Therapeutic pCRISPRi Delivery to Lung Squamous Cell Carcinoma by Combining Nanobubbles and Ultrasound
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Preparation of NBs
2.3. Cell Culture
2.4. Preparation of Tumor-Bearing Mice
2.5. US Imaging
2.6. pDNA
2.7. Luciferase Assay
2.8. Real-Time PCR
2.9. Terminal Deoxynucleotidyl Transferase-Mediated dUTP–Biotin Nick End Labeling (TUNEL) Staining
2.10. Animals
2.11. Statistical Analysis
3. Results
3.1. NBs Preparation and Characterization
3.2. Intratumoral Distribution of NBs via US Imaging
3.3. pDNA Delivery into Tumors via NBs and US Irradiation
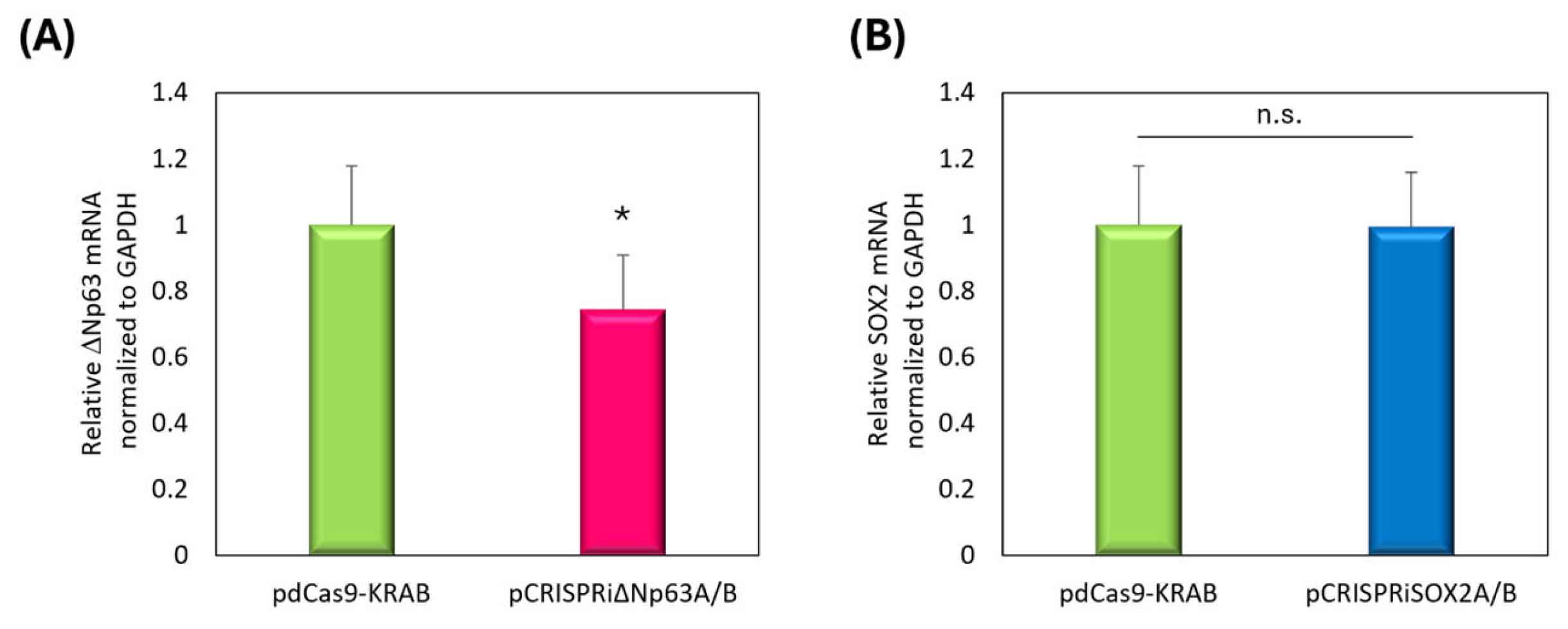
3.4. Evaluation of Target Gene Downregulation Using pCRISPRi Delivery via NBs and US
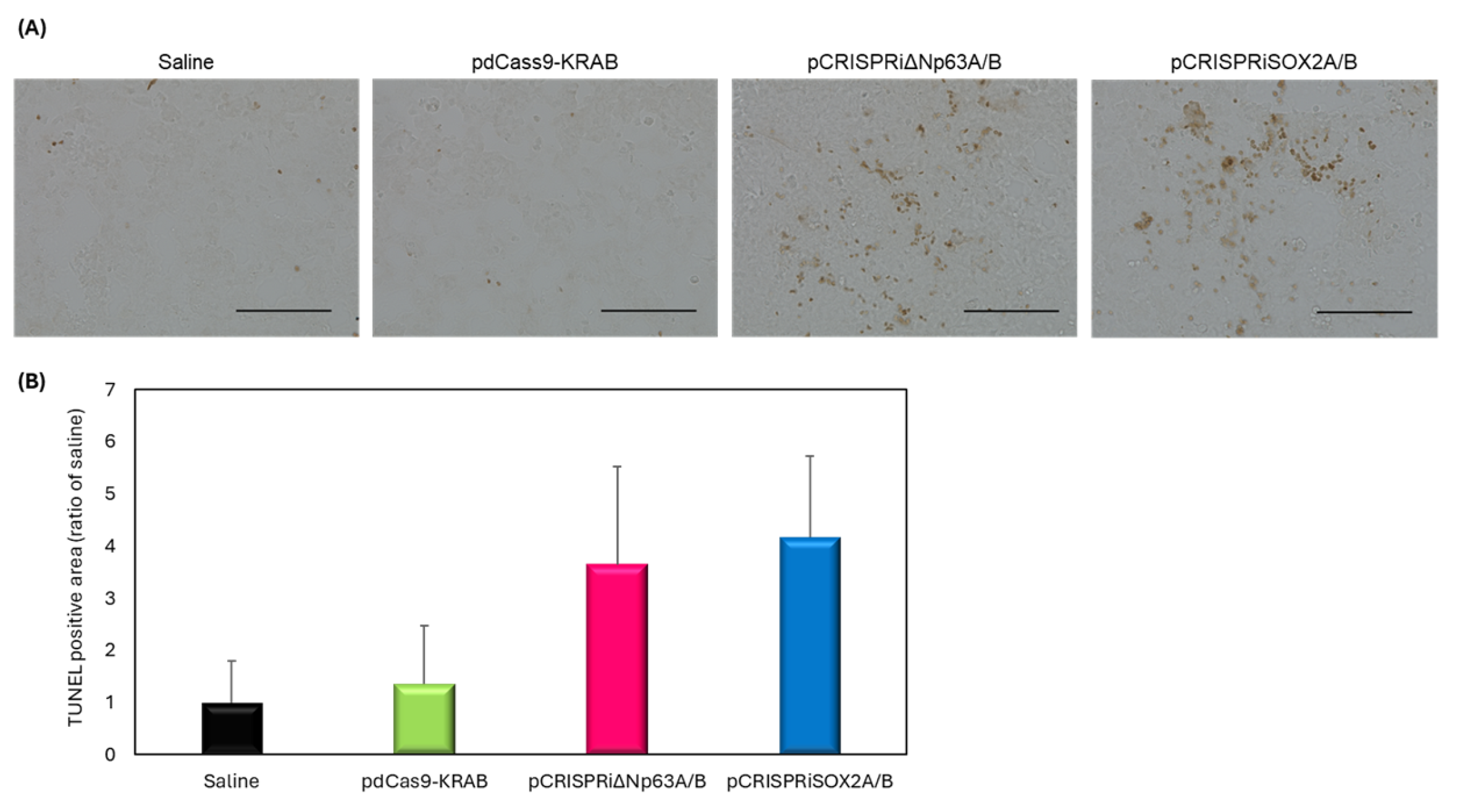
3.5. pCRISPRi-Induced Apoptosis
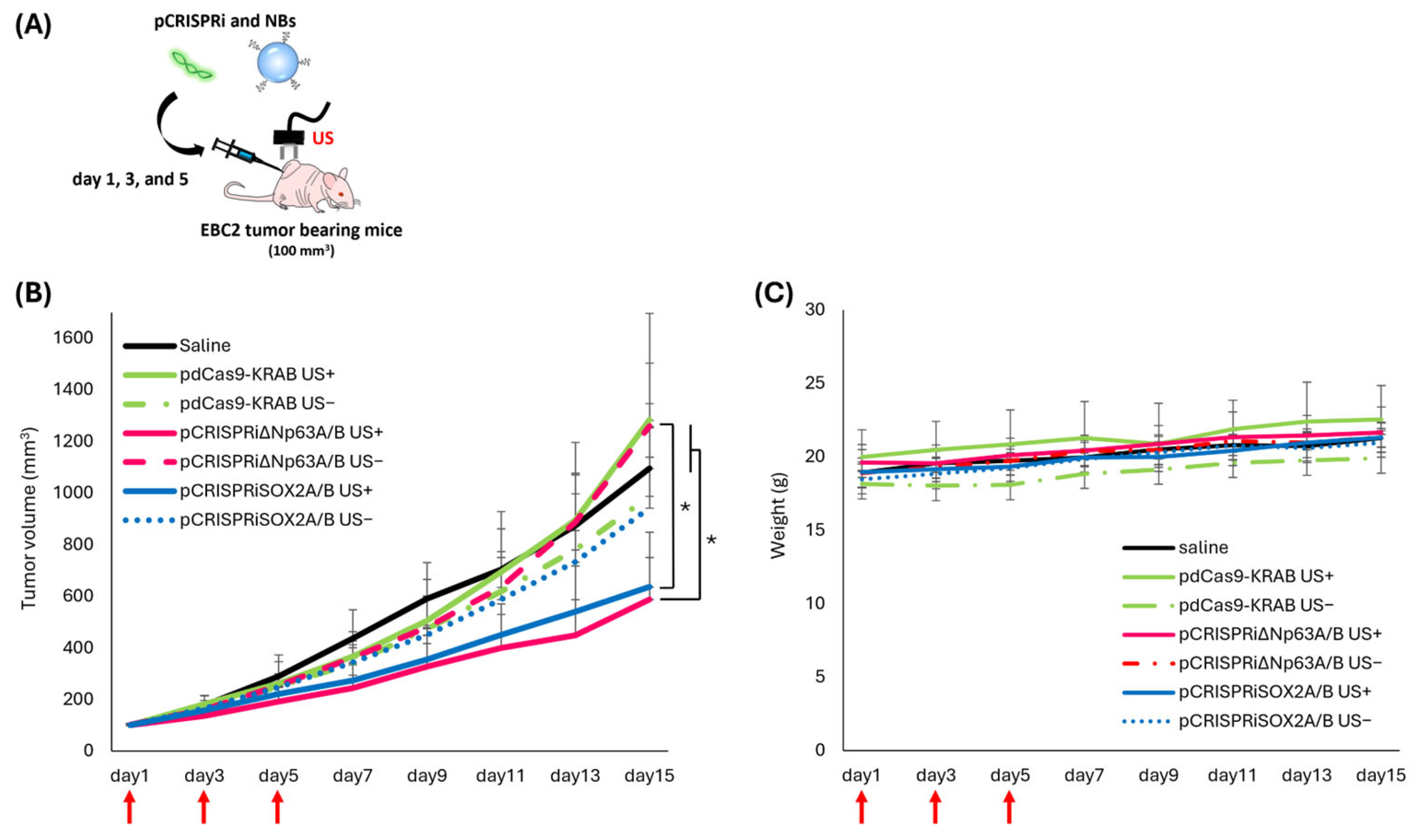
3.6. Antitumor Effect of pCRISPRi via NBs and US Irradiation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| SCC | squamous cell carcinoma |
| CRISPRi | CRISPR interference |
| NBs | nanobubbles |
| US | ultrasound |
References
- Zhang, Y.; Vaccarella, S.; Morgan, E.; Li, M.; Etxeberria, J.; Chokunonga, E.; Manraj, S.S.; Kamate, B.; Omonisi, A.; Bray, F. Global Variations in Lung Cancer Incidence by Histological Subtype in 2020: A Population-Based Study. Lancet Oncol. 2023, 24, 1206–1218. [Google Scholar] [CrossRef]
- Bray, F.; Laversanne, M.; Sung, H.; Ferlay, J.; Siegel, R.L.; Soerjomataram, I.; Jemal, A. Global Cancer Statistics 2022: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2024, 74, 229–263. [Google Scholar] [CrossRef] [PubMed]
- Hammerman, P.S.; Voet, D.; Lawrence, M.S.; Voet, D.; Jing, R.; Cibulskis, K.; Sivachenko, A.; Stojanov, P.; McKenna, A.; Lander, E.S.; et al. Comprehensive Genomic Characterization of Squamous Cell Lung Cancers. Nature 2012, 489, 519–525. [Google Scholar] [CrossRef]
- Araghi, M.; Mannani, R.; Heidarnejad Maleki, A.; Hamidi, A.; Rostami, S.; Safa, S.H.; Faramarzi, F.; Khorasani, S.; Alimohammadi, M.; Tahmasebi, S.; et al. Recent Advances in Non-Small Cell Lung Cancer Targeted Therapy; an Update Review. Cancer Cell Int. 2023, 23, 162. [Google Scholar] [CrossRef]
- Lau, S.C.M.; Pan, Y.; Velcheti, V.; Wong, K.K. Squamous Cell Lung Cancer: Current Landscape and Future Therapeutic Options. Cancer Cell 2022, 40, 1279–1293. [Google Scholar] [CrossRef]
- Chen, L.; Chen, F.; Li, J.; Pu, Y.; Yang, C.; Wang, Y.; Lei, Y.; Huang, Y. CAR-T Cell Therapy for Lung Cancer: Potential and Perspective. Thorac. Cancer 2022, 13, 889–899. [Google Scholar] [CrossRef]
- Qi, L.S.; Larson, M.H.; Gilbert, L.A.; Doudna, J.A.; Weissman, J.S.; Arkin, A.P.; Lim, W.A. Repurposing CRISPR as an RNA-Guided Platform for Sequence-Specific Control of Gene Expression. Cell 2013, 152, 1173–1183. [Google Scholar] [CrossRef]
- La Russa, M.F.; Qi, L.S. The New State of the Art: Cas9 for Gene Activation and Repression. Mol. Cell Biol. 2015, 35, 3800–3809. [Google Scholar] [CrossRef] [PubMed]
- Bendixen, L.; Jensen, T.I.; Bak, R.O. CRISPR-Cas-Mediated Transcriptional Modulation: The Therapeutic Promises of CRISPRa and CRISPRi. Mol. Ther. 2023, 31, 1920–1937. [Google Scholar] [CrossRef] [PubMed]
- Sakuma, T.; Nishikawa, A.; Kume, S.; Chayama, K.; Yamamoto, T. Multiplex Genome Engineering in Human Cells Using All-in-One CRISPR/Cas9 Vector System. Sci. Rep. 2014, 4, 5400. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, M.; Yokota, E.; Sakuma, T.; Yamatsuji, T.; Takigawa, N.; Ushijima, T.; Yamamoto, T.; Fukazawa, T.; Naomoto, Y. Development of an Integrated CRISPRi Targeting ΔNp63 for Treatment of Squamous Cell Carcinoma. Oncotarget 2018, 9, 29220–29232. [Google Scholar] [CrossRef]
- Liu, Q.; Zhao, K.; Wang, C.; Zhang, Z.; Zheng, C.; Zhao, Y.; Zheng, Y.; Liu, C.; An, Y.; Shi, L.; et al. Multistage Delivery Nanoparticle Facilitates Efficient CRISPR/DCas9 Activation and Tumor Growth Suppression In Vivo. Adv. Sci. 2019, 6, 1801423. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Chooi, W.H.; Liu, S.; Chin, J.S.; Murray, A.; Nizetic, D.; Cheng, D.; Chew, S.Y. Localized Delivery of CRISPR/DCas9 via Layer-by-Layer Self-Assembling Peptide Coating on Nanofibers for Neural Tissue Engineering. Biomaterials 2020, 256, 120225. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Zhang, L.; Tong, S.; Lee, C.M.; Deshmukh, H.; Bao, G. Spatial Control of in Vivo CRISPR–Cas9 Genome Editing via Nanomagnets. Nat. Biomed. Eng. 2019, 3, 126–136. [Google Scholar] [CrossRef] [PubMed]
- Pu, Y.; Yin, H.; Dong, C.; Xiang, H.; Wu, W.; Zhou, B.; Du, D.; Chen, Y.; Xu, H. Sono-Controllable and ROS-Sensitive CRISPR-Cas9 Genome Editing for Augmented/Synergistic Ultrasound Tumor Nanotherapy. Adv. Mater. 2021, 33, 2104641. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Li, F.; Lyu, Y.; Wang, F.; Lee, L.T.O.; He, S.; Guo, Z.; Li, J. A Semiconducting Polymer NanoCRISPR for Near-Infrared Photoactivatable Gene Editing and Cancer Gene Therapy. Nano Lett. 2025, 25, 4518–4525. [Google Scholar] [CrossRef]
- Moradi Kashkooli, F.; Jakhmola, A.; Hornsby, T.K.; Tavakkoli, J.; Kolios, M.C. Ultrasound-Mediated Nano Drug Delivery for Treating Cancer: Fundamental Physics to Future Directions. J. Control. Release 2023, 355, 552–578. [Google Scholar] [CrossRef]
- Unger, E.C.; Hersh, E.; Vannan, M.; McCreery, T. Gene Delivery Using Ultrasound Contrast Agents. Echocardiography 2001, 18, 355–361. [Google Scholar] [CrossRef]
- Hernot, S.; Klibanov, A.L. Microbubbles in Ultrasound-Triggered Drug and Gene Delivery. Adv. Drug Deliv. Rev. 2008, 60, 1153–1166. [Google Scholar] [CrossRef]
- Kwak, G.; Grewal, A.; Slika, H.; Mess, G.; Li, H.; Kwatra, M.; Poulopoulos, A.; Woodworth, G.F.; Eberhart, C.G.; Ko, H.S.; et al. Brain Nucleic Acid Delivery and Genome Editing via Focused Ultrasound-Mediated Blood-Brain Barrier Opening and Long-Circulating Nanoparticles. ACS Nano 2024, 18, 24139–24153. [Google Scholar] [CrossRef]
- He, S.; Singh, D.; Helfield, B. Flow Rate Modulates Focused Ultrasound-Mediated Vascular Delivery of MicroRNA. Mol. Ther. Nucleic Acids 2025, 36, 102426. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wu, P.; Zhu, M.; Liang, M.; Zhang, L.; Zong, Y.; Wan, M. High-Performance Delivery of a CRISPR Interference System via Lipid-Polymer Hybrid Nanoparticles Combined with Ultrasound-Mediated Microbubble Destruction for Tumor-Specific Gene Repression. Adv. Healthc. Mater. 2023, 12, 2203082. [Google Scholar] [CrossRef] [PubMed]
- Kong, W.; Li, X.; Guo, X.; Sun, Y.; Chai, W.; Chang, Y.; Huang, Q.; Wang, P.; Wang, X. Ultrasound-Assisted CRISPRi-Exosome for Epigenetic Modification of α-Synuclein Gene in a Mouse Model of Parkinson’s Disease. ACS Nano 2024, 18, 7837–7851. [Google Scholar] [CrossRef] [PubMed]
- Hazel, K.; Singh, D.; He, S.; Guertin, Z.; Husser, M.C.; Helfield, B. Focused Ultrasound and Microbubble-Mediated Delivery of CRISPR-Cas9 Ribonucleoprotein to Human Induced Pluripotent Stem Cells. Mol. Ther. 2025, 33, 986–996. [Google Scholar] [CrossRef]
- Sheth, M.; Knight, C.; Wu, Q.; Vasilyeva, A.; Upadhyay, A.; Bau, L.; Ruan, J.-L.; Ovenden, N.; Stride, E. Size Matters: Micro- versus Nanobubbles in Ultrasound Imaging and Therapy. Sci. Adv. 2025, 11, 2177. [Google Scholar] [CrossRef]
- Pellow, C.; Pellow, C.; Pellow, C.; O’Reilly, M.A.; O’Reilly, M.A.; Hynynen, K.; Hynynen, K.; Zheng, G.; Zheng, G.; Goertz, D.E.; et al. Simultaneous Intravital Optical and Acoustic Monitoring of Ultrasound-Triggered Nanobubble Generation and Extravasation. Nano Lett. 2020, 20, 4512–4519. [Google Scholar] [CrossRef]
- Helfield, B.L.; Yoo, K.; Liu, J.; Williams, R.; Sheeran, P.S.; Goertz, D.E.; Burns, P.N. Investigating the Accumulation of Submicron Phase-Change Droplets in Tumors. Ultrasound Med. Biol. 2020, 46, 2861–2870. [Google Scholar] [CrossRef]
- Suzuki, R.; Takizawa, T.; Negishi, Y.; Hagisawa, K.; Tanaka, K.; Sawamura, K.; Utoguchi, N.; Nishioka, T.; Maruyama, K. Gene Delivery by Combination of Novel Liposomal Bubbles with Perfluoropropane and Ultrasound. J. Control. Release 2007, 117, 130–136. [Google Scholar] [CrossRef]
- Negishi, Y.; Endo, Y.; Fukuyama, T.; Suzuki, R.; Takizawa, T.; Omata, D.; Maruyama, K.; Aramaki, Y. Delivery of SiRNA into the Cytoplasm by Liposomal Bubbles and Ultrasound. J. Control. Release 2008, 132, 124–130. [Google Scholar] [CrossRef]
- Negishi, Y.; Tsunoda, Y.; Endo-Takahashi, Y.; Oda, Y.; Suzuki, R.; Maruyama, K.; Yamamoto, M.; Aramaki, Y. Local Gene Delivery System by Bubble Liposomes and Ultrasound Exposure into Joint Synovium. J. Drug Deliv. 2011, 2011, 203986. [Google Scholar] [CrossRef]
- Negishi, Y.; Matsuo, K.; Endo-Takahashi, Y.; Suzuki, K.; Matsuki, Y.; Takagi, N.; Suzuki, R.; Maruyama, K.; Aramaki, Y. Delivery of an Angiogenic Gene into Ischemic Muscle by Novel Bubble Liposomes Followed by Ultrasound Exposure. Pharm. Res. 2011, 28, 712–719. [Google Scholar] [CrossRef]
- Sugano, M.; Negishi, Y.; Endo-Takahashi, Y.; Suzuki, R.; Maruyama, K.; Yamamoto, M.; Aramaki, Y. Gene Delivery System Involving Bubble Liposomes and Ultrasound for the Efficient in Vivo Delivery of Genes into Mouse Tongue Tissue. Int. J. Pharm. 2012, 422, 332–337. [Google Scholar] [CrossRef]
- Suzuki, R.; Namai, E.; Oda, Y.; Nishiie, N.; Otake, S.; Koshima, R.; Hirata, K.; Taira, Y.; Utoguchi, N.; Negishi, Y.; et al. Cancer Gene Therapy by IL-12 Gene Delivery Using Liposomal Bubbles and Tumoral Ultrasound Exposure. J. Control. Release 2010, 142, 245–250. [Google Scholar] [CrossRef] [PubMed]
- Fukazawa, T.; Guo, M.; Ishida, N.; Yamatsuji, T.; Takaoka, M.; Yokota, E.; Haisa, M.; Miyake, N.; Ikeda, T.; Okui, T.; et al. SOX2 Suppresses CDKN1A to Sustain Growth of Lung Squamous Cell Carcinoma. Sci. Rep. 2016, 6, 20113. [Google Scholar] [CrossRef] [PubMed]
- Yokota, E.; Yamatsuji, T.; Takaoka, M.; Haisa, M.; Takigawa, N.; Miyake, N.; Ikeda, T.; Mori, T.; Ohno, S.; Sera, T.; et al. Targeted Silencing of SOX2 by an Artificial Transcription Factor Showed Antitumor Effect in Lung and Esophageal Squamous Cell Carcinoma. Oncotarget 2017, 8, 103063–103076. [Google Scholar] [CrossRef] [PubMed]
- Yu, V.Z.; So, S.S.; Lung, B.C.C.; Hou, G.Z.; Wong, C.W.y.; Chow, L.K.y.; Chung, M.K.y.; Wong, I.Y.h.; Wong, C.L.y.; Chan, D.K.k.; et al. ΔNp63-Restricted Viral Mimicry Response Impedes Cancer Cell Viability and Remodels Tumor Microenvironment in Esophageal Squamous Cell Carcinoma. Cancer Lett. 2024, 595, 216999. [Google Scholar] [CrossRef]
- Endo-Takahashi, Y.; Sakurai, A.; Oguri, Y.; Katagiri, F.; Akiyama, S.; Sashida, S.; Yamaguchi, T.; Marunouchi, T.; Suzuki, R.; Maruyama, K.; et al. Phosphorodiamidate Morpholino Oligomers-Loaded Nanobubbles for Ultrasound-Mediated Delivery to the Myocardium in Muscular Dystrophy. ACS Omega 2025, 10, 9639–9648. [Google Scholar] [CrossRef]
- Yamaguchi, T.; Endo-Takahashi, Y.; Awaji, K.; Numazawa, S.; Onishi, Y.; Tada, R.; Ogasawara, M.; Takizawa, Y.; Kurumizaka, H.; Negishi, Y. Microfluidic Nanobubbles Produced Using a Micromixer for Ultrasound Imaging and Gene Delivery. Sci. Rep. 2025, 15, 14871. [Google Scholar] [CrossRef]
- Nakagawa, Y.; Sakuma, T.; Sakamoto, T.; Ohmuraya, M.; Nakagata, N.; Yamamoto, T. Production of Knockout Mice by DNA Microinjection of Various CRISPR/Cas9 Vectors into Freeze-Thawed Fertilized Oocytes. BMC Biotechnol. 2015, 15, 33. [Google Scholar] [CrossRef]
- Gilbert, L.A.; Horlbeck, M.A.; Adamson, B.; Villalta, J.E.; Chen, Y.; Whitehead, E.H.; Guimaraes, C.; Panning, B.; Ploegh, H.L.; Bassik, M.C.; et al. Genome-Scale CRISPR-Mediated Control of Gene Repression and Activation. Cell 2014, 159, 647–661. [Google Scholar] [CrossRef]
- Wang, J.P.; Zhou, X.L.; Yan, J.P.; Zheng, R.Q.; Wang, W. Nanobubbles as Ultrasound Contrast Agent for Facilitating Small Cell Lung Cancer Imaging. Oncotarget 2017, 8, 78153. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Hu, Z.; Wang, C.; Yang, J.; Zeng, C.; Fan, R.; Guo, J. PD-L1-Targeted Microbubbles Loaded with Docetaxel Produce a Synergistic Effect for the Treatment of Lung Cancer under Ultrasound Irradiation. Biomater. Sci. 2020, 8, 1418–1430. [Google Scholar] [CrossRef] [PubMed]
- Negishi, Y.; Endo-Takahashi, Y.; Matsuki, Y.; Kato, Y.; Takagi, N.; Suzuki, R.; Maruyama, K.; Aramaki, Y. Systemic Delivery Systems of Angiogenic Gene by Novel Bubble Liposomes Containing Cationic Lipid and Ultrasound Exposure. Mol. Pharm. 2012, 9, 1834–1840. [Google Scholar] [CrossRef] [PubMed]
- Endo-Takahashi, Y.; Negishi, Y.; Nakamura, A.; Suzuki, D.; Ukai, S.; Sugimoto, K.; Moriyasu, F.; Takagi, N.; Suzuki, R.; Maruyama, K.; et al. PDNA-Loaded Bubble Liposomes as Potential Ultrasound Imaging and Gene Delivery Agents. Biomaterials 2013, 34, 2807–2813. [Google Scholar] [CrossRef]
- Endo-Takahashi, Y.; Negishi, Y.; Kato, Y.; Suzuki, R.; Maruyama, K.; Aramaki, Y. Efficient SiRNA Delivery Using Novel SiRNA-Loaded Bubble Liposomes and Ultrasound. Int. J. Pharm. 2012, 422, 504–509. [Google Scholar] [CrossRef]
- Endo-Takahashi, Y.; Negishi, Y.; Nakamura, A.; Ukai, S.; Ooaku, K.; Oda, Y.; Sugimoto, K.; Moriyasu, F.; Takagi, N.; Suzuki, R.; et al. Systemic Delivery of MiR-126 by MiRNA-Loaded Bubble Liposomes for the Treatment of Hindlimb Ischemia. Sci. Rep. 2014, 4, 3883. [Google Scholar] [CrossRef]
- Safai Zadeh, E.; Görg, C.; Dietrich, C.F.; Görlach, J.; Alhyari, A.; Trenker, C. Contrast-Enhanced Ultrasound for Evaluation of Pleural Effusion: A Pictorial Essay. J. Ultrasound Med. 2022, 41, 485–503. [Google Scholar] [CrossRef]
- Sugiyama, M.G.; Mintsopoulos, V.; Raheel, H.; Goldenberg, N.M.; Batt, J.E.; Brochard, L.; Kuebler, W.M.; Leong-Poi, H.; Karshafian, R.; Lee, W.L. Lung Ultrasound and Microbubbles Enhance Aminoglycoside Efficacy and Delivery to the Lung in Escherichia coli –Induced Pneumonia and Acute Respiratory Distress Syndrome. Am. J. Respir. Crit. Care Med. 2018, 198, 404–408. [Google Scholar] [CrossRef]
- Pioche, M.; Lafon, C.; Constanciel, E.; Vignot, A.; Birer, A.; Gincul, R.; Lépilliez, V.; Prat, F.; Roman, S.; Chapelon, J.Y.; et al. High-Intensity Focused Ultrasound Liver Destruction through the Gastric Wall under Endoscopic Ultrasound Control: First Experience in Living Pigs. Endoscopy 2012, 44, E376–E377. [Google Scholar] [CrossRef]
- Ashida, R.; Kawabata, K.I.; Asami, R.; Kitano, M. Novel Treatment System Using Endoscopic Ultrasound-Guided High-Intensity Focused Ultrasound: A Proof-of-Concept Study. Pancreatology 2024, 24, 88–92. [Google Scholar] [CrossRef]
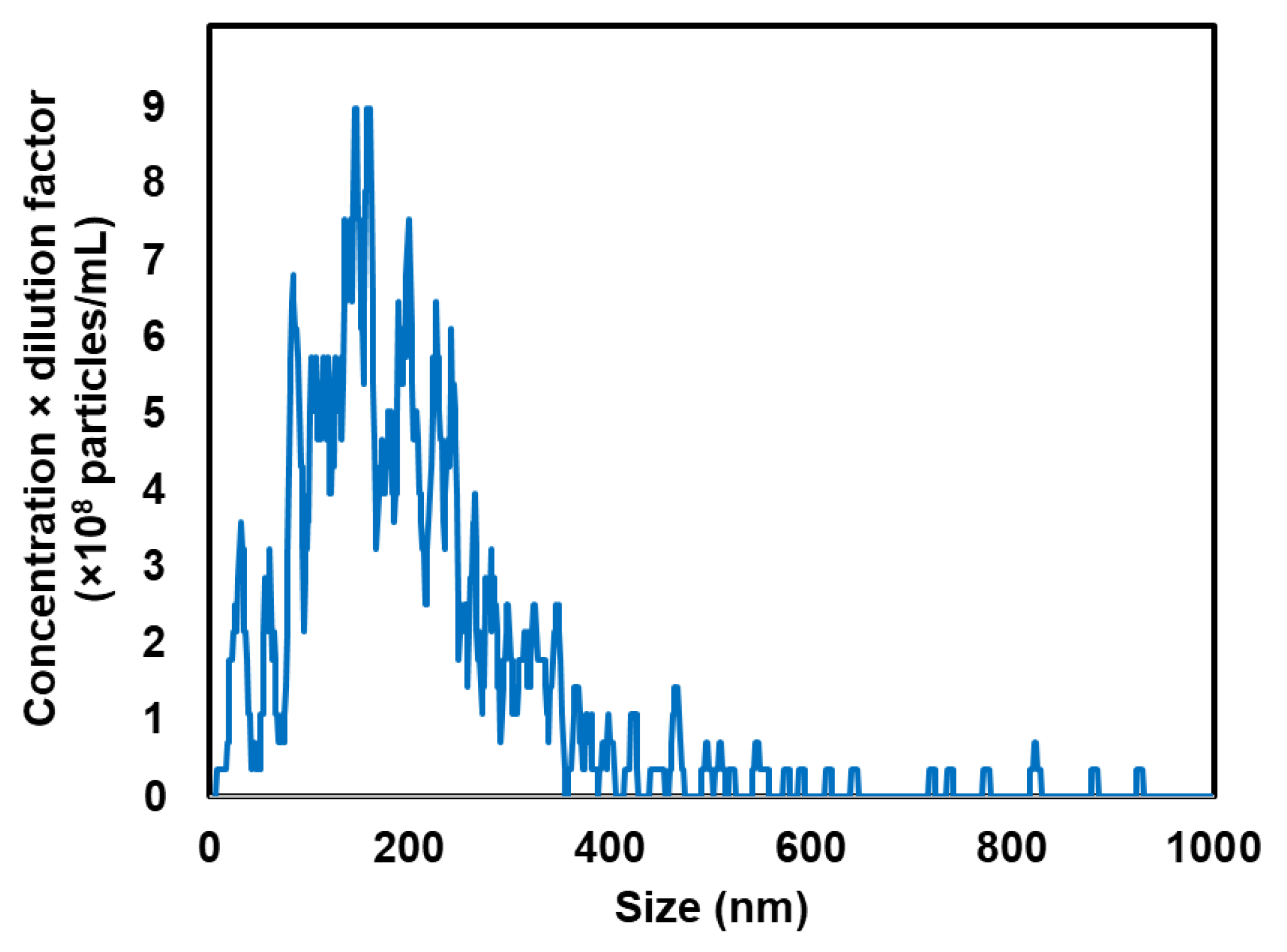


| Mean Size (nm) | Span (nm) | Concentration (×1011 Particles/mL) | Zeta Potential (mV) |
|---|---|---|---|
| 198.0 ± 17.9 | 1.47 ± 0.09 | 1.65 ± 0.38 | −20.0 ± 0.41 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yamaguchi, T.; Endo-Takahashi, Y.; Amano, T.; Ihara, A.; Sakuma, T.; Yamamoto, T.; Fukazawa, T.; Negishi, Y. Therapeutic pCRISPRi Delivery to Lung Squamous Cell Carcinoma by Combining Nanobubbles and Ultrasound. Pharmaceutics 2025, 17, 1053. https://doi.org/10.3390/pharmaceutics17081053
Yamaguchi T, Endo-Takahashi Y, Amano T, Ihara A, Sakuma T, Yamamoto T, Fukazawa T, Negishi Y. Therapeutic pCRISPRi Delivery to Lung Squamous Cell Carcinoma by Combining Nanobubbles and Ultrasound. Pharmaceutics. 2025; 17(8):1053. https://doi.org/10.3390/pharmaceutics17081053
Chicago/Turabian StyleYamaguchi, Taiki, Yoko Endo-Takahashi, Takumi Amano, Arina Ihara, Tetsushi Sakuma, Takashi Yamamoto, Takuya Fukazawa, and Yoichi Negishi. 2025. "Therapeutic pCRISPRi Delivery to Lung Squamous Cell Carcinoma by Combining Nanobubbles and Ultrasound" Pharmaceutics 17, no. 8: 1053. https://doi.org/10.3390/pharmaceutics17081053
APA StyleYamaguchi, T., Endo-Takahashi, Y., Amano, T., Ihara, A., Sakuma, T., Yamamoto, T., Fukazawa, T., & Negishi, Y. (2025). Therapeutic pCRISPRi Delivery to Lung Squamous Cell Carcinoma by Combining Nanobubbles and Ultrasound. Pharmaceutics, 17(8), 1053. https://doi.org/10.3390/pharmaceutics17081053







