Nanoparticle-Mediated Therapeutic Application for Modulation of Lysosomal Ion Channels and Functions
Abstract
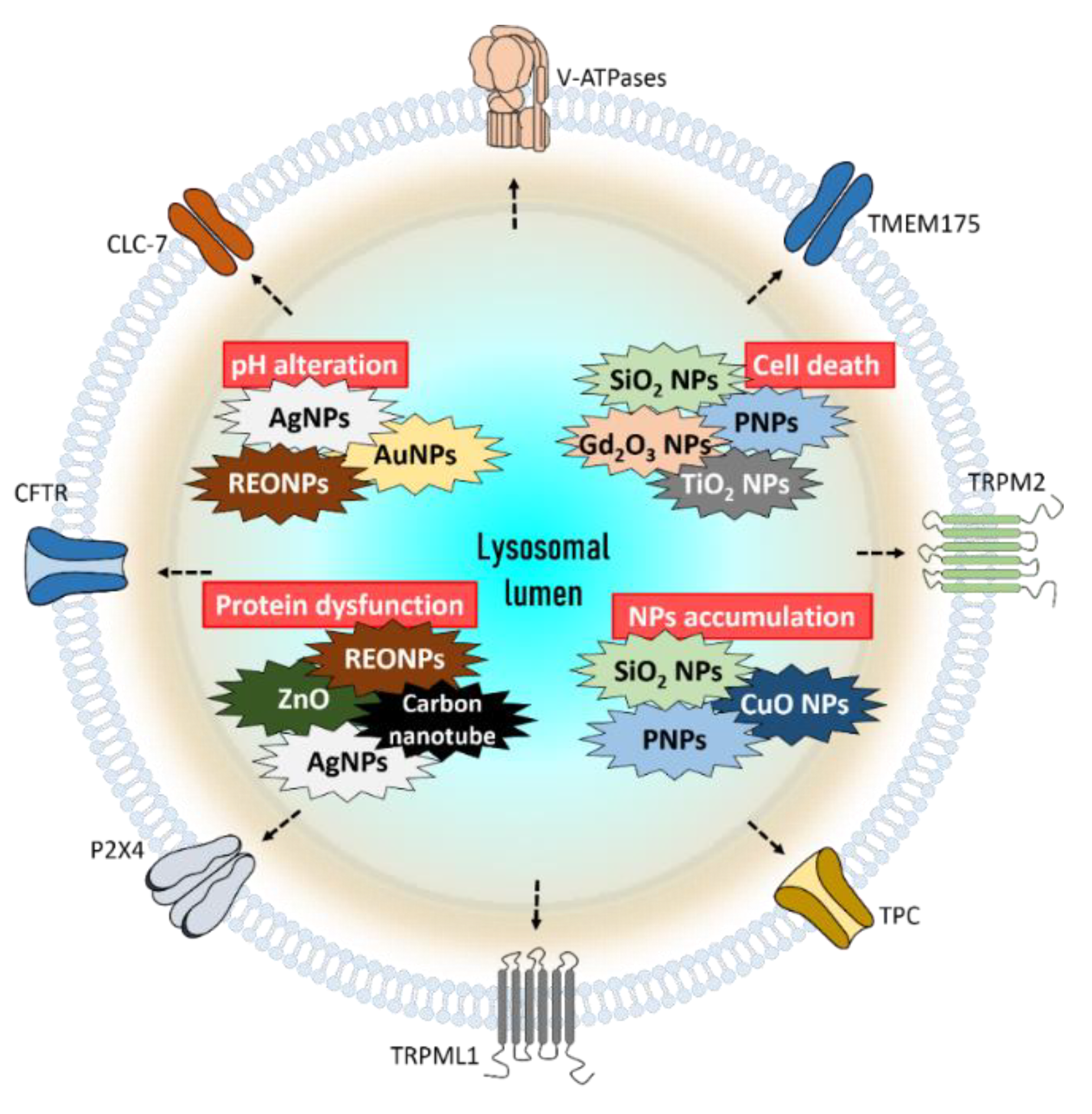
1. Lysosomal Target of Nanoparticles (NPs) and Modulation of NPs for Lysosomal Function
1.1. pH Alteration
1.2. Cell Viability
1.3. Protein Activity and Expression
1.4. Accumulation of NPs
2. Regulation of Lysosomal pH and Its Physiological Function
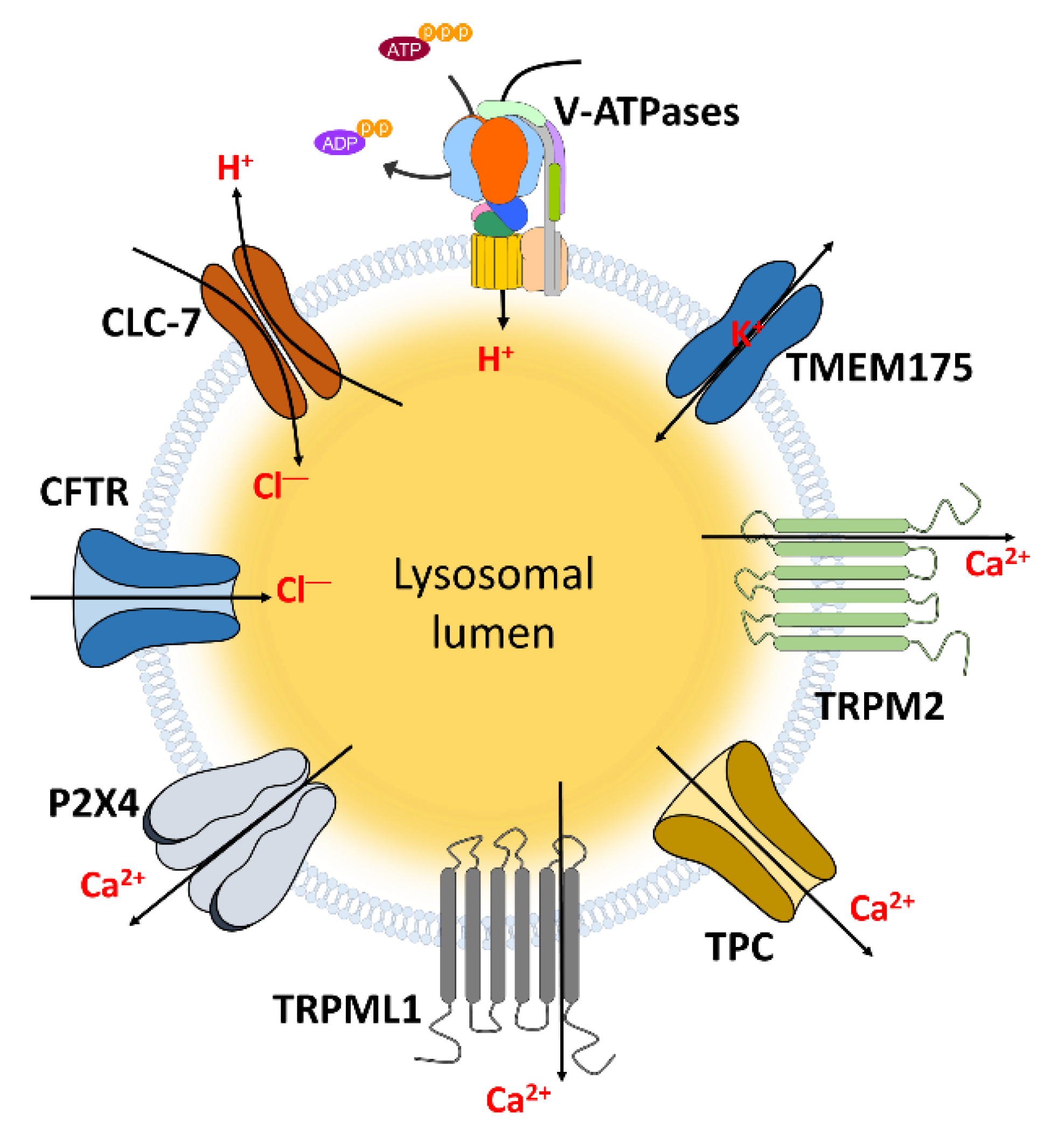
3. Lysosome-Associated Ion Channels for Lysosomal Function
3.1. CLC
3.2. Cystic Fibrosis (CF) Transmembrane Conductance Regulator (CFTR)
3.3. TRPs
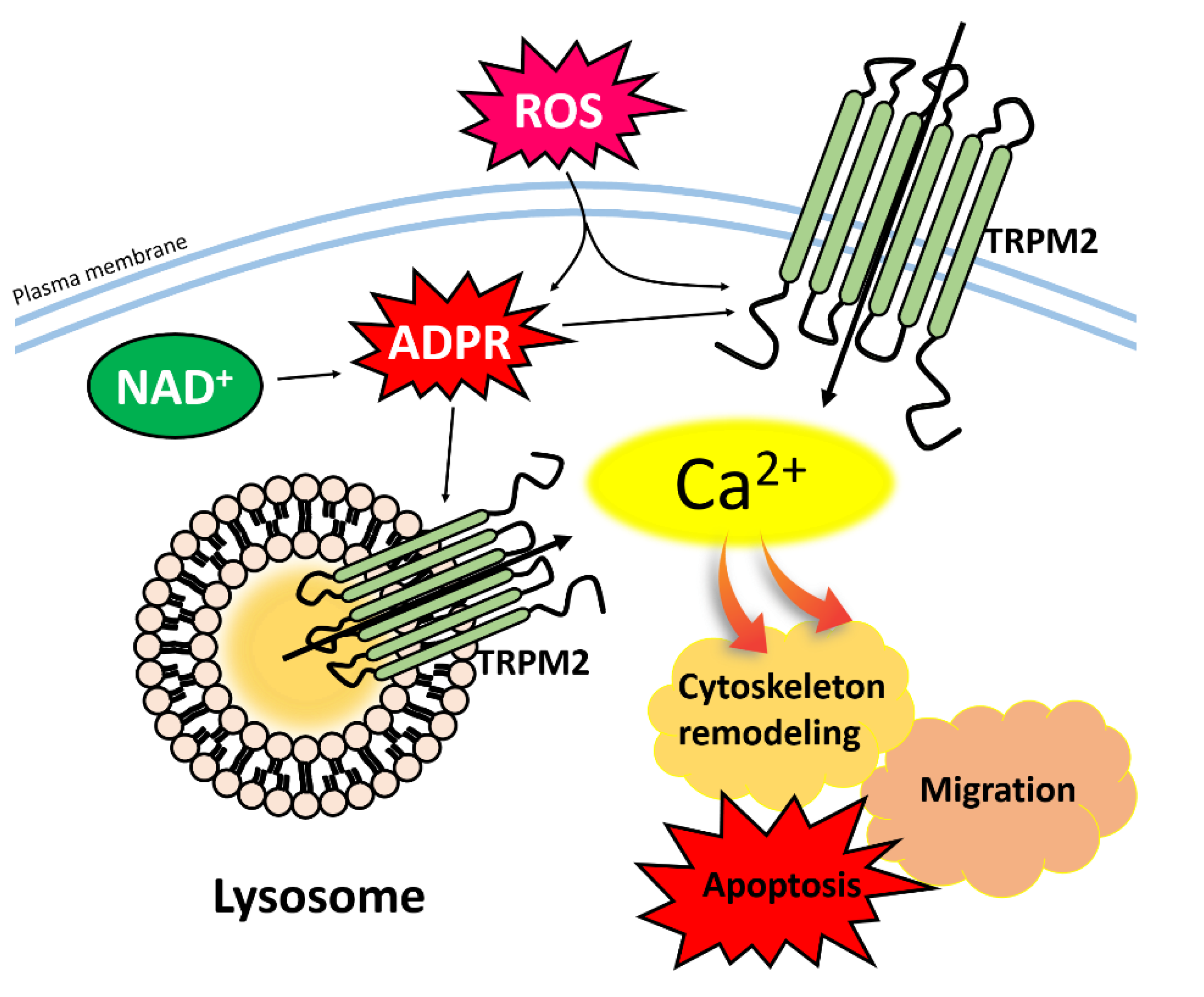
3.3.1. TRPM2
3.3.2. TRPMLs
3.4. TMEM175
3.5. Other Ca2+ Channels
3.5.1. Two Pore Channels (TPCs)
3.5.2. P2X4
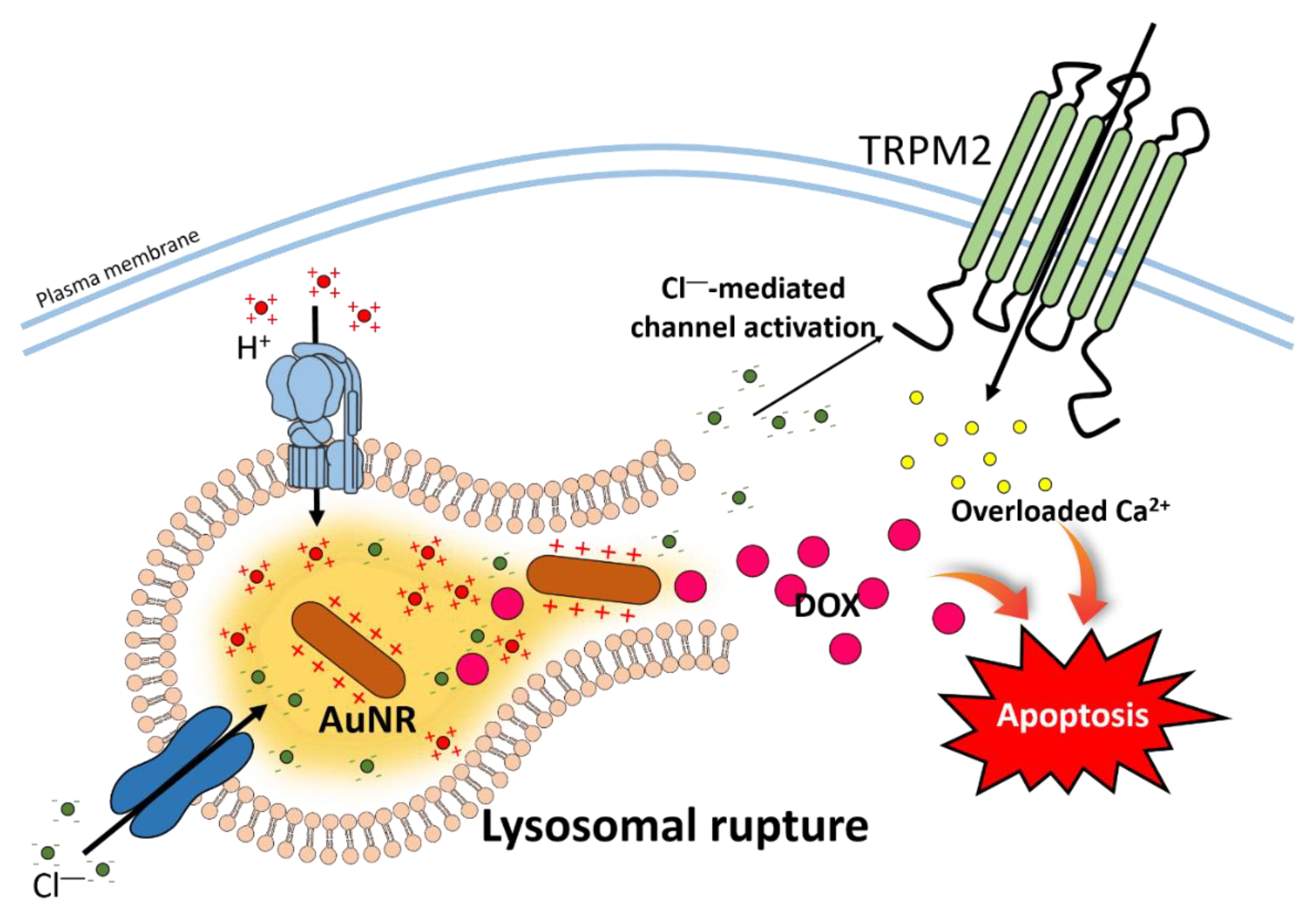
4. NP-Induced Proton Sponge Effect through Ion Channels in the Tumor System
5. Clinical Application and Limitation of Nanomaterials
6. Future Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- De Duve, C.; Wattiaux, R. Functions of lysosomes. Annu. Rev. Physiol. 1966, 28, 435–492. [Google Scholar] [CrossRef]
- de Duve, C. The lysosome turns fifty. Nat. Cell Biol. 2005, 7, 847–849. [Google Scholar] [CrossRef] [PubMed]
- Perera, R.M.; Zoncu, R. The Lysosome as a Regulatory Hub. Annu. Rev. Cell Dev. Biol. 2016, 32, 223–253. [Google Scholar] [CrossRef] [PubMed]
- Pu, J.; Guardia, C.M.; Keren-Kaplan, T.; Bonifacino, J.S. Mechanisms and functions of lysosome positioning. J. Cell Sci. 2016, 129, 4329–4339. [Google Scholar] [CrossRef]
- Mindell, J.A. Lysosomal acidification mechanisms. Annu. Rev. Physiol. 2012, 74, 69–86. [Google Scholar] [CrossRef]
- Walkley, S.U. Pathogenic mechanisms in lysosomal disease: A reappraisal of the role of the lysosome. Acta Paediatr. 2007, 96, 26–32. [Google Scholar] [CrossRef] [PubMed]
- Hipolito, V.E.B.; Ospina-Escobar, E.; Botelho, R.J. Lysosome remodelling and adaptation during phagocyte activation. Cell Microbiol. 2018, 20. [Google Scholar] [CrossRef]
- Herb, M.; Gluschko, A.; Schramm, M. LC3-associated phagocytosis—The highway to hell for phagocytosed microbes. Semin. Cell Dev. Biol. 2019. [Google Scholar] [CrossRef]
- Folts, C.J.; Scott-Hewitt, N.; Proschel, C.; Mayer-Proschel, M.; Noble, M. Lysosomal Re-acidification Prevents Lysosphingolipid-Induced Lysosomal Impairment and Cellular Toxicity. PLoS Biol. 2016, 14, e1002583. [Google Scholar] [CrossRef]
- Stern, S.T.; Adiseshaiah, P.P.; Crist, R.M. Autophagy and lysosomal dysfunction as emerging mechanisms of nanomaterial toxicity. Part. Fibre Toxicol. 2012, 9, 20. [Google Scholar] [CrossRef]
- Woldemichael, T.; Rosania, G.R. The physiological determinants of drug-induced lysosomal stress resistance. PLoS ONE 2017, 12, e0187627. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Wu, Y.; Jin, S.; Tian, Y.; Zhang, X.; Zhao, Y.; Yu, L.; Liang, X.J. Gold nanoparticles induce autophagosome accumulation through size-dependent nanoparticle uptake and lysosome impairment. ACS Nano 2011, 5, 8629–8639. [Google Scholar] [CrossRef] [PubMed]
- Miyayama, T.; Fujiki, K.; Matsuoka, M. Silver nanoparticles induce lysosomal-autophagic defects and decreased expression of transcription factor EB in A549 human lung adenocarcinoma cells. Toxicol. Vitr. 2018, 46, 148–154. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Ji, Z.; Qin, H.; Kang, X.; Sun, B.; Wang, M.; Chang, C.H.; Wang, X.; Zhang, H.; Zou, H.; et al. Interference in autophagosome fusion by rare earth nanoparticles disrupts autophagic flux and regulation of an interleukin-1beta producing inflammasome. ACS Nano 2014, 8, 10280–10292. [Google Scholar] [CrossRef]
- Winchester, B.G. Lysosomal membrane proteins. Eur. J. Paediatr. Neurol. 2001, 5, 11–19. [Google Scholar] [CrossRef]
- Schwake, M.; Schroder, B.; Saftig, P. Lysosomal membrane proteins and their central role in physiology. Traffic 2013, 14, 739–748. [Google Scholar] [CrossRef]
- Yu, S.; Melia, T.J. The coordination of membrane fission and fusion at the end of autophagosome maturation. Curr. Opin. Cell Biol. 2017, 47, 92–98. [Google Scholar] [CrossRef]
- Nascimbeni, A.C.; Codogno, P.; Morel, E. Phosphatidylinositol-3-phosphate in the regulation of autophagy membrane dynamics. FEBS J. 2017, 284, 1267–1278. [Google Scholar] [CrossRef]
- Wong, Y.C.; Kim, S.; Peng, W.; Krainc, D. Regulation and Function of Mitochondria-Lysosome Membrane Contact Sites in Cellular Homeostasis. Trends Cell Biol. 2019. [Google Scholar] [CrossRef]
- Helle, S.C.; Kanfer, G.; Kolar, K.; Lang, A.; Michel, A.H.; Kornmann, B. Organization and function of membrane contact sites. Biochim. Biophys. Acta 2013, 1833, 2526–2541. [Google Scholar] [CrossRef]
- Zoncu, R.; Bar-Peled, L.; Efeyan, A.; Wang, S.; Sancak, Y.; Sabatini, D.M. mTORC1 senses lysosomal amino acids through an inside-out mechanism that requires the vacuolar H(+)-ATPase. Science 2011, 334, 678–683. [Google Scholar] [CrossRef] [PubMed]
- Johnson, D.E.; Ostrowski, P.; Jaumouille, V.; Grinstein, S. The position of lysosomes within the cell determines their luminal pH. J. Cell Biol. 2016, 212, 677–692. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Gu, M.; Xu, H. Lysosomal Ion Channels as Decoders of Cellular Signals. Trends Biochem. Sci. 2019, 44, 110–124. [Google Scholar] [CrossRef] [PubMed]
- Nishino, I.; Fu, J.; Tanji, K.; Yamada, T.; Shimojo, S.; Koori, T.; Mora, M.; Riggs, J.E.; Oh, S.J.; Koga, Y.; et al. Primary LAMP-2 deficiency causes X-linked vacuolar cardiomyopathy and myopathy (Danon disease). Nature 2000, 406, 906–910. [Google Scholar] [CrossRef] [PubMed]
- Kornak, U.; Kasper, D.; Bosl, M.R.; Kaiser, E.; Schweizer, M.; Schulz, A.; Friedrich, W.; Delling, G.; Jentsch, T.J. Loss of the ClC-7 chloride channel leads to osteopetrosis in mice and man. Cell 2001, 104, 205–215. [Google Scholar] [CrossRef]
- Berkovic, S.F.; Dibbens, L.M.; Oshlack, A.; Silver, J.D.; Katerelos, M.; Vears, D.F.; Lullmann-Rauch, R.; Blanz, J.; Zhang, K.W.; Stankovich, J.; et al. Array-based gene discovery with three unrelated subjects shows SCARB2/LIMP-2 deficiency causes myoclonus epilepsy and glomerulosclerosis. Am. J. Hum. Genet. 2008, 82, 673–684. [Google Scholar] [CrossRef]
- Wang, F.; Gomez-Sintes, R.; Boya, P. Lysosomal membrane permeabilization and cell death. Traffic 2018, 19, 918–931. [Google Scholar] [CrossRef]
- Mrschtik, M.; Ryan, K.M. Lysosomal proteins in cell death and autophagy. FEBS J. 2015, 282, 1858–1870. [Google Scholar] [CrossRef]
- Serrano-Puebla, A.; Boya, P. Lysosomal membrane permeabilization in cell death: New evidence and implications for health and disease. Ann. N. Y. Acad. Sci. 2016, 1371, 30–44. [Google Scholar] [CrossRef]
- Micsenyi, M.C.; Sikora, J.; Stephney, G.; Dobrenis, K.; Walkley, S.U. Lysosomal membrane permeability stimulates protein aggregate formation in neurons of a lysosomal disease. J. Neurosci. 2013, 33, 10815–10827. [Google Scholar] [CrossRef]
- Venkatesan, R.; Park, Y.U.; Ji, E.; Yeo, E.J.; Kim, S.Y. Malathion increases apoptotic cell death by inducing lysosomal membrane permeabilization in N2a neuroblastoma cells: A model for neurodegeneration in Alzheimer’s disease. Cell Death Discov. 2017, 3, 17007. [Google Scholar] [CrossRef] [PubMed]
- Song, W.; Popp, L.; Yang, J.; Kumar, A.; Gangoli, V.S.; Segatori, L. The autophagic response to polystyrene nanoparticles is mediated by transcription factor EB and depends on surface charge. J. Nanobiotechnol. 2015, 13, 87. [Google Scholar] [CrossRef] [PubMed]
- Lunova, M.; Prokhorov, A.; Jirsa, M.; Hof, M.; Olzynska, A.; Jurkiewicz, P.; Kubinova, S.; Lunov, O.; Dejneka, A. Nanoparticle core stability and surface functionalization drive the mTOR signaling pathway in hepatocellular cell lines. Sci. Rep. 2017, 7, 16049. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Salvati, A.; Boya, P. Lysosome-dependent cell death and deregulated autophagy induced by amine-modified polystyrene nanoparticles. Open Biol. 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Jessop, F.; Hamilton, R.F., Jr.; Rhoderick, J.F.; Fletcher, P.; Holian, A. Phagolysosome acidification is required for silica and engineered nanoparticle-induced lysosome membrane permeabilization and resultant NLRP3 inflammasome activity. Toxicol. Appl. Pharm. 2017, 318, 58–68. [Google Scholar] [CrossRef]
- Popp, L.; Tran, V.; Patel, R.; Segatori, L. Autophagic response to cellular exposure to titanium dioxide nanoparticles. Acta Biomater. 2018, 79, 354–363. [Google Scholar] [CrossRef]
- Jin, Y.; Chen, S.; Duan, J.; Jia, G.; Zhang, J. Europium-doped Gd2O3 nanotubes cause the necrosis of primary mouse bone marrow stromal cells through lysosome and mitochondrion damage. J. Inorg. Biochem. 2015, 146, 28–36. [Google Scholar] [CrossRef]
- Korolchuk, V.I.; Saiki, S.; Lichtenberg, M.; Siddiqi, F.H.; Roberts, E.A.; Imarisio, S.; Jahreiss, L.; Sarkar, S.; Futter, M.; Menzies, F.M.; et al. Lysosomal positioning coordinates cellular nutrient responses. Nat. Cell Biol. 2011, 13, 453–460. [Google Scholar] [CrossRef]
- Noda, T.; Ohsumi, Y. Tor, a phosphatidylinositol kinase homologue, controls autophagy in yeast. J. Biol. Chem. 1998, 273, 3963–3966. [Google Scholar] [CrossRef]
- Eskelinen, E.L. Roles of LAMP-1 and LAMP-2 in lysosome biogenesis and autophagy. Mol. Asp. Med. 2006, 27, 495–502. [Google Scholar] [CrossRef]
- Chen, Y.; Wang, M.; Zhang, T.; Du, E.; Liu, Y.; Qi, S.; Xu, Y.; Zhang, Z. Autophagic effects and mechanisms of silver nanoparticles in renal cells under low dose exposure. Ecotoxicol. Environ. Saf. 2018, 166, 71–77. [Google Scholar] [CrossRef] [PubMed]
- Roy, R.; Singh, S.K.; Chauhan, L.K.; Das, M.; Tripathi, A.; Dwivedi, P.D. Zinc oxide nanoparticles induce apoptosis by enhancement of autophagy via PI3K/Akt/mTOR inhibition. Toxicol. Lett. 2014, 227, 29–40. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Shi, S.S.; Zhang, J.Q.; Zhang, Y.J.; Zhang, L.; Liu, Y.; Jin, P.P.; Wei, P.F.; Shi, R.H.; Zhou, W.; et al. Giant Cellular Vacuoles Induced by Rare Earth Oxide Nanoparticles are Abnormally Enlarged Endo/Lysosomes and Promote mTOR-Dependent TFEB Nucleus Translocation. Small 2016, 12, 5759–5768. [Google Scholar] [CrossRef] [PubMed]
- Qin, X.; Zhang, J.; Wang, B.; Xu, G.; Zou, Z. LAMP-2 mediates oxidative stress-dependent cell death in Zn(2+)-treated lung epithelium cells. Biochem. Biophys. Res. Commun. 2017, 488, 177–181. [Google Scholar] [CrossRef]
- Cohignac, V.; Landry, M.J.; Ridoux, A.; Pinault, M.; Annangi, B.; Gerdil, A.; Herlin-Boime, N.; Mayne, M.; Haruta, M.; Codogno, P.; et al. Carbon nanotubes, but not spherical nanoparticles, block autophagy by a shape-related targeting of lysosomes in murine macrophages. Autophagy 2018, 14, 1323–1334. [Google Scholar] [CrossRef]
- Hirokawa, N.; Noda, Y. Intracellular transport and kinesin superfamily proteins, KIFs: Structure, function, and dynamics. Physiol. Rev. 2008, 88, 1089–1118. [Google Scholar] [CrossRef]
- Hollenbeck, P.J.; Swanson, J.A. Radial extension of macrophage tubular lysosomes supported by kinesin. Nature 1990, 346, 864–866. [Google Scholar] [CrossRef]
- Paschal, B.M.; Vallee, R.B. Retrograde transport by the microtubule-associated protein MAP 1C. Nature 1987, 330, 181–183. [Google Scholar] [CrossRef]
- Harada, A.; Takei, Y.; Kanai, Y.; Tanaka, Y.; Nonaka, S.; Hirokawa, N. Golgi vesiculation and lysosome dispersion in cells lacking cytoplasmic dynein. J. Cell Biol. 1998, 141, 51–59. [Google Scholar] [CrossRef]
- Yuzaki, M. Snapin snaps into the dynein complex for late endosome-lysosome trafficking and autophagy. Neuron 2010, 68, 4–6. [Google Scholar] [CrossRef][Green Version]
- Chun, Y.; Kim, J. Autophagy: An Essential Degradation Program for Cellular Homeostasis and Life. Cells 2018, 7, 278. [Google Scholar] [CrossRef] [PubMed]
- Settembre, C.; Ballabio, A. Lysosome: Regulator of lipid degradation pathways. Trends Cell Biol. 2014, 24, 743–750. [Google Scholar] [CrossRef] [PubMed]
- Lawrence, R.E.; Zoncu, R. The lysosome as a cellular centre for signalling, metabolism and quality control. Nat. Cell Biol. 2019, 21, 133–142. [Google Scholar] [CrossRef]
- Ferguson, S.M. Neuronal lysosomes. Neurosci. Lett. 2019, 697, 1–9. [Google Scholar] [CrossRef]
- Miyayama, T.; Matsuoka, M. Involvement of lysosomal dysfunction in silver nanoparticle-induced cellular damage in A549 human lung alveolar epithelial cells. J. Occup. Med. Toxicol. 2016, 11, 1. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zou, Z.; Wang, B.; Xu, G.; Wu, Q.; Zhang, Y.; Yuan, Z.; Yang, X.; Yu, C. Lysosomal deposition of copper oxide nanoparticles triggers HUVEC cells death. Biomaterials 2018, 161, 228–239. [Google Scholar] [CrossRef] [PubMed]
- Schutz, I.; Lopez-Hernandez, T.; Gao, Q.; Puchkov, D.; Jabs, S.; Nordmeyer, D.; Schmudde, M.; Ruhl, E.; Graf, C.M.; Haucke, V. Lysosomal Dysfunction Caused by Cellular Accumulation of Silica Nanoparticles. J. Biol. Chem. 2016, 291, 14170–14184. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Bexiga, M.G.; Anguissola, S.; Boya, P.; Simpson, J.C.; Salvati, A.; Dawson, K.A. Time resolved study of cell death mechanisms induced by amine-modified polystyrene nanoparticles. Nanoscale 2013, 5, 10868–10876. [Google Scholar] [CrossRef]
- Ohkuma, S.; Moriyama, Y.; Takano, T. Identification and characterization of a proton pump on lysosomes by fluorescein-isothiocyanate-dextran fluorescence. Proc. Natl. Acad. Sci. USA 1982, 79, 2758–2762. [Google Scholar] [CrossRef]
- Cipriano, D.J.; Wang, Y.; Bond, S.; Hinton, A.; Jefferies, K.C.; Qi, J.; Forgac, M. Structure and regulation of the vacuolar ATPases. Biochim. Biophys. Acta 2008, 1777, 599–604. [Google Scholar] [CrossRef]
- Hirata, T.; Iwamoto-Kihara, A.; Sun-Wada, G.H.; Okajima, T.; Wada, Y.; Futai, M. Subunit rotation of vacuolar-type proton pumping ATPase: Relative rotation of the G and C subunits. J. Biol. Chem. 2003, 278, 23714–23719. [Google Scholar] [CrossRef]
- Yokoyama, K.; Nakano, M.; Imamura, H.; Yoshida, M.; Tamakoshi, M. Rotation of the proteolipid ring in the V-ATPase. J. Biol. Chem. 2003, 278, 24255–24258. [Google Scholar] [CrossRef] [PubMed]
- Ishida, Y.; Nayak, S.; Mindell, J.A.; Grabe, M. A model of lysosomal pH regulation. J. Gen. Physiol. 2013, 141, 705–720. [Google Scholar] [CrossRef] [PubMed]
- Kasper, D.; Planells-Cases, R.; Fuhrmann, J.C.; Scheel, O.; Zeitz, O.; Ruether, K.; Schmitt, A.; Poet, M.; Steinfeld, R.; Schweizer, M.; et al. Loss of the chloride channel ClC-7 leads to lysosomal storage disease and neurodegeneration. EMBO J. 2005, 24, 1079–1091. [Google Scholar] [CrossRef] [PubMed]
- Checchetto, V.; Teardo, E.; Carraretto, L.; Leanza, L.; Szabo, I. Physiology of intracellular potassium channels: A unifying role as mediators of counterion fluxes? Biochim. Biophys. Acta 2016, 1857, 1258–1266. [Google Scholar] [CrossRef] [PubMed]
- Betz, C.; Hall, M.N. Where is mTOR and what is it doing there? J. Cell Biol. 2013, 203, 563–574. [Google Scholar] [CrossRef] [PubMed]
- Sundler, R. Lysosomal and cytosolic pH as regulators of exocytosis in mouse macrophages. Acta Physiol. Scand. 1997, 161, 553–556. [Google Scholar] [CrossRef]
- Tapper, H.; Sundler, R. Role of lysosomal and cytosolic pH in the regulation of macrophage lysosomal enzyme secretion. Biochem. J. 1990, 272, 407–414. [Google Scholar] [CrossRef]
- Camargo, M.J.; Sumpio, B.E.; Maack, T. Renal hydrolysis of absorbed protein: Influence of load and lysosomal pH. Am. J. Physiol. 1984, 247, F656–F664. [Google Scholar] [CrossRef]
- Smith, M.L.; Greene, A.A.; Potashnik, R.; Mendoza, S.A.; Schneider, J.A. Lysosomal cystine transport. Effect of intralysosomal pH and membrane potential. J. Biol. Chem. 1987, 262, 1244–1253. [Google Scholar]
- Lee, D.; Hong, J.H. Physiological application of nanoparticles in calcium-related proteins and channels. Nanomed. 2019. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wang, T.; Zhao, Z.; Weinman, S.A. The ClC-3 chloride channel promotes acidification of lysosomes in CHO-K1 and Huh-7 cells. Am. J. Physiol. Cell Physiol. 2002, 282, C1483–C1491. [Google Scholar] [CrossRef] [PubMed]
- Okamoto, F.; Kajiya, H.; Toh, K.; Uchida, S.; Yoshikawa, M.; Sasaki, S.; Kido, M.A.; Tanaka, T.; Okabe, K. Intracellular ClC-3 chloride channels promote bone resorption in vitro through organelle acidification in mouse osteoclasts. Am. J. Physiol. Cell Physiol. 2008, 294, C693–C701. [Google Scholar] [CrossRef] [PubMed]
- Poet, M.; Kornak, U.; Schweizer, M.; Zdebik, A.A.; Scheel, O.; Hoelter, S.; Wurst, W.; Schmitt, A.; Fuhrmann, J.C.; Planells-Cases, R.; et al. Lysosomal storage disease upon disruption of the neuronal chloride transport protein ClC-6. Proc. Natl. Acad. Sci. USA 2006, 103, 13854–13859. [Google Scholar] [CrossRef] [PubMed]
- Henriksen, K.; Gram, J.; Neutzsky-Wulff, A.V.; Jensen, V.K.; Dziegiel, M.H.; Bollerslev, J.; Karsdal, M.A. Characterization of acid flux in osteoclasts from patients harboring a G215R mutation in ClC-7. Biochem. Biophys. Res. Commun. 2009, 378, 804–809. [Google Scholar] [CrossRef]
- Henriksen, K.; Sorensen, M.G.; Jensen, V.K.; Dziegiel, M.H.; Nosjean, O.; Karsdal, M.A. Ion transporters involved in acidification of the resorption lacuna in osteoclasts. Calcif. Tissue Int. 2008, 83, 230–242. [Google Scholar] [CrossRef]
- Wen, X.; Lacruz, R.S.; Paine, M.L. Dental and Cranial Pathologies in Mice Lacking the Cl(-) /H(+) -Exchanger ClC-7. Anat. Rec. 2015, 298, 1502–1508. [Google Scholar] [CrossRef]
- Guo, J.; Bervoets, T.J.; Henriksen, K.; Everts, V.; Bronckers, A.L. Null mutation of chloride channel 7 (Clcn7) impairs dental root formation but does not affect enamel mineralization. Cell Tissue Res. 2016, 363, 361–370. [Google Scholar] [CrossRef][Green Version]
- Murphy, M.P.; LeVine, H., 3rd. Alzheimer’s disease and the amyloid-beta peptide. J. Alzheimers Dis. 2010, 19, 311–323. [Google Scholar] [CrossRef]
- Majumdar, A.; Capetillo-Zarate, E.; Cruz, D.; Gouras, G.K.; Maxfield, F.R. Degradation of Alzheimer’s amyloid fibrils by microglia requires delivery of ClC-7 to lysosomes. Mol. Biol. Cell 2011, 22, 1664–1676. [Google Scholar] [CrossRef]
- Jentsch, T.J. Chloride and the endosomal-lysosomal pathway: Emerging roles of CLC chloride transporters. J. Physiol. 2007, 578, 633–640. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Wei, Q.; He, A.; Jia, R.; Xiao, Y. CLC-7: A potential therapeutic target for the treatment of osteoporosis and neurodegeneration. Biochem. Biophys. Res. Commun. 2009, 384, 277–279. [Google Scholar] [CrossRef] [PubMed]
- Weinert, S.; Jabs, S.; Supanchart, C.; Schweizer, M.; Gimber, N.; Richter, M.; Rademann, J.; Stauber, T.; Kornak, U.; Jentsch, T.J. Lysosomal pathology and osteopetrosis upon loss of H+-driven lysosomal Cl- accumulation. Science 2010, 328, 1401–1403. [Google Scholar] [CrossRef]
- Sartelet, A.; Stauber, T.; Coppieters, W.; Ludwig, C.F.; Fasquelle, C.; Druet, T.; Zhang, Z.; Ahariz, N.; Cambisano, N.; Jentsch, T.J.; et al. A missense mutation accelerating the gating of the lysosomal Cl-/H+-exchanger ClC-7/Ostm1 causes osteopetrosis with gingival hamartomas in cattle. Dis. Model. Mech. 2014, 7, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Haggie, P.M.; Verkman, A.S. Unimpaired lysosomal acidification in respiratory epithelial cells in cystic fibrosis. J. Biol. Chem. 2009, 284, 7681–7686. [Google Scholar] [CrossRef] [PubMed]
- Di, A.; Brown, M.E.; Deriy, L.V.; Li, C.; Szeto, F.L.; Chen, Y.; Huang, P.; Tong, J.; Naren, A.P.; Bindokas, V.; et al. CFTR regulates phagosome acidification in macrophages and alters bactericidal activity. Nat. Cell Biol. 2006, 8, 933–944. [Google Scholar] [CrossRef]
- Sumoza-Toledo, A.; Lange, I.; Cortado, H.; Bhagat, H.; Mori, Y.; Fleig, A.; Penner, R.; Partida-Sanchez, S. Dendritic cell maturation and chemotaxis is regulated by TRPM2-mediated lysosomal Ca2+ release. FASEB J. 2011, 25, 3529–3542. [Google Scholar] [CrossRef]
- Li, F.; Abuarab, N.; Sivaprasadarao, A. Reciprocal regulation of actin cytoskeleton remodelling and cell migration by Ca2+ and Zn2+: Role of TRPM2 channels. J. Cell Sci. 2016, 129, 2016–2029. [Google Scholar] [CrossRef]
- Lange, I.; Yamamoto, S.; Partida-Sanchez, S.; Mori, Y.; Fleig, A.; Penner, R. TRPM2 functions as a lysosomal Ca2+-release channel in beta cells. Sci. Signal. 2009, 2, ra23. [Google Scholar] [CrossRef]
- Abuarab, N.; Munsey, T.S.; Jiang, L.H.; Li, J.; Sivaprasadarao, A. High glucose-induced ROS activates TRPM2 to trigger lysosomal membrane permeabilization and Zn(2+)-mediated mitochondrial fission. Sci. Signal. 2017, 10. [Google Scholar] [CrossRef]
- Katsnelson, M.A.; Lozada-Soto, K.M.; Russo, H.M.; Miller, B.A.; Dubyak, G.R. NLRP3 inflammasome signaling is activated by low-level lysosome disruption but inhibited by extensive lysosome disruption: Roles for K+ efflux and Ca2+ influx. Am. J. Physiol. Cell Physiol. 2016, 311, C83–C100. [Google Scholar] [CrossRef] [PubMed]
- Soyombo, A.A.; Tjon-Kon-Sang, S.; Rbaibi, Y.; Bashllari, E.; Bisceglia, J.; Muallem, S.; Kiselyov, K. TRP-ML1 regulates lysosomal pH and acidic lysosomal lipid hydrolytic activity. J. Biol. Chem. 2006, 281, 7294–7301. [Google Scholar] [CrossRef] [PubMed]
- Chandra, M.; Zhou, H.; Li, Q.; Muallem, S.; Hofmann, S.L.; Soyombo, A.A. A role for the Ca2+ channel TRPML1 in gastric acid secretion, based on analysis of knockout mice. Gastroenterology 2011, 140, 857–867. [Google Scholar] [CrossRef] [PubMed]
- Erkhembaatar, M.; Gu, D.R.; Lee, S.H.; Yang, Y.M.; Park, S.; Muallem, S.; Shin, D.M.; Kim, M.S. Lysosomal Ca2+ Signaling is Essential for Osteoclastogenesis and Bone Remodeling. J. Bone Min. Res. 2017, 32, 385–396. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Li, X.; Walsh, S.W.; Zhang, Y.; Abais, J.M.; Boini, K.M.; Li, P.L. Intracellular two-phase Ca2+ release and apoptosis controlled by TRP-ML1 channel activity in coronary arterial myocytes. Am. J. Physiol. Cell Physiol. 2013, 304, C458–C466. [Google Scholar] [CrossRef] [PubMed]
- Samie, M.; Wang, X.; Zhang, X.; Goschka, A.; Li, X.; Cheng, X.; Gregg, E.; Azar, M.; Zhuo, Y.; Garrity, A.G.; et al. A TRP channel in the lysosome regulates large particle phagocytosis via focal exocytosis. Dev. Cell 2013, 26, 511–524. [Google Scholar] [CrossRef]
- Shen, D.; Wang, X.; Li, X.; Zhang, X.; Yao, Z.; Dibble, S.; Dong, X.P.; Yu, T.; Lieberman, A.P.; Showalter, H.D.; et al. Lipid storage disorders block lysosomal trafficking by inhibiting a TRP channel and lysosomal calcium release. Nat. Commun. 2012, 3, 731. [Google Scholar] [CrossRef]
- Weiss, N. Cross-talk between TRPML1 channel, lipids and lysosomal storage diseases. Commun. Integr. Biol. 2012, 5, 111–113. [Google Scholar] [CrossRef]
- Zeevi, D.A.; Frumkin, A.; Bach, G. TRPML and lysosomal function. Biochim. Biophys. Acta 2007, 1772, 851–858. [Google Scholar] [CrossRef]
- Dong, X.P.; Cheng, X.; Mills, E.; Delling, M.; Wang, F.; Kurz, T.; Xu, H. The type IV mucolipidosis-associated protein TRPML1 is an endolysosomal iron release channel. Nature 2008, 455, 992–996. [Google Scholar] [CrossRef]
- Miedel, M.T.; Rbaibi, Y.; Guerriero, C.J.; Colletti, G.; Weixel, K.M.; Weisz, O.A.; Kiselyov, K. Membrane traffic and turnover in TRP-ML1-deficient cells: A revised model for mucolipidosis type IV pathogenesis. J. Exp. Med. 2008, 205, 1477–1490. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Jin, S.; Yi, F.; Li, P.L. TRP-ML1 functions as a lysosomal NAADP-sensitive Ca2+ release channel in coronary arterial myocytes. J. Cell Mol. Med. 2009, 13, 3174–3185. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; McBrayer, M.K.; Wolfe, D.M.; Haslett, L.J.; Kumar, A.; Sato, Y.; Lie, P.P.; Mohan, P.; Coffey, E.E.; Kompella, U.; et al. Presenilin 1 Maintains Lysosomal Ca(2+) Homeostasis via TRPML1 by Regulating vATPase-Mediated Lysosome Acidification. Cell Rep. 2015, 12, 1430–1444. [Google Scholar] [CrossRef]
- Sterea, A.M.; Almasi, S.; El Hiani, Y. The hidden potential of lysosomal ion channels: A new era of oncogenes. Cell Calcium. 2018, 72, 91–103. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.; Zhao, Z.; Li, Q.; Tan, Z. Lysosomal Potassium Channels: Potential Roles in Lysosomal Function and Neurodegenerative Diseases. CNS Neurol. Disord. Drug Targets 2018, 17, 261–266. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Yue, J. TPC2 mediates autophagy progression and extracellular vesicle secretion in cancer cells. Exp. Cell Res. 2018, 370, 478–489. [Google Scholar] [CrossRef] [PubMed]
- Grimm, C.; Bartel, K.; Vollmar, A.M.; Biel, M. Endolysosomal Cation Channels and Cancer-A Link with Great Potential. Pharmaceuticals 2018, 11, 4. [Google Scholar] [CrossRef]
- Lin-Moshier, Y.; Keebler, M.V.; Hooper, R.; Boulware, M.J.; Liu, X.; Churamani, D.; Abood, M.E.; Walseth, T.F.; Brailoiu, E.; Patel, S.; et al. The Two-pore channel (TPC) interactome unmasks isoform-specific roles for TPCs in endolysosomal morphology and cell pigmentation. Proc. Natl. Acad. Sci. USA 2014, 111, 13087–13092. [Google Scholar] [CrossRef]
- Hockey, L.N.; Kilpatrick, B.S.; Eden, E.R.; Lin-Moshier, Y.; Brailoiu, G.C.; Brailoiu, E.; Futter, C.E.; Schapira, A.H.; Marchant, J.S.; Patel, S. Dysregulation of lysosomal morphology by pathogenic LRRK2 is corrected by TPC2 inhibition. J. Cell Sci. 2015, 128, 232–238. [Google Scholar] [CrossRef]
- Rivero-Rios, P.; Gomez-Suaga, P.; Fernandez, B.; Madero-Perez, J.; Schwab, A.J.; Ebert, A.D.; Hilfiker, S. Alterations in late endocytic trafficking related to the pathobiology of LRRK2-linked Parkinson’s disease. Biochem Soc. Trans. 2015, 43, 390–395. [Google Scholar] [CrossRef]
- Cao, Q.; Zhong, X.Z.; Zou, Y.; Murrell-Lagnado, R.; Zhu, M.X.; Dong, X.P. Calcium release through P2X4 activates calmodulin to promote endolysosomal membrane fusion. J. Cell Biol. 2015, 209, 879–894. [Google Scholar] [CrossRef] [PubMed]
- Fois, G.; Winkelmann, V.E.; Bareis, L.; Staudenmaier, L.; Hecht, E.; Ziller, C.; Ehinger, K.; Schymeinsky, J.; Kranz, C.; Frick, M. ATP is stored in lamellar bodies to activate vesicular P2X4 in an autocrine fashion upon exocytosis. J. Gen. Physiol. 2018, 150, 277–291. [Google Scholar] [CrossRef] [PubMed]
- Le Guilcher, C.; Garcin, I.; Dellis, O.; Cauchois, F.; Tebbi, A.; Doignon, I.; Guettier, C.; Julien, B.; Tordjmann, T. The P2X4 purinergic receptor regulates hepatic myofibroblast activation during liver fibrogenesis. J. Hepatol. 2018, 69, 644–653. [Google Scholar] [CrossRef] [PubMed]
- Poroca, D.R.; Pelis, R.M.; Chappe, V.M. ClC Channels and Transporters: Structure, Physiological Functions, and Implications in Human Chloride Channelopathies. Front. Pharm. 2017, 8, 151. [Google Scholar] [CrossRef]
- Stauber, T.; Jentsch, T.J. Sorting motifs of the endosomal/lysosomal CLC chloride transporters. J. Biol. Chem. 2010, 285, 34537–34548. [Google Scholar] [CrossRef]
- Zifarelli, G. A tale of two CLCs: Biophysical insights toward understanding ClC-5 and ClC-7 function in endosomes and lysosomes. J. Physiol. 2015, 593, 4139–4150. [Google Scholar] [CrossRef]
- Graves, A.R.; Curran, P.K.; Smith, C.L.; Mindell, J.A. The Cl-/H+ antiporter ClC-7 is the primary chloride permeation pathway in lysosomes. Nature 2008, 453, 788–792. [Google Scholar] [CrossRef]
- Braun, A.P. Identification of ClC-7 as a major pathway for Cl- movement in lysosomes. Channels 2008, 2, 309. [Google Scholar] [CrossRef][Green Version]
- Lange, P.F.; Wartosch, L.; Jentsch, T.J.; Fuhrmann, J.C. ClC-7 requires Ostm1 as a beta-subunit to support bone resorption and lysosomal function. Nature 2006, 440, 220–223. [Google Scholar] [CrossRef]
- Fuller, C.M.; Benos, D.J. Cftr! Am. J. Physiol. 1992, 263, C267–C286. [Google Scholar] [CrossRef]
- Meng, X.; Clews, J.; Kargas, V.; Wang, X.; Ford, R.C. The cystic fibrosis transmembrane conductance regulator (CFTR) and its stability. Cell Mol. Life Sci. 2017, 74, 23–38. [Google Scholar] [CrossRef] [PubMed]
- Moran, O. The gating of the CFTR channel. Cell Mol. Life Sci. 2017, 74, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Borowitz, D. CFTR, bicarbonate, and the pathophysiology of cystic fibrosis. Pediatr. Pulmonol. 2015, 50, S24–S30. [Google Scholar] [CrossRef] [PubMed]
- Csanady, L.; Vergani, P.; Gadsby, D.C. Structure, Gating, and Regulation of the Cftr Anion Channel. Physiol. Rev. 2019, 99, 707–738. [Google Scholar] [CrossRef] [PubMed]
- Riordan, J.R. CFTR function and prospects for therapy. Annu. Rev. Biochem. 2008, 77, 701–726. [Google Scholar] [CrossRef]
- Bradbury, N.A. Intracellular CFTR: Localization and function. Physiol. Rev. 1999, 79, S175–S191. [Google Scholar] [CrossRef]
- Swanson, J. CFTR: Helping to acidify macrophage lysosomes. Nat. Cell Biol. 2006, 8, 908–909. [Google Scholar] [CrossRef]
- Liu, J.; Lu, W.; Guha, S.; Baltazar, G.C.; Coffey, E.E.; Laties, A.M.; Rubenstein, R.C.; Reenstra, W.W.; Mitchell, C.H. Cystic fibrosis transmembrane conductance regulator contributes to reacidification of alkalinized lysosomes in RPE cells. Am. J. Physiol. Cell Physiol. 2012, 303, C160–C169. [Google Scholar] [CrossRef]
- Clapham, D.E.; Runnels, L.W.; Strubing, C. The TRP ion channel family. Nat. Rev. Neurosci. 2001, 2, 387–396. [Google Scholar] [CrossRef]
- Ramsey, I.S.; Delling, M.; Clapham, D.E. An introduction to TRP channels. Annu. Rev. Physiol. 2006, 68, 619–647. [Google Scholar] [CrossRef]
- Bootman, M.D.; Chehab, T.; Bultynck, G.; Parys, J.B.; Rietdorf, K. The regulation of autophagy by calcium signals: Do we have a consensus? Cell Calcium. 2018, 70, 32–46. [Google Scholar] [CrossRef] [PubMed]
- Sano, Y.; Inamura, K.; Miyake, A.; Mochizuki, S.; Yokoi, H.; Matsushime, H.; Furuichi, K. Immunocyte Ca2+ influx system mediated by LTRPC2. Science 2001, 293, 1327–1330. [Google Scholar] [CrossRef] [PubMed]
- Perraud, A.L.; Fleig, A.; Dunn, C.A.; Bagley, L.A.; Launay, P.; Schmitz, C.; Stokes, A.J.; Zhu, Q.; Bessman, M.J.; Penner, R.; et al. ADP-ribose gating of the calcium-permeable LTRPC2 channel revealed by Nudix motif homology. Nature 2001, 411, 595–599. [Google Scholar] [CrossRef]
- Fleig, A.; Penner, R. Emerging roles of TRPM channels. Novartis. Found. Symp. 2004, 258, 248–258; discussion 258–266. [Google Scholar] [PubMed]
- Wehage, E.; Eisfeld, J.; Heiner, I.; Jungling, E.; Zitt, C.; Luckhoff, A. Activation of the cation channel long transient receptor potential channel 2 (LTRPC2) by hydrogen peroxide. A splice variant reveals a mode of activation independent of ADP-ribose. J. Biol. Chem. 2002, 277, 23150–23156. [Google Scholar] [CrossRef]
- Hara, Y.; Wakamori, M.; Ishii, M.; Maeno, E.; Nishida, M.; Yoshida, T.; Yamada, H.; Shimizu, S.; Mori, E.; Kudoh, J.; et al. LTRPC2 Ca2+-permeable channel activated by changes in redox status confers susceptibility to cell death. Mol. Cell 2002, 9, 163–173. [Google Scholar] [CrossRef]
- Kolisek, M.; Beck, A.; Fleig, A.; Penner, R. Cyclic ADP-ribose and hydrogen peroxide synergize with ADP-ribose in the activation of TRPM2 channels. Mol. Cell 2005, 18, 61–69. [Google Scholar] [CrossRef]
- McHugh, D.; Flemming, R.; Xu, S.Z.; Perraud, A.L.; Beech, D.J. Critical intracellular Ca2+ dependence of transient receptor potential melastatin 2 (TRPM2) cation channel activation. J. Biol. Chem. 2003, 278, 11002–11006. [Google Scholar] [CrossRef]
- Starkus, J.; Beck, A.; Fleig, A.; Penner, R. Regulation of TRPM2 by extra- and intracellular calcium. J. Gen. Physiol. 2007, 130, 427–440. [Google Scholar] [CrossRef]
- Csanady, L.; Torocsik, B. Four Ca2+ ions activate TRPM2 channels by binding in deep crevices near the pore but intracellularly of the gate. J. Gen. Physiol. 2009, 133, 189–203. [Google Scholar] [CrossRef]
- Perraud, A.L.; Schmitz, C.; Scharenberg, A.M. TRPM2 Ca2+ permeable cation channels: From gene to biological function. Cell Calcium. 2003, 33, 519–531. [Google Scholar] [CrossRef]
- Sumoza-Toledo, A.; Penner, R. TRPM2: A multifunctional ion channel for calcium signalling. J. Physiol 2011, 589, 1515–1525. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, N.; Kozai, D.; Kobayashi, R.; Ebert, M.; Mori, Y. Roles of TRPM2 in oxidative stress. Cell Calcium. 2011, 50, 279–287. [Google Scholar] [CrossRef] [PubMed]
- Puertollano, R.; Kiselyov, K. TRPMLs: In sickness and in health. Am. J. Physiol Ren. Physiol. 2009, 296, F1245–F1254. [Google Scholar] [CrossRef]
- Qian, F.; Noben-Trauth, K. Cellular and molecular function of mucolipins (TRPML) and polycystin 2 (TRPP2). Pflug. Arch. 2005, 451, 277–285. [Google Scholar] [CrossRef]
- Dong, X.P.; Wang, X.; Xu, H. TRP channels of intracellular membranes. J. Neurochem. 2010, 113, 313–328. [Google Scholar] [CrossRef]
- Abe, K.; Puertollano, R. Role of TRP channels in the regulation of the endosomal pathway. Physiology 2011, 26, 14–22. [Google Scholar] [CrossRef]
- Grimm, C.; Hassan, S.; Wahl-Schott, C.; Biel, M. Role of TRPML and two-pore channels in endolysosomal cation homeostasis. J. Pharm. Exp. 2012, 342, 236–244. [Google Scholar] [CrossRef]
- Venkatachalam, K.; Wong, C.O.; Zhu, M.X. The role of TRPMLs in endolysosomal trafficking and function. Cell Calcium 2015, 58, 48–56. [Google Scholar] [CrossRef]
- Patel, S.; Cai, X. Evolution of acidic Ca(2)(+) stores and their resident Ca(2)(+)-permeable channels. Cell Calcium 2015, 57, 222–230. [Google Scholar] [CrossRef]
- Zhang, X.; Hu, M.; Yang, Y.; Xu, H. Organellar TRP channels. Nat. Struct. Mol. Biol. 2018, 25, 1009–1018. [Google Scholar] [CrossRef] [PubMed]
- Karacsonyi, C.; Miguel, A.S.; Puertollano, R. Mucolipin-2 localizes to the Arf6-associated pathway and regulates recycling of GPI-APs. Traffic 2007, 8, 1404–1414. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Hua, Y.; Vergarajauregui, S.; Diab, H.I.; Puertollano, R. Novel Role of TRPML2 in the Regulation of the Innate Immune Response. J. Immunol. 2015, 195, 4922–4932. [Google Scholar] [CrossRef] [PubMed]
- Lindvall, J.M.; Blomberg, K.E.; Valiaho, J.; Vargas, L.; Heinonen, J.E.; Berglof, A.; Mohamed, A.J.; Nore, B.F.; Vihinen, M.; Smith, C.I. Bruton’s tyrosine kinase: Cell biology, sequence conservation, mutation spectrum, siRNA modifications, and expression profiling. Immunol. Rev. 2005, 203, 200–215. [Google Scholar] [CrossRef]
- Miao, Y.; Li, G.; Zhang, X.; Xu, H.; Abraham, S.N. A TRP Channel Senses Lysosome Neutralization by Pathogens to Trigger Their Expulsion. Cell 2015, 161, 1306–1319. [Google Scholar] [CrossRef]
- Guo, Z.; Grimm, C.; Becker, L.; Ricci, A.J.; Heller, S. A novel ion channel formed by interaction of TRPML3 with TRPV5. PLoS ONE 2013, 8, e58174. [Google Scholar] [CrossRef]
- Di Palma, F.; Belyantseva, I.A.; Kim, H.J.; Vogt, T.F.; Kachar, B.; Noben-Trauth, K. Mutations in Mcoln3 associated with deafness and pigmentation defects in varitint-waddler (Va) mice. Proc. Natl. Acad. Sci. USA 2002, 99, 14994–14999. [Google Scholar] [CrossRef]
- Kim, H.J.; Soyombo, A.A.; Tjon-Kon-Sang, S.; So, I.; Muallem, S. The Ca(2+) channel TRPML3 regulates membrane trafficking and autophagy. Traffic 2009, 10, 1157–1167. [Google Scholar] [CrossRef]
- Medina, D.L.; Di Paola, S.; Peluso, I.; Armani, A.; De Stefani, D.; Venditti, R.; Montefusco, S.; Scotto-Rosato, A.; Prezioso, C.; Forrester, A.; et al. Lysosomal calcium signalling regulates autophagy through calcineurin and TFEB. Nat. Cell Biol. 2015, 17, 288–299. [Google Scholar] [CrossRef]
- Settembre, C.; Di Malta, C.; Polito, V.A.; Garcia Arencibia, M.; Vetrini, F.; Erdin, S.; Erdin, S.U.; Huynh, T.; Medina, D.; Colella, P.; et al. TFEB links autophagy to lysosomal biogenesis. Science 2011, 332, 1429–1433. [Google Scholar] [CrossRef]
- Scotto Rosato, A.; Montefusco, S.; Soldati, C.; Di Paola, S.; Capuozzo, A.; Monfregola, J.; Polishchuk, E.; Amabile, A.; Grimm, C.; Lombardo, A.; et al. TRPML1 links lysosomal calcium to autophagosome biogenesis through the activation of the CaMKKbeta/VPS34 pathway. Nat. Commun. 2019, 10, 5630. [Google Scholar] [CrossRef]
- Capurro, M.I.; Greenfield, L.K.; Prashar, A.; Xia, S.; Abdullah, M.; Wong, H.; Zhong, X.Z.; Bertaux-Skeirik, N.; Chakrabarti, J.; Siddiqui, I.; et al. VacA generates a protective intracellular reservoir for Helicobacter pylori that is eliminated by activation of the lysosomal calcium channel TRPML1. Nat. Microbiol. 2019, 4, 1411–1423. [Google Scholar] [CrossRef]
- Qi, X.; Man, S.M.; Malireddi, R.K.; Karki, R.; Lupfer, C.; Gurung, P.; Neale, G.; Guy, C.S.; Lamkanfi, M.; Kanneganti, T.D. Cathepsin B modulates lysosomal biogenesis and host defense against Francisella novicida infection. J. Exp. Med. 2016, 213, 2081–2097. [Google Scholar] [CrossRef] [PubMed]
- Cao, Q.; Yang, Y.; Zhong, X.Z.; Dong, X.P. The lysosomal Ca(2+) release channel TRPML1 regulates lysosome size by activating calmodulin. J. Biol. Chem. 2017, 292, 8424–8435. [Google Scholar] [CrossRef] [PubMed]
- Cang, C.; Aranda, K.; Seo, Y.J.; Gasnier, B.; Ren, D. TMEM175 Is an Organelle K(+) Channel Regulating Lysosomal Function. Cell 2015, 162, 1101–1112. [Google Scholar] [CrossRef]
- Jinn, S.; Drolet, R.E.; Cramer, P.E.; Wong, A.H.; Toolan, D.M.; Gretzula, C.A.; Voleti, B.; Vassileva, G.; Disa, J.; Tadin-Strapps, M.; et al. TMEM175 deficiency impairs lysosomal and mitochondrial function and increases alpha-synuclein aggregation. Proc. Natl. Acad. Sci. USA 2017, 114, 2389–2394. [Google Scholar] [CrossRef] [PubMed]
- Tugba Durlu-Kandilci, N.; Ruas, M.; Chuang, K.T.; Brading, A.; Parrington, J.; Galione, A. TPC2 proteins mediate nicotinic acid adenine dinucleotide phosphate (NAADP)- and agonist-evoked contractions of smooth muscle. J. Biol. Chem. 2010, 285, 24925–24932. [Google Scholar] [CrossRef] [PubMed]
- Zhu, M.X.; Evans, A.M.; Ma, J.; Parrington, J.; Galione, A. Two-pore channels for integrative Ca signaling. Commun. Integr. Biol. 2010, 3, 12–17. [Google Scholar] [CrossRef] [PubMed]
- Grimm, C.; Butz, E.; Chen, C.C.; Wahl-Schott, C.; Biel, M. From mucolipidosis type IV to Ebola: TRPML and two-pore channels at the crossroads of endo-lysosomal trafficking and disease. Cell Calcium. 2017, 67, 148–155. [Google Scholar] [CrossRef]
- Calcraft, P.J.; Ruas, M.; Pan, Z.; Cheng, X.; Arredouani, A.; Hao, X.; Tang, J.; Rietdorf, K.; Teboul, L.; Chuang, K.T.; et al. NAADP mobilizes calcium from acidic organelles through two-pore channels. Nature 2009, 459, 596–600. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, X.; Dong, X.P.; Samie, M.; Li, X.; Cheng, X.; Goschka, A.; Shen, D.; Zhou, Y.; Harlow, J.; et al. TPC proteins are phosphoinositide- activated sodium-selective ion channels in endosomes and lysosomes. Cell 2012, 151, 372–383. [Google Scholar] [CrossRef] [PubMed]
- Cang, C.; Zhou, Y.; Navarro, B.; Seo, Y.J.; Aranda, K.; Shi, L.; Battaglia-Hsu, S.; Nissim, I.; Clapham, D.E.; Ren, D. mTOR regulates lysosomal ATP-sensitive two-pore Na(+) channels to adapt to metabolic state. Cell 2013, 152, 778–790. [Google Scholar] [CrossRef] [PubMed]
- Ogunbayo, O.A.; Zhu, Y.; Shen, B.; Agbani, E.; Li, J.; Ma, J.; Zhu, M.X.; Evans, A.M. Organelle-specific subunit interactions of the vertebrate two-pore channel family. J. Biol. Chem. 2015, 290, 1086–1095. [Google Scholar] [CrossRef] [PubMed]
- Cang, C.; Aranda, K.; Ren, D. A non-inactivating high-voltage-activated two-pore Na(+) channel that supports ultra-long action potentials and membrane bistability. Nat. Commun. 2014, 5, 5015. [Google Scholar] [CrossRef] [PubMed]
- Ruas, M.; Davis, L.C.; Chen, C.C.; Morgan, A.J.; Chuang, K.T.; Walseth, T.F.; Grimm, C.; Garnham, C.; Powell, T.; Platt, N.; et al. Expression of Ca(2)(+)-permeable two-pore channels rescues NAADP signalling in TPC-deficient cells. EMBO J. 2015, 34, 1743–1758. [Google Scholar] [CrossRef] [PubMed]
- Patel, S. Function and dysfunction of two-pore channels. Sci. Signal. 2015, 8, re7. [Google Scholar] [CrossRef]
- Marchant, J.S.; Patel, S. Two-pore channels at the intersection of endolysosomal membrane traffic. Biochem Soc. Trans. 2015, 43, 434–441. [Google Scholar] [CrossRef]
- Grimm, C.; Chen, C.C.; Wahl-Schott, C.; Biel, M. Two-Pore Channels: Catalyzers of Endolysosomal Transport and Function. Front. Pharm. 2017, 8, 45. [Google Scholar] [CrossRef]
- Nguyen, O.N.; Grimm, C.; Schneider, L.S.; Chao, Y.K.; Atzberger, C.; Bartel, K.; Watermann, A.; Ulrich, M.; Mayr, D.; Wahl-Schott, C.; et al. Two-Pore Channel Function Is Crucial for the Migration of Invasive Cancer Cells. Cancer Res. 2017, 77, 1427–1438. [Google Scholar] [CrossRef]
- Garcia-Rua, V.; Feijoo-Bandin, S.; Rodriguez-Penas, D.; Mosquera-Leal, A.; Abu-Assi, E.; Beiras, A.; Maria Seoane, L.; Lear, P.; Parrington, J.; Portoles, M.; et al. Endolysosomal two-pore channels regulate autophagy in cardiomyocytes. J. Physiol. 2016, 594, 3061–3077. [Google Scholar] [CrossRef]
- Qureshi, O.S.; Paramasivam, A.; Yu, J.C.; Murrell-Lagnado, R.D. Regulation of P2X4 receptors by lysosomal targeting, glycan protection and exocytosis. J. Cell Sci. 2007, 120, 3838–3849. [Google Scholar] [CrossRef] [PubMed]
- Suurvali, J.; Boudinot, P.; Kanellopoulos, J.; Ruutel Boudinot, S. P2X4: A fast and sensitive purinergic receptor. Biomed. J. 2017, 40, 245–256. [Google Scholar] [CrossRef] [PubMed]
- Murrell-Lagnado, R.D.; Frick, M. P2X4 and lysosome fusion. Curr. Opin. Pharmacol. 2019, 47, 126–132. [Google Scholar] [CrossRef] [PubMed]
- Toyomitsu, E.; Tsuda, M.; Yamashita, T.; Tozaki-Saitoh, H.; Tanaka, Y.; Inoue, K. CCL2 promotes P2X4 receptor trafficking to the cell surface of microglia. Purinergic Signal. 2012, 8, 301–310. [Google Scholar] [CrossRef] [PubMed]
- Huang, P.; Zou, Y.; Zhong, X.Z.; Cao, Q.; Zhao, K.; Zhu, M.X.; Murrell-Lagnado, R.; Dong, X.P. P2X4 forms functional ATP-activated cation channels on lysosomal membranes regulated by luminal pH. J. Biol. Chem. 2014, 289, 17658–17667. [Google Scholar] [CrossRef]
- Gofman, L.; Cenna, J.M.; Potula, R. P2X4 receptor regulates alcohol-induced responses in microglia. J. Neuroimmune Pharm. 2014, 9, 668–678. [Google Scholar] [CrossRef]
- Moore, M.N.; Allen, J.I.; McVeigh, A.; Shaw, J. Lysosomal and autophagic reactions as predictive indicators of environmental impact in aquatic animals. Autophagy 2006, 2, 217–220. [Google Scholar] [CrossRef]
- Meng Lin, M.; Kim, H.H.; Kim, H.; Muhammed, M.; Kyung Kim, D. Iron oxide-based nanomagnets in nanomedicine: Fabrication and applications. Nano Rev. 2010, 1. [Google Scholar] [CrossRef]
- Zhu, J.; He, K.; Dai, Z.; Gong, L.; Zhou, T.; Liang, H.; Liu, J. Self-Assembly of Luminescent Gold Nanoparticles with Sensitive pH-Stimulated Structure Transformation and Emission Response toward Lysosome Escape and Intracellular Imaging. Anal. Chem. 2019, 91, 8237–8243. [Google Scholar] [CrossRef]
- Lee, D.U.; Park, J.Y.; Kwon, S.; Park, J.Y.; Kim, Y.H.; Khang, D.; Hong, J.H. Apoptotic lysosomal proton sponge effect in tumor tissue by cationic gold nanorods. Nanoscale 2019, 11, 19980–19993. [Google Scholar] [CrossRef]
- Bhowmik, T.; Gomes, A. NKCT1 (purified Naja kaouthia protein toxin) conjugated gold nanoparticles induced Akt/mTOR inactivation mediated autophagic and caspase 3 activated apoptotic cell death in leukemic cell. Toxicon 2016, 121, 86–97. [Google Scholar] [CrossRef] [PubMed]
- Totten, J.D.; Wongpinyochit, T.; Seib, F.P. Silk nanoparticles: Proof of lysosomotropic anticancer drug delivery at single-cell resolution. J. Drug Target. 2017, 25, 865–872. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.F.; Gurunathan, S. Combination of salinomycin and silver nanoparticles enhances apoptosis and autophagy in human ovarian cancer cells: An effective anticancer therapy. Int. J. Nanomed. 2016, 11, 3655–3675. [Google Scholar] [CrossRef]
- Martin-Banderas, L.; Holgado, M.A.; Duran-Lobato, M.; Infante, J.J.; Alvarez-Fuentes, J.; Fernandez-Arevalo, M. Role of Nanotechnology for Enzyme Replacement Therapy in Lysosomal Diseases. A Focus on Gaucher’s Disease. Curr. Med. Chem. 2016, 23, 929–952. [Google Scholar] [CrossRef] [PubMed]
- Muhlstein, A.; Gelperina, S.; Shipulo, E.; Maksimenko, O.; Kreuter, J. Arylsulfatase A bound to poly(butyl cyanoacrylate) nanoparticles for enzyme replacement therapy—Physicochemical evaluation. Pharmazie 2014, 69, 518–524. [Google Scholar] [PubMed]
- Tan, Y.L.; Ho, H.K. Navigating albumin-based nanoparticles through various drug delivery routes. Drug Discov. Today 2018, 23, 1108–1114. [Google Scholar] [CrossRef]
- Elzoghby, A.O.; Samy, W.M.; Elgindy, N.A. Albumin-based nanoparticles as potential controlled release drug delivery systems. J. Control. Release 2012, 157, 168–182. [Google Scholar] [CrossRef]
- An, F.F.; Zhang, X.H. Strategies for Preparing Albumin-based Nanoparticles for Multifunctional Bioimaging and Drug Delivery. Theranostics 2017, 7, 3667–3689. [Google Scholar] [CrossRef]
- Tang, W.L.; Tang, W.H.; Li, S.D. Cancer theranostic applications of lipid-based nanoparticles. Drug Discov. Today 2018, 23, 1159–1166. [Google Scholar] [CrossRef]
- Guven, E. Lipid-based nanoparticles in the treatment of erectile dysfunction. Int J. Impot Res. 2020. [Google Scholar] [CrossRef]
- Zhou, Y.; Fang, X.; Gong, Y.; Xiao, A.; Xie, Y.; Liu, L.; Cao, Y. The Interactions between ZnO Nanoparticles (NPs) and alpha-Linolenic Acid (LNA) Complexed to BSA Did Not Influence the Toxicity of ZnO NPs on HepG2 Cells. Nanomaterials 2017, 7, 91. [Google Scholar] [CrossRef]
- Peynshaert, K.; Soenen, S.J.; Manshian, B.B.; Doak, S.H.; Braeckmans, K.; De Smedt, S.C.; Remaut, K. Coating of Quantum Dots strongly defines their effect on lysosomal health and autophagy. Acta Biomater. 2017, 48, 195–205. [Google Scholar] [CrossRef] [PubMed]
- Pankiv, S.; Clausen, T.H.; Lamark, T.; Brech, A.; Bruun, J.A.; Outzen, H.; Overvatn, A.; Bjorkoy, G.; Johansen, T. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J. Biol. Chem. 2007, 282, 24131–24145. [Google Scholar] [CrossRef] [PubMed]
- Thurston, T.L.; Ryzhakov, G.; Bloor, S.; von Muhlinen, N.; Randow, F. The TBK1 adaptor and autophagy receptor NDP52 restricts the proliferation of ubiquitin-coated bacteria. Nat. Immunol. 2009, 10, 1215–1221. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.W.; Hong, S.B.; Kim, J.H.; Kwon, D.H.; Song, H.K. Structural basis for recognition of autophagic receptor NDP52 by the sugar receptor galectin-8. Nat. Commun. 2013, 4, 1613. [Google Scholar] [CrossRef] [PubMed]
- Wild, P.; Farhan, H.; McEwan, D.G.; Wagner, S.; Rogov, V.V.; Brady, N.R.; Richter, B.; Korac, J.; Waidmann, O.; Choudhary, C.; et al. Phosphorylation of the autophagy receptor optineurin restricts Salmonella growth. Science 2011, 333, 228–233. [Google Scholar] [CrossRef]
- Zeng, J.; Shirihai, O.S.; Grinstaff, M.W. Degradable Nanoparticles Restore Lysosomal pH and Autophagic Flux in Lipotoxic Pancreatic Beta Cells. Adv. Healthc. Mater. 2019, 8, e1801511. [Google Scholar] [CrossRef]
- Jimeno-Romero, A.; Oron, M.; Cajaraville, M.P.; Soto, M.; Marigomez, I. Nanoparticle size and combined toxicity of TiO2 and DSLS (surfactant) contribute to lysosomal responses in digestive cells of mussels exposed to TiO2 nanoparticles. Nanotoxicology 2016, 10, 1168–1176. [Google Scholar] [CrossRef]
| Channels | Mechanisms and Related Diseases | Ref. |
|---|---|---|
| CLC-3 | Promotion of lysosomal acidification | [72,73] |
| CLC-6 | LSD in CLC-6 mutated neuronal cells | [74] |
| CLC-7 | Maintenance of acidic pH of lysosomes | [75,76] |
| Decrease of dentinogenesis and dental bone formation in CLC-7 deficient mice | [77,78] | |
| Degradation of fAβ which drives AD | [79,80] | |
| Osteopetrosis in CLC-7 mutation | [81,82,83,84] | |
| LSD and neurodegeneration in CLC-7-deficient mice | [64,82] | |
| CFTR | Support lysosomal acidification | [85] |
| Decrease of bacteria killing function and phago-lysosomal fusion in macrophage | [86] | |
| TRPM2 | Induce DC maturation and migration | [87] |
| Increase of actin remodeling | [88] | |
| Increase of pancreatic β cell apoptosis | [89] | |
| Increase LMP, NLRP3 inflammasome, and mitochondrial fission on the plasma membrane | [90,91] | |
| TRPML1 | Maintenance of acidic pH of lysosomes | [92] |
| Increase of large particle phagocytosis, bone remodeling, gastric acid secretion, and myocytes apoptosis | [93,94,95,96] | |
| Stomach hypertrophy, hypergastrinemia, LSD, mucolipidosis, NPC, and AD in TRPML1 deficiency | [93,97,98,99,100,101,102,103] | |
| TMEM175 | Support lysosomal Ca2+ signaling and pH regulation | [104] |
| Related in LSD | [105] | |
| TPC | Related in autophagy, cancer cell migration, and cellular pigmentation | [106,107,108] |
| Related in Parkinson’s disease | [109,110] | |
| P2X4 | Promotion of endo-lysosomal fusion | [111,112] |
| Related in liver fibrogenesis | [113] |
| Related Cellular Function | NPs | Details | Reference |
|---|---|---|---|
| pH alteration (alkalization of lysosome) | AuNPs | Increase of oxidative stress, mitochondrial damage, and decrease cell migration/invasion | [11] |
| Accumulation of LC3 and block p62 degradation | [12] | ||
| AgNPs | Decrease of TFEB protein expression | [13] | |
| REONPs | Activation of IL-1β inflammasome | [14] | |
| Cell viability (cell death) | PNPs | Decrease of autophagic flux | [32] |
| Decrease of cathepsin release | [34] | ||
| SiO2 NPs | Increase of membrane damage and NLRP inflammasome | [35,44] | |
| TiO2 NPs | Increase of membrane damage | [36] | |
| Gd2O3 NPs | Increase of membrane damage and necrosis | [37] | |
| Protein activity and expression | AgNPs | Decrease of lysosomal protease activities | [41] |
| REONPs | Induce lysosomal imbalance by inhibiting mTORC1 pathway | [43] | |
| ZnO NPs | Increase of macrophage cell death by inhibiting mTORC1 pathway | [42] | |
| Deglycosylation of LAMP-2 | [44] | ||
| Carbon nanotube | Decrease of SNAP | [50] | |
| Accumulation of NPs | CuO NPs | Subsequent cellular damage leading to cell death by agglomeration of lysosomes | [55,56] |
| SiO2 NPs, PNPs | Induce lysosomal swelling leading to apoptosis | [57,58] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, D.; Hong, J.H. Nanoparticle-Mediated Therapeutic Application for Modulation of Lysosomal Ion Channels and Functions. Pharmaceutics 2020, 12, 217. https://doi.org/10.3390/pharmaceutics12030217
Lee D, Hong JH. Nanoparticle-Mediated Therapeutic Application for Modulation of Lysosomal Ion Channels and Functions. Pharmaceutics. 2020; 12(3):217. https://doi.org/10.3390/pharmaceutics12030217
Chicago/Turabian StyleLee, Dongun, and Jeong Hee Hong. 2020. "Nanoparticle-Mediated Therapeutic Application for Modulation of Lysosomal Ion Channels and Functions" Pharmaceutics 12, no. 3: 217. https://doi.org/10.3390/pharmaceutics12030217
APA StyleLee, D., & Hong, J. H. (2020). Nanoparticle-Mediated Therapeutic Application for Modulation of Lysosomal Ion Channels and Functions. Pharmaceutics, 12(3), 217. https://doi.org/10.3390/pharmaceutics12030217






