Advanced Static and Dynamic Fluorescence Microscopy Techniques to Investigate Drug Delivery Systems
Abstract
1. Introduction
2. Advanced Imaging of Drug Delivery Systems
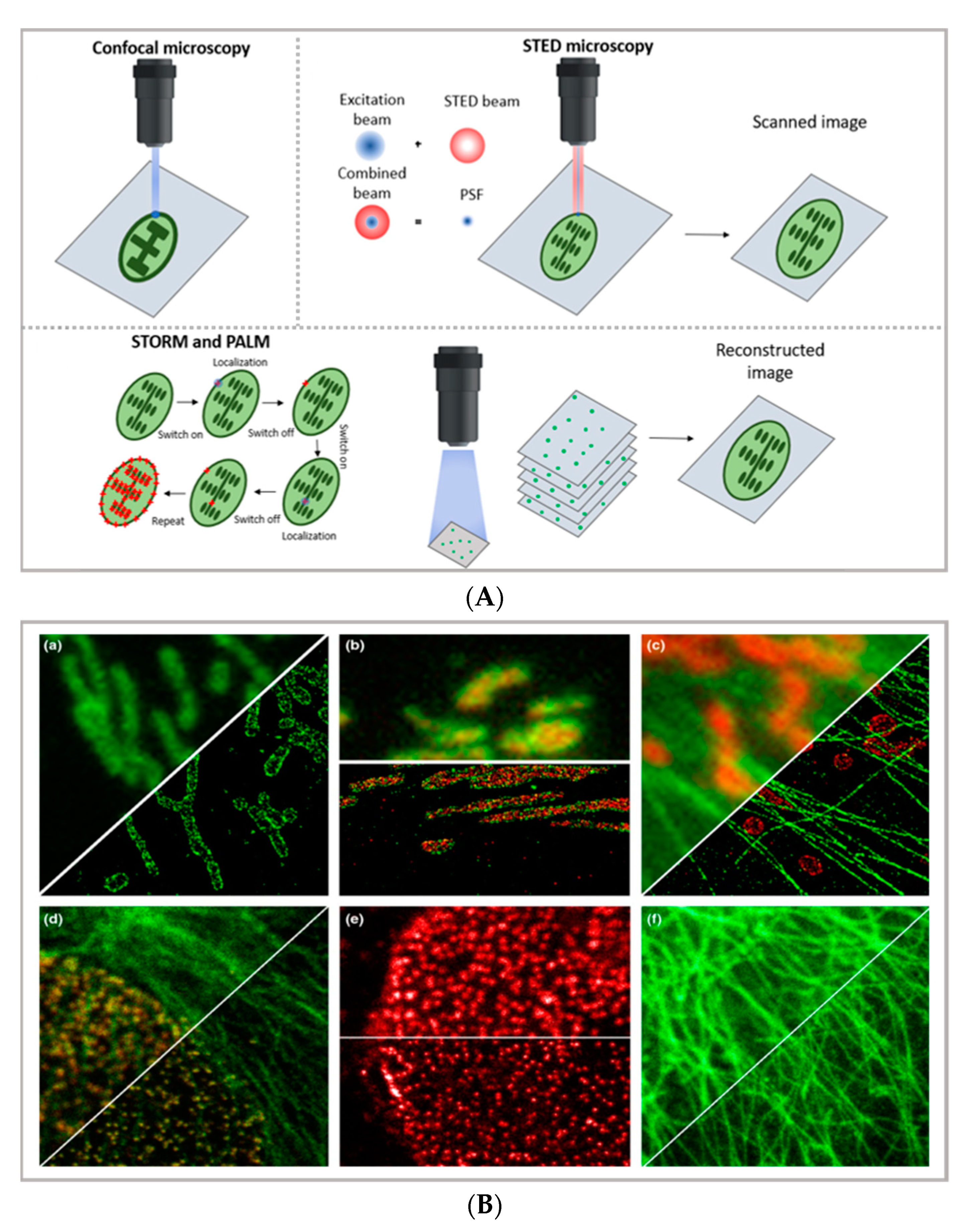
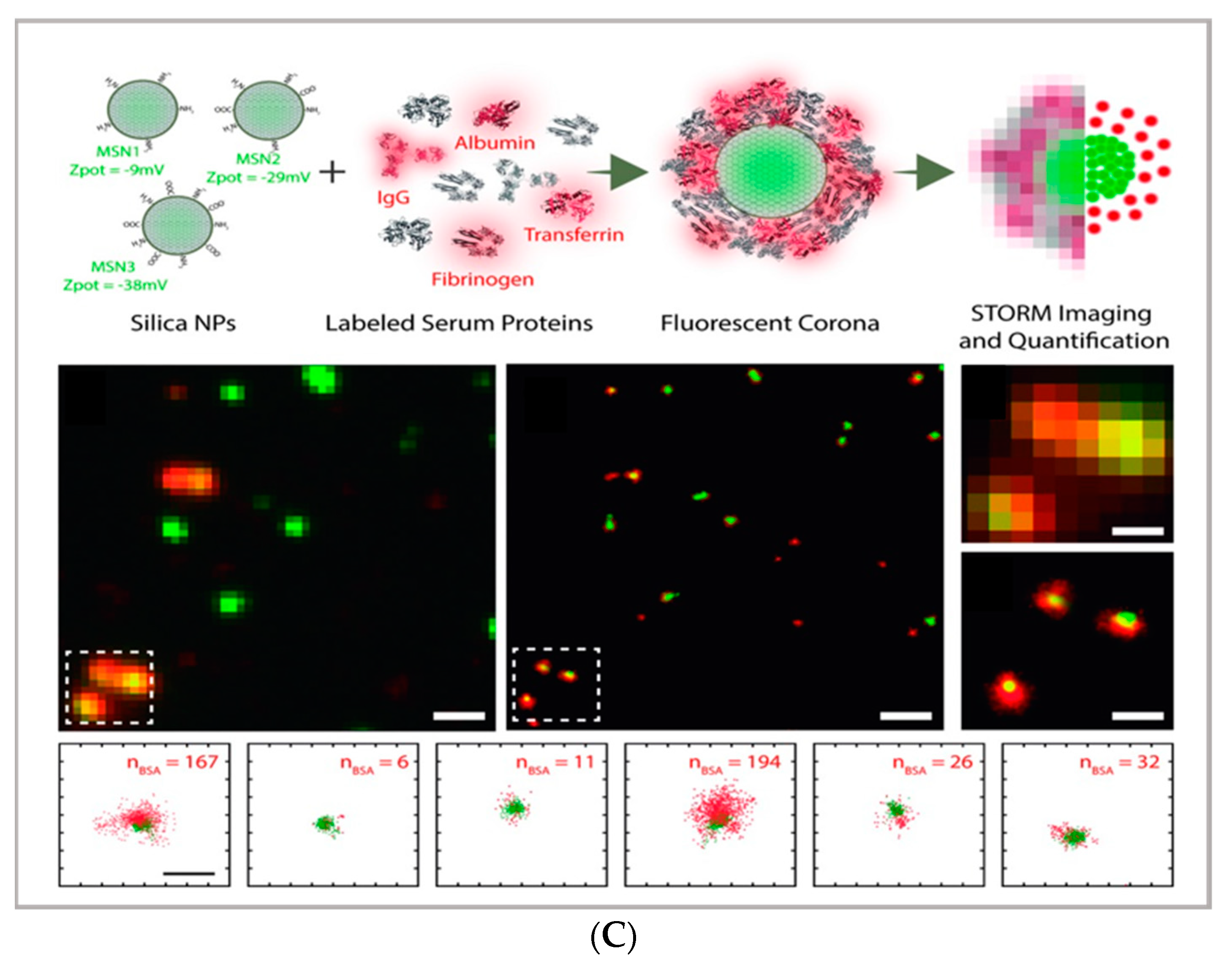
2.1. Super-Resolution Imaging—STED, PALM, STORM
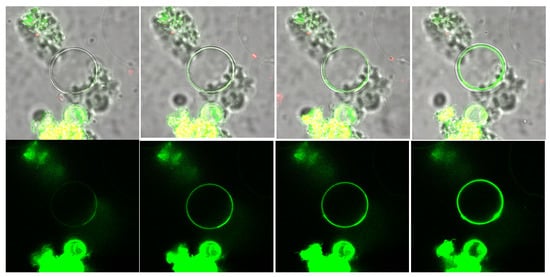
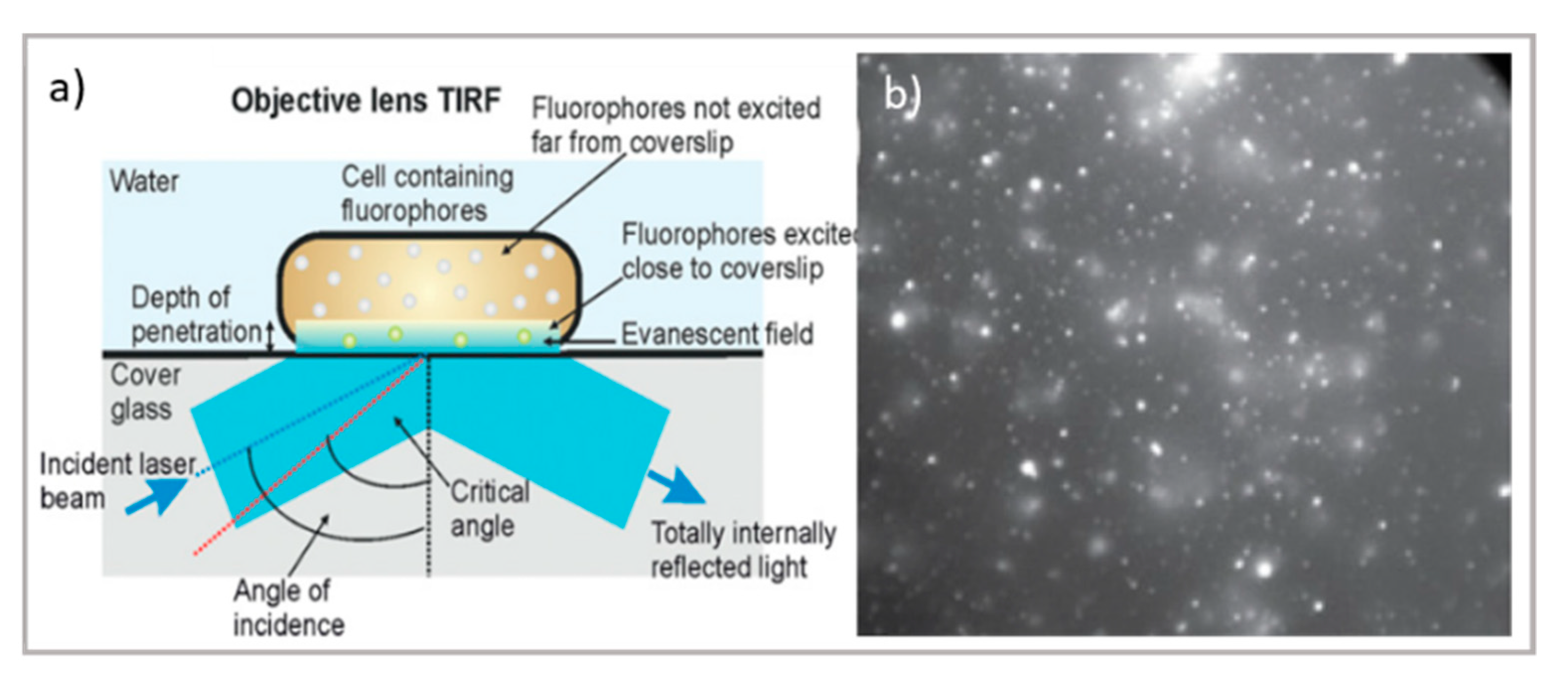
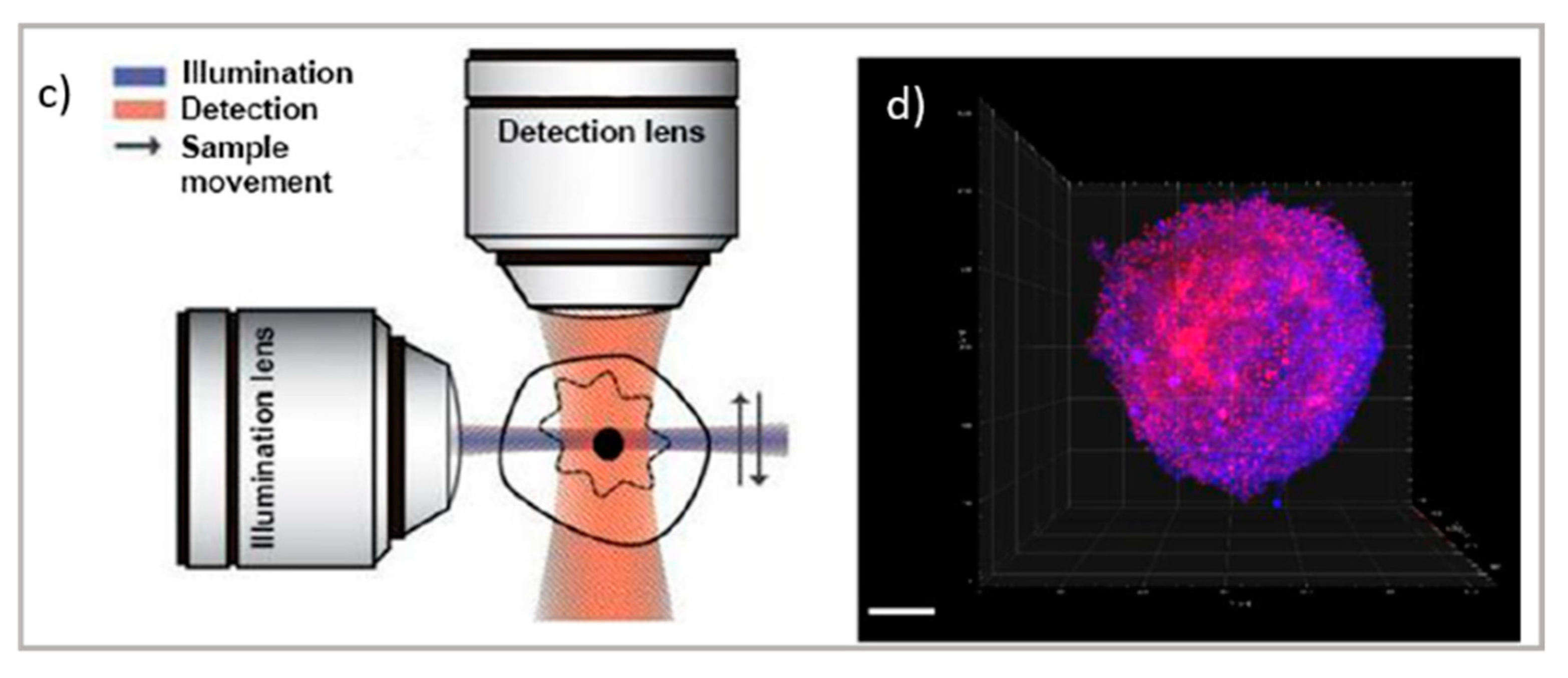
2.2. Imaging Thin Layers and Thick Samples: TIRF, LSFM
3. Dynamics of Drug Delivery System
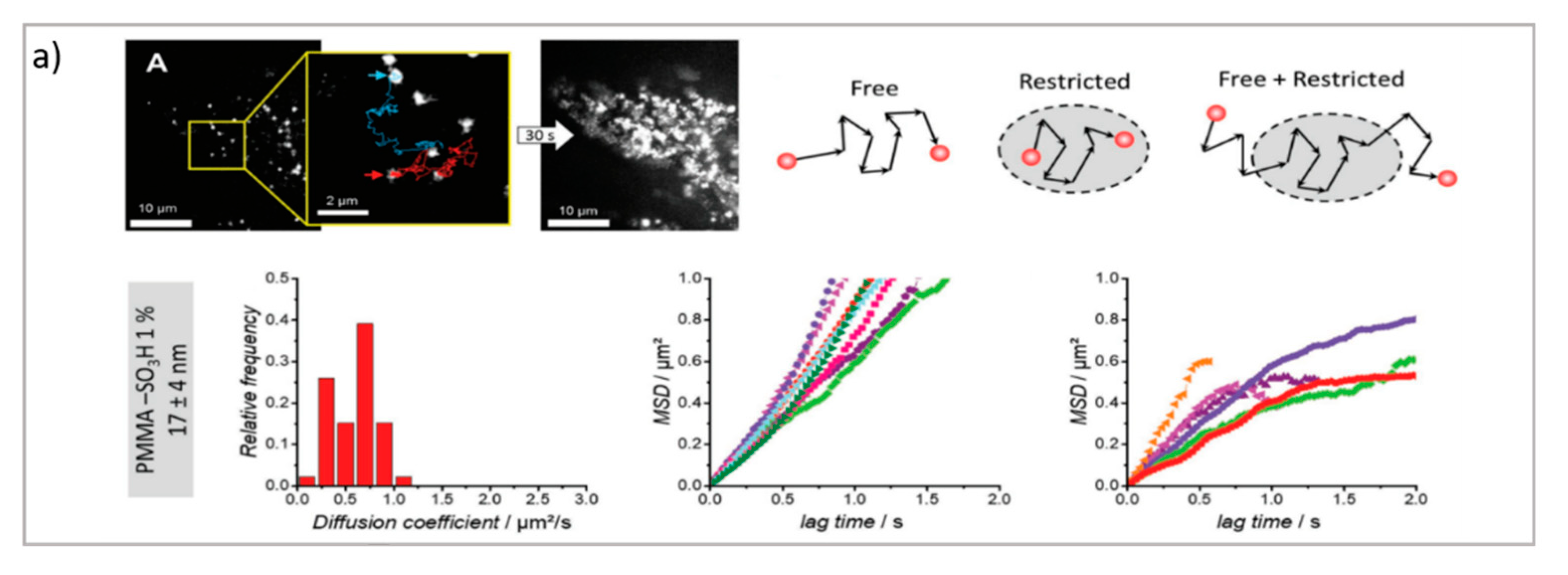
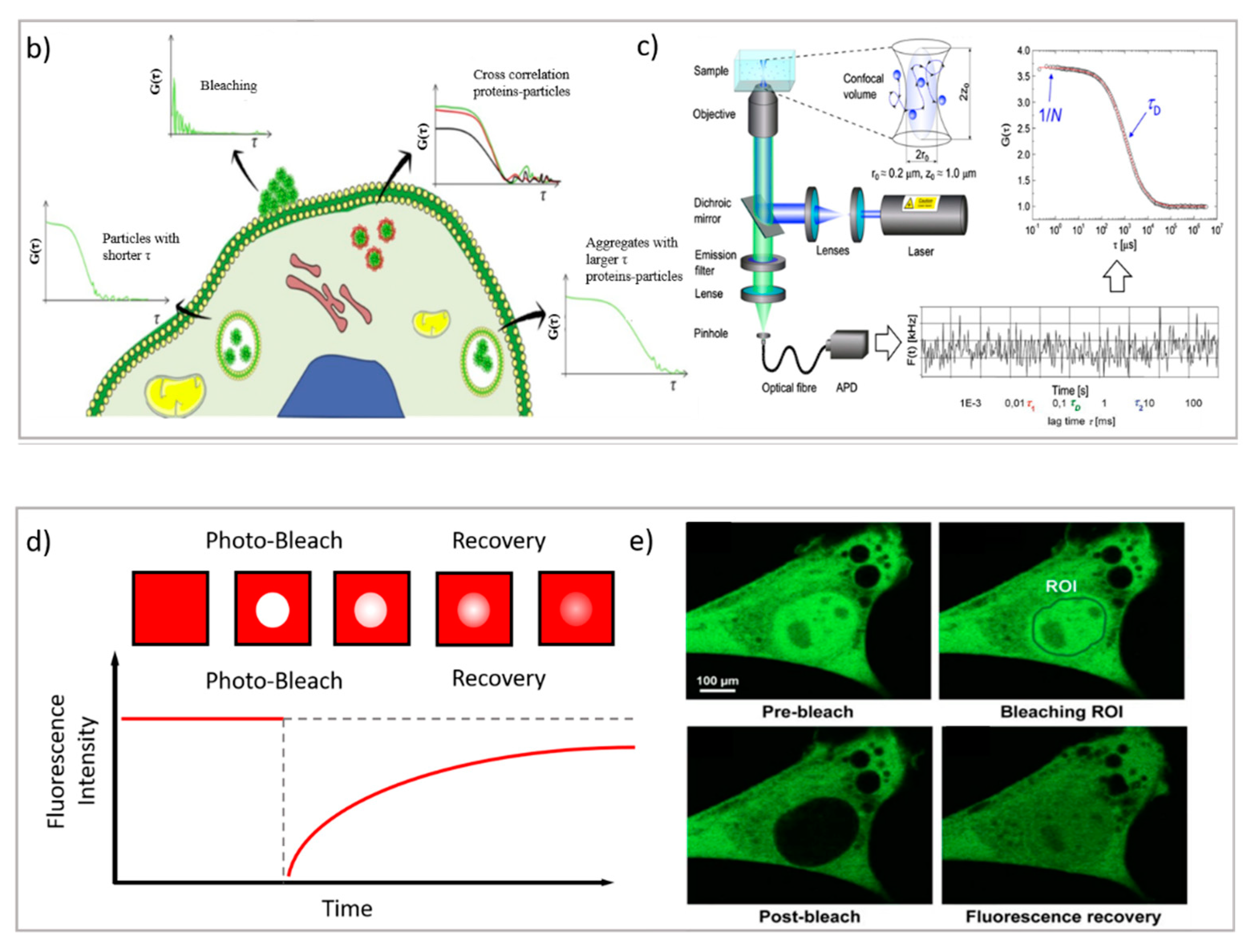
Dynamic Techniques (PT, FCS, FRAP)
4. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Cui, W.; Li, J.; Decher, G. Self-Assembled Smart Nanocarriers for Targeted Drug Delivery. Adv. Mater. 2016, 28, 1302–1311. [Google Scholar] [CrossRef]
- Li, Y.; Tenchov, R.; Smoot, J.; Liu, C.; Watkins, S.; Zhou, Q. A Comprehensive Review of the Global Efforts on COVID-19 Vaccine Development. ACS Cent. Sci. 2021, 2. [Google Scholar] [CrossRef]
- Pujals, S.; Albertazzi, L. Super-resolution Microscopy for Nanomedicine Research. ACS Nano 2019, 13, 9707–9712. [Google Scholar] [CrossRef]
- Murphy, C.J.; Vartanian, A.M.; Geiger, F.M.; Hamers, R.J.; Pedersen, J.; Cui, Q.; Haynes, C.L.; Carlson, E.E.; Hernandez, R.; Klaper, R.D.; et al. Biological responses to engineered nanomaterials: Needs for the next decade. ACS Cent. Sci. 2015, 1, 117–123. [Google Scholar] [CrossRef]
- Chen, K.L.; Bothun, G.D. Nanoparticles meet cell membranes: Probing nonspecific interactions using model membranes. Environ. Sci. Technol. 2014, 48, 873–880. [Google Scholar] [CrossRef] [PubMed]
- Beddoes, C.M.; Case, C.P.; Briscoe, W.H. Understanding nanoparticle cellular entry: A physicochemical perspective. Adv. Colloid Interface Sci. 2015, 218, 48–68. [Google Scholar] [CrossRef] [PubMed]
- Mendozza, M.; Caselli, L.; Salvatore, A.; Montis, C.; Berti, D. Nanoparticles and organized lipid assemblies: From interaction to design of hybrid soft devices. Soft Matter 2019, 15, 8951–8970. [Google Scholar] [CrossRef]
- Giansanti, L.; Condello, M.; Altieri, B.; Galantini, L.; Meschini, S.; Mancini, G. Influence of lipid composition on the ability of liposome loaded voacamine to improve the reversion of doxorubicin resistant osteosarcoma cells. Chem. Phys. Lipids 2019, 223, 104781. [Google Scholar] [CrossRef]
- Xing, Y.; Cheng, Z.; Wang, R.; Lv, C.; James, T.D.; Yu, F. Analysis of extracellular vesicles as emerging theranostic nanoplatforms. Coord. Chem. Rev. 2020, 424, 213506. [Google Scholar] [CrossRef]
- Montis, C.; Generini, V.; Boccalini, G.; Bergese, P.; Bani, D.; Berti, D. Model lipid bilayers mimic non-specific interactions of gold nanoparticles with macrophage plasma membranes. J. Colloid Interface Sci. 2018, 516, 284–294. [Google Scholar] [CrossRef]
- Marín-Menéndez, A.; Montis, C.; Díaz-Calvo, T.; Carta, D.; Hatzixanthis, K.; Morris, C.J.; McArthur, M.; Berti, D. Antimicrobial Nanoplexes meet Model Bacterial Membranes: The key role of Cardiolipin. Sci. Rep. 2017, 7, 41242. [Google Scholar] [CrossRef]
- Tasca, E.; Andreozzi, P.; Del Giudice, A.; Galantini, L.; Schillén, K.; Maria Giuliani, A.; Ramirez, M.d.l.A.; Moya, S.E.; Giustini, M. Poloxamer/sodium cholate co-formulation for micellar encapsulation of doxorubicin with high efficiency for intracellular delivery: An in-vitro bioavailability study. J. Colloid Interface Sci. 2020, 579, 551–561. [Google Scholar] [CrossRef]
- Storti, B.; Civita, S.; Faraci, P.; Maroni, G.; Krishnan, I.; Levantini, E.; Bizzarri, R. Fluorescence imaging of biochemical relationship between ubiquitinated histone 2A and Polycomb complex protein BMI1. Biophys. Chem. 2019, 253, 106225. [Google Scholar] [CrossRef] [PubMed]
- Suh, J.; Wirtz, D.; Hanes, J. Efficient active transport of gene nanocarriers to the cell nucleus. Proc. Natl. Acad. Sci. USA 2003, 100, 3878–3882. [Google Scholar] [CrossRef] [PubMed]
- Maffre, P.; Nienhaus, K.; Amin, F.; Parak, W.J.; Nienhaus, G.U. Characterization of protein adsorption onto fept nanoparticles using dual-focus fluorescence correlation spectroscopy. Beilstein J. Nanotechnol. 2011, 2, 374–383. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Moro, M.; Di Silvio, D.; Moya, S.E. Fluorescence correlation spectroscopy as a tool for the study of the intracellular dynamics and biological fate of protein corona. Biophys. Chem. 2019, 253, 106218. [Google Scholar] [CrossRef]
- Bednarski, M.; Dudek, M.; Knutelska, J.; Nowiński, L.; Sapa, J.; Zygmunt, M.; Nowak, G.; Luty-Błocho, M.; Wojnicki, M.; Fitzner, K.; et al. The influence of the route of administration of gold nanoparticles on their tissue distribution and basic biochemical parameters: In vivo studies. Pharmacol. Rep. 2015, 67, 405–409. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Frei, M.S.; Salim, A.; Johnsson, K. Small-Molecule Fluorescent Probes for Live-Cell Super-Resolution Microscopy. J. Am. Chem. Soc. 2019, 141, 2770–2781. [Google Scholar] [CrossRef] [PubMed]
- Van De Linde, S.; Heilemann, M.; Sauer, M. Live-cell super-resolution imaging with synthetic fluorophores. Annu. Rev. Phys. Chem. 2012, 63, 519–540. [Google Scholar] [CrossRef]
- Lavis, L.D. Teaching old dyes new tricks: Biological probes built from fluoresceins and rhodamines. Annu. Rev. Biochem. 2017, 86, 825–843. [Google Scholar] [CrossRef] [PubMed]
- Bates, M.; Huang, B.; Zhuang, X. Super-resolution microscopy by nanoscale localization of photo-switchable fluorescent probes. Curr. Opin. Chem. Biol. 2008, 12, 505–514. [Google Scholar] [CrossRef] [PubMed]
- Montis, C.; Salvatore, A.; Valle, F.; Paolini, L.; Carlà, F.; Bergese, P.; Berti, D. Biogenic supported lipid bilayers as a tool to investigate nano-bio interfaces. J. Colloid Interface Sci. 2020, 570, 340–349. [Google Scholar] [CrossRef]
- Montis, C.; Busatto, S.; Valle, F.; Zendrini, A.; Salvatore, A.; Gerelli, Y.; Berti, D.; Bergese, P. Biogenic Supported Lipid Bilayers from Nanosized Extracellular Vesicles. Adv. Biosyst. 2018, 2, 1700200. [Google Scholar] [CrossRef]
- Balestri, A.; Chiappisi, L.; Montis, C.; Micciulla, S.; Lonetti, B.; Berti, D. Organized Hybrid Molecular Films from Natural Phospholipids and Synthetic Block Copolymers: A Physicochemical Investigation. Langmuir 2020, 36, 10941–10951. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Li, K.; Hart-Smith, G.; Xu, Y.D.; Jiang, Y.; Lu, H.; Fok, S.; Macmillian, A.; Pandzic, E.; Stenzel, M. Light-sheet microscopy as a tool to understanding the behaviour of Polyion complex micelles for drug delivery. Chem. Commun. 2018, 54, 12618–12621. [Google Scholar] [CrossRef]
- Hell, S.W.; Sahl, S.J.; Bates, M.; Zhuang, X.; Heintzmann, R.; Booth, M.J.; Bewersdorf, J.; Shtengel, G.; Hess, H.; Tinnefeld, P.; et al. The 2015 super-resolution microscopy roadmap. J. Phys. D Appl. Phys. 2015, 48, 443001. [Google Scholar] [CrossRef]
- Hein, B.; Willig, K.I.; Hell, S.W. Stimulated emission depletion (STED) nanoscopy of a fluorescent protein-labeled organelle inside a living cell. Proc. Natl. Acad. Sci. USA 2008. [Google Scholar] [CrossRef]
- Vicidomini, G.; Bianchini, P.; Diaspro, A. STED super-resolved microscopy. Nat. Methods 2018, 15, 173–182. [Google Scholar] [CrossRef]
- Rust, M.J.; Bates, M.; Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 2006, 3, 793–795. [Google Scholar] [CrossRef] [PubMed]
- Henriques, R.; Griffiths, C.; Rego, E.H.; Mhlanga, M.M. PALM and STORM: Unlocking live-cell super-resolution. Biopolymers 2011, 95, 322–331. [Google Scholar] [CrossRef]
- Tam, J.; Merino, D. Stochastic optical reconstruction microscopy (STORM) in comparison with stimulated emission depletion (STED) and other imaging methods. J. Neurochem. 2015, 135, 643–658. [Google Scholar] [CrossRef]
- Egger, B.; Sprecher, S.G. Super-Resolution STED and STORM/PALM Microscopy for Brain Imaging. In Progress in Optical Science and Photonics; Springer: Berlin/Heidelberg, Germany, 2019; Volume 5, pp. 245–258. ISBN 978-981-10-9019-6. [Google Scholar]
- Feiner-Gracia, N.; Beck, M.; Pujals, S.; Tosi, S.; Mandal, T.; Buske, C.; Linden, M.; Albertazzi, L. Super-Resolution Microscopy Unveils Dynamic Heterogeneities in Nanoparticle Protein Corona. Small 2017, 13, 1–11. [Google Scholar] [CrossRef]
- Hell, S.W.; Wichmann, J. Breaking the diffraction resolution limit by stimulated emission: Stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 1994, 19, 780. [Google Scholar] [CrossRef] [PubMed]
- Blom, H.; Widengren, J. Stimulated Emission Depletion Microscopy. Chem. Rev. 2017, 117, 7377–7427. [Google Scholar] [CrossRef] [PubMed]
- Baker, M.B.; Gosens, R.P.J.; Albertazzi, L.; Matsumoto, N.M.; Palmans, A.R.A.; Meijer, E.W. Exposing Differences in Monomer Exchange Rates of Multicomponent Supramolecular Polymers in Water. ChemBioChem 2016, 17, 207–213. [Google Scholar] [CrossRef] [PubMed]
- Karathanasis, C.; Fricke, F.; Hummer, G.; Heilemann, M. Molecule Counts in Localization Microscopy with Organic Fluorophores. ChemPhysChem 2017, 18, 942–948. [Google Scholar] [CrossRef]
- Post, R.A.J.; van der Zwaag, D.; Bet, G.; Wijnands, S.P.W.; Albertazzi, L.; Meijer, E.W.; van der Hofstad, R.W. A stochastic view on surface inhomogeneity of nanoparticles. Nat. Commun. 2019, 10, 1–11. [Google Scholar] [CrossRef]
- Belfiore, L.; Spenkelink, L.M.; Ranson, M.; van Oijen, A.M.; Vine, K.L. Quantification of ligand density and stoichiometry on the surface of liposomes using single-molecule fluorescence imaging. J. Control. Release 2018, 278, 80–86. [Google Scholar] [CrossRef]
- Grimm, F.; Nizamov, S.; Belov, V.N. Green-Emitting Rhodamine Dyes for Vital Labeling of Cell Organelles Using STED Super-Resolution Microscopy. ChemBioChem 2019, 20, 2248–2254. [Google Scholar] [CrossRef]
- Xu, H.; Zhang, H.; Liu, G.; Kong, L.; Zhu, X.; Tian, X.; Zhang, Z.; Zhang, R.; Wu, Z.; Tian, Y.; et al. Coumarin-Based Fluorescent Probes for Super-resolution and Dynamic Tracking of Lipid Droplets. Anal. Chem. 2019, 91, 977–982. [Google Scholar] [CrossRef]
- Han, Y.; Li, M.; Qiu, F.; Zhang, M.; Zhang, Y.H. Cell-permeable organic fluorescent probes for live-cell long-term super-resolution imaging reveal lysosome-mitochondrion interactions. Nat. Commun. 2017, 8. [Google Scholar] [CrossRef]
- Wu, Q.; Jing, Y.; Zhao, T.; Gao, J.; Cai, M.; Xu, H.; Liu, Y.; Liang, F.; Chen, J.; Wang, H. Development of small molecule inhibitor-based fluorescent probes for highly specific super-resolution imaging. Nanoscale 2020, 12, 21591–21598. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Song, X.; Yu, F.; Chen, L. A ratiometric fluorescent probe for imaging and quantifying anti-apoptotic effects of GSH under temperature stress. Chem. Sci. 2017, 8, 6991–7002. [Google Scholar] [CrossRef] [PubMed]
- Song, X.; Han, X.; Yu, F.; Zhang, X.; Chen, L.; Lv, C. Polyamine-targeting Gefitinib prodrug and its near-infrared fluorescent theranostic derivative for monitoring drug delivery and lung cancer therapy. Theranostics 2018, 8, 2217–2228. [Google Scholar] [CrossRef] [PubMed]
- Jin, D.; Xi, P.; Wang, B.; Zhang, L.; Enderlein, J.; Van Oijen, A.M. Nanoparticles for super-resolution microscopy and single-molecule tracking. Nat. Methods 2018, 15, 415–423. [Google Scholar] [CrossRef]
- Li, D.; Qin, W.; Xu, B.; Qian, J.; Tang, B.Z. AIE Nanoparticles with High Stimulated Emission Depletion Efficiency and Photobleaching Resistance for Long-Term Super-Resolution Bioimaging. Adv. Mater. 2017, 29, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Van Der Zwaag, D.; Vanparijs, N.; Wijnands, S.; De Rycke, R.; De Geest, B.G.; Albertazzi, L. Super Resolution Imaging of Nanoparticles Cellular Uptake and Trafficking. ACS Appl. Mater. Interfaces 2016, 8, 6391–6399. [Google Scholar] [CrossRef] [PubMed]
- He, H.; Li, S.; Shi, X.; Wang, X.; Liu, X.; Wang, Q.; Guo, A.; Ge, B.; Khan, N.U.; Huang, F. Quantitative Nanoscopy of Small Blinking Graphene Nanocarriers in Drug Delivery. Bioconjug. Chem. 2018, 29, 3658–3666. [Google Scholar] [CrossRef]
- Chen, C.; Zong, S.; Wang, Z.; Lu, J.; Zhu, D.; Zhang, Y.; Cui, Y. Imaging and Intracellular Tracking of Cancer-Derived Exosomes Using Single-Molecule Localization-Based Super-Resolution Microscope. ACS Appl. Mater. Interfaces 2016, 8, 25825–25833. [Google Scholar] [CrossRef] [PubMed]
- Wojnilowicz, M.; Glab, A.; Bertucci, A.; Caruso, F.; Cavalieri, F. Super-resolution Imaging of Proton Sponge-Triggered Rupture of Endosomes and Cytosolic Release of Small Interfering RNA. ACS Nano 2019, 13, 187–202. [Google Scholar] [CrossRef]
- Riera, R.; Feiner-Gracia, N.; Fornaguera, C.; Cascante, A.; Borrós, S.; Albertazzi, L. Tracking the DNA complexation state of pBAE polyplexes in cells with super resolution microscopy. Nanoscale 2019, 11, 17869–17877. [Google Scholar] [CrossRef]
- Nödling, A.R.; Mills, E.M.; Li, X.; Cardella, D.; Sayers, E.J.; Wu, S.H.; Jones, A.T.; Luk, L.Y.P.; Tsai, Y.H. Cyanine dye mediated mitochondrial targeting enhances the anti-cancer activity of small-molecule cargoes. Chem. Commun. 2020, 56, 4672–4675. [Google Scholar] [CrossRef]
- Jiang, Z.; Liu, H.; He, H.; Yadava, N.; Chambers, J.J.; Thayumanavan, S. Anionic Polymers Promote Mitochondrial Targeting of Delocalized Lipophilic Cations. Bioconjug. Chem. 2020, 31, 1344–1353. [Google Scholar] [CrossRef] [PubMed]
- Iachina, I.; Antonescu, I.E.; Dreier, J.; Sørensen, J.A.; Brewer, J.R. The nanoscopic molecular pathway through human skin. Biochim. Biophys. Acta Gen. Subj. 2019, 1863, 1226–1233. [Google Scholar] [CrossRef] [PubMed]
- May, J.N.; Golombek, S.K.; Baues, M.; Dasgupta, A.; Drude, N.; Rix, A.; Rommel, D.; Von Stillfried, S.; Appold, L.; Pola, R.; et al. Multimodal and multiscale optical imaging of nanomedicine delivery across the blood-brain barrier upon sonopermeation. Theranostics 2020, 10, 1948–1959. [Google Scholar] [CrossRef]
- Fu, Y.; Winter, P.W.; Rojas, R.; Wang, V.; McAuliffe, M.; Patterson, G.H. Axial superresolution via multiangle TIRF microscopy with sequential imaging and photobleaching. Proc. Natl. Acad. Sci. USA 2016, 113, 4368–4373. [Google Scholar] [CrossRef] [PubMed]
- Jaykumar, A.B.; Caceres, P.S.; Sablaban, I.; Tannous, B.A.; Ortiz, P.A. Real-time monitoring of NKCC2 endocytosis by total internal reflection fluorescence (TIRF) microscopy. Am. J. Physiol. Ren. Physiol. 2016, 310, F183–F191. [Google Scholar] [CrossRef] [PubMed]
- Soohoo, A.L.; Bowersox, S.L.; Puthenveedu, M.A. Visualizing clathrin-mediated endocytosis of G protein-coupled receptors at single-event resolution via TIRF microscopy. J. Vis. Exp. 2014, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Dyett, B.P.; Yu, H.; Strachan, J.; Drummond, C.J.; Conn, C.E. Fusion dynamics of cubosome nanocarriers with model cell membranes. Nat. Commun. 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Joyce, P.; Jõemetsa, S.; Isaksson, S.; Hossain, S.; Larsson, P.; Bergström, C.; Höök, F. TIRF Microscopy-Based Monitoring of Drug Permeation Across a Lipid Membrane Supported on Mesoporous Silica. Angew. Chemie 2021, 133, 2097–2101. [Google Scholar] [CrossRef]
- Elisa, Z.; Toon, B.; De Smedt, S.C.; Katrien, R.; Kristiaan, N.; Kevin, B. Technical implementations of light sheet microscopy. Microsc. Res. Tech. 2018, 81, 941–958. [Google Scholar] [CrossRef]
- Lazzari, G.; Vinciguerra, D.; Balasso, A.; Nicolas, V.; Goudin, N.; Garfa-Traore, M.; Fehér, A.; Dinnyés, A.; Nicolas, J.; Couvreur, P.; et al. Light sheet fluorescence microscopy versus confocal microscopy: In quest of a suitable tool to assess drug and nanomedicine penetration into multicellular tumor spheroids. Eur. J. Pharm. Biopharm. 2019, 142, 195–203. [Google Scholar] [CrossRef]
- Friedrich, M.; Gan, Q.; Ermolayev, V.; Harms, G.S. STED-SPIM: Stimulated emission depletion improves sheet illumination microscopy resolution. Biophys. J. 2011, 100, L43–L45. [Google Scholar] [CrossRef]
- Scheul, T.; Wang, I.; Vial, J.-C. STED-SPIM made simple. Opt. Express 2014, 22, 30852. [Google Scholar] [CrossRef]
- Shashkova, S.; Leake, M.C. Single-molecule fluorescence microscopy review: Shedding new light on old problems. Biosci. Rep. 2017, 37, 1–19. [Google Scholar] [CrossRef]
- Reisch, A.; Heimburger, D.; Ernst, P.; Runser, A.; Didier, P.; Dujardin, D.; Klymchenko, A.S. Protein-Sized Dye-Loaded Polymer Nanoparticles for Free Particle Diffusion in Cytosol. Adv. Funct. Mater. 2018, 28, 1–10. [Google Scholar] [CrossRef]
- Koynov, K.; Butt, H.J. Fluorescence correlation spectroscopy in colloid and interface science. Curr. Opin. Colloid Interface Sci. 2012, 17, 377–387. [Google Scholar] [CrossRef]
- Carman, C.V. Overview: Imaging in the Study of Integrins; Springer: Berlin/Heidelberg, Germany, 2011; Volume 757, ISBN 9781617791659. [Google Scholar]
- Ishikawa-Ankerhold, H.C.; Ankerhold, R.; Drummen, G.P.C. Advanced fluorescence microscopy techniques-FRAP, FLIP, FLAP, FRET and FLIM. Molecules 2012, 17, 4047–4132. [Google Scholar] [CrossRef] [PubMed]
- Palazzo, G.; Berti, D. Diffusion and Aggregation; Elsevier B.V.: Amsterdam, The Netherlands, 2014; ISBN 9780444595416. [Google Scholar]
- Ruthardt, N.; Lamb, D.C.; Bräuchle, C. Single-particle tracking as a quantitative microscopy-based approach to unravel cell entry mechanisms of viruses and pharmaceutical nanoparticles. Mol. Ther. 2011, 19, 1199–1211. [Google Scholar] [CrossRef] [PubMed]
- Schuster, B.S.; Ensign, L.M.; Allan, D.B.; Suk, J.S.; Hanes, J. Particle tracking in drug and gene delivery research: State-of-the-art applications and methods. Adv. Drug Deliv. Rev. 2015, 91, 70–91. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Li, X.; Chen, H.; Hu, X.; Luo, Y.; Wang, T.; Wang, Z.; Li, Q.; Fan, C.; Shi, J.; et al. Real-Time Imaging of Endocytosis and Intracellular Trafficking of Semiconducting Polymer Dots. ACS Appl. Mater. Interfaces 2017, 9, 21200–21208. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Han, Y.; Hu, X.; Yin, M.; Wu, C.; Li, Q.; Chen, N.; Zhao, Y. Live-cell imaging of octaarginine-modified polymer dots via single particle tracking. Cell Prolif. 2019, 52. [Google Scholar] [CrossRef]
- Wang, T.; Wang, L.; Li, X.; Hu, X.; Han, Y.; Luo, Y.; Wang, Z.; Li, Q.; Aldalbahi, A.; Wang, L.; et al. Size-Dependent Regulation of Intracellular Trafficking of Polystyrene Nanoparticle-Based Drug-Delivery Systems. ACS Appl. Mater. Interfaces 2017, 9, 18619–18625. [Google Scholar] [CrossRef]
- Zhang, R.L.; Pratiwi, F.W.; Chen, B.C.; Chen, P.; Wu, S.H.; Mou, C.Y. Simultaneous Single-Particle Tracking and Dynamic pH Sensing Reveal Lysosome-Targetable Mesoporous Silica Nanoparticle Pathways. ACS Appl. Mater. Interfaces 2020, 12, 42472–42484. [Google Scholar] [CrossRef]
- Lv, C.; Yang, C.; Ding, D.; Sun, Y.; Wang, R.; Han, D.; Tan, W. Endocytic Pathways and Intracellular Transport of Aptamer-Drug Conjugates in Live Cells Monitored by Single-Particle Tracking. Anal. Chem. 2019, 91, 13818–13823. [Google Scholar] [CrossRef] [PubMed]
- Liang, L.; Li, J.; Li, Q.; Huang, Q.; Shi, J.; Yan, H.; Fan, C. Single-particle tracking and modulation of cell entry pathways of a tetrahedral DNA nanostructure in live cells. Angew. Chemie Int. Ed. 2014. [Google Scholar] [CrossRef]
- Deschout, H.; Raemdonck, K.; Demeester, J.; De Smedt, S.C.; Braeckmans, K. FRAP in pharmaceutical research: Practical guidelines and applications in drug delivery. Pharm. Res. 2014, 31, 255–270. [Google Scholar] [CrossRef]
- Vermonden, T.; Jena, S.S.; Barriet, D.; Censi, R.; Van Der Gucht, J.; Hennink, W.E.; Siegel, R.A. Macromolecular diffusion in self-assembling biodegradable thermosensitive hydrogels. Macromolecules 2010. [Google Scholar] [CrossRef]
- Cardarelli, F.; Tosti, L.; Serresi, M.; Beltram, F.; Bizzarri, R. Fluorescent recovery after photobleaching (FRAP) analysis of nuclear export rates identifies intrinsic features of nucleocytoplasmic transport. J. Biol. Chem. 2012, 287, 5554–5561. [Google Scholar] [CrossRef]
- Starigazdová, J.; Nešporová, K.; Čepa, M.; Šínová, R.; Šmejkalová, D.; Huerta-Angeles, G.; Velebný, V. In vitro investigation of hyaluronan-based polymeric micelles for drug delivery into the skin: The internalization pathway. Eur. J. Pharm. Sci. 2020. [Google Scholar] [CrossRef]
- von Haartman, E.; Lindberg, D.; Prabhakar, N.; Rosenholm, J.M. On the intracellular release mechanism of hydrophobic cargo and its relation to the biodegradation behavior of mesoporous silica nanocarriers. Eur. J. Pharm. Sci. 2016. [Google Scholar] [CrossRef]
- Mudumbi, K.C.; Schirmer, E.C.; Yang, W. Single-point single-molecule FRAP distinguishes inner and outer nuclear membrane protein distribution. Nat. Commun. 2016, 7, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Tønnesen, J.; Katona, G.; Rózsa, B.; Nägerl, U.V. Spine neck plasticity regulates compartmentalization of synapses. Nat. Neurosci. 2014, 17, 678–685. [Google Scholar] [CrossRef]
- Montis, C.; Zendrini, A.; Valle, F.; Busatto, S.; Paolini, L.; Radeghieri, A.; Salvatore, A.; Berti, D.; Bergese, P. Size distribution of extracellular vesicles by optical correlation techniques. Colloids Surfaces B Biointerfaces 2017. [Google Scholar] [CrossRef] [PubMed]
- Ries, J.; Schwille, P. Fluorescence correlation spectroscopy. Bioessays 2012, 34, 361–368. [Google Scholar] [CrossRef]
- Di Silvio, D.; Martínez-Moro, M.; Salvador, C.; de los Angeles Ramirez, M.; Caceres-Velez, P.R.; Ortore, M.G.; Dupin, D.; Andreozzi, P.; Moya, S.E. Self-assembly of poly(allylamine)/siRNA nanoparticles, their intracellular fate and siRNA delivery. J. Colloid Interface Sci. 2019, 557, 757–766. [Google Scholar] [CrossRef] [PubMed]
- Salvatore, A.; Montis, C.; Berti, D.; Baglioni, P. Multifunctional Magnetoliposomes for Sequential Controlled Release. ACS Nano 2016, 10. [Google Scholar] [CrossRef]
- Dabkowska, A.P.; Hirst, C.; Valldeperas, M.; Clifton, L.A.; Montis, C.; Nöjd, S.; Gentile, L.; Wang, M.; Pálsson, G.K.; Lages, S.; et al. Temperature responsive lipid liquid crystal layers with embedded nanogels. Chem. Commun. 2017, 53. [Google Scholar] [CrossRef]
- Montis, C.; Castroflorio, B.; Mendozza, M.; Salvatore, A.; Berti, D.; Baglioni, P. Magnetocubosomes for the delivery and controlled release of therapeutics. J. Colloid Interface Sci. 2015, 449. [Google Scholar] [CrossRef]
- Montis, C.; Baglioni, P.; Berti, D. Monitoring the interaction of nucleolipoplexes with model membranes. Soft Matter 2014, 10, 39–43. [Google Scholar] [CrossRef]
- Vagias, A.; Košovan, P.; Koynov, K.; Holm, C.; Butt, H.-J.; Fytas, G. Dynamics in Stimuli-Responsive Poly(N -isopropylacrylamide) Hydrogel Layers As Revealed by Fluorescence Correlation Spectroscopy. Macromolecules 2014, 47, 5303–5312. [Google Scholar] [CrossRef]
- Sheth, S.; Barnard, E.; Hyatt, B.; Rathinam, M.; Zustiak, S.P. Predicting Drug Release from Degradable Hydrogels Using Fluorescence Correlation Spectroscopy and Mathematical Modeling. Front. Bioeng. Biotechnol. 2019, 7, 1–12. [Google Scholar] [CrossRef]
- Shang, L.; Nienhaus, G.U. In Situ Characterization of Protein Adsorption onto Nanoparticles by Fluorescence Correlation Spectroscopy. Acc. Chem. Res. 2017, 50, 387–395. [Google Scholar] [CrossRef]
- Milani, S.; Baldelli Bombelli, F.; Pitek, A.S.; Dawson, K.A.; Rädler, J. Reversible versus Irreversible Binding of Transferrin to Polystyrene Nanoparticles: Soft and Hard Corona. ACS Nano 2012, 1–7. [Google Scholar] [CrossRef]
- Eriksen, A.Z.; Brewer, J.; Andresen, T.L.; Urquhart, A.J. The diffusion dynamics of PEGylated liposomes in the intact vitreous of the ex vivo porcine eye: A fluorescence correlation spectroscopy and biodistribution study. Int. J. Pharm. 2017, 522, 90–97. [Google Scholar] [CrossRef]
- Mittag, J.J.; Kneidl, B.; Preiβ, T.; Hossann, M.; Winter, G.; Wuttke, S.; Engelke, H.; Rädler, J.O. Impact of plasma protein binding on cargo release by thermosensitive liposomes probed by fluorescence correlation spectroscopy. Eur. J. Pharm. Biopharm. 2017, 119, 215–223. [Google Scholar] [CrossRef]
- Negwer, I.; Best, A.; Schinnerer, M.; Schäfer, O.; Capeloa, L.; Wagner, M.; Schmidt, M.; Mailänder, V.; Helm, M.; Barz, M.; et al. Monitoring drug nanocarriers in human blood by near-infrared fluorescence correlation spectroscopy. Nat. Commun. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Rezgui, R.; Blumer, K.; Yeoh-Tan, G.; Trexler, A.J.; Magzoub, M. Precise quantification of cellular uptake of cell-penetrating peptides using fluorescence-activated cell sorting and fluorescence correlation spectroscopy. Biochim. Biophys. Acta Biomembr. 2016, 1858, 1499–1506. [Google Scholar] [CrossRef] [PubMed]
- Montis, C.; Joseph, P.; Magnani, C.; Marín-Menéndez, A.; Barbero, F.; Estrada, A.R.; Nepravishta, R.; Angulo, J.; Checcucci, A.; Mengoni, A.; et al. Multifunctional nanoassemblies target bacterial lipopolysaccharides for enhanced antimicrobial DNA delivery. Colloids Surfaces B Biointerfaces 2020, 195, 111266. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; De Smedt, S.C.; Remaut, K. Fluorescence Correlation Spectroscopy to find the critical balance between extracellular association and intracellular dissociation of mRNA complexes. Acta Biomater. 2018, 75, 358–370. [Google Scholar] [CrossRef] [PubMed]
- Fu, X.; Sompol, P.; Brandon, J.A.; Norris, C.M.; Wilkop, T.; Johnson, L.A.; Richards, C.I. In Vivo Single-Molecule Detection of Nanoparticles for Multiphoton Fluorescence Correlation Spectroscopy to Quantify Cerebral Blood Flow. Nano Lett. 2020, 20, 6135–6141. [Google Scholar] [CrossRef] [PubMed]
- Petrov, E.P.; Petrosyan, R.; Schwille, P. Translational and rotational diffusion of micrometer-sized solid domains in lipid membranes. Soft Matter 2012, 8, 3552–3555. [Google Scholar] [CrossRef]
- Smeazzetto, S.; Tadini-Buoninsegni, F.; Thiel, G.; Berti, D.; Montis, C. Phospholamban spontaneously reconstitutes into giant unilamellar vesicles where it generates a cation selective channel. Phys. Chem. Chem. Phys. 2016. [Google Scholar] [CrossRef] [PubMed]
- Bianchini, P.; Cardarelli, F.; Di Luca, M.; Diaspro, A.; Bizzarri, R. Nanoscale protein diffusion by STED-based pair correlation analysis. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Lanzanò, L.; Scipioni, L.; Di Bona, M.; Bianchini, P.; Bizzarri, R.; Cardarelli, F.; Diaspro, A.; Vicidomini, G. Measurement of nanoscale three-dimensional diffusion in the interior of living cells by STED-FCS. Nat. Commun. 2017, 8, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Clausen, M.P.; Sezgin, E.; Bernardino de la Serna, J.; Waithe, D.; Lagerholm, B.C.; Eggeling, C. A straightforward approach for gated STED-FCS to investigate lipid membrane dynamics. Methods 2015, 88, 67–75. [Google Scholar] [CrossRef] [PubMed]
- Andrade, D.M.; Clausen, M.P.; Keller, J.; Mueller, V.; Wu, C.; Bear, J.E.; Hell, S.W.; Lagerholm, B.C.; Eggeling, C. Cortical actin networks induce spatio-temporal confinement of phospholipids in the plasma membrane—A minimally invasive investigation by STED-FCS. Sci. Rep. 2015, 5, 1–12. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cardellini, J.; Balestri, A.; Montis, C.; Berti, D. Advanced Static and Dynamic Fluorescence Microscopy Techniques to Investigate Drug Delivery Systems. Pharmaceutics 2021, 13, 861. https://doi.org/10.3390/pharmaceutics13060861
Cardellini J, Balestri A, Montis C, Berti D. Advanced Static and Dynamic Fluorescence Microscopy Techniques to Investigate Drug Delivery Systems. Pharmaceutics. 2021; 13(6):861. https://doi.org/10.3390/pharmaceutics13060861
Chicago/Turabian StyleCardellini, Jacopo, Arianna Balestri, Costanza Montis, and Debora Berti. 2021. "Advanced Static and Dynamic Fluorescence Microscopy Techniques to Investigate Drug Delivery Systems" Pharmaceutics 13, no. 6: 861. https://doi.org/10.3390/pharmaceutics13060861
APA StyleCardellini, J., Balestri, A., Montis, C., & Berti, D. (2021). Advanced Static and Dynamic Fluorescence Microscopy Techniques to Investigate Drug Delivery Systems. Pharmaceutics, 13(6), 861. https://doi.org/10.3390/pharmaceutics13060861