Genetically Encoded Self-Assembling Iron Oxide Nanoparticles as a Possible Platform for Cancer-Cell Tracking
Abstract
1. Introduction
2. Materials and Methods
2.1. Genetic Constructs
2.2. Cell Culture and Protein Expression
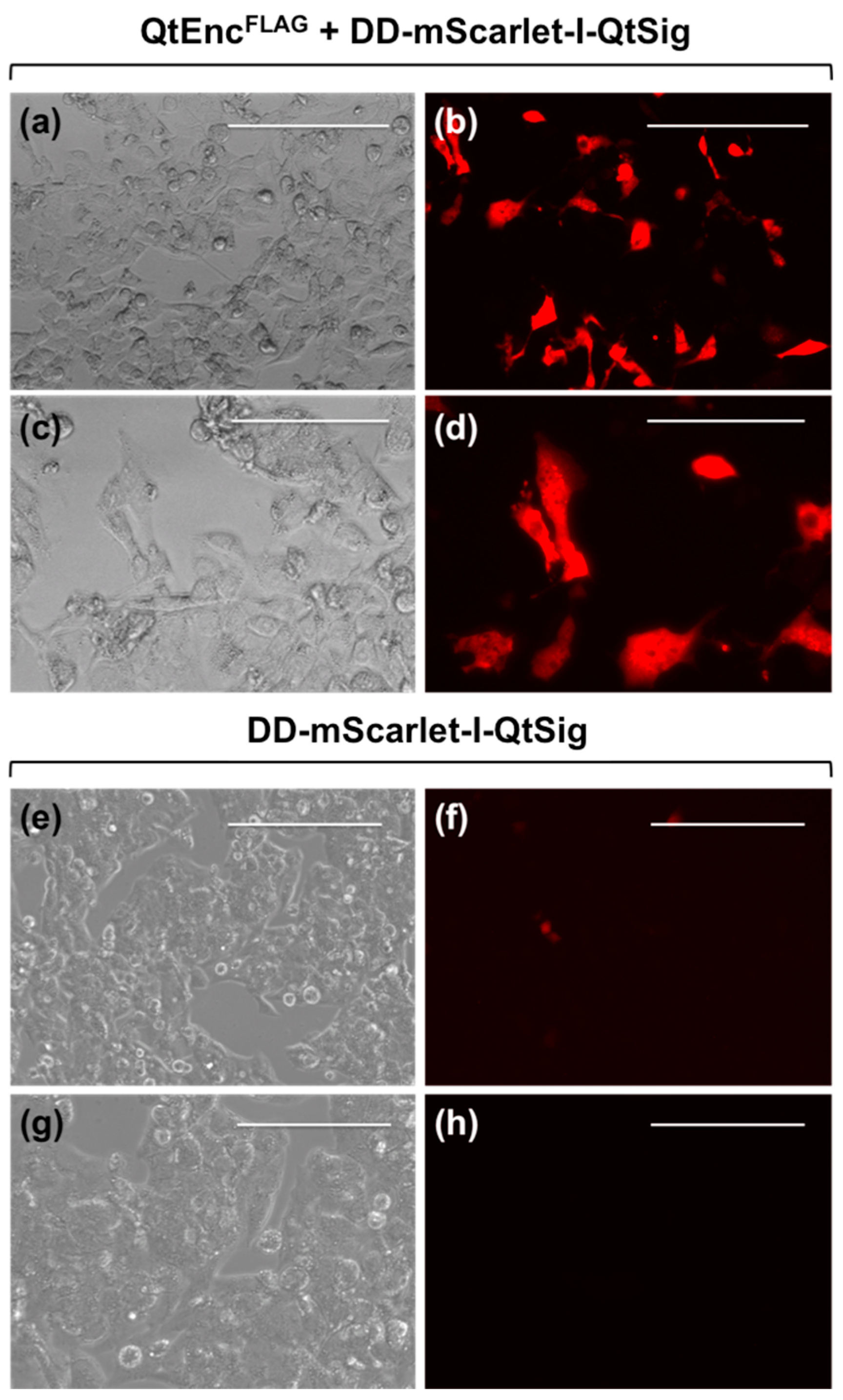
2.3. Optical Microscopy
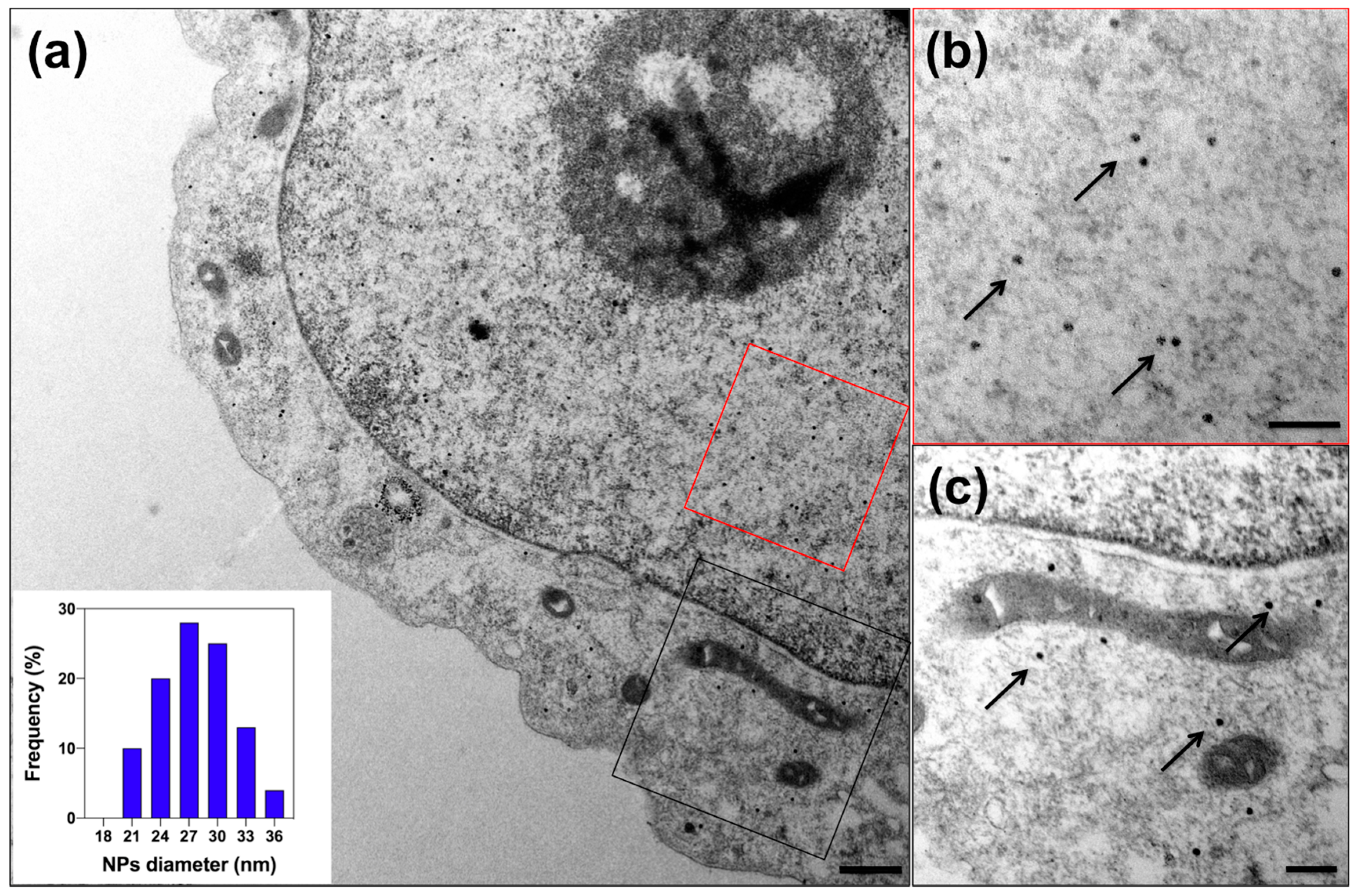
2.4. Transmission Electron Microscopy (TEM)
2.5. Blue Native Polyacrylamide Gel Electrophoresis (PAGE)
2.6. Cell-Viability Assay
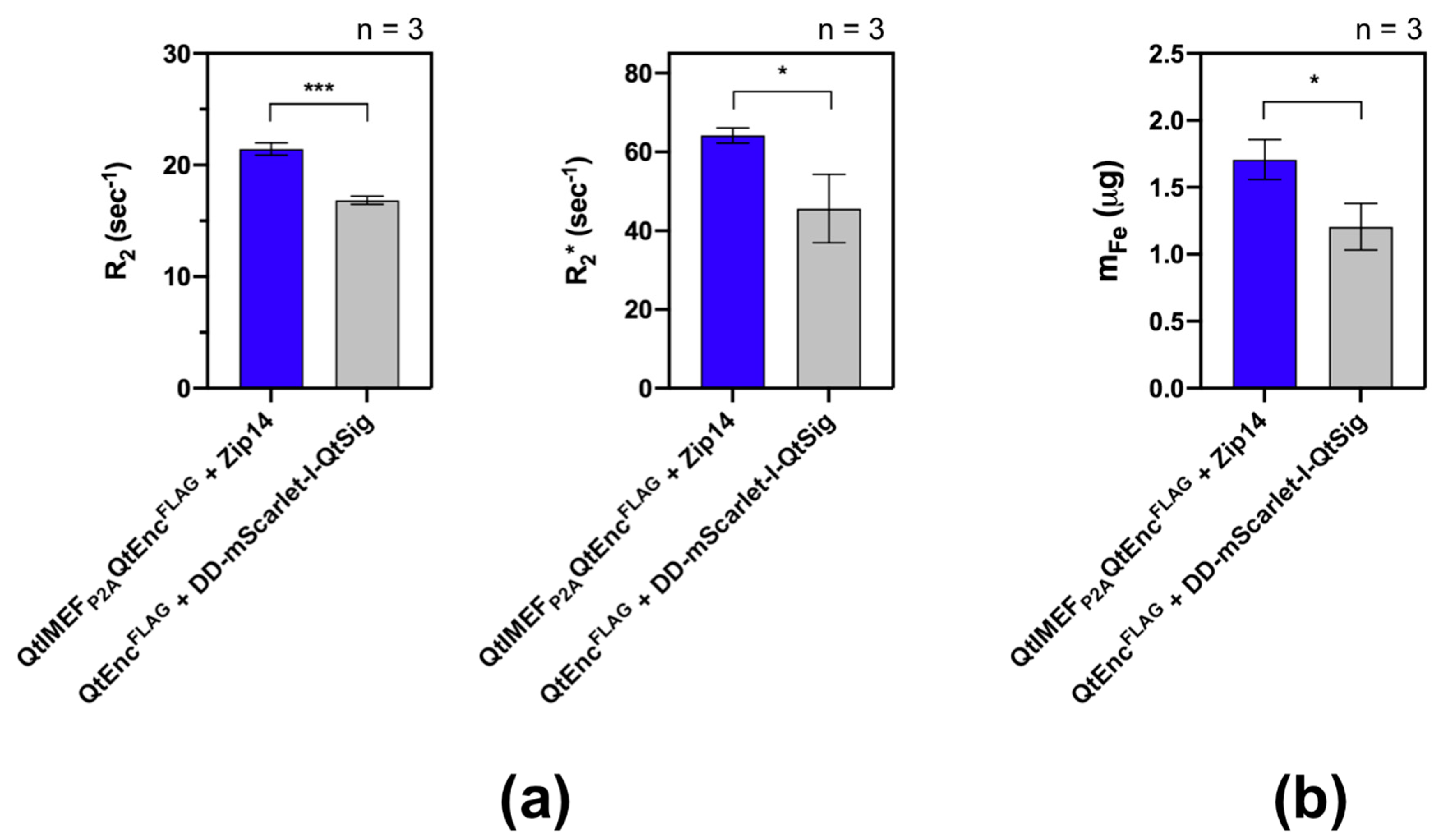
2.7. Magnetic Resonance Imaging of Cells
2.8. Inductively Coupled Plasma Mass Spectrometry (ICP-MS)
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Altorki, N.K.; Markowitz, G.J.; Gao, D.; Port, J.L.; Saxena, A.; Stiles, B.; McGraw, T.; Mittal, V. The lung microenvironment: An important regulator of tumour growth and metastasis. Nat. Rev. Cancer 2019, 19, 9–31. [Google Scholar] [CrossRef]
- Hojman, P.; Gehl, J.; Christensen, J.F.; Pedersen, B.K. Molecular Mechanisms Linking Exercise to Cancer Prevention and Treatment. Cell Metab. 2018, 27, 10–21. [Google Scholar] [CrossRef] [PubMed]
- Cherry, S.R. In vivo molecular and genomic imaging: New challenges for imaging physics. Phys. Med. Biol. 2004, 49, R13–R48. [Google Scholar] [CrossRef] [PubMed]
- Kitamura, T.; Qian, B.-Z.; Pollard, J.W. Immune cell promotion of metastasis. Nat. Rev. Immunol. 2015, 15, 73–86. [Google Scholar] [CrossRef] [PubMed]
- Guzman, R.; Uchida, N.; Bliss, T.M.; He, D.; Christopherson, K.K.; Stellwagen, D.; Capela, A.; Greve, J.; Malenka, R.C.; Moseley, M.E.; et al. Long-term monitoring of transplanted human neural stem cells in developmental and pathological contexts with MRI. Proc. Natl. Acad. Sci. USA 2007, 104, 10211–10216. [Google Scholar] [CrossRef] [PubMed]
- Rice, C.M.; Halfpenny, C.A.; Scolding, N.J. Stem cells for the treatment of neurological disease. Transfus. Med. 2003, 13, 351–361. [Google Scholar] [CrossRef] [PubMed]
- Yang, B.; Chen, B.; He, M.; Hu, B. Quantum Dots Labeling Strategy for “Counting and Visualization” of HepG2 Cells. Anal. Chem. 2017, 89, 1879–1886. [Google Scholar] [CrossRef] [PubMed]
- Kuo, C.W.; Chueh, D.-Y.; Chen, P. Real-time in vivo imaging of subpopulations of circulating tumor cells using antibody conjugated quantum dots. J. Nanobiotechnol. 2019, 17, 26. [Google Scholar] [CrossRef]
- Devaraj, N.K.; Upadhyay, R.; Haun, J.B.; Hilderbrand, S.A.; Weissleder, R. Fast and sensitive pretargeted labeling of cancer cells through a tetrazine/trans-cyclooctene cycloaddition. Angew. Chem. Weinheim Bergstr. Ger. 2009, 121, 7147–7150. [Google Scholar] [CrossRef]
- Chung, J.-K. Sodium iodide symporter: Its role in nuclear medicine. J. Nucl. Med. 2002, 43, 1188–1200. [Google Scholar]
- Kircher, M.F.; Gambhir, S.S.; Grimm, J. Noninvasive cell-tracking methods. Nat. Rev. Clin. Oncol. 2011, 8, 677–688. [Google Scholar] [CrossRef] [PubMed]
- Wahsner, J.; Gale, E.M.; Rodríguez-Rodríguez, A.; Caravan, P. Chemistry of MRI Contrast Agents: Current Challenges and New Frontiers. Chem. Rev. 2019, 119, 957–1057. [Google Scholar] [CrossRef]
- Budde, M.D.; Frank, J.A. Magnetic tagging of therapeutic cells for MRI. J. Nucl. Med. 2009, 50, 171–174. [Google Scholar] [CrossRef] [PubMed]
- Herschman, H.R. Noninvasive imaging of reporter gene expression in living subjects. Adv. Cancer Res. 2004, 92, 29–80. [Google Scholar] [CrossRef] [PubMed]
- Massoud, T.F.; Gambhir, S.S. Molecular imaging in living subjects: Seeing fundamental biological processes in a new light. Genes Dev. 2003, 17, 545–580. [Google Scholar] [CrossRef] [PubMed]
- Aoyama, N.; Miyoshi, H.; Miyachi, H.; Sonoshita, M.; Okabe, M.; Taketo, M.M. Transgenic mice that accept Luciferase- or GFP-expressing syngeneic tumor cells at high efficiencies. Genes Cells 2018, 23, 580–589. [Google Scholar] [CrossRef] [PubMed]
- Taylor, A.; Sharkey, J.; Plagge, A.; Wilm, B.; Murray, P. Multicolour In Vivo Bioluminescence Imaging Using a NanoLuc-Based BRET Reporter in Combination with Firefly Luciferase. Contrast Media Mol. Imaging 2018, 2018, 2514796. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Jing, M.; Li, Y. Lighting up the brain: Genetically encoded fluorescent sensors for imaging neurotransmitters and neuromodulators. Curr. Opin. Neurobiol. 2018, 50, 171–178. [Google Scholar] [CrossRef] [PubMed]
- Ntziachristos, V. Going deeper than microscopy: The optical imaging frontier in biology. Nat. Methods 2010, 7, 603–614. [Google Scholar] [CrossRef]
- Matlashov, M.E.; Shcherbakova, D.M.; Alvelid, J.; Baloban, M.; Pennacchietti, F.; Shemetov, A.A.; Testa, I.; Verkhusha, V.V. A set of monomeric near-infrared fluorescent proteins for multicolor imaging across scales. Nat. Commun. 2020, 11, 239. [Google Scholar] [CrossRef]
- Mishin, A.S.; Belousov, V.V.; Solntsev, K.M.; Lukyanov, K.A. Novel uses of fluorescent proteins. Curr. Opin. Chem. Biol. 2015, 27, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Volpe, A.; Man, F.; Lim, L.; Khoshnevisan, A.; Blower, J.; Blower, P.J.; Fruhwirth, G.O. Radionuclide-fluorescence Reporter Gene Imaging to Track Tumor Progression in Rodent Tumor Models. J. Vis. Exp. 2018. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, A.; Davis, H.C.; Ramesh, P.; Lu, G.J.; Shapiro, M.G. Biomolecular MRI reporters: Evolution of new mechanisms. Prog. Nucl. Magn. Reson. Spectrosc. 2017, 102–103, 32–42. [Google Scholar] [CrossRef] [PubMed]
- Farhadi, A.; Sigmund, F.; Westmeyer, G.G.; Shapiro, M.G. Genetically encodable materials for non-invasive biological imaging. Nat. Mater. 2021. [Google Scholar] [CrossRef]
- Duewell, S.; Kasserra, C.E.; Jezzard, P.; Balaban, R.S. Evaluation of methemoglobin as an autologous intravascular MRI contrast agent. Magn. Reson. Med. 1996, 35, 787–789. [Google Scholar] [CrossRef] [PubMed]
- Naumova, A.V.; Vande Velde, G. Genetically encoded iron-associated proteins as MRI reporters for molecular and cellular imaging. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2018, 10. [Google Scholar] [CrossRef] [PubMed]
- Vande Velde, G.; Rangarajan, J.R.; Toelen, J.; Dresselaers, T.; Ibrahimi, A.; Krylychkina, O.; Vreys, R.; Van der Linden, A.; Maes, F.; Debyser, Z.; et al. Evaluation of the specificity and sensitivity of ferritin as an MRI reporter gene in the mouse brain using lentiviral and adeno-associated viral vectors. Gene Ther. 2011, 18, 594–605. [Google Scholar] [CrossRef]
- Genove, G.; DeMarco, U.; Xu, H.; Goins, W.F.; Ahrens, E.T. A new transgene reporter for in vivo magnetic resonance imaging. Nat. Med. 2005, 11, 450–454. [Google Scholar] [CrossRef]
- Cohen, B.; Dafni, H.; Meir, G.; Harmelin, A.; Neeman, M. Ferritin as an endogenous MRI reporter for noninvasive imaging of gene expression in C6 glioma tumors. Neoplasia 2005, 7, 109–117. [Google Scholar] [CrossRef] [PubMed]
- Gabashvili, A.N.; Chmelyuk, N.S.; Efremova, M.V.; Malinovskaya, J.A.; Semkina, A.S.; Abakumov, M.A. Encapsulins-Bacterial Protein Nanocompartments: Structure, Properties, and Application. Biomolecules 2020, 10, 966. [Google Scholar] [CrossRef] [PubMed]
- Sutter, M.; Boehringer, D.; Gutmann, S.; Günther, S.; Prangishvili, D.; Loessner, M.J.; Stetter, K.O.; Weber-Ban, E.; Ban, N. Structural basis of enzyme encapsulation into a bacterial nanocompartment. Nat. Struct. Mol. Biol. 2008, 15, 939–947. [Google Scholar] [CrossRef]
- Giessen, T.W. Encapsulins: Microbial nanocompartments with applications in biomedicine, nanobiotechnology and materials science. Curr. Opin. Chem. Biol. 2016, 34, 1–10. [Google Scholar] [CrossRef]
- Sigmund, F.; Pettinger, S.; Kube, M.; Schneider, F.; Schifferer, M.; Schneider, S.; Efremova, M.V.; Pujol-Martí, J.; Aichler, M.; Walch, A.; et al. Iron-Sequestering Nanocompartments as Multiplexed Electron Microscopy Gene Reporters. ACS Nano 2019, 13, 8114–8123. [Google Scholar] [CrossRef]
- Sigmund, F.; Massner, C.; Erdmann, P.; Stelzl, A.; Rolbieski, H.; Desai, M.; Bricault, S.; Wörner, T.P.; Snijder, J.; Geerlof, A.; et al. Bacterial encapsulins as orthogonal compartments for mammalian cell engineering. Nat. Commun. 2018, 9, 1990. [Google Scholar] [CrossRef] [PubMed]
- Cheung, J.Y.-N.; Ong, R.C.-Y.; Suen, Y.-K.; Ooi, V.; Wong, H.N.-C.; Mak, T.C.-W.; Fung, K.-P.; Yu, B.; Kong, S.-K. Polyphyllin D is a potent apoptosis inducer in drug-resistant HepG2 cells. Cancer Lett. 2005, 217, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Yip, D.; Cho, C.H. A multicellular 3D heterospheroid model of liver tumor and stromal cells in collagen gel for anti-cancer drug testing. Biochem. Biophys. Res. Commun. 2013, 433, 327–332. [Google Scholar] [CrossRef] [PubMed]
- Maynard-Smith, L.A.; Chen, L.-C.; Banaszynski, L.A.; Ooi, A.G.L.; Wandless, T.J. A directed approach for engineering conditional protein stability using biologically silent small molecules. J. Biol. Chem. 2007, 282, 24866–24872. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Filippi, M.-D. An Alternative Approach for Sample Preparation with Low Cell Number for TEM Analysis. J. Vis. Exp. 2016. [Google Scholar] [CrossRef] [PubMed]
- Bindels, D.S.; Haarbosch, L.; van Weeren, L.; Postma, M.; Wiese, K.E.; Mastop, M.; Aumonier, S.; Gotthard, G.; Royant, A.; Hink, M.A.; et al. mScarlet: A bright monomeric red fluorescent protein for cellular imaging. Nat. Methods 2017, 14, 53–56. [Google Scholar] [CrossRef]
- McCullock, T.W.; MacLean, D.M.; Kammermeier, P.J. Comparing the performance of mScarlet-I, mRuby3, and mCherry as FRET acceptors for mNeonGreen. PLoS ONE 2020, 15, e0219886. [Google Scholar] [CrossRef]
- Duan, Q.; Jia, P.; Zhuang, Z.; Liu, C.; Zhang, X.; Wang, Z.; Sheng, W.; Li, Z.; Zhu, H.; Zhu, B.; et al. Rational Design of a Hepatoma-Specific Fluorescent Probe for HOCl and Its Bioimaging Applications in Living HepG2 Cells. Anal. Chem. 2019, 91, 2163–2168. [Google Scholar] [CrossRef] [PubMed]
- Lai, Z.; Tan, J.; Wan, R.; Tan, J.; Zhang, Z.; Hu, Z.; Li, J.; Yang, W.; Wang, Y.; Jiang, Y.; et al. An “activatable” aptamer-based fluorescence probe for the detection of HepG2 cells. Oncol. Rep. 2017, 37, 2688–2694. [Google Scholar] [CrossRef] [PubMed]
- Kozenkova, E.; Levada, K.; Efremova, M.V.; Omelyanchik, A.; Nalench, Y.A.; Garanina, A.S.; Pshenichnikov, S.; Zhukov, D.G.; Lunov, O.; Lunova, M.; et al. Multifunctional Fe3O4-Au Nanoparticles for the MRI Diagnosis and Potential Treatment of Liver Cancer. Nanomaterials 2020, 10, 1646. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Shang, W.; Zeng, C.; Wang, K.; Liang, X.; Chi, C.; Liang, X.; Yang, J.; Fang, C.; Tian, J. Theranostic imaging of liver cancer using targeted optical/MRI dual-modal probes. Oncotarget 2017, 8, 32741–32751. [Google Scholar] [CrossRef]
- Chen, C.; Wang, S.; Li, L.; Wang, P.; Chen, C.; Sun, Z.; Song, T. Bacterial magnetic nanoparticles for photothermal therapy of cancer under the guidance of MRI. Biomaterials 2016, 104, 352–360. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Efremova, M.V.; Bodea, S.-V.; Sigmund, F.; Semkina, A.; Westmeyer, G.G.; Abakumov, M.A. Genetically Encoded Self-Assembling Iron Oxide Nanoparticles as a Possible Platform for Cancer-Cell Tracking. Pharmaceutics 2021, 13, 397. https://doi.org/10.3390/pharmaceutics13030397
Efremova MV, Bodea S-V, Sigmund F, Semkina A, Westmeyer GG, Abakumov MA. Genetically Encoded Self-Assembling Iron Oxide Nanoparticles as a Possible Platform for Cancer-Cell Tracking. Pharmaceutics. 2021; 13(3):397. https://doi.org/10.3390/pharmaceutics13030397
Chicago/Turabian StyleEfremova, Maria V., Silviu-Vasile Bodea, Felix Sigmund, Alevtina Semkina, Gil G. Westmeyer, and Maxim A. Abakumov. 2021. "Genetically Encoded Self-Assembling Iron Oxide Nanoparticles as a Possible Platform for Cancer-Cell Tracking" Pharmaceutics 13, no. 3: 397. https://doi.org/10.3390/pharmaceutics13030397
APA StyleEfremova, M. V., Bodea, S.-V., Sigmund, F., Semkina, A., Westmeyer, G. G., & Abakumov, M. A. (2021). Genetically Encoded Self-Assembling Iron Oxide Nanoparticles as a Possible Platform for Cancer-Cell Tracking. Pharmaceutics, 13(3), 397. https://doi.org/10.3390/pharmaceutics13030397





