Small Interfering RNA Carriers for Oncotherapy: A Preclinical Overview
Abstract
1. Introduction
2. Biological Mechanism of siRNA
3. Fundamental Principles for siRNA Design
4. siRNA Modifications and Delivery
4.1. Chemical Modifications to siRNA Strands
- Modifications to the rest of the ribose. Ribose modifications, particularly at the 2′ position, have been the most widely used to protect siRNA from attack by ribonucleases, improve affinity for the target mRNA, and decrease the immune response. They have the advantage of not altering backbone conformation or gene silencing efficiency. The most common is the substitution of the 2′-OH group with a less nucleophilic group. The most used group is 2′-O-methyl (2′-O-Me), from which derivatives such as 2′-O-methoxyethyl (2′-O-MOE) and 2′-deoxy-2′-fluoro (2′-F) have been developed [32,33,34]. Substitutions can be made at different sites on the double strand or in the central part of the antisense strand [35]. The best results are obtained by alternating 2′-O-Me and 2′-F substitutions [36,37].
- Modifications to the ribose-phosphate backbone. These are designed to minimize off-target effects and immune responses, enhance cell uptake, and increase the bioavailability of siRNA. The most used is the replacement of a non-bridging oxygen in the phosphodiester group linking two consecutive riboses with a sulfur to form a phosphorothioate (PS) [36,49,50,51], a boron to form a borane-phosphate (PB) [52,53], or an acetate to form a phosphonoacetate [54]. These modifications are used to link the siRNA to different molecules [55]. The most prevalent is the formation of PS, where the sulfur atom preserves the negative charge of the siRNA. The principal benefits of this modification are the improvement of siRNA resistance to nucleases, hydrophobicity, stability, and affinity to plasma proteins, resulting in a longer circulation time [56,57]. However, the number of PS should be limited. This can reduce the silencing effect and induce cytotoxicity [49,58,59]. The best PS substitution variant is at the end of the strands. Some authors have synthesized siRNAs with more than one simultaneous modification, with attractive properties for in vivo use [60]. Alternatives with different phosphate derivatives include phosphorodithioate, PS2 [50,61,62], methylphosphonate [58], 5′-(E)-vinyl-phosphonate [63,64,65]. The phosphotriester groups have also been modified [66].
- Modifications to nucleobases. To a lesser extent than ribose and ribose-phosphate backbone modifications, modifications to uridine, cytidine, and adenosine are made to improve thermal stability, nuclease resistance, cell uptake, and reducing immune response [67,68,69,70]. However, concerns exist about the safety of metabolizing modified siRNAs via this route. The modifications described are used to obtain siRNAs with different structures, conjugates, and nanosystems for transport and delivery. There are numerous recent publications that summarize these modifications and their impact on siRNA properties [9,11,39,71]. Table 2 summarizes the advantages and disadvantages of the modifications described, as well as their applications. A common disadvantage is that, depending on the site and extent of the modification, the physicochemical and biological properties of the siRNA may deteriorate.
4.2. siRNA Transporters and Delivery Nanosystems
4.2.1. siRNA Conjugates with Biomolecules and Small Molecules
- (i)
- siRNA Conjugates with Biomolecules
- Ab-siRNA and FAb-siRNA conjugates. Abs are ideal carrier systems due to their high affinity and specificity, long half-life in blood, and relatively low immunogenicity. Experience gained with antibody-drug conjugates (ADCs) has enabled the development of antibody-siRNA conjugates (ARCs), which combine the precision of siRNA for target gene silencing with the high sensitivity and specificity of Antibodies for binding to their antigens/receptors on the cell surface. They are an advantageous option for targeting siRNA to antigen-expressing tumor cells because they accumulate and internalize more readily than naked siRNA [76,79,80].
- Peptide-siRNA conjugates. The small size of peptides, their relatively low molecular weight, ease of synthesis, and low cost of production have facilitated the development of peptide-siRNA conjugates (Figure 3C) with lower immunogenicity and toxicity, better pharmacokinetic properties, high cellular uptake, and more efficient endocytosis than Ab-siRNA conjugates [102,103]. Amino acids (AA) with acidic, hydrophilic, hydrophobic, or aromatic residues can be combined to generate several peptides with siRNA delivery potential, called cell-penetrating peptides (CPP) or membrane transduction peptides (MTP). These peptides are generally composed of <30 AA, often including lysine, histidine, and arginine. They readily cross the anionic surface of cell membranes and reach intracellular compartments without interacting with receptors or altering their functions [102,104,105]. They can be chemically modified to enhance endosomal escape and decrease the influence of endocytic proteases [105,106]. They have been used to internalize, through passive diffusion or endocytosis, various types of macromolecules, including siRNA. Some are internalized by both mechanisms, depending on factors such as the AA sequence and its structure, the CPP/siRNA concentration ratio, cell line and others [102,107,108].
- Aptamer-siRNA conjugates: Aptamers are single-stranded DNA or RNA molecules known for their high affinity and specificity for cell receptors overexpressed in tumors. Their large-scale production and low cost have driven the development of aptamer-siRNA conjugates (Figure 3D), also known as chimeras, as alternatives to Ab-siRNA and peptide-siRNA conjugates [153,154]. Unlike antibodies and peptides, aptamers maintain their biological activity when linked with siRNA. Since aptamers are smaller (6–30 kDa) compared to antibodies (150 kDa), their conjugates with siRNA are more easily internalized, and their endosomal escape is more efficient [155,156,157].
- Toxins-siRNA conjugates. Toxins are poisonous substances produced by living organisms such as plants, animals, and microorganisms. The AB-type toxin has a domain that can be engineered to reduce (attenuate) toxicity, a translocation domain that enables endosomal escape, and a receptor domain that induces endocytosis mediated by cell receptors [173]. Diphtheria and anthrax are examples of AB-type toxins, which have been used to create conjugates for delivering siRNA in vitro to cancer cells [173,174]. siRNA conjugates with attenuated AB-toxins become a promising delivery system because they address some of the challenges of siRNA delivery, especially endosomal escape, and improve upon some disadvantages of polycationic delivery systems. Although these engineered modified toxins have been used for protein delivery [175], few studies have reported on siRNA delivery. This field is likely to grow in the future as methods for modifying toxins and conjugating siRNA continue to advance.
- (ii)
- siRNA Conjugates with Small Molecules
- FA-siRNA conjugates. FA is a crucial component for cell growth and proliferation, serving as a transporter for several therapeutic agents. It is especially useful in oncology because FA receptors (FR), particularly FRα and FRβ, are overexpressed in many tumor cells, while they are expressed at very low or no levels in normal tissues [179,180]. The uptake of FA via FR is specific and exhibits high affinity (KD 10−9 M) [181]. The small size of FA-siRNA conjugates allows them to easily reach solid tumors [179,180]. These conjugates are produced by direct covalent conjugation or by hybridization [177]. However, direct conjugation has limitations. Since folate conjugates bind to FRs through the pteridine residue, conjugation must occur through the glutamate residue, which, having two carboxyl groups, results in mixtures of α and γ isomers [182]. Although folate can be incorporated into the 3′ or 5′ ends of the sense strand of siRNA, many researchers prefer central positions [179]. Common covalent conjugation methods include: (i) linking siRNA to a bifunctional amide thiol linker followed by attachment of folate [181]; (ii) connecting the 5′-end of an oligodeoxynucleotide (ODN) to a folate molecule (ODN-FA), then extending the sense strand of the siRNA to couple it with the ODN-FA adduct [183]; and azide-alkyne reactions with various variants [177,184]. The same reactions are used for conjugation via hybridization [183].
- Cholesterol-siRNA conjugates were developed to increase hydrophobicity and improve siRNA cell uptake [185,186,187]. They are created through direct covalent bonding or hybridization. The most commonly used reactions are thiol-maleimide and azide-alkyne. For the linkage, the 3′ or 5′ ends of the sense strand are modified, followed by adding cholesterol bound to a linker [188,189,190]. The common reaction methods include: (i) attaching the cholesterol connector to a terminal -OH group of the sense strand of the siRNA [188]; (ii) performing PS modifications at the 3′-end of both strands, adding modified nucleotides (2′-OMe) to the antisense strand, and attaching the cholesterol linker to the sense strand [188,190]; (iii) similar to the previous method but alternating modifications with 2′-OMe and 2′F, then attaching nucleotides to the antisense strand [191]; (iv) truncating one or both strands to produce short asymmetric siRNAs, then adding nucleotides with PS modifications to the antisense strand and attaching the linker-cholesterol adduct to the sense strand [189]. These variants enhance thermodynamic stability, pharmacokinetics, internalization, and silencing efficiency. The linker’s nature and the siRNA binding site significantly influence the biological properties of the conjugates [185,189,192]. The most mentioned connectors include triethylene glycol (TEG) [190,192], 2-amino butyl-1-3-propanediol (C7) [188], trans-4-hydroxyprolinol [78,190], and hexamethylenediamine [189].
- Fatty acid conjugates with siRNA. siRNA has been conjugated to fatty acids such as docosanoic acid (DCA, C22H44O2), docosahexaenoic acid (DHA, C22H32O2), lithocholic acid (LCA, C24H40O3), palmitic acid (C16H32O2), among others [198,199]. These conjugates are obtained by methods similar to those described for cholesterol-siRNA conjugates. Their physicochemical properties, biodistribution, and silencing capacity depend on the specific characteristics of each fatty acid [198,199,200,201,202]. However, their direct use is also limited. For this reason, it is preferred to associate them with lipid nanosystems.
- Calcium-siRNA complexes. The spontaneous formation of calcium ion with siRNA produces nanocomplexes that are stable, have a uniform size (~100 nm), and carry a negative surface charge (−8 mV). The reversible nature of the electrostatic interactions between Ca2+ and siRNA has been effectively used for in vitro siRNA transfection [203]. These complexes serve as the foundation for developing highly efficient siRNA nanocarrier systems [204].
- Anisamide-siRNA conjugates. One and two receptors are polypeptide chains located in the endoplasmic reticulum and as transmembrane proteins in nervous system cells. They have neuromodulatory and ion-channeling functions and bind to a wide range of psychoactive drugs [205]. These receptors are also expressed in several tumor cell types [205] and exhibit high affinity for anisamide (2-(4′-methoxy benzamido) ethyleneamide) and its derivatives [206]. Anisamide conjugates with drugs demonstrate high internalization efficiency via endocytosis mediated by these receptors [207,208], which has driven the development of mono- and multivalent anisamide-siRNA conjugates. Monovalent conjugates are formed by direct covalent bonding between an anisamide phosphoramidate and the 5′ end of the sense strand of the siRNA, which has been previously modified with 2′O-Me. For multivalent conjugation, multifunctional linkers are used to first bind to the siRNA and then to the modified anisamide. These conjugates show a high transfection capacity in prostate cancer cells [206]. However, the use of anisamide-siRNA conjugates remains limited, likely due to concerns about potential side effects. Some have been incorporated into nanocarrier systems.
- N-acetyl galactosamine-siRNA conjugates. These are designed to silence specific mRNAs of liver proteins by binding to asialoglycoprotein receptors ASGP-R [209,210]. These conjugates consist of three molecules of N-acetyl galactosamine (GaINaC) linked via a spacer to the siRNA. By using different spacers and modifications of the siRNA, a wide variety of these conjugates have been developed, and they are incorporated into nanocarrier systems. Some of these formulations have received approval for treating various diseases [8].
4.2.2. siRNA Nanocarrier Systems
- (i)
- Organic Nanocarriers
- (a)
- Polymeric Nanosystems
- PEI-siRNA polyplexes. PEI is a widely used cationic polymer which forms polyplexes with siRNA because of its high cation density and ability to buffer protons across a broad pH range [238,239]. Its structure includes repetitive ethylenimine groups, which confer extensive buffering capacity within the pH range of the endosomal/lysosomal pathway [240,241]. Chemically, it is highly versatile, easily functionalized and branched [136,240,242]. The transfection efficiency of PEI-siRNA polyplexes is high because, under acidic conditions, their amines facilitate endosomal escape, helping siRNA reach the cytoplasm [136,241]. Branched PEI generally outperforms linear PEI in transfection efficiency [241,243,244]. Additionally, some PEIs can form covalent complexes with siRNA [241,245].
- CS-siRNA polyplexes. Chitosan (CS) is the most widely used natural polymer for preparing polyplexes that directly deliver siRNA to cells or form part of other nanosystems. It is a linear polysaccharide derived by deacetylation of chitin, composed of repeated units of N-acetyl-D-glucosamine and D-glucosamine linked via β-1,4 bonds. The proportion of these units determines the polymer’s degree of deacetylation (DD) [141,227]. CS is biocompatible, biodegradable, non-toxic, and easily modified [227,249,250]. siRNA can be incorporated into CS through: (i) encapsulation; (ii) adsorption; or (iii) electrostatic interactions [249,251]. The most common method is electrostatic complexation, as the high positive charge of CS facilitates binding with the negatively charged phosphate groups of siRNA [137,227,250].
- PBAE-siRNA polyplexes involve polymers prepared by polymerizing diacrylate and amino compounds. These polymers were designed to enhance biodegradability and reduce cytotoxicity compared to PEI and PLL [262,263,264]. They come in diverse structures and shapes, such as linear, spherical, and multi-layered films [231,265]. Their chemical modifications allow for control over size, surface charge, hydrophobicity, degradability, and stimulus responsiveness, influenced by factors like chain length, structure, end groups, solution pH, and N/P ratio [12,231,262,266]. Typically, these polyplexes are serum-stable and readily taken up by cells [12,262]. Inside the cell, they quickly degrade, releasing siRNA and PBAE is hydrolyzed into biocompatible products [12,231]. Their transfection efficiency is influenced by terminal groups, while toxicity relates to hydrophobicity levels and the spacing of amino groups [267]. Adding groups like amines or hydroxyls to the PBAE end chains notably enhances transfection, and the alkyl chain length and end-group hydrophobicity directly impact toxicity [264].
- (b)
- Lipid Nanosystems
- (c)
- Lipoprotein-Based Nanosystems
- (d)
- Cell-Derived Nanosystems
- (ii)
- Inorganic and Hybrid Nanocarriers
- Au nanoparticles. These are colloidal suspensions of Au with unique chemical and physicochemical properties that have been used in biomedicine since ancient times. They absorb light and X-rays, and their absorption maxima can be adjusted during synthesis. They can disperse and convert absorbed light into heat, enhance the Raman spectra of molecules near their surface, produce fluorescence, and carry different molecules with high loading capacity. They are bioinert, low in toxicity, and easy to obtain [406,407,408]. They easily conjugate to biomolecules due to their high affinity for thiols, disulfides, and amine groups [384,409,410]. They efficiently transfect siRNA without needing additional transfection agents.
- Magnetic nanoparticles are used to develop NMR theranostic systems and for gene silencing. They consist of materials containing ferromagnetic, paramagnetic, diamagnetic, antiferromagnetic, and ferrimagnetic elements, with sizes ranging from 10 to 20 nm. These particles exhibit different responses to external magnetic fields [426,427]. Superparamagnetic iron oxide nanoparticles (Fe3O4 and γ-Fe2O3), known as SPIONs, are the most commonly used in medicine due to their versatility, biocompatibility, strong magnetization, and low production costs [428,429,430].
- Inorganic semiconductor crystals, known as quantum dots (QDs). They are nanocrystals ranging from 2 to 20 nm that are created from binary combinations such as CdSe, CdTe, CdS, ZnS, ZnHgSe, PbS, CdHgTe, and CdxPb1-xTe alloys. Their optical properties vary with changes in composition, size, and shape [439]. These QDs offer several advantages: (i) tunable emission; (ii) high fluorescence quantum yield; (iii) resistance to photobleaching; (iv) a high surface-to-volume ratio; and (v) ease of functionalization, which facilitates the transport of molecules of diverse types [440]).
- Carbon-based nanoparticles leverage the chemical properties of carbon to create carrier NPs with sizes akin to many biological structures. Hybridization of sp2 carbon enables the production of various graphitic materials—such as nanodiamonds (3D), graphene (2D), nanotubes (1D), fullerenes, and quantum dots (0D)—which are used to fabricate NPs with diverse structures. These materials offer exceptional mechanical strength, high stability, a large surface area-to-volume ratio, unique optical features, excellent thermal and caloric conductivity, biocompatibility, ease of functionalization, and antibacterial properties [454,455]. For siRNA coupling, the primary options include: (i) carbon nanotubes, (ii) graphene oxide nanosheets, (iii) fullerenes, and (iv) carbon quantum dots.
- Mesoporous silica nanoparticles (MSNP). These are silica nanoparticles with various morphologies and numerous pores. Their structure, shape, nanoparticle size (50–200 nm), and pore size (2–50 nm) can vary depending on the production method [485,486,487]. They are highly stable under physiological conditions, biocompatible, biodegradable, low in toxicity, and considered safe for in vivo applications by the FDA [488,489]. Adjustable pore sizes enable the loading of small molecules into small nanoparticles [486]. Surface silanol groups (Si-OH) facilitate functionalization with amines, thiols, chlorides, phosphates, carboxyl groups, and others, allowing the attachment of a variety of molecules [488,489,490,491]. This versatility supports the development of controlled release systems [486], targeted therapy [489,490], stimuli-responsive systems [492,493], theranostics [494,495], and multimodal therapy [496,497,498]. They can be either conventional (MSN) or hollow (HMS), with different synthesis methods utilized depending on the specific application, as summarized in several publications [487,499].
4.2.3. siRNA as a Tool for Multimodal Cancer Treatment
5. siRNA-Targeted Genes Under Clinical Investigation
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Morris, K.V. SiRNA-Mediated Transcriptional Gene Silencing: The Potential Mechanism and a Possible Role in the Histone Code. Cell Mol. Life Sci. 2005, 62, 3057–3066. [Google Scholar] [CrossRef]
- Morris, K.V.; Chan, S.W.L.; Jacobsen, S.E.; Looney, D.J. Small Interfering RNA-Induced Transcriptional Gene Silencing in Human Cells. Science (1979) 2004, 305, 1289–1292. [Google Scholar] [CrossRef]
- Ting, A.H.; Schuebel, K.E.; Herman, J.G.; Baylin, S.B. Short Double-Stranded RNA Induces Transcriptional Gene Silencing in Human Cancer Cells in the Absence of DNA Methylation. Nat. Genet. 2005, 37, 906–910. [Google Scholar] [CrossRef]
- Zhang, X.; Rossi, J.J. Phylogenetic Comparison of Small RNA-Triggered Transcriptional Gene Silencing. J. Biol. Chem. 2011, 286, 29443–29448. [Google Scholar] [CrossRef]
- Zhang, K.; Wang, Q.; Xie, Y.; Mor, G.; Sega, E.; Low, P.S.; Huang, Y. Receptor-Mediated Delivery of SiRNAs by Tethered Nucleic Acid Base-Paired Interactions. RNA 2008, 14, 577–583. [Google Scholar] [CrossRef]
- Weinberg, M.S.; Morris, K.V. Transcriptional Gene Silencing in Humans. Nucleic. Acids. Res. 2016, 44, 6505–6517. [Google Scholar] [CrossRef] [PubMed]
- Motamedi, H.; Ari, M.M.; Alvandi, A.; Abiri, R. Principle, Application and Challenges of Development SiRNA-Based Therapeutics against Bacterial and Viral Infections: A Comprehensive Review. Front. Microbiol. 2024, 15, 1393646. [Google Scholar] [CrossRef] [PubMed]
- Guo, F.; Li, Y.; Yu, W.; Fu, Y.; Zhang, J.; Cao, H. Recent Progress of Small Interfering RNA Delivery on the Market and Clinical Stage. Mol. Pharm. 2024, 21, 2081–2096. [Google Scholar] [CrossRef]
- Friedrich, M.; Aigner, A. Therapeutic SiRNA: State-of-the-Art and Future Perspectives. BioDrugs 2022, 36, 549–571. [Google Scholar] [CrossRef]
- Ebenezer, O.; Oyebamiji, A.K.; Olanlokun, J.O.; Tuszynski, J.A.; Wong, G.K.-S. Recent Update on SiRNA Therapeutics. Int. J. Mol. Sci. 2025, 26, 3456. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Zhong, L.; Weng, Y.; Peng, L.; Huang, Y.; Zhao, Y.; Liang, X.-J. Therapeutic SiRNA: State of the Art. Signal Transduct. Target. Ther. 2020, 5, 101. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Chen, B.; Gan, C.; Sun, H.; Zhang, J.; Feng, L. A Comprehensive Review of Small Interfering RNAs (SiRNAs): Mechanism, Therapeutic Targets, and Delivery Strategies for Cancer Therapy. Int. J. Nanomed. 2023, 18, 7605–7635. [Google Scholar] [CrossRef] [PubMed]
- Lisowiec-Wąchnicka, J.; Bartyś, N.; Pasternak, A. A Systematic Study on the Influence of Thermodynamic Asymmetry of 5′-Ends of SiRNA Duplexes in Relation to Their Silencing Potency. Sci. Rep. 2019, 9, 255. [Google Scholar] [CrossRef]
- Ye, X.; Huang, N.; Liu, Y.; Paroo, Z.; Huerta, C.; Li, P.; Chen, S.; Liu, Q.; Zhang, H. Structure of C3PO and Mechanism of Human RISC Activation. Nat. Struct. Mol. Biol. 2011, 18, 650–657. [Google Scholar] [CrossRef]
- Sahu, S.; Williams, L.; Perez, A.; Philip, F.; Caso, G.; Zurawsky, W.; Scarlata, S. Regulation of the Activity of the Promoter of RNA-Induced Silencing, C3PO. Protein Sci. 2017, 26, 1807–1818. [Google Scholar] [CrossRef]
- Dong, Y.; Siegwart, D.J.; Anderson, D.G. Strategies, Design, and Chemistry in SiRNA Delivery Systems. Adv. Drug Deliv. Rev. 2019, 144, 133–147. [Google Scholar] [CrossRef]
- Elbashir, S.M. Functional Anatomy of SiRNAs for Mediating Efficient RNAi in Drosophila Melanogaster Embryo Lysate. EMBO J. 2001, 20, 6877–6888. [Google Scholar] [CrossRef]
- Martinelli, D.D. Machine Learning for SiRNA Efficiency Prediction: A Systematic Review. Health Sci. Rev. 2024, 11, 100157. [Google Scholar] [CrossRef]
- Fakhr, E.; Zare, F.; Teimoori-Toolabi, L. Precise and Efficient SiRNA Design: A Key Point in Competent Gene Silencing. Cancer Gene Ther. 2016, 23, 73–82. [Google Scholar] [CrossRef] [PubMed]
- Setten, R.L.; Rossi, J.J.; Han, S. The Current State and Future Directions of RNAi-Based Therapeutics. Nat. Rev. Drug Discov. 2019, 18, 421–446. [Google Scholar] [CrossRef]
- Lam, J.K.-W.; Liang, W.; Chan, H.-K. Pulmonary Delivery of Therapeutic SiRNA. Adv. Drug Deliv. Rev. 2012, 64, 1–15. [Google Scholar] [CrossRef]
- Lück, S.; Kreszies, T.; Strickert, M.; Schweizer, P.; Kuhlmann, M.; Douchkov, D. SiRNA-Finder (Si-Fi) Software for RNAi-Target Design and Off-Target Prediction. Front. Plant Sci. 2019, 10, 1023. [Google Scholar] [CrossRef]
- Birmingham, A.; Anderson, E.; Sullivan, K.; Reynolds, A.; Boese, Q.; Leake, D.; Karpilow, J.; Khvorova, A. A Protocol for Designing SiRNAs with High Functionality and Specificity. Nat. Protoc. 2007, 2, 2068–2078. [Google Scholar] [CrossRef]
- Piekna-Przybylska, D.; DiChiacchio, L.; Mathews, D.H.; Bambara, R.A. A Sequence Similar to TRNA3Lys Gene Is Embedded in HIV-1 U3–R and Promotes Minus-Strand Transfer. Nat. Struct. Mol. Biol. 2010, 17, 83–90. [Google Scholar] [CrossRef] [PubMed]
- Mysara, M.; Garibaldi, J.M.; ElHefnawi, M. Mysirna-Designer: A Workflow for Efficient SiRNA Design. PLoS ONE 2011, 6, e25642. [Google Scholar] [CrossRef] [PubMed]
- Ayyagari, V.S. Design of SiRNA Molecules for Silencing of Membrane Glycoprotein, Nucleocapsid Phosphoprotein, and Surface Glycoprotein Genes of SARS-CoV2. J. Genet. Eng. Biotechnol. 2022, 20, 113. [Google Scholar] [CrossRef]
- Dar, S.A.; Gupta, A.K.; Thakur, A.; Kumar, M. SMEpred Workbench: A Web Server for Predicting Efficacy of Chemicallymodified SiRNAs. RNA Biol. 2016, 13, 1144–1151. [Google Scholar] [CrossRef]
- Dar, S.A.; Thakur, A.; Qureshi, A.; Kumar, M. SiRNAmod: A Database of Experimentally Validated Chemically Modified SiRNAs. Sci. Rep. 2016, 6, 20031. [Google Scholar] [CrossRef] [PubMed]
- Shtykalova, S.; Deviatkin, D.; Freund, S.; Egorova, A.; Kiselev, A. Non-Viral Carriers for Nucleic Acids Delivery: Fundamentals and Current Applications. Life 2023, 13, 903. [Google Scholar] [CrossRef]
- Paul, A.; Muralidharan, A.; Biswas, A.; Kamath, B.V.; Joseph, A.; Alex, A.T. SiRNA Therapeutics and Its Challenges: Recent Advances in Effective Delivery for Cancer Therapy. OpenNano 2022, 7, 100063. [Google Scholar] [CrossRef]
- Davis, S.M.; Sousa, J.; Vangjeli, L.; Hassler, M.R.; Echeverria, D.; Knox, E.; Turanov, A.A.; Alterman, J.F.; Khvorova, A. 2′-O-Methyl at 20-Mer Guide Strand 3′ Termini May Negatively Affect Target Silencing Activity of Fully Chemically Modified SiRNA. Mol. Ther. Nucleic. Acids. 2020, 21, 266–277. [Google Scholar] [CrossRef]
- Basila, M.; Kelley, M.L.; Smith, A.V.B. Minimal 2′-O-Methyl Phosphorothioate Linkage Modification Pattern of Synthetic Guide RNAs for Increased Stability and Efficient CRISPR-Cas9 Gene Editing Avoiding Cellular Toxicity. PLoS ONE 2017, 12, e0188593. [Google Scholar] [CrossRef]
- Gore, K.R.; Nawale, G.N.; Harikrishna, S.; Chittoor, V.G.; Pandey, S.K.; Höbartner, C.; Patankar, S.; Pradeepkumar, P.I. Synthesis, Gene Silencing, and Molecular Modeling Studies of 4′C-Aminomethyl-2′-O-Methyl Modified Small Interfering RNAs. J. Org. Chem. 2012, 77, 3233–3245. [Google Scholar] [CrossRef]
- Harp, J.M.; Guenther, D.C.; Bisbe, A.; Perkins, L.; Matsuda, S.; Bommineni, G.R.; Zlatev, I.; Foster, D.J.; Taneja, N.; Charisse, K.; et al. Structural Basis for the Synergy of 4ʹ- and 2ʹ-Modifications on SiRNA Nuclease Resistance, Thermal Stability and RNAi Activity. Nucleic. Acids. Res. 2018, 46, 8090–8104. [Google Scholar] [CrossRef]
- Song, X.; Wang, X.; Ma, Y.; Liang, Z.; Yang, Z.; Cao, H. Site-Specific Modification Using the 2′-Methoxyethyl Group Improves the Specificity and Activity of SiRNAs. Mol. Ther. Nucleic. Acids. 2017, 9, 242–250. [Google Scholar] [CrossRef] [PubMed]
- Ly, S.; Echeverria, D.; Sousa, J.; Khvorova, A. Single-Stranded Phosphorothioated Regions Enhance Cellular Uptake of Cholesterol-Conjugated SiRNA but Not Silencing Efficacy. Mol. Ther. Nucleic. Acids. 2020, 21, 991–1005. [Google Scholar] [CrossRef] [PubMed]
- Hassler, M.R.; Turanov, A.A.; Alterman, J.F.; Haraszti, R.A.; Coles, A.H.; Osborn, M.F.; Echeverria, D.; Nikan, M.; Salomon, W.E.; Roux, L.; et al. Comparison of Partially and Fully Chemically-Modified SiRNA in Conjugate-Mediated Delivery in Vivo. Nucleic. Acids. Res. 2018, 46, 2185–2196. [Google Scholar] [CrossRef]
- Czauderna, F.; Fechtner, M.; Dames, S.; Aygün, H.; Klippel, A.; Pronk, G.J.; Giese, K.; Kaufmann, J. Structural Variations and Stabilising Modifications of Synthetic SiRNAs in Mammalian Cells. Nucleic. Acids. Res. 2003, 31, 2705–2716. [Google Scholar] [CrossRef] [PubMed]
- Chernikov, I.V.; Vlassov, V.V.; Chernolovskaya, E.L. Current Development of SiRNA Bioconjugates: From Research to the Clinic. Front. Pharmacol. 2019, 10, 444. [Google Scholar] [CrossRef]
- Prakash, T.P.; Allerson, C.R.; Dande, P.; Vickers, T.A.; Sioufi, N.; Jarres, R.; Baker, B.F.; Swayze, E.E.; Griffey, R.H.; Bhat, B. Positional Effect of Chemical Modifications on Short Interference RNA Activity in Mammalian Cells. J. Med. Chem. 2005, 48, 4247–4253. [Google Scholar] [CrossRef]
- Gangopadhyay, S.; Gore, K.R. Advances in SiRNA Therapeutics and Synergistic Effect on SiRNA Activity Using Emerging Dual Ribose Modifications. RNA Biol. 2022, 19, 452–467. [Google Scholar] [CrossRef] [PubMed]
- Esparza-Garrido, R.; Velázquez-Flores, M. Activation of Toll-like Receptors by Non-coding RNAs and Their Fragments (Review). Mol. Med. Rep. 2025, 32, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Pallan, P.S.; Greene, E.M.; Jicman, P.A.; Pandey, R.K.; Manoharan, M.; Rozners, E.; Egli, M. Unexpected Origins of the Enhanced Pairing Affinity of 2′-Fluoro-Modified RNA. Nucleic. Acids. Res. 2011, 39, 3482–3495. [Google Scholar] [CrossRef]
- Ge, Q.; Dallas, A.; Ilves, H.; Shorenstein, J.; Behlke, M.A.; Johnston, B.H. Effects of Chemical Modification on the Potency, Serum Stability, and Immunostimulatory Properties of Short ShRNAs. RNA 2010, 16, 118–130. [Google Scholar] [CrossRef][Green Version]
- Cochran, M.; Arias, D.; Burke, R.; Chu, D.; Erdogan, G.; Hood, M.; Kovach, P.; Kwon, H.W.; Chen, Y.; Moon, M.; et al. Structure-Activity Relationship of Antibody-Oligonucleotide Conjugates: Evaluating Bioconjugation Strategies for Antibody-SiRNA Conjugates for Drug Development. J. Med. Chem. 2024, 67, 14852–14867. [Google Scholar] [CrossRef]
- Langkjær, N.; Pasternak, A.; Wengel, J. UNA (Unlocked Nucleic Acid): A Flexible RNA Mimic That Allows Engineering of Nucleic Acid Duplex Stability. Bioorg Med. Chem. 2009, 17, 5420–5425. [Google Scholar] [CrossRef]
- Snead, N.M.; Escamilla-Powers, J.R.; Rossi, J.J.; McCaffrey, A.P. 5′ Unlocked Nucleic Acid Modification Improves SiRNA Targeting. Mol. Ther. Nucleic. Acids. 2013, 2, e103. [Google Scholar] [CrossRef]
- Laursen, M.B.; Pakula, M.M.; Gao, S.; Fluiter, K.; Mook, O.R.; Baas, F.; Langklær, N.; Wengel, S.L.; Wengel, J.; Kjems, J.; et al. Utilization of Unlocked Nucleic Acid (UNA) to Enhance SiRNA Performance in Vitro and in Vivo. Mol. Biosyst. 2010, 6, 862–870. [Google Scholar] [CrossRef]
- Crooke, S.T.; Seth, P.P.; Vickers, T.A.; Liang, X. The Interaction of Phosphorothioate-Containing RNA Targeted Drugs with Proteins Is a Critical Determinant of the Therapeutic Effects of These Agents. J. Am. Chem. Soc. 2020, 142, 14754–14771. [Google Scholar] [CrossRef]
- Yang, X.; Sierant, M.; Janicka, M.; Peczek, L.; Martinez, C.; Hassell, T.; Li, N.; Li, X.; Wang, T.; Nawrot, B. Gene Silencing Activity of SiRNA Molecules Containing Phosphorodithioate Substitutions. ACS Chem. Biol. 2012, 7, 1214–1220. [Google Scholar] [CrossRef] [PubMed]
- Zharkov, T.D.; Markov, O.V.; Zhukov, S.A.; Khodyreva, S.N.; Kupryushkin, M.S. Influence of Combinations of Lipophilic and Phosphate Backbone Modifications on Cellular Uptake of Modified Oligonucleotides. Molecules 2024, 29, 452. [Google Scholar] [CrossRef]
- Hall, A.H.S.; Wan, J.; Spesock, A.; Sergueeva, Z.; Shaw, B.R.; Alexander, K.A. High Potency Silencing by Single-Stranded Boranophosphate SiRNA. Nucleic. Acids. Res. 2006, 34, 2773–2781. [Google Scholar] [CrossRef]
- Hall, A.H.S. RNA Interference Using Boranophosphate SiRNAs: Structure-Activity Relationships. Nucleic. Acids. Res. 2004, 32, 5991–6000. [Google Scholar] [CrossRef]
- Ryan, D.E.; Diamant-Levi, T.; Steinfeld, I.; Taussig, D.; Visal-Shah, S.; Thakker, S.; Lunstad, B.D.; Kaiser, R.J.; McCaffrey, R.; Ortiz, M.; et al. Phosphonoacetate Modifications Enhance the Stability and Editing Yields of Guide RNAs for Cas9 Editors. Biochemistry 2023, 62, 3512–3520. [Google Scholar] [CrossRef]
- Hawner, M.; Ducho, C. Cellular Targeting of Oligonucleotides by Conjugation with Small Molecules. Molecules 2020, 25, 5963. [Google Scholar] [CrossRef]
- Berk, C.; Civenni, G.; Wang, Y.; Steuer, C.; Catapano, C.V.; Hall, J. Pharmacodynamic and Pharmacokinetic Properties of Full Phosphorothioate Small Interfering RNAs for Gene Silencing in Vivo. Nucleic. Acid. Ther. 2021, 31, 237–244. [Google Scholar] [CrossRef]
- Crooke, S.T.; Wang, S.; Vickers, T.A.; Shen, W.; Liang, X. Cellular Uptake and Trafficking of Antisense Oligonucleotides. Nat. Biotechnol. 2017, 35, 230–237. [Google Scholar] [CrossRef]
- Migawa, M.T.; Shen, W.; Wan, W.B.; Vasquez, G.; Oestergaard, M.E.; Low, A.; De Hoyos, C.L.; Gupta, R.; Murray, S.; Tanowitz, M.; et al. Site-Specific Replacement of Phosphorothioate with Alkyl Phosphonate Linkages Enhances the Therapeutic Profile of Gapmer ASOs by Modulating Interactions with Cellular Proteins. Nucleic. Acids. Res. 2019, 47, 5465–5479. [Google Scholar] [CrossRef] [PubMed]
- Shen, W.; De Hoyos, C.L.; Migawa, M.T.; Vickers, T.A.; Sun, H.; Low, A.; Bell, T.A.; Rahdar, M.; Mukhopadhyay, S.; Hart, C.E.; et al. Chemical Modification of PS-ASO Therapeutics Reduces Cellular Protein-Binding and Improves the Therapeutic Index. Nat. Biotechnol. 2019, 37, 640–650. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, Y.; Sato, K.; Wada, T. Solid-Phase Synthesis of Boranophosphate/Phosphorothioate/Phosphate Chimeric Oligonucleotides and Their Potential as Antisense Oligonucleotides. J. Org. Chem. 2022, 87, 3895–3909. [Google Scholar] [CrossRef] [PubMed]
- Duschmalé, J.; Schäublin, A.; Funder, E.; Schmidt, S.; Kiełpiński, Ł.J.; Nymark, H.; Jensen, K.; Koch, T.; Duschmalé, M.; Koller, E.; et al. Investigating Discovery Strategies and Pharmacological Properties of Stereodefined Phosphorodithioate LNA Gapmers. Mol. Ther. Nucleic. Acids. 2022, 29, 176–188. [Google Scholar] [CrossRef]
- Wu, S.Y.; Yang, X.; Gharpure, K.M.; Hatakeyama, H.; Egli, M.; McGuire, M.H.; Nagaraja, A.S.; Miyake, T.M.; Rupaimoole, R.; Pecot, C.V.; et al. 2′-OMe-Phosphorodithioate-Modified SiRNAs Show Increased Loading into the RISC Complex and Enhanced Anti-Tumour Activity. Nat. Commun. 2014, 5, 3455. [Google Scholar] [CrossRef]
- Elkayam, E.; Parmar, R.; Brown, C.R.; Willoughby, J.L.; Theile, C.S.; Manoharan, M.; Joshua-Tor, L. SiRNA Carrying an (E)-Vinylphosphonate Moiety at the 5′ End of the Guide Strand Augments Gene Silencing by Enhanced Binding to Human Argonaute-2. Nucleic. Acids. Res. 2017, 45, 3528–3536. [Google Scholar] [CrossRef] [PubMed]
- Parmar, R.; Willoughby, J.L.S.; Liu, J.; Foster, D.J.; Brigham, B.; Theile, C.S.; Charisse, K.; Akinc, A.; Guidry, E.; Pei, Y.; et al. 5′-(E)-Vinylphosphonate: A Stable Phosphate Mimic Can Improve the RNAi Activity of SiRNA-GalNAc Conjugates. ChemBioChem 2016, 17, 985–989. [Google Scholar] [CrossRef]
- Parmar, R.G.; Brown, C.R.; Matsuda, S.; Willoughby, J.L.S.; Theile, C.S.; Charissé, K.; Foster, D.J.; Zlatev, I.; Jadhav, V.; Maier, M.A.; et al. Facile Synthesis, Geometry, and 2′-Substituent-Dependent in Vivo Activity of 5′-(E)- and 5′-(Z)-Vinylphosphonate-Modified SiRNA Conjugates. J. Med. Chem. 2018, 61, 734–744. [Google Scholar] [CrossRef]
- Tsubaki, K.; Hammill, M.L.; Varley, A.J.; Kitamura, M.; Okauchi, T.; Desaulniers, J.-P. Synthesis and Evaluation of Neutral Phosphate Triester Backbone-Modified SiRNAs. ACS Med. Chem. Lett. 2020, 11, 1457–1462. [Google Scholar] [CrossRef] [PubMed]
- Sipa, K.; Sochacka, E.; Kazmierczak-Baranska, J.; Maszewska, M.; Janicka, M.; Nowak, G.; Nawrot, B. Effect of Base Modifications on Structure, Thermodynamic Stability, and Gene Silencing Activity of Short Interfering RNA. RNA 2007, 13, 1301–1316. [Google Scholar] [CrossRef] [PubMed]
- Peacock, H.; Kannan, A.; Beal, P.A.; Burrows, C.J. Chemical Modification of SiRNA Bases to Probe and Enhance RNA Interference. J. Org. Chem. 2011, 76, 7295–7300. [Google Scholar] [CrossRef]
- Shinohara, F.; Oashi, T.; Harumoto, T.; Nishikawa, T.; Takayama, Y.; Miyagi, H.; Takahashi, Y.; Nakajima, T.; Sawada, T.; Koda, Y.; et al. SiRNA Potency Enhancement via Chemical Modifications of Nucleotide Bases at the 5 ′-End of the SiRNA Guide Strand. RNA 2021, 27, 163–173. [Google Scholar] [CrossRef]
- Valenzuela, R.A.P.; Suter, S.R.; Ball-Jones, A.A.; Ibarra-Soza, J.M.; Zheng, Y.; Beal, P.A. Base Modification Strategies to Modulate Immune Stimulation by an SiRNA. ChemBioChem 2015, 16, 262–267. [Google Scholar] [CrossRef]
- Li, Q.; Dong, M.; Chen, P. Advances in Structural-Guided Modifications of SiRNA. Bioorg. Med. Chem. 2024, 110, 117825. [Google Scholar] [CrossRef]
- Chernikov, I.V.; Ponomareva, U.A.; Chernolovskaya, E.L. Structural Modifications of SiRNA Improve Its Performance In Vivo. Int. J. Mol. Sci. 2023, 24, 956. [Google Scholar] [CrossRef]
- Li, Z.; Wang, X.; Zhou, X.; Wang, J.; Guan, Z.; Yang, Z. Optimization in Chemical Modification of Single-Stranded SiRNA Encapsulated by Neutral Cytidinyl/Cationic Lipids. Front. Chem. 2022, 10, 843181. [Google Scholar] [CrossRef]
- Bulcha, J.T.; Wang, Y.; Ma, H.; Tai, P.W.L.; Gao, G. Viral Vector Platforms within the Gene Therapy Landscape. Signal Transduct. Target. Ther. 2021, 6, 53. [Google Scholar] [CrossRef]
- Lundstrom, K. Viral Vectors in Gene Therapy: Where Do We Stand in 2023? Viruses 2023, 15, 698. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.W.; Choi, J.; Kim, E.H.; Choi, J.; Kim, S.H.; Yang, Y. Design of SiRNA Bioconjugates for Efficient Control of Cancer-Associated Membrane Receptors. ACS Omega 2023, 8, 36435–36448. [Google Scholar] [CrossRef]
- Lee, J.W.; Choi, J.; Choi, Y.; Kim, K.; Yang, Y.; Kim, S.H.; Yoon, H.Y.; Kwon, I.C. Molecularly Engineered SiRNA Conjugates for Tumor-Targeted RNAi Therapy. J. Control. Release 2022, 351, 713–726. [Google Scholar] [CrossRef] [PubMed]
- Tai, W. Current Aspects of SiRNA Bioconjugate for In Vitro and In Vivo Delivery. Molecules 2019, 24, 2211. [Google Scholar] [CrossRef]
- Im, S.-W.; Pravinsagar, P.; Im, S.-R.; Jang, Y.-J. Variable Heavy Chain Domain Derived from a Cell-Penetrating Anti-DNA Monoclonal Antibody for the Intracellular Delivery of Biomolecules. Immunol. Investig. 2017, 46, 500–517. [Google Scholar] [CrossRef] [PubMed]
- Cao, W.; Li, R.; Pei, X.; Chai, M.; Sun, L.; Huang, Y.; Wang, J.; Barth, S.; Yu, F.; He, H. Antibody–SiRNA Conjugates (ARC): Emerging SiRNA Drug Formulation. Med. Drug Discov. 2022, 15, 100128. [Google Scholar] [CrossRef]
- Li, G.; Moellering, R.E. A Concise, Modular Antibody–Oligonucleotide Conjugation Strategy Based on Disuccinimidyl Ester Activation Chemistry. ChemBioChem 2019, 20, 1599–1605. [Google Scholar] [CrossRef]
- Dugal-Tessier, J.; Thirumalairajan, S.; Jain, N. Antibody-Oligonucleotide Conjugates: A Twist to Antibody-Drug Conjugates. J. Clin. Med. 2021, 10, 838. [Google Scholar] [CrossRef]
- Bäumer, N.; Berdel, W.E.; Bäumer, S. Immunoprotein-Mediated SiRNA Delivery. Mol. Pharm. 2017, 14, 1339–1351. [Google Scholar] [CrossRef]
- Ma, Y.; Kowolik, C.M.; Swiderski, P.M.; Kortylewski, M.; Yu, H.; Horne, D.A.; Jove, R.; Caballero, O.L.; Simpson, A.J.G.; Lee, F.T.; et al. Humanized Lewis-Y Specific Antibody Based Delivery of STAT3 SiRNA. ACS Chem. Biol. 2011, 6, 962–970. [Google Scholar] [CrossRef]
- Cuellar, T.L.; Barnes, D.; Nelson, C.; Tanguay, J.; Yu, S.-F.; Wen, X.; Scales, S.J.; Gesch, J.; Davis, D.; van Brabant Smith, A.; et al. Systematic Evaluation of Antibody-Mediated SiRNA Delivery Using an Industrial Platform of THIOMAB–SiRNA Conjugates. Nucleic. Acids. Res. 2015, 43, 1189–1203. [Google Scholar] [CrossRef] [PubMed]
- Hammond, S.M.; Abendroth, F.; Goli, L.; Stoodley, J.; Burrell, M.; Thom, G.; Gurrell, I.; Ahlskog, N.; Gait, M.J.; Wood, M.J.A.; et al. Antibody-Oligonucleotide Conjugate Achieves CNS Delivery in Animal Models for Spinal Muscular Atrophy. JCI Insight 2022, 7, e154142. [Google Scholar] [CrossRef] [PubMed]
- Sugo, T.; Terada, M.; Oikawa, T.; Miyata, K.; Nishimura, S.; Kenjo, E.; Ogasawara-Shimizu, M.; Makita, Y.; Imaichi, S.; Murata, S.; et al. Development of Antibody-SiRNA Conjugate Targeted to Cardiac and Skeletal Muscles. J. Control. Release 2016, 237, 1–13. [Google Scholar] [CrossRef]
- Yu, Z.; Zhang, X.; Pei, X.; Cao, W.; Ye, J.; Wang, J.; Sun, L.; Yu, F.; Wang, J.; Li, N.; et al. Antibody-SiRNA Conjugates (ARCs) Using Multifunctional Peptide as a Tumor Enzyme Cleavable Linker Mediated Effective Intracellular Delivery of SiRNA. Int. J. Pharm. 2021, 606, 120940. [Google Scholar] [CrossRef] [PubMed]
- Dovgan, I.; Koniev, O.; Kolodych, S.; Wagner, A. Antibody–Oligonucleotide Conjugates as Therapeutic, Imaging, and Detection Agents. Bioconjug. Chem. 2019, 30, 2483–2501. [Google Scholar] [CrossRef]
- Sreedurgalakshmi, K.; Srikar, R.; Harikrishnan, K.; Srinivasan, L.; Rajkumari, R. Cetuximab–SiRNA Conjugate Linked Through Cationized Gelatin Knocks Down KRAS G12C Mutation in NSCLC Sensitizing the Cells Toward Gefitinib. Technol. Cancer. Res. Treat. 2021, 20, 15330338211041453. [Google Scholar] [CrossRef]
- Asiaei, S.; Smith, B.; Nieva, P. Enhancing Conjugation Rate of Antibodies to Carboxylates: Numerical Modeling of Conjugation Kinetics in Microfluidic Channels and Characterization of Chemical over-Exposure in Conventional Protocols by Quartz Crystal Microbalance. Biomicrofluidics 2015, 9, 024122. [Google Scholar] [CrossRef]
- Nanna, A.R.; Kel’in, A.V.; Theile, C.; Pierson, J.M.; Voo, Z.X.; Garg, A.; Nair, J.K.; Maier, M.A.; Fitzgerald, K.; Rader, C. Generation and Validation of Structurally Defined Antibody-SiRNA Conjugates. Nucleic. Acids. Res. 2020, 48, 5281–5293. [Google Scholar] [CrossRef] [PubMed]
- Satake, N.; Duong, C.; Yoshida, S.; Oestergaard, M.; Chen, C.; Peralta, R.; Guo, S.; Seth, P.P.; Li, Y.; Beckett, L.; et al. Novel Targeted Therapy for Precursor B-Cell Acute Lymphoblastic Leukemia: Anti-CD22 Antibody-MXD3 Antisense Oligonucleotide Conjugate. Mol. Med. 2016, 22, 632–642. [Google Scholar] [CrossRef]
- Honcharenko, D.; Druceikaite, K.; Honcharenko, M.; Bollmark, M.; Tedebark, U.; Strömberg, R. New Alkyne and Amine Linkers for Versatile Multiple Conjugation of Oligonucleotides. ACS Omega 2021, 6, 579–593. [Google Scholar] [CrossRef]
- Perrone, D.; Marchesi, E.; Preti, L.; Navacchia, M.L. Modified Nucleosides, Nucleotides and Nucleic Acids via Click Azide-Alkyne Cycloaddition for Pharmacological Applications. Molecules 2021, 26, 3100. [Google Scholar] [CrossRef]
- Kumar, P.; Ban, H.-S.; Kim, S.-S.; Wu, H.; Pearson, T.; Greiner, D.L.; Laouar, A.; Yao, J.; Haridas, V.; Habiro, K.; et al. T Cell-Specific SiRNA Delivery Suppresses HIV-1 Infection in Humanized Mice. Cell 2008, 134, 577–586. [Google Scholar] [CrossRef] [PubMed]
- Bäumer, N.; Rehkämper, J.; Appel, N.; Terheyden, L.; Hartmann, W.; Wardelmann, E.; Buchholz, F.; Müller-Tidow, C.; Berdel, W.E.; Bäumer, S. Downregulation of PIK3CA via Antibody-EsiRNA-Complexes Suppresses Human Xenograft Tumor Growth. PLoS ONE 2018, 13, e0200163. [Google Scholar] [CrossRef]
- Shi, S.-J.; Wang, L.-J.; Han, D.-H.; Wu, J.-H.; Jiao, D.; Zhang, K.-L.; Chen, J.-W.; Li, Y.; Yang, F.; Zhang, J.-L.; et al. Therapeutic Effects of Human Monoclonal PSMA Antibody-Mediated TRIM24 SiRNA Delivery in PSMA-Positive Castration-Resistant Prostate Cancer. Theranostics 2019, 9, 1247–1263. [Google Scholar] [CrossRef]
- Bäumer, N.; Tiemann, J.; Scheller, A.; Meyer, T.; Wittmann, L.; Suburu, M.E.G.; Greune, L.; Peipp, M.; Kellmann, N.; Gumnior, A.; et al. Targeted SiRNA Nanocarrier: A Platform Technology for Cancer Treatment. Oncogene 2022, 41, 2210–2224. [Google Scholar] [CrossRef] [PubMed]
- Xia, C.F.; Boado, R.J.; Pardridge, W.M. Antibody-Mediated Targeting of SiRNA via the Human Insulin Receptor Using Avidin-Biotin Technology. Mol. Pharm. 2009, 6, 747–751. [Google Scholar] [CrossRef]
- Xia, C.F.; Zhang, Y.; Zhang, Y.; Boado, R.J.; Pardridge, W.M. Intravenous SiRNA of Brain Cancer with Receptor Targeting and Avidin-Biotin Technology. Pharm. Res. 2007, 24, 2309–2316. [Google Scholar] [CrossRef]
- Falato, L.; Gestin, M.; Langel, Ü. Cell-Penetrating Peptides Delivering SiRNAs: An Overview. In Design and Delivery of SiRNA Therapeutics; Methods in Molecular Biology; Humana Press: New York, NY, USA, 2021; Volume 2282, pp. 329–352. [Google Scholar]
- Lopuszynski, J.; Agrawal, V.; Zahid, M. Tissue-Specific Cell Penetrating Peptides for Targeted Delivery of Small Interfering RNAs. Med. Res. Arch. 2022, 10, 2998. [Google Scholar] [CrossRef]
- Walrant, A.; Cardon, S.; Burlina, F.; Sagan, S. Membrane Crossing and Membranotropic Activity of Cell-Penetrating Peptides: Dangerous Liaisons? Acc. Chem. Res. 2017, 50, 2968–2975. [Google Scholar] [CrossRef] [PubMed]
- Sajid, M.I.; Mandal, D.; El-Sayed, N.S.; Lohan, S.; Moreno, J.; Tiwari, R.K. Oleyl Conjugated Histidine-Arginine Cell-Penetrating Peptides as Promising Agents for SiRNA Delivery. Pharmaceutics 2022, 14, 881. [Google Scholar] [CrossRef]
- Jagrosse, M.L.; Baliga, U.K.; Jones, C.W.; Russell, J.J.; García, C.I.; Najar, R.A.; Rahman, A.; Dean, D.A.; Nilsson, B.L. Impact of Peptide Sequence on Functional SiRNA Delivery and Gene Knockdown with Cyclic Amphipathic Peptide Delivery Agents. Mol. Pharm. 2023, 20, 6090–6103. [Google Scholar] [CrossRef] [PubMed]
- Ruseska, I.; Zimmer, A. Internalization Mechanisms of Cell-Penetrating Peptides. Beilstein J. Nanotechnol. 2020, 11, 101–123. [Google Scholar] [CrossRef]
- Beloor, J.; Zeller, S.; Choi, C.S.; Lee, S.-K.; Kumar, P. Cationic Cell-Penetrating Peptides as Vehicles for SiRNA Delivery. Ther. Deliv. 2015, 6, 491–507. [Google Scholar] [CrossRef]
- Moschos, S.A.; Jones, S.W.; Perry, M.M.; Williams, A.E.; Erjefalt, J.S.; Turner, J.J.; Barnes, P.J.; Sproat, B.S.; Gait, M.J.; Lindsay, M.A. Lung Delivery Studies Using SiRNA Conjugated to TAT(48-60) and Penetratin Reveal Peptide Induced Reduction in Gene Expression and Induction of Innate Immunity. Bioconjug. Chem. 2007, 18, 1450–1459. [Google Scholar] [CrossRef]
- Tai, W.; Gao, X. Functional Peptides for SiRNA Delivery. Adv. Drug Deliv. Rev. 2017, 110–111, 157–168. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Huang, Y.; Liu, Y.; Chen, Y.; Jin, H.; Zheng, Y.; Du, Q.; Yang, Z.; Zhang, L. Synthesis and Biological Evaluation of Peptide-SiRNA Conjugates with Phosphodiester Unit as Linker. Sci. China Chem. 2013, 56, 1542–1549. [Google Scholar] [CrossRef]
- Desale, K.; Kuche, K.; Jain, S. Cell-Penetrating Peptides (CPPs): An Overview of Applications for Improving the Potential of Nanotherapeutics. Biomater. Sci. 2021, 9, 1153–1188. [Google Scholar] [CrossRef]
- Muratovska, A.; Eccles, M.R. Conjugate for Efficient Delivery of Short Interfering RNA (SiRNA) into Mammalian Cells. FEBS Lett. 2004, 558, 63–68. [Google Scholar] [CrossRef]
- Yu, Z.; Ye, J.; Pei, X.; Sun, L.; Liu, E.; Wang, J.; Huang, Y.; Lee, S.J.; He, H. Improved Method for Synthesis of Low Molecular Weight Protamine–SiRNA Conjugate. Acta Pharm. Sin. B 2018, 8, 116–126. [Google Scholar] [CrossRef]
- Lin, L.; Chi, J.; Yan, Y.; Luo, R.; Feng, X.; Zheng, Y.; Xian, D.; Li, X.; Quan, G.; Liu, D.; et al. Membrane-Disruptive Peptides/Peptidomimetics-Based Therapeutics: Promising Systems to Combat Bacteria and Cancer in the Drug-Resistant Era. Acta Pharm. Sin. B 2021, 11, 2609–2644. [Google Scholar] [CrossRef] [PubMed]
- Pan, R.; Xu, W.; Yuan, F.; Chu, D.; Ding, Y.; Chen, B.; Jafari, M.; Yuan, Y.; Chen, P. A Novel Peptide for Efficient SiRNA Delivery in Vitro and Therapeutics in Vivo. Acta Biomater. 2015, 21, 74–84. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.; Dussouillez, C.; Padilla, B.; Frisch, B.; Mason, A.J.; Kichler, A. Design of a New Cell Penetrating Peptide for DNA, SiRNA and MRNA Delivery. J. Gene Med. 2022, 24, e3401. [Google Scholar] [CrossRef]
- He, S.; Cen, B.; Liao, L.; Wang, Z.; Qin, Y.; Wu, Z.; Liao, W.; Zhang, Z.; Ji, A. A Tumor-Targeting CRGD-EGFR SiRNA Conjugate and Its Anti-Tumor Effect on Glioblastoma in Vitro and in Vivo. Drug Deliv. 2017, 24, 471–481. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Xu, S.; Xiang, T.; Liu, H.; Chen, L.; Jiang, B.; Yao, J.; Zhu, H.; Hu, R.; Chen, Z. Multifunctional Building Elements for the Construction of Peptide Drug Conjugates. Eng. Regen. 2022, 3, 92–109. [Google Scholar] [CrossRef]
- Zeller, S.; Choi, C.S.; Uchil, P.D.; Ban, H.-S.; Siefert, A.; Fahmy, T.M.; Mothes, W.; Lee, S.-K.; Kumar, P. Attachment of Cell-Binding Ligands to Arginine-Rich Cell-Penetrating Peptides Enables Cytosolic Translocation of Complexed SiRNA. Chem. Biol. 2015, 22, 50–62. [Google Scholar] [CrossRef]
- Cen, B.; Wei, Y.; Huang, W.; Teng, M.; He, S.; Li, J.; Wang, W.; He, G.; Bai, X.; Liu, X.; et al. An Efficient Bivalent Cyclic RGD-PIK3CB SiRNA Conjugate for Specific Targeted Therapy against Glioblastoma In Vitro and In Vivo. Mol. Ther. Nucleic. Acids. 2018, 13, 220–232. [Google Scholar] [CrossRef]
- Huang, Y.; Wang, X.; Huang, W.; Cheng, Q.; Zheng, S.; Guo, S.; Cao, H.; Liang, X.J.; Du, Q.; Liang, Z. Systemic Administration of SiRNA via CRGD-Containing Peptide. Sci. Rep. 2015, 5, 12458. [Google Scholar] [CrossRef]
- Juretić, D. Designed Multifunctional Peptides for Intracellular Targets. Antibiotics 2022, 11, 1196. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Liu, E.; Gong, J.; Wang, J.; Huang, Y.; He, H.; Yang, V.C. High-Yield Synthesis of Monomeric LMWP(CPP)-SiRNA Covalent Conjugate for Effective Cytosolic Delivery of SiRNA. Theranostics 2017, 7, 2495–2508. [Google Scholar] [CrossRef] [PubMed]
- Park, J.W.; Bang, E.-K.; Jeon, E.M.; Kim, B.H. Complexation and Conjugation Approaches to Evaluate SiRNA Delivery Using Cationic, Hydrophobic and Amphiphilic Peptides. Org. Biomol. Chem. 2012, 10, 96–102. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, X.F.; Chen, Y.; Zhang, L.H.; Yang, Z.J. A Solid-Phase Method for Peptide-SiRNA Covalent Conjugates Based on Click Chemistry. Medchemcomm 2012, 3, 506–511. [Google Scholar] [CrossRef]
- Matsubara, M.; Honda, K.; Ozaki, K.; Kajino, R.; Kakisawa, Y.; Maeda, Y.; Ueno, Y. Synthesis of SiRNAs Incorporated with Cationic Peptides R8G7 and R8A7 and the Effect of the Modifications on SiRNA Properties. RSC Adv. 2020, 10, 34815–34824. [Google Scholar] [CrossRef]
- Smidt, J.M.; Lykke, L.; Stidsen, C.E.; Pristovšek, N.; Gothelf, K.V. Synthesis of Peptide–SiRNA Conjugates via Internal Sulfonylphosphoramidate Modifications and Evaluation of Their in Vitro Activity. Nucleic. Acids. Res. 2024, 52, 49–58. [Google Scholar] [CrossRef]
- Ando, Y.; Nakazawa, H.; Miura, D.; Otake, M.; Umetsu, M. Enzymatic Ligation of an Antibody and Arginine 9 Peptide for Efficient and Cell-Specific SiRNA Delivery. Sci. Rep. 2021, 11, 10035. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.W.; Kim, N.Y.; Choi, Y.B.; Park, S.H.; Yang, J.M.; Shin, S. RNA Interference in Vitro and in Vivo Using an Arginine Peptide/SiRNA Complex System. J. Control. Release 2010, 143, 335–343. [Google Scholar] [CrossRef]
- Maeda, Y.; Iwata Hara, R.; Nishina, K.; Yoshida-Tanaka, K.; Sakamoto, T.; Yokota, T.; Wada, T. Artificial Cationic Peptides That Increase Nuclease Resistance of SiRNA without Disturbing RNAi Activity. Nucleosides Nucleotides Nucleic Acids 2019, 38, 338–348. [Google Scholar] [CrossRef] [PubMed]
- Crombez, L.; Aldrian-Herrada, G.; Konate, K.; Nguyen, Q.N.; McMaster, G.K.; Brasseur, R.; Heitz, F.; Divita, G. A New Potent Secondary Amphipathic Cell-Penetrating Peptide for SiRNA Delivery into Mammalian Cells. Mol. Ther. 2009, 17, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Crowet, J.M.; Lins, L.; Deshayes, S.; Divita, G.; Morris, M.; Brasseur, R.; Thomas, A. Modeling of Non-Covalent Complexes of the Cell-Penetrating Peptide CADY and Its SiRNA Cargo. Biochim. Biophys. Acta Biomembr. 2013, 1828, 499–509. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ali Zaidi, S.S.; Fatima, F.; Ali Zaidi, S.A.; Zhou, D.; Deng, W.; Liu, S. Engineering SiRNA Therapeutics: Challenges and Strategies. J. Nanobiotechnol. 2023, 21, 381. [Google Scholar] [CrossRef]
- Crombez, L.; Divita, G. A Non-Covalent Peptide-Based Strategy for SiRNA Delivery. In Methods in Molecular Biology; Humana Press Inc.: Totowa, NJ, USA, 2011; Volume 683, pp. 349–360. [Google Scholar]
- Wang, F.; Gao, L.; Meng, L.Y.; Xie, J.M.; Xiong, J.W.; Luo, Y. A Neutralized Noncharged Polyethylenimine-Based System for Efficient Delivery of SiRNA into Heart without Toxicity. ACS Appl. Mater. Interfaces 2016, 8, 33529–33538. [Google Scholar] [CrossRef]
- Farra, R.; Musiani, F.; Perrone, F.; Čemažar, M.; Kamenšek, U.; Tonon, F.; Abrami, M.; Ručigaj, A.; Grassi, M.; Pozzato, G.; et al. Polymer-Mediated Delivery of SiRNAs to Hepatocellular Carcinoma: Variables Affecting Specificity and Effectiveness. Molecules 2018, 23, 777. [Google Scholar] [CrossRef] [PubMed]
- Patil, Y.; Panyam, J. Polymeric Nanoparticles for SiRNA Delivery and Gene Silencing. Int. J. Pharm. 2009, 367, 195–203. [Google Scholar] [CrossRef]
- Cavallaro, G.; Sardo, C.; Craparo, E.F.; Porsio, B.; Giammona, G. Polymeric Nanoparticles for SiRNA Delivery: Production and Applications. Int. J. Pharm. 2017, 525, 313–333. [Google Scholar] [CrossRef]
- Zhang, L.; Lou, W.; Wang, J. Advances in Nucleic Acid Therapeutics: Structures, Delivery Systems, and Future Perspectives in Cancer Treatment. Clin. Exp. Med. 2024, 24, 200. [Google Scholar] [CrossRef]
- Guţoaia, A.; Schuster, L.; Margutti, S.; Laufer, S.; Schlosshauer, B.; Krastev, R.; Stoll, D.; Hartmann, H. Fine-Tuned PEGylation of Chitosan to Maintain Optimal SiRNA-Nanoplex Bioactivity. Carbohydr. Polym. 2016, 143, 25–34. [Google Scholar] [CrossRef]
- Jiang, Z.; Thayumanavan, S. Noncationic Material Design for Nucleic Acid Delivery. Adv. Ther. 2020, 3, 1900206. [Google Scholar] [CrossRef]
- Zamboni, C.G.; Kozielski, K.L.; Vaughan, H.J.; Nakata, M.M.; Kim, J.; Higgins, L.J.; Pomper, M.G.; Green, J.J. Polymeric Nanoparticles as Cancer-Specific DNA Delivery Vectors to Human Hepatocellular Carcinoma. J. Control. Release 2017, 263, 18–28. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Mundra, V.; Peng, Y.; Wang, Y.; Tan, C.; Mahato, R.I. Pharmacokinetics and Biodistribution of Polymeric Micelles Containing MiRNA and Small-Molecule Drug in Orthotopic Pancreatic Tumor-Bearing Mice. Theranostics 2018, 8, 4033–4049. [Google Scholar] [CrossRef]
- Ahmad, A.; Khan, J.M. PH-Sensitive Endosomolytic Peptides in Gene and Drug Delivery: Endosomal Escape and Current Challenges. J. Drug Deliv. Sci. Technol. 2022, 76, 103786. [Google Scholar] [CrossRef]
- Xie, L.; Liu, R.; Chen, X.; He, M.; Zhang, Y.; Chen, S. Micelles Based on Lysine, Histidine, or Arginine: Designing Structures for Enhanced Drug Delivery. Front. Bioeng. Biotechnol. 2021, 9, 744657. [Google Scholar] [CrossRef]
- Dutta, K.; Das, R.; Medeiros, J.; Kanjilal, P.; Thayumanavan, S. Charge-Conversion Strategies for Nucleic Acid Delivery. Adv. Funct. Mater. 2021, 31, 2011103. [Google Scholar] [CrossRef]
- Guo, J.; Cheng, W.P.; Gu, J.; Ding, C.; Qu, X.; Yang, Z.; O’Driscoll, C. Systemic Delivery of Therapeutic Small Interfering RNA Using a PH-Triggered Amphiphilic Poly-l-Lysine Nanocarrier to Suppress Prostate Cancer Growth in Mice. Eur. J. Pharm. Sci. 2012, 45, 521–532. [Google Scholar] [CrossRef]
- Gorzkiewicz, M.; Kopeć, O.; Janaszewska, A.; Konopka, M.; Pędziwiatr-Werbicka, E.; Tarasenko, I.I.; Bezrodnyi, V.V.; Neelov, I.M.; Klajnert-Maculewicz, B. Poly(Lysine) Dendrimers Form Complexes with Sirna and Provide Its Effcient Uptake by Myeloid Cells: Model Studies for Therapeutic Nucleic Acid Delivery. Int. J. Mol. Sci. 2020, 21, 3138. [Google Scholar] [CrossRef]
- Liyanage, W.; Wu, T.; Kannan, S.; Kannan, R.M. Dendrimer–SiRNA Conjugates for Targeted Intracellular Delivery in Glioblastoma Animal Models. ACS Appl. Mater. Interfaces 2022, 14, 46290–46303. [Google Scholar] [CrossRef]
- Osipova, O.; Zakharova, N.; Pyankov, I.; Egorova, A.; Kislova, A.; Lavrentieva, A.; Kiselev, A.; Tennikova, T.; Korzhikova-Vlakh, E. Amphiphilic PH-Sensitive Polypeptides for SiRNA Delivery. J. Drug Deliv. Sci. Technol. 2022, 69, 103135. [Google Scholar] [CrossRef]
- Egorova, A.A.; Kiselev, A.V.; Tarasenko, I.I.; Il’ina, P.L.; Pankova, G.A.; Il’ina, I.E.; Baranov, V.C.; Vlasov, G.P. Hyperbranched Polylysines Modified with Histidine and Arginine: The Optimization of Their DNA Compacting and Endosomolytic Properties. Russ. J. Bioorg. Chem. 2009, 35, 437–445. [Google Scholar] [CrossRef] [PubMed]
- Kruspe, S.; Giangrande, P. Aptamer-SiRNA Chimeras: Discovery, Progress, and Future Prospects. Biomedicines 2017, 5, 45. [Google Scholar] [CrossRef]
- Mahmoudian, F.; Ahmari, A.; Shabani, S.; Sadeghi, B.; Fahimirad, S.; Fattahi, F. Aptamers as an Approach to Targeted Cancer Therapy. Cancer Cell Int. 2024, 24, 108. [Google Scholar] [CrossRef]
- Sivakumar, P.; Kim, S.; Kang, H.C.; Shim, M.S. Targeted SiRNA Delivery Using Aptamer-siRNA Chimeras and Aptamer-conjugated Nanoparticles. WIREs Nanomed. Nanobiotechnol. 2019, 11, e1543. [Google Scholar] [CrossRef]
- Safarzadeh Kozani, P.; Safarzadeh Kozani, P.; Rahbarizadeh, F. Aptamer-Assisted Delivery of Nucleotides with Tumor-Suppressing Properties for Targeted Cancer Therapies. Trends Med. Sci. 2021, 1, 114909. [Google Scholar] [CrossRef]
- McNamara, J.O.; Andrechek, E.R.; Wang, Y.; Viles, K.D.; Rempel, R.E.; Gilboa, E.; Sullenger, B.A.; Giangrande, P.H. Cell Type–Specific Delivery of SiRNAs with Aptamer-SiRNA Chimeras. Nat. Biotechnol. 2006, 24, 1005–1015. [Google Scholar] [CrossRef]
- Odeh, F.; Nsairat, H.; Alshaer, W.; Ismail, M.A.; Esawi, E.; Qaqish, B.; Al Bawab, A.; Ismail, S.I. Aptamers Chemistry: Chemical Modifications and Conjugation Strategies. Molecules 2019, 25, 3. [Google Scholar] [CrossRef]
- Lai, W.Y.; Wang, W.Y.; Chang, Y.C.; Chang, C.J.; Yang, P.C.; Peck, K. Synergistic Inhibition of Lung Cancer Cell Invasion, Tumor Growth and Angiogenesis Using Aptamer-SiRNA Chimeras. Biomaterials 2014, 35, 2905–2914. [Google Scholar] [CrossRef]
- Yoo, H.; Jung, H.; Kim, S.A.; Mok, H. Multivalent Comb-Type Aptamer–SiRNA Conjugates for Efficient and Selective Intracellular Delivery. Chem. Commun. 2014, 50, 6765–6767. [Google Scholar] [CrossRef] [PubMed]
- Dassie, J.P.; Liu, X.Y.; Thomas, G.S.; Whitaker, R.M.; Thiel, K.W.; Stockdale, K.R.; Meyerholz, D.K.; McCaffrey, A.P.; McNamara, J.O.; Giangrande, P.H. Systemic Administration of Optimized Aptamer-SiRNA Chimeras Promotes Regression of PSMA-Expressing Tumors. Nat. Biotechnol. 2009, 27, 839–846. [Google Scholar] [CrossRef] [PubMed]
- Puplampu-Dove, Y.; Gefen, T.; Rajagopalan, A.; Muheramagic, D.; Schrand, B.; Gilboa, E. Potentiating Tumor Immunity Using Aptamer-Targeted RNAi to Render CD8+ T Cells Resistant to TGFβ Inhibition. Oncoimmunology 2018, 7, e1349588. [Google Scholar] [CrossRef] [PubMed]
- Esposito, C.L.; Nuzzo, S.; Catuogno, S.; Romano, S.; de Nigris, F.; de Franciscis, V. STAT3 Gene Silencing by Aptamer-SiRNA Chimera as Selective Therapeutic for Glioblastoma. Mol. Ther. Nucleic. Acids. 2018, 10, 398–411. [Google Scholar] [CrossRef] [PubMed]
- Herrmann, A.; Priceman, S.J.; Kujawski, M.; Xin, H.; Cherryholmes, G.A.; Zhang, W.; Zhang, C.; Lahtz, C.; Kowolik, C.; Forman, S.J.; et al. CTLA4 Aptamer Delivers STAT3 SiRNA to Tumor-Associated and Malignant T Cells. J. Clin. Investig. 2014, 124, 2977–2987. [Google Scholar] [CrossRef]
- Zhou, J.; Tiemann, K.; Chomchan, P.; Alluin, J.; Swiderski, P.; Burnett, J.; Zhang, X.; Forman, S.; Chen, R.; Rossi, J. Dual Functional BAFF Receptor Aptamers Inhibit Ligand-Induced Proliferation and Deliver SiRNAs to NHL Cells. Nucleic. Acids. Res. 2013, 41, 4266–4283. [Google Scholar] [CrossRef]
- Liu, H.Y.; Yu, X.; Liu, H.; Wu, D.; She, J.-X. Co-Targeting EGFR and Survivin with a Bivalent Aptamer-Dual SiRNA Chimera Effectively Suppresses Prostate Cancer. Sci. Rep. 2016, 6, 30346. [Google Scholar] [CrossRef]
- Xue, L.; Maihle, N.J.; Yu, X.; Tang, S.-C.; Liu, H.Y. Synergistic Targeting HER2 and EGFR with Bivalent Aptamer-SiRNA Chimera Efficiently Inhibits HER2-Positive Tumor Growth. Mol. Pharm. 2018, 15, 4801–4813. [Google Scholar] [CrossRef]
- Aaldering, L.J.; Tayeb, H.; Krishnan, S.; Fletcher, S.; Wilton, S.D.; Veedu, R.N. Smart Functional Nucleic Acid Chimeras: Enabling Tissue Specific RNA Targeting Therapy. RNA Biol. 2015, 12, 412–425. [Google Scholar] [CrossRef]
- Zhou, J.; Rossi, J.J. Cell-Type-Specific, Aptamer-Functionalized Agents for Targeted Disease Therapy. Mol. Ther. Nucleic. Acids. 2014, 3, e169. [Google Scholar] [CrossRef]
- Pastor, F.; Kolonias, D.; Giangrande, P.H.; Gilboa, E. Induction of Tumour Immunity by Targeted Inhibition of Nonsense-Mediated MRNA Decay. Nature 2010, 465, 227–230. [Google Scholar] [CrossRef] [PubMed]
- Quirico, L.; Orso, F.; Esposito, C.L.; Bertone, S.; Coppo, R.; Conti, L.; Catuogno, S.; Cavallo, F.; de Franciscis, V.; Taverna, D. Axl-148b Chimeric Aptamers Inhibit Breast Cancer and Melanoma Progression. Int. J. Biol. Sci. 2020, 16, 1238–1251. [Google Scholar] [CrossRef] [PubMed]
- Hao, Y.; Yang, J.; Liu, D.; Zhang, H.; Ou, T.; Xiao, L.; Chen, W. Construction of Aptamer-SiRNA Chimera and Glutamine Modified Carboxymethyl-β-Cyclodextrin Nanoparticles for the Combination Therapy against Lung Squamous Cell Carcinoma. Biomed. Pharmacother. 2024, 174, 116506. [Google Scholar] [CrossRef]
- Arnold, A.E.; Smith, L.J.; Beilhartz, G.L.; Bahlmann, L.C.; Jameson, E.; Melnyk, R.A.; Shoichet, M.S. Attenuated Diphtheria Toxin Mediates SiRNA Delivery. Sci. Adv. 2020, 6, 4848. [Google Scholar] [CrossRef]
- Dyer, P.D.R.; Shepherd, T.R.; Gollings, A.S.; Shorter, S.A.; Gorringe-Pattrick, M.A.M.; Tang, C.K.; Cattoz, B.N.; Baillie, L.; Griffiths, P.C.; Richardson, S.C.W. Disarmed Anthrax Toxin Delivers Antisense Oligonucleotides and SiRNA with High Efficiency and Low Toxicity. J. Control. Release 2015, 220, 316–328. [Google Scholar] [CrossRef]
- Zhao, S.; Karp, J.M.; Joshi, N. Toxin-Mediated SiRNA Delivery. Trends Pharmacol. Sci. 2020, 41, 511–513. [Google Scholar] [CrossRef]
- Yamada, T.; Peng, C.G.; Matsuda, S.; Addepalli, H.; Jayaprakash, K.N.; Alam, M.R.; Mills, K.; Maier, M.A.; Charisse, K.; Sekine, M.; et al. Versatile Site-Specific Conjugation of Small Molecules to SiRNA Using Click Chemistry. J. Org. Chem. 2011, 76, 1198–1211. [Google Scholar] [CrossRef]
- Salim, L.; Desaulniers, J.-P. To Conjugate or to Package? A Look at Targeted SiRNA Delivery Through Folate Receptors. Nucleic. Acid. Ther. 2021, 31, 21–38. [Google Scholar] [CrossRef]
- Abdelaal, A.M.; Kasinski, A.L. Ligand-Mediated Delivery of RNAi-Based Therapeutics for the Treatment of Oncological Diseases. NAR Cancer 2021, 3, 030. [Google Scholar] [CrossRef] [PubMed]
- Salim, L.; Islam, G.; Desaulniers, J.-P. Targeted Delivery and Enhanced Gene-Silencing Activity of Centrally Modified Folic Acid–SiRNA Conjugates. Nucleic. Acids. Res. 2020, 48, 75–85. [Google Scholar] [CrossRef] [PubMed]
- Gangopadhyay, S.; Nikam, R.R.; Gore, K.R. Folate Receptor-Mediated SiRNA Delivery: Recent Developments and Future Directions for RNAi Therapeutics. Nucleic. Acid. Ther. 2021, 31, 245–270. [Google Scholar] [CrossRef]
- Thomas, M.; Kularatne, S.A.; Qi, L.; Kleindl, P.; Leamon, C.P.; Hansen, M.J.; Low, P.S. Ligand-Targeted Delivery of Small Interfering RNAs to Malignant Cells and Tissues. Ann. N. Y. Acad. Sci. 2009, 1175, 32–39. [Google Scholar] [CrossRef] [PubMed]
- Boss, S.D.; Betzel, T.; Müller, C.; Fischer, C.R.; Haller, S.; Reber, J.; Groehn, V.; Schibli, R.; Ametamey, S.M. Comparative Studies of Three Pairs of α- and γ-Conjugated Folic Acid Derivatives Labeled with Fluorine-18. Bioconjug Chem. 2016, 27, 74–86. [Google Scholar] [CrossRef]
- Zhang, L.; Chan, J.M.; Gu, F.X.; Rhee, J.W.; Wang, A.Z.; Radovic-Moreno, A.F.; Alexis, F.; Langer, R.; Farokhzad, O.C. Self-Assembled Lipid-Polymer Hybrid Nanoparticles: A Robust Drug Delivery Platform. ACS Nano 2008, 2, 1696–1702. [Google Scholar] [CrossRef]
- Dohmen, C.; Fröhlich, T.; Lächelt, U.; Röhl, I.; Vornlocher, H.P.; Hadwiger, P.; Wagner, E. Defined Folate-PEG-SiRNA Conjugates for Receptor-Specific Gene Silencing. Mol. Ther. Nucleic. Acids. 2012, 1, e7. [Google Scholar] [CrossRef] [PubMed]
- Osborn, M.F.; Khvorova, A. Improving SiRNA Delivery in Vivo Through Lipid Conjugation. Nucleic. Acid. Ther. 2018, 28, 128–136. [Google Scholar] [CrossRef]
- Chen, M.; Chen, M.; He, J. Cancer Cell Membrane Cloaking Nanoparticles for Targeted Co-Delivery of Doxorubicin and PD-L1 SiRNA. Artif. Cells Nanomed. Biotechnol. 2019, 47, 1635–1641. [Google Scholar] [CrossRef]
- Zheng, Y.; Tai, W. Insight into the SiRNA Transmembrane Delivery—From Cholesterol Conjugating to Tagging. WIREs Nanomed. Nanobiotechnol. 2020, 12, e1606. [Google Scholar] [CrossRef]
- Petrova, N.S.; Chernikov, I.V.; Meschaninova, M.I.; Dovydenko, I.S.; Venyaminova, A.G.; Zenkova, M.A.; Vlassov, V.V.; Chernolovskaya, E.L. Carrier-Free Cellular Uptake and the Gene-Silencing Activity of the Lipophilic SiRNAs Is Strongly Affected by the Length of the Linker between SiRNA and Lipophilic Group. Nucleic. Acids. Res. 2012, 40, 2330–2344. [Google Scholar] [CrossRef] [PubMed]
- Chernikov, I.V.; Ponomareva, U.A.; Meschaninova, M.I.; Bachkova, I.K.; Vlassov, V.V.; Zenkova, M.A.; Chernolovskaya, E.L. Cholesterol Conjugates of Small Interfering RNA: Linkers and Patterns of Modification. Molecules 2024, 29, 786. [Google Scholar] [CrossRef] [PubMed]
- Shmushkovich, T.; Monopoli, K.R.; Homsy, D.; Leyfer, D.; Betancur-Boissel, M.; Khvorova, A.; Wolfson, A.D. Functional Features Defining the Efficacy of Cholesterol-Conjugated, Self-Deliverable, Chemically Modified SiRNAs. Nucleic. Acids. Res. 2018, 46, 10905–10916. [Google Scholar] [CrossRef]
- Alterman, J.F.; Hall, L.M.; Coles, A.H.; Hassler, M.R.; Didiot, M.C.; Chase, K.; Abraham, J.; Sottosanti, E.; Johnson, E.; Sapp, E.; et al. Hydrophobically Modified SiRNAs Silence Huntingtin MRNA in Primary Neurons and Mouse Brain. Mol. Ther. Nucleic. Acids. 2015, 4, e266. [Google Scholar] [CrossRef]
- Wada, S.; Yasuhara, H.; Wada, F.; Sawamura, M.; Waki, R.; Yamamoto, T.; Harada-Shiba, M.; Obika, S. Evaluation of the Effects of Chemically Different Linkers on Hepatic Accumulations, Cell Tropism and Gene Silencing Ability of Cholesterol-Conjugated Antisense Oligonucleotides. J. Control. Release 2016, 226, 57–65. [Google Scholar] [CrossRef]
- Tai, W.; Gao, X. Noncovalent Tagging of SiRNA with Steroids for Transmembrane Delivery. Biomaterials 2018, 178, 720–727. [Google Scholar] [CrossRef]
- Kim, J.B.; Lee, Y.M.; Ryu, J.; Lee, E.; Kim, W.J.; Keum, G.; Bang, E.K. Coordinative Amphiphiles as Tunable SiRNA Transporters. Bioconjug. Chem. 2016, 27, 1850–1856. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Li, F.; Xu, Y.; Zhang, W.; Hu, Y.; Fu, Y.; Xu, W.; Ge, S.; Fan, X.; Lu, L. Cholesterol Modification of SDF-1-Specific SiRNA Enables Therapeutic Targeting of Angiogenesis through Akt Pathway Inhibition. Exp. Eye. Res. 2019, 184, 64–71. [Google Scholar] [CrossRef]
- Biscans, A.; Ly, S.; McHugh, N.; Cooper, D.A.; Khvorova, A. Engineered Ionizable Lipid SiRNA Conjugates Enhance Endosomal Escape but Induce Toxicity in Vivo. J. Control. Release 2022, 349, 831–843. [Google Scholar] [CrossRef]
- Østergaard, M.E.; Jackson, M.; Low, A.; Chappell, A.E.; Lee, R.G.; Peralta, R.Q.; Yu, J.; Kinberger, G.A.; Dan, A.; Carty, R.; et al. Conjugation of Hydrophobic Moieties Enhances Potency of Antisense Oligonucleotides in the Muscle of Rodents and Non-Human Primates. Nucleic. Acids. Res. 2019, 47, 6045–6058. [Google Scholar] [CrossRef]
- Biscans, A.; Coles, A.; Echeverria, D.; Khvorova, A. The Valency of Fatty Acid Conjugates Impacts SiRNA Pharmacokinetics, Distribution, and Efficacy in Vivo. J. Control. Release 2019, 302, 116–125. [Google Scholar] [CrossRef]
- Biscans, A.; Caiazzi, J.; McHugh, N.; Hariharan, V.; Muhuri, M.; Khvorova, A. Docosanoic Acid Conjugation to SiRNA Enables Functional and Safe Delivery to Skeletal and Cardiac Muscles. Mol. Ther. 2021, 29, 1382–1394. [Google Scholar] [CrossRef] [PubMed]
- Alexander, P.; Kucera, G.; Pardee, T.S. Improving Nucleoside Analogs via Lipid Conjugation: Is Fatter Any Better? Crit. Rev. Oncol. Hematol. 2016, 100, 46–56. [Google Scholar] [CrossRef] [PubMed]
- Osborn, M.F.; Coles, A.H.; Biscans, A.; Haraszti, R.A.; Roux, L.; Davis, S.; Ly, S.; Echeverria, D.; Hassler, M.R.; Godinho, B.M.D.C.; et al. Hydrophobicity Drives the Systemic Distribution of Lipid-Conjugated SiRNAs via Lipid Transport Pathways. Nucleic. Acids. Res. 2019, 47, 1070–1081. [Google Scholar] [CrossRef] [PubMed]
- Tran, P.; Weldemichael, T.; Liu, Z.; Li, H. Delivery of Oligonucleotides: Efficiency with Lipid Conjugation and Clinical Outcome. Pharmaceutics 2022, 14, 342. [Google Scholar] [CrossRef]
- Ruvinov, E.; Kryukov, O.; Forti, E.; Korin, E.; Goldstein, M.; Cohen, S. Calcium–SiRNA Nanocomplexes: What Reversibility Is All About. J. Control. Release 2015, 203, 150–160. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Li, Z.; Zhao, X.; Keen, L.; Kong, X. Calcium Phosphate Nanoparticles-Based Systems for SiRNA Delivery. Regen. Biomater. 2016, 3, 187–195. [Google Scholar] [CrossRef] [PubMed]
- Oyer, H.M.; Sanders, C.M.; Kim, F.J. Small-Molecule Modulators of Sigma1 and Sigma2/TMEM97 in the Context of Cancer: Foundational Concepts and Emerging Themes. Front. Pharmacol. 2019, 10, 1141. [Google Scholar] [CrossRef]
- Alam, M.R.; Ming, X.; Nakagawa, O.; Jin, J.; Juliano, R.L. Covalent Conjugation of Oligonucleotides with Cell-Targeting Ligands. Bioorg. Med. Chem. 2013, 21, 6217–6223. [Google Scholar] [CrossRef]
- Qu, D.; Jiao, M.; Lin, H.; Tian, C.; Qu, G.; Xue, J.; Xue, L.; Ju, C.; Zhang, C. Anisamide-Functionalized PH-Responsive Amphiphilic Chitosan-Based Paclitaxel Micelles for Sigma-1 Receptor Targeted Prostate Cancer Treatment. Carbohydr. Polym. 2020, 229, 115498. [Google Scholar] [CrossRef]
- Yao, W.; Liu, C.; Wang, N.; Zhou, H.; Chen, H.; Qiao, W. Anisamide-Modified Dual-Responsive Drug Delivery System with MRI Capacity for Cancer Targeting Therapy. J. Mol. Liq. 2021, 340, 116889. [Google Scholar] [CrossRef]
- Springer, A.D.; Dowdy, S.F. GalNAc-SiRNA Conjugates: Leading the Way for Delivery of RNAi Therapeutics. Nucleic. Acid. Ther. 2018, 28, 109–118. [Google Scholar] [CrossRef]
- Biessen, E.A.L.; Van Berkel, T.J.C. N-Acetyl Galactosamine Targeting: Paving the Way for Clinical Application of Nucleotide Medicines in Cardiovascular Diseases. Arterioscler. Thromb. Vasc. Biol. 2021, 41, 2855–2865. [Google Scholar] [CrossRef]
- Willibald, J.; Harder, J.; Sparrer, K.; Conzelmann, K.-K.; Carell, T. Click-Modified Anandamide SiRNA Enables Delivery and Gene Silencing in Neuronal and Immune Cells. J. Am. Chem. Soc. 2012, 134, 12330–12333. [Google Scholar] [CrossRef] [PubMed]
- Nishina, K.; Unno, T.; Uno, Y.; Kubodera, T.; Kanouchi, T.; Mizusawa, H.; Yokota, T. Efficient In Vivo Delivery of SiRNA to the Liver by Conjugation of α-Tocopherol. Mol. Ther. 2008, 16, 734–740. [Google Scholar] [CrossRef]
- Tang, S.; Kapoor, E.; Ding, L.; Yu, A.; Tang, W.; Hang, Y.; Smith, L.M.; Sil, D.; Oupický, D. Effect of Tocopherol Conjugation on Polycation-Mediated SiRNA Delivery to Orthotopic Pancreatic Tumors. Biomater. Adv. 2023, 145, 213236. [Google Scholar] [CrossRef]
- Yagublu, V.; Karimova, A.; Hajibabazadeh, J.; Reissfelder, C.; Muradov, M.; Bellucci, S.; Allahverdiyev, A. Overview of Physicochemical Properties of Nanoparticles as Drug Carriers for Targeted Cancer Therapy. J. Funct. Biomater. 2022, 13, 196. [Google Scholar] [CrossRef]
- Yuan, Z.; Yan, R.; Fu, Z.; Wu, T.; Ren, C. Impact of Physicochemical Properties on Biological Effects of Lipid Nanoparticles: Are They Completely Safe. Sci. Total Environ. 2024, 927, 172240. [Google Scholar] [CrossRef]
- Hui, Y.; Yi, X.; Hou, F.; Wibowo, D.; Zhang, F.; Zhao, D.; Gao, H.; Zhao, C.-X. Role of Nanoparticle Mechanical Properties in Cancer Drug Delivery. ACS Nano 2019, 13, 7410–7424. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Yuk, S.A.; Dieterly, A.M.; Kwon, S.; Park, J.; Meng, F.; Gadalla, H.H.; Cadena, M.J.; Lyle, L.T.; Yeo, Y. Nanosac, a Noncationic and Soft Polyphenol Nanocapsule, Enables Systemic Delivery of SiRNA to Solid Tumors. ACS Nano 2021, 15, 4576–4593. [Google Scholar] [CrossRef]
- Zein, R.; Sharrouf, W.; Selting, K. Physical Properties of Nanoparticles That Result in Improved Cancer Targeting. J. Oncol. 2020, 2020, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Li, L.; Li, L.; Song, F.; Song, F. Interplay of Nanoparticle Properties during Endocytosis. Crystals 2021, 11, 728. [Google Scholar] [CrossRef]
- Palmieri, V.; Caracciolo, G. Tuning the Immune System by Nanoparticle–Biomolecular Corona. Nanoscale Adv. 2022, 4, 3300–3308. [Google Scholar] [CrossRef]
- Lu, Z.-R.; Sun, D. Mechanism of PH-Sensitive Amphiphilic Endosomal Escape of Ionizable Lipid Nanoparticles for Cytosolic Nucleic Acid Delivery. Pharm. Res. 2025, 42, 1065–1077. [Google Scholar] [CrossRef]
- Oyewumi, M.O.; Kumar, A.; Cui, Z. Nano-Microparticles as Immune Adjuvants: Correlating Particle Sizes and the Resultant Immune Responses. Expert. Rev. Vaccines 2010, 9, 1095–1107. [Google Scholar] [CrossRef]
- Morales-Becerril, A.; Aranda-Lara, L.; Isaac-Olivé, K.; Ocampo-García, B.E.; Morales-ávila, E. Nanocarriers for delivery of sirna as gene silencing mediator. Excli J. 2022, 21, 1028–1052. [Google Scholar] [PubMed]
- Zhang, T.; Yin, H.; Li, Y.; Yang, H.; Ge, K.; Zhang, J.; Yuan, Q.; Dai, X.; Naeem, A.; Weng, Y.; et al. Optimized Lipid Nanoparticles (LNPs) for Organ-Selective Nucleic Acids Delivery in Vivo. iScience 2024, 27, 109804. [Google Scholar] [CrossRef] [PubMed]
- Sinani, G.; Durgun, M.E.; Cevher, E.; Özsoy, Y. Polymeric-Micelle-Based Delivery Systems for Nucleic Acids. Pharmaceutics 2023, 15, 2021. [Google Scholar] [CrossRef] [PubMed]
- Serrano-Sevilla, I.; Artiga, Á.; Mitchell, S.G.; De Matteis, L.; de la Fuente, J.M. Natural Polysaccharides for SiRNA Delivery: Nanocarriers Based on Chitosan, Hyaluronic Acid, and Their Derivatives. Molecules 2019, 24, 2570. [Google Scholar] [CrossRef]
- Cao, Y.; Tan, Y.F.; Wong, Y.S.; Liew, M.W.J.; Venkatraman, S. Recent Advances in Chitosan-Based Carriers for Gene Delivery. Mar. Drugs 2019, 17, 381. [Google Scholar] [CrossRef]
- Oh, Y.-K.; Park, T.G. SiRNA Delivery Systems for Cancer Treatment. Adv. Drug Deliv. Rev. 2009, 61, 850–862. [Google Scholar] [CrossRef]
- Singha, K.; Namgung, R.; Kim, W.J. Polymers in Small-Interfering RNA Delivery. Nucleic. Acid. Ther. 2011, 21, 133–147. [Google Scholar] [CrossRef]
- Ewe, A.; Höbel, S.; Heine, C.; Merz, L.; Kallendrusch, S.; Bechmann, I.; Merz, F.; Franke, H.; Aigner, A. Optimized Polyethylenimine (PEI)-Based Nanoparticles for SiRNA Delivery, Analyzed in Vitro and in an Ex Vivo Tumor Tissue Slice Culture Model. Drug Deliv. Transl. Res. 2017, 7, 206–216. [Google Scholar] [CrossRef]
- Karlsson, J.; Rhodes, K.R.; Green, J.J.; Tzeng, S.Y. Poly(Beta-Amino Ester)s as Gene Delivery Vehicles: Challenges and Opportunities. Expert. Opin. Drug Deliv. 2020, 17, 1395–1410. [Google Scholar] [CrossRef]
- Zielińska, A.; Carreiró, F.; Oliveira, A.M.; Neves, A.; Pires, B.; Venkatesh, D.N.; Durazzo, A.; Lucarini, M.; Eder, P.; Silva, A.M.; et al. Polymeric Nanoparticles: Production, Characterization, Toxicology and Ecotoxicology. Molecules 2020, 25, 3731. [Google Scholar] [CrossRef]
- Fus-Kujawa, A.; Teper, P.; Botor, M.; Klarzyńska, K.; Sieroń, Ł.; Verbelen, B.; Smet, M.; Sieroń, A.L.; Mendrek, B.; Kowalczuk, A. Functional Star Polymers as Reagents for Efficient Nucleic Acids Delivery into HT-1080 Cells. Int. J. Polym. Mater. Polym. Biomater. 2021, 70, 356–370. [Google Scholar] [CrossRef]
- Nguyen, J.; Steele, T.W.J.; Merkel, O.; Reul, R.; Kissel, T. Fast Degrading Polyesters as SiRNA Nano-Carriers for Pulmonary Gene Therapy. J. Control. Release 2008, 132, 243–251. [Google Scholar] [CrossRef]
- Zhu, J.; Qiao, M.; Wang, Q.; Ye, Y.; Ba, S.; Ma, J.; Hu, H.; Zhao, X.; Chen, D. Dual-Responsive Polyplexes with Enhanced Disassembly and Endosomal Escape for Efficient Delivery of SiRNA. Biomaterials 2018, 162, 47–59. [Google Scholar] [CrossRef] [PubMed]
- Bholakant, R.; Qian, H.; Zhang, J.; Huang, X.; Huang, D.; Feijen, J.; Zhong, Y.; Chen, W. Recent Advances of Polycationic SiRNA Vectors for Cancer Therapy. Biomacromolecules 2020, 21, 2966–2982. [Google Scholar] [CrossRef]
- Yao, Z.; Liu, T.; Wang, J.; Fu, Y.; Zhao, J.; Wang, X.; Li, Y.; Yang, X.; He, Z. Targeted Delivery Systems of SiRNA Based on Ionizable Lipid Nanoparticles and Cationic Polymer Vectors. Biotechnol. Adv. 2025, 81, 108546. [Google Scholar] [CrossRef]
- Hibbitts, A.J.; Ramsey, J.M.; Barlow, J.; Macloughlin, R.; Cryan, S.A. In Vitro and in Vivo Assessment of Pegylated Pei for Anti-Il-8/Cxcl-1 Sirna Delivery to the Lungs. Nanomaterials 2020, 10, 1248. [Google Scholar] [CrossRef]
- Cho, S.K.; Dang, C.; Wang, X.; Ragan, R.; Kwon, Y.J. Mixing-Sequence-Dependent Nucleic Acid Complexation and Gene Transfer Efficiency by Polyethylenimine. Biomater. Sci. 2015, 3, 1124–1133. [Google Scholar] [CrossRef]
- Kazemi Oskuee, R.; Dabbaghi, M.; Gholami, L.; Taheri-Bojd, S.; Balali-Mood, M.; Mousavi, S.H.; Malaekeh-Nikouei, B. Investigating the Influence of Polyplex Size on Toxicity Properties of Polyethylenimine Mediated Gene Delivery. Life Sci. 2018, 197, 101–108. [Google Scholar] [CrossRef]
- Höbel, S.; Aigner, A. Polyethylenimines for SiRNA and MiRNA Delivery in Vivo. WIREs Nanomed. Nanobiotechnol. 2013, 5, 484–501. [Google Scholar] [CrossRef]
- Ewe, A.; Panchal, O.; Pinnapireddy, S.R.; Bakowsky, U.; Przybylski, S.; Temme, A.; Aigner, A. Liposome-Polyethylenimine Complexes (DPPC-PEI Lipopolyplexes) for Therapeutic SiRNA Delivery in Vivo. Nanomedicine 2017, 13, 209–218. [Google Scholar] [CrossRef] [PubMed]
- Parhamifar, L.; Larsen, A.K.; Hunter, A.C.; Andresen, T.L.; Moghimi, S.M. Polycation Cytotoxicity: A Delicate Matter for Nucleic Acid Therapy—Focus on Polyethylenimine. Soft Matter. 2010, 6, 4001. [Google Scholar] [CrossRef]
- Hall, A.; Lächelt, U.; Bartek, J.; Wagner, E.; Moghimi, S.M. Polyplex Evolution: Understanding Biology, Optimizing Performance. Mol. Ther. 2017, 25, 1476–1490. [Google Scholar] [CrossRef]
- Liu, S.; Huang, W.; Jin, M.J.; Fan, B.; Xia, G.M.; Gao, Z.G. Inhibition of Murine Breast Cancer Growth and Metastasis by Survivin-Targeted SiRNA Using Disulfide Cross-Linked Linear PEI. Eur. J. Pharm. Sci. 2016, 82, 171–182. [Google Scholar] [CrossRef]
- Zhang, Q.Y.; Ho, P.Y.; Tu, M.J.; Jilek, J.L.; Chen, Q.X.; Zeng, S.; Yu, A.M. Lipidation of Polyethylenimine-Based Polyplex Increases Serum Stability of Bioengineered RNAi Agents and Offers More Consistent Tumoral Gene Knockdown in Vivo. Int. J. Pharm. 2018, 547, 537–544. [Google Scholar] [CrossRef]
- Merkel, O.M.; Beyerle, A.; Librizzi, D.; Pfestroff, A.; Behr, T.M.; Sproat, B.; Barth, P.J.; Kissel, T. Nonviral SiRNA Delivery to the Lung: Investigation of PEG−PEI Polyplexes and Their In Vivo Performance. Mol. Pharm. 2009, 6, 1246–1260. [Google Scholar] [CrossRef]
- Zhupanyn, P.; Ewe, A.; Büch, T.; Malek, A.; Rademacher, P.; Müller, C.; Reinert, A.; Jaimes, Y.; Aigner, A. Extracellular Vesicle (ECV)-Modified Polyethylenimine (PEI) Complexes for Enhanced SiRNA Delivery in Vitro and in Vivo. J. Control. Release 2020, 319, 63–76. [Google Scholar] [CrossRef] [PubMed]
- Chuan, D.; Jin, T.; Fan, R.; Zhou, L.; Guo, G. Chitosan for Gene Delivery: Methods for Improvement and Applications. Adv. Colloid. Interface Sci. 2019, 268, 25–38. [Google Scholar] [CrossRef] [PubMed]
- Al-Absi, M.Y.; Caprifico, A.E.; Calabrese, G. Chitosan and Its Structural Modifications for SiRNA Delivery. Adv. Pharm. Bull. 2023, 13, 275–282. [Google Scholar] [CrossRef]
- Karayianni, M.; Sentoukas, T.; Skandalis, A.; Pippa, N.; Pispas, S. Chitosan-Based Nanoparticles for Nucleic Acid Delivery: Technological Aspects, Applications, and Future Perspectives. Pharmaceutics 2023, 15, 1849. [Google Scholar] [CrossRef]
- Şalva, E.; Özkan, N.; Kabasakal, L.; Turan, S.Ö.; Akbuğa, J. The Effect of Chitosan Complexes on Biodistribution of SiRNA. Clin. Exp. Health Sci. 2011, 1, 1. [Google Scholar]
- Holzerny, P.; Ajdini, B.; Heusermann, W.; Bruno, K.; Schuleit, M.; Meinel, L.; Keller, M. Biophysical Properties of Chitosan/SiRNA Polyplexes: Profiling the Polymer/SiRNA Interactions and Bioactivity. J. Control. Release 2012, 157, 297–304. [Google Scholar] [CrossRef]
- Cao, Y.; Tan, Y.F.; Wong, Y.S.; Aminuddin, M.; Ramya, B.; Liew, M.W.J.; Liu, J.; Venkatraman, S.S. Designing SiRNA/Chitosan-Methacrylate Complex Nanolipogel for Prolonged Gene Silencing Effects. Sci. Rep. 2022, 12, 5335. [Google Scholar] [CrossRef]
- Liu, X.; Ma, L.; Qin, W.; Gao, C. Effect of N/P Ratios on Physicochemical Stability, Cellular Association, and Gene Silencing Efficiency for Trimethyl Chitosan/Small Interfering RNA Complexes. J. Bioact. Compat. Polym. 2013, 28, 590–604. [Google Scholar] [CrossRef]
- Veilleux, D.; Gopalakrishna Panicker, R.K.; Chevrier, A.; Biniecki, K.; Lavertu, M.; Buschmann, M.D. Lyophilisation and Concentration of Chitosan/SiRNA Polyplexes: Influence of Buffer Composition, Oligonucleotide Sequence, and Hyaluronic Acid Coating. J. Colloid. Interface Sci. 2018, 512, 335–345. [Google Scholar] [CrossRef]
- Grząbka-Zasadzińska, A.; Amietszajew, T.; Borysiak, S. Thermal and Mechanical Properties of Chitosan Nanocomposites with Cellulose Modified in Ionic Liquids. J. Therm. Anal. Calorim. 2017, 130, 143–154. [Google Scholar] [CrossRef]
- Prakash, S.; Tomaro-Duchesneau, C.; Saha, S.; Kahouli; Malhotra, M. Development and Characterization of Chitosan-PEG-TAT Nanoparticles for the Intracellular Delivery of SiRNA. Int. J. Nanomed. 2013, 8, 2041. [Google Scholar] [CrossRef] [PubMed]
- Sun, P.; Huang, W.; Kang, L.; Jin, M.; Fan, B.; Jin, H.; Wang, Q.; Gao, Z. SiRNA-Loaded Poly(Histidine-Arginine)6-Modified Chitosan Nanoparticle with Enhanced Cell-Penetrating and Endosomal Escape Capacities for Suppressing Breast Tumor Metastasis. Int. J. Nanomed. 2017, 12, 3221–3234. [Google Scholar] [CrossRef] [PubMed]
- Muddineti, O.S.; Shah, A.; Rompicharla, S.V.K.; Ghosh, B.; Biswas, S. Cholesterol-Grafted Chitosan Micelles as a Nanocarrier System for Drug-SiRNA Co-Delivery to the Lung Cancer Cells. Int. J. Biol. Macromol. 2018, 118, 857–863. [Google Scholar] [CrossRef] [PubMed]
- Paredes-Hernández, U.; Aguilar-Peña, L.V.; Isaac-Olivé, K.; Ocampo-García, B.; Contreras, I.; Estrada, J.A.; Izquierdo, G.; Morales-Avila, E.; Aranda-Lara, L. Enhancing Photodynamic and Radionuclide Therapy by Small Interfering RNA (SiRNA)-RAD51 Transfection via Self-Emulsifying Delivery Systems (SNEDDS). Cytotherapy 2025, 27, 66–77. [Google Scholar] [CrossRef]
- Green, J.J.; Shi, J.; Chiu, E.; Leshchiner, E.S.; Langer, R.; Anderson, D.G. Biodegradable Polymeric Vectors for Gene Delivery to Human Endothelial Cells. Bioconjug. Chem. 2006, 17, 1162–1169. [Google Scholar] [CrossRef]
- Kozielski, K.L.; Tzeng, S.Y.; Green, J.J. A Bioreducible Linear Poly(β-Amino Ester) for SiRNA Delivery. Chem. Commun. 2013, 49, 5319–5321. [Google Scholar] [CrossRef]
- Zhang, J.; Cai, X.; Dou, R.; Guo, C.; Tang, J.; Hu, Y.; Chen, H.; Chen, J. Poly(β-Amino Ester)s-Based Nanovehicles: Structural Regulation and Gene Delivery. Mol. Ther. Nucleic. Acids. 2023, 32, 568–581. [Google Scholar] [CrossRef]
- Chou, J.J.; Berger, A.G.; Jalili-Firoozinezhad, S.; Hammond, P.T. A Design Approach for Layer-by-Layer Surface-Mediated SiRNA Delivery. Acta Biomater. 2021, 135, 331–341. [Google Scholar] [CrossRef]
- Rui, Y.; Wilson, D.R.; Choi, J.; Varanasi, M.; Sanders, K.; Karlsson, J.; Lim, M.; Green, J.J. Carboxylated Branched Poly(β-Amino Ester) Nanoparticles Enable Robust Cytosolic Protein Delivery and CRISPR-Cas9 Gene Editing. Sci. Adv. 2019, 5, 3255. [Google Scholar] [CrossRef] [PubMed]
- Berger, A.G.; DeLorenzo, C.; Vo, C.; Kaskow, J.A.; Nabar, N.; Hammond, P.T. Poly(β-Aminoester) Physicochemical Properties Govern the Delivery of SiRNA from Electrostatically Assembled Coatings. Biomacromolecules 2024, 25, 2934–2952. [Google Scholar] [CrossRef]
- Mohan, T.; Kleinschek, K.S.; Kargl, R. Polysaccharide Peptide Conjugates: Chemistry, Properties and Applications. Carbohydr. Polym. 2022, 280, 118875. [Google Scholar] [CrossRef]
- Zhou, J.; Patel, T.R.; Fu, M.; Bertram, J.P.; Saltzman, W.M. Octa-Functional PLGA Nanoparticles for Targeted and Efficient SiRNA Delivery to Tumors. Biomaterials 2012, 33, 583–591. [Google Scholar] [CrossRef]
- Abedi-Gaballu, F.; Dehghan, G.; Ghaffari, M.; Yekta, R.; Abbaspour-Ravasjani, S.; Baradaran, B.; Ezzati Nazhad Dolatabadi, J.; Hamblin, M.R. PAMAM Dendrimers as Efficient Drug and Gene Delivery Nanosystems for Cancer Therapy. Appl. Mater. Today 2018, 12, 177–190. [Google Scholar] [CrossRef] [PubMed]
- Kulkarni, J.A.; Cullis, P.R.; Van Der Meel, R. Lipid Nanoparticles Enabling Gene Therapies: From Concepts to Clinical Utility. Nucleic. Acid. Ther. 2018, 28, 146–157. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Zhang, H.; Li, G.; Su, J.; Wei, Y.; Xu, C. Lipid Nanovehicles Overcome Barriers to Systemic RNA Delivery: Lipid Components, Fabrication Methods, and Rational Design. Acta. Pharm. Sin. B 2024, 14, 579–601. [Google Scholar] [CrossRef]
- Kotelianski, V.; Zatsepin, T.; Kotelevtsev, Y. Lipid Nanoparticles for Targeted SiRNA Delivery—Going from Bench to Bedside. Int. J. Nanomed. 2016, 11, 3077–3086. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Jiang, Y.; He, Y.; Boucetta, H.; Wu, J.; Chen, Z.; He, W. Lipid Carriers for MRNA Delivery. Acta Pharm. Sin. B 2023, 13, 4105–4126. [Google Scholar] [CrossRef]
- Ponti, F.; Campolungo, M.; Melchiori, C.; Bono, N.; Candiani, G. Cationic Lipids for Gene Delivery: Many Players, One Goal. Chem. Phys. Lipids 2021, 235, 105032. [Google Scholar] [CrossRef]
- Tenchov, R.; Bird, R.; Curtze, A.E.; Zhou, Q. Lipid Nanoparticles─From Liposomes to MRNA Vaccine Delivery, a Landscape of Research Diversity and Advancement. ACS Nano 2021, 15, 16982–17015. [Google Scholar] [CrossRef]
- Kang, M.; Kim, H.; Leal, C. Self-Organization of Nucleic Acids in Lipid Constructs. Curr. Opin. Colloid Interface Sci. 2016, 26, 58–65. [Google Scholar] [CrossRef]
- Han, X.; Mitchell, M.J.; Nie, G. Nanomaterials for Therapeutic RNA Delivery. Matter 2020, 3, 1948–1975. [Google Scholar] [CrossRef]
- Zheng, L.I.; Bandara, S.R.; Tan, Z.I.; Leal, C. Lipid Nanoparticle Topology Regulates Endosomal Escape and Delivery of RNA to the Cytoplasm. Proc. Natl. Acad. Sci. USA 2023, 120, e2303567120. [Google Scholar] [CrossRef]
- Hattori, Y.; Arai, S.; Kikuchi, T.; Hamada, M.; Okamoto, R.; Machida, Y.; Kawano, K. Optimization of SiRNA Delivery Method into the Liver by Sequential Injection of Polyglutamic Acid and Cationic Lipoplex. Pharmacol. Pharm. 2015, 6, 302–310. [Google Scholar] [CrossRef]
- Mendonça, M.C.P.; Kont, A.; Kowalski, P.S.; O’Driscoll, C.M. Design of Lipid-Based Nanoparticles for Delivery of Therapeutic Nucleic Acids. Drug Discov. Today 2023, 28, 103505. [Google Scholar] [CrossRef] [PubMed]
- Mihailescu, M.; Worcester, D.L.; Carroll, C.L.; Chamberlin, A.R.; White, S.H. DOTAP: Structure, Hydration, and the Counterion Effect. Biophys. J. 2023, 122, 1086–1093. [Google Scholar] [CrossRef]
- Lechanteur, A.; Sanna, V.; Duchemin, A.; Evrard, B.; Mottet, D.; Piel, G. Cationic Liposomes Carrying SiRNA: Impact of Lipid Composition on Physicochemical Properties, Cytotoxicity and Endosomal Escape. Nanomaterials 2018, 8, 270. [Google Scholar] [CrossRef] [PubMed]
- Ni, R.; Feng, R.; Chau, Y. Synthetic Approaches for Nucleic Acid Delivery: Choosing the Right Carriers. Life 2019, 9, 59. [Google Scholar] [CrossRef] [PubMed]
- Sharma, D.; Arora, S.; Singh, J.; Layek, B. A Review of the Tortuous Path of Nonviral Gene Delivery and Recent Progress. Int. J. Biol. Macromol. 2021, 183, 2055–2073. [Google Scholar] [CrossRef]
- Kurakula, H.; Vaishnavi, S.; Sharif, M.Y.; Ellipilli, S. Emergence of Small Interfering RNA-Based Gene Drugs for Various Diseases. ACS Omega 2023, 8, 20234–20250. [Google Scholar] [CrossRef]
- Mai, Y.; Guo, J.; Zhao, Y.; Ma, S.; Hou, Y.; Yang, J. Intranasal Delivery of Cationic Liposome-Protamine Complex MRNA Vaccine Elicits Effective Anti-Tumor Immunity. Cell Immunol. 2020, 354, 104143. [Google Scholar] [CrossRef]
- Habrant, D.; Peuziat, P.; Colombani, T.; Dallet, L.; Gehin, J.; Goudeau, E.; Evrard, B.; Lambert, O.; Haudebourg, T.; Pitard, B. Design of Ionizable Lipids To Overcome the Limiting Step of Endosomal Escape: Application in the Intracellular Delivery of MRNA, DNA, and SiRNA. J. Med. Chem. 2016, 59, 3046–3062. [Google Scholar] [CrossRef]
- Kulkarni, J.A.; Darjuan, M.M.; Mercer, J.E.; Chen, S.; Van Der Meel, R.; Thewalt, J.L.; Tam, Y.Y.C.; Cullis, P.R. On the Formation and Morphology of Lipid Nanoparticles Containing Ionizable Cationic Lipids and SiRNA. ACS Nano 2018, 12, 4787–4795. [Google Scholar] [CrossRef]
- Fedorovskiy, A.G.; Antropov, D.N.; Dome, A.S.; Puchkov, P.A.; Makarova, D.M.; Konopleva, M.V.; Matveeva, A.M.; Panova, E.A.; Shmendel, E.V.; Maslov, M.A.; et al. Novel Efficient Lipid-Based Delivery Systems Enable a Delayed Uptake and Sustained Expression of MRNA in Human Cells and Mouse Tissues. Pharmaceutics 2024, 16, 684. [Google Scholar] [CrossRef]
- Pozzi, D.; Marchini, C.; Cardarelli, F.; Amenitsch, H.; Garulli, C.; Bifone, A.; Caracciolo, G. Transfection Efficiency Boost of Cholesterol-Containing Lipoplexes. Biochim. Biophys. Acta. Biomembr. 2012, 1818, 2335–2343. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Ahn, H.J. PEGylated DC-Chol/DOPE Cationic Liposomes Containing KSP SiRNA as a Systemic SiRNA Delivery Carrier for Ovarian Cancer Therapy. Biochem. Biophys. Res. Commun. 2018, 503, 1716–1722. [Google Scholar] [CrossRef]
- Semple, S.C.; Klimuk, S.K.; Harasym, T.O.; Dos Santos, N.; Ansell, S.M.; Wong, K.F.; Maurer, N.; Stark, H.; Cullis, P.R.; Hope, M.J.; et al. Efficient Encapsulation of Antisense Oligonucleotides in Lipid Vesicles Using Ionizable Aminolipids: Formation of Novel Small Multilamellar Vesicle Structures. Biochim. Biophys. Acta (BBA) Biomembr. 2001, 1510, 152–166. [Google Scholar] [CrossRef]
- Lin, P.J.C.; Tam, Y.Y.C.; Hafez, I.; Sandhu, A.; Chen, S.; Ciufolini, M.A.; Nabi, I.R.; Cullis, P.R. Influence of Cationic Lipid Composition on Uptake and Intracellular Processing of Lipid Nanoparticle Formulations of SiRNA. Nanomedicine 2013, 9, 233–246. [Google Scholar] [CrossRef]
- Dong, Y.; Love, K.T.; Dorkin, J.R.; Sirirungruang, S.; Zhang, Y.; Chen, D.; Bogorad, R.L.; Yin, H.; Chen, Y.; Vegas, A.J.; et al. Lipopeptide Nanoparticles for Potent and Selective SiRNA Delivery in Rodents and Nonhuman Primates. Proc. Natl. Acad. Sci. USA 2014, 111, 3955–3960. [Google Scholar] [CrossRef]
- Maier, M.A.; Jayaraman, M.; Matsuda, S.; Liu, J.; Barros, S.; Querbes, W.; Tam, Y.K.; Ansell, S.M.; Kumar, V.; Qin, J.; et al. Biodegradable Lipids Enabling Rapidly Eliminated Lipid Nanoparticles for Systemic Delivery of RNAi Therapeutics. Mol. Ther. 2013, 21, 1570–1578. [Google Scholar] [CrossRef]
- Buyens, K.; De Smedt, S.C.; Braeckmans, K.; Demeester, J.; Peeters, L.; van Grunsven, L.A.; de Mollerat du Jeu, X.; Sawant, R.; Torchilin, V.; Farkasova, K.; et al. Liposome Based Systems for Systemic SiRNA Delivery: Stability in Blood Sets the Requirements for Optimal Carrier Design. J. Control. Release 2012, 158, 362–370. [Google Scholar] [CrossRef]
- Cox, A.; Lim, S.A.; Chung, E.J. Strategies to Deliver RNA by Nanoparticles for Therapeutic Potential. Mol. Aspects Med. 2022, 83, 100991. [Google Scholar] [CrossRef]
- John, R.; Monpara, J.; Swaminathan, S.; Kalhapure, R. Chemistry and Art of Developing Lipid Nanoparticles for Biologics Delivery: Focus on Development and Scale-Up. Pharmaceutics 2024, 16, 131. [Google Scholar] [CrossRef] [PubMed]
- Nsairat, H.; Khater, D.; Sayed, U.; Odeh, F.; Al Bawab, A.; Alshaer, W. Liposomes: Structure, Composition, Types, and Clinical Applications. Heliyon 2022, 8, e09394. [Google Scholar] [CrossRef] [PubMed]
- Ong, S.G.M.; Ming, L.C.; Lee, K.S.; Yuen, K.H. Influence of the Encapsulation Efficiency and Size of Liposome on the Oral Bioavailability of Griseofulvin-Loaded Liposomes. Pharmaceutics 2016, 8, 25. [Google Scholar] [CrossRef]
- Fan, Y.; Marioli, M.; Zhang, K. Analytical Characterization of Liposomes and Other Lipid Nanoparticles for Drug Delivery. J. Pharm. Biomed. Anal. 2021, 192, 113642. [Google Scholar] [CrossRef] [PubMed]
- Rengaswamy, V.; Zimmer, D.; Süss, R.; Rössler, J. RGD Liposome-Protamine-SiRNA (LPR) Nanoparticles Targeting PAX3-FOXO1 for Alveolar Rhabdomyosarcoma Therapy. J. Control. Release 2016, 235, 319–327. [Google Scholar] [CrossRef]
- Jovanović, A.A.; Balanč, B.D.; Ota, A.; Ahlin Grabnar, P.; Djordjević, V.B.; Šavikin, K.P.; Bugarski, B.M.; Nedović, V.A.; Poklar Ulrih, N. Comparative Effects of Cholesterol and β-Sitosterol on the Liposome Membrane Characteristics. Eur. J. Lipid Sci. Technol. 2018, 120, 1800039. [Google Scholar] [CrossRef]
- Nosova, A.S.; Koloskova, O.O.; Nikonova, A.A.; Simonova, V.A.; Smirnov, V.V.; Kudlay, D.; Khaitov, M.R. Diversity of PEGylation Methods of Liposomes and Their Influence on RNA Delivery. Medchemcomm 2019, 10, 369–377. [Google Scholar] [CrossRef]
- Kulkarni, J.A.; Witzigmann, D.; Chen, S.; Cullis, P.R.; Van Der Meel, R. Lipid Nanoparticle Technology for Clinical Translation of SiRNA Therapeutics. Acc. Chem. Res. 2019, 52, 2435–2444. [Google Scholar] [CrossRef]
- Noble, G.T.; Stefanick, J.F.; Ashley, J.D.; Kiziltepe, T.; Bilgicer, B. Ligand-Targeted Liposome Design: Challenges and Fundamental Considerations. Trends Biotechnol. 2014, 32, 32–45. [Google Scholar] [CrossRef]
- Aranda-Lara, L.; Morales-Avila, E.; Luna-Gutiérrez, M.A.; Olivé-Alvarez, E.; Isaac-Olivé, K. Radiolabeled Liposomes and Lipoproteins as Lipidic Nanoparticles for Imaging and Therapy. Chem. Phys. Lipids 2020, 230, 104934. [Google Scholar] [CrossRef]
- Liang, Z.; Du, L.; Zhang, E.; Zhao, Y.; Wang, W.; Ma, P.; Dai, M.; Zhao, Q.; Xu, H.; Zhang, S.; et al. Targeted-Delivery of SiRNA via a Polypeptide-Modified Liposome for the Treatment of Gp96 over-Expressed Breast Cancer. Mater. Sci. Eng. C 2021, 121, 111847. [Google Scholar] [CrossRef]
- Belletti, D.; Tonelli, M.; Forni, F.; Tosi, G.; Vandelli, M.A.; Ruozi, B. AFM and TEM Characterization of SiRNAs Lipoplexes: A Combinatory Tools to Predict the Efficacy of Complexation. Colloids Surf. A Physicochem. Eng. Asp. 2013, 436, 459–466. [Google Scholar] [CrossRef]
- Park, J.; Park, J.; Pei, Y.; Xu, J.; Yeo, Y. Pharmacokinetics and Biodistribution of Recently-Developed SiRNA Nanomedicines. Adv. Drug Deliv. Rev. 2016, 104, 93–109. [Google Scholar] [CrossRef] [PubMed]
- Chono, S.; Li, S.D.; Conwell, C.C.; Huang, L. An Efficient and Low Immunostimulatory Nanoparticle Formulation for Systemic SiRNA Delivery to the Tumor. J. Control. Release 2008, 131, 64–69. [Google Scholar] [CrossRef]
- Buyens, K.; Demeester, J.; De Smedt, S.S.; Sanders, N.N. Elucidating the Encapsulation of Short Interfering RNA in PEGylated Cationic Liposomes. Langmuir 2009, 25, 4886–4891. [Google Scholar] [CrossRef]
- Chen, S.; Tam, Y.Y.C.; Lin, P.J.C.; Sung, M.M.H.; Tam, Y.K.; Cullis, P.R. Influence of Particle Size on the in Vivo Potency of Lipid Nanoparticle Formulations of SiRNA. J. Control. Release 2016, 235, 236–244. [Google Scholar] [CrossRef] [PubMed]
- Mokhtarieh, A.A.; Lee, J.; Kim, S.; Lee, M.K. Preparation of SiRNA Encapsulated Nanoliposomes Suitable for SiRNA Delivery by Simply Discontinuous Mixing. Biochim. Biophys. Acta Biomembr. 2018, 1860, 1318–1325. [Google Scholar] [CrossRef]
- Kim, H.K.; Davaa, E.; Myung, C.S.; Park, J.S. Enhanced SiRNA Delivery Using Cationic Liposomes with New Polyarginine-Conjugated PEG-Lipid. Int. J. Pharm. 2010, 392, 141–147. [Google Scholar] [CrossRef]
- Vader, P.; Crielaard, B.J.; Van Dommelen, S.M.; Van Der Meel, R.; Storm, G.; Schiffelers, R.M. Targeted Delivery of Small Interfering RNA to Angiogenic Endothelial Cells with Liposome-Polycation-DNA Particles. J. Control. Release 2012, 160, 211–216. [Google Scholar] [CrossRef]
- Grijalvo, S.; Alagia, A.; Jorge, A.; Eritja, R. Covalent Strategies for Targeting Messenger and Non-Coding RNAs: An Updated Review on SiRNA, MiRNA and AntimiR Conjugates. Genes 2018, 9, 74. [Google Scholar] [CrossRef] [PubMed]
- Kalita, T.; Dezfouli, S.A.; Pandey, L.M.; Uludag, H. SiRNA Functionalized Lipid Nanoparticles (LNPs) in Management of Diseases. Pharmaceutics 2022, 14, 2520. [Google Scholar] [CrossRef]
- Yonezawa, S.; Koide, H.; Asai, T. Recent Advances in SiRNA Delivery Mediated by Lipid-Based Nanoparticles. Adv. Drug Deliv. Rev. 2020, 154–155, 64–78. [Google Scholar] [CrossRef]
- Xue, H.Y.; Wong, H.L. Tailoring Nanostructured Solid-Lipid Carriers for Time-Controlled Intracellular SiRNA Kinetics to Sustain RNAi-Mediated Chemosensitization. Biomaterials 2011, 32, 2662–2672. [Google Scholar] [CrossRef]
- Cortesi, R.; Campioni, M.; Ravani, L.; Drechsler, M.; Pinotti, M.; Esposito, E. Cationic Lipid Nanosystems as Carriers for Nucleic Acids. N. Biotechnol. 2014, 31, 44–54. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.H.; Kim, E.; Park, D.E.; Shim, G.; Lee, S.; Kim, Y.B.; Kim, C.-W.; Oh, Y.-K. Cationic Solid Lipid Nanoparticles for Co-Delivery of Paclitaxel and SiRNA. Eur. J. Pharm. Biopharm. 2012, 80, 268–273. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, M.C.; Carbone, C.; Souto, E.B. Beyond Liposomes: Recent Advances on Lipid Based Nanostructures for Poorly Soluble/Poorly Permeable Drug Delivery. Prog. Lipid Res. 2017, 68, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, T.S.; Lee, A.C.H.; Akinc, A.; Bramlage, B.; Bumcrot, D.; Fedoruk, M.N.; Harborth, J.; Heyes, J.A.; Jeffs, L.B.; John, M.; et al. RNAi-Mediated Gene Silencing in Non-Human Primates. Nature 2006, 441, 111–114. [Google Scholar] [CrossRef]
- Sarisozen, C.; Salzano, G.P.; Torchilin, V. Lipid-Based SiRNA Delivery Systems: Challenges, Promises and Solutions Along the Long Journey. Curr. Pharm. Biotechnol. 2016, 17, 728–740. [Google Scholar] [CrossRef]
- Jia, Z.; Gong, Y.; Pi, Y.; Liu, X.; Gao, L.; Kang, L.; Wang, J.; Yang, F.; Tang, J.; Lu, W.; et al. PPB Peptide-Mediated SiRNA-Loaded Stable Nucleic Acid Lipid Nanoparticles on Targeting Therapy of Hepatic Fibrosis. Mol. Pharm. 2018, 15, 53–62. [Google Scholar] [CrossRef]
- Ge, X.; Chen, L.; Zhao, B.; Yuan, W. Rationale and Application of PEGylated Lipid-Based System for Advanced Target Delivery of SiRNA. Front. Pharmacol. 2021, 11, 598175. [Google Scholar] [CrossRef]
- Chen, S.; Tam, Y.Y.C.; Lin, P.J.C.; Leung, A.K.K.; Tam, Y.K.; Cullis, P.R. Development of Lipid Nanoparticle Formulations of SiRNA for Hepatocyte Gene Silencing Following Subcutaneous Administration. J. Control. Release 2014, 196, 106–112. [Google Scholar] [CrossRef]
- Gao, L.Y.; Liu, X.Y.; Chen, C.J.; Wang, J.C.; Feng, Q.; Yu, M.Z.; Ma, X.F.; Pei, X.W.; Niu, Y.J.; Qiu, C.; et al. Core-Shell Type Lipid/RPAA-Chol Polymer Hybrid Nanoparticles for Invivo SiRNA Delivery. Biomaterials 2014, 35, 2066–2078. [Google Scholar] [CrossRef]
- Parveen, S.; Gupta, P.; Kumar, S.; Banerjee, M. Lipid Polymer Hybrid Nanoparticles as Potent Vehicles for Drug Delivery in Cancer Therapeutics. Med. Drug Discov. 2023, 20, 100165. [Google Scholar] [CrossRef]
- Yang, X.Z.; Dou, S.; Wang, Y.C.; Long, H.Y.; Xiong, M.H.; Mao, C.Q.; Yao, Y.D.; Wang, J. Single-Step Assembly of Cationic Lipid-Polymer Hybrid Nanoparticles for Systemic Delivery of SiRNA. ACS Nano 2012, 6, 4955–4965. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Zhao, P.; Su, W.; Wang, S.; Liao, Z.; Niu, R.; Chang, J. PLGA/Polymeric Liposome for Targeted Drug and Gene Co-Delivery. Biomaterials 2010, 31, 8741–8748. [Google Scholar] [CrossRef]
- Zhao, X.; Li, F.; Li, Y.; Wang, H.; Ren, H.; Chen, J.; Nie, G.; Hao, J. Co-Delivery of HIF1α SiRNA and Gemcitabine via Biocompatible Lipid-Polymer Hybrid Nanoparticles for Effective Treatment of Pancreatic Cancer. Biomaterials 2015, 46, 13–25. [Google Scholar] [CrossRef]
- Thanki, K.; Zeng, X.; Foged, C. Preparation, Characterization, and In Vitro Evaluation of Lipidoid–Polymer Hybrid Nanoparticles for SiRNA Delivery to the Cytosol. In Methods in Molecular Biology; Humana Press Inc.: Totowa, NJ, USA, 2019; Volume 1943, pp. 141–152. [Google Scholar]
- Li, J.; He, Y.Z.; Li, W.; Shen, Y.Z.; Li, Y.R.; Wang, Y.F. A Novel Polymer-Lipid Hybrid Nanoparticle for Efficient Nonviral Gene Delivery. Acta Pharmacol. Sin. 2010, 31, 509–514. [Google Scholar] [CrossRef]
- Hasan, W.; Chu, K.; Gullapalli, A.; Dunn, S.S.; Enlow, E.M.; Luft, J.C.; Tian, S.; Napier, M.E.; Pohlhaus, P.D.; Rolland, J.P.; et al. Delivery of Multiple SiRNAs Using Lipid-Coated PLGA Nanoparticles for Treatment of Prostate Cancer. Nano Lett. 2012, 12, 287–292. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves, G.; Vila, M.; Portolés, M.T.; Vallet-Regi, M.; Gracio, J.; Marques, P.A.A.P. Nano-Graphene Oxide: A Potential Multifunctional Platform for Cancer Therapy. Adv. Healthc. Mater. 2013, 2, 1072–1090. [Google Scholar] [CrossRef] [PubMed]
- Bose, R.J.C.; Arai, Y.; Ahn, J.C.; Park, H.; Lee, S.H. Influence of Cationic Lipid Concentration on Properties of Lipid–Polymer Hybrid Nanospheres for Gene Delivery. Int. J. Nanomed. 2015, 10, 5367–5382. [Google Scholar] [CrossRef] [PubMed]
- Persano, F.; Gigli, G.; Leporatti, S. Lipid-Polymer Hybrid Nanoparticles in Cancer Therapy: Current Overview and Future Directions. Nano Express 2021, 2, 012006. [Google Scholar] [CrossRef]
- Wang, J.; Xie, L.; Wang, T.; Wu, F.; Meng, J.; Liu, J.; Xu, H. Visible Light-Switched Cytosol Release of SiRNA by Amphiphilic Fullerene Derivative to Enhance RNAi Efficacy in Vitro and in Vivo. Acta. Biomater. 2017, 59, 158–169. [Google Scholar] [CrossRef]
- Yang, Z.; Luo, X.; Zhang, X.; Liu, J.; Jiang, Q. Targeted Delivery of 10-Hydroxycamptothecin to Human Breast Cancers by Cyclic RGD-Modified Lipid-Polymer Hybrid Nanoparticles. Biomed. Mater. 2013, 8, 025012. [Google Scholar] [CrossRef]
- Krishnamurthy, S.; Vaiyapuri, R.; Zhang, L.; Chan, J.M. Lipid-Coated Polymeric Nanoparticles for Cancer Drug Delivery. Biomater. Sci. 2015, 3, 923–936. [Google Scholar] [CrossRef]
- Triantafyllopoulou, E.; Perinelli, D.R.; Forys, A.; Pantelis, P.; Gorgoulis, V.G.; Lagopati, N.; Trzebicka, B.; Bonacucina, G.; Valsami, G.; Pippa, N.; et al. Unveiling the Performance of Co-Assembled Hybrid Nanocarriers: Moving towards the Formation of a Multifunctional Lipid/Random Copolymer Nanoplatform. Pharmaceutics 2024, 16, 1204. [Google Scholar] [CrossRef]
- Rezaee, M.; Oskuee, R.K.; Nassirli, H.; Malaekeh-Nikouei, B. Progress in the Development of Lipopolyplexes as Efficient Non-Viral Gene Delivery Systems. J. Control. Release 2016, 236, 1–14. [Google Scholar] [CrossRef]
- Jain, S.; Kumar, M.; Kumar, P.; Verma, J.; Rosenholm, J.M.; Bansal, K.K.; Vaidya, A. Lipid–Polymer Hybrid Nanosystems: A Rational Fusion for Advanced Therapeutic Delivery. J. Funct. Biomater. 2023, 14, 437. [Google Scholar] [CrossRef]
- Morakul, B. Self-Nanoemulsifying Drug Delivery Systems (SNEDDS): An Advancement Technology for Oral Drug Delivery. Pharm. Sci. Asia 2020, 47, 205–220. [Google Scholar] [CrossRef]
- Kubackova, J.; Holas, O.; Zbytovska, J.; Vranikova, B.; Zeng, G.; Pavek, P.; Mullertz, A. Oligonucleotide Delivery across the Caco-2 Monolayer: The Design and Evaluation of Self-Emulsifying Drug Delivery Systems (Sedds). Pharmaceutics 2021, 13, 459. [Google Scholar] [CrossRef]
- Reyna-Lázaro, L.; Morales-Becerril, A.; Aranda-Lara, L.; Isaac-Olivé, K.; Ocampo-García, B.; Gibbens-Bandala, B.; Olea-Mejía, O.; Morales-Avila, E. Pharmaceutical Nanoplatforms Based on Self-Nanoemulsifying Drug Delivery Systems for Optimal Transport and Co-Delivery of SiRNAs and Anticancer Drugs. J. Pharm. Sci. 2024, 113, 1907–1918. [Google Scholar] [CrossRef] [PubMed]
- Attri, N.; Das, S.; Banerjee, J.; Shamsuddin, S.H.; Dash, S.K.; Pramanik, A. Liposomes to Cubosomes: The Evolution of Lipidic Nanocarriers and Their Cutting-Edge Biomedical Applications. ACS Appl. Bio. Mater. 2024, 7, 2677–2694. [Google Scholar] [CrossRef] [PubMed]
- Sivadasan, D.; Sultan, M.H.; Alqahtani, S.S.; Javed, S. Cubosomes in Drug Delivery—A Comprehensive Review on Its Structural Components, Preparation Techniques and Therapeutic Applications. Biomedicines 2023, 11, 1114. [Google Scholar] [CrossRef]
- Sarkar, S.; Tran, N.; Soni, S.K.; Nasa, Z.; Drummond, C.J.; Conn, C.E. Cuboplex-Mediated Nonviral Delivery of Functional SiRNA to Chinese Hamster Ovary (CHO) Cells. ACS Appl. Mater. Interfaces 2021, 13, 2336–2345. [Google Scholar] [CrossRef] [PubMed]
- Yaghmur, A.; Østergaard, J.; Mu, H. Lipid Nanoparticles for Targeted Delivery of Anticancer Therapeutics: Recent Advances in Development of SiRNA and Lipoprotein-Mimicking Nanocarriers. Adv. Drug Deliv. Rev. 2023, 203, 115136. [Google Scholar] [CrossRef]
- Busatto, S.; Walker, S.A.; Grayson, W.; Pham, A.; Tian, M.; Nesto, N.; Barklund, J.; Wolfram, J. Lipoprotein-Based Drug Delivery. Adv. Drug Deliv. Rev. 2020, 159, 377–390. [Google Scholar] [CrossRef]
- Dai, L.; Li, S.; Hao, Q.; Zhou, R.; Zhou, H.; Lei, W.; Kang, H.; Wu, H.; Li, Y.; Ma, X. Low-Density Lipoprotein: A Versatile Nanoscale Platform for Targeted Delivery. Nanoscale Adv. 2023, 5, 1011–1022. [Google Scholar] [CrossRef]
- McMahon, K.M.; Plebanek, M.P.; Thaxton, C.S. Properties of Native High-Density Lipoproteins Inspire Synthesis of Actively Targeted In Vivo SiRNA Delivery Vehicles. Adv. Funct. Mater. 2016, 26, 7824–7835. [Google Scholar] [CrossRef]
- McMahon, K.M.; Thaxton, C.S. High-Density Lipoproteins for the Systemic Delivery of Short Interfering RNA. Expert. Opin. Drug Deliv. 2014, 11, 231–247. [Google Scholar] [CrossRef] [PubMed]
- Lacko, A.G.; Sabnis, N.A.; Nagarajan, B.; McConathy, W.J. HDL as a Drug and Nucleic Acid Delivery Vehicle. Front. Pharmacol. 2015, 6, 247. [Google Scholar] [CrossRef] [PubMed]
- Lacko, A.G.; Nair, M.; Prokai, L.; McConathy, W.J. Prospects and Challenges of the Development of Lipoprotein-Based Formulations for Anti-Cancer Drugs. Expert. Opin. Drug Deliv. 2007, 4, 665–675. [Google Scholar] [CrossRef] [PubMed]
- Almer, G.; Mangge, H.; Zimmer, A.; Prassl, R. Lipoprotein-Related and Apolipoprotein-Mediated Delivery Systems for Drug Targeting and Imaging. Curr. Med. Chem. 2015, 22, 3631–3651. [Google Scholar] [CrossRef]
- Aranda-Lara, L.; Isaac-Olivé, K.; Ocampo-García, B.; Ferro-Flores, G.; González-Romero, C.; Mercado-López, A.; García-Marín, R.; Santos-Cuevas, C.; Estrada, J.A.; Morales-Avila, E. Engineered RHDL Nanoparticles as a Suitable Platform for Theranostic Applications. Molecules 2022, 27, 7046. [Google Scholar] [CrossRef]
- Ng, K.K.; Lovell, J.F.; Zheng, G. Lipoprotein-Inspired Nanoparticles for Cancer Theranostics. Acc. Chem. Res. 2011, 44, 1105–1113. [Google Scholar] [CrossRef]
- Yang, S.D.; Zhu, W.J.; Zhu, Q.L.; Chen, W.L.; Ren, Z.X.; Li, F.; Yuan, Z.Q.; Li, J.Z.; Liu, Y.; Zhou, X.F.; et al. Binary-Copolymer System Base on Low-Density Lipoprotein-Coupled N-Succinyl Chitosan Lipoic Acid Micelles for Co-Delivery MDR1 SiRNA and Paclitaxel, Enhances Antitumor Effects via Reducing Drug. J. Biomed. Mater. Res. B Appl. Biomater. 2017, 105, 1114–1125. [Google Scholar] [CrossRef]
- Quintos-Meneses, H.A.; Aranda-Lara, L.; Morales-Ávila, E.; Ocampo-García, B.; Contreras, I.; Ramírez-Nava, G.J.; Santos-Cuevas, C.L.; Estrada, J.A.; Luna-Gutiérrez, M.A.; Ferro-Flores, G.; et al. A Multimodal Theranostic System Prepared from High-Density Lipoprotein Carrier of Doxorubicin and 177 Lu. J. Biomed. Nanotechnol. 2021, 17, 2125–2141. [Google Scholar] [CrossRef]
- He, B.; Yang, Q. Recent Development of LDL-Based Nanoparticles for Cancer Therapy. Pharmaceuticals 2022, 16, 18. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.; Li, Z.; Zheng, Q.; Yao, Q. Correlation Study of Serum Lipid Levels and Lipid Metabolism-Related Genes in Cervical Cancer. Front. Oncol. 2024, 14, 1384778. [Google Scholar] [CrossRef]
- Deng, C.-F.; Zhu, N.; Zhao, T.-J.; Li, H.-F.; Gu, J.; Liao, D.-F.; Qin, L. Involvement of LDL and Ox-LDL in Cancer Development and Its Therapeutical Potential. Front. Oncol. 2022, 12, 803473. [Google Scholar] [CrossRef]
- Jin, H.; Lovell, J.F.; Chen, J.; Lin, Q.; Ding, L.; Ng, K.K.; Pandey, R.K.; Manoharan, M.; Zhang, Z.; Zheng, G. Mechanistic Insights into LDL Nanoparticle-Mediated SiRNA Delivery. Bioconjug Chem. 2012, 23, 33–41. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Duan, X.; Lv, Y.; Zhao, Y. Low Density Lipoprotein Receptor (LDLR)-Targeted Lipid Nanoparticles for the Delivery of Sorafenib and Dihydroartemisinin in Liver Cancers. Life Sci. 2019, 239, 117013. [Google Scholar] [CrossRef]
- Zhu, C.; Pradhan, P.; Huo, D.; Xue, J.; Shen, S.; Roy, K.; Xia, Y. Reconstitution of Low-Density Lipoproteins with Fatty Acids for the Targeted Delivery of Drugs into Cancer Cells. Angew. Chem. Int. Ed. 2017, 56, 10399–10402. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.; Wang, Y.; Huang, Y.; Zi, Y.; Yan, S.; Shi, X.; Cai, J.; Zhang, H.; Sang, J.; Zhang, W.; et al. Bioinspired Low-Density Lipoprotein Co-Delivery System for Targeting and Synergistic Cancer Therapy. Nanomedicine 2023, 48, 102641. [Google Scholar] [CrossRef]
- McMahon, K.M.; Foit, L.; Angeloni, N.L.; Giles, F.J.; Gordon, L.I.; Thaxton, C.S. Synthetic High-Density Lipoprotein-like Nanoparticles as Cancer Therapy. Cancer Treat. Res. 2015, 166, 129–150. [Google Scholar] [CrossRef]
- Wolfrum, C.; Shi, S.; Jayaprakash, K.N.; Jayaraman, M.; Wang, G.; Pandey, R.K.; Rajeev, K.G.; Nakayama, T.; Charrise, K.; Ndungo, E.M.; et al. Mechanisms and Optimization of in Vivo Delivery of Lipophilic SiRNAs. Nat. Biotechnol. 2007, 25, 1149–1157. [Google Scholar] [CrossRef]
- Kuwahara, H.; Nishina, K.; Yoshida, K.; Nishina, T.; Yamamoto, M.; Saito, Y.; Piao, W.; Yoshida, M.; Mizusawa, H.; Yokota, T. Efficient in Vivo Delivery of SiRNA into Brain Capillary Endothelial Cells along with Endogenous Lipoprotein. Mol. Ther. 2011, 19, 2213–2221. [Google Scholar] [CrossRef]
- Tanaka, M.; Hasegawa, M.; Yoshimoto, N.; Hoshikawa, K.; Mukai, T. Preparation of Lipid Nanodisks Containing Apolipoprotein E-Derived Synthetic Peptides for Biocompatible Delivery Vehicles Targeting Low-Density Lipoprotein Receptor. Biol. Pharm. Bull. 2019, 42, 1376–1383. [Google Scholar] [CrossRef]
- Zhu, Q.L.; Zhou, Y.; Guan, M.; Zhou, X.-F.; Yang, S.-D.; Liu, Y.; Chen, W.-L.; Zhang, C.-G.; Yuan, Z.-Q.; Liu, C.; et al. Low-Density Lipoprotein-Coupled N-Succinyl Chitosan Nanoparticles Co-Delivering SiRNA and Doxorubicin for Hepatocyte-Targeted Therapy. Biomaterials 2014, 35, 5965–5976. [Google Scholar] [CrossRef]
- Glickson, J.D.; Lund-Katz, S.; Zhou, R.; Choi, H.; Chen, I.-W.; Li, H.; Corbin, I.; Popov, A.V.; Cao, W.; Song, L.; et al. Lipoprotein Nanoplatform for Targeted Delivery of Diagnostic and Therapeutic Agents. In Oxygen Transport to Tissue XXX; Springer: Boston, MA, USA, 2009; Volume 645, pp. 227–239. [Google Scholar]
- Mei, Y.; Tang, L.; Xiao, Q.; Zhang, Z.; Zhang, Z.; Zang, J.; Zhou, J.; Wang, Y.; Wang, W.; Ren, M. Reconstituted High Density Lipoprotein (RHDL), a Versatile Drug Delivery Nanoplatform for Tumor Targeted Therapy. J. Mater. Chem. B 2021, 9, 612–633. [Google Scholar] [CrossRef]
- Shahzad, M.M.K.; Mangala, L.S.; Han, H.D.; Lu, C.; Bottsford-Miller, J.; Nishimura, M.; Mora, E.M.; Lee, J.W.; Stone, R.L.; Pecot, C.V.; et al. Targeted Delivery of Small Interfering RNA Using Reconstituted High-Density Lipoprotein Nanoparticles. Neoplasia 2011, 13, 309–319. [Google Scholar] [CrossRef] [PubMed]
- Pottash, A.E.; Kuffner, C.; Noonan-Shueh, M.; Jay, S.M. Protein-Based Vehicles for Biomimetic RNAi Delivery. J. Biol. Eng. 2019, 13, 30. [Google Scholar] [CrossRef] [PubMed]
- Zamanian-Daryoush, M.; Lindner, D.J.; Buffa, J.; Gopalan, B.; Na, J.; Hazen, S.L.; Didonato, J.A. Apolipoprotein A-I Anti-Tumor Activity Targets Cancer Cell Metabolism. Oncotarget 2020, 11, 1777–1796. [Google Scholar]
- Kannan Mutharasan, R.; Foit, L.; Shad Thaxton, C. High-Density Lipoproteins for Therapeutic Delivery Systems. J. Mater. Chem. B 2015, 4, 188–197. [Google Scholar] [CrossRef]
- Raut, S.; Mooberry, L.; Sabnis, N.; Garud, A.; Dossou, A.S.; Lacko, A. Reconstituted HDL: Drug Delivery Platform for Overcoming Biological Barriers to Cancer Therapy. Front. Pharmacol. 2018, 9, 1154. [Google Scholar] [CrossRef]
- Shen, J.; Zhang, W.; Qi, R.; Mao, Z.-W.; Shen, H. Engineering Functional Inorganic–Organic Hybrid Systems: Advances in SiRNA Therapeutics. Chem. Soc. Rev. 2018, 47, 1969–1995. [Google Scholar] [CrossRef] [PubMed]
- Lin, Q.; Chen, J.; Ng, K.K.; Cao, W.; Zhang, Z.; Zheng, G. Imaging the Cytosolic Drug Delivery Mechanism of HDL-like Nanoparticles. Pharm. Res. 2014, 31, 1438–1449. [Google Scholar] [CrossRef] [PubMed]
- Cruz, W.; Huang, H.; Barber, B.; Pasini, E.; Ding, L.; Zheng, G.; Chen, J.; Bhat, M. Lipoprotein-Like Nanoparticle Carrying Small Interfering RNA Against Spalt-Like Transcription Factor 4 Effectively Targets Hepatocellular Carcinoma Cells and Decreases Tumor Burden. Hepatol. Commun. 2020, 4, 2020. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.; Han, Y.; Wang, R.; Wang, Y.; Chi, C.; Zhao, Z.; Zhang, H.; Wang, W.; Yin, L.; Zhou, J. Rerouting Native HDL to Predetermined Receptors for Improved Tumor-Targeted Gene Silencing Therapy. ACS Appl. Mater. Interfaces 2017, 9, 30488–30501. [Google Scholar] [CrossRef] [PubMed]
- Uno, Y.; Piao, W.; Miyata, K.; Nishina, K.; Mizusawa, H.; Yokota, T. High-Density Lipoprotein Facilitates in Vivo Delivery of α-Tocopherol-Conjugated Short-Interfering RNA to the Brain. Hum. Gene Ther. 2011, 22, 711–719. [Google Scholar] [CrossRef]
- Rui, M.; Tang, H.; Li, Y.; Wei, X.; Xu, Y. Recombinant High Density Lipoprotein Nanoparticles for Target-Specific Delivery of SiRNA. Pharm. Res. 2013, 30, 1203–1214. [Google Scholar] [CrossRef]
- Yang, M.; Jin, H.; Chen, J.; Ding, L.; Ng, K.K.; Lin, Q.; Lovell, J.F.; Zhang, Z.; Zheng, G. Efficient Cytosolic Delivery of SiRNA Using HDL-Mimicking Nanoparticles. Small 2011, 7, 568–573. [Google Scholar] [CrossRef]
- Nakayama, T.; Butler, J.S.; Sehgal, A.; Severgnini, M.; Racie, T.; Sharman, J.; Ding, F.; Morskaya, S.S.; Brodsky, J.; Tchangov, L.; et al. Harnessing a Physiologic Mechanism for SiRNA Delivery with Mimetic Lipoprotein Particles. Mol. Ther. 2012, 20, 1582–1589. [Google Scholar] [CrossRef]
- Ding, Y.; Wang, Y.; Zhou, J.; Gu, X.; Wang, W.; Liu, C.; Bao, X.; Wang, C.; Li, Y.; Zhang, Q. Direct Cytosolic SiRNA Delivery by Reconstituted High Density Lipoprotein for Target-Specific Therapy of Tumor Angiogenesis. Biomaterials 2014, 35, 7214–7227. [Google Scholar] [CrossRef]
- Malajczuk, C.J.; Gandhi, N.S.; Mancera, R.L. Structure and Intermolecular Interactions in Spheroidal High-Density Lipoprotein Subpopulations. J. Struct. Biol. X 2021, 5, 100042. [Google Scholar] [CrossRef]
- Corbin, I.R. Ligand-Coupled Lipoprotein for Ovarian Cancer-Specific Drug Delivery. Methods Mol. Biol. 2013, 1049, 467–480. [Google Scholar] [CrossRef]
- Pérez-Medina, C.; Binderup, T.; Lobatto, M.E.; Tang, J.; Calcagno, C.; Giesen, L.; Wessel, C.H.; Witjes, J.; Ishino, S.; Baxter, S.; et al. In Vivo PET Imaging of HDL in Multiple Atherosclerosis Models. JACC Cardiovasc. Imaging 2016, 9, 950–961. [Google Scholar] [CrossRef] [PubMed]
- Kuai, R.; Li, D.; Chen, Y.E.; Moon, J.J.; Schwendeman, A. High-Density Lipoproteins: Nature’s Multifunctional Nanoparticles. ACS Nano 2016, 10, 3015–3041. [Google Scholar] [CrossRef]
- Ávila-Sánchez, M.A.; Isaac-Olivé, K.; Aranda-Lara, L.; Morales-Ávila, E.; Plata-Becerril, A.; Jiménez-Mancilla, N.P.; Ocampo-García, B.; Estrada, J.A.; Santos-Cuevas, C.L.; Torres-García, E.; et al. Targeted Photodynamic Therapy Using Reconstituted High-Density Lipoproteins as Rhodamine Transporters. Photodiagnosis Photodyn. Ther. 2022, 37, 102630. [Google Scholar] [CrossRef]
- Chuang, S.T.; Cruz, S.; Narayanaswami, V. Reconfiguring Nature’s Cholesterol Accepting Lipoproteins as Nanoparticle Platforms for Transport and Delivery of Therapeutic and Imaging Agents. Nanomaterials 2020, 10, 906. [Google Scholar] [CrossRef] [PubMed]
- Ubanako, P.; Mirza, S.; Ruff, P.; Penny, C. Exosome-Mediated Delivery of SiRNA Molecules in Cancer Therapy: Triumphs and Challenges. Front. Mol. Biosci. 2024, 11, 1447953. [Google Scholar] [CrossRef]
- Chen, J.; Fei, X.; Wang, J.; Cai, Z. Tumor-Derived Extracellular Vesicles: Regulators of Tumor Microenvironment and the Enlightenment in Tumor Therapy. Pharmacol. Res. 2020, 159, 105041. [Google Scholar] [CrossRef]
- Alvarez-Erviti, L.; Seow, Y.; Yin, H.; Betts, C.; Lakhal, S.; Wood, M.J.A. Delivery of SiRNA to the Mouse Brain by Systemic Injection of Targeted Exosomes. Nat. Biotechnol. 2011, 29, 341–345. [Google Scholar] [CrossRef]
- Kang, J.Y.; Mun, D.; Chun, Y.; Park, D.S.; Kim, H.; Yun, N.; Joung, B. Engineered Small Extracellular Vesicle-mediated NOX4 SiRNA Delivery for Targeted Therapy of Cardiac Hypertrophy. J. Extracell. Vesicles 2023, 12, e12371. [Google Scholar] [CrossRef] [PubMed]
- Ahmadi-Kashani, M.; Dehghani, H.; Zarrabi, A. A Biocompatible Nanoplatform Formed by MgAl-Layered Double Hydroxide Modified Mn3O4/N-Graphene Quantum Dot Conjugated-Polyaniline for PH-Triggered Release of Doxorubicin. Mater. Sci. Eng. C 2020, 114, 111055. [Google Scholar] [CrossRef]
- Jiang, Y.; Huo, S.; Hardie, J.; Liang, X.-J.; Rotello, V.M. Progress and Perspective of Inorganic Nanoparticle-Based SiRNA Delivery Systems. Expert. Opin. Drug Deliv. 2016, 13, 547–559. [Google Scholar] [CrossRef]
- Mobarakeh, V.; Modarressi, M.H.; Rahimi, P.; Bolhassani, A.; Arefian, E. Modification of SPION Nanocarriers for SiRNA Delivery: A Therapeutic Strategy against HIV Infection. Vaccine Res. 2019, 6, 43–49. [Google Scholar] [CrossRef]
- Chhour, P.; Naha, P.C.; Cheheltani, R.; Benardo, B.; Mian, S.; Cormode, D.P. Gold Nanoparticles for Biomedical Applications: Synthesis and In Vitro Evaluation. In Methods in Pharmacology and Toxicology; Humana Press Inc.: Totowa, NJ, USA, 2016; Volume 39, pp. 87–111. [Google Scholar]
- Wu, J.; Liu, B.; Wu, H.; Wu, Y.; Zhang, W.; Zhao, S.; Zhang, L.; Pan, X.; Gao, W.; Wang, X.; et al. A Gold Nanoparticle Platform for the Delivery of Functional TGF-Β1 SiRNA into Cancer Cells. J. Biomed. Nanotechnol. 2016, 12, 800–810. [Google Scholar] [CrossRef]
- Liu, X.Y.; Wang, J.Q.; Ashby, C.R.; Zeng, L.; Fan, Y.F.; Chen, Z.S. Gold Nanoparticles: Synthesis, Physiochemical Properties and Therapeutic Applications in Cancer. Drug Discov. Today 2021, 26, 1284–1292. [Google Scholar] [CrossRef]
- Cruz-Rodríguez, J.C.; Camacho-López, M.A.; Torres-García, E.; Aranda-Lara, L.; Morales-Avila, E.; Díaz-Sánchez, L.E.; Jiménez-Mancilla, N.P.; Isaac-Olivé, K. Characterization of the Absorption Properties of 5 Nm Spherical Gold Nanoparticles Functionalized with Dodecanothiol and without Functionalization with Potential Therapeutic Applications. Phys. Scr. 2023, 98, 065005. [Google Scholar] [CrossRef]
- DeLong, R.K.; Akhtar, U.; Sallee, M.; Parker, B.; Barber, S.; Zhang, J.; Craig, M.; Garrad, R.; Hickey, A.J.; Engstrom, E. Characterization and Performance of Nucleic Acid Nanoparticles Combined with Protamine and Gold. Biomaterials 2009, 30, 6451–6459. [Google Scholar] [CrossRef]
- Kong, L.; Wu, Y.; Alves, C.S.; Shi, X. Efficient Delivery of Therapeutic SiRNA into Glioblastoma Cells Using Multifunctional Dendrimer-Entrapped Gold Nanoparticles. Nanomedicine 2016, 12, 3103–3115. [Google Scholar] [CrossRef]
- Melamed, J.R.; Riley, R.S.; Valcourt, D.M.; Billingsley, M.M.; Kreuzberger, N.L.; Day, E.S. Quantification of SiRNA Duplexes Bound to Gold Nanoparticle Surfaces. In Methods in Molecular Biology; Humana Press Inc.: Totowa, NJ, USA, 2017; Volume 1570, pp. 1–15. [Google Scholar]
- Bloise, N.; Strada, S.; Dacarro, G.; Visai, L. Gold Nanoparticles Contact with Cancer Cell: A Brief Update. Int. J. Mol. Sci. 2022, 23, 7683. [Google Scholar] [CrossRef]
- Artiga, Á.; Serrano-Sevilla, I.; De Matteis, L.; Mitchell, S.G.; de la Fuente, J.M. Current Status and Future Perspectives of Gold Nanoparticle Vectors for SiRNA Delivery. J. Mater. Chem. B 2019, 7, 876–896. [Google Scholar] [CrossRef]
- Ekin, A.; Karatas, O.F.; Culha, M.; Ozen, M. Designing a Gold Nanoparticle-Based Nanocarrier for MicroRNA Transfection into the Prostate and Breast Cancer Cells. J. Gene Med. 2014, 16, 331–335. [Google Scholar] [CrossRef]
- Cho, T.J.; Gorham, J.M.; Pettibone, J.M.; Liu, J.; Tan, J.; Hackley, V.A. Parallel Multi-Parameter Study of PEI-Functionalized Gold Nanoparticle Synthesis for Bio-Medical Applications: Part 1—A Critical Assessment of Methodology, Properties, and Stability. J. Nanoparticle Res. 2019, 21, 148. [Google Scholar] [CrossRef]
- Elbakry, A.; Zaky, A.; Liebl, R.; Rachel, R.; Goepferich, A.; Breunig, M. Layer-by-Layer Assembled Gold Nanoparticles for Sirna Delivery. Nano Lett. 2009, 9, 2059–2064. [Google Scholar] [CrossRef]
- Kim, S.T.; Chompoosor, A.; Yeh, Y.; Agasti, S.S.; Solfiell, D.J.; Rotello, V.M. Dendronized Gold Nanoparticles for SiRNA Delivery. Small 2012, 8, 3253–3256. [Google Scholar] [CrossRef]
- Labala, S.; Jose, A.; Venuganti, V.V.K. Transcutaneous Iontophoretic Delivery of STAT3 SiRNA Using Layer-by-Layer Chitosan Coated Gold Nanoparticles to Treat Melanoma. Colloids Surf. B Biointerfaces 2016, 146, 188–197. [Google Scholar] [CrossRef]
- Shaabani, E.; Sharifiaghdam, M.; De Keersmaecker, H.; De Rycke, R.; De Smedt, S.; Faridi-Majidi, R.; Braeckmans, K.; Fraire, J.C. Layer by Layer Assembled Chitosan-Coated Gold Nanoparticles for Enhanced SiRNA Delivery and Silencing. Int. J. Mol. Sci. 2021, 22, 831. [Google Scholar] [CrossRef]
- Baghani, L.; Noroozi Heris, N.; Khonsari, F.; Dinarvand, S.; Dinarvand, M.; Atyabi, F. Trimethyl-Chitosan Coated Gold Nanoparticles Enhance Delivery, Cellular Uptake and Gene Silencing Effect of EGFR-SiRNA in Breast Cancer Cells. Front. Mol. Biosci. 2022, 9, 871541. [Google Scholar] [CrossRef]
- Elizarova, T.N.; Antopolsky, M.L.; Novichikhin, D.O.; Skirda, A.M.; Orlov, A.V.; Bragina, V.A.; Nikitin, P.I. A Straightforward Method for the Development of Positively Charged Gold Nanoparticle-Based Vectors for Effective SiRNA Delivery. Molecules 2023, 28, 3318. [Google Scholar] [CrossRef]
- Wu, L.; Zhou, W.; Lin, L.; Chen, A.; Feng, J.; Qu, X.; Zhang, H.; Yue, J. Delivery of Therapeutic Oligonucleotides in Nanoscale. Bioact. Mater. 2022, 7, 292–323. [Google Scholar] [CrossRef]
- Pędziwiatr-Werbicka, E.; Gorzkiewicz, M.; Michlewska, S.; Ionov, M.; Shcharbin, D.; Klajnert-Maculewicz, B.; Peña-González, C.E.; Sánchez-Nieves, J.; Gómez, R.; de la Mata, F.J.; et al. Evaluation of Dendronized Gold Nanoparticles as SiRNAs Carriers into Cancer Cells. J. Mol. Liq. 2021, 324, 114726. [Google Scholar] [CrossRef]
- Huang, N.; Liu, Y.; Fang, Y.; Zheng, S.; Wu, J.; Wang, M.; Zhong, W.; Shi, M.; Xing, M.; Liao, W. Gold Nanoparticles Induce Tumor Vessel Normalization and Impair Metastasis by Inhibiting Endothelial Smad2/3 Signaling. ACS Nano 2020, 14, 7940–7958. [Google Scholar] [CrossRef]
- Issa, B.; Obaidat, I.; Albiss, B.; Haik, Y. Magnetic Nanoparticles: Surface Effects and Properties Related to Biomedicine Applications. Int. J. Mol. Sci. 2013, 14, 21266–21305. [Google Scholar] [CrossRef]
- Abdelrahman, M.; Douziech Eyrolles, L.; Alkarib, S.Y.; Hervé-Aubert, K.; Ben Djemaa, S.; Marchais, H.; Chourpa, I.; David, S. SiRNA Delivery System Based on Magnetic Nanovectors: Characterization and Stability Evaluation. Eur. J. Pharm. Sci. 2017, 106, 287–293. [Google Scholar] [CrossRef]
- Gozuacik, D.; Yagci-Acar, H.F.; Akkoc, Y.; Kosar, A.; Dogan-Ekici, A.I.; Ekici, S. Anticancer Use of Nanoparticles as Nucleic Acid Carriers. J. Biomed. Nanotechnol. 2014, 10, 1751–1783. [Google Scholar] [CrossRef]
- Dalmina, M.; Pittella, F.; Sierra, J.A.; Souza, G.R.R.; Silva, A.H.; Pasa, A.A.; Creczynski-Pasa, T.B. Magnetically Responsive Hybrid Nanoparticles for in Vitro SiRNA Delivery to Breast Cancer Cells. Mater. Sci. Eng. C 2019, 99, 1182–1190. [Google Scholar] [CrossRef]
- Arango, D.; Cifuentes, J.; Puentes, P.R.; Beltran, T.; Bittar, A.; Ocasión, C.; Muñoz-Camargo, C.; Bloch, N.I.; Reyes, L.H.; Cruz, J.C. Tailoring Magnetite-Nanoparticle-Based Nanocarriers for Gene Delivery: Exploiting CRISPRa Potential in Reducing Conditions. Nanomaterials 2023, 13, 1782. [Google Scholar] [CrossRef]
- Dobson Magnetic Nanoparticles for Gene and Drug Delivery. Int. J. Nanomed. 2008, 3, 169. [CrossRef]
- Bruniaux, J.; Allard-Vannier, E.; Aubrey, N.; Lakhrif, Z.; Ben Djemaa, S.; Eljack, S.; Marchais, H.; Hervé-Aubert, K.; Chourpa, I.; David, S. Magnetic Nanocarriers for the Specific Delivery of SiRNA: Contribution of Breast Cancer Cells Active Targeting for down-Regulation Efficiency. Int. J. Pharm. 2019, 569, 118572. [Google Scholar] [CrossRef]
- Mahajan, U.M.; Teller, S.; Sendler, M.; Palankar, R.; van den Brandt, C.; Schwaiger, T.; Kühn, J.-P.; Ribback, S.; Glöckl, G.; Evert, M.; et al. Tumour-Specific Delivery of SiRNA-Coupled Superparamagnetic Iron Oxide Nanoparticles, Targeted against PLK1, Stops Progression of Pancreatic Cancer. Gut 2016, 65, 1838–1849. [Google Scholar] [CrossRef]
- Panday, R.; Abdalla, A.M.E.; Yu, M.; Li, X.; Ouyang, C.; Yang, G. Functionally Modified Magnetic Nanoparticles for Effective SiRNA Delivery to Prostate Cancer Cells in Vitro. J. Biomater. Appl. 2020, 34, 952–964. [Google Scholar] [CrossRef]
- Yang, Z.; Duan, J.; Wang, J.; Liu, Q.; Shang, R.; Yang, X.; Lu, P.; Xia, C.; Wang, L.; Dou, K. Superparamagnetic Iron Oxide Nanoparticles Modified with Polyethylenimine and Galactose for SiRNA Targeted Delivery in Hepatocellular Carcinoma Therapy. Int. J. Nanomed. 2018, 13, 1851–1865. [Google Scholar] [CrossRef]
- Lin, G.; Huang, J.; Zhang, M.; Chen, S.; Zhang, M. Chitosan-Crosslinked Low Molecular Weight PEI-Conjugated Iron Oxide Nanoparticle for Safe and Effective DNA Delivery to Breast Cancer Cells. Nanomaterials 2022, 12, 584. [Google Scholar] [CrossRef]
- Licciardi, M.; Li Volsi, A.; Sardo, C.; Mauro, N.; Cavallaro, G.; Giammona, G. Inulin-Ethylenediamine Coated SPIONs Magnetoplexes: A Promising Tool for Improving SiRNA Delivery. Pharm. Res. 2015, 32, 3674–3687. [Google Scholar] [CrossRef]
- Bi, Q.; Song, X.; Hu, A.; Luo, T.; Jin, R.; Ai, H.; Nie, Y. Magnetofection: Magic Magnetic Nanoparticles for Efficient Gene Delivery. Chin. Chem. Lett. 2020, 31, 3041–3046. [Google Scholar] [CrossRef]
- Mondal, S.; Raut, J.; Sahoo, P. Gene Silencing and Gene Delivery in Therapeutics: Insights Using Quantum Dots. Front. Biosci. 2023, 28, 364. [Google Scholar] [CrossRef]
- Alizadeh-Ghodsi, M.; Pourhassan-Moghaddam, M.; Zavari-Nematabad, A.; Walker, B.; Annabi, N.; Akbarzadeh, A. State-of-the-Art and Trends in Synthesis, Properties, and Application of Quantum Dots-Based Nanomaterials. Part. Part. Syst. Charact. 2019, 36, 1900197. [Google Scholar] [CrossRef]
- Banerjee, A.; Pons, T.; Lequeux, N.; Dubertret, B. Quantum Dots–DNA Bioconjugates: Synthesis to Applications. Interface Focus. 2016, 6, 20160064. [Google Scholar] [CrossRef]
- Li, J.-M.; Zhao, M.-X.; Su, H.; Wang, Y.-Y.; Tan, C.-P.; Ji, L.-N.; Mao, Z.-W. Multifunctional Quantum-Dot-Based SiRNA Delivery for HPV18 E6 Gene Silence and Intracellular Imaging. Biomaterials 2011, 32, 7978–7987. [Google Scholar] [CrossRef]
- Li, J.M.; Wang, Y.Y.; Zhao, M.X.; Tan, C.P.; Li, Y.Q.; Le, X.Y.; Ji, L.N.; Mao, Z.W. Multifunctional QD-Based Co-Delivery of SiRNA and Doxorubicin to HeLa Cells for Reversal of Multidrug Resistance and Real-Time Tracking. Biomaterials 2012, 33, 2780–2790. [Google Scholar] [CrossRef]
- Bae, K.H.; Lee, J.Y.; Lee, S.H.; Park, T.G.; Nam, Y.S. Optically Traceable Solid Lipid Nanoparticles Loaded with Sirna and Paclitaxel for Synergistic Chemotherapy with in Situ Imaging. Adv. Healthc. Mater. 2013, 2, 576–584. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Dai, W.; Ju, H.; Lu, H.; Wang, S.; Xu, L.; Zhou, S.F.; Zhang, Y.; Zhang, X. Multifunctional Poly(l-Lactide)-Polyethylene Glycol-Grafted Graphene Quantum Dots for Intracellular MicroRNA Imaging and Combined Specific-Gene-Targeting Agents Delivery for Improved Therapeutics. ACS Appl. Mater. Interfaces 2015, 7, 11015–11023. [Google Scholar] [CrossRef] [PubMed]
- Jung, J.; Solanki, A.; Memoli, K.A.; Kamei, K.; Kim, H.; Drahl, M.A.; Williams, L.J.; Tseng, H.; Lee, K. Selective Inhibition of Human Brain Tumor Cells through Multifunctional Quantum-Dot-Based SiRNA Delivery. Angew. Chem. Int. Ed. 2010, 49, 103–107. [Google Scholar] [CrossRef] [PubMed]
- Endres, T.; Zheng, M.; Kiliç, A.; Turowska, A.; Beck-Broichsitter, M.; Renz, H.; Merkel, O.M.; Kissel, T. Amphiphilic Biodegradable PEG-PCL-PEI Triblock Copolymers for FRET-Capable in Vitro and in Vivo Delivery of SiRNA and Quantum Dots. Mol. Pharm. 2014, 11, 1273–1281. [Google Scholar] [CrossRef]
- Zhu, H.; Zhang, S.; Ling, Y.; Meng, G.; Yang, Y.; Zhang, W. PH-Responsive Hybrid Quantum Dots for Targeting Hypoxic Tumor SiRNA Delivery. J. Control. Release 2015, 220, 529–544. [Google Scholar] [CrossRef]
- Lee, H.; Kim, I.-K.; Park, T.G. Intracellular Trafficking and Unpacking of SiRNA/Quantum Dot-PEI Complexes Modified with and without Cell Penetrating Peptide: Confocal and Flow Cytometric FRET Analysis. Bioconjug Chem. 2010, 21, 289–295. [Google Scholar] [CrossRef] [PubMed]
- Lin, G.; Chen, T.; Zou, J.; Wang, Y.; Wang, X.; Li, J.; Huang, Q.; Fu, Z.; Zhao, Y.; Lin, M.C.M.; et al. Quantum Dots-SiRNA Nanoplexes for Gene Silencing in Central Nervous System Tumor Cells. Front. Pharmacol. 2017, 8, 182. [Google Scholar] [CrossRef]
- Derfus, A.M.; Chen, A.A.; Min, D.H.; Ruoslahti, E.; Bhatia, S.N. Targeted Quantum Dot Conjugates for SiRNA Delivery. Bioconjug Chem. 2007, 18, 1391–1396. [Google Scholar] [CrossRef]
- Bagalkot, V.; Gao, X. SiRNA-Aptamer Chimeras on Nanoparticles: Preserving Targeting Functionality for Effective Gene Silencing. ACS Nano 2011, 5, 8131–8139. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Yang, C.; Hu, R.; Toh, H.T.; Liu, X.; Lin, G.; Yin, F.; Yoon, H.S.; Yong, K.T. Assembling Mn:ZnSe Quantum Dots-SiRNA Nanoplexes for Gene Silencing in Tumor Cells. Biomater. Sci. 2015, 3, 192–202. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, D.; Sil, M.; Goswami, A.; Lahiri, D.; Nag, M. Antibiofilm Activities of Carbon-Based Nanoparticles and Nanocomposites: A Comparative Review. J. Inorg. Organomet. Polym. Mater. 2023, 33, 3961–3983. [Google Scholar] [CrossRef]
- Yazdani, S.; Mozaffarian, M.; Pazuki, G.; Hadidi, N.; Villate-Beitia, I.; Zárate, J.; Puras, G.; Pedraz, J.L. Carbon-Based Nanostructures as Emerging Materials for Gene Delivery Applications. Pharmaceutics 2024, 16, 288. [Google Scholar] [CrossRef]
- Riley, P.R.; Narayan, R.J. Recent Advances in Carbon Nanomaterials for Biomedical Applications: A Review. Curr. Opin. Biomed. Eng. 2021, 17, 100262. [Google Scholar] [CrossRef]
- Siu, K.S.; Zheng, X.; Liu, Y.; Zhang, Y.; Zhang, X.; Chen, D.; Yuan, K.; Gillies, E.R.; Koropatnick, J.; Min, W.-P. Single-Walled Carbon Nanotubes Noncovalently Functionalized with Lipid Modified Polyethylenimine for SiRNA Delivery in Vitro and in Vivo. Bioconjug Chem. 2014, 25, 1744–1751. [Google Scholar] [CrossRef]
- Siu, K.S.; Zhang, Y.; Zheng, X.; Koropatnick, J.; Min, W.P. Non-Covalently Functionalized of Single-Walled Carbon Nanotubes by DSPE-PEG-PEI for SiRNA Delivery. In Methods in Molecular Biology; Humana Press Inc.: Totowa, NJ, USA, 2016; Volume 1364, pp. 151–163. [Google Scholar]
- Varkouhi, A.K.; Foillard, S.; Lammers, T.; Schiffelers, R.M.; Doris, E.; Hennink, W.E.; Storm, G. SiRNA Delivery with Functionalized Carbon Nanotubes. Int. J. Pharm. 2011, 416, 419–425. [Google Scholar] [CrossRef]
- Zhang, L.; Zheng, W.; Tang, R.; Wang, N.; Zhang, W.; Jiang, X. Gene Regulation with Carbon-Based SiRNA Conjugates for Cancer Therapy. Biomaterials 2016, 104, 269–278. [Google Scholar] [CrossRef]
- Meran, M.; Akkus, P.D.; Kurkcuoglu, O.; Baysak, E.; Hizal, G.; Haciosmanoglu, E.; Unlu, A.; Karatepe, N.; Güner, F.S. Noncovalent Pyrene-Polyethylene Glycol Coatings of Carbon Nanotubes Achieve in Vitro Biocompatibility. Langmuir 2018, 34, 12071–12082. [Google Scholar] [CrossRef]
- Imani, R.; Shao, W.; Taherkhani, S.; Emami, S.H.; Prakash, S.; Faghihi, S. Dual-Functionalized Graphene Oxide for Enhanced SiRNA Delivery to Breast Cancer Cells. Colloids Surf. B Biointerfaces 2016, 147, 315–325. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Liu, Z.; Welsher, K.; Robinson, J.T.; Goodwin, A.; Zaric, S.; Dai, H. Nano-Graphene Oxide for Cellular Imaging and Drug Delivery. Nano Res. 2008, 1, 203–212. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Guo, Z.; Huang, D.; Liu, Z.; Guo, X.; Zhong, H. Synergistic Effect of Chemo-Photothermal Therapy Using PEGylated Graphene Oxide. Biomaterials 2011, 32, 8555–8561. [Google Scholar] [CrossRef]
- Yin, F.; Hu, K.; Chen, Y.; Yu, M.; Wang, D.; Wang, Q.; Yong, K.T.; Lu, F.; Liang, Y.; Li, Z. SiRNA Delivery with PEGylated Graphene Oxide Nan Osheets for Combined Photothermal and Genetherapy for Pancreatic Cancer. Theranostics 2017, 7, 1133–1148. [Google Scholar] [CrossRef]
- Liu, J.; Cui, L.; Losic, D. Graphene and Graphene Oxide as New Nanocarriers for Drug Delivery Applications. Acta Biomater. 2013, 9, 9243–9257. [Google Scholar] [CrossRef]
- Feng, L.; Yang, X.; Shi, X.; Tan, X.; Peng, R.; Wang, J.; Liu, Z. Polyethylene Glycol and Polyethylenimine Dual-Functionalized Nano-Graphene Oxide for Photothermally Enhanced Gene Delivery. Small 2013, 9, 1989–1997. [Google Scholar] [CrossRef] [PubMed]
- Georgakilas, V.; Tiwari, J.N.; Kemp, K.C.; Perman, J.A.; Bourlinos, A.B.; Kim, K.S.; Zboril, R. Noncovalent Functionalization of Graphene and Graphene Oxide for Energy Materials, Biosensing, Catalytic, and Biomedical Applications. Chem. Rev. 2016, 116, 5464–5519. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Ding, R.; Zhao, X.; Li, Y.; Qu, L.; Pei, H.; Yildirimer, L.; Wu, Z.; Zhang, W. Graphene-Based Nanomaterials for Drug and/or Gene Delivery, Bioimaging, and Tissue Engineering. Drug Discov. Today 2017, 22, 1302–1317. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.Y.; Zhang, W.; Liu, H.; Jiang, J.J.; Wang, W.J.; Jia, Z.Y. Cell-Penetrating Peptide-Modified Graphene Oxide Nanoparticles Loaded with Rictor SiRNA for the Treatment of Triple-Negative Breast Cancer. Drug Devel. Ther. 2021, 15, 4961–4972. [Google Scholar] [CrossRef]
- Montellano, A.; Da Ros, T.; Bianco, A.; Prato, M. Fullerene C60 as a Multifunctional System for Drug and Gene Delivery. Nanoscale 2011, 3, 4035–4041. [Google Scholar] [CrossRef] [PubMed]
- Minami, K.; Okamoto, K.; Doi, K.; Harano, K.; Noiri, E.; Nakamura, E. SiRNA Delivery Targeting to the Lung via Agglutination-Induced Accumulation and Clearance of Cationic Tetraamino Fullerene. Sci. Rep. 2014, 4, 4916. [Google Scholar] [CrossRef]
- Korzuch, J.; Rak, M.; Balin, K.; Zubko, M.; Głowacka, O.; Dulski, M.; Musioł, R.; Madeja, Z.; Serda, M. Towards Water-Soluble [60]Fullerenes for the Delivery of SiRNA in a Prostate Cancer Model. Sci. Rep. 2021, 11, 10565. [Google Scholar] [CrossRef]
- Minami, K.; Okamoto, K.; Harano, K.; Noiri, E.; Nakamura, E. Hierarchical Assembly of SiRNA with Tetraamino Fullerene in Physiological Conditions for Efficient Internalization into Cells and Knockdown. ACS Appl. Mater. Interfaces 2018, 10, 19347–19354. [Google Scholar] [CrossRef]
- Liu, S.; Sun, X.; Lu, H.; Chen, D.; Li, X.; Li, L.; Su, S.; Zhao, Z.; Cao, X.; Liu, L.; et al. Fullerene-Based Nanocomplex Assists Pulmonary Delivery of SiRNA for Treating Metastatic Lung Cancer. Nano Today 2023, 50, 101878. [Google Scholar] [CrossRef]
- Chen, F.; Gao, W.; Qiu, X.; Zhang, H.; Liu, L.; Liao, P.; Fu, W.; Luo, Y. Graphene Quantum Dots in Biomedical Applications: Recent Advances and Future Challenges. Front. Lab. Med. 2017, 1, 192–199. [Google Scholar] [CrossRef]
- Das, S.; Mondal, S.; Ghosh, D. Carbon Quantum Dots in Bioimaging and Biomedicines. Front. Bioeng. Biotechnol. 2024, 11, 1333752. [Google Scholar] [CrossRef]
- Mansuriya, B.D.; Altintas, Z. Carbon Dots: Classification, Properties, Synthesis, Characterization, and Applications in Health Care-An Updated Review (2018–2021). Nanomaterials 2021, 11, 2525. [Google Scholar] [CrossRef]
- Yang, H.-L.; Bai, L.-F.; Geng, Z.-R.; Chen, H.; Xu, L.-T.; Xie, Y.-C.; Wang, D.-J.; Gu, H.-W.; Wang, X.-M. Carbon Quantum Dots: Preparation, Optical Properties, and Biomedical Applications. Mater. Today Adv. 2023, 18, 100376. [Google Scholar] [CrossRef]
- Xia, C.; Zhu, S.; Feng, T.; Yang, M.; Yang, B. Evolution and Synthesis of Carbon Dots: From Carbon Dots to Carbonized Polymer Dots. Adv. Sci. 2019, 6, 1901316. [Google Scholar] [CrossRef]
- Pierrat, P.; Wang, R.; Kereselidze, D.; Lux, M.; Didier, P.; Kichler, A.; Pons, F.; Lebeau, L. Efficient Invitro and Invivo Pulmonary Delivery of Nucleic Acid by Carbon Dot-Based Nanocarriers. Biomaterials 2015, 51, 290–302. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Zhang, C.; Shen, G.; Liu, H.; Fu, H.; Cui, D. Fluorescent Carbon Dots as an Efficient SiRNA Nanocarrier for Its Interference Therapy in Gastric Cancer Cells. J. Nanobiotechnol. 2014, 12, 58. [Google Scholar] [CrossRef]
- Wang, L.; Wang, X.; Bhirde, A.; Cao, J.; Zeng, Y.; Huang, X.; Sun, Y.; Liu, G.; Chen, X. Carbon-Dot-Based Two-Photon Visible Nanocarriers for Safe and Highly Efficient Delivery of SiRNA and DNA. Adv. Healthc. Mater. 2014, 3, 1203–1209. [Google Scholar] [CrossRef]
- Adrita, S.; Tasnim, K.; Ryu, J.; Sharker, S. Nanotheranostic Carbon Dots as an Emerging Platform for Cancer Therapy. J. Nanotheranostics 2020, 58–77. [Google Scholar] [CrossRef]
- Vallet-Regí, M.; Colilla, M.; Izquierdo-Barba, I.; Manzano, M. Mesoporous Silica Nanoparticles for Drug Delivery: Current Insights. Molecules 2018, 23, 47. [Google Scholar] [CrossRef]
- Jafari, S.; Derakhshankhah, H.; Alaei, L.; Fattahi, A.; Varnamkhasti, B.S.; Saboury, A.A. Mesoporous Silica Nanoparticles for Therapeutic/Diagnostic Applications. Biomed. Pharmacother. 2019, 109, 1100–1111. [Google Scholar] [CrossRef] [PubMed]
- Rastegari, E.; Hsiao, Y.-J.; Lai, W.-Y.; Lai, Y.-H.; Yang, T.-C.; Chen, S.-J.; Huang, P.-I.; Chiou, S.-H.; Mou, C.-Y.; Chien, Y. An Update on Mesoporous Silica Nanoparticle Applications in Nanomedicine. Pharmaceutics 2021, 13, 1067. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Shi, S.; Goel, S.; Shen, X.; Xie, X.; Chen, Z.; Zhang, H.; Li, S.; Qin, X.; Yang, H.; et al. Recent Advancements in Mesoporous Silica Nanoparticles towards Therapeutic Applications for Cancer. Acta Biomater. 2019, 89, 1–13. [Google Scholar] [CrossRef]
- Xie, X.; Yue, T.; Gu, W.; Cheng, W.; He, L.; Ren, W.; Li, F.; Piao, J.-G. Recent Advances in Mesoporous Silica Nanoparticles Delivering SiRNA for Cancer Treatment. Pharmaceutics 2023, 15, 2483. [Google Scholar] [CrossRef]
- Ma, X.; Zhao, Y.; Ng, K.W.; Zhao, Y. Integrated Hollow Mesoporous Silica Nanoparticles for Target Drug/SiRNA Co-Delivery. Chem. Eur. J. 2013, 19, 15593–15603. [Google Scholar] [CrossRef]
- Wang, Y.; Zhao, Q.; Han, N.; Bai, L.; Li, J.; Liu, J.; Che, E.; Hu, L.; Zhang, Q.; Jiang, T.; et al. Mesoporous Silica Nanoparticles in Drug Delivery and Biomedical Applications. Nanomedicine 2015, 11, 313–327. [Google Scholar] [CrossRef]
- Chen, X.; Zhang, Y.; Tang, C.; Tian, C.; Sun, Q.; Su, Z.; Xue, L.; Yin, Y.; Ju, C.; Zhang, C. Co-Delivery of Paclitaxel and Anti-Survivin SiRNA via Redox-Sensitive Oligopeptide Liposomes for the Synergistic Treatment of Breast Cancer and Metastasis. Int. J. Pharm. 2017, 529, 102–115. [Google Scholar] [CrossRef]
- Xiao, D.; Hu, J.-J.; Zhu, J.-Y.; Wang, S.-B.; Zhuo, R.-X.; Zhang, X.-Z. A Redox-Responsive Mesoporous Silica Nanoparticle with a Therapeutic Peptide Shell for Tumor Targeting Synergistic Therapy. Nanoscale 2016, 8, 16702–16709. [Google Scholar] [CrossRef]
- Chan, M.-H.; Lin, H.-M. Preparation and Identification of Multifunctional Mesoporous Silica Nanoparticles for in Vitro and in Vivo Dual-Mode Imaging, Theranostics, and Targeted Tracking. Biomaterials 2015, 46, 149–158. [Google Scholar] [CrossRef] [PubMed]
- Girija, A.R.; Balasubramanian, S. Theragnostic Potentials of Core/Shell Mesoporous Silica Nanostructures. Nanotheranostics 2019, 3, 1–40. [Google Scholar] [CrossRef] [PubMed]
- Hanafi-Bojd, M.Y.; Ansari, L.; Malaekeh-Nikouei, B. Codelivery of Anticancer Drugs and SiRNA By Mesoporous Silica Nanoparticles. Ther. Deliv. 2016, 7, 649–655. [Google Scholar] [CrossRef]
- Meng, H.; Mai, W.X.; Zhang, H.; Xue, M.; Xia, T.; Lin, S.; Wang, X.; Zhao, Y.; Ji, Z.; Zink, J.I.; et al. Codelivery of an Optimal Drug/SiRNA Combination Using Mesoporous Silica Nanoparticles To Overcome Drug Resistance in Breast Cancer in Vitro and in Vivo. ACS Nano 2013, 7, 994–1005. [Google Scholar] [CrossRef] [PubMed]
- Paris, J.L.; Vallet-Regí, M. Mesoporous Silica Nanoparticles for Co-Delivery of Drugs and Nucleic Acids in Oncology: A Review. Pharmaceutics 2020, 12, 526. [Google Scholar] [CrossRef]
- Carvalho, A.M.; Cordeiro, R.A.; Faneca, H. Silica-Based Gene Delivery Systems: From Design to Therapeutic Applications. Pharmaceutics 2020, 12, 649. [Google Scholar] [CrossRef]
- Jin, M.; Hou, Y.; Quan, X.; Chen, L.; Gao, Z.; Huang, W. Smart Polymeric Nanoparticles with PH-Responsive and Peg-Detachable Properties (II): Co-Delivery of Paclitaxel and VEGF SiRNA for Synergistic Breast Cancer Therapy in Mice. Int. J. Nanomed. 2021, 16, 5479–5494. [Google Scholar] [CrossRef] [PubMed]
- Lv, W.; Xu, C.; Wu, H.; Zhu, Y.; Akakuru, O.U.; Du, H.; Nie, F.; Wu, A.; Li, J. Ultrasound-Visualized Nanocarriers with SiRNA for Targeted Inhibition of M2-like TAM Polarization to Enhance Photothermal Therapy in NSCLC. Nano Res. 2023, 16, 882–893. [Google Scholar] [CrossRef]
- Dang, J.; Ye, H.; Li, Y.; Liang, Q.; Li, X.; Yin, L. Multivalency-Assisted Membrane-Penetrating SiRNA Delivery Sensitizes Photothermal Ablation via Inhibition of Tumor Glycolysis Metabolism. Biomaterials 2019, 223, 119437. [Google Scholar] [CrossRef] [PubMed]
- Zheng, N.; Luo, X.; Zhang, Z.; Wang, A.; Song, W. Cationic Polyporphyrins as SiRNA Delivery Vectors for Photodynamic and Gene Synergistic Anticancer Therapy. ACS App.l Mater. Interfaces 2021, 13, 27513–27521. [Google Scholar] [CrossRef]
- Camorani, S.; Tortorella, S.; Agnello, L.; Spanu, C.; d’Argenio, A.; Nilo, R.; Zannetti, A.; Locatelli, E.; Fedele, M.; Comes Franchini, M.; et al. Aptamer-Functionalized Nanoparticles Mediate PD-L1 SiRNA Delivery for Effective Gene Silencing in Triple-Negative Breast Cancer Cells. Pharmaceutics 2022, 14, 2225. [Google Scholar] [CrossRef]
- Tan, S.; Chen, Z.; Mironchik, Y.; Mori, N.; Penet, M.F.; Si, G.; Krishnamachary, B.; Bhujwalla, Z.M. VEGF Overexpression Significantly Increases Nanoparticle-Mediated SiRNA Delivery and Target-Gene Downregulation. Pharmaceutics 2022, 14, 1260. [Google Scholar] [CrossRef]
- Ding, L.; Tang, S.; Yu, A.; Wang, A.; Tang, W.; Jia, H.; Oupický, D. Nanoemulsion-Assisted SiRNA Delivery to Modulate the Nervous Tumor Microenvironment in the Treatment of Pancreatic Cancer. ACS Appl. Mater. Interfaces 2022, 14, 10015–10029. [Google Scholar] [CrossRef]
- Vaidya, S.; Jeengar, M.K.; Wadaan, M.A.; Mahboob, S.; Kumar, P.; Reece, L.M.; Bathula, S.R.; Dutta, M. Design and In Vitro Evaluation of Novel Cationic Lipids for SiRNA Delivery in Breast Cancer Cell Lines. Evid. Based Complement. Altern. Med. 2022, 2022, 1–9. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, T.; Gao, J. Biocompatible Iron Oxide Nanoparticles for Targeted Cancer Gene Therapy: A Review. Nanomaterials 2022, 12, 3323. [Google Scholar] [CrossRef]
- Yu, S.; Bi, X.; Yang, L.; Wu, S.; Yu, Y.; Jiang, B.; Zhang, A.; Lan, K.; Duan, S. Co-Delivery of Paclitaxel and PLK1-Targeted SiRNA Using Aptamer-Functionalized Cationic Liposome for Synergistic Anti-Breast Cancer Effects in Vivo. J. Biomed. Nanotechnol. 2019, 15, 1135–1148. [Google Scholar] [CrossRef]
- Lee, J.; Cho, Y.J.; Lee, J.W.; Ahn, H.J. KSP SiRNA/Paclitaxel-Loaded PEGylated Cationic Liposomes for Overcoming Resistance to KSP Inhibitors: Synergistic Antitumor Effects in Drug-Resistant Ovarian Cancer. J. Control. Release 2020, 321, 184–197. [Google Scholar] [CrossRef]
- Xie, Y.; Lu, X.; Wang, Z.; Liu, M.; Liu, L.; Wang, R.; Yang, K.; Xiao, H.; Li, J.; Tang, X.; et al. A Hypoxia-Dissociable SiRNA Nanoplatform for Synergistically Enhanced Chemo-Radiotherapy of Glioblastoma. Biomater. Sci. 2022, 10, 6791–6803. [Google Scholar] [CrossRef]
- Çağdaş Tunalı, B.; Çelik, E.; Budak Yıldıran, F.A.; Türk, M. Delivery of SiRNA Using Hyaluronic Acid-Guided Nanoparticles for Downregulation of CXCR4. Biopolymers 2023, 114, e23535. [Google Scholar] [CrossRef] [PubMed]
- Park, J.; Kim, S.; Kim, T. Il Polyethylenimine-Conjugated Hydroxyethyl Cellulose for Doxorubicin/Bcl-2 SiRNA Co-Delivery Systems. Pharmaceutics 2023, 15, 708. [Google Scholar] [CrossRef] [PubMed]
- Aranda-Lara, L.; Trujillo-Nolasco, M.; López-Marmolejo, C.M.; Cárdenas-Rodríguez, K.; Gómez-Pulido, V.O.; Morales-Avila, E.; Ocampo-García, B.; Jiménez-Mancilla, N.P.; Estrada, J.A.; Otero, G.; et al. Chitosan/SiRNA Complex in Mimetic High-Density Lipoproteins as Specific/Hybrid Transfection Technology for SR-B1 Expressing Cancer Cells. J. Drug Deliv. Sci. Technol. 2024, 101, 106274. [Google Scholar] [CrossRef]
- Abbasi, A.; Lindqvist-Reis, P.; Eriksson, L.; Sandström, D.; Lidin, S.; Persson, I.; Sandström, M. Highly Hydrated Cations: Deficiency, Mobility, and Coordination of Water in Crystalline Nonahydrated Scandium(III), Yttrium(III), and Lanthanoid(III) Trifluoromethanesulfonates. Chem. Eur. J. 2005, 11, 4065–4077. [Google Scholar] [CrossRef]
- Hattab, D.; Gazzali, A.M.; Bakhtiar, A. Clinical Advances of Sirna-Based Nanotherapeutics for Cancer Treatment. Pharmaceutics 2021, 13, 1009. [Google Scholar] [CrossRef]
- Drago, S.E.; Utzeri, M.A.; Mauro, N.; Cavallaro, G. Polyamidoamine-Carbon Nanodot Conjugates with Bioreducible Building Blocks: Smart Theranostic Platforms for Targeted SiRNA Delivery. Biomacromolecules 2024, 25, 1191–1204. [Google Scholar] [CrossRef]
- Ranjbar, S.; Zhong, X.; Manautou, J.; Lu, X. A Holistic Analysis of the Intrinsic and Delivery-Mediated Toxicity of SiRNA Therapeutics. Adv. Drug Deliv. Rev. 2023, 201, 115052. [Google Scholar] [CrossRef]
- Liu, Z.; Sun, Q.; Wang, X. PLK1, A Potential Target for Cancer Therapy. Transl. Oncol. 2017, 10, 22–32. [Google Scholar] [CrossRef]
- El Dika, I.; Lim, H.Y.; Yong, W.P.; Lin, C.-C.; Yoon, J.-H.; Modiano, M.; Freilich, B.; Choi, H.J.; Chao, T.-Y.; Kelley, R.K.; et al. An Open-Label, Multicenter, Phase I, Dose Escalation Study with Phase II Expansion Cohort to Determine the Safety, Pharmacokinetics, and Preliminary Antitumor Activity of Intravenous TKM-080301 in Subjects with Advanced Hepatocellular Carcinoma. Oncologist 2019, 24, 747-e218. [Google Scholar] [CrossRef] [PubMed]
- Chiappa, M.; Petrella, S.; Damia, G.; Broggini, M.; Guffanti, F.; Ricci, F. Present and Future Perspective on PLK1 Inhibition in Cancer Treatment. Front. Oncol. 2022, 12, 903016. [Google Scholar] [CrossRef]
- Mukai, H.; Muramatsu, A.; Mashud, R.; Kubouchi, K.; Tsujimoto, S.; Hongu, T.; Kanaho, Y.; Tsubaki, M.; Nishida, S.; Shioi, G.; et al. PKN3 Is the Major Regulator of Angiogenesis and Tumor Metastasis in Mice. Sci. Rep. 2016, 6, 18979. [Google Scholar] [CrossRef]
- Schultheis, B.; Strumberg, D.; Kuhlmann, J.; Wolf, M.; Link, K.; Seufferlein, T.; Kaufmann, J.; Feist, M.; Gebhardt, F.; Khan, M.; et al. Safety, Efficacy and Pharcacokinetics of Targeted Therapy with the Liposomal RNA Interference Therapeutic Atu027 Combined with Gemcitabine in Patients with Pancreatic Adenocarcinoma. A Randomized Phase Ib/IIa Study. Cancers 2020, 12, 3130. [Google Scholar] [CrossRef]
- Temme, L.; Laitinen, T.; Asquith, C.R.M. PKN3: A Target in Cancer Metastasis. Nat. Rev. Drug Discov. 2022, 21, 704. [Google Scholar] [CrossRef] [PubMed]
- Unsal-Kacmaz, K.; Ragunathan, S.; Rosfjord, E.; Dann, S.; Upeslacis, E.; Grillo, M.; Hernandez, R.; Mack, F.; Klippel, A. The Interaction of PKN3 with RhoC Promotes Malignant Growth. Mol. Oncol. 2012, 6, 284–298. [Google Scholar] [CrossRef]
- London, M.; Gallo, E. The EphA2 and Cancer Connection: Potential for Immune-Based Interventions. Mol. Biol. Rep. 2020, 47, 8037–8048. [Google Scholar] [CrossRef]
- Oner, E.; Kotmakci, M.; Baird, A.M.; Gray, S.G.; Debelec Butuner, B.; Bozkurt, E.; Kantarci, A.G.; Finn, S.P. Development of EphA2 SiRNA-Loaded Lipid Nanoparticles and Combination with a Small-molecule Histone Demethylase Inhibitor in Prostate Cancer Cells and Tumor Spheroids. J. Nanobiotechnol. 2021, 19, 40. [Google Scholar] [CrossRef] [PubMed]
- Wagner, M.J.; Mitra, R.; Mcarthur, M.J.; Baze, W.; Barnhart, K.; Wu, S.Y.; Rodriguez-Aguayo, C.; Zhang, X.; Coleman, R.L.; Lopez-Berestein, G.; et al. Preclinical Mammalian Safety Studies of EPHARNA (DOPC Nanoliposomal EphA2-Targeted SiRNA). Mol. Cancer Ther. 2017, 16, 1114–1123. [Google Scholar] [CrossRef]
- Golan, T.; Khvalevsky, E.Z.; Hubert, A.; Gabai, R.M.; Hen, N.; Segal, A.; Domb, A.; Harari, G.; David, E.B.; Raskin, S.; et al. RNAi Therapy Targeting KRAS in Combination with Chemotherapy for Locally Advanced Pancreatic Cancer Patients. Oncotarget 2015, 6, 24560–24570. [Google Scholar] [CrossRef] [PubMed]
- Hunter, J.C.; Manandhar, A.; Carrasco, M.A.; Gurbani, D.; Gondi, S.; Westover, K.D. Biochemical and Structural Analysis of Common Cancer-Associated KRAS Mutations. Mol. Cancer Res. 2015, 13, 1325–1335. [Google Scholar] [CrossRef]
- Huang, L.; Guo, Z.; Wang, F.; Fu, L. KRAS Mutation: From Undruggable to Druggable in Cancer. Signal Transduct. Target. Ther. 2021, 6, 386. [Google Scholar] [CrossRef] [PubMed]
- Jani, V.; Sonavane, U.; Joshi, R. Insight into Structural Dynamics Involved in Activation Mechanism of Full Length KRAS Wild Type and P-Loop Mutants. Heliyon 2024, 10, e36161. [Google Scholar] [CrossRef]
- Baba, A.B.; Rah, B.; Bhat, G.R.; Mushtaq, I.; Parveen, S.; Hassan, R.; Hameed Zargar, M.; Afroze, D. Transforming Growth Factor-Beta (TGF-β) Signaling in Cancer-A Betrayal Within. Front. Pharmacol. 2022, 13, 791272. [Google Scholar] [CrossRef] [PubMed]
- Derynck, R.; Turley, S.J.; Akhurst, R.J. TGFβ Biology in Cancer Progression and Immunotherapy. Nat. Rev. Clin. Oncol. 2021, 18, 9–34. [Google Scholar] [CrossRef]
- Hashemi Goradel, N.; Najafi, M.; Salehi, E.; Farhood, B.; Mortezaee, K. Cyclooxygenase-2 in Cancer: A Review. J. Cell Physiol. 2019, 234, 5683–5699. [Google Scholar] [CrossRef]
- Li, S.; Jiang, M.; Wang, L.; Yu, S. Combined Chemotherapy with Cyclooxygenase-2 (COX-2) Inhibitors in Treating Human Cancers: Recent Advancement. Biomed. Pharmacother. 2020, 129, 110389. [Google Scholar] [CrossRef]
- Liu, S.; Zhang, Y.; Chen, Y.; Su, Y.; Chen, B.; Wang, Y.; Xu, M.; Qiao, K.; Li, S.; Liu, Z. Isolation and Purification of Protamine from the Cultured Takifugu Flavidus and Its Physicochemical Properties. Molecules 2024, 29, 263. [Google Scholar] [CrossRef]
- Massagué, J. TGFβ in Cancer. Cell 2008, 134, 215–230. [Google Scholar] [CrossRef]
- Chien, S.T.; Yang, T.F.; Yang, M.C.; Hsu, C.M.; Hong, Y.R.; Lee, T.M. Differential Roles of Bcl2L12 and Its Short Variant in Breast Cancer Lymph Node Metastasis. Oncol. Rep. 2015, 34, 961–971. [Google Scholar] [CrossRef]
- Kouri, F.M.; Jensen, S.A.; Stegh, A.H. The Role of Bcl-2 Family Proteins in Therapy Responses of Malignant Astrocytic Gliomas: Bcl2L12 and Beyond. Sci. World J. 2012, 2012, 1–8. [Google Scholar] [CrossRef]
- Kumthekar, P.; Ko, C.H.; Paunesku, T.; Dixit, K.; Sonabend, A.M.; Bloch, O.; Tate, M.; Schwartz, M.; Zuckerman, L.; Lezon, R.; et al. A First-in-Human Phase 0 Clinical Study of RNA Interference-Based Spherical Nucleic Acids in Patients with Recurrent Glioblastoma. Sci. Transl. Med. 2021, 13, eabb3945. [Google Scholar] [CrossRef] [PubMed]
- Stegh, A.H.; Brennan, C.; Mahoney, J.A.; Forloney, K.L.; Jenq, H.T.; Luciano, J.P.; Protopopov, A.; Chin, L.; DePinho, R.A. Glioma Oncoprotein Bc12L12 Inhibits the P53 Tumor Suppressor. Genes. Dev. 2010, 24, 2194–2204. [Google Scholar] [CrossRef]
- Apte, R.S.; Chen, D.S.; Ferrara, N. VEGF in Signaling and Disease: Beyond Discovery and Development. Cell 2019, 176, 1248–1264. [Google Scholar] [CrossRef]
- Ahmad, A.; Nawaz, M.I. Molecular Mechanism of VEGF and Its Role in Pathological Angiogenesis. J. Cell Biochem. 2022, 123, 1938–1965. [Google Scholar] [CrossRef]
- Kang, Y.; Li, H.; Liu, Y.; Li, Z. Regulation of VEGF-A Expression and VEGF-A-Targeted Therapy in Malignant Tumors. J. Cancer Res. Clin. Oncol. 2024, 150, 221. [Google Scholar] [CrossRef] [PubMed]
- Elebiyo, T.C.; Rotimi, D.; Evbuomwan, I.O.; Maimako, R.F.; Iyobhebhe, M.; Ojo, O.A.; Oluba, O.M.; Adeyemi, O.S. Reassessing Vascular Endothelial Growth Factor (VEGF) in Anti-Angiogenic Cancer Therapy. Cancer Treat. Res. Commun. 2022, 32, 100620. [Google Scholar] [CrossRef]
- Tabernero, J.; Shapiro, G.I.; LoRusso, P.M.; Cervantes, A.; Schwartz, G.K.; Weiss, G.J.; Paz-Ares, L.; Cho, D.C.; Infante, J.R.; Alsina, M.; et al. First-in-Humans Trial of an RNA Interference Therapeutic Targeting VEGF and KSP in Cancer Patients with Liver Involvement. Cancer Discov. 2013, 3, 406–417. [Google Scholar] [CrossRef] [PubMed]
- Tolcher, A.W.; Spira, A.I.; Nemunaitis, J.J.; Vandross, L.; Mamdani, H.; O’brien, Z.; Huynh, A.; Albaugh, Z.; Gullbo, J.; Zabludoff, S. First-in-Human Dose-Escalation Study of NBF-006, a Novel Investigational SiRNA Targeting GSTP, in Patients with Non-Small Cell Lung, Pancreatic, or Colorectal Cancer. J. Clin. Oncol. 2023, 41, 3084. [Google Scholar] [CrossRef]
- Kennedy, L.; Sandhu, J.K.; Harper, M.-E.; Cuperlovic-Culf, M. Role of Glutathione in Cancer: From Mechanisms to Therapies. Biomolecules 2020, 10, 1429. [Google Scholar] [CrossRef]
- Singh, R.R.; Reindl, K.M. Glutathione S-Transferases in Cancer. Antioxidants 2021, 10, 701. [Google Scholar] [CrossRef]
- Belete, T.M. The Current Status of Gene Therapy for the Treatment of Cancer. Biologics 2021, 15, 67–77. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Kang, G.; Wang, T.; Huang, H. Tumor Angiogenesis and Anti-angiogenic Gene Therapy for Cancer (Review). Oncol. Lett. 2018, 16, 687–702. [Google Scholar] [CrossRef]
- Younis, M.; Harashima, H. Understanding Gene Involvement in Hepatocellular Carcinoma: Implications for Gene Therapy and Personalized Medicine. Pharmgenomics Pers. Med. 2024, 17, 193–213. [Google Scholar] [CrossRef] [PubMed]
- Nussinov, R.; Yavuz, B.R.; Jang, H. Molecular Principles Underlying Aggressive Cancers. Signal Transduct. Target. Ther. 2025, 10, 42. [Google Scholar] [CrossRef]
- Man, R.C.H.; Qiu, Y.; Leung, S.W.S.; Fruhwirth, G.O.; Lam, J.K.W. Co-Delivery of PD-L1- and EGFR-Targeting SiRNAs by Synthetic PEG12-KL4 Peptide to the Lungs as Potential Strategy against Non-Small Cell Lung Cancer. Eur. J. Pharm. Biopharm. 2024, 195, 114177. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.J.; Kim, H.; Gao, X.; Ryu, J.H.; Yang, Y.; Kwon, I.C.; Roberts, T.M.; Kim, S.H. Multi-Targeting SiRNA Nanoparticles for Simultaneous Inhibition of PI3K and Rac1 in PTEN-Deficient Prostate Cancer. J. Ind. Eng. Chem. 2021, 99, 196–203. [Google Scholar] [CrossRef]
- Jamalinia, M.; Weiskirchen, R. Advances in Personalized Medicine: Translating Genomic Insights into Targeted Therapies for Cancer Treatment. Ann. Transl. Med. 2025, 13, 18. [Google Scholar] [CrossRef]
- Sousa, C.; Videira, M. Dual Approaches in Oncology: The Promise of SiRNA and Chemotherapy Combinations in Cancer Therapies. Oncology 2025, 5, 2. [Google Scholar] [CrossRef]


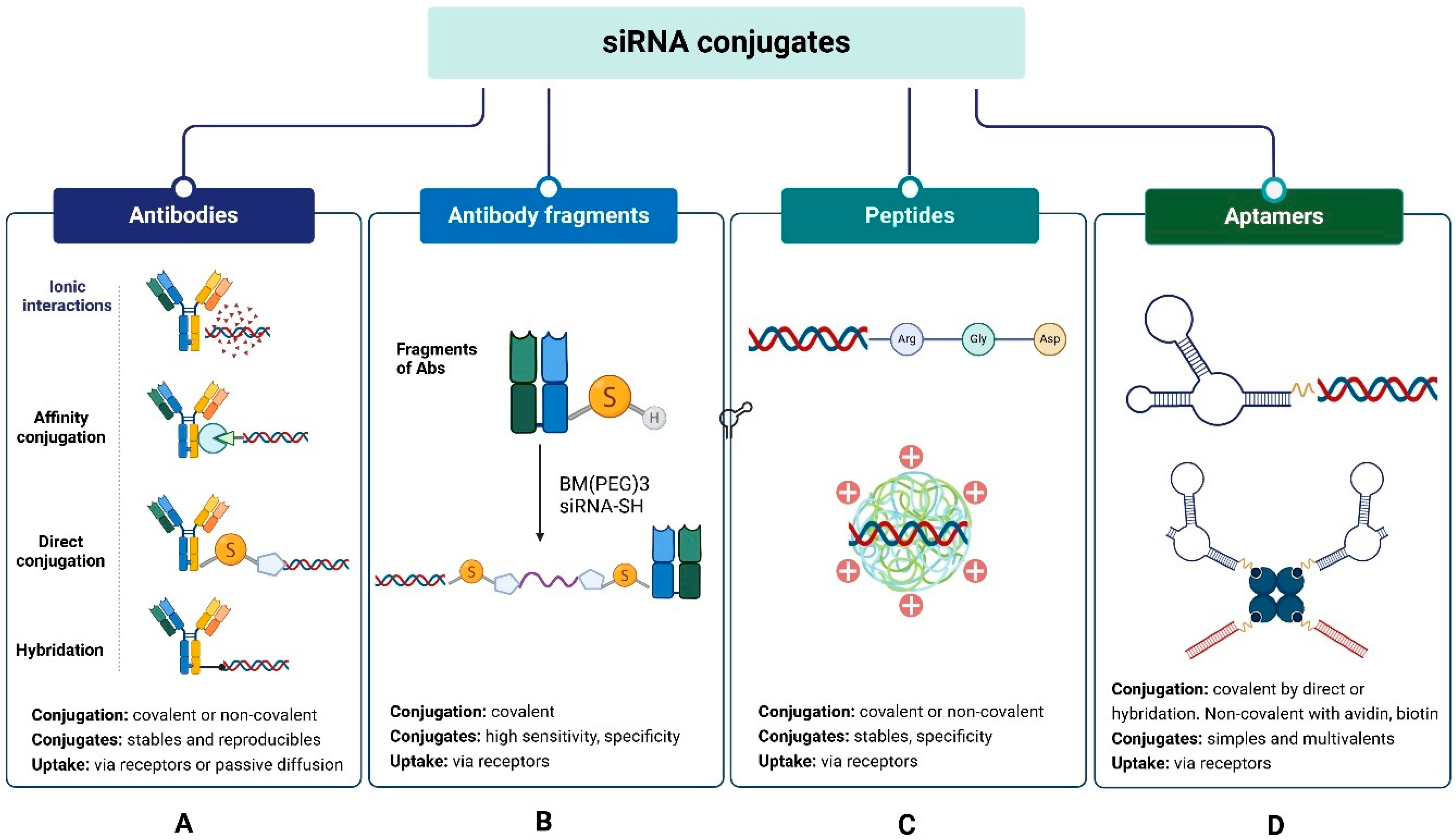

| Tool | URL | Reference |
|---|---|---|
| DuplexFold | https://rna.urmc.rochester.edu/RNAstructureWeb/Servers/DuplexFold/DuplexFold.html | [24] |
| MysiRNA-Designer | https://sourceforge.net/projects/mysirna/ | [25] |
| Genscript Package | https://www.genscript.com/sslbin/app/rnai | [19] |
| ThermoFisher BLOCK-It RNAi Designer | https://rnaidesigner.thermofisher.com/rnaiexpress/ | [19] |
| siRNA design | https://www.idtdna.com/page | [19] |
| siDESIGN Center | https://horizondiscovery.com/en/ordering-and-calculation-tools/sidesign-center | [26] |
| BLAST | https://blast.ncbi.nlm.nih.gov/Blast.cgi | [26] |
| SMEPred | https://bioinfo.imtech.res.in/manojk/smepred/ | [26] |
| Modification | Advantage | Disadvantage | Application | References |
|---|---|---|---|---|
| Modifications to the rest of the ribose | ||||
| Substitution of an -OH group by less nucleophilic groups: 2′-O-Me, 2′-O-MOE, 2′-F | Increases nuclease resistance, specificity, and affinity for the target mRNA. Decreases immunogenicity | Decreases the power of the silencing | Biomolecule and small molecule conjugates | [11,31,45] |
| LNA training | Increases stability and specificity. Decreases toxicity | The extent and site of modification influence the physico-chemical and biological properties | Conjugates with biomolecules and small molecules | [46] |
| UNA training | Increases silencing potency and structural flexibility. Decreases toxicity. | Physico-chemical and biological properties depend on the extent and site of modification. | Conjugates to biomolecules and small molecules | [46,47,48] |
| Modifications to the phosphate backbone | ||||
| Substitution of a non-bridging oxygen of the phosphodiester by a sulfur to form a PS | Increases nuclease resistance, improves pharmacokinetics, and cell internalization | Efficacy and toxicity depend on the extent and site of the modification | Conjugates with biomolecules and small molecules. Incorporation into transport nanosystems | [36,51,56] |
| Substitution of a non-bridging oxygen by phosphate derivatives | Increases stability, reduces off-target effects and toxicity | Effectiveness depends on the extent and site of the modification | Conjugates with biomolecules and small molecules. Incorporation into transport nanosystems. | [54,58,65] |
| Nucleobase modifications | ||||
| Nucleobase modifications | Increases thermal stability, power, and selectivity | The properties can be worsened depending on the base being modified and the type of modification. Unnatural residues could be incorporated into the genome. | [11,67,69] | |
| Multiple modifications | ||||
| A combination of ribose and backbone modifications | Improve biological properties and silencing efficacy. | ---- | ---- | [37,54,62] |
| Parameter | LDL | HDL |
|---|---|---|
| Diameter (nm) | 18–25 | 5–12 |
| Density (g/mL) | 1.006–1.019 | 1.063–1.210 |
| Molecular weight (Da) | 2.3 × 106 | (0.17–0.36) × 106 |
| Origin | VLDL/IDL | Liver and intestine |
| Composition | ||
| Triglycerides (%) | 10–15 | 2–3 |
| Free cholesterol (%) | 8–10 | 3–5 |
| Cholesterol esters (%) | 37–48 | 14–18 |
| Phospholipids (%) | 19–21 | 17–24 |
| Apolipoproteins (%) | Apo B-100: (20–22) | Apo A-I, Apo A-II, Apo A-IV, Apo A-V, Apo C-I, Apo C-II, Apo C-III, Apo E, and others: (~10) |
| Carrier System | Delivery Strategy | siRNA | Effect of Gene Silencing on Cancer Signaling Pathways | Combined Therapy | Type of Cancer | Advantages | Reference |
|---|---|---|---|---|---|---|---|
| Single treatments | |||||||
| siPD-L1@PNPs-sTN145 | PLGA-based polymeric nanoparticles functionalized with sTN145 aptamer | siRNA targeting programmed cell death-ligand 1 (siPD-L1) | Disabling tumor immune evasion to enable T cell–mediated cancer destruction | - | Triple-negative breast cancer (TNBC) | siPD-L1@PNPs-sTN145 effectively delivered PD-L1 siRNA into TNBC target cells. | [504] |
| Chkα siRNA PEG-PEI NPs | Polymeric nanoparticles with Polyethylene glycol (PEG) conjugated polyethylenimine (PEI) | siRNA targeting Choline kinase alpha (Chkα) | Inhibition of malignant transformation and tumor progression | - | Triple negative breast cancer (TNBC) | A significant decrease in Chkα and phosphocholine was achieved in xenografts with VEGF overexpression. Demonstrated importance of tumor vascularization in achieving effective siRNA delivery | [505] |
| P@P EPs (PFOB/PCX/siNGF) /emulsion polyplexes (EPs) | Cationic perfluorocarbon nanoemulsions Perfluorooctyl bromide (PFOB)/Polymeric CXCR4 antagonists (PCX)/siNGF | siRNA targeting nerve growth factor | Inhibition of tumor nerve fiber formation related remodeling tumor microenvironment (TME) and promotion of the proliferation, metastasis, and chemoresistance | - | Pancreatic cancer (PC) | P@P Eps shows better tumor penetration, together with siNGF shows enhanced gene inhibition in vitro and in vivo, by inhibition of tumor nerve fiber formation reducing metastatic potential. Leading to the effective and safe suppression of tumor growth in orthotopic PC. | [506] |
| C12-DMA lipoplexes | Cationic lipoplexes with cholesterol/PEG/ 3,4-dimethoxyaniline lipid (DMA) | siRNA targeting Survivin | Promoted cell apoptosis | - | Breast cancer | A quaternary amine-based liposome with DMA cationic lipoplexes, successfully delivered siRNA to reduce surviving mRNA expression, indicating potential for siRNA therapeutics in breast cancer treatment. | [507] |
| Nanocarriers NCAPAMAM/siRNA polycaprolactone-graft-poly(amidoamine) (PCL-g-PAMAM) | Polymeric dendrimer nanocarrier | siRNA targeting PLK-1 siRNA targeting PD-L1 | siRNA-PLK1 blocked proliferation; siRNA-PD-L1 boosted immunity. | - | Breast cancer and colorectal cancer | The nanoconfined loading strategy enhanced the electrostatic interaction between siRNA, and the cationic moieties of PAMAM. The system improved tumor cell uptake by releasing the complex in the acidic tumor microenvironment. The system enabled effective PD-L1 silencing. | [508] |
| Multimodal treatments | |||||||
| AS1411/Lipo-PTX-siPLK1 | Cationic liposomes functionalized with AS1411 aptamers. It specifically binds to nucleolin, overexpressed on the surface of tumor cells. | siRNA targeting Polo-like kinase 1 (PLK-1) | Inhibition of cell cycle progression | Chemotherapy with Paclitaxel (PTX): induces mitotic arrest via microtubule targeting. | Breast cancer | Paclitaxel and PLK1-targeted siRNA using AS1411 aptamer-functionalized cationic liposomes. Functionalized with a targeting aptamer (AS1411) to further enhance targeting and tumor accumulation. The PLK-1-siRNA reduces resistance PTX. The combined therapy increased the number of apoptotic cells and reduced angiogenesis, limiting the progression of breast cancer. | [509] |
| PEGylated DC-Chol/DOPE | Cationic Liposomes loaded with paclitaxel | siRNA targeting kinesin spindle protein (KSP) | Cell cycle arrest during mitosis | Chemotherapy with PTX | Ovarian cancer | PEG (Polyethylene glycol) prolonged circulation time in the blood by preventing potential aggregation of cationic liposomes and avoiding nonspecific uptake by the reticuloendothelial system. DOPE (Dioleoylphosphatidylethanolamine) improved endosomal escape by destabilizing the endosomal membrane. PTX suppressed Kif15 motility, which overcame KSP resistance and enhanced the effectiveness of PTX. | [510] |
| RDPP(Met)/TMZ/siMGMT | Hypoxia-radiosensitive nanoparticle | siRNA targeting O6-methylguanine-DNA-methyltransferase (siMGMT) | Inhibition of DNA-damage repair | Chemotherapy with Temozolomide (TMZ) & Radiotherapy | Glioblastoma (GBM) | A hypoxia-radiosensitive nanoparticle for co-delivering TMZ and siMGMT was synthesized to sensitize chemotherapy and radiotherapy for glioblastoma (GBM). Efficient combinatorial GBM therapy. Penetrate the blood–brain barrier, target GBM cells and effectively inhibit GBM proliferation. Effective therapy for overcoming temozolomide resistance in GBM. Together enhanced the sensitivity of TMZ as well as radiotherapy. | [511] |
| AuPEI-HA-DOX/siRNA | Gold nanoparticles | siRNA targeting CXCR4 | Inhibition of tumor microenvironment (TME) remodeling and suppression of proliferation, metastasis, and chemoresistance. | Phototherma therapy via irradiation AuNP & Chemotherapy with doxorubicin | Breast cancer | Gold nanoparticles with low-molecular weight polyethyleneimine (PEI) and hyaluronic acid (HA) transported doxorubicin (DOX) and siRNA. Combination of gene silencing, photothermal therapy and chemotherapy increased cytotoxic effect on cancer cells. | [512] |
| HECP2k/doxorubicin/Bcl-2 siRNA | Polymeric nanoparticles Polyethylenimine2k (PEI2k)-conjugated to hydroxyethyl cellulose (HECP2k) | siRNA targeted Bcl-2 (B-cell lymphoma 2) | Apoptosis induction | Chemotherapy with Doxorubicin | Cervical carcinoma | Endosome buffering ability of PEI2k and osmotic stress caused by HEC, facilitated cellular uptake and endosome escape, leading to efficient transfection. Induced apoptosis, enhanced chemosensitivity, high siRNA loading capacity, controlled release of siRNA. Improved tumor regression and chemosensitivity. | [513] |
| mHDL(CS/siRNA) | Mimetic high-density lipoproteins were prepared with Apo-A1 mimetic peptides and chitosan/siRNA complex | siRNA targeted RAD51 | Inhibition of DNA repair | Radionuclide therapy with DOTA-Bombesin labeled with Actinium-225 | Breast cancer | The system proved to be stable in serum, provided protection against RNases, and transfects via specific recognition by SR-B1. mHDL(CS/siRNA-RAD51) pretreatment prevented homologous repair of DBSs caused by 225Ac-DOTA-Bombesin, enhancing the therapeutic response. | [514] |
| SNEDDS (siRNA-RAD51) | Self-nanoemulsifying systems (SNEDDS) based on Phospholipon-90G, Labrafil-M1944-CS and Cremophor-RH40. In the strategy a chitosan/siRNA complex was used. | siRNA targeted RAD51 | Inhibition of DNA repair to promote cancer sensitivity | Radionuclide therapy with Lutetium-177 or Photodynamic Therapy with doxorubicin irradiated with 450 nm | Breast cancer | SNEDDS (siRNA-RAD51) yielded an emulsion of 50 nm and good homogeneity (PDI 0.41) capable of efficiently transfecting siRNA-RAD51 into T47D cells. The SNEDDS(siRNA-RAD51) pretreatment in PDT or radionuclide therapy increased cell sensitivity. | [261,349] |
| Target | Candidate | Family/Type | Main Function | Alterations in Cancer | Effect of Inhibition | Reference |
|---|---|---|---|---|---|---|
| Phase II completed | ||||||
| PLK1 | TKM-080301 (CTN: NCT01262235; NCT02191878) | Serine/threonine kinase (polokinase) | Regulates several phases of the cell cycle | Overexpressed in multiple cancers; associated with poor prognosis, chemo- and radioresistance | Induces apoptosis | [7,519,520,521] |
| PKN3 | Atu027 (CTN: NCT01808638) | Protein kinase | Proliferation, migration, invasion and angiogenesis | Overexpressed in different cancers | Prevents tumor growth, metastasis and induces apoptosis | [7,12,522,523,524,525] |
| Phase II in progress | ||||||
| EphA2 | siRNA-EphA2-DOPC (CTN: NCT01591356) | Receptor tyrosine kinase | Proliferation, survival and differentiation | Overexpressed in several cancers; almost absent in healthy tissues; associated with high malignancy, metastasis and low survival | Inhibits carcinogenesis and angiogenesis; promotes apoptosis | [11,12,526,527,528] |
| KRAS G12 D mutation | siG12D LODER (CTN: NCT01676259) | Oncogene of the RAS family | Signaling for growth, differentiation, survival and apoptosis | Frequent mutations in cancer (codon 12: lung, colon, pancreas); G12D predominant in PDAC; conferring therapy resistance | Specific inhibition (e.g., G12D) blocks proliferation and tumor survival | [7,8,12,529,530,531,532] |
| TGF-β1, COX-2 | STP705 (CTN: NCT04844983) | Cytokine (TGF-β1) Membrane enzyme (COX-2) | TGF-β1: homeostasis, apoptosis; COX-2: carcinogenesis, angiogenesis, resistance | Dysregulated TGF-β1 → epithelial–mesenchymal transition, proliferation, immune evasion; COX-2 overexpressed in aggressive tumors | Co-inhibition blocks tumor progression, angiogenesis and metastasis | [7,8,12,533,534,535,536,537,538] |
| Phase I completed | ||||||
| BCL2L12 | NU-0129 (CTN: NCT03020017) | Anti-apoptotic oncoprotein | Potent apoptosis inhibitor in glial cells | Overexpressed in various cancers; poor prognosis and therapy resistance | Inhibition restores apoptosis in tumor cells | [7,12,539,540,541,542] |
| VEGF and KSP | ALN-VSP02 (CTN: NCT01158079) | VEGF: glycoprotein RTK; KSP: microtubule-associated motor protein | VEGF: angiogenesis; KSP: mitosis and proliferation | VEGF overexpressed by tumor hypoxia → metastasis; KSP overexpressed in mitotic tumors | Co-inhibition reduces proliferation | [7,543,544,545,546,547] |
| Phase I in progress | ||||||
| GSTP | NBF-06 (CTN: NCT03819387) | Detoxifying enzyme (glutathione S-transferase) | Regulation of cellular oxidation, proliferation and apoptosis | Overexpressed in aggressive tumors; contributes to therapeutic resistance | Inhibition favors apoptosis and sensitizes to therapy | [7,548,549,550] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aranda-Lara, L.; Escudero-Castellanos, A.; Trujillo-Nolasco, M.; Morales-Avila, E.; Ocampo-García, B.; Oros-Pantoja, R.; Sánchez-Monroy, V.; Isaac-Olivé, K. Small Interfering RNA Carriers for Oncotherapy: A Preclinical Overview. Pharmaceutics 2025, 17, 1408. https://doi.org/10.3390/pharmaceutics17111408
Aranda-Lara L, Escudero-Castellanos A, Trujillo-Nolasco M, Morales-Avila E, Ocampo-García B, Oros-Pantoja R, Sánchez-Monroy V, Isaac-Olivé K. Small Interfering RNA Carriers for Oncotherapy: A Preclinical Overview. Pharmaceutics. 2025; 17(11):1408. https://doi.org/10.3390/pharmaceutics17111408
Chicago/Turabian StyleAranda-Lara, Liliana, Alondra Escudero-Castellanos, Maydelid Trujillo-Nolasco, Enrique Morales-Avila, Blanca Ocampo-García, Rigoberto Oros-Pantoja, Virginia Sánchez-Monroy, and Keila Isaac-Olivé. 2025. "Small Interfering RNA Carriers for Oncotherapy: A Preclinical Overview" Pharmaceutics 17, no. 11: 1408. https://doi.org/10.3390/pharmaceutics17111408
APA StyleAranda-Lara, L., Escudero-Castellanos, A., Trujillo-Nolasco, M., Morales-Avila, E., Ocampo-García, B., Oros-Pantoja, R., Sánchez-Monroy, V., & Isaac-Olivé, K. (2025). Small Interfering RNA Carriers for Oncotherapy: A Preclinical Overview. Pharmaceutics, 17(11), 1408. https://doi.org/10.3390/pharmaceutics17111408









