Dual-Targeted Hyaluronic Acid/Albumin Micelle-Like Nanoparticles for the Vectorization of Doxorubicin
Abstract
1. Introduction
2. Materials and Methods
2.1. Synthesis of HAcys
2.2. Synthesis of HAssHSA Conjugate
2.3. Preparation of Labeled HAssHSA Conjugate
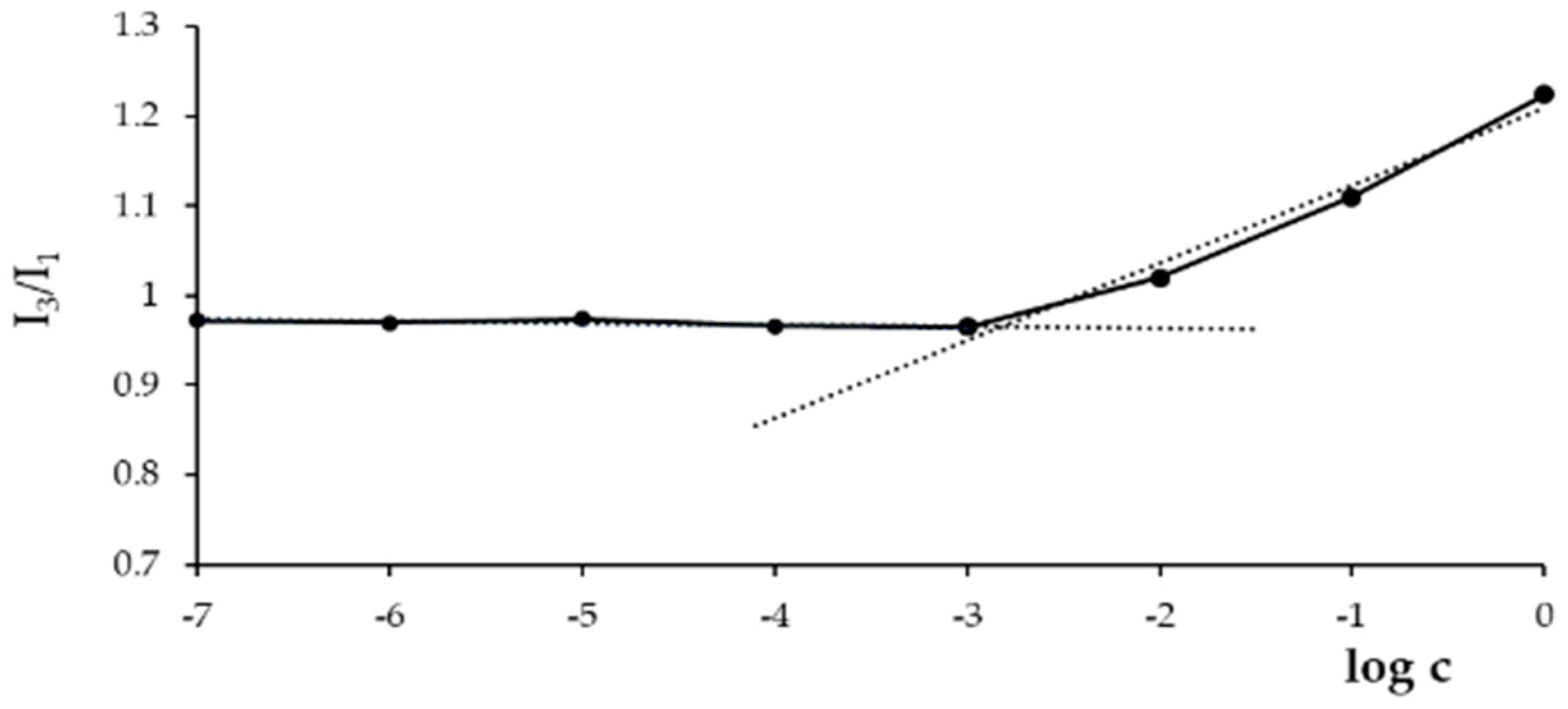
2.4. Determination of the Critical Aggregation Concentration (CAC)
2.5. Preparation and Characterization of Micelle-Like Nanoparticles
2.6. Destabilization Experiments
2.7. DOX Loading
2.8. Release Experiments
2.9. Cytotoxicity Experiments
2.10. Visualization of CD44-Mediated Cellular Uptake
2.11. Statistical Analysis
3. Results and Discussion
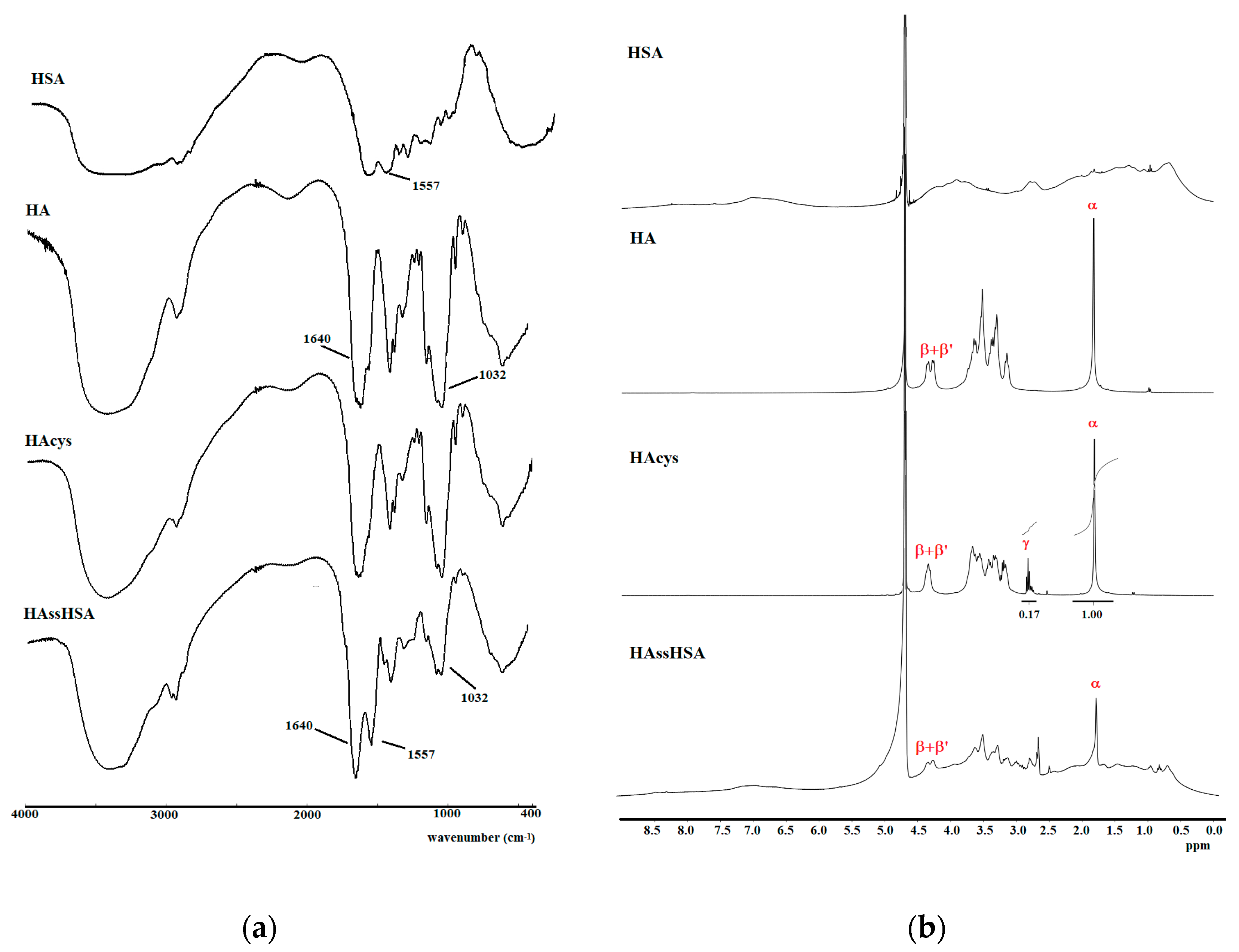
3.1. Synthesis of HAssHSA Conjugate
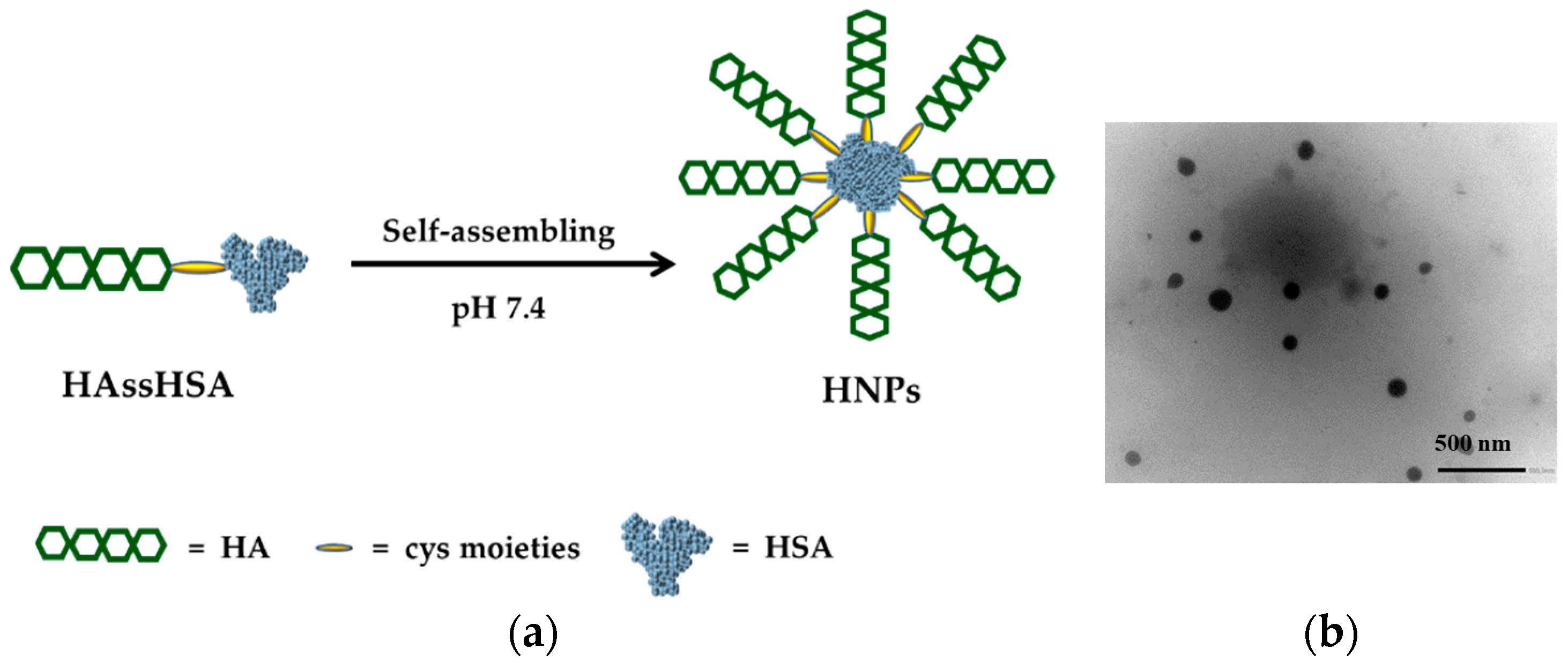
3.2. Preparation of HNPs and DOX Loading
3.3. Destabilization Experiments
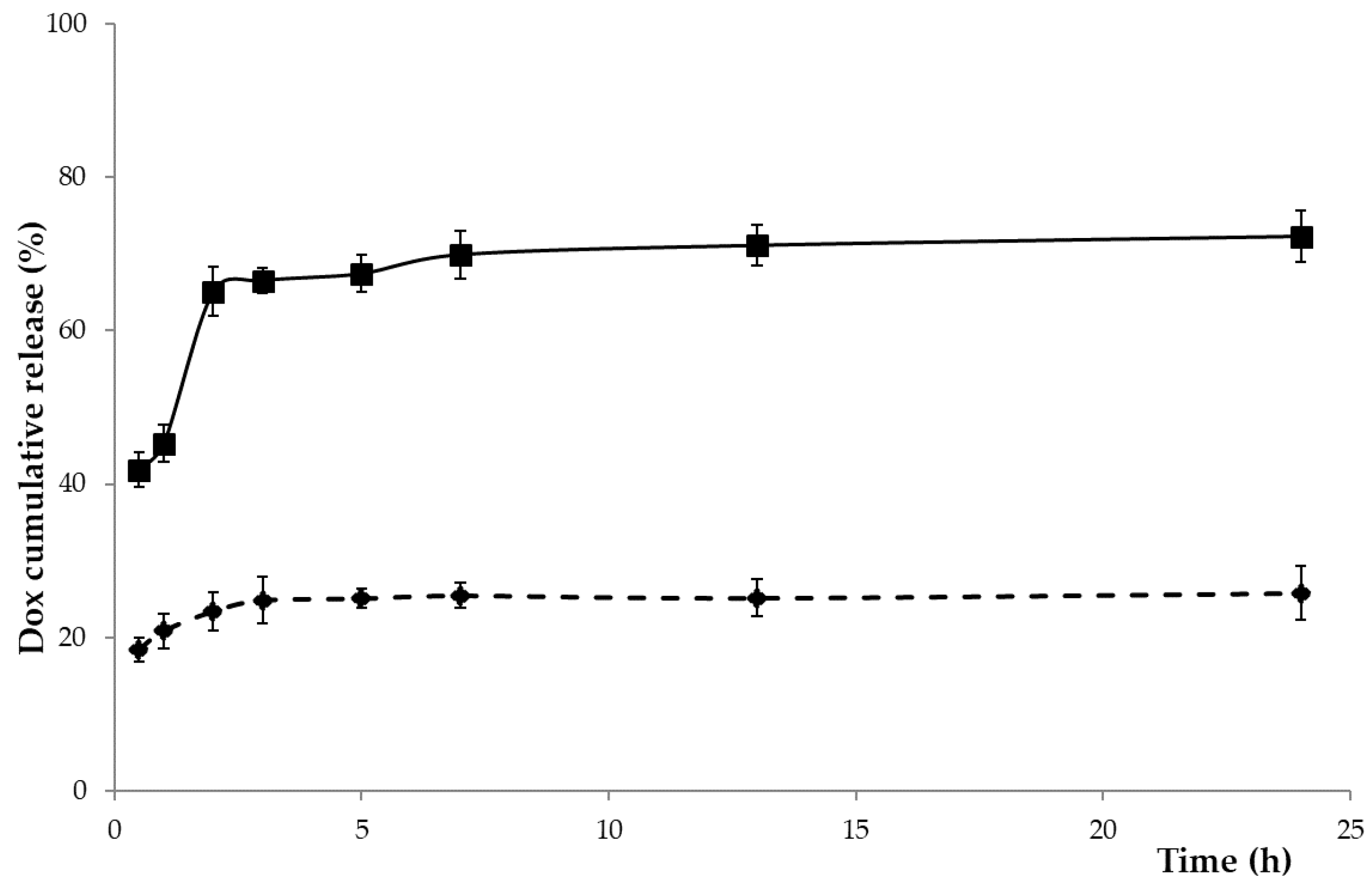
3.4. DOX-Loading and Release Experiments
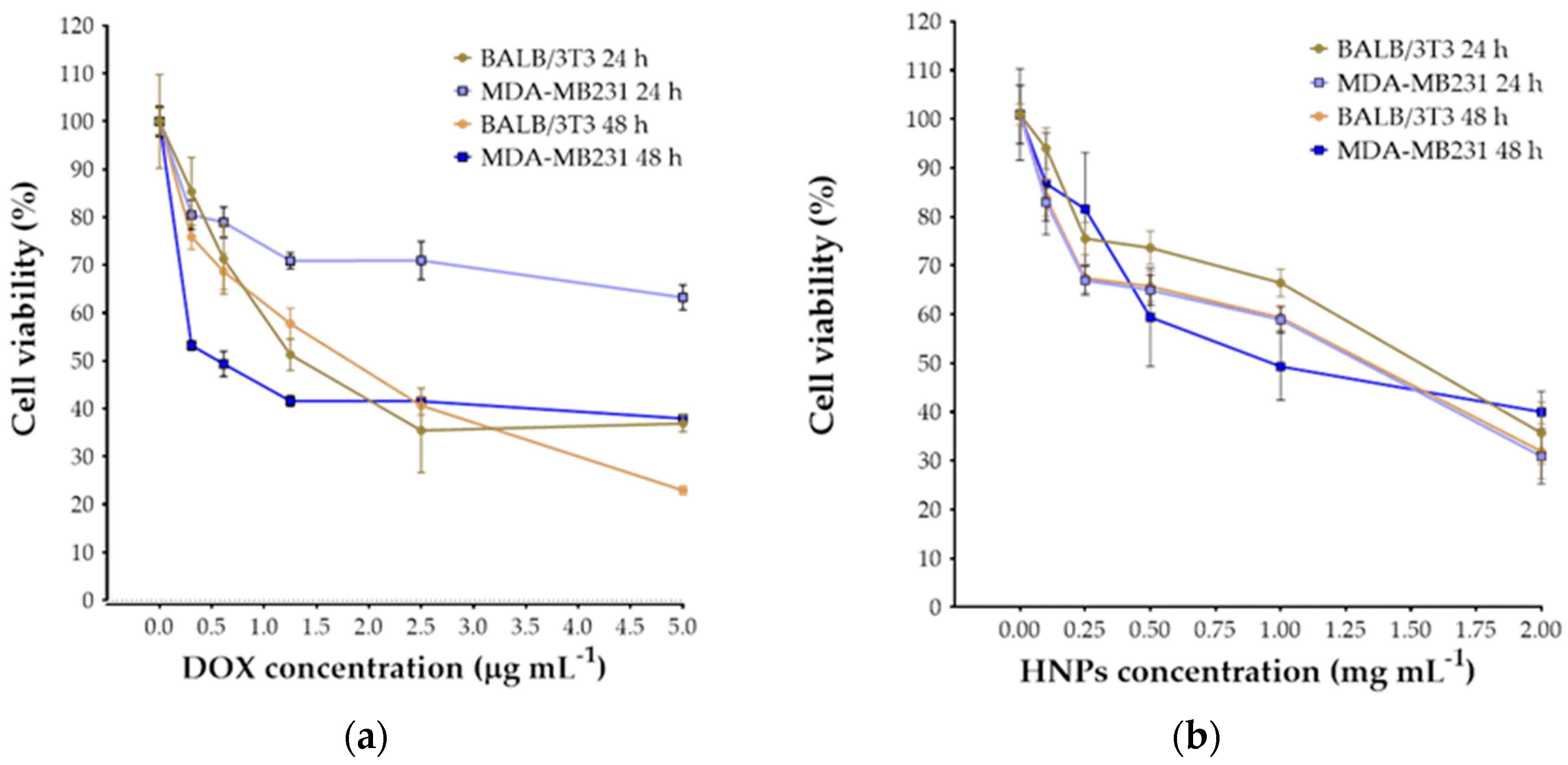
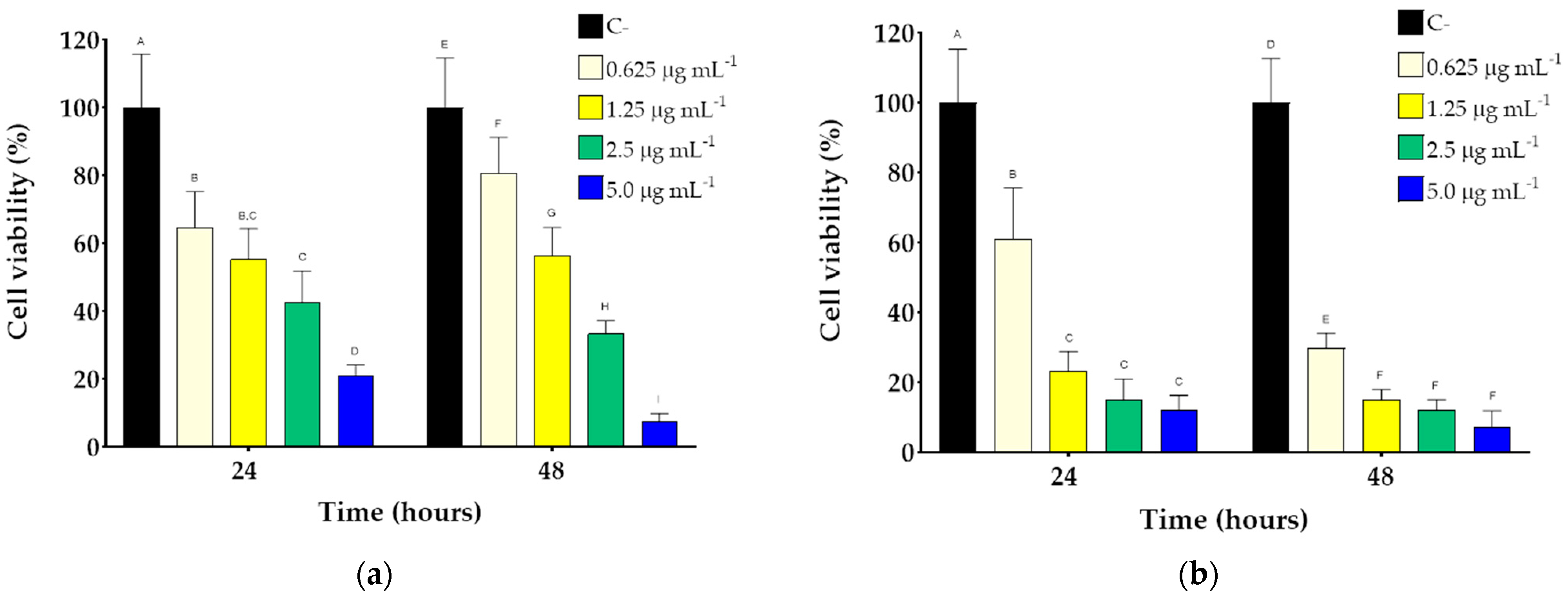
3.5. Cytotoxicity Studies
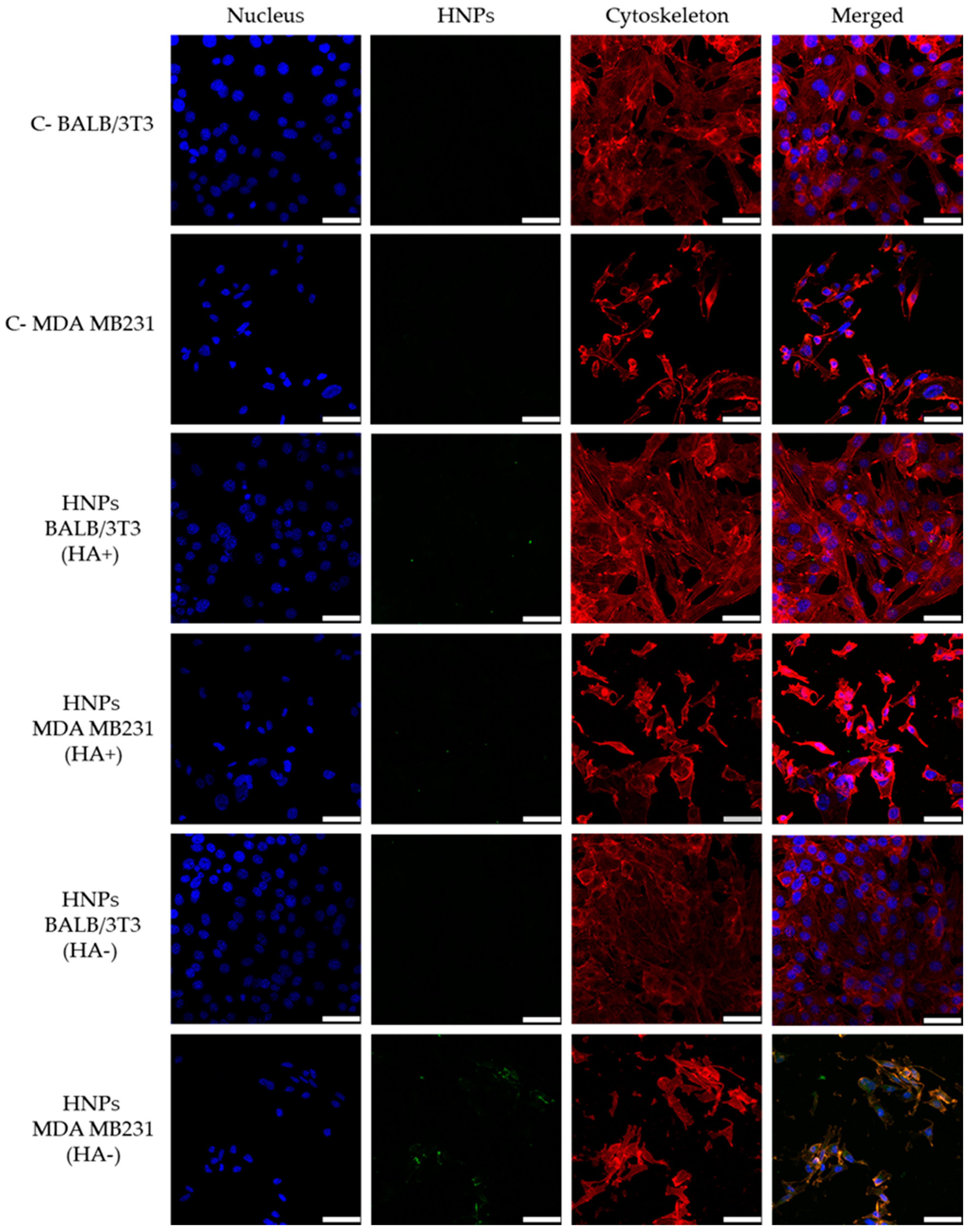
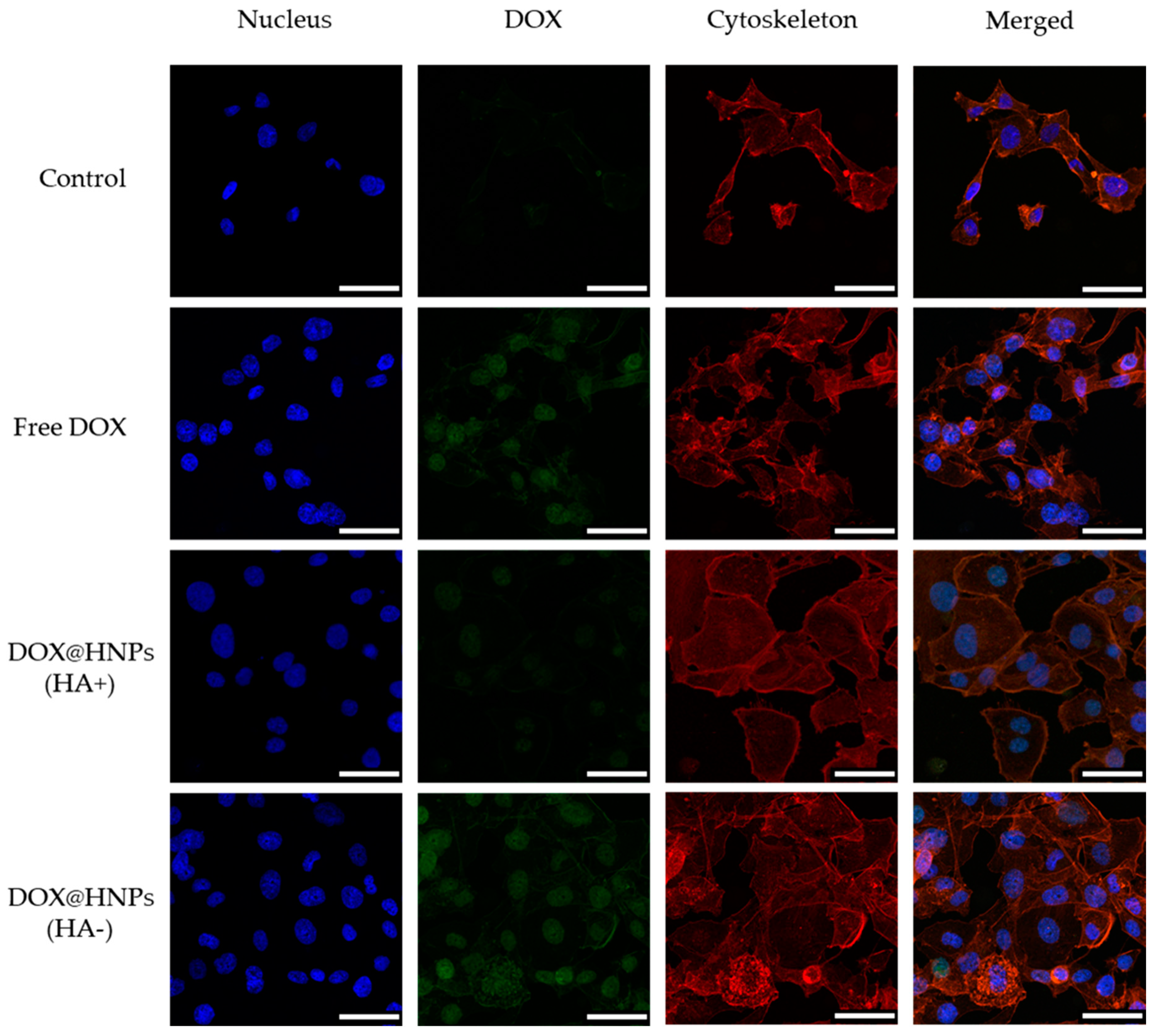
3.6. Cellular Uptake
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Attia, M.F.; Anton, N.; Wallyn, J.; Omran, Z.; Vandamme, T.F. An overview of active and passive targeting strategies to improve the nanocarriers efficiency to tumour sites. J. Pharm. Pharmacol. 2019, 71, 1185–1198. [Google Scholar] [CrossRef]
- Matsumura, Y.; Maeda, H. A New Concept for Macromolecular Therapeutics in Cancer-Chemotherapy—Mechanism of Tumoritropic Accumulation of Proteins and the Antitumor Agent Smancs. Cancer Res. 1986, 46, 6387–6392. [Google Scholar]
- Yoo, J.; Park, C.; Yi, G.; Lee, D.; Koo, H. Active Targeting Strategies Using Biological Ligands for Nanoparticle Drug Delivery Systems. Cancers 2019, 11, 640. [Google Scholar] [CrossRef] [PubMed]
- Curcio, M.; Altimari, I.; Spizzirri, U.G.; Cirillo, G.; Vittorio, O.; Puoci, F.; Picci, N.; Iemma, F. Biodegradable gelatin-based nanospheres as pH-responsive drug delivery systems. J. Nanopart. Res. 2013, 15, 1–11. [Google Scholar] [CrossRef]
- Mi, P. Stimuli-responsive nanocarriers for drug delivery, tumor imaging, therapy and theranostics. Theranostics 2020, 10, 4557–4588. [Google Scholar] [CrossRef]
- Rahman, M.; Ahmad, M.Z.; Kazmi, I.; Akhter, S.; Afzal, M.; Gupta, G.; Sinha, V.R. Emergence of nanomedicine as cancer targeted magic bullets: Recent development and need to address the toxicity apprehension. Curr. Drug Discov. Technol. 2012, 9, 319–329. [Google Scholar] [CrossRef] [PubMed]
- Lv, Y.J.; Cao, Y.; Li, P.; Liu, J.X.; Chen, H.L.; Hu, W.J.; Zhang, L.K. Ultrasound-Triggered Destruction of Folate-Functionalized Mesoporous Silica Nanoparticle-Loaded Microbubble for Targeted Tumor Therapy. Adv. Healthc. Mater. 2017, 6, 1–10. [Google Scholar] [CrossRef]
- Clark, A.J.; Davis, M.E. Increased brain uptake of targeted nanoparticles by adding an acid-cleavable linkage between transferrin and the nanoparticle core. Proc. Natl. Acad. Sci. USA 2015, 112, 12486–12491. [Google Scholar] [CrossRef]
- Curcio, M.; Diaz-Gomez, L.; Cirillo, G.; Concheiro, A.; Iemma, F.; Alvarez-Lorenzo, C. pH/redox dual-sensitive dextran nanogels for enhanced intracellular drug delivery. Eur. J. Pharm. Biopharm. 2017, 117, 324–332. [Google Scholar] [CrossRef]
- Schafer, F.Q.; Buettner, G.R. Redox environment of the cell as viewed through the redox state of the glutathione disulfide/glutathione couple. Free Radic. Biol. Med. 2001, 30, 1191–1212. [Google Scholar] [CrossRef]
- Curcio, M.; Avena, P.; Cirillo, G.; Casaburi, I.; Spizzirri, U.G.; Nicoletta, F.P.; Iemma, F.; Pezzi, V. Functional Albumin Nanoformulations to Fight Adrenocortical Carcinoma: A Redox-Responsive Approach. Pharm. Res. 2020, 37, 55. [Google Scholar] [CrossRef]
- Idrees, H.; Zaidi, S.Z.J.; Sabir, A.; Khan, R.U.; Zhang, X.L.; Hassan, S.U. A Review of Biodegradable Natural Polymer-Based Nanoparticles for Drug Delivery Applications. Nanomaterials 2020, 10, 1970. [Google Scholar] [CrossRef]
- Catanzaro, G.; Curcio, M.; Cirillo, G.; Spizzirri, U.G.; Besharat, Z.M.; Abballe, L.; Vacca, A.; Iemma, F.; Picci, N.; Ferretti, E. Albumin nanoparticles for glutathione-responsive release of cisplatin: New opportunities for medulloblastoma. Int. J. Pharm. 2017, 517, 168–174. [Google Scholar] [CrossRef]
- Lohcharoenkal, W.; Wang, L.Y.; Chen, Y.C.; Rojanasakul, Y. Protein Nanoparticles as Drug Delivery Carriers for Cancer Therapy. Biomed. Res. Int. 2014, 2014, 180549. [Google Scholar] [CrossRef]
- Swierczewska, M.; Han, H.S.; Kim, K.; Park, J.H.; Lee, S. Polysaccharide-based nanoparticles for theranostic nanomedicine. Adv. Drug Deliv. Rev. 2016, 99, 70–84. [Google Scholar] [CrossRef]
- Xu, A.Y.; Melton, L.D.; Williams, M.A.K.; McGillivray, D.J. Protein and polysaccharide conjugates as emerging scaffolds for drug delivery systems. Int. J. Nanotechnol. 2017, 14, 470–480. [Google Scholar] [CrossRef]
- Deng, W.; Li, J.A.; Yao, P.; He, F.; Huang, C. Green Preparation Process, Characterization and Antitumor Effects of Doxorubicin-BSA-Dextran Nanoparticles. Macromol. Biosci. 2010, 10, 1224–1234. [Google Scholar] [CrossRef]
- Chen, Q.; Liang, C.; Wang, C.; Liu, Z. An Imagable and Photothermal “Abraxane-Like” Nanodrug for Combination Cancer Therapy to Treat Subcutaneous and Metastatic Breast Tumors. Adv. Mater. 2015, 27, 903–910. [Google Scholar] [CrossRef]
- An, F.F.; Zhang, X.H. Strategies for Preparing Albumin-based Nanoparticles for Multifunctional Bioimaging and Drug Delivery. Theranostics 2017, 7, 3667–3689. [Google Scholar] [CrossRef]
- Zsila, F. Subdomain IB Is the Third Major Drug Binding Region of Human Serum Albumin: Toward the Three-Sites Model. Mol. Pharmaceut 2013, 10, 1668–1682. [Google Scholar] [CrossRef]
- Agudelo, D.; Bourassa, P.; Bruneau, J.; Berube, G.; Asselin, E.; Tajmir-Riahi, H.A. Probing the Binding Sites of Antibiotic Drugs Doxorubicin and N-(trifluoroacetyl) Doxorubicin with Human and Bovine Serum Albumins. PLoS ONE 2012, 7, e43814. [Google Scholar] [CrossRef]
- Choi, S.H.; Byeon, H.J.; Choi, J.S.; Thao, L.; Kim, I.; Lee, E.S.; Shin, B.S.; Lee, K.C.; Youn, Y.S. Inhalable self-assembled albumin nanoparticles for treating drug-resistant lung cancer. J. Control. Release 2015, 197, 199–207. [Google Scholar] [CrossRef]
- Zhang, B.Y.; Wan, S.Y.; Peng, X.Y.; Zhao, M.Y.; Li, S.; Pu, Y.J.; He, B. Human serum albumin-based doxorubicin prodrug nanoparticles with tumor pH-responsive aggregation-enhanced retention and reduced cardiotoxicity. J. Mater. Chem B 2020, 8, 3939–3948. [Google Scholar] [CrossRef] [PubMed]
- Rao, N.V.; Rho, J.G.; Um, W.; Ek, P.K.; Nguyen, V.Q.; Oh, B.H.; Kim, W.; Park, J.H. Hyaluronic Acid Nanoparticles as Nanomedicine for Treatment of Inflammatory Diseases. Pharmaceutics 2020, 12, 931. [Google Scholar] [CrossRef]
- Kim, J.H.; Moon, M.J.; Kim, D.Y.; Heo, S.H.; Jeong, Y.Y. Hyaluronic Acid-Based Nanomaterials for Cancer Therapy. Polymers 2018, 10, 1133. [Google Scholar] [CrossRef]
- Park, H.K.; Lee, S.J.; Oh, J.S.; Lee, S.G.; Jeong, Y.I.; Lee, H.C. Smart Nanoparticles Based on Hyaluronic Acid for Redox-Responsive and CD44 Receptor-Mediated Targeting of Tumor. Nanoscale Res. Lett. 2015, 10, 981. [Google Scholar] [CrossRef]
- Rivankar, S. An overview of doxorubicin formulations in cancer therapy. J. Cancer Res. Ther. 2014, 10, 853–858. [Google Scholar] [CrossRef]
- Volokhova, A.S.; Edgar, K.J.; Matson, J.B. Polysaccharide-containing block copolymers: Synthesis and applications. Mater. Chem. Front. 2020, 4, 99–112. [Google Scholar] [CrossRef]
- Bosker, W.T.E.; Agoston, K.; Stuart, M.A.C.; Norde, W.; Timmermans, J.W.; Slaghek, T.M. Synthesis and interfacial behavior of polystyrene-polysaccharide diblock copolymers. Macromolecules 2003, 36, 1982–1987. [Google Scholar] [CrossRef]
- Ke, C.Y.; Wu, Y.T.; Tseng, W.L. Fluorescein-5-isothiocyanate-conjugated protein-directed synthesis of gold nanoclusters for fluorescent ratiometric sensing of an enzyme-substrate system. Biosens. Bioelectron. 2015, 69, 46–53. [Google Scholar] [CrossRef]
- Besheer, A.; Hause, G.; Kressler, J.; Mader, K. Hydrophobically modified hydroxyethyl starch: Synthesis, characterization and aqueous self-assembly into nano-sized polymeric micelles and vesicles. Biomacromolecules 2007, 8, 359–367. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.L.; Liu, S.; Chen, X.S.; Mo, G.J.; Xie, Z.G.; Jing, X.B. Biodegradable amphiphilic block copolymers bearing protected hydroxyl groups: Synthesis and characterization. Biomacromolecules 2008, 9, 553–560. [Google Scholar] [CrossRef] [PubMed]
- Provencher, S.W. A constrained regularization method for inverting data represented by linear algebraic or integral equations. Comput. Phys. Commun. 1982, 27, 213–227. [Google Scholar] [CrossRef]
- Liu, C.X.; Jiang, T.T.; Yuan, Z.X.; Lu, Y. Self-Assembled Casein Nanoparticles Loading Triptolide for the Enhancement of Oral Bioavailability. Nat. Prod. Commun. 2020, 15, 1–9. [Google Scholar] [CrossRef]
- Tavano, L.; Muzzalupo, R.; Mauro, L.; Pellegrino, M.; Ando, S.; Picci, N. Transferrin-Conjugated Pluronic Niosomes as a New Drug Delivery System for Anticancer Therapy. Langmuir 2013, 29, 12638–12646. [Google Scholar] [CrossRef]
- Maeda, H.; Ueda, M.; Morinaga, T.; Matsumoto, T. Conjugation of Poly(Styrene-Co-Maleic Acid) Derivatives to the Antitumor Protein Neocarzinostatin—Pronounced Improvements in Pharmacological Properties. J. Med. Chem. 1985, 28, 455–461. [Google Scholar] [CrossRef]
- Dong, X.M.; Liu, C.G. Preparation and Characterization of Self-Assembled Nanoparticles of Hyaluronic Acid-Deoxycholic Acid Conjugates. J. Nanomater. 2010, 2010, 906936. [Google Scholar] [CrossRef]
- Liu, Y.H.; Sun, J.; Cao, W.; Yang, J.H.; Lian, H.; Li, X.; Sun, Y.H.; Wang, Y.J.; Wang, S.L.; He, Z.G. Dual targeting folate-conjugated hyaluronic acid polymeric micelles for paclitaxel delivery. Int. J. Pharm. 2011, 421, 160–169. [Google Scholar] [CrossRef]
- He, X.M.; Carter, D.C. Atomic-Structure and Chemistry of Human Serum Albumin. Nature 1992, 358, 209–215. [Google Scholar] [CrossRef]
- Galantini, L.; Leggio, C.; Konarev, P.V.; Pavel, N.V. Human serum albumin binding ibuprofen: A 3D description of the unfolding pathway in urea. Biophys. Chem. 2010, 147, 111–122. [Google Scholar] [CrossRef]
- Ding, D.W.; Tang, X.L.; Cao, X.L.; Wu, J.H.; Yuan, A.H.; Qiao, Q.; Pan, J.; Hu, Y.Q. Novel Self-assembly Endows Human Serum Albumin Nanoparticles with an Enhanced Antitumor Efficacy. AAPS PharmSciTech 2014, 15, 213–222. [Google Scholar] [CrossRef] [PubMed]
- Edelman, R.; Assaraf, Y.G.; Levitzky, I.; Shahar, T.; Livney, Y.D. Hyaluronic acid-serum albumin conjugate-based nanoparticles for targeted cancer therapy. Oncotarget 2017, 8, 24337–24353. [Google Scholar] [CrossRef]
- Lian, H.B.; Wu, J.H.; Hu, Y.Q.; Guo, H.Q. Self-assembled albumin nanoparticles for combination therapy in prostate cancer. Int. J. Nanomed. 2017, 12, 7777–7787. [Google Scholar] [CrossRef]
- Lee, J.E.; Kim, M.G.; Jang, Y.L.; Lee, M.S.; Kim, N.W.; Yin, Y.; Lee, J.H.; Lim, S.Y.; Park, J.W.; Kim, J.; et al. Self-assembled PEGylated albumin nanoparticles (SPAN) as a platform for cancer chemotherapy and imaging. Drug Deliv. 2018, 25, 1570–1578. [Google Scholar] [CrossRef]
- Jeong, H.; Huh, M.; Lee, S.J.; Koo, H.; Kwon, I.C.; Jeong, S.Y.; Kim, K. Photosensitizer-Conjugated Human Serum Albumin Nanoparticles for Effective Photodynamic Therapy. Theranostics 2011, 1, 230–239. [Google Scholar] [CrossRef]
- Dreis, S.; Rothweller, F.; Michaelis, A.; Cinatl, J.; Kreuter, J.; Langer, K. Preparation, characterisation and maintenance of drug efficacy of doxorubicin-loaded human serum albumin (HSA) nanoparticles. Int. J. Pharm. 2007, 341, 207–214. [Google Scholar] [CrossRef]
- Liu, W.; Dai, J.; Xue, W. Design and self-assembly of albumin nanoclusters as a dynamic-covalent targeting co-delivery and stimuli-responsive controlled release platform. J. Mater. Chem. B 2018, 6, 6817–6830. [Google Scholar] [CrossRef]
- Bae, S.; Ma, K.; Kim, T.H.; Lee, E.S.; Oh, K.T.; Park, E.S.; Lee, K.C.; Youn, Y.S. Doxorubicin-loaded human serum albumin nanoparticles surface-modified with TNF-related apoptosis-inducing ligand and transferrin for targeting multiple tumor types. Biomaterials 2012, 33, 1536–1546. [Google Scholar] [CrossRef]
- Wang, H.B.; Li, Y.; Bai, H.S.; Shen, J.; Chen, X.; Ping, Y.; Tang, G.P. A Cooperative Dimensional Strategy for Enhanced Nucleus-Targeted Delivery of Anticancer Drugs. Adv. Funct. Mater. 2017, 27, 1700339. [Google Scholar] [CrossRef]
- Lin, C.; Zhong, Z.Y.; Lok, M.C.; Jiang, X.L.; Hennink, W.E.; Feijen, J.; Engbersen, J.F.J. Novel bioreducible poly(amido amine)s for highly efficient gene delivery. Bioconjugate Chem. 2007, 18, 138–145. [Google Scholar] [CrossRef]
- Zhang, J.; Ren, X.L.; Tian, X.H.; Zhang, P.; Chen, Z.H.; Hu, X.; Mei, X.F. GSH and enzyme responsive nanospheres based on self-assembly of green tea polyphenols and BSA used for target cancer chemotherapy. Colloids Surf. B Biointerfaces 2019, 173, 654–661. [Google Scholar] [CrossRef]
- Xu, L.; Wang, S.B.; Xu, C.; Han, D.; Ren, X.H.; Zhang, X.Z.; Cheng, S.X. Multifunctional Albumin-Based Delivery System Generated by Programmed Assembly for Tumor-Targeted Multimodal Therapy and Imaging. ACS Appl. Mater. Inter. 2019, 11, 38385–38394. [Google Scholar] [CrossRef]
- Gun’ko, V.M.; Turov, V.V.; Krupska, T.V.; Tsapko, M.D. Interactions of human serum albumin with doxorubicin in different media. Chem. Phys. 2017, 483, 26–34. [Google Scholar] [CrossRef]
- Chen, B.; Wu, C.; Zhuo, R.X.; Cheng, S.X. A self-assembled albumin based multiple drug delivery nanosystem to overcome multidrug resistance. RSC Adv. 2015, 5, 6807–6814. [Google Scholar] [CrossRef]
- Wang, Y.C.; Wang, F.; Sun, T.M.; Wang, J. Redox-Responsive Nanoparticles from the Single Disulfide Bond-Bridged Block Copolymer as Drug Carriers for Overcoming Multidrug Resistance in Cancer Cells. Bioconjugate Chem. 2011, 22, 1939–1945. [Google Scholar] [CrossRef]
- Curcio, M.; Cirillo, G.; Paoli, A.; Naimo, G.D.; Mauro, L.; Amantea, D.; Leggio, A.; Nicoletta, F.P.; Lemma, F. Self-assembling Dextran prodrug for redox- and pH-responsive co-delivery of therapeutics in cancer cells. Colloids Surf. B Biointerfaces 2020, 185, 110537. [Google Scholar] [CrossRef]
- Kibria, G.; Hatakeyama, H.; Akiyama, K.; Hida, K.; Harashima, H. Comparative Study of the Sensitivities of Cancer Cells to Doxorubicin, and Relationships between the Effect of the Drug-Efflux Pump P-gp. Biol. Pharm. Bull. 2014, 37, 1926–1935. [Google Scholar] [CrossRef] [PubMed]
- Hermawan, A.; Wagner, E.; Roidl, A. Consecutive salinomycin treatment reduces doxorubicin resistance of breast tumor cells by diminishing drug efflux pump expression and activity. Oncol. Rep. 2016, 35, 1732–1740. [Google Scholar] [CrossRef]
- Almalik, A.; Karimi, S.; Ouasti, S.; Donno, R.; Wandrey, C.; Day, P.J.; Tirelli, N. Hyaluronic acid (HA) presentation as a tool to modulate and control the receptor-mediated uptake of HA-coated nanoparticles. Biomaterials 2013, 34, 5369–5380. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.J.; Lee, W.C.; Shieh, M.J. Hyaluronic acid conjugated micelles possessing CD44 targeting potential for gene delivery. Carbohyd. Polym. 2017, 155, 101–108. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Curcio, M.; Diaz-Gomez, L.; Cirillo, G.; Nicoletta, F.P.; Leggio, A.; Iemma, F. Dual-Targeted Hyaluronic Acid/Albumin Micelle-Like Nanoparticles for the Vectorization of Doxorubicin. Pharmaceutics 2021, 13, 304. https://doi.org/10.3390/pharmaceutics13030304
Curcio M, Diaz-Gomez L, Cirillo G, Nicoletta FP, Leggio A, Iemma F. Dual-Targeted Hyaluronic Acid/Albumin Micelle-Like Nanoparticles for the Vectorization of Doxorubicin. Pharmaceutics. 2021; 13(3):304. https://doi.org/10.3390/pharmaceutics13030304
Chicago/Turabian StyleCurcio, Manuela, Luis Diaz-Gomez, Giuseppe Cirillo, Fiore Pasquale Nicoletta, Antonella Leggio, and Francesca Iemma. 2021. "Dual-Targeted Hyaluronic Acid/Albumin Micelle-Like Nanoparticles for the Vectorization of Doxorubicin" Pharmaceutics 13, no. 3: 304. https://doi.org/10.3390/pharmaceutics13030304
APA StyleCurcio, M., Diaz-Gomez, L., Cirillo, G., Nicoletta, F. P., Leggio, A., & Iemma, F. (2021). Dual-Targeted Hyaluronic Acid/Albumin Micelle-Like Nanoparticles for the Vectorization of Doxorubicin. Pharmaceutics, 13(3), 304. https://doi.org/10.3390/pharmaceutics13030304







