Antiproliferative Copper(II) Complexes Bearing Mixed Chelating Ligands: Structural Characterization, ROS Scavenging, In Silico Studies, and Anti-Melanoma Activity
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials and Physical Measurements
2.2. Synthesis of Complexes
2.3. Crystal Structural Analysis
2.4. In Vitro Cytotoxicity Assay
2.4.1. Cell Culture Conditions
2.4.2. Cell Viability Assay
2.4.3. Cell Cycle Analysis
2.5. Computational Strategy
2.5.1. Molecular Modeling of Complexes
2.5.2. Assessment of Compounds’ Drug- and Lead-Likeness Features
2.5.3. Computational Pharmacokinetics and Pharmacogenomics Profiles of Copper(II) Complexes
2.5.4. Compounds’ Computational Pharmacodynamic Profiles
3. Results and Discussion
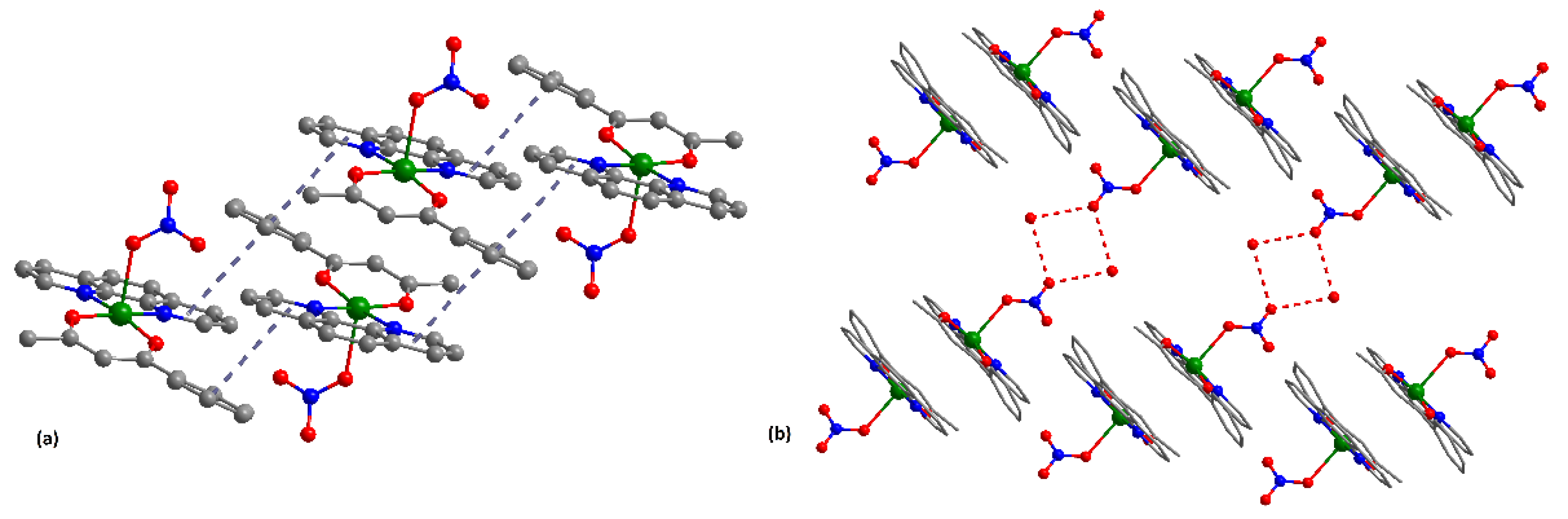
3.1. Description of the Crystal Structure of Complex (2)
3.2. Physico-Chemical Characterization of Complexes
3.2.1. FT-IR Spectra
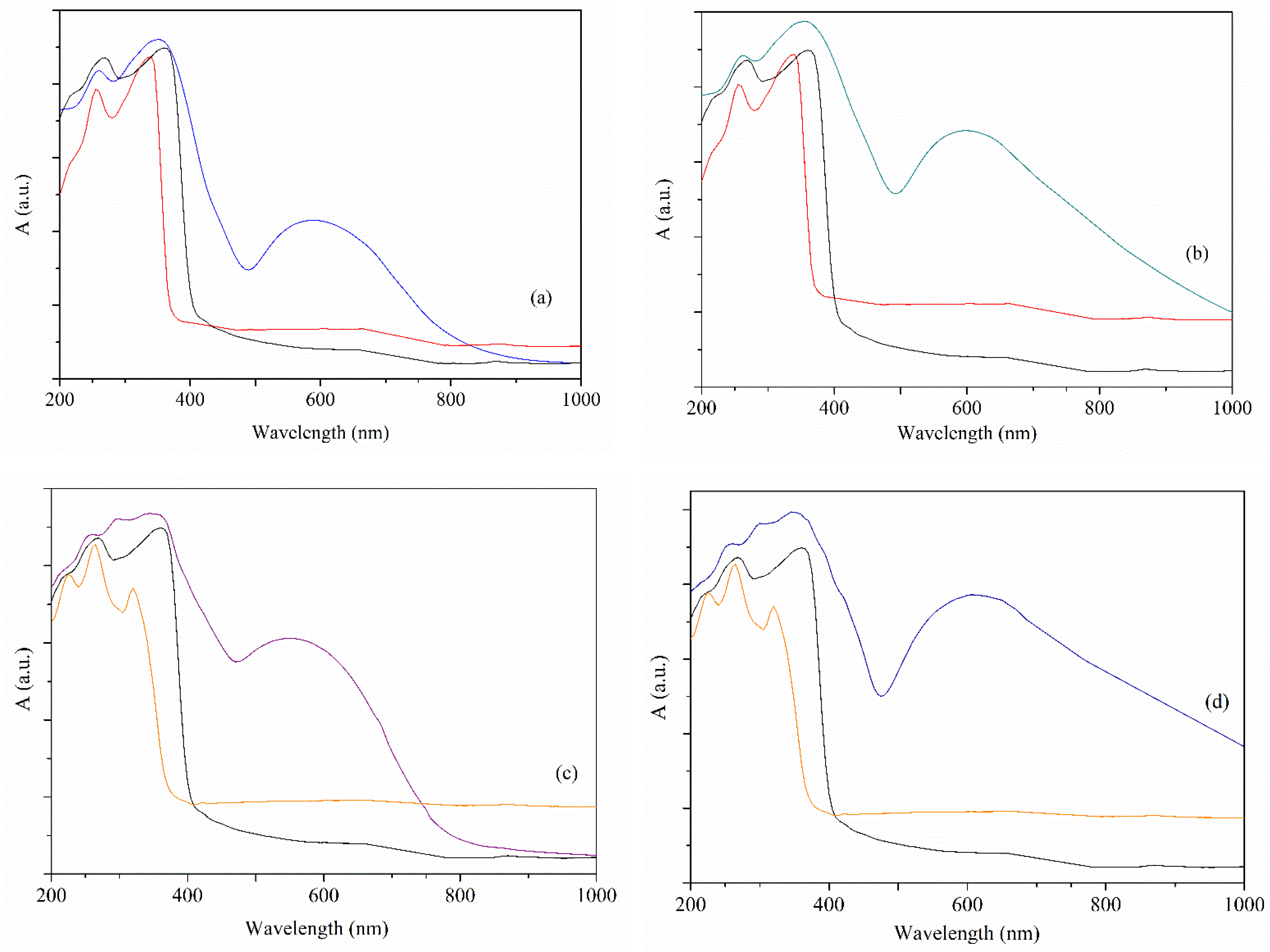
3.2.2. UV-Vis Spectra
3.2.3. EPR Spectroscopy
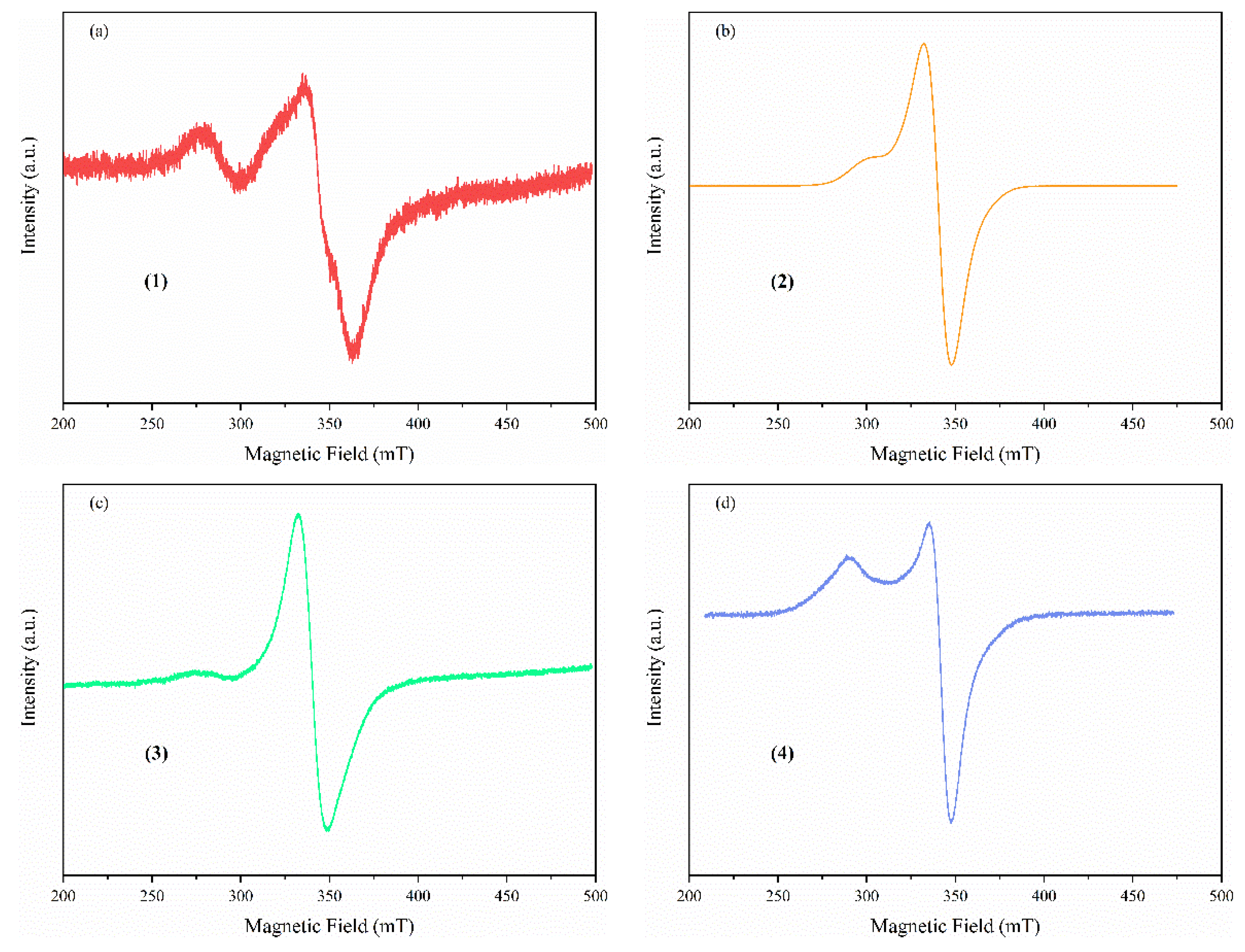
Solid State EPR Spectroscopy
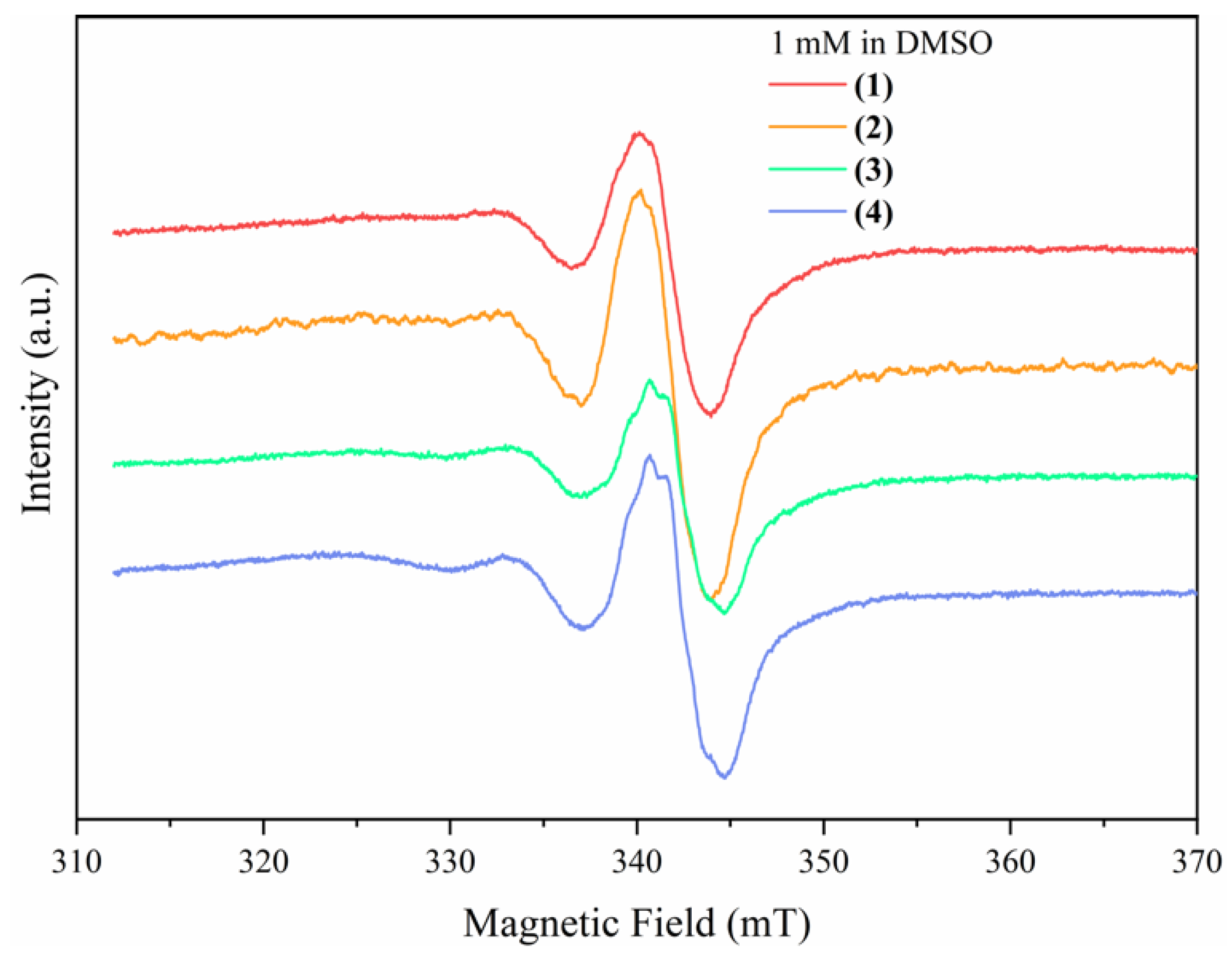
Solution EPR Spectroscopy
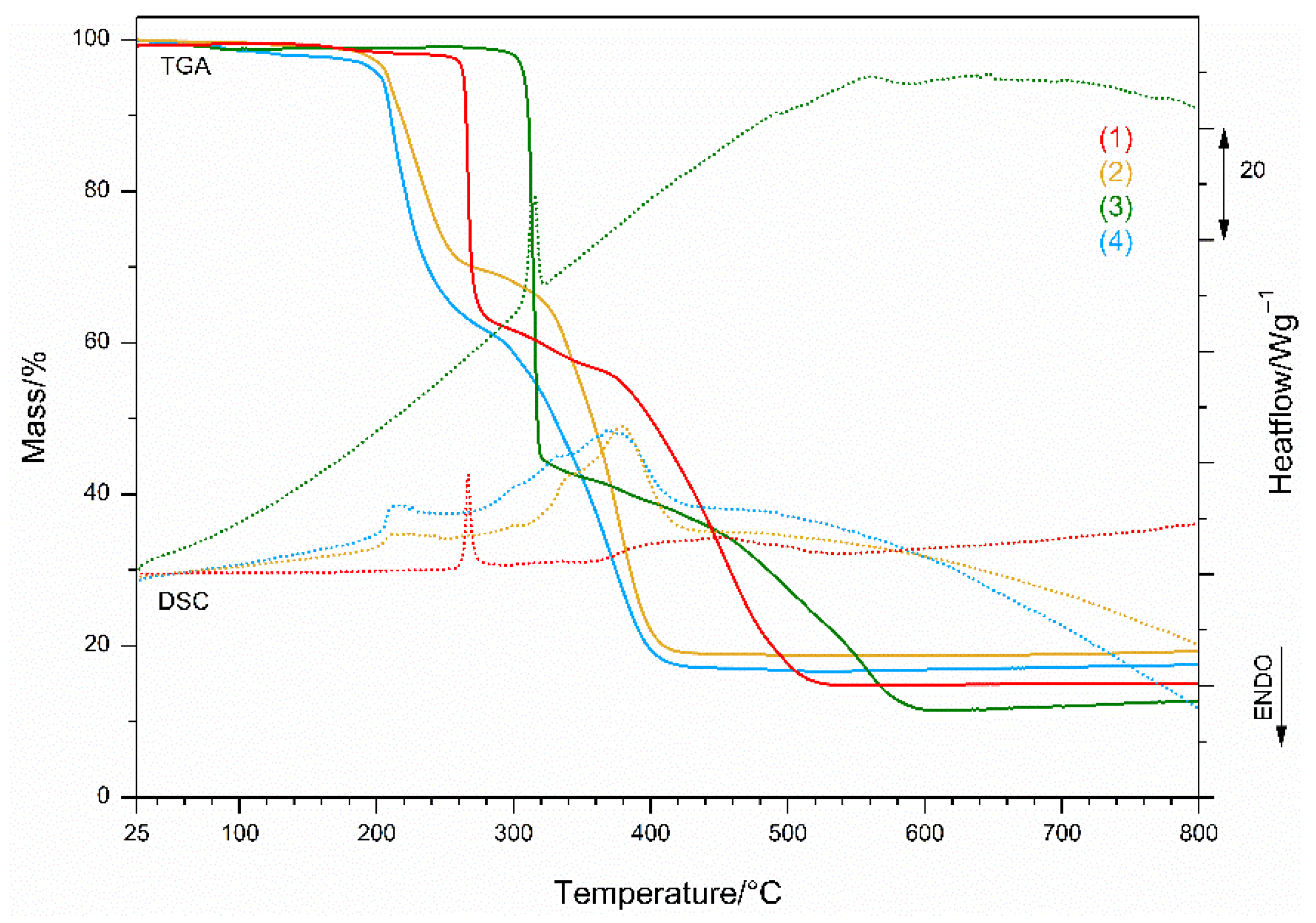
3.2.4. Thermal Behavior
3.3. Complexes Interaction with Cells and Biological Species
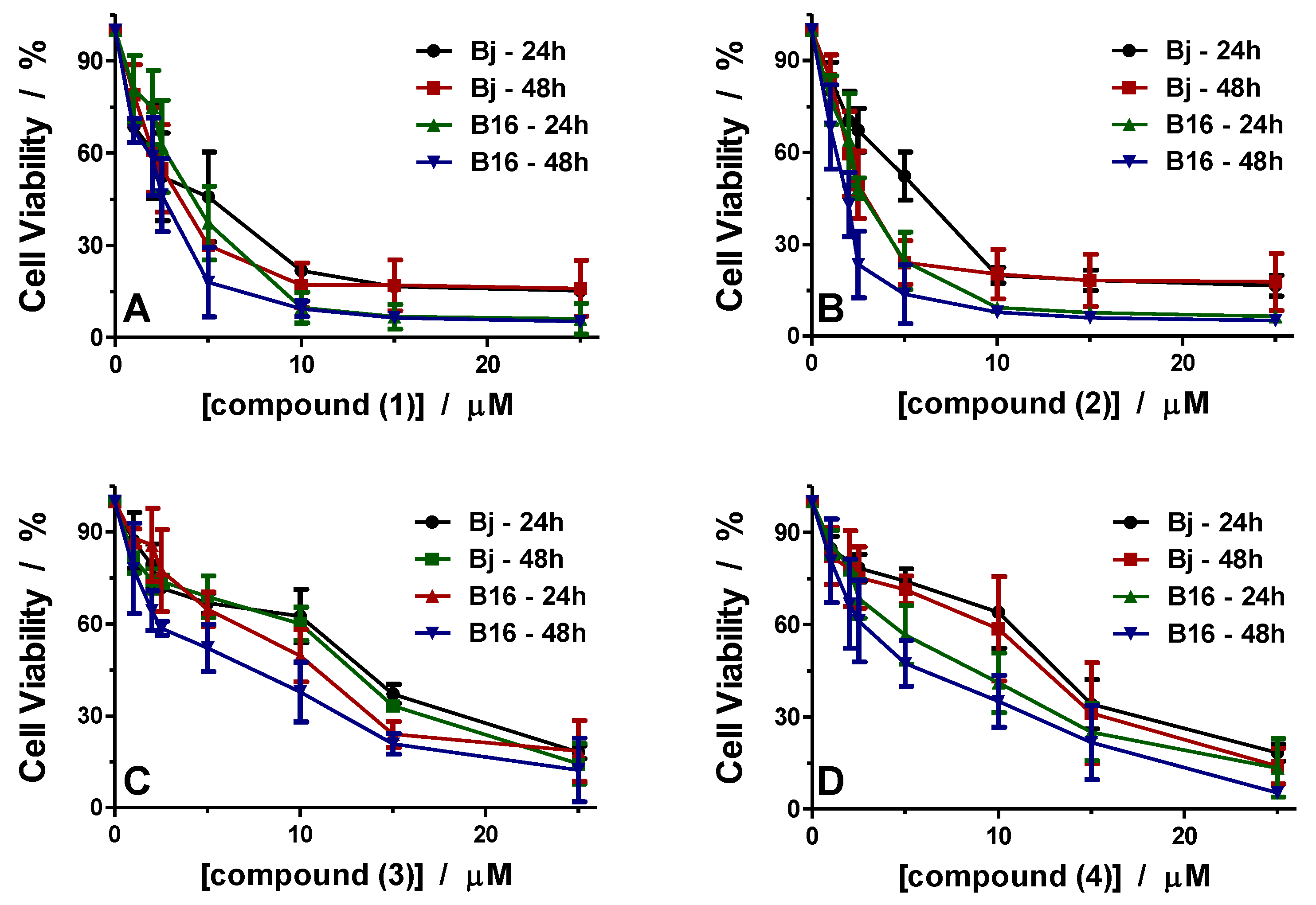
3.3.1. Antiproliferative Activity
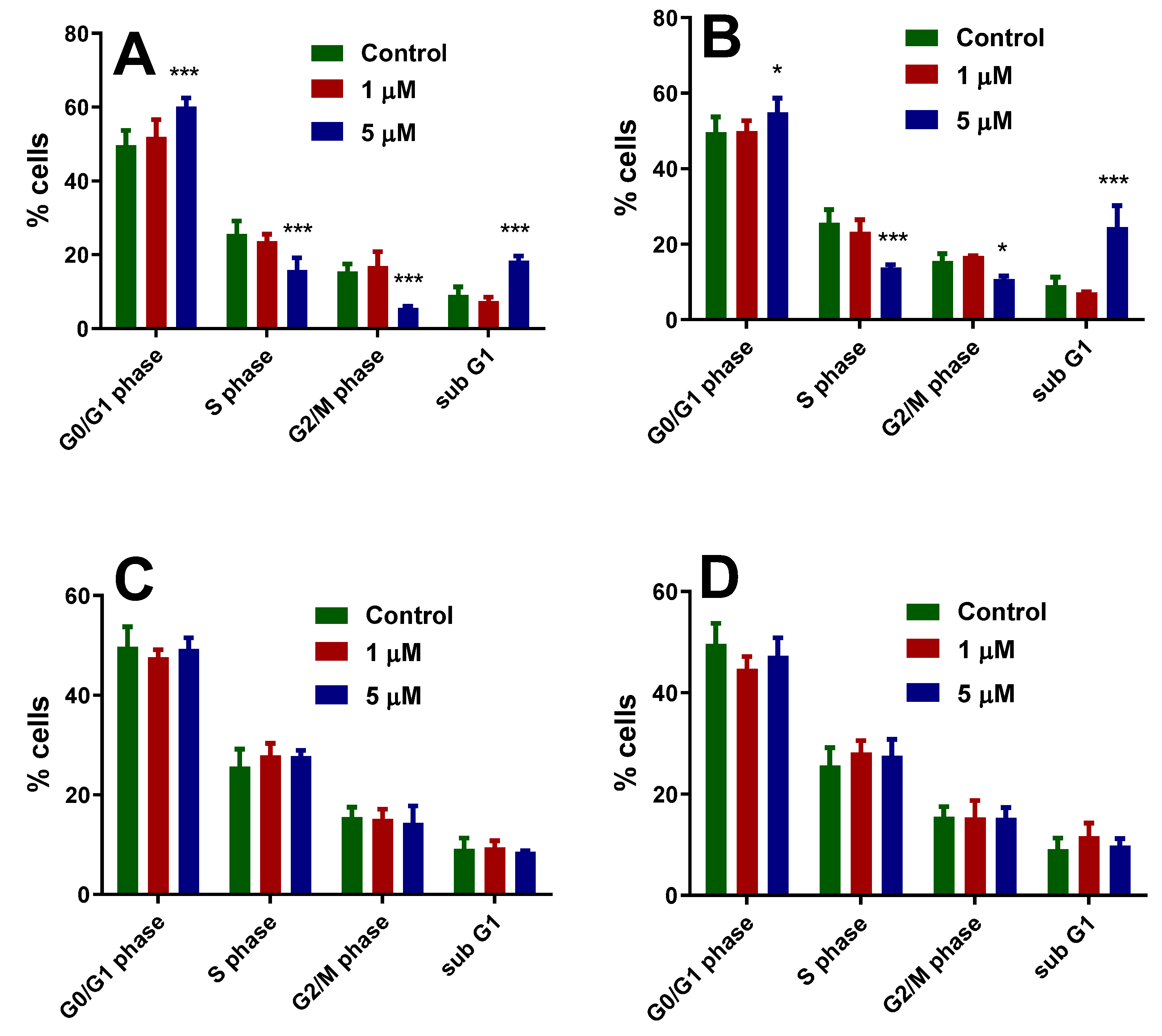
3.3.2. Effect of Copper(II) Complexes on Cell Cycle Distribution
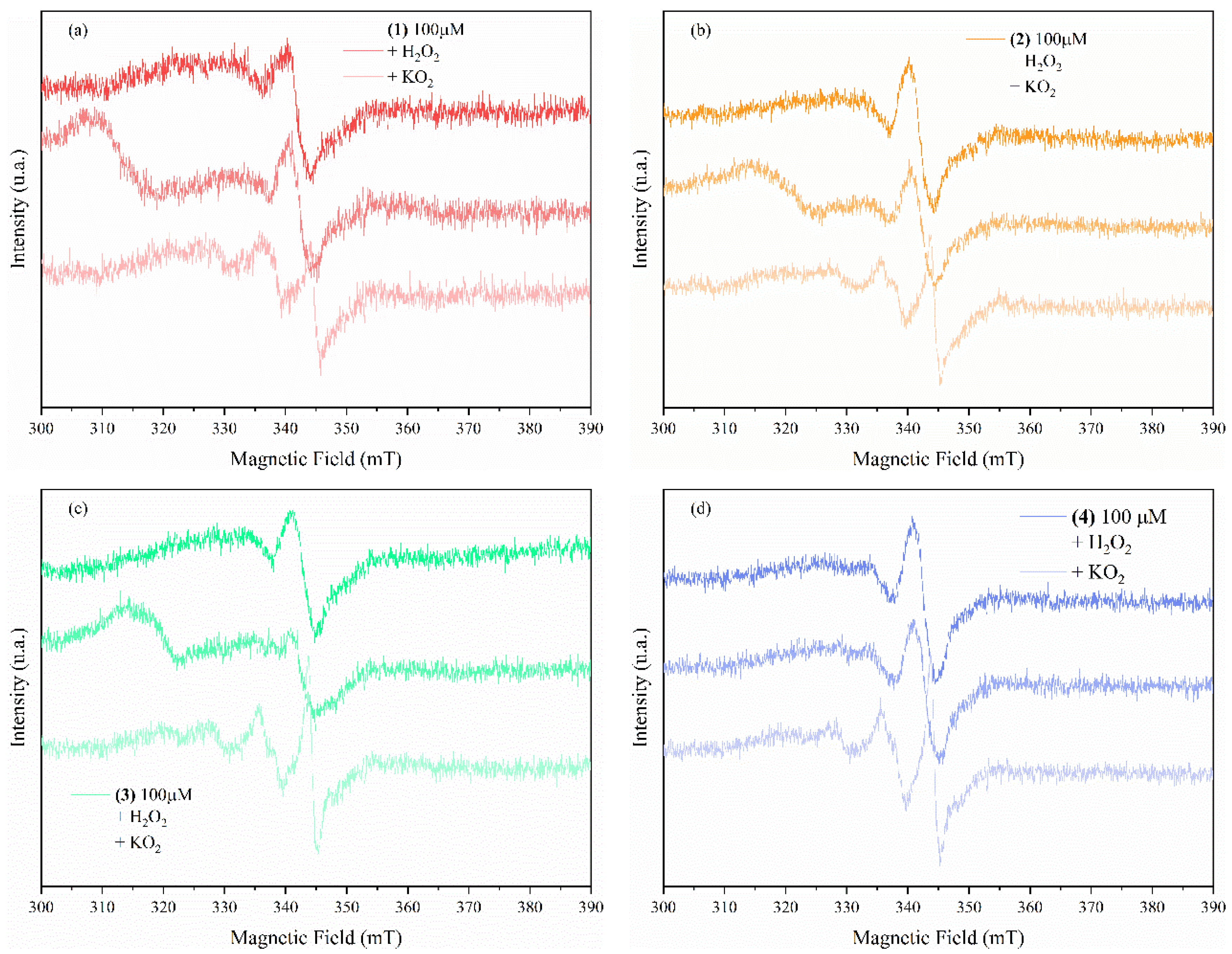
3.3.3. Complexes Interaction with ROS
3.4. In Silico Studies
3.4.1. Drug-Likeness, Pharmacokinetics, and Pharmacogenomics Profiles of the Compounds
3.4.2. Computational Pharmacodynamic Profiles of the Complexes
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Manzano, C.M.; Nakahata, D.H.; de Paiva, R.E.F. Revisiting metallodrugs for the treatment of skin cancers. Coord. Chem. Rev. 2022, 462, 214506. [Google Scholar] [CrossRef]
- Sander, C.S.; Hamm, F.; Elsner, P.; Thiele, J.J. Oxidative stress in malignant melanoma and non-melanoma skin cancer. Br. J. Dermatol. 2003, 148, 913–922. [Google Scholar] [CrossRef]
- Gupta, A.; Gomes, F.; Lorigan, P. The role for chemotherapy in the modernmanagement of melanoma. Melanoma Manag. 2017, 4, 125. [Google Scholar] [CrossRef] [PubMed]
- Denoyer, D.; Masaldan, S.; La Fontaine, S.; Cater, M.A. Targeting copper in cancer therapy: ‘Copper That Cancer’. Metallomics 2015, 7, 1459–1476. [Google Scholar] [CrossRef]
- Leon, I.E.; Cadavid-Vargas, J.F.; Di Virgilio, A.L.; Etcheverry, S.B. Vanadium, Ruthenium and Copper Compounds: A New Class of Nonplatinum Metallodrugs with Anticancer Activity. Curr. Med. Chem. 2017, 24, 112–148. [Google Scholar] [CrossRef]
- Anthony, E.J.; Bolitho, E.M.; Bridgewater, H.E.; Carter, O.W.L.; Donnelly, J.M.; Imberti, C.; Lant, E.C.; Lermyte, F.; Needham, R.J.; Palau, M.; et al. Metallodrugs are unique: Opportunities and challenges of discovery and development. Chem. Sci. 2020, 11, 12879–13154. [Google Scholar] [CrossRef] [PubMed]
- Badea, M.; Uivarosi, V.; Olar, R. Improvement in the Pharmacological Profile of Copper Biological Active Complexes by Their Incorporation into Organic or Inorganic Matrix. Molecules 2020, 25, 5830. [Google Scholar] [CrossRef]
- Balsa, L.M.; Baran, E.J.; León, I.E. Copper complexes as antitumor agents: In vitro and in vivo evidences. Curr. Med. Chem. 2021. [Google Scholar] [CrossRef]
- Hasinoff, B.B.; Yadav, A.A.; Patel, D.; Wu, X. The cytotoxicity of the anticancer drug elesclomol is due to oxidative stress indirectly mediated through its complex with Cu(II). J. Inorg. Biochem. 2014, 137, 22–30. [Google Scholar] [CrossRef]
- Křikavová, R.; Vančo, J.; Trávníček, Z.; Hutyra, J.; Dvořák, Z. Design and characterization of highly in vitro antitumor active ternary copper(II) complexes containing 2’-hydroxychalcone ligands. J. Inorg. Biochem. 2016, 163, 8–17. [Google Scholar] [CrossRef] [PubMed]
- Mroueh, M.; Daher, C.; Hariri, E.; Demirdjian, S.; Isber, S.; Choi, E.S.; Mirtamizdoust, B.; Hammud, H.H. Magnetic property, DFT calculation, and biological activity of bis [(μ2-chloro) chloro (1,10-phenanthroline)copper (II)] complex. Chem. Biol. Interact. 2015, 231, 53–60. [Google Scholar] [CrossRef] [PubMed]
- Hammud, H.H.; Kortz, U.; Bhattacharya, S.; Demirdjian, S.; Hariri, E.; Isber, S.; Sang Choi, E.; Mirtamizdoust, B.; Mroueh, M.; Daher, C.F. Structure, DFT studies, Magnetism and Biological activity of Bis[(μ-azido)-chloro-(1,10-phenanthroline)-copper(II)] complex. Inorg. Chim. Acta. 2020, 506, 119533. [Google Scholar] [CrossRef]
- Slator, C.; Molphy, Z.; McKee, V.; Long, C.; Brown, T.; Kellett, A. Di-copper metallodrugs promote NCI-60 chemotherapy via singlet oxygen and superoxide production with tandem TA/TA and AT/AT oligonucleotide discrimination. Nucleic Acids Res. 2018, 46, 2733–2750. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, M.; Suhaimi, S.-N.; Chu, T.-L.; Abdul Aziz, N.; Mohd Kornain, N.-K.; Samiulla, D.S.; Lo, K.-W.; Ng, C.-H.; Khoo, A.S.-B. Ternary copper(II) complex: NCI60 screening, toxicity studies, and evaluation of efficacy in xenograft models of nasopharyngeal carcinoma. PLoS ONE 2018, 13, 0191295. [Google Scholar] [CrossRef] [PubMed]
- Nakahata, D.H.; de Paiva, R.E.F.; Lustri, W.R.; Ribeiro, C.M.; Pavan, F.R.; da Silva, G.G.; Ruiz, A.L.T.G.; de Carvalho, J.E.; Corbi, P.P. Sulfonamide-containing copper(II) metallonucleases: Correlations with in vitro antimycobacterial and antiproliferative activities. J. Inorg. Biochem. 2018, 187, 85–96. [Google Scholar] [CrossRef] [PubMed]
- Olar, R.; Badea, M.; Bacalum, M.; Raileanu, M.; Ruta, L.L.; Farcasanu, I.C.; Rostas, A.M.; Vlaicu, I.D.; Popa, M.; Chifiriuc, M.C. Antiproliferative and antibacterial properties of biocompatible copper(II) complexes bearing chelating N,N-heterocycle ligands and potential mechanisms of action. Biometals 2021, 34, 1155–1172. [Google Scholar] [CrossRef] [PubMed]
- Rostas, A.M.; Badea, M.; Ruță, L.L.; Farcașanu, I.C.; Maxim, C.; Chifiriuc, M.C.; Popa, M.; Luca, M.; Celan Korošin, N.; Cerč Korošec, R.; et al. Copper(II) complexes with mixed heterocycle ligands as promising antibacterial and antitumor species. Molecules 2020, 25, 3777. [Google Scholar] [CrossRef]
- Ruta, L.L.; Farcasanu, I.C.; Bacalum, M.; Raileanu, M.; Rostas, A.M.; Daniliuc, C.G.; Chifiriuc, M.C.; Marutescu, L.; Popa, M.; Badea, M.; et al. Biological activity of triazolopyrimidine copper(II) complexes modulated by an auxiliary N-N-chelating heterocycle ligands. Molecules 2021, 26, 6772. [Google Scholar] [CrossRef]
- Ruan, B.-F.; Liang, Y.-K.; Liu, W.-D.; Wu, J.-Y.; Tian, Y.-P. Synthesis, characterization, and antitumor activities of two copper(II) complexes with pyrazole derivatives. J. Coord. Chem. 2012, 65, 2127–2134. [Google Scholar] [CrossRef]
- Borges, L.J.H.; Bull, É.S.; Fernandes, C.; Horn, A.; Azeredo, N.F.; Resende, J.A.L.C.; Freitas, W.R.; Carvalho, E.C.Q.; Lemos, L.S.; Jerdy, H.; et al. In vitro and in vivo studies of the antineoplastic activity of copper (II) compounds against human leukemia THP-1 and murine melanoma B16-F10 cell lines. Eur. J. Med. Chem. 2016, 123, 128–140. [Google Scholar] [CrossRef]
- Kalinowska-Lis, U.; Szabłowska-Gadomska, I.; Lisowska, K.; Ochocki, J.; Małecki, M.; Felczak, A. Cytotoxic and Antimicrobial Properties of Copper (II) Complexes of Pyridine and Benzimidazole Derivatives. Z. Anorg. Allg. Chem. 2017, 643, 993–998. [Google Scholar] [CrossRef]
- Mariani, D.; Ghasemishahrestani, Z.; Freitas, W.; Pezzuto, P.; Costa-da-Silva, A.C.; Tanuri, A.; Kanashiro, M.M.; Fernandes, C.; Horn, A., Jr.; Pereira, M.D. Antitumoral synergism between a copper(II) complex and cisplatin improves in vitro and in vivo anticancer activity against melanoma, lung and breast cancer cells. Biochim. Biophys. Acta 2021, 1865, 129963. [Google Scholar] [CrossRef] [PubMed]
- Gurudevaru, C.; Gopalakrishnan, M.; Senthilkumar, K.; Hemachandran, H.; Siva, R.; Srinivasan, T.; Velmurugan, D.; Shanmugan, S.; Palanisami, N. Synthesis and structural and DNA binding studies of mono and dinuclear copper (II) complexes constructed with –O and –N donor ligands: Potential anti-skin cancer drugs. Appl. Organometal. Chem. 2017, 31, e3998. [Google Scholar] [CrossRef]
- Monti, E.; Paracchini, L.; Piccinini, F.; Malatesta, V.; Morazzoni, F.; Supino, R. Cardiotoxicity and antitumor activity of a copper (II)-doxorubicin chelate. Cancer Chemother. Pharmacol. 1990, 25, 333–336. [Google Scholar] [CrossRef]
- Eshaghi Malekshah, R.; Salehi, M.; Kubicki, M.; Khaleghian, A. New mononuclear copper(II) complexes from β-diketone and β-keto ester N-donor heterocyclic ligands: Structure, bioactivity and molecular simulation studies. J. Coord. Chem. 2018, 71, 952–968. [Google Scholar] [CrossRef]
- Zhang, L.; Xu, D.; Xu, Y.; Gu, J. (Benzoylacetonato-O,O’)(2,2’-bipyridine-N,N’)(nitrato-O)copper(II). Acta Cryst. 1997, C53, 299–301. [Google Scholar] [CrossRef]
- Avram, S.; Mernea, M.; Bagci, E.; Hritcu, L.; Borcan, L.C.; Mihailescu, D.F. Advanced Structure-activity Relationships Applied to Mentha spicata L. Subsp. spicata Essential Oil Compounds as AChE and NMDA Ligands, in Comparison with Donepezil, Galantamine and Memantine—New Approach in Brain Disorders Pharmacology. CNS Neurol. Disord. 2022, 16, 800–811. [Google Scholar] [CrossRef]
- Udrea, A.M.; Puia, A.; Shaposhnikov, S.; Avram, S. Computational approaches of new perspectives in the treatment of depression during pregnancy. Farmacia 2018, 66, 680–687. [Google Scholar] [CrossRef]
- Avram, S.; Bologa, C.; Flonta, M.L. Quantitative structure-activity relationship by CoMFA for cyclic urea and nonpeptidecyclic cyanoguanidine derivatives on the wild type and mutants HIV-1 protease. J. Mol. Model. 2005, 11, 105–115. [Google Scholar] [CrossRef]
- Sheldrick, G.M. SHELXL-97, Program for Crystal Structure Refinement; University of Göttingen: Göttingen, Germany, 1998. [Google Scholar]
- Bordei Telehoiu, A.T.; Nuță, D.C.; Căproiu, M.T.; Dumitrascu, F.; Zarafu, I.; Ioniță, P.; Bădiceanu, C.D.; Avram, S.; Chifiriuc, M.C.; Bleotu, C.; et al. Design, Synthesis and In Vitro Characterization of Novel Antimicrobial Agents Based on 6-Chloro-9H-carbazol Derivatives and 1,3,4-Oxadiazole Scaffolds. Molecules 2020, 25, 266. [Google Scholar] [CrossRef]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and Computational Approaches to Estimate Solubility and Permeability in Drug Discovery and Development Settings 1PII of Original Article: S0169-409X9600423-1. Adv. Drug Deliv. Rev. 2001, 46, 3–26. [Google Scholar] [CrossRef]
- Ghose, A.K.; Viswanathan, V.N.; Wendoloski, J.J. A Knowledge-Based Approach in Designing Combinatorial or Medicinal Chemistry Libraries for Drug Discovery. 1. A Qualitative and Quantitative Characterization of Known Drug Databases. J. Comb. Chem. 1999, 1, 55–68. [Google Scholar] [CrossRef] [PubMed]
- Veber, D.F.; Johnson, S.R.; Cheng, H.-Y.; Smith, B.R.; Ward, K.W.; Kopple, K.D. Molecular Properties That Influence the Oral Bioavailability of Drug Candidates. J. Med. Chem. 2002, 45, 2615–2623. [Google Scholar] [CrossRef] [PubMed]
- Egan, W.J.; Merz, K.M.; Baldwin, J.J. Prediction of Drug Absorption Using Multivariate Statistics. J. Med. Chem. 2000, 43, 3867–3877. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A Free Web Tool to Evaluate Pharmacokinetics, Drug-Likeness and Medicinal Chemistry Friendliness of Small Molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef]
- Available online: http://biosig.unimelb.edu.au/pkcsm/ (accessed on 22 June 2022).
- Available online: https://prediction.charite.de/ (accessed on 22 June 2022).
- Available online: https://prediction.charite.de/subpages/target_result.php (accessed on 22 June 2022).
- Madalan, A.M.; Avarvari, N.; Andruh, M. Metal complexes as second-sphere ligands. New J. Chem. 2006, 30, 521–523. [Google Scholar] [CrossRef]
- Dumitru, I.; Ene, C.D.; Ofiteru, A.M.; Paraschivescu, C.; Madalan, A.M.; Baciu, I.; Farcasanu, I.C. Identification of [CuCl(acac)(tmed)], a copper(II) complex with mixed ligands, as a modulator of Cu, Zn superoxide dismutase (Sod1p) activity in yeast. J. Biol. Inorg. Chem. 2012, 17, 961–974. [Google Scholar] [CrossRef]
- Nakamoto, K. Infrared and Raman Spectra of Inorganic and Coordination Compounds, Part B, Applications in Coordination, Organometallic, and Bioinorganic Chemistry, 6th ed.; John Wiley & Sons: Hoboken, NJ, USA, 2009; pp. 57–61. [Google Scholar]
- Hathaway, B.J. Oxyanions. In Comprehensive Coordination Chemistry, 1st ed.; Wilkinson, G., Gillard, R.D., McCleverty, J.A., Eds.; Pergamon Press: Oxford, UK, 1987; pp. 413–434. [Google Scholar]
- Addison, A.W.; Rao, T.N.; Reedijk, J.; van Rijn, J.; Verschor, G.C. Synthesis, structure, and spectroscopic properties of copper(II) compounds containing nitrogen–sulphur donor ligands; the crystal and molecular structure of aqua[1,7-bis(N-methylbenzimidazol-2′-yl)-2,6-dithiaheptane]copper(II) perchlorate. J. Chem. Soc. Dalton Trans. 1984, 1349–1356. [Google Scholar] [CrossRef]
- Lever, A.B.P. Inorganic Electronic Spectroscopy; Elsevier: Amsterdam, The Netherlands, 1986; pp. 555–572. [Google Scholar]
- Anatole, A.; Bleaney, B. Electron Paramagnetic Resonance of Transition Ion; Oxford University Press: Oxford, UK, 2012; pp. 455–466. [Google Scholar]
- Maxim, C.; Badea, M.; Rostas, A.M.; Chifiriuc, M.C.; Pircalabioru, G.G.; Avram, S.; Olar, R. Copper (II) species with 1-(o-tolyl) biguanide: Structural characterization, ROS scavenging, antibacterial activity, biocompatibility and in silico studies. Appl. Organomet. Chem. 2022, 36, 6471. [Google Scholar] [CrossRef]
- Garriba, E.; Micera, G. The Determination of the Geometry of Cu(II) Complexes An EPR Spectroscopy Experiment. J. Chem Ed. 2006, 83, 1229–1232. [Google Scholar] [CrossRef]
- Machado, P.H.A.; Paixão, D.A.; Campos Lino, R.; de Souza, T.R.; de Souza Bontempo, N.J.; Sousa, L.M.; Van Petten de Vasconcelos Azevedo, F.; Capelari Orsolin, P.; Alves Pereira Lima, P.M.; Castro Martins, I.; et al. A selective CuII complex with 4-fluorophenoxyacetic acid hydrazide and phenanthroline displays DNA-cleaving and pro-apoptotic properties in cancer cells. Sci. Rep. 2021, 11, 24450. [Google Scholar] [CrossRef] [PubMed]
- Levín, P.; Ruiz, M.C.; Romo, A.I.B.; Nascimento, O.R.; Di Virgilio, A.L.; Oliver, A.G.; Ayala, A.P.; Diógenes, I.C.N.; León, I.E.; Lemus, L. Water-mediated reduction of [Cu(dmp)2(CH3CN)]2+: Implications of the structure of a classical complex on its activity as an anticancer drug. Inorg. Chem. Front. 2021, 8, 3238–3252. [Google Scholar] [CrossRef]
- Ruiz, M.C.; Perelmulter, K.; Levín, P.; Romo, A.I.B.; Lemus, L.; Bollati -Fogolín, M.; León, I.E.; Di Virgilio, A.L. Antiproliferative activity of two copper (II) complexes on colorectal cancer cell models: Impact on ROS production, apoptosis induction and NF-κB inhibition. Eur. J. Pharm. Sci. 2022, 169, 106092. [Google Scholar] [CrossRef]
- Almeida, J.C.; Paixão, D.A.; Marzano, I.M.; Ellena, J.; Pivatto, M.; Lopes, N.P.; Ferreira, A.M.D.C.; Pereira-Maia, E.C.; Guilardi, S.; Guerra, W. Copper(II) complexes with β-diketones and N-donor heterocyclic ligands: Crystal structure, spectral properties, and cytotoxic activity. Polyhedron 2015, 89, 1–8. [Google Scholar] [CrossRef]
- Burgos-López, Y.; Balsa, L.M.; Piro, O.E.; León, I.E.; García-Tojal, J.; Echeverría, G.A.; González-Baró, A.C.; Parajón-Costa, B.S. Tridentate acylhydrazone copper(II) complexes with heterocyclic bases as coligands. Synthesis, spectroscopic studies, crystal structure and cytotoxicity assays. Polyhedron 2022, 213, 115621. [Google Scholar] [CrossRef]
- Elsayed, S.A.; Elnabky, I.M.; di Biase, A.; El-Hendawy, A.M. New mixed ligand copper(II) hydrazone-based complexes: Synthesis, characterization, crystal structure, DNA/RNA/BSA binding, in vitro anticancer, apoptotic activity, and cell cycle analysis. Appl. Organomet. Chem. 2022, 36, 6481. [Google Scholar] [CrossRef]
- Barnum, K.J.; O’Connell, M.J. Cell Cycle Regulation by Checkpoints. Methods Mol. Biol. 2014, 1170, 29–40. [Google Scholar]
- Shaltiel, I.A.; Krenning, L.; Bruinsma, W.; Medema, R.H. The same, only different—DNA damage checkpoints and their reversal throughout the cell cycle. J. Cell Sci. 2015, 128, 607–620. [Google Scholar] [CrossRef]
- Hajrezaie, M.; Paydar, M.; Zorofchian Moghadamtousi, S.; Hassandarvish, P.; Gwaram, N.S.; Zahedifard, M.; Rouhollahi, E.; Karimian, H.; Looi, C.Y.; Ali, H.M.; et al. A Schiff Base-Derived Copper (II) Complex Is a Potent Inducer of Apoptosis in Colon Cancer Cells by Activating the Intrinsic Pathway. Sci. World J. 2014, 2014, 540463. [Google Scholar] [CrossRef]
- Gu, Y.-Q.; Zhong, Y.-J.; Hu, M.-Q.; Li, H.-Q.; Yang, K.; Dong, Q.; Liang, H.; Chen, Z.-F. Terpyridine copper(II) complexes as potential anticancer agents by inhibiting cell proliferation, blocking the cell cycle and inducing apoptosis in BEL-7402 cells. Dalton Trans. 2022, 51, 1968–1978. [Google Scholar] [CrossRef]
- Wehbe, M.; Leung, A.W.Y.; Abrams, M.J.; Orvig, C.; Bally, M.B. A Perspective—Can copper complexes be developed as a novel class of therapeutics? Dalton Trans. 2017, 46, 10758–10773. [Google Scholar] [CrossRef]
- Esmaeili, L.; Perez, M.G.; Jafari, M.; Paquin, J.; Ispas-Szabo, P.; Pop, V.; Andruh, M.; Byers, J.; Mateescu, M.A. Copper complexes for biomedical applications: Structural insights, antioxidant activity and neuron compatibility. J. Inorg. Biochem. 2019, 192, 87–97. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.H.; Yin, Y.H.; Chen, H.Z.; Feng, S.Y.; Liu, J.L.; Chen, L.; Fu, W.L.; Sun, G.C.; Yu, X.G.; Xu, D.G. TRIM24 Promotes Stemness and Invasiveness of Glioblastoma Cells via Activating SOX2 Expression. Neuro-Oncol. 2020, 22, 1797–1808. [Google Scholar] [CrossRef] [PubMed]
- Yu, T.; Gan, S.; Zhu, Q.; Dai, D.; Li, N.; Wang, H.; Chen, X.; Hou, D.; Wang, Y.; Pan, Q.; et al. Modulation of M2 macrophage polarization by the crosstalk between Stat6 and Trim24. Nat. Commun. 2019, 10, 4353. [Google Scholar] [CrossRef]
- Klapper, L.; Idel, C.; Kuppler, P.; Jagomast, T.; von Bernuth, A.; Bruchhage, K.L.; Rades, D.; Offermann, A.; Kirfel, J.; Perner, S.; et al. TRIM24 Expression as an Independent Biomarker for Prognosis and Tumor Recurrence in HNSCC. J. Pers. Med. 2022, 12, 991. [Google Scholar] [CrossRef]
- Luo, Y.; Chen, C. The roles and regulation of the KLF5 transcription factor in cancers. Cancer Sci. 2021, 112, 2097–2117. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.K.; Saxena, M.; Maharjan, K.; Song, J.J.; Shroyer, K.R.; Bialkowska, A.B.; Shivdasani, R.A.; Yang, V.W. Krüppel-like Factor 5 Regulates Stemness, Lineage Specification, and Regeneration of Intestinal Epithelial Stem Cells. Cell. Mol. Gastroenterol. Hepatol. 2020, 9, 587–609. [Google Scholar] [CrossRef] [PubMed]

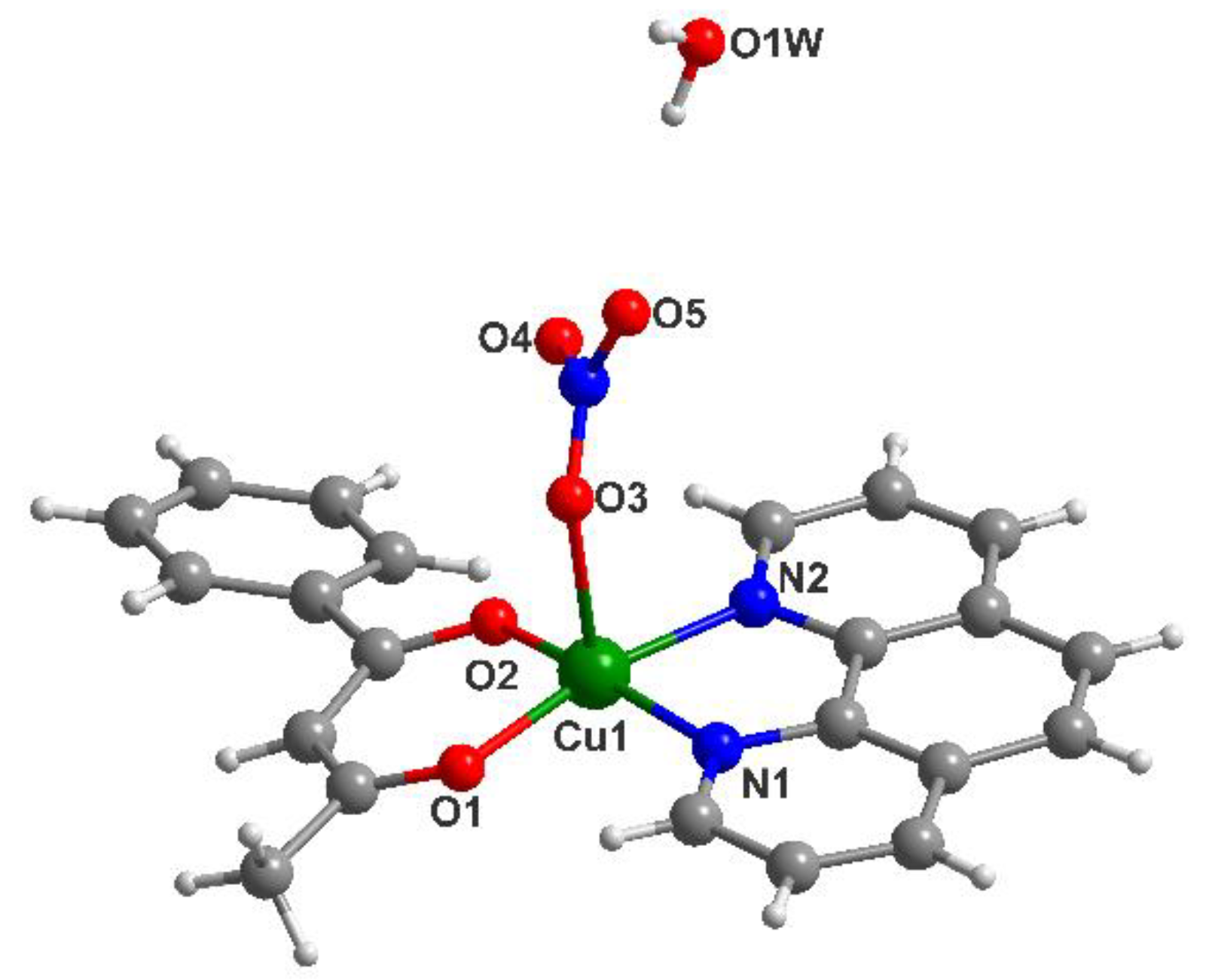
| Empirical Formula | C22H19CuN3O6 |
|---|---|
| Formula weight | 484.94 |
| Temperature/K | 293(2) |
| Crystal system | triclinic |
| Space group | P-1 |
| a/Å | 8.7851(7) |
| b/Å | 10.1855(7) |
| c/Å | 12.6653(9) |
| α/° | 74.179(6) |
| β/° | 70.089(7) |
| γ/° | 75.645(6) |
| Volume/Å3 | 1010.01(14) |
| Z | 2 |
| ρcalcg/cm3 | 1.595 |
| μ/mm−1 | 1.128 |
| F(000) | 498 |
| Radiation | MoK\a |
| Reflections collected | 9422 |
| Data/restraints/parameters | 330/3/296 |
| Goodness-of-fit on F2 | 1.018 |
| Final R indexes [I ≥ 2σ (I)] | 0.0702 |
| Final R indexes [all data] | 0.0775 |
| Largest diff. peak/hole/e Å−3 | −1.112, 0.924 |
| (2) | ||
|---|---|---|
| Cu1 | O1 | 1.901(3) |
| Cu1 | O2 | 1.909(3) |
| Cu1 | N1 | 1.997(4) |
| Cu1 | N2 | 2.000(3) |
| Cu1 | O3 | 2.370(4) |
| N3 | O4 | 1.206(5) |
| N3 | O5 | 1.251(5) |
| N1 | C1 | 1.341(5) |
| N1 | C5 | 1.357(5) |
| N2 | C1 | 1.314(4) |
| N1 | C2 | 1.341(3) |
| Compound | IC50 BJ/μM | IC50 B16/μM | TI | |||
|---|---|---|---|---|---|---|
| 24 h | 48 h | 24 h | 48 h | 24 h | 48 h | |
| (1) | 3.95 | 2.80 | 3.48 | 2.20 | 1.13 | 1.27 |
| (2) | 4.85 | 2.48 | 2.65 | 1.44 | 1.83 | 1.72 |
| (3) | 10.96 | 10.43 | 8.28 | 4.65 | 1.32 | 2.24 |
| (4) | 11.71 | 10.33 | 6.43 | 4.70 | 1.82 | 2.19 |
| Compound | Lipinski | Veber | Ghose | Egan | MW g/mol | Log Po/w (XLOGP3) | Bioavailability Score |
|---|---|---|---|---|---|---|---|
| (1) | Yes; one violation: MW > 500 | Yes | No; one violation: MW > 480 | Yes | 505.39 | 4.77 | 0.55 |
| (2) | Yes | Yes | Yes | Yes | 467.94 | 5.82 | 0.55 |
| (3) | Yes | Yes | No; one violation: MW > 480 | Yes | 482.37 | 3.94 | 0.55 |
| (4) | Yes | Yes | Yes | Yes | 443.92 | 5.52 | 0.55 |
| Compound | (1) | (2) | (3) | (4) |
|---|---|---|---|---|
| HIA | 88.33 | 96.747 | 84.765 | 93.026 |
| Log BBB | −0.988 | −0.714 | −0.94 | −0.882 |
| CNS Permeability | −3.09 | −2.309 | −3.118 | −1.976 |
| OCT2 substrate | yes | yes | no | no |
| AMES toxicity | yes | no | no | no |
| Immunotoxicity | inactive | active | inactive | inactive |
| hepatotoxicity | no | yes | yes | yes |
| Carcinogenicity | inactive | active | inactive | active |
| LD50 | 2.672 | 2.606 | 2.535 | 2.968 |
| Compound | CYP2D6 Substrate/Inhibitor | CYP3A4 Substrate/Inhibitor | CYP1A2 Inhibitor | CYP2C19 Inhibitor | CYP2C9 Inhibitor |
|---|---|---|---|---|---|
| (1) | no/no | yes/no | yes | no | no |
| (2) | no/no | yes/no | yes | yes | yes |
| (3) | no/no | yes/no | no | no | no |
| (4) | no/no | yes/yes | no | no | no |
| Compound (1) | ||||
| Target Name | UniProt ID | PDB Visualization | Probability (%) | Model accuracy (%) |
| Monoamine oxidase A | P21397 | 2Z5Y | 95 | 91 |
| Tyrosyl-DNA phosphodiesterase 1 | Q9NUW8 | 6N0D | 95 | 71 |
| Transcription intermediary factor 1-alpha | O15164 | 4YBM | 92 | 96 |
| Kruppel-like factor 5 | Q13887 | Not Available | 90 | 86 |
| Compound (2) | ||||
| Pregnane X receptor | O75469 | 6TFI | 94 | 95 |
| Transcription intermediary factor 1-alpha | O15164 | 4YBM | 93 | 96 |
| Kruppel-like factor 5 | Q13887 | Not Available | 92 | 86 |
| Monoamine oxidase A | P21397 | 2Z5Y | 91.2 | 91 |
| Compound (3) | ||||
| Tyrosyl-DNA phosphodiesterase 1 | Q9NUW8 | 6N0D | 96 | 71 |
| Endoplasmic reticulum-associated amyloid beta-peptide-binding protein | Q99714 | 2O23 | 95 | 70 |
| Transcription intermediary factor 1-alpha | O15164 | 4YBM | 93 | 96 |
| Nuclear factor NF-kappa-B p105 subunit | P19838 | 1SVC | 93 | 96 |
| Kruppel-like factor 5 | Q13887 | Not Available | 91 | 86 |
| Compound (4) | ||||
| Tyrosyl-DNA phosphodiesterase 1 | Q9NUW8 | 6N0D | 96 | 71 |
| HERG | Q12809 | 5VA1 | 96 | 90 |
| Transcription intermediary factor 1-alpha | O15164 | 4YBM | 96 | 96 |
| Kruppel-like factor 5 | Q13887 | Not Available | 95 | 86 |
| Monoamine oxidase A | P21397 | 2Z5Y | 93 | 91 |
| Cathepsin D | P07339 | 4OD9 | 90 | 99 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Olar, R.; Maxim, C.; Badea, M.; Bacalum, M.; Raileanu, M.; Avram, S.; Korošin, N.Č.; Burlanescu, T.; Rostas, A.M. Antiproliferative Copper(II) Complexes Bearing Mixed Chelating Ligands: Structural Characterization, ROS Scavenging, In Silico Studies, and Anti-Melanoma Activity. Pharmaceutics 2022, 14, 1692. https://doi.org/10.3390/pharmaceutics14081692
Olar R, Maxim C, Badea M, Bacalum M, Raileanu M, Avram S, Korošin NČ, Burlanescu T, Rostas AM. Antiproliferative Copper(II) Complexes Bearing Mixed Chelating Ligands: Structural Characterization, ROS Scavenging, In Silico Studies, and Anti-Melanoma Activity. Pharmaceutics. 2022; 14(8):1692. https://doi.org/10.3390/pharmaceutics14081692
Chicago/Turabian StyleOlar, Rodica, Catalin Maxim, Mihaela Badea, Mihaela Bacalum, Mina Raileanu, Speranta Avram, Nataša Čelan Korošin, Teodora Burlanescu, and Arpad Mihai Rostas. 2022. "Antiproliferative Copper(II) Complexes Bearing Mixed Chelating Ligands: Structural Characterization, ROS Scavenging, In Silico Studies, and Anti-Melanoma Activity" Pharmaceutics 14, no. 8: 1692. https://doi.org/10.3390/pharmaceutics14081692
APA StyleOlar, R., Maxim, C., Badea, M., Bacalum, M., Raileanu, M., Avram, S., Korošin, N. Č., Burlanescu, T., & Rostas, A. M. (2022). Antiproliferative Copper(II) Complexes Bearing Mixed Chelating Ligands: Structural Characterization, ROS Scavenging, In Silico Studies, and Anti-Melanoma Activity. Pharmaceutics, 14(8), 1692. https://doi.org/10.3390/pharmaceutics14081692