Pompa, J.L. Notch Signaling in Cardiac Valve Development and Disease. Birth Defects Res. Part A Clin. Mol. Teratol. 2011, 91, 449–459. [Google Scholar] [CrossRef]
- Niessen, K.; Karsan, A. Notch signaling in cardiac development. Circ. Res. 2008, 102, 1169–1181. [Google Scholar] [CrossRef] [PubMed]
- Fahed, A.C.; Gelb, B.D.; Seidman, J.G.; Seidman, C.E. Genetics of Congenital Heart Disease. The Glass Half Empty. Circ. Res. 2013, 112, 707–720. [Google Scholar] [CrossRef] [PubMed]
- George, R.M.; Maldonado-Velez, G.; Firulli, A.B. The heart of the neural crest: Cardiac neural crest cells in development and regeneration. Development 2020, 147, dev188706. [Google Scholar] [CrossRef] [PubMed]
- Hong, H.K.; Lass, J.H.; Chakravarti, A. Pleiotropic skeletal and ocular phenotypes of the mouse mutation congenital hydro-cephalus (ch/Mf1) arise from a winged helix/forkhead transcriptionfactor gene. Hum. Mol. Genet. 1999, 8, 625–637. [Google Scholar] [CrossRef]
- Seo, S.; Kume, T. Forkhead transcription factors, Foxc1 and Foxc2, are required for the morphogenesis of the cardiac outflow tract. Dev. Biol. 2006, 296, 421–436. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lo-A-Njoe, S.M.; Verberne, E.A.; van der Veken, L.T.; Arends, E.; van Tintelen, J.P.; Postma, A.V.; van Haelst, M.M. GMDS Intragenic Deletions Associate with Congenital Heart Disease including Ebstein Anomaly. Cardiogenetics 2023, 13, 106-112. https://doi.org/10.3390/cardiogenetics13030010
Lo-A-Njoe SM, Verberne EA, van der Veken LT, Arends E, van Tintelen JP, Postma AV, van Haelst MM. GMDS Intragenic Deletions Associate with Congenital Heart Disease including Ebstein Anomaly. Cardiogenetics. 2023; 13(3):106-112. https://doi.org/10.3390/cardiogenetics13030010
Chicago/Turabian StyleLo-A-Njoe, Shirley M., Eline A. Verberne, Lars T. van der Veken, Eric Arends, J. Peter van Tintelen, Alex V. Postma, and Mieke M. van Haelst. 2023. "GMDS Intragenic Deletions Associate with Congenital Heart Disease including Ebstein Anomaly" Cardiogenetics 13, no. 3: 106-112. https://doi.org/10.3390/cardiogenetics13030010
APA StyleLo-A-Njoe, S. M., Verberne, E. A., van der Veken, L. T., Arends, E., van Tintelen, J. P., Postma, A. V., & van Haelst, M. M. (2023). GMDS Intragenic Deletions Associate with Congenital Heart Disease including Ebstein Anomaly. Cardiogenetics, 13(3), 106-112. https://doi.org/10.3390/cardiogenetics13030010






