Abstract
Automated food and drink recognition methods connect to cloud-based lookup databases (e.g., food item barcodes, previously identified food images, or previously classified NIR (Near Infrared) spectra of food and drink items databases) to match and identify a scanned food or drink item, and report the results back to the user. However, these methods remain of limited value if we cannot further reason with the identified food and drink items, ingredients and quantities/portion sizes in a proposed meal in various contexts; i.e., understand from a semantic perspective their types, properties, and interrelationships in the context of a given user’s health condition and preferences. In this paper, we review a number of “food ontologies”, such as the Food Products Ontology/FOODpedia (by Kolchin and Zamula), Open Food Facts (by Gigandet et al.), FoodWiki (Ontology-driven Mobile Safe Food Consumption System by Celik), FOODS-Diabetes Edition (A Food-Oriented Ontology-Driven System by Snae Namahoot and Bruckner), and AGROVOC multilingual agricultural thesaurus (by the UN Food and Agriculture Organization—FAO). These food ontologies, with appropriate modifications (or as a basis, to be added to and further expanded) and together with other relevant non-food ontologies (e.g., about diet-sensitive disease conditions), can supplement the aforementioned lookup databases to enable progression from the mere automated identification of food and drinks in our meals to a more useful application whereby we can automatically reason with the identified food and drink items and their details (quantities and ingredients/bromatological composition) in order to better assist users in making the correct, healthy food and drink choices for their particular health condition, age, body weight/BMI (Body Mass Index), lifestyle and preferences, etc.
Keywords:
food ontologies; Food Informatics; health; diabetes; diet; food scanning; Internet of Things; Semantic Web; linked data 1. Introduction
In this age of the “Quantified Self” [1], a new breed of “automated food scanner” apps, devices and methods is emerging that aims at identifying the exact nature (quantity and bromatological composition) of food and drinks in our diet (and even identify the medicines in our pillboxes), much like a “Shazam” service [2] for all items we might ingest.
Methods used include the following: (i) before ingesting food, food/drink item barcode scanning [3,4], food weighing by portable electronic scales [5], VBM (vision-based measurement) quantification of food volume/weight/portion size by smartphone camera photos [5,6,7,8], remote food and drink recognition by human crowdsourced volunteers or dieticians using smartphone photos of meals sent over the Internet [9], and/or NIR (Near Infrared) spectroscopy (using a handheld sensor/scanner communicating wirelessly with a specialized smartphone app) [10,11]; and (ii) after ingesting food, bioimpedance [12].
Used singly, these methods vary widely in their accuracy and comprehensiveness/suitability in different meal identification scenarios. A smart triangulation of more than one of these methods is likely to produce better and more reliable results in a wider range of meal identification scenarios (as described in “ADAMILO—Automated Diet and Activity Monitoring for Intelligent Lifestyle Optimisation” EU H2020 Research Proposal [13]).
Many of these methods connect to cloud-based (sometimes also crowdsourced) lookup databases (e.g., of food item barcodes, of previously identified food images, or of previously classified NIR spectra of food and drink items) to match and identify a scanned food or drink item, and report the results back to the user. In this paper, we review a number of existing “food ontologies” that can, with appropriate modifications (or as a basis, to be added to and expanded) and together with other relevant non-food ontologies, supplement these lookup databases, in an attempt to progress from the mere automated identification of food and drinks in our meals to a more useful and advanced application, whereby we can automatically reason with the identified food and drink items and details (quantities and ingredients/composition) in order to better assist users in making the correct, healthy food and drink choices that are tailored and personalized for their particular health condition, age, body weight/BMI (Body Mass Index), lifestyle and preferences, etc. (by “reason with” we mean “infer and offer intelligent, personalized and context-sensitive dietary recommendations”).
Such an application can, for example and among other possibilities, help with personalized food menu planning (particularly useful for people with special dietary requirements dictated by their health condition and/or other factors such as meal cost/affordability), and advise users about any essential ingredients lacking in their diet (under-nutrition) or about their intake of substances with cumulative toxicity in daily food, so that they can always stay within the recommended limits. Users can also be warned regarding allergens or any other food intolerances in food they are about to ingest, based on their known medical history of such conditions (e.g., peanut allergy). People with diabetes would benefit from a closer look at the carbohydrates (amounts and composition) profile of their diet, while those with hypertension and cardiovascular disease would appreciate the automated checks on the amounts of salt and saturated fats in their meals. Furthermore, “measured doses of calorie burning via exercise” can be prescribed in the form of tailored and personalized physical activities/exercise regimes, as necessary, based on not just users’ BMI, current calorie intake and expenditure/activity patterns, but also their general health condition and other individual requirements such as hip/knee joint issues, old age and frailty, pollen allergy (workout routes prescribed based on latest pollen forecasts and where needed alternative routes are suggested), etc.
This kind of advanced application and reasoning requires a detailed knowledge of (or ontologies about): (i) individual user’s characteristics and health status/medical history; and (ii) the best, current clinical evidence about nutrition/dietetics and the various disease conditions to be covered; coupled with (iii) knowledge about different foods and drinks, local cuisine characteristics/cooking habits, commercial food/drink product offerings and variants, etc., all of which can vary by locale/country and region of the world. This variation by locale should also be taken into consideration, given that people frequently travel these days for short or prolonged periods for work or leisure.
These complementary types of knowledge and the corresponding ontologies are key to delivering a smart “Internet of Food” that can serve situation, context and user-specific diet insights and “intelligent” recommendations to users based on their actual individual and health needs, circumstances and profiles at any given time, similar to the approach described in [14,15,16] combining multiple ontologies and reasoning with them together. For example, if gluten is identified as part of the composition of a scanned food item (from a food ontology) and we know that the user who is about to ingest that food suffers from coeliac disease (from user’s health profile/medical history instance of a user model ontology) and we know that eating foods containing gluten can trigger a range of mild to severe symptoms in people with celiac disease (from best current clinical evidence in a disease ontology), we can then warn the user that s/he should avoid eating that food item.
An ontology is a formal naming of a set of concepts within a domain and it is used to infer about the properties of that domain [16,17]. As we are rapidly moving towards the Internet of Things (IoT) paradigm, where the deployment of various devices in smart e-health systems faces interoperability and flexibility challenges, ontologies will play an important role in facilitating knowledge sharing and understanding among various interrelated subsystems and knowledge silos. In smart e-health systems, food ontologies offer efficient and flexible means to capture knowledge about dietary concepts, and would therefore provide us with a key required component for designing and implementing the said advanced application.
In the rest of this paper, we start by reviewing a number of existing “food ontologies”, followed by a discussion of their coverage gaps and their potential reuse or adaptation, (as a model to follow, add to, further expand, and combine with other relevant non-food ontologies), in a universal application to automatically reason with scanned food and drink items and their details (quantities and ingredients/bromatological composition) in order to better assist users in making the correct, healthy food and drink choices for their particular health condition, age, body weight/BMI, lifestyle and preferences.
2. Food Ontologies
In this section, we review the following food ontologies: FoodWiki [18], AGROVOC [19], Open Food Facts [20], Food Product Ontology [21], and FOODS (Diabetes Edition) [22]. Table 1 below presents a summary of these methods in terms of their various properties and characteristics. It is obvious that some ontologies are designed to serve a specific purpose, while others are intended to be an open project for collaboration and contribution. AGROVOC is the most mature database/ontology, covering almost all kind of food products. This is not surprising since AGROVOC started more than three decades ago and is supported by world organizations and government agencies. Open Food Facts is becoming a “Wikipedia of food” and follows the same structure of contribution, where contributors manage their own inputs and edits; thus, making it very dynamic with updates added almost every day. FoodWiki and Food Product Ontology are designed to integrate with national databases to track food products produced in their corresponding countries. While these ontologies are helpful, their use is only limited to people tracking packaged food included in some ministerial database. Finally, FOODS has recently evolved to a service targeting people with diabetes and their dietary needs.

Table 1.
Comparison of the studied “food ontologies”.
| Ontology name | Type of food | National/ international coverage | Technology | Availability | Advantages and disadvantages |
|---|---|---|---|---|---|
| FoodWiki [18] | Packaged food | National food products (Turkey) | OWL, RDF | Mobile App for end users | Comprehensive coverage for national packaged food products. Limited only to food products included in the ministry’s database. |
| AGROVOC [19] | All kind of food | Worldwide | OWL, RDF, RDFS, OWL SKOS and SKOS-XL | Free and can be integrated with any application | Comprehensive coverage for international food products (over 50,000 products). Covers all major languages. Only experts and organizations can use it. |
| Open Food Facts [20] | Packaged food | Worldwide | OWL, RDF | Free and can be integrated with any application | Comprehensive coverage of international food product. Data may contain errors. |
| Food Product Ontology [21] | Packaged food | National food products (Russia) | OWL, RDF | NA | Product coverage is not clear. |
| FOODS (A Food-Oriented Ontology-Driven System) [22] | All kind of food | Thailand, but approach can work worldwide | RDF, linked data (from trusted sources) and SPARQL (SPARQL Protocol and RDF Query Language) | Web-based | Has newer version specifically developed for people with diabetes. |
With the above summary in mind, let us now take a look at the details of each of these ontologies.
2.1. FoodWiki
FoodWiki [18] is an ontology-driven, safe food consumption mobile e-health system. It allows users/patients in risk groups to monitor and control their food intake. Specifically, patients will be able to avoid unhealthy ingredients that could worsen their health condition. The mobile e-health system is equipped with a smart knowledge base component and a search mechanism that takes users’ queries and suggests appropriate food consumption relevant to the individual. The intuition is that when people make informed choices about their food consumption, they can maximize their life quality and minimize the usage of unhealthy substances in their meals. Furthermore, many people eat packaged food without paying attention to the ingredients or even worse without proper understanding of the used terminologies in the ingredients section. FoodWiki considers only packaged food products from the market shelves. The system compares and suggests appropriate packaged food products according to the individual’s food intolerance/allergy and other relevant health information. It uses food ontology to share knowledge among mobile smart devices of food consumers and the product database of the Republic of Turkey Ministry of Food, Agriculture and Livestock via ontology parser Web services. The application supports various platforms, such as mobile applications of iOS, Android, and Windows. The aim of FoodWiki is to help and give instant detailed information to food sensitive people about their about-to-be-consumed nutrients and food items from packaged products, and to do so while users are buying these packaged products from store shelves. FoodWiki does not suggest recipes for healthy eating applications, shopping applications, cooking robots and smart fridges.
FoodWiki works as follows: The user, through his/her mobile device, wants to check a packaged product in the supermarket to see if it is suitable for him/her considering his/her health conditions such as food allergy, heart disease, high cholesterol, etc. The system requires scanning the barcode of the packaged food, which provides the entire nutritional information, food additives and energy details of the scanned product through a back-end connection to a product database. This database has a unique International Article Number (EAN—originally called European Article Number, hence EAN, now renamed to International Article Number) for each product. The system starts by searching its ontology knowledgebase and tries to retrieve the nutritional information for the selected product. In addition, FoodWiki compares that nutritional information with the specific nutritional concepts that have negative effects for the consumer’s health. In order to perform this comparison, it needs to reason with the concepts and relationships defined in three foundation ontologies: Human, Disease and Food Ontologies. The system presents three different colors as a result to the consumer: red, green, and yellow. The green light indicates “the product is safe” for the consumer, while the red light is an “objectionable food product”. The yellow light indicates a warning sign, which alerts the user to seek medical advice from a health professional before consuming the product. It must be noted that FoodWiki operates on a national base; i.e., it identifies packaged products that are tracked and included in the database of the Turkish Ministry of Agriculture only.
The ontology design of FoodWiki is based on the well-known Web Ontology Language (OWL). The system uses OWL to express food concepts in ontological form with specific spatial terms and features. The food ontology knowledgebase involves four subsections: Person, Disease, Product, and Food Ingredients/Compounds. Subsections “Person” and “Disease” cover contents related to common allergies such as lactose, nuts, egg etc. The current food ontology contains four main classes, 58 subclasses, 15 object type properties with 17 sub-object type properties, 12 data type properties, 1530 individuals with annotation type properties, and 210 semantic rules. The ontology starts with a “Thing” class that contains four main classes: “Diseases”, “Person”, “Ingredients”, and “Product”. Figure 1 provides a simplified graphical relationship of the class ingredient, which contains two subclasses, food items and food additives.
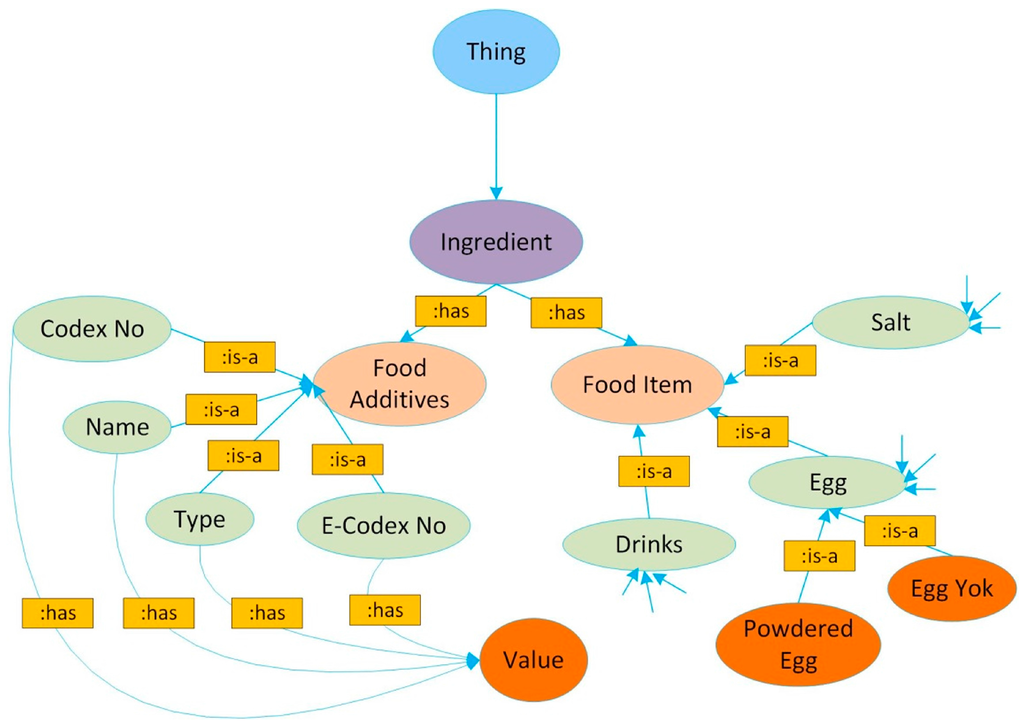
Figure 1.
An example of the “Ingredient Class” of FoodWiki ontology, based on [18].
The philosophy underlying the use of food ontology is to provide a unified vocabulary for an integrated environment that enables the interaction of different ubiquitous devices, such as mobile phones and medical devices, with the aim of providing users with personalized advice. The food concepts described in the FoodWiki ontology encapsulate a subset of instance data from the domain. The ontology knowledgebase provides a standardized method for data matching from the database. Thus, FoodWiki’s aim is to represent an abstract model of the different types of foods available to the users, together with their nutritional information, including the type and amount of nutrients, and the recommended daily intake. This is done by integrating information in natural language so that its interpretation becomes easier. The ontology is also structured in a way that healthy and unhealthy ingredients are categorized in relation to risk groups.
Concept Matching Engine (CME): FoodWiki keeps personal and health data as profile information of users in the Consumer Intolerance List. It uses a content matching engine to generate a new list from the terms in the Consumer Intolerance List. The ontology establishes relations between the concepts in each list using synonyms such as “is_a” and “has additive type”. In addition, the database of the system includes the barcode numbers with the information about the food additives and ingredient or nutrient details of the selected products in the supermarket. Therefore, the system has two lists as inputs to the concept matching engine to perform the semantic enhancement step: The Consumer Intolerance List and the selected Product Ingredients List. The CME generates another list called the Semantically Enhanced Consumer Intolerance List from the Consumer Intolerance List. In order to perform the semantic enhancement task, the system parses appropriate concepts and relationships defined in the food ingredients section of the ontology. Specifically, concepts related to food ingredients and their properties are processed according to the terms in the Consumer Intolerance List [18].
2.2. AGROVOC
AGROVOC [19] is a large and mature multilingual thesaurus covering all areas of interest to the Food and Agriculture Organisation (FAO) of the United Nations. It includes terminology widely used in the practice of subject fields in agriculture, fisheries, forestry, food and related domains. The vocabulary consists of over 32,000 concepts, with up to 40,000 terms in 23 languages. AGROVOC was first developed in 1980 by FAO and the Commission of the European Communities, and is currently maintained by an active international community of experts and institutions. AGROVOC represents a great degree of consensus regarding terminology, and is used by specialized digital libraries and digital repositories for indexing and retrieving data. The structured and controlled vocabulary terms in AGROVOC allow for unambiguous identification of resources, standardization of indexing processes, and efficient searching.
Since its existence, AGROVOC has gone through a series of improvements. It was first deployed using a relational database, but the managing community soon realized the limitation of this approach given the distributed nature of the contributors and the availability of the data to third parties only by means of database dumps or through Web services. Such process of information sharing imposed a huge burden on the developers and maintainers of the applications. In order to overcome this shortcoming, AGROVOC moved to linked data, which offered a standard model that is both human and machine-readable.
Since 2003, several efforts attempted to convert the AGROVOC thesaurus into an ontology. Fisseha, Liang, and Keizer [23] sketched the first concepts to transform AGROVOC to ontology. This was a major milestone towards a proper representation of the AGROVOC vocabulary, but it came short of addressing important aspects related to query processing and completeness of domain-relevant entity types and relationship types. Later, in 2004, Soergel et al. [24] developed an OWL-based ontology for AGROVOC, which was the best option available to share information through the World Wide Web. After that, several ad-hoc solutions were added to the OWL structure to conveniently express terms in all the languages available in AGROVOC. The conversion of AGROVOC to OWL ontology offered a simplified mechanism to model multilingual resources, but was not efficient to deal with a thesaurus resource. In 2009, the World Wide Web Consortium (W3C) recommended the use of Simple Knowledge Organization System (SKOS) [25] for developing specifications and standards to support the use of thesauri and classification schemes within the framework of the Semantic Web. This was the right choice: to represent AGROVOC in SKOS, which is a vocabulary for Resource Description Framework (RDF) developed explicitly to support thesauri. SKOS offers much more flexible semantics compared to OWL and provides specific relationships between concepts and also relationships between their multilingual lexicalizations. AGROVOC is currently represented using SKOS-eXtention of Labels (SKOS-XL). The model was structured with concept schemes, where metadata descriptors about edited resources are provided. It also puts emphasis on multilingualism and the natural language description of resources. Figure 2 shows an example of AGROVOC using RDF/SKOS.
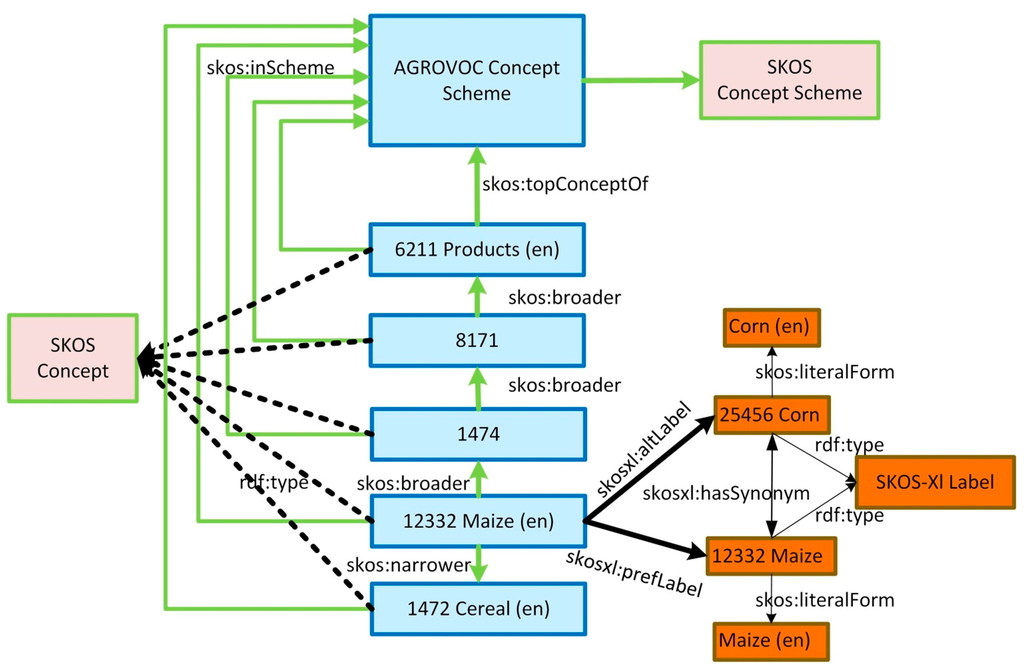
Figure 2.
AGROVOC Simple Knowledge Organization System (SKOS) model, modified from [19].
Old ontology editing tools suffered from several drawbacks, such as poor usability and lack of a collaborative environment supporting roles-based authentication, editorial workflow and multilingual search, among others, which led AGROVOC designers to develop their own editor called VocBench [26], a Web application specifically tailored to the AGROVOC Vocabulary. By using this tool, authenticated users can add or translate AGROVOC terms in their languages. VocBench is a distributed editing framework for thesauri, glossaries, authority lists, code lists, taxonomies and classifications. Changes can be easily tracked with VocBench, which gives the system an important feature to allow individuals and organizations to contribute to AGROVOC while maintaining information about the provenance of their authorship. Moreover, multilingual search, visualization and editing are fundamentals to VocBench. Today, the VocBench user community has grown, and includes FAO’s Fisheries and Aquaculture Department with their data.fao.org project, the European Commission Publications Office, the European Environment Agency, and the Italian Senate, with more organizations joining. The latest production release is VocBench 2.0, released in November 2013 with new features and improvements, such as native SKOS support, support for multiple triple-stores, and OSGi (Open Services Gateway initiative) compliance. Currently, supported vocabularies are RDF, RDFS, OWL (1st version), SKOS and SKOS-XL. A sandbox server for testing is available online, thanks to the Malaysia’s national R&D (Research & Development) center MIMOS Berhad. Today, there is a large community using VocBench, especially public organizations that were interested in open source solutions for maintaining their thesauri, code lists and authority resources.
AGROVOC is now available as a linked data set published, aligned (linked) with several vocabularies. The linked data version of AGROVOC is in RDF/SKOS-XL, and is stored in AllegroGraph TripleStore (a closed source triplestore designed to store RDF triples, a standard format for linked data). In order to support legacy applications and users who are still using old editing tools, all contributions coming from different sources through VocBench are converted to relational database and used for applications based on it. In addition to converting to rational database, an SKOS-XL version is produced from the finalized VocBench version and enriched with metadata descriptors from the VoID (Vocabulary of Interlinked datasets) vocabulary to feed the Linked Open Data (LOD).
AGROVOC uses the linked data Cloud which has links to ten resources, vocabularies, thesauri and ontologies in areas covered by AGROVOC. There are six linked resources covering general areas: The Library of Congress Subject Headings (LCSH), the NAL Thesaurus (US National Agricultural Library Agricultural Thesaurus), RAMEAU (Repertoire d’Autorite-Matiere Encyclopedique et Alphabetique Unifie) Eurovoc, DBpedia, and an experimental linked data version of the Dewey Decimal Classification. The remaining four resources are specific to various areas of interest. The linked resources are mostly thesauri. Since linked data are important, there is continuous effort for enhancing VocBench so that it may natively support RDF/SKOS. This will have several beneficial effects. Firstly, linked data can be distributed from a single triple store, thus minimizing the tedious conversions. Secondly, the tool will be useful to any community that wants to utilize SKOS for its data. In addition, there are efforts for integrating VocBench functionalities with Eclipse (an integrated development environment) and to extract and validate links to other resources. By so doing, VocBench workflow will be aligned with the overall AGROVOC editing workflow. From the content viewpoint, there are plans to continue linking AGROVOC to other resources and to start using skos:closeMatch in addition to skos:exactMatch.
2.3. Open Food Facts
Open Food Facts [20] is a global food database based on contributions from individuals around the world. As of July 2015, the Open Food Facts database covers 50,448 food products from 123 countries. People add products to the database and have the privilege to edit, improve and propose applications ideas through the Idea Forum. The Open Food Facts app [27] allows users to learn about food nutritional information, and compare products from around the world. This is simply done by scanning the barcode of the food product or searching the database. The aim is to connect users with smart tools to help them make better choices about their food consumption. The idea is primarily to encourage users to understand the labels and to better select healthy foods, which contain fewer potentially harmful substances such as saturated fats, free sugars, salts, additives or allergens. A search function is also under consideration, and the data listed on the site can optionally be crossed with other studies to make the link between certain ingredients and certain diseases. Open Food Facts is also beneficial for the food industry to track, monitor, and strategically plan for its food production.
The contributors to the Open Food Facts are usually volunteers who send pictures of the food (mainly packaged food), its labels, ingredients lists and nutrition facts tables. If there are errors in the submitted information, the users may correct the information by themselves. In order to detect potential errors more easily, the Open Food Facts project is considering the addition of automated checks. For example, when there is contradiction between the nutrition facts of a product among several other different products of the same category, the system should be able to detect the error automatically. Contributors can also add or edit food items based on the information explicitly shown on the package. Not only individual users can contribute to the database, but also manufacturers can add their food products after agreeing to the open license agreement.
Another advantage of Open Food Facts is that data can be used freely by individuals, associations, companies, and researchers from all around the world to brainstorm and develop applications for the greater welfare of the people. This means that everyone can share the contents of Open Food Facts through Web sites, services, software, mobile applications, or to write articles and studies. They are free to make the resulting work either freely available or to sell it, as long as they respect the terms of reuse. Open Food Facts supports several technologies for researchers and developers to acquire the data. The data can be exported using MongoDB dumps, CSV (comma-separated values) exports, and RDF data export. The popular and easy-to-use JavaScript Object Notation (JSON) is also available to read the data of a product. JSON is in particular used in the Open Food Facts mobile app for iPhone and Android [27]. Developers can also download the Android App codes from GitHub. As mentioned above, the app allows users to scan the barcode of products, to view the product information, and to take and send pictures and data for missing products.
2.4. Food Product Ontology
The Food Product Ontology [21] describes food products with common representation, vocabulary and language for the food product domain, to help manufacturers, retailers, governments and institutions to publish their data related to this domain in a way that maximizes the reuse of data. The food product ontology allows for better integration, sharing, and collaborative processing of food information among several stakeholders. It extends a widely used standardized ontology for product, price, store and company data, called GoodRelations [28], which is an ontology describing tangible goods and commodity services using structured data in RDF and microformats. The Food Product Ontology was initially built for MneMojno [29], a mobile application that provides users with additional information about a food product that cannot be found on the package, and can assist them in selecting better products.
The food product ontology can be used to serve various parties as follows: On a retailer Web site, food products have only name, description, image and price. A manufacturer’s Web site generally provides more information about their food products such as name, description, images, information about food nutritional value data, contents/ingredients, and other specifications, which are within the domain name space of the respective manufacturer. There may exist actual instances of this food product, or it may inherit its properties from the specifications of food products, which are produced by the same manufacturer. An institution’s Web site such as the UK Food Standards Agency [30] publishes a list of approved additives that can be used in food manufacturing. Usually the list has a label and a brief description of the additives. The above three examples (retailer, manufacturer and institution) show possible relationship among these entities. To model this relationship, the Food Product Ontology extended the classes and properties of GoodRelations to include concepts from the food product domain such as Food and Ingredient, and properties such as energy per 100 grams and carbohydrates per 100 grams. GoodRelations has all the representations of a product and its specification, but usually does not provide a relation to a particular domain. The Food Product Ontology is needed to add a food product domain to describe the new classes and properties within that domain. For example, Food is expressed as a subclass of Product or Service class from the GoodRelations that represents any product. The name of a food product can be annotated with gr:name or rdfs:label properties. Ingredient is a subclass of Thing that is the parent class of any class in OWL language. The rdfs:label property is used to describe a label of an ingredient. Food and Ingredient classes are related to each other through the ingredient object property. “Carbohydrates per 100 g” is a data property that represents amount of carbohydrates (in grams) per 100 grams a food product has. Properties such as carbohydrates per 100 g, energy per 100 g, proteins per 100 g and fat per 100 g are added to the ontology to simplify a food product representation in RDF. Another addition to the Food Product Ontology is the categories of food products (e.g., eggs and egg products). The food product categories are similar to those defined by the CODEX Alimentarius [31]. The CODEX is an organization established by the World Health Organization and FAO. The category system consists of 16 top categories and more than 300 subcategories, with maximum depth equal to 4. An example of the Food Ontology Classes is shown in Figure 3.
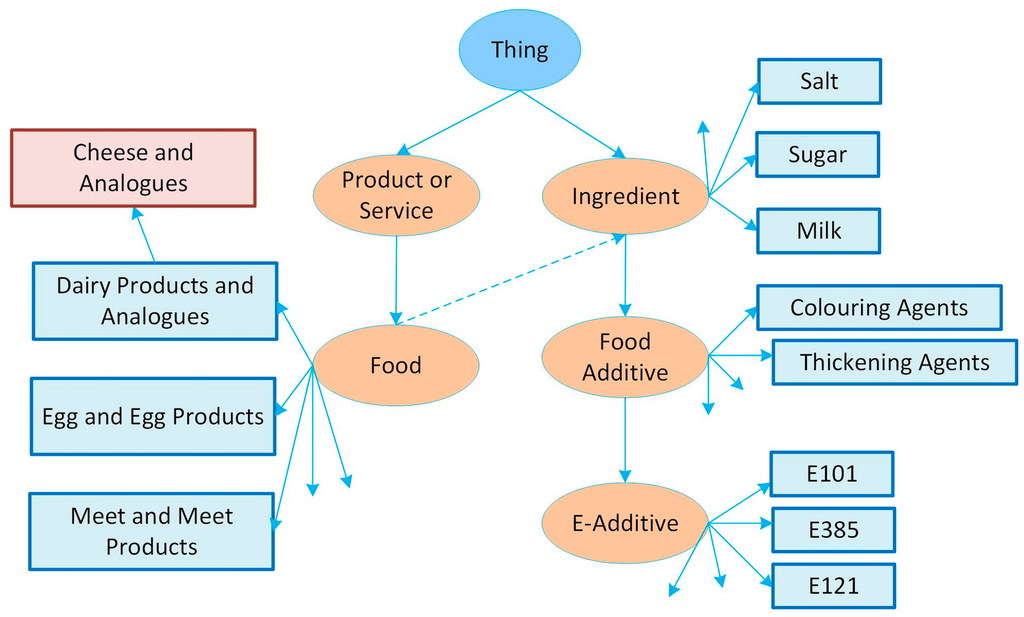
Figure 3.
An example of the Food Ontology Classes, modified from [21].
It is worth noting that Kolchin, one of the developers of the Food Product Ontology, is also working with others on a related FOODpedia, a DBpedia of Food Products [32,33].
2.5. FOODS: A Food-Oriented Ontology-Driven System—Diabetes Edition
This “food ontology” example builds on earlier work by Snae Namahoot and Bruckner [22] to deliver a Web-Based Food Menu Recommender System for Patients with Diabetes in Thailand.
In order to treat patients with type 1 and type 2 diabetes nutrition therapy used within the overall treatment plan. In particular, type 1 diabetes needs patients to follow a meal planning approach, which involves carbohydrate counting and can improve glycemic control (for a recent study, see Evert et al. [34]).
Furthermore, Evert et al. state that “a simple diabetes meal planning approach such as portion control or healthy food choices may be better suited to individuals with type 2 diabetes identified with health and numeracy literacy concerns. This may also be an effective meal planning strategy for older adults”. As a consequence, meal planning is an essential task for individuals (at home) as well as dietitians and physicians (in a clinical setting), and supporting this task with a recommender system would greatly improve the smoothness of the clinical workflow, especially if the recommender system is connected to the clinical system holding appropriate data about the patients.
The food menu planning system described in the rest of this section focuses on adult patients with diabetes, whose dietary and management needs are obviously different from those of children and adolescents. Personal diet recommender systems have to consider the health profiles of the users/patients (e.g., by accessing their Electronic Health Record (EHR)) and their food preferences, representing the objective and the subjective levels of satisfaction with food, respectively. An Electronic Health Record contains medical information collected in a systematic way, thereby covering various individual patient healthcare data and settings (demographics, medical history, medications, immunization status, laboratory test results, among others).
Personal recommender systems consist of domains, user profiles, and items. The domain covered by the system described below is hospital food planning for in-patients with diabetes. The user profile includes all the data about an individual patient, which are taken into account in the process of deriving the recommendation; typical examples are sex, age, place of birth, diabetes type (clinical diagnosis), and blood sugar level. Items comprise the pieces of information that users can search for in the system and get results on. In the setting of this research, the users can be dietitians, physicians and patients.
Setting up a food plan is not an easy task in the context of hospital management and is mainly the joint effort of physicians and clinical dietitians. It is basically the sum of all individual diet plans of patients in active treatment for a given period of time, e.g., a week of the year. The diet plans can be derived from patients’ data.
The patients’ data are stored in the Electronic Health Record (EHR), which can be accessed by authorized staff in the hospital. The data comprise such elements as personal data (full name, sex, address, height, blood group, allergies), temporal medical data (weight, Body Mass Index (BMI), blood pressure, pregnancy, symptoms and conditions, diagnostic details, treatments, medication, attending doctor) and clinical data (blood sugar level, data of A1c (glycohemoglobin) and fructosamine tests, and OGTT (Oral Glucose Tolerance Test)), among others. All of these are used as input for the recommendation process for the meals, which is described later in this section.
The required calories are calculated on a daily basis, and the meals may include breakfast, snack before lunch, lunch, snack after lunch, dinner and snack before bed. The daily intake of nutrients and calories is then calculated and allocated to the preferred number of meals, which typically contain a mix of the six main ingredients flour, meat, vegetable, fruits, fats, and milk.
Use scenario and the knowledge base: Consider an internal medicine ward specialized in patients with diabetes. The ward comprises 11 persons as inpatients. One of the patients is Chatchai, a 38-year-old man, with a height of 173 cm and a current weight of 86 kg. He maintains a sedentary lifestyle, walks for 30 min at moderate speed and strolls with his child for 20 min. His general BMI is 28.7, which might be considered healthy if the percentage body fat is lower than average. The daily energy expenditure can be calculated as 2586 kcal (calculated after the revised FAO/WHO/UNU (Food and Agriculture Organization/World Health Organization/United Nations University Joint) equations, see [35]), plus 194 kcal used for the exercises [36]. Such a calculation is being performed on a weekly (or even daily) basis supervised by the clinical dietitian for Chatchai. The recommended minimum nutritional intake is shown in Figure 4.
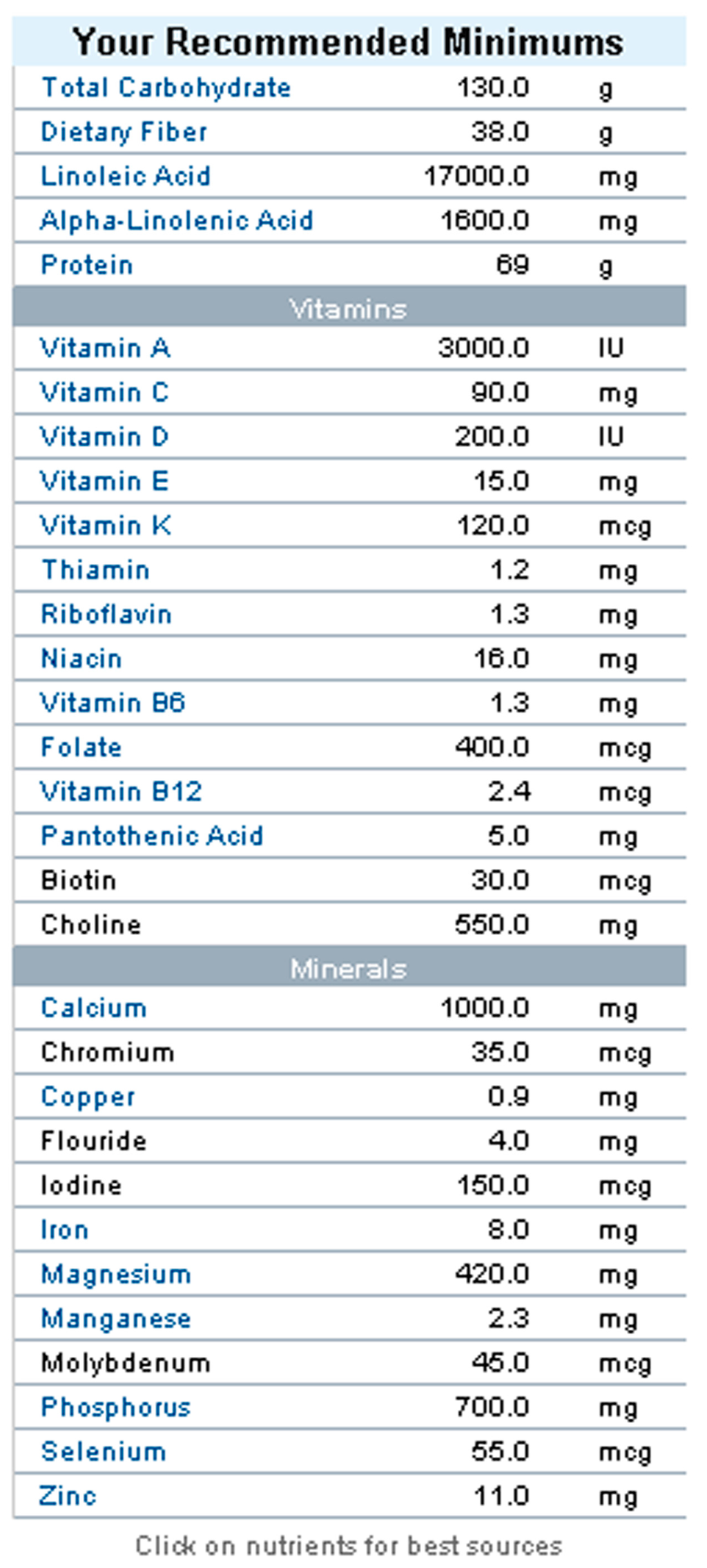
Figure 4.
Recommended nutritional minimums for the person of the use case (values according to [37]).
The next step is to convert these values into a food menu taking into account Chatchai’s preferred foods (Thai meals). This process has to be performed for all other 10 patients with diabetes in the ward. It leads to a well-balanced food menu plan, and the system presented here aims at effectively supporting this process.
The Web-based system can also be used by adults with diabetes who want to get online advice on appropriate foods without the need to wait for seeing a dietitian (Figure 5). As an example, for breakfast Chatchai is recommended to have 3/8 flour products, 1/8 meat, 2/8 vegetables, 1/8 fruits and 1/8 milk products. Also note that the actual user interface is in Thai language, but was translated into English for the purpose of this paper (as can be seen in Figure 5). For adult people with diabetes, access to data input is restricted to personal data and such temporal medical data as weight and blood sugar level, which the patients in the UC scheme (the Thai government’s universal health coverage (UC) scheme for citizens of the country since 2002) can usually measure at home.
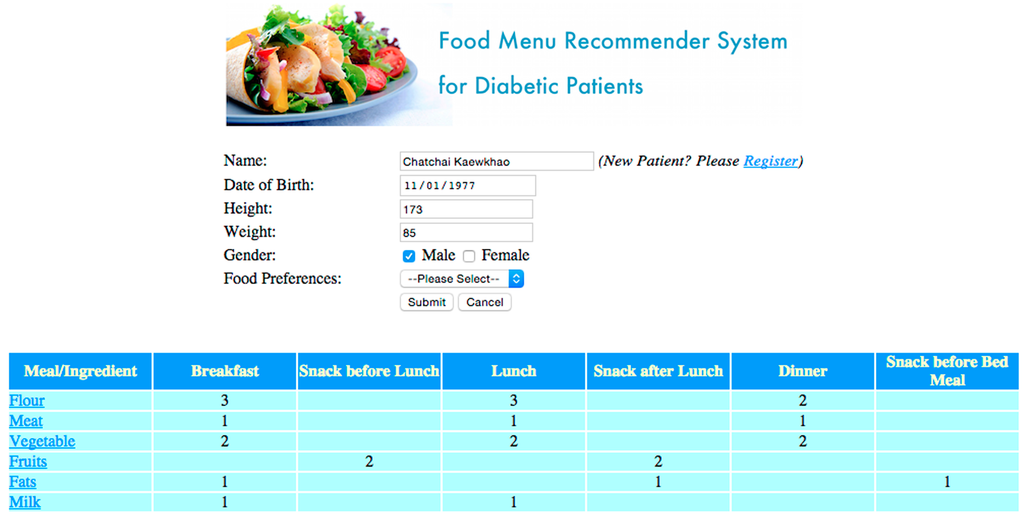
Figure 5.
Screenshot for a registered patient accessing the Food Menu Recommender System and getting the basic recommendations on the intake of flour, meat, vegetables, fruits, fats and milk products.
Research dietitians and physicians continuously work on improving diets for various nutritionally based/diet sensitive disorders, including diabetes. As a consequence, not only the temporal medical data and the lab data have to be updated on demand but also the knowledge base has to be adapted to new recommendations (best current clinical and dietetics evidence). It is, therefore, necessary to design a smart infrastructure for the knowledge base, which adapts to trusted and current data automatically. This problem can be tackled by harnessing appropriate linked data (LD) about required diets and nutritional information for diabetes intervention. We use SPARQL to retrieve acceptable LD from trusted sources.
Food menu planning: The default distribution of caloric intake per day and per meal is breakfast (20%), snack before lunch (10%), lunch (25%), evening snack (10%), dinner (25%), bed tea (10%). The snippet in Figure 6 shows the input and calories calculation process used here, while Figure 7 shows the processing section for the output.
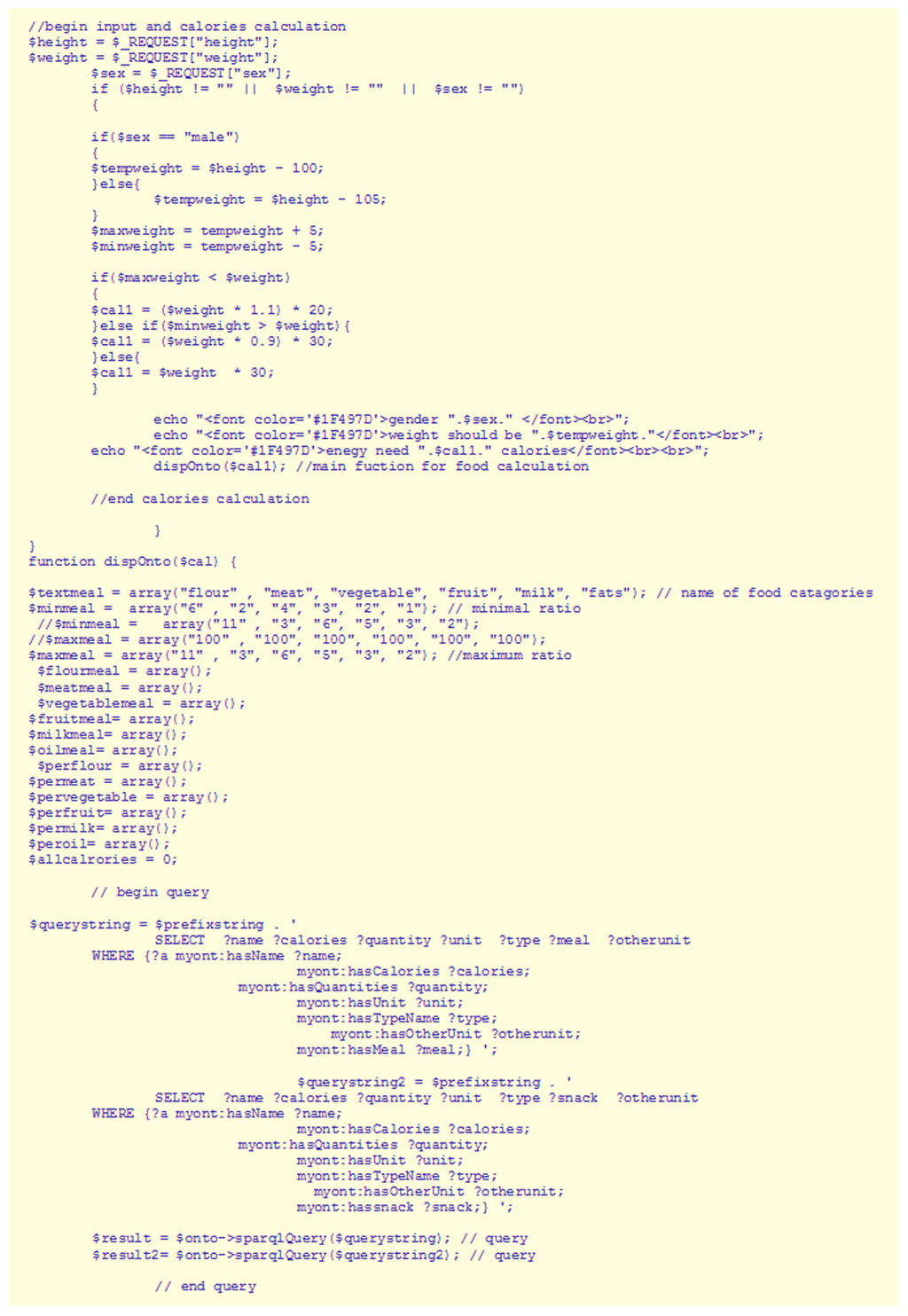
Figure 6.
Code snippet showing the input and calories calculation process in FOODS—Diabetes Edition.

Figure 7.
Code snippet showing the processing section for the output in FOODS—Diabetes Edition.
The output data can be used to set up a weekly dietary plan as has been proposed Lin et al. [38]. This results in a more concrete food plan, which instead of just recommending “1/8 fruits” could say “1 apple” (or, in line with the stated preferences of the individual patient, “1 bunch of grapes”). A further improvement would most likely include monitoring and storing in the EHR the amount per kind of food that was actually consumed by a patient on a daily or weekly basis. This would offer insight into the optimal amount of each ingredient to serve, thereby offering the opportunity to reduce plate waste in health institutions.
3. Discussion and Conclusions
There is a growing demand for healthcare services to help people make informed choices about their wellbeing, including the foods and drinks they consume. Improper eating can contribute to or precipitate diseases, such as diabetes, hypertension, some types of cancer, and some types of allergies, among others [18]. In order to enable lay users and healthcare professionals to gain access to relevant medical knowledge about food products in e-health systems that integrate different technologies and data sources, semantic frameworks containing machine-readable annotations (ontology) about food and other relevant and related domains (e.g., clinical medicine, dietetics, individual user profiles, etc.) are critical for the successful delivery of such smart (context-aware) e-health systems. The ultimate goal is to progress beyond the mere identification of the full details of what is on the user’s plate (what food scanning methods connected to cloud-based lookup databases can do) to answering the key question about “how good or healthy is the food on the user’s plate (or in his/her supermarket shopping basket), given the user’s individual health condition, personal needs and preferences” and making recommendations for specific, personalized dietary improvements as necessary.
In this paper, we reviewed some “food ontologies” and the associated methods and tools they employ, as well as some of their applications. In the same vein as the afore described “Web-Based Food Menu Recommender System for Patients with Diabetes in Thailand”, Tumnark et al. [39] proposed an ontology-based personalized dietary recommendation system for weightlifting athletes, while Chi et al. [40] presented a chronic disease dietary consultation system using ontologies and semantic rules. These ontologies and their applications hint at the potential of the “powerful and smarter semantic reasoning with food data” that can be achieved if such ontologies are combined with other relevant non-food ontologies (e.g., ontologies about diet-sensitive disease conditions—cf. [14,15,16]).
Although none of the reviewed ontologies is fully comprehensive in terms of scope and coverage (some are meant for very specific uses, e.g., in diabetes, and locales, e.g., Thailand), they remain good examples to learn from and might also form a basis, to be added to and further expanded, for future developments towards a universal comprehensive “smart diet assessment and recommendation” engine/application for use by consumers and patients with various diet-sensitive conditions worldwide (i.e., international coverage). Indeed, one of our main reasons for embarking on a review of existing food ontologies was to document their coverage gaps (the incompleteness or limited scope of existing food ontologies could negatively affect the performance of any wider-scope reasoning engine that relies on them). These coverage gaps justify the need, as a future research direction, for developing a universal, comprehensive food ontology (and the appropriate mechanisms for maintaining and updating it), while learning from previous attempts and approaches in this domain. Future developments should also cover other relevant and equally comprehensive non-food ontologies (e.g., user model ontology, clinical dietetics and disease ontology, etc.) to properly reason with food details in the context of individual users and disease conditions.
Although differing but overlapping in coverage (each being more fit for its specific purposes), the five main ontologies presented in this article can also be seen as complementing each other in various ways. For example, Open Food Facts ontology [20] covers 50,448 packaged food products from 123 countries, while FoodWiki [18] focuses on packaged products in the Turkish market only. Both ontologies use OWL language, which means they can be integrated to extend the coverage of food products. Open Food Facts ontology allows for many ways of integration, including MongoDB dumps, CSV exports, and RDF data export, and also uses JavaScript Object Notation (JSON) to read data. This means data about packaged food can be exported to the Open Food Facts database easily. FoodWiki will also be able to use its mobile app to access the information in the Open Food Facts database, in addition to the Turkish Food, Agriculture and Livestock database. Such integration may also motivate other ontologies that track packaged food at the national level to become accessible from a global platform.
FOODS—Diabetes Edition (based on [22]) is an example of such ontologies. It consists of an ontology specifically tailored to Thai food and aims at helping people with diabetes to monitor their food intake through a Web-based system. FOODS covers all kind of Thai food, and not only packaged food items. It similarly uses OWL and can be integrated with Open Food Facts. Furthermore, using the concept of Food Product Ontology [21], the coverage can be extended to help manufacturers, retailers, governments and institutions publish their data related to the food domain in a way that maximizes the reuse of data. Food Product Ontology fosters sharing and collaboration among several stakeholders. The main advantage of Food Product Ontology is that it extends the knowledge to contents and ingredients as described by the manufacturer. Food Product Ontology also uses OWL language to describe the classes and objects of food products.
AGROVOC [19] covers a large and mature multilingual thesaurus of all kinds of food. AGROVOC represents a great degree of consensus regarding terminology, and is used by specialized digital libraries and digital repositories for indexing and retrieving data. From this perspective AGROVOC can be considered as the best candidate (basis) for a unified food ontology that has worldwide coverage of all kinds of food. However, the main issue with AGROVOC is that it is accessed and controlled by experts under the administration of UN agencies. This means there is a conceptual difference on how people interact with AGROVOC and Open Food Facts. In Open Food Facts, a passionate individual may instantly scan a barcode of a packaged food item from any place in the world and add it to Open Food Facts database (crowdsourced input). This option is not available in AGROVOC. From this perspective, the direct integration of both ontologies is not possible, but both of them complement each other and each has its merits regarding the approaches employed. They can thus together form a helpful basis for developing a new, universal and more comprehensive food ontology in the future.
Critical to the success, scalability and long-term sustainability of the above mentioned “smart diet assessment and recommendation” engine will be the quality and accuracy/up-to-datedness of the food products data and the clinical/dietetics evidence in the ontologies powering it. Mechanisms for regular updates (e.g., as medical and dietetics knowledge advances and new research results become available and/or as food manufacturers modify their existing products or release new ones) and for integrity checking/quality assurance (particularly for any crowdsourced input) will become an essential ingredient of such an engine and the associated ontologies. The use of appropriate clinical terminologies (which are themselves ontologies) will also prove indispensable when describing and reasoning with disease conditions and patients’ medical profiles in the context of the “smart diet assessment and recommendation” engine [14,15,16].
Acknowledgments
The authors are grateful for the most valuable support offered by members of the Department of Nutritional Science, Naresuan University, Thailand, who have detailed the basic planning methods for people with diabetes, as applied by Naresuan University Hospital on a regular basis.
Author Contributions
M.N.K.B. conceived and drafted the manuscript with equal contributions from all named coauthors. M.N.K.B. established the rationale for the article, designed the literature review and wrote the larger parts of the “Introduction” and “Discussion and Conclusions” sections. A.Y. and S.S. contributed to the literature review and summary of the selected food ontologies, as well as Figure 1, Figure 2 and Figure 3. The FOODS—Diabetes Edition ontology was developed and described by C.S.N. and M.B., who also contributed Figure 4, Figure 5, Figure 6 and Figure 7. All authors read and approved the final version.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Swan, M. Health 2050: The Realization of Personalized Medicine through Crowdsourcing, the Quantified Self, and the Participatory Biocitizen. J. Pers. Med. 2012, 2, 93–118. [Google Scholar] [CrossRef] [PubMed]
- Shazam (Wikipedia, 15 July 2015). Available online: https://en.wikipedia.org/w/index.php?title=Shazam_(service)&oldid=671402622 (accessed on 4 September 2015).
- ShopWell. Available online: http://www.shopwell.com/ (accessed on 4 September 2015).
- Food Barcode Scanning Apps (Fooducate and FoodScanner). Available online: http://www.webcitation.org/6bIFp0VLR (accessed on 4 September 2015).
- SmartPlate. Available online: http://getsmartplate.com/ (accessed on 4 September 2015).
- Google’s, A.I. Is Training Itself to Count Calories in Food Photos. Available online: http://www.popsci.com/google-using-ai-count-calories-food-photos (accessed on 4 September 2015).
- SRI. Food Recognition Technology. Available online: http://www.sri.com/engage/products-solutions/food-recognition-technology (accessed on 4 September 2015).
- Pouladzadeh, P.; Shirmohammadi, S.; Al-Maghrabi, R. Measuring Calorie and Nutrition from Food Image. IEEE Trans. Instrum. Meas. 2014, 63, 1947–1956. [Google Scholar] [CrossRef]
- Rise. Available online: https://www.rise.us/ (accessed on 4 September 2015).
- Strickland, E. Hand-Held Spectroscopy Tool Lets You Examine the Molecular Composition of Your Food. IEEE Spectrum. 30 April 2014. Available online: http://spectrum.ieee.org/tech-talk/consumer-electronics/gadgets/handheld-spectroscopy-tool-lets-you-examine-the-molecular-composition-of-your-food (accessed on 4 September 2015).
- SCiO Molecular Sensor. Available online: https://www.consumerphysics.com/myscio/scio (accessed on 4 September 2015).
- HEALBE. Available online: http://healbe.com/ (accessed on 4 September 2015).
- Kamel Boulos, M.N. ADAMILO—Automated Diet and Activity Monitoring for Intelligent Lifestyle Optimisation. EU H2020 Research. Proposal 2015. [Google Scholar] [CrossRef]
- Kamel Boulos, M.N.; Harvey, F.E.; Roudsari, A.V.; Bellazzi, R.; Hernando Perez, M.E.; Deutsch, T.; Cramp, D.G.; Carson, E.R. A Proposed Semantic Framework for Diabetes Education Content Management, Customisation and Delivery within the M2DM Project. Comput. Methods Programs Biomed. 2006, 83, 188–197. [Google Scholar] [CrossRef] [PubMed]
- Kamel Boulos, M.N. A first look at HealthCyberMap medical semantic subject search engine. Technol. Health Care 2004, 12, 33–41. [Google Scholar]
- Kamel Boulos, M.N.; Roudsari, A.V.; Carson, E.R. Towards a Semantic Medical Web: HealthCyberMap’s Tool for Building an RDF Metadata Base of Health Information Resources Based on the Qualified Dublin Core Metadata Set. Med. Sci. Monit. 2002, 8, MT124–MT136. [Google Scholar]
- Igor, J.; John, M.; Eric, Y. Using Ontologies for Knowledge Management: An Information Systems Perspective. In Proceedings of the Annual Conference of the American Society for Information Science, Washington, DC, USA, 31 October–4 November 1999.
- Çelik, D. FoodWiki: Ontology-Driven Mobile Safe Food Consumption System. Sci. World J. 2015, 2015. [Google Scholar] [CrossRef] [PubMed]
- Caracciolo, C.; Stellato, A.; Rajbahndari, S.; Morshed, A.; Johannsen, G.; Keizer, J.; Jaques, Y. Thesaurus maintenance, alignment and publication as linked data: The AGROVOC use case. Int. J. Metadata Semant. Ontol. 2012, 7, 65–75. [Google Scholar] [CrossRef]
- Open Food Facts. Available online: http://world.openfoodfacts.org/who-we-are (accessed on 4 September 2015).
- Kolchin, M.; Zamula, D. Food Product Ontology: Initial Implementation of a Vocabulary for Describing Food Products. In Proceeding of the 14th Conference of Open Innovations Association FRUCT, Helsinki, Finland, 11–15 November 2013; pp. 191–196.
- Snae, C.; Bruckner, M. FOODS: A Food-Oriented Ontology-Driven System. In Proceedings of the 2nd IEEE International Conference on Digital Ecosystems and Technologies (DEST 2008), Phitsanulok, Thailand, 26–29 February 2008; pp. 168–176.
- Fisseha, F.; Liang, A.; Keizer, J. Reengineering AGROVOC to Ontologies: Step towards better semantic structure. In Proceedings of the NKOS Workshop, Rice University, Houston, TX, USA, 31 May 2003.
- Soergel, D.; Lauser, B.; Liang, A.; Fisseha, F.; Keizer, J.; Katz, S. Reengineering thesauri for new applications: The AGROVOC example. J. Digit. Inf. JODI 2004, 4, 1–23. [Google Scholar]
- Miles, A.; Bechhofer, S. W3Cb SKOS Simple Knowledge Organization System. (World Wide Web Consortium (W3C), 5 July 2015). Available online: http://www.w3.org/TR/skos-reference/ (accessed on 4 September 2015).
- Sánchez-Alonso, S.; Sicilia, M.-A. Using an AGROVOC-based ontology for the description of learning resources on organic agriculture. Metadata Semant. 2009, 481–492. [Google Scholar]
- Open Food Facts app (Android version). Available online: https://play.google.com/store/apps/details?id=org.openfoodfacts.scanner (accessed on 4 September 2015).
- Hepp, M. GoodRelations: An Ontology for Describing Products and Services Offers on the Web. In Proceedings of the 16th International Conference, EKAW 2008, Acitrezza, Italy, 29 September–2 October 2008; pp. 329–346.
- MneMojno. Available online: http://mnemojno.ru (accessed on 4 September 2015).
- Food Standards Agency. Available online: http://www.food.gov.uk (accessed on 4 September 2015).
- FAO/WHO Food Standards: CODEX Alimentarius Food Category System. Available online: http://www.codexalimentarius.net/gsfaonline/foods/index.html (accessed on 4 September 2015).
- FOODpedia—A DBpedia of Food Products (SlideShare Presentation). Available online: http://www.slideshare.net/kolchinmax/kesw-2014-foo-dpedia-a-dbpedia-of-food-products (accessed on 4 September 2015).
- FOODpedia. Available online: http://foodpedia.tk/?lang=en (accessed on 4 September 2015).
- Evert, A.B.; Boucher, J.L.; Cypress, M.; Dunbar, S.A.; Franz, M.J.; Mayer-Davis, E.J.; Neumiller, J.J.; Nwankwo, R.; Verdi, C.L.; Urbanski, P.; et al. American Diabetes Association. Nutrition therapy recommendations for the management of adults with diabetes. Diabetes Care 2013, 36, 3821–3842. [Google Scholar] [CrossRef] [PubMed]
- Henry, C.J. Basal metabolic rate studies in humans: Measurements and development of new equations. Public Health Nutr. 2005, 8, 1133–1152. [Google Scholar] [CrossRef] [PubMed]
- Ainsworth, B.E. The Compendium of Physical Activities Tracking Guide. Prevention Research Center, Norman J. Arnold School of Public Health, University of South Carolina, 2002. Available online: http://prevention.sph.sc.edu/tools/docs/documents_compendium.pdf (accessed on 4 September 2015).
- Food and Nutrition Board, Institute of Medicine. Dietary Reference Intakes for Energy, Carbohydrate, Fiber, Fat, Fatty Acids, Cholesterol, Protein, and Amino Acids. 2002. Available online: http://www.nap.edu/openbook.php?isbn=0309085373 (accessed on 4 September 2015).
- Lin, E.-Y.; Yang, D.-L.; Hung, M.-C. System Design of an Intelligent Nutrition Consultation and Recommendation Model. In Proceedings of the 9th International Conference on Ubiquitous Intelligence and Computing and 9th International Conference on Autonomic and Trusted Computing, Fukuoka, Japan, 4–7 September 2012; pp. 740–745.
- Tumnark, P.; Oliveira, L.; Santibutr, N. Ontology-Based Personalized Dietary Recommendation for Weightlifting. In Proceedings of the International Workshop on Computer Science in Sports (IWCSS 2013), Wuhan, China, 1–2 August 2013.
- Chi, Y.L.; Chen, T.Y.; Tsai, W.T. A chronic disease dietary consultation system using OWL-based ontologies and semantic rules. J. Biomed. Inform. 2015, 53, 208–219. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).