Abstract
In the past few years, there has been increasing focus on the use of messenger RNA (mRNA) as a new therapeutic modality. Current clinical efforts encompassing mRNA-based drugs are directed toward infectious disease vaccines, cancer immunotherapies, therapeutic protein replacement therapies, and treatment of genetic diseases. However, challenges that impede the successful translation of these molecules into drugs are that (i) mRNA is a very large molecule, (ii) it is intrinsically unstable and prone to degradation by nucleases, and (iii) it activates the immune system. Although some of these challenges have been partially solved by means of chemical modification of the mRNA, intracellular delivery of mRNA still represents a major hurdle. The clinical translation of mRNA-based therapeutics requires delivery technologies that can ensure stabilization of mRNA under physiological conditions. Here, we (i) review opportunities and challenges in the delivery of mRNA-based therapeutics with a focus on non-viral delivery systems, (ii) present the clinical status of mRNA vaccines, and (iii) highlight perspectives on the future of this promising new type of medicine.
Keywords:
mRNA; vaccines; therapeutic; prophylactic; drug delivery systems; lipids; polymers; nanoparticles; nanomedicine 1. Introduction
Vaccination has had a tremendous impact on global health and the quality of human life by reducing the mortality and morbidity caused by infectious diseases. The development of vaccines is predicated on the classical 3I’s paradigm of “isolating, inactivating and injecting” the causative microorganism, coined by Louis Pasteur [1]. Vaccines can be prophylactic or therapeutic and can broadly be classified as live attenuated vaccines (weakened microorganisms), inactivated vaccines (killed microorganisms), subunit vaccines (purified antigens), or toxoid vaccines (inactivated bacterial toxins). As opposed to the conventional concept of injecting live-attenuated or inactivated pathogens, modern vaccine approaches, i.e., subunit vaccines, focus on exhibiting efficacy similar to conventional vaccines while obviating the safety risks associated with whole-cell vaccines. However, subunit antigens often display lower immunogenicity, which can be rectified by employing delivery systems and/or immunopotentiating compounds as adjuvants to boost immunogenicity. The modern genome-based rational vaccine design offers tremendous potential over conventional whole-organism-based vaccine approaches.
Nucleic acid-based vaccines, i.e., DNA (as plasmids) and RNA (as messenger RNA (mRNA)) vaccines, pave the way for safe and efficacious biologics to mimic inoculation with live organism-based vaccines, particularly for stimulation of cell-mediated immunity [2]. This technology exhibits promising potential for the development of novel vaccines against a wide variety of indications and diseases, extending from prophylactics to therapeutics for infectious diseases, cancer, autoimmune diseases, and hypersensitivities. While nucleic-acid-based vaccines demonstrate significant advantages over traditional vaccines [3] in terms of safety, efficacy, induction of both B- and T-cell responses and specificity, it is noteworthy to mention that mRNA vaccines have advantages when compared to vaccines based on other types of nucleic acids. A technical challenge associated with DNA vaccines is to ensure delivery into the cell nucleus, where antigen transcription takes place prior to nuclear export and translation into protein in the cytoplasm. In addition, DNA vaccines carry a potential risk of integration into the host genome, which may result in insertional mutagenesis. In contrast, mRNA vaccines are only targeted for cytoplasmic delivery, circumventing the risk of genomic integration [4]. The relatively short half-life results in transient and more controlled expression of the encoded antigen. Moreover, mRNA can be produced in a cell-free environment by in vitro transcription (IVT), thereby eschewing the use of microbes or cultured cells for production, and avoiding the associated quality and safety issues in the production. This permits simple downstream purification and rapid and cost-effective manufacturing [5]. However, mRNA is often promulgated on the grounds of the popular opinion that when using mRNA, unlike DNA, the stringent gene-therapy regulations are bypassed because mRNA does not integrate into the host genome. However, in reality, this only holds true in the US since in Europe, any active pharmaceutical ingredient, which contains or consists of a recombinant nucleic acid, used in or administered to human beings, falls under the scope of the regulation for advanced therapy medicinal products [6]. Therefore, mRNA-based therapeutics are categorized as gene therapy. The burgeoning field of mRNA vaccines is very exciting [3,7] and considerable amounts of relevant preclinical data have been generated, and several clinical trials have been initiated during the last decade. This gives rise to the vision of translating the mRNA vaccines into human application for prophylaxis and therapy. In this review, we discuss the current trends in mRNA vaccine approaches and various strategies and systems for delivering mRNA vaccines.
2. mRNA as Vaccines
The central dogma of molecular biology states that DNA is transcribed into mRNA, which is subsequently translated into protein [8]. The flow of genetic information in time and space is orchestrated by complex regulatory mechanisms. Gene therapy represents the introduction of genetic material into an individual’s cells and biological tissues. Techniques like insertion, alteration, or removal of genes are employed for correcting defective genes responsible for disease development, which then cures a disease or ameliorates the clinical status of a patient [9]. Several vectors have been utilized for gene therapy and they are generally classified as non-viral and viral vectors. Non-viral vectors possess several advantages compared to the viral vectors, including low host immunogenicity and potential for scale-up [10]. However, the success of non-viral gene therapy has been very limited in the past, primarily due to the barriers existing for plasmid DNA (pDNA) delivery, e.g., the necessity to cross the nuclear membrane before translation, the presence of antibiotic resistance genes in pDNA, and most importantly, the difficulty in controlling and regulating long-term expression. The lack of control of long-term expression of pDNA poses a huge disadvantage in terms of the duration of treatment and possible side effects, which is in contrast to conventional drugs, where the treatment can be stopped instantaneously. These disadvantages of pDNA can possibly be overcome by using mRNA [11]. The mRNA carries genetic information from the DNA in the nucleus to the cytosol, where it is used by the ribosomes as a template for protein synthesis. As opposed to pDNA, the mRNA is efficacious in both mitotic and non-mitotic cells because mRNA exerts its function in the cytoplasm, hence its function is not dependent on active cell division. Furthermore, unlike pDNA or viral vectors, mRNA does not contain additional foreign genes, which makes mRNA a safer vector. The challenge of long-term expression posed by pDNA can also be overcome by using mRNA, since mRNA mediates a rapid, transient expression of the encoded protein and the duration of the production is well-defined (usually a few days or weeks, depending on the specific mRNA platform). This makes mRNA expression easier to control than the gene expression from pDNA and viral vectors [11]. In addition, the manufacturing of mRNA is cell-free, which strongly reduces the chance of mRNA contamination with bacterial components. This makes it easier to produce mRNA than pDNA under good manufacturing practice conditions [12]. Finally, vector-induced immunogenicity can be avoided for mRNA therapeutics, unlike for viral vectors or virus-like particles, which may elicit a specific immune response against the exposed viral proteins [13]. Specific therapeutic applications of mRNA, which are currently being explored include (i) vaccination against cancer and infectious diseases, (ii) protein-replacement therapy, and (iii) gene editing. Table 1 summarizes examples of ongoing clinical trials of mRNA-based therapeutic and prophylactic vaccine candidates.

Table 1.
Examples of ongoing clinical trials of mRNA-based vaccines.
Two classes of mRNAs, i.e., non-replicating and self-amplifying mRNA, are commonly used as vaccine vectors. Non-replicating mRNA encodes only the protein antigen(s) of interest, while self-amplifying mRNA also encodes proteins enabling RNA replication [14]. Vaccines based on self-amplifying mRNA encode the RNA genome of a single-stranded RNA virus, e.g., an alphavirus, a flavivirus [15], or a picornavirus [7]. They are engineered to increase the duration and level of expression, as well as the subsequent immune response induced by the encoded antigen(s). They efficiently amplify the production of sub-genomic mRNA encoding antigen(s) of interest subsequent to a single round of replication. While both self-amplifying mRNA and non-replicating mRNA find application in prophylactic vaccines for infectious diseases, non-replicating mRNA is used for cancer vaccines.
3. Fundamental Pharmacology of mRNA Vaccines
In vitro transcribed (IVT) mRNA is employed therapeutically as it mimics fully mature native mRNA present in the eukaryotic cytosol [16]. This may be achieved either by ex vivo transfection of cells with mRNA that are then adoptively transferred or by direct in vivo delivery of the IVT mRNA to the cytosol [17]. These approaches are explored for genome engineering, genetic reprogramming, adoptive T cell and dendritic cell (DC) based cancer and infectious disease immunotherapies, tolerization regimens to treat allergies, and protein replacement therapies. Both ex vivo transfection and direct in vivo transfection enable the target cells to synthesize the encoded protein(s) in situ, where mRNA is used as a template and the protein(s) represents the active product. The open reading frame (ORF) of mature mRNA encoding the protein(s) of interest (the active product) marked by start and stop codons, respectively, is flanked by untranslated regions (UTRs), and ideally consists of a 5’ cap and a poly(A) tail [3].
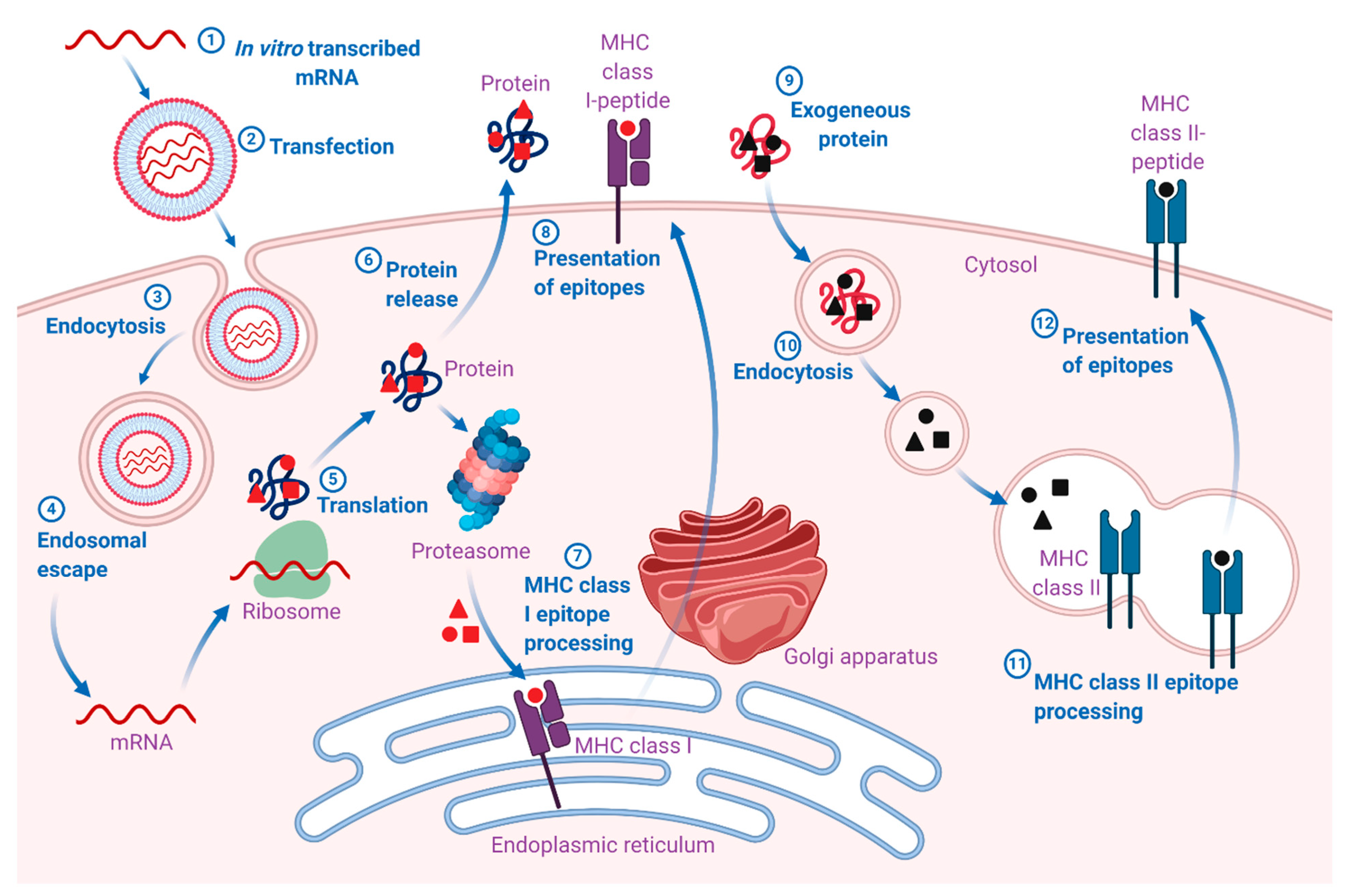
The pharmacodynamic activity of both native and IVT mRNA takes place in the cytosol (Figure 1). However, in contrast to endogenous mRNA, which is transcribed from DNA in the nucleus and enters the cytosol through nuclear export, IVT mRNA enters the cytosol from an extracellular source [18]. Once the IVT mRNA is delivered to the cytosol, its pharmacology is governed by the same complex cellular mechanisms that regulate the stability and translation of endogenous mRNA. The engineered IVT mRNA resembles the endogenous mRNA so closely that the cellular translation machinery is seamlessly utilized to synthesize a protein that may undergo post-translational modifications, eventually resulting in mature protein product(s). In the case of vaccines, this mature protein product(s) represents the antigen(s), which may elicit potent pathogen-specific humoral and cell-mediated immune responses. However, the final intracellular destination is determined by the natural or engineered sequence(s) of the signal peptide or the transmembrane domain [7]. Therefore, mRNA vaccines can be designed for the delivery of the encoded protein(s) to the desired cellular compartment for proper presentation and/or function [19].
Figure 1.
Mechanism of action of mRNA vaccines. 1. The mRNA is in vitro transcribed (IVT) from a DNA template in a cell-free system. 2. IVT mRNA is subsequently transfected into dendritic cells (DCs) via (3) endocytosis. 4. Entrapped mRNA undergoes endosomal escape and is released into the cytosol. 5. Using the translational machinery of host cells (ribosomes), the mRNA is translated into antigenic proteins. The translated antigenic protein undergoes post-translational modification and can act in the cell where it is generated. 6. Alternatively, the protein is secreted from the host cell. 7. Antigen protein is degraded by the proteasome in the cytoplasm. The generated antigenic peptide epitopes are transported into the endoplasmic reticulum and loaded onto major histocompatibility complex (MHC) class I molecules (MHC I). 8. The loaded MHC I-peptide epitope complexes are presented on the surface of cells, eventually leading to the induction of antigen-specific CD8+ T cell responses after T-cell receptor recognition and appropriate co-stimulation. 9. Exogenous proteins are taken up DCs. 10. They are degraded in endosomes and presented via the MHC II pathway. Moreover, to obtain cognate T-cell help in antigen-presenting cells, the protein should be routed through the MHC II pathway. 11. The generated antigenic peptide epitopes are subsequently loaded onto MHC II molecules. 12. The loaded MHC II-peptide epitope complexes are presented on the surface of cells, leading to the induction of the antigen-specific CD4+ T cell responses. Exogenous antigens can also be processed and loaded onto MHC class I molecules via a mechanism known as cross-presentation (not shown in the figure). The figure was created with BioRender.com.
The pharmacokinetics of IVT mRNA is determined by the half-life of the mRNA and the resulting mature protein after post-translational modification. The two major factors influencing the bioavailability of exogenous mRNA in the cytosol are (i) rapid RNase-mediated degradation and (ii) lack of passive diffusion across the plasma membrane owing to high molecular weight and electrostatic repulsion between the negative charges of the proteoglycan-coated cell membrane and the negatively-charged mRNA molecules [20]. Naked mRNA is rapidly degraded by extracellular RNases, thus hindering its efficient delivery and efficacy. A wide range of in vitro and in vivo transfection reagents have been shown to protect the mRNA against degradation and facilitate the cellular uptake and endosomal escape of mRNA. Major efforts have been dedicated to improving the enzymatic RNA stability, as discussed further below [3]. Ultimately, the IVT mRNA composed of natural nucleotides is metabolized by inherent physiological mechanisms, hence reducing the risk of metabolite-induced toxicity. Therefore, the delivery issues can be overcome by approaches including encapsulation of mRNA in drug delivery systems consisting of cationic molecules, lipids, polymers, and nanoparticles [21] as well as targeting DCs [22]. In addition, physical transfection methods like electroporation have been shown to enhance the delivery efficiency of large, self-amplifying mRNA in vivo, upon measuring reporter gene expression and immunogenicity of genes encoding HIV envelope proteins [23].
4. Approaches for Enhancing mRNA Stability
The development of mRNA-based drugs dates back to 1990 with the successful expression of a number of different proteins upon injecting mRNA encoding these proteins directly into the muscles of mice [24]. This led to (i) the testing of the first mRNA-based vaccine in 1993, which was shown to induce an anti-influenza cytotoxic T-lymphocyte response in mice [25], and (ii) the first vaccination with mRNA-encoding cancer antigens in 1995 [26]. These inceptive demonstrations ratified the potential of mRNA for (i) in situ expression of specific proteins and (ii) for induction of protective antigen-specific cellular and humoral immunity. However, the field was neglected for almost ten years until the potential of in vivo application of mRNA, i.e., induction of specific cytotoxic T lymphocytes and antibodies, was discovered [27]. Advances in the mRNA field were slow due to the labile nature of mRNA, which makes experiments employing unmodified mRNA very challenging unless precautions to handle mRNA are strictly adhered to [28]. Instead, the focus was directed towards DNA-based drugs, since DNA is more stable than RNA.
In a cell-free system, mRNA can be synthesized by IVT of a DNA template (e.g., a linearized plasmid or a PCR product), which encodes all the structural elements of a functional mRNA. To perform an IVT reaction, all the elements of the natural transcription process are required, i.e., a DNA template, an RNA polymerase, and nucleotide building blocks. During the subsequent purification of the mRNA, the DNA template is often degraded by the addition of DNases, followed by purification by means of other conventional methods for isolating mRNA, e.g., precipitation and chromatography. This process results in highly pure mRNA products ready for use [29,30,31]. Several strategies have been pursued to cope with mRNA’s inherent lack of stability and potential immunogenicity, which are discussed further below.
4.1. Molecular Stabilization
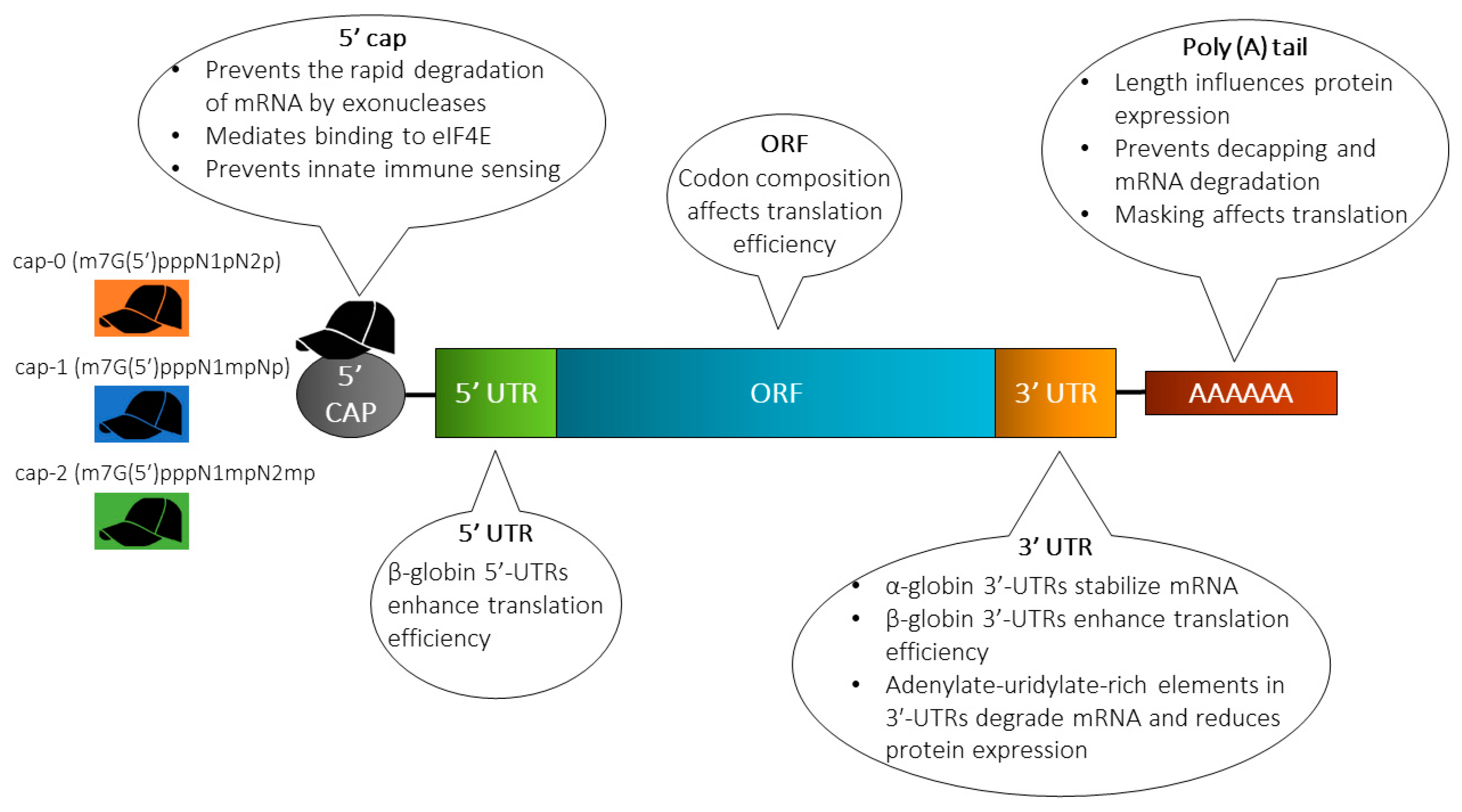
Strategies including engineering of sequences and/or structure to enhance mRNA stability (extend the half-life) and translation are often instrumental in increasing the protein expression levels. Techniques employed to achieve this includes elongation of the poly(A) tail, modification of the 5’ cap, engineering of the UTRs and the sequence patterns in the ORF, and/or incorporation of modified nucleotides (Figure 2).
Figure 2.
Structure of in vitro transcribed (IVT) mRNA and commonly used modification strategies. The design of IVT mRNA is based on the blueprint of eukaryotic mRNA, and it consists of a 5’ cap, 5’ and 3’ untranslated regions (UTRs), an open reading frame (ORF) encoding antigen(s), and a 3’ poly(A) tail. The IVT mRNA can be modified in one or multiple sites, e.g., by modification of the caps, the UTRs and/or the poly(A) tail, to modulate the duration and kinetic profile of protein expression. eIF4E, eukaryotic translation initiation factor 4E.
4.1.1. Cap Analog
A synthetic cap analogue can readily be added to the mRNA because the 5’ end cap is not encoded by the DNA template. Natural eukaryotic mRNA has a 7-methylguanosine (m7G) cap coupled to the mRNA during the transcription process via a 5’-5’-triphosphate bridge (ppp) [32]. The m7GpppN structure at the 5’ end of the mRNA cap serves several functions [33]. First, it protects the mRNA from rapid degradation by exonucleases. Second, it plays an indispensable role during translation because the eukaryotic initiation factor (eIF) 4E recognizes and binds to the cap of the mRNA. It further plays a role in preventing innate immune sensors from recognizing the mRNA [34]. The mRNA may contain one of three distinct caps, i.e., cap-0 [m7G(5’)pppN1pN2p], cap-1 [m7G(5’)pppN1mpNp] and cap-2 [m7G(5’)pppN1mpN2mp], respectively [11]. Capping of IVT mRNA can be performed using two different approaches: The first approach includes the addition of a second step with recombinant vaccinia virus-derived capping enzymes after the transcription, resulting in a cap identical to the most frequent endogenous eukaryotic cap structure i.e., 7-methylguanosine (m7G) cap [30]. Alternatively, a synthetic cap analogue may be added during the in vitro transcription reaction, hence capping and in vitro transcription is performed in a single step. This approach is referred to as co-transcriptional capping [35]. A major drawback of this approach is the competition between the cap analog and the GTP nucleotide required for in vitro transcription, which ultimately results in a fraction of uncapped and translationally inactive mRNA [36]. The fraction of uncapped mRNA containing 5’ ppp groups is more immune-stimulating, which can be rectified by treatment with phosphatase to remove the ppp group at the 5’ end of uncapped mRNA [37].
Three different classes of m7GpppG cap analogs are used [38]: (i) anti-reverse cap analogs (ARCAs) [39], (ii) 3’-O-Me-m7GpppG [40], and (iii) modified ARCAs [41]. Initial mRNA research was performed using mRNA containing the m7 cap analog (GpppG) [42], and it is currently the most commonly used mRNA cap in clinical trials. Unfortunately, a fraction of the m7GpppG cap analog used during in vitro transcription becomes incorporated in the opposite orientation and is therefore not recognized by the ribosomes, eventually resulting in lower translational activity. To avoid this, the so-called ARCA with only one 3’-OH group instead of two 3’-OH groups (ARCAs; m27,3’−OGpppG) has been introduced to prevent incorporation in the opposite orientation [43]. ARCAs juxtaposing the traditional cap analogues have been shown to exhibit more than four times RNA transcription efficiency [44]. In addition, the duration and the levels of protein expression have been found to be enhanced in cells transfected with ARCA-capped IVT RNA [41]. Recently, new types of chemically-modified cap analogues have been introduced, e.g., of phosphorothioate, phosphorothiolate [35], imidiphosphate [45], locked nucleic acid [46], boranophosphate bonds [47] and other types of modifications, which provide the mRNA with resistance to decapping by the mRNA-decapping enzyme 2, eventually resulting in a longer half-life of the mRNA [48].
Conventionally, the synthetic 5’-Cap 0-capped RNA is transcribed by performing an in vitro transcription, where more than 80% of the GTP added to the reaction is substituted with a dinucleotide cap analog (i.e., m7G[5’]ppp[5’]G), resulting in initiation of transcription with the cap analog [49]. However, this approach has been shown to exhibit a number of efficiency-related drawbacks, which have been overcome by the introduction of ScriptCap™. ScriptCap™ involves the addition of enzymatically built cap-0 structures onto the RNA transcripts by employing a capping enzyme with 100% reaction efficiency [50]. Native eukaryotic mRNA may contain a cap-1 or cap-2 but never a cap-0 [11]. Therefore, IVT mRNA should contain a cap-1 or a cap-2 in order to be less immune-stimulating [51]. A hallmark of these cap structures is the methylation status of the 2’ position of the 5’ second last and third last nucleoside. Prior to the advent of the novel technology of CleanCap™ introduced by TriLink, synthetic mRNA with a cap-1 could only be prepared by enzymatic capping [52,53]. However, cap-1 or cap-2 can now be incorporated during co-transcriptional capping at a capping efficiency of approximately 94% [52]. Notably, the capping efficiency has been shown to be significantly higher than the efficiency achieved by traditional co-transcriptional capping with cap-0 or ARCA [54].
4.1.2. 5’ and 3’ Untranslated Regions (UTRs)
The importance of incorporation of 5’- and 3’-UTRs has been noted during in vitro post-transcriptional regulation of gene expression [55]. The numerous roles that UTRs play include (i) regulation of mRNA export from the nucleus, (ii) regulation of translation efficiency [56], (iii) orchestration of subcellular localization [57], and (iv) mRNA stability [58]. Introduction of α-globin 3’ end UTRs results in stabilization of mRNA, while the incorporation of beta-globin 5’ end and 3’ end UTRs leads to enhanced translational efficiency [59]. The optimal outcome is achieved by using two β-globin 3’-UTRs aligned in a head-to-tail configuration. α-globin and β-globin UTRs have been incorporated for tweaking the RNA for optimized in vitro transcription followed by mRNA electroporation of autologous T cells [60] and intranodal injection of naked antigen-encoding RNA [61]. Moreover, DCs transfected with antigen-encoding UTR-optimized mRNA have been used in a study involving immunization of cytomegalovirus-seropositive individuals and cancer patients [62]. In some situations, destabilizing the mRNA might be a viable approach to reduce the duration of protein synthesis. This may be accomplished by introducing adenylate-uridylate-rich elements in the 3’-UTRs of the mRNA, eventually resulting in faster mRNA degradation and shortening of the duration of protein expression [63].
4.1.3. Poly(A) Tail
The poly(A) tail plays a significant role in mRNA translation as well as for the enzymatic stability of mRNA. The poly(A) tail binds to several polyadenosyl binding proteins (PABPs) while working synergistically with 5’m7Gcap sequences to regulate translational efficiency [64]. Eukaryotic translation initiation factor eIF4E binds to the 5’m7G cap, which in turn complexes with eIF4G and eIF4A. PABP then interacts with the N-terminus of the eukaryotic translation initiation factor eIF4G, which forms an mRNP (messenger ribonucleoprotein) or a polysome complex [65]. The former depicts the mRNA-protein complex not yet involved in protein synthesis, while the latter is one that is already being translated. An adequately long poly(A) tail is required to circularize the mRNA via binding of PABPs to the poly(A) tail and the cap [55,66]. It has been observed that increasing the poly(A) tail length improves the efficiency of polysome generation and consequently influences the protein expression levels [67].
It has been shown that a gradual increase in the poly(A) tail length of IVT mRNA to 120 bases commensurately increases the protein expression level, while an increase in the number of bases beyond 120 does not further enhance protein expression [68]. Poly(A) tails can be added to mRNA by encoding the poly(A) tail in the DNA template, or by extension of the IVT RNA after transcription using recombinant poly(A) polymerase. However, polyadenylation with recombinant poly(A) polymerase results in variable poly(A) tail length, thereby yielding polyadenylated mRNA with varying lengths. Therefore, the preferred approach is the generation of poly(A) tails with well-defined length from the mRNAs transcribed from poly(A) tail-encoding DNA templates [69]. The physical interactions between the 5’ and 3’ ends of mRNA take place between the cap and the poly(A) tail [70]. The poly(A) tail also plays a role in preventing decapping and mRNA degradation because removal or shortening of the poly(A) tail to less than 12 residues results in degradation of the mRNA through cleavage of the 5’ cap structure and 5’ to 3’ exonucleolytic digestion or 3’ to 5’ degradation [71].
4.2. Formulation Strategies
Despite the promising potential of mRNA-based vaccines, efficient intracellular delivery of mRNA to the cytosol continues to pose a major hurdle, especially for mRNA administered systemically. The large molecular weight (105–106 Da) [21] and high negative charge density of mRNA impair the permeation of mRNA across cellular membranes. It is well known that the absorption of mRNA in the absence of a delivery system is extremely low, and the half-life of mRNA is approximately 7 h [72]. Moreover, mRNA is an inherently unstable molecule, which is highly prone to degradation by 5’ exonucleases, 3’ exonucleases, and endonucleases [73]. Consequently, delivery systems are imperative for intracellular delivery of mRNA to the therapeutic site of action in vitro as well as in vivo [8,74]. Different strategies have been investigated to improve RNA delivery, including improved injection strategies such as microinjections [75], RNA patches [76], gene gun-based administration [77], protamine condensation [78], RNA adjuvants [79], and encapsulation of RNA in nanoparticles consisting of lipids and/or polymers [80]. Generally, IVT mRNA for cytosolic delivery is formulated with a delivery system by mixing with a complexing agent [81], which can protect the mRNA against rapid degradation and facilitate cellular uptake. Although the general dogma in the field is that efficient carriers are needed for substantially enhancing the in vivo transfection of mRNA, naked mRNA have been applied in many in vivo studies. Hence, the following section discusses the delivery of naked mRNA, followed by sections discussing vector-based mRNA delivery [82,83,84,85].
4.2.1. Naked RNA
The simplest administration strategy comprises intramuscular (i.m.) injection of naked mRNA, and proof of concept was originally demonstrated by in vivo reporter gene expression in mice [24]. Since then, the effectiveness of naked mRNA has been confirmed upon i.m. [86], subcutaneous (s.c.) [87] or intradermal (i.d.) [88] injections. I.d. and s.c. administration of mRNA has been shown to mediate healing of various skin diseases and to ameliorate wound healing by in situ expression of specific proteins in the skin [75,89]. Adopting this approach circumvents several obstacles otherwise associated with systemic administration of mRNA, e.g., clearance from the bloodstream via the liver, the kidneys, and the spleen [75]. Very efficient translation of the encoded protein has been shown for mRNA administered s.c. [90,91,92]. Interestingly, more efficient translation has at times been measured when compared to mRNA-loaded nanoparticle-based delivery systems [90,93]. Hence, it circumvents the need to employ carriers, eventually contributing to reduced cost and potential risk. Another advantage of administering mRNA-based vaccines via the s.c. route is that both cellular and humoral immune responses are induced [94] because the mRNA is expressed by both skin-resident DCs [95] and non-immune cells [96]. However, the outermost stratum corneum layer of the epidermis serves as a tight barrier to the absorption of topically administered drugs [97]. Heretofore, various approaches have been adopted to overcome this barrier, including physical (e.g., microporation [98], microneedles [99], and jet injection [100,101]), active (e.g., electroporation [102], iontophoresis [103], and sonophoresis [104]), and passive methods (e.g., nanoparticles [105] and liposomes [106]). Electroporation and sonoporation can transiently permeabilize the skin by means of electric pulses and low-frequency ultrasound, respectively, for efficient delivery of genes into the skin [107]. Modified mRNA-encoding vascular endothelial growth factor-A was formulated in citrate-buffered saline without the use of a delivery system and was used for i.d. vaccination of patients with type 2 diabetes [108]. However, in spite of pronounced, sustained, dose-dependent and cargo-specific vasodilation, blood flow increase, oxygen-metabolic upregulation, angiogenesis and neovessel formation in animal models, the vasodilatory and angiogenic activity did not translate into humans [108,109]. Microneedle-based delivery is also an efficient technique where micron-sized needle patch/arrays composed of water-soluble polymeric or sugar excipients are employed, into which the mRNA is incorporated [76]. The patches/arrays provide mechanical strength needed for the needle to permeate the stratum corneum and penetrate into the viable skin layers. Following injection, depending upon the type of microneedles, the patches/arrays degrade/dissolve and the encapsulated drug is released in the interstitial fluid of the skin [75]. Administration of mRNA by using dissolvable microneedles provides an important advantage, i.e., delivery in a solid dosage form circumvents the necessity of dealing with mRNA in a liquid dosage form [110], which therefore eliminates the menace emanating from RNase contamination, along with increasing mRNA stability, and shelf life [76]. Scavenger receptor-mediated endocytosis and micropinocytosis have been shown to be the active uptake mechanisms for naked mRNA in immature DCs [16]. However, naked mRNA displays a short plasma half-life, is prone to ribonuclease degradation, and faces difficulties in entering the cell. Therefore, delivery systems have been propounded to protect the mRNA and shield its negative charge.
4.2.2. Viral Vectors
Delivery of mRNA can be mediated by viral and non-viral vectors. Non-viral vectors can further be categorized into lipid-based delivery systems, polymer-based delivery systems, and lipid-polymer hybrid systems [111]. For viral RNA delivery, there has been a great deal of interest in the engineering of adeno-associated viruses to carry nucleic acid cargoes [112]. Genetically-modified viruses are usually employed for mRNA/gene delivery. The genes of these viruses are partially or fully substituted with model or therapeutic genes. A benefit of RNA viruses is that they are replicated and expressed locally in the cytoplasm. Positive strand RNA viruses are distinguished by a genomic sequence that can be translated directly into proteins of interest by host ribosomes. Notably, alphaviruses (e.g., Sindbis and Semliki Forest virus) [113], picornaviruses [114], and flavivirus [115] (e.g., Kunjin virus) have been employed for mRNA delivery. Various alphavirus vectors can be used to express high levels of exogenous protein in a wide spectrum of hosts [116]. Commonly used approaches include the direct substitution of structural genes with heterologous expression or placing the nonstructural genes downstream of the RNA sub-genomic promoter [117]. However, alphaviruses induce severe cytopathogenic effects, which restrict their application in gene therapy, although different strategies can be employed to surmount this challenge [118]. Some of these strategies include engineering of mutant vectors with mitigated cytotoxicity and temperature-inducibility, and self-inactivating vectors with point mutations in the nsP2 gene (especially at position 726, 259 and 650) [119]. Sendai virus (SeV) (murine parainfluenza virus type 1 or hemagglutinating virus of Japan) that belongs to the Paramyxoviridae family is worth mentioning owing to its popular application as a vector. It is favored for its high but transient gene expression levels, wide host cell specificity, low pathogenicity, and strong immunogenicity [120]. As a vaccine platform, the Venezuelan encephalitis virus is of particular interest [121]. These vaccines encompass the live-attenuated viral vaccine TC-83 and a formalin-inactivated variety of it is referred to as C-84, which boosts the efficacy and increases the duration of immunity upon administering the TC-83 vaccine via different administration routes, intranasal being the most widely used [122]. However, using viral vectors embodies crucial drawbacks associated with genome integration, and possible host rejection (immunogenicity and cytotoxicity) among others [123], hence provoking the need for non-viral vectors for mRNA delivery [10].
4.2.3. Polymer-Based Vectors
Diethylaminoethyl (DEAE) dextran was the first polymer to be tested as a delivery reagent for IVT mRNA [124]. Later, it was shown that lipid-mediated mRNA transfection is 100 to 1000 times more efficient than DEAE-dextran [125]. This discovery stalled the progress of polymeric carriers and paved the way for lipid-based transfection reagents for nucleic acids, including mRNA. A comprehensive study compared the polymers poly-beta-amino-esters (PBAE) and polyethylenimine (PEI) with commercial transfection reagent Lipofectamine™ 2000 and 1,2-dioleoyl-3-trimethylammonium propane (DOTAP)/ 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE) for functional, antigen-specific T-cell responses after mRNA delivery [126]. All carriers were complexed with mRNA encoding the HIV-1 antigen gag. Gag-specific, IFN-γ secreting T cells were measured in the spleen and lymph nodes of mice immunized with gag mRNA complexed with cationic lipids but not in mice immunized with naked and polymer-complexed mRNA. PEI and its derivatives are among the most commonly employed cationic polymers [127]. They are water-soluble, display a high density of positive charge associated with the amino groups, and are proven mRNA carriers for in vitro transfection [128]. However, PEI displays toxicity issues owing to the high molecular weight (>25 kDa), which may arise from the adsorption of anionic serum proteins onto the polyplex surface between cationic polymers and anionic serum plasma proteins. However, the resultant increase in size is only transient as the proteins adsorbed on the surface of polyplexes prevent particle–particle aggregation in the long run [129]. Various efforts have been made to mitigate these challenges. The first proof of concept for safe and efficacious mRNA vaccine transfection employing cationic polymer was obtained by intranasal administration of 2 kDa PEI conjugated to cyclodextrin. Cyclodextrin conjugated to PEI enabled delocalization of the charge density on the polyamine backbone, hence reducing cytotoxicity and at the same time maintaining protonatable groups, resulting in improved transfection [130]. Polymeric nanoparticles composed of biodegradable polymers, e.g., poly(lactic-co-glycolic acid) (PLGA), are well suited for incorporation of hydrophobic and positively-charged molecules. They provide good colloidal stability, low toxicity, and the possibility of sustained release. However, due to the anionic nature of PLGA at physiological pH [131], the mRNA encapsulation efficiency is very low. Polymer-based carriers exhibit considerable potential for gene therapy owing to the substantial transfection efficiency and tolerable toxicity [132]. A series of multifunctional block copolymers, i.e., dimethylaminoethyl methacrylate (DEAMA), poly(ethylene glycol) methacrylate, and DEAEMA-co-n-butyl methacrylate, demonstrated a transfection efficiency of 77% and 50% in RAW 264.7 macrophages and DC2.4 dendritic cells, respectively, thereby exhibiting potential as a carrier for mRNA-based intracellular vaccine delivery [133]. While different types of polymers and copolymers have been tested, the correlation between the structure of polymers and their biological response, e.g., transfection and toxicity, was found to be poor and thus, design of various polymer-based delivery systems relies on empirical, rather than rational approaches [134]. Despite the advantages mentioned above, polymer-based delivery systems are not as clinically advanced as lipid-based delivery systems owing to their polydispersity and challenges pertaining to metabolism of large molecular weight polymers [21].
4.2.4. Lipid-Based Vectors
Vectors based on lipids or lipid-like compounds (lipidoids) represent the most commonly used non-viral gene carriers [21]. Various synthetic and naturally-derived lipids have been employed to form liposomes (complexes of liposomes and nucleic acids are referred to as lipoplexes) or lipid nanoparticles (LNPs), both of which have been reported to efficiently deliver mRNA-based vaccines (Table 2). LNPs are often formulated by using cationic lipids displaying tertiary or quaternary amines to encapsulate the polyanionic mRNA. Cationic lipids spontaneously encapsulate negatively-charged mRNA, mediated by a combination of attractive electrostatic interactions with RNA and hydrophobic interactions, and thus have been used alone or in combination for lipofection of mRNA. The first reported use of LNPs as delivery system for mRNA came in 2015, with the delivery system consisting of ionizable cationic lipid/phosphatidylcholine/cholesterol/PEG-lipid in the ratio of (50:10:38.5:1.5 mol/mol) [86]. Examples of cationic lipids include e.g., 1,2-di-O-octadecenyl-3-trimethylammonium propane (DOTMA) [135], DOTAP [136], and zwitterionic DOPE [137,138]. They are structurally denoted by a cationic headgroup, a hydrophobic tail group, and a linking group in between [139]. However, cationic lipids have been observed to exhibit pro-inflammatory reactions and undesirable side effects [140]. Therefore, neutral lipids are also incorporated into cationic liposomes to decrease toxicity and attain high transfection levels in vivo [106]. The mechanism of LNP-mediated delivery of mRNA is not fully understood, but LNPs are suggested to be internalized by endocytosis and are attached electrostatically and fused with the cell membrane via inverted non-bilayer lipid phases [21].

Table 2.
Examples of nanoparticulate drug delivery systems for mRNA delivery.
Liposomes are closed membrane structures, which are formed by self-assembly when phospholipids are dispersed in aqueous systems [141]. They consist of at least one phospholipid bilayer, which mimics the cell membrane structure enclosing an aqueous core [8]. DOTAP/DOPE at a 1:1 molar ratio has been reported as an effective transfection agent for mRNA encoding the HIV-1 antigen Gag, which successfully induced an antigen-specific immune response in vivo in mice [126]. Additionally, 3β-[N-(N’,N’-dimethylaminoethane) carbamoyl](DC)-Cholesterol)/DOPE-based liposomes in a [1:2] ratio achieved high encapsulation efficiency of enhanced green fluorescent protein (eGFP) mRNA, along with high eGFP expression in vitro [106]. Furthermore, an additional multi-component LNP displayed a tumor-suppressant effect when loaded with herpes simplex virus I (HSV I) thymidine kinase encoding mRNA. The LNPs were composed of DOTAP/Cholesterol [1:1] liposomes along with 1,2-distearoyl-phosphatidylethanolamine (DSPE)-polyethylene glycol (PEG) and DSPE-PEG-anisamide (AA) [142]. The principle behind their effectiveness may be summarized as a combination of its electrostatic interactions attributable to opposite charges and hydrophobic interactions with mRNA. Additionally, the endosomal escape capabilities and self-assembling properties resulting in uniform layers enclosing polymeric cores also contribute to the wide application of cationic lipids [143]. However, in vivo studies are more challenging due to the fast elimination of cationic lipids by the mononuclear phagocytic system [144]. Cationic lipids consisting of only one quaternary ammonium headgroup pose safety issues such as toxicity and immunogenicity in vitro [145] and in vivo [146]. For instance, cationic liposomes when administered via the intravenous route may induce hepatotoxicity [147] and can trigger a strong IFN-γ response in mice resulting in inflammation [148,149]. Furthermore, positively-charged lipids, e.g., DOTAP and DOTMA, can be neutralized by anionic serum proteins, leading to toxicity and reduced efficacy [150]. Moreover, challenges like unrestricted protein binding, colloidal instability, and drug leakage may arise [151].
Alternatively, new gene delivery vectors containing ionizable lipids [152] and lipid-like materials termed lipidoids [153] have been introduced to overcome challenges posed by conventional cationic lipids while retaining their advantageous transfection properties. Ionizable lipids for mRNA transfection are positively-charged at low pH (which aids in mRNA complexation when it is carried out in acidic buffer) but are neutral at physiological pH (for reduced toxicity post-injection) [154]. Unlike conventional cationic lipids, these lipidoids display a series of secondary and tertiary amines allowing for more efficient interactions with mRNA without remarkably increasing the overall charge of the delivery system [155]. Encapsulation of mRNA in nanoparticles serves to physically protect nucleic acids from degradation and depending on the specific chemistry can aid in cellular uptake and endosomal escape [156]. The combination of the ionizable lipid C12-200, cholesterol, DOPE and C14-PEG2000 at a 3.5:4.65:1.6:0.25 molar ratio, respectively, encapsulating erythropoietin mRNA (EPO-mRNA) displayed high efficacy in vivo when injected into mice, measured as the cellular expression of EPO [157]. The emphasis on the nanoparticle platform for mRNA delivery is in part due to the application of established DNA and siRNA delivery systems.
4.2.5. Lipid-Polymer Hybrid Nanoparticles
Lipid-polymer hybrid nanoparticles (LPNs) have been demonstrated previously to exhibit effective functional delivery of siRNA in vitro [158] and therapeutic delivery of siRNA in vitro and in vivo [159]. This hybrid delivery system has also shown promising results for delivery of mRNA, with the mRNA being encapsulated in a hybrid nanoparticle composed of the lipid-like material N1, N3, N5-tris(2-aminoethyl)benzene-1,3,5-tricarboxamide (TT) in TT3:DOPE:Cholesterol:DMG-PEG2000 (1,2-dimyristoyl-sn-glycerol, methoxypolyethylene glycol) with a polymeric PLGA core [160]. In addition, optimized LPNs consisting of the degradable polymer PBAE, formulated with PEG-Lipid C14-2000, showed successful delivery of mRNA to the lungs [161]. This, along with reported co-delivery of siRNA and mRNA with lipidoid polymer hybrid nanoparticles [162], shows that LPNs are an emerging nucleic acid delivery system. The hybrid formulation is thermodynamically favorable, with respect to hydrophobic, van der Waal, and electrostatic interactions [80]. Several lipids and polymers have been investigated to formulate stable nucleic acid lipid particles using this delivery system. Most common polymers employed are PLGA, polycaprolactone, polylactic acid, or their combinations, whereas the lipids used include DOTAP, 1,2-dilauroyl-sn-glycero-3-phosphocholine, 1,2-distearoyl-sn-glycero-3-phosphocholine, lecithin, DSPE, and PEG, among others. Structurally, based on small-angle X-ray scattering and cryogenic transmission electron microscopy, these nanoparticles are suggested to entail a polymeric matrix core with lamellar lipid structures with the nucleic acid localized in the core and in the corona [163].
4.2.6. Peptide-Based Vectors
Peptide-based systems for mRNA delivery are gaining momentum due to the versatility peptides can offer. Peptide-based delivery systems, both alone and in combination with other materials such as polymers, have been reported in the literature. In a study concerning ovarian cancer therapy, the commercially available cell-penetrating peptide (CPP) PepFect14 was complexed with eGFP mRNA via attractive electrostatic interactions [164]. This nanoparticulate formulation was more efficient in transfecting eGFP mRNA into cells associated with ovarian cancer than commercially available lipofectamine MessengerMAX. Similarly, the CPP RALA has been used to effectively deliver both eGFP and OVA mRNA and has been demonstrated to outperform the cationic lipid DOTAP and the fusogenic lipid DOPE [165]. However, current limitations include targeted cell delivery [164] and short circulation half-life due to the low stability in serum-containing medium [166].
Recently, a novel polymer-peptide hybrid mRNA delivery nanoplatform was introduced [167] combining both polymer (PLA) based micelles and a cationic fusogenic peptide (RALA) to achieve appropriate degradability, mRNA stability, and endosomolytic properties for translation. It was reported to protect eGFP as well as FLuc mRNA against serum nuclease degradation and achieve DC transfection. Indeed, peptide-based vectors and hybrids are promising and interesting additions to the various existing non-viral carriers for the delivery of mRNA.
4.3. Cell-Specific mRNA Delivery
Cell-specific delivery of mRNA would be beneficial for the development of mRNA-based therapeutics. This can enhance the delivery of mRNA molecules to the targeted cells and hence reduce the required mRNA dose, as well as reducing potential off-target effects. It has been reported that lymphoid organs can be targeted by adjusting the net charge of the formulation [177]. This is based on the principle of APCs being in the vicinity of T cells in these organs, thus providing optimal conditions for efficient priming and amplification of T-cell responses. Site-specific delivery of mRNA-loaded nanoparticles via active targeting has been shown to result in induction of strong effector and memory T-cell responses, and mediation of potent IFN-α-dependent rejection of progressive tumors, as observed with RNAs. In another study, cell-specific delivery of FLuc and IL-10 mRNA to leukocytes (Ly6c+) was achieved by coating the formulated mRNA-containing LNPs with anti-L6c+ monoclonal antibodies [178]. Alternatively, DCs and macrophages express receptors with the ability to present antigens, e.g., C-type lectin receptors [179], which recognize sugar groups such as mannose- and fucose-terminated glycans [180] and mediate the endocytosis of mannose-modified nanoparticles. This has been exploited for transfection of GFP mRNA into DCs by self-assembly of mannose-cholesterol conjugates with varying PEG units as linkers [181].
5. Conclusions and Future Perspectives
The field of mRNA-based therapeutics spans from protein replacement therapy and gene editing to vaccination. With the dozens of mRNA-based vaccine candidates currently in pre-clinical and clinical phases of development, it is evident that the mRNA-based vaccine technology is a promising tool for the development of novel therapeutic and prophylactic vaccines against infectious diseases and cancer. However, the multifarious obstacles associated with mRNA’s extremely large size, charge, intrinsic instability, and high susceptibility to enzymatic degradation hamper the translation of mRNA-based therapeutics from the bench to the bedside. Therefore, the wider application of mRNA-based therapeutics is still limited by the need for improved vectors or drug delivery systems. Advanced delivery systems can be applied to overcome the poor stability, cell targeting, and translational efficiency of naked mRNA. However, many clinically tested mRNA vaccine candidates are formulated without any delivery system, which suggests a need for further improvement of delivery systems for mRNA vaccines. Presently, lipoplexes and lipid-based nanoparticles are mostly used for delivering mRNA. Additionally, polymers and lipid-polymer hybrid nanoparticles offer great promise in terms of safety, stability, high transfection efficiency, and low price. Continued advancement in mRNA formulation and delivery using different nanomaterials can improve the wider use of mRNA for the treatment and prevention of infectious diseases and cancers.
Author Contributions
A.W., A.A., A.L., and A.T. wrote the review article and C.F. and A.T. critically revised all versions of the article. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Independent Research Fund, Denmark, Technology and Production Sciences (grant number 9041-00198B).
Acknowledgments
We gratefully acknowledge the financial support from the Independent Research Fund, Denmark, Technology and Production Sciences (grant number 9041-00198B). The funder had no role in literature search, decision to publish, or preparation of the manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Bragazzi, N.L.; Gianfredi, V.; Villarini, M.; Rosselli, R.; Nasr, A.; Hussein, A.; Martini, M.; Behzadifar, M. Vaccines Meet Big Data: State-of-the-Art and Future Prospects. From the Classical 3Is (“Isolate-Inactivate-Inject”) Vaccinology 1.0 to Vaccinology 3.0, Vaccinomics, and Beyond: A Historical Overview. Front. Pub. Health 2018, 6, 62. [Google Scholar] [CrossRef] [PubMed]
- Chandler, M.; Johnson, M.B.; Panigaj, M.; Afonin, K.A. Innate immune responses triggered by nucleic acids inspire the design of immunomodulatory nucleic acid nanoparticles (NANPs). Curr. Opin. Biotechnol. 2020, 63, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. mRNA vaccines - a new era in vaccinology. Nat. Rev. Drug Discov. 2018, 17, 261–279. [Google Scholar] [CrossRef] [PubMed]
- Jahanafrooz, Z.; Baradaran, B.; Mosafer, J.; Hashemzaei, M.; Rezaei, T.; Mokhtarzadeh, A.; Hamblin, M.R. Comparison of DNA and mRNA vaccines against cancer. Drug Discov. Today 2019. [Google Scholar] [CrossRef]
- Iavarone, C.; O’Hagan, D.T.; Yu, D.; Delahaye, N.F.; Ulmer, J.B. Mechanism of action of mRNA-based vaccines. Expert Rev. Vaccines 2017, 16, 871–881. [Google Scholar] [CrossRef]
- Iglesias-Lopez, C.; Agusti, A.; Obach, M.; Vallano, A. Regulatory Framework for Advanced Therapy Medicinal Products in Europe and United States. Front. Pharmacol. 2019, 10, 921. [Google Scholar] [CrossRef]
- Maruggi, G.; Zhang, C.; Li, J.; Ulmer, J.B.; Yu, D. mRNA as a Transformative Technology for Vaccine Development to Control Infectious Diseases. Mol. Ther. 2019, 27, 757–772. [Google Scholar] [CrossRef]
- Hajj, K.A.; Whitehead, K.A. Tools for translation: Non-viral materials for therapeutic mRNA delivery. Nat. Rev. Mat. 2017, 2, 1–17. [Google Scholar] [CrossRef]
- Lin, Y.; Li, Z.; Wang, T.; Wang, X.; Wang, L.; Dong, W.; Jing, C.; Yang, X. MAR characteristic motifs mediate episomal vector in CHO cells. Gene 2015, 559, 137–143. [Google Scholar] [CrossRef]
- Ramamoorth, M.; Narvekar, A. Non viral vectors in gene therapy- an overview. J. Clin. Diagn. Res. 2015, 9, GE01-06. [Google Scholar] [CrossRef]
- Zhong, Z.; Mc Cafferty, S.; Combes, F.; Huysmans, H.; De Temmerman, J.; Gitsels, A.; Vanrompay, D.; Portela Catani, J.; Sanders, N.N. mRNA therapeutics deliver a hopeful message. Nano. Today 2018, 23, 16–39. [Google Scholar] [CrossRef]
- Yamamoto, A.; Kormann, M.; Rosenecker, J.; Rudolph, C. Current prospects for mRNA gene delivery. Eur. J. Pharm. Biopharm. 2009, 71, 484–489. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Hagedorn, C.; Schulz, E.; Lipps, H.J.; Ehrhardt, A. Viral hybrid-vectors for delivery of autonomous replicons. Curr. Gene Ther. 2014, 14, 10–23. [Google Scholar] [CrossRef] [PubMed]
- Vogel, A.B.; Lambert, L.; Kinnear, E.; Busse, D.; Erbar, S.; Reuter, K.C.; Wicke, L.; Perkovic, M.; Beissert, T.; Haas, H.; et al. Self-Amplifying RNA Vaccines Give Equivalent Protection against Influenza to mRNA Vaccines but at Much Lower Doses. Mol. Ther. 2018, 26, 446–455. [Google Scholar] [CrossRef]
- Lundstrom, K. Self-Replicating RNA Viruses for RNA Therapeutics. Molecules 2018, 23. [Google Scholar] [CrossRef]
- Van Hoecke, L.; Roose, K. How mRNA therapeutics are entering the monoclonal antibody field. J. Transl. Med. 2019, 17, 54. [Google Scholar] [CrossRef]
- Sahin, U.; Kariko, K.; Tureci, O. mRNA-based therapeutics--developing a new class of drugs. Nat. Rev. Drug Discov. 2014, 13, 759–780. [Google Scholar] [CrossRef]
- Androulla, M.N.; Lefkothea, P.C. In Vitro-Transcribed (IVT)-mRNA CAR Therapy Development. Methods Mol. Biol. 2020, 2086, 87–117. [Google Scholar] [CrossRef]
- Reichmuth, A.M.; Oberli, M.A.; Jaklenec, A.; Langer, R.; Blankschtein, D. mRNA vaccine delivery using lipid nanoparticles. Ther. Deliv. 2016, 7, 319–334. [Google Scholar] [CrossRef]
- Schlake, T.; Thess, A.; Thran, M.; Jordan, I. mRNA as novel technology for passive immunotherapy. Cell Mol. Life Sci. 2019, 76, 301–328. [Google Scholar] [CrossRef]
- Kowalski, P.S.; Rudra, A.; Miao, L.; Anderson, D.G. Delivering the Messenger: Advances in Technologies for Therapeutic mRNA Delivery. Mol. Ther. 2019, 27, 710–728. [Google Scholar] [CrossRef] [PubMed]
- Cohn, L.; Delamarre, L. Dendritic cell-targeted vaccines. Front Immunol. 2014, 5, 255. [Google Scholar] [CrossRef] [PubMed]
- Cu, Y.; Broderick, K.E.; Banerjee, K.; Hickman, J.; Otten, G.; Barnett, S.; Kichaev, G.; Sardesai, N.Y.; Ulmer, J.B.; Geall, A. Enhanced Delivery and Potency of Self-Amplifying mRNA Vaccines by Electroporation in Situ. Vaccines 2013, 1, 367–383. [Google Scholar] [CrossRef] [PubMed]
- Wolff, J.A.; Malone, R.W.; Williams, P.; Chong, W.; Acsadi, G.; Jani, A.; Felgner, P.L. Direct gene transfer into mouse muscle in vivo. Science 1990, 247, 1465–1468. [Google Scholar] [CrossRef] [PubMed]
- Martinon, F.; Krishnan, S.; Lenzen, G.; Magne, R.; Gomard, E.; Guillet, J.G.; Levy, J.P.; Meulien, P. Induction of virus-specific cytotoxic T lymphocytes in vivo by liposome-entrapped mRNA. Eur. J. Immunol. 1993, 23, 1719–1722. [Google Scholar] [CrossRef] [PubMed]
- Kallen, K.J.; Thess, A. A development that may evolve into a revolution in medicine: mRNA as the basis for novel, nucleotide-based vaccines and drugs. Ther. Adv. Vaccines 2014, 2, 10–31. [Google Scholar] [CrossRef]
- Hoerr, I.; Obst, R.; Rammensee, H.G.; Jung, G. In vivo application of RNA leads to induction of specific cytotoxic T lymphocytes and antibodies. Eur. J. Immunol. 2000, 30, 1–7. [Google Scholar] [CrossRef]
- Guan, S.; Rosenecker, J. Nanotechnologies in delivery of mRNA therapeutics using nonviral vector-based delivery systems. Gene Ther. 2017, 24, 133–143. [Google Scholar] [CrossRef]
- Baiersdörfer, M.; Boros, G.; Muramatsu, H.; Mahiny, A.; Vlatkovic, I.; Sahin, U.; Karikó, K. A Facile Method for the Removal of dsRNA Contaminant from In Vitro-Transcribed mRNA. Mol. Ther. Nucleic Acids 2019, 15, 26–35. [Google Scholar] [CrossRef]
- Pardi, N.; Muramatsu, H.; Weissman, D.; Karikó, K. In Vitro Transcription of Long RNA Containing Modified Nucleosides. In Synthetic Messenger RNA and Cell Metabolism Modulation: Methods and Protocols; Rabinovich, P.M., Ed.; Humana Press: Totowa, NJ, USA, 2013; pp. 29–42. [Google Scholar]
- Weissman, D.; Pardi, N.; Muramatsu, H.; Karikó, K. HPLC Purification of In Vitro Transcribed Long RNA. In Synthetic Messenger RNA and Cell Metabolism Modulation: Methods and Protocols; Rabinovich, P.M., Ed.; Humana Press: Totowa, NJ, USA, 2013; pp. 43–54. [Google Scholar]
- Li, Y.; Kiledjian, M. Regulation of mRNA decapping. Wiley Interdiscip. Rev. RNA 2010, 1, 253–265. [Google Scholar] [CrossRef]
- Cowling, V.H.; Cole, M.D. Myc Regulation of mRNA Cap Methylation. Genes Cancer 2010, 1, 576–579. [Google Scholar] [CrossRef] [PubMed]
- Roers, A.; Hiller, B.; Hornung, V. Recognition of Endogenous Nucleic Acids by the Innate Immune System. Immunity 2016, 44, 739–754. [Google Scholar] [CrossRef] [PubMed]
- Wojtczak, B.A.; Sikorski, P.J.; Fac-Dabrowska, K.; Nowicka, A.; Warminski, M.; Kubacka, D.; Nowak, E.; Nowotny, M.; Kowalska, J.; Jemielity, J. 5’-Phosphorothiolate Dinucleotide Cap Analogues: Reagents for Messenger RNA Modification and Potent Small-Molecular Inhibitors of Decapping Enzymes. J. Am. Chem. Soc. 2018, 140, 5987–5999. [Google Scholar] [CrossRef] [PubMed]
- Ramanathan, A.; Robb, G.B.; Chan, S.H. mRNA capping: Biological functions and applications. Nucleic. Acids Res. 2016, 44, 7511–7526. [Google Scholar] [CrossRef]
- Zimmermann, O.; Homann, J.M.; Bangert, A.; Müller, A.M.; Hristov, G.; Goeser, S.; Wiehe, J.M.; Zittrich, S.; Rottbauer, W.; Torzewski, J.; et al. Successful use of mRNA-nucleofection for overexpression of interleukin-10 in murine monocytes/macrophages for anti-inflammatory therapy in a murine model of autoimmune myocarditis. J. Am. Heart Association 2012. [Google Scholar] [CrossRef]
- Hamm, J.; Mattaj, I.W. Monomethylated cap structures facilitate RNA export from the nucleus. Cell 1990, 63, 109–118. [Google Scholar] [CrossRef]
- Rydzik, A.M.; Warminski, M.; Sikorski, P.J.; Baranowski, M.R.; Walczak, S.; Kowalska, J.; Zuberek, J.; Lukaszewicz, M.; Nowak, E.; TD, W.C.; et al. mRNA cap analogues substituted in the tetraphosphate chain with CX2: Identification of O-to-CCl2 as the first bridging modification that confers resistance to decapping without impairing translation. Nucleic Acids Res. 2017, 45, 8661–8675. [Google Scholar] [CrossRef]
- Kariko, K.; Muramatsu, H.; Welsh, F.A.; Ludwig, J.; Kato, H.; Akira, S.; Weissman, D. Incorporation of pseudouridine into mRNA yields superior nonimmunogenic vector with increased translational capacity and biological stability. Mol. Ther. 2008, 16, 1833–1840. [Google Scholar] [CrossRef]
- Kocmik, I.; Piecyk, K.; Rudzinska, M.; Niedzwiecka, A.; Darzynkiewicz, E.; Grzela, R.; Jankowska-Anyszka, M. Modified ARCA analogs providing enhanced translational properties of capped mRNAs. Cell Cycle 2018, 17, 1624–1636. [Google Scholar] [CrossRef]
- Kiriakidou, M.; Tan, G.S.; Lamprinaki, S.; De Planell-Saguer, M.; Nelson, P.T.; Mourelatos, Z. An mRNA m7G cap binding-like motif within human Ago2 represses translation. Cell 2007, 129, 1141–1151. [Google Scholar] [CrossRef]
- Jemielity, J.; Fowler, T.; Zuberek, J.; Stepinski, J.; Lewdorowicz, M.; Niedzwiecka, A.; Stolarski, R.; Darzynkiewicz, E.; Rhoads, R.E. Novel “anti-reverse” cap analogs with superior translational properties. RNA 2003, 9, 1108–1122. [Google Scholar] [CrossRef] [PubMed]
- Grudzien-Nogalska, E.; Jemielity, J.; Kowalska, J.; Darzynkiewicz, E.; Rhoads, R.E. Phosphorothioate cap analogs stabilize mRNA and increase translational efficiency in mammalian cells. RNA 2007, 13, 1745–1755. [Google Scholar] [CrossRef] [PubMed]
- Ziemniak, M.; Kowalska, J.; Lukaszewicz, M.; Zuberek, J.; Wnek, K.; Darzynkiewicz, E.; Jemielity, J. Phosphate-modified analogues of m(7)GTP and m(7)Gppppm(7)G-Synthesis and biochemical properties. Bioorg. Med. Chem. 2015, 23, 5369–5381. [Google Scholar] [CrossRef] [PubMed]
- Kore, A.R.; Shanmugasundaram, M.; Charles, I.; Vlassov, A.V.; Barta, T.J. Locked nucleic acid (LNA)-modified dinucleotide mRNA cap analogue: Synthesis, enzymatic incorporation, and utilization. J. Am. Chem. Soc. 2009, 131, 6364–6365. [Google Scholar] [CrossRef] [PubMed]
- Kowalska, J.; Wypijewska del Nogal, A.; Darzynkiewicz, Z.M.; Buck, J.; Nicola, C.; Kuhn, A.N.; Lukaszewicz, M.; Zuberek, J.; Strenkowska, M.; Ziemniak, M.; et al. Synthesis, properties, and biological activity of boranophosphate analogs of the mRNA cap: Versatile tools for manipulation of therapeutically relevant cap-dependent processes. Nucleic Acids Res. 2014, 42, 10245–10264. [Google Scholar] [CrossRef]
- Grudzien-Nogalska, E.; Kiledjian, M. New insights into decapping enzymes and selective mRNA decay. Wiley Interdiscip Rev RNA 2017, 8. [Google Scholar] [CrossRef]
- Shanmugasundaram, M.; Charles, I.; Kore, A.R. Design, synthesis and biological evaluation of dinucleotide mRNA cap analog containing propargyl moiety. Bioorg. Med. Chem. 2016, 24, 1204–1208. [Google Scholar] [CrossRef]
- Yunus, M.A.; Chung, L.M.; Chaudhry, Y.; Bailey, D.; Goodfellow, I. Development of an optimized RNA-based murine norovirus reverse genetics system. J. Virol. Methods 2010, 169, 112–118. [Google Scholar] [CrossRef]
- Devoldere, J.; Dewitte, H.; De Smedt, S.C.; Remaut, K. Evading innate immunity in nonviral mRNA delivery: don’t shoot the messenger. Drug Discov. Today 2016, 21, 11–25. [Google Scholar] [CrossRef]
- Vaidyanathan, S.; Azizian, K.T.; Haque, A.; Henderson, J.M.; Hendel, A.; Shore, S.; Antony, J.S.; Hogrefe, R.I.; Kormann, M.S.D.; Porteus, M.H.; et al. Uridine Depletion and Chemical Modification Increase Cas9 mRNA Activity and Reduce Immunogenicity without HPLC Purification. Mol. Ther. Nucleic Acids 2018, 12, 530–542. [Google Scholar] [CrossRef]
- Muttach, F.; Muthmann, N.; Rentmeister, A. Synthetic mRNA capping. Beilstein J. Org. Chem. 2017, 13, 2819–2832. [Google Scholar] [CrossRef] [PubMed]
- Shin, D.; Azizian, K.T.; Henderson, J.M.; Lebedev, A.; Hogrefe, R.I.; Houston, M.; McCaffrey, A.P.; BioTechnologies, T. Exploring the Messenger RNA Capping Code: CleanCap Co-Transcriptional Capping Allows Synthesis of Cap 0, Cap 1, Cap 2 and (M6) A (M) Capped RNAs; Cell Press: Cambridge, MA, USA, 2018; p. 91. [Google Scholar]
- Diebold, S.S.; Kaisho, T.; Hemmi, H.; Akira, S.; Reis e Sousa, C. Innate antiviral responses by means of TLR7-mediated recognition of single-stranded RNA. Science 2004, 303, 1529–1531. [Google Scholar] [CrossRef] [PubMed]
- Suknuntha, K.; Tao, L.; Brok-Volchanskaya, V.; D’Souza, S.S.; Kumar, A.; Slukvin, I. Optimization of Synthetic mRNA for Highly Efficient Translation and its Application in the Generation of Endothelial and Hematopoietic Cells from Human and Primate Pluripotent Stem Cells. Stem Cell Rev. Rep. 2018, 14, 525–534. [Google Scholar] [CrossRef] [PubMed]
- Creusot, R.J.; Chang, P.; Healey, D.G.; Tcherepanova, I.Y.; Nicolette, C.A.; Fathman, C.G. A short pulse of IL-4 delivered by DCs electroporated with modified mRNA can both prevent and treat autoimmune diabetes in NOD mice. Mol. Ther. 2010, 18, 2112–2120. [Google Scholar] [CrossRef] [PubMed]
- Asrani, K.H.; Farelli, J.D.; Stahley, M.R.; Miller, R.L.; Cheng, C.J.; Subramanian, R.R.; Brown, J.M. Optimization of mRNA untranslated regions for improved expression of therapeutic mRNA. RNA Biol. 2018, 15, 756–762. [Google Scholar] [CrossRef] [PubMed]
- Adibzadeh, S.; Fardaei, M.; Takhshid, M.A.; Miri, M.R.; Rafiei Dehbidi, G.; Farhadi, A.; Ranjbaran, R.; Alavi, P.; Nikouyan, N.; Seyyedi, N.; et al. Enhancing Stability of Destabilized Green Fluorescent Protein Using Chimeric mRNA Containing Human Beta-Globin 5’ and 3’ Untranslated Regions. Avicenna J. Med. Biotechnol. 2019, 11, 112–117. [Google Scholar]
- Zhao, Y.; Moon, E.; Carpenito, C.; Paulos, C.M.; Liu, X.; Brennan, A.L.; Chew, A.; Carroll, R.G.; Scholler, J.; Levine, B.L.; et al. Multiple injections of electroporated autologous T cells expressing a chimeric antigen receptor mediate regression of human disseminated tumor. Cancer Res. 2010, 70, 9053–9061. [Google Scholar] [CrossRef]
- Kreiter, S.; Selmi, A.; Diken, M.; Koslowski, M.; Britten, C.M.; Huber, C.; Tureci, O.; Sahin, U. Intranodal vaccination with naked antigen-encoding RNA elicits potent prophylactic and therapeutic antitumoral immunity. Cancer Res. 2010, 70, 9031–9040. [Google Scholar] [CrossRef]
- Kreiter, S.; Selmi, A.; Diken, M.; Sebastian, M.; Osterloh, P.; Schild, H.; Huber, C.; Tureci, O.; Sahin, U. Increased antigen presentation efficiency by coupling antigens to MHC class I trafficking signals. J. Immunol. 2008, 180, 309–318. [Google Scholar] [CrossRef]
- Fotin-Mleczek, M.; Duchardt, K.M.; Lorenz, C.; Pfeiffer, R.; Ojkic-Zrna, S.; Probst, J.; Kallen, K.J. Messenger RNA-based vaccines with dual activity induce balanced TLR-7 dependent adaptive immune responses and provide antitumor activity. J. Immunother. 2011, 34, 1–15. [Google Scholar] [CrossRef]
- Goss, D.J.; Kleiman, F.E. Poly(A) binding proteins: Are they all created equal? Wiley Interdiscip. Rev. RNA 2013, 4, 167–179. [Google Scholar] [CrossRef] [PubMed]
- Hershey, J.W. Regulation of protein synthesis and the role of eIF3 in cancer. Braz. J. Med. Biol. Res. 2010, 43, 920–930. [Google Scholar] [CrossRef]
- Anderson, B.R.; Muramatsu, H.; Nallagatla, S.R.; Bevilacqua, P.C.; Sansing, L.H.; Weissman, D.; Kariko, K. Incorporation of pseudouridine into mRNA enhances translation by diminishing PKR activation. Nucleic Acids Res. 2010, 38, 5884–5892. [Google Scholar] [CrossRef] [PubMed]
- Munroe, D.; Jacobson, A. mRNA poly (A) tail, a 3’enhancer of translational initiation. Mol. Cell. Biol. 1990, 10, 3441–3455. [Google Scholar] [CrossRef] [PubMed]
- Kormann, M.S.; Hasenpusch, G.; Aneja, M.K.; Nica, G.; Flemmer, A.W.; Herber-Jonat, S.; Huppmann, M.; Mays, L.E.; Illenyi, M.; Schams, A.; et al. Expression of therapeutic proteins after delivery of chemically modified mRNA in mice. Nat. Biotechnol. 2011, 29, 154–157. [Google Scholar] [CrossRef]
- Holtkamp, S.; Kreiter, S.; Selmi, A.; Simon, P.; Koslowski, M.; Huber, C.; Tureci, O.; Sahin, U. Modification of antigen-encoding RNA increases stability, translational efficacy, and T-cell stimulatory capacity of dendritic cells. Blood 2006, 108, 4009–4017. [Google Scholar] [CrossRef]
- Coller, J.; Parker, R. Eukaryotic mRNA decapping. Annu. Rev. Biochem. 2004, 73, 861–890. [Google Scholar] [CrossRef]
- Mugridge, J.S.; Coller, J.; Gross, J.D. Structural and molecular mechanisms for the control of eukaryotic 5’-3’ mRNA decay. Nat. Struct. Mol. Biol. 2018, 25, 1077–1085. [Google Scholar] [CrossRef]
- Sharova, L.V.; Sharov, A.A.; Nedorezov, T.; Piao, Y.; Shaik, N.; Ko, M.S.H. Database for mRNA half-life of 19 977 genes obtained by DNA microarray analysis of pluripotent and differentiating mouse embryonic stem cells. DNA Res. 2008, 16, 45–58. [Google Scholar] [CrossRef]
- Houseley, J.; Tollervey, D. The many pathways of RNA degradation. Cell 2009, 136, 763–776. [Google Scholar] [CrossRef]
- Deering, R.P.; Kommareddy, S.; Ulmer, J.B.; Brito, L.A.; Geall, A.J. Nucleic acid vaccines: Prospects for non-viral delivery of mRNA vaccines. Expert Opin. Drug Deliv. 2014, 11, 885–899. [Google Scholar] [CrossRef] [PubMed]
- Golombek, S.; Pilz, M.; Steinle, H.; Kochba, E.; Levin, Y.; Lunter, D.; Schlensak, C.; Wendel, H.P.; Avci-Adali, M. Intradermal Delivery of Synthetic mRNA Using Hollow Microneedles for Efficient and Rapid Production of Exogenous Proteins in Skin. Mol. Ther. Nucleic Acids 2018, 11, 382–392. [Google Scholar] [CrossRef] [PubMed]
- Koh, K.J.; Liu, Y.; Lim, S.H.; Loh, X.J.; Kang, L.; Lim, C.Y.; Phua, K.K.L. Formulation, characterization and evaluation of mRNA-loaded dissolvable polymeric microneedles (RNApatch). Sci. Rep. 2018, 8, 11842. [Google Scholar] [CrossRef] [PubMed]
- Tavernier, G.; Andries, O.; Demeester, J.; Sanders, N.N.; De Smedt, S.C.; Rejman, J. mRNA as gene therapeutic: How to control protein expression. J. Control. Release 2011, 150, 238–247. [Google Scholar] [CrossRef]
- Zhang, R.; Men, K.; Zhang, X.; Huang, R.; Tian, Y.; Zhou, B.; Yu, C.; Wang, Y.; Ji, X.; Hu, Q.; et al. Delivery of a Modified mRNA Encoding IL-22 Binding Protein (IL-22BP) for Colon Cancer Gene Therapy. J. Biomed. Nanotechnol. 2018, 14, 1239–1251. [Google Scholar] [CrossRef]
- Schlake, T.; Thess, A.; Fotin-Mleczek, M.; Kallen, K.J. Developing mRNA-vaccine technologies. RNA Biol. 2012, 9, 1319–1330. [Google Scholar] [CrossRef]
- Mukherjee, A.; Waters, A.K.; Kalyan, P.; Achrol, A.S.; Kesari, S.; Yenugonda, V.M. Lipid-polymer hybrid nanoparticles as a next-generation drug delivery platform: State of the art, emerging technologies, and perspectives. Int. J. Nanomed. 2019, 14, 1937–1952. [Google Scholar] [CrossRef]
- Sahin, U.; Karikó, K.; Türeci, Ö. mRNA-based therapeutics — developing a new class of drugs. Nat. Pub. Gr. 2014. [Google Scholar] [CrossRef]
- Takahashi, M.; Elbarbary, R.A.; Nakashima, A.; Abe, M.; Watanabe, N.; Narita, M.; Takahashi, M.; Tamura, M.; Yoshida, T.; Nashimoto, M. A naked RNA heptamer targeting the human Bcl-2 mRNA induces apoptosis of HL60 leukemia cells. Cancer Lett. 2013, 328, 362–368. [Google Scholar] [CrossRef]
- Weide, B.; Carralot, J.P.; Reese, A.; Scheel, B.; Eigentler, T.K.; Hoerr, I.; Rammensee, H.G.; Garbe, C.; Pascolo, S. Results of the first phase I/II clinical vaccination trial with direct injection of mRNA. J. Immunother. 2008, 31, 180–188. [Google Scholar] [CrossRef]
- Lorenz, C.; Fotin-Mleczek, M.; Roth, G.; Becker, C.; Dam, T.C.; Verdurmen, W.P.; Brock, R.; Probst, J.; Schlake, T. Protein expression from exogenous mRNA: Uptake by receptor-mediated endocytosis and trafficking via the lysosomal pathway. RNA Biol. 2011, 8, 627–636. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.Y.; Kim, M.K.; Kim-Han, T.H.; Indik, Z.K.; Schreiber, A.D. Effect of locally administered Syk siRNA on allergen-induced arthritis and asthma. Mol. Immunol. 2013, 53, 52–59. [Google Scholar] [CrossRef] [PubMed]
- Pardi, N.; Tuyishime, S.; Muramatsu, H.; Kariko, K.; Mui, B.L.; Tam, Y.K.; Madden, T.D.; Hope, M.J.; Weissman, D. Expression kinetics of nucleoside-modified mRNA delivered in lipid nanoparticles to mice by various routes. J. Control. Release 2015, 217, 345–351. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Maruggi, G.; Shan, H.; Li, J. Advances in mRNA Vaccines for Infectious Diseases. Front. Immunol. 2019, 10, 594. [Google Scholar] [CrossRef] [PubMed]
- Pardi, N.; Hogan, M.J.; Pelc, R.S.; Muramatsu, H.; Andersen, H.; DeMaso, C.R.; Dowd, K.A.; Sutherland, L.L.; Scearce, R.M.; Parks, R.; et al. Zika virus protection by a single low-dose nucleoside-modified mRNA vaccination. Nature 2017, 543, 248–251. [Google Scholar] [CrossRef] [PubMed]
- Zakrewsky, M.; Kumar, S.; Mitragotri, S. Nucleic acid delivery into skin for the treatment of skin disease: Proofs-of-concept, potential impact, and remaining challenges. J. Control. Release 2015, 219, 445–456. [Google Scholar] [CrossRef]
- Phua, K.K.; Leong, K.W.; Nair, S.K. Transfection efficiency and transgene expression kinetics of mRNA delivered in naked and nanoparticle format. J. Control. Release 2013, 166, 227–233. [Google Scholar] [CrossRef]
- Van Lint, S.; Goyvaerts, C.; Maenhout, S.; Goethals, L.; Disy, A.; Benteyn, D.; Pen, J.; Bonehill, A.; Heirman, C.; Breckpot, K.; et al. Preclinical evaluation of TriMix and antigen mRNA-based antitumor therapy. Cancer Res. 2012, 72, 1661–1671. [Google Scholar] [CrossRef]
- Probst, J.; Weide, B.; Scheel, B.; Pichler, B.J.; Hoerr, I.; Rammensee, H.G.; Pascolo, S. Spontaneous cellular uptake of exogenous messenger RNA in vivo is nucleic acid-specific, saturable and ion dependent. Gene Ther. 2007, 14, 1175–1180. [Google Scholar] [CrossRef]
- Sultana, N.; Magadum, A.; Hadas, Y.; Kondrat, J.; Singh, N.; Youssef, E.; Calderon, D.; Chepurko, E.; Dubois, N.; Hajjar, R.J.; et al. Optimizing Cardiac Delivery of Modified mRNA. Mol. Ther. 2017, 25, 1306–1315. [Google Scholar] [CrossRef]
- McNamara, M.A.; Nair, S.K.; Holl, E.K. RNA-Based Vaccines in Cancer Immunotherapy. J. Immunol. Res. 2015, 2015, 794528. [Google Scholar] [CrossRef] [PubMed]
- Selmi, A.; Vascotto, F.; Kautz-Neu, K.; Tureci, O.; Sahin, U.; von Stebut, E.; Diken, M.; Kreiter, S. Uptake of synthetic naked RNA by skin-resident dendritic cells via macropinocytosis allows antigen expression and induction of T-cell responses in mice. Cancer Immunol. Immunother. 2016, 65, 1075–1083. [Google Scholar] [CrossRef] [PubMed]
- De Beuckelaer, A.; Grooten, J.; De Koker, S. Type I Interferons Modulate CD8(+) T Cell Immunity to mRNA Vaccines. Trends Mol. Med. 2017, 23, 216–226. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharya, N.; Ganguli-Indra, G.; Indra, A.K. Transcriptional control and transcriptomic analysis of lipid metabolism in skin barrier formation and atopic dermatitis (AD). Expert Rev. Proteomics 2019, 16, 627–645. [Google Scholar] [CrossRef]
- Ahlemeyer, B.; Vogt, J.F.; Michel, V.; Hahn-Kohlberger, P.; Baumgart-Vogt, E. Microporation is an efficient method for siRNA-induced knockdown of PEX5 in HepG2 cells: Evaluation of the transfection efficiency, the PEX5 mRNA and protein levels and induction of peroxisomal deficiency. Histochem. Cell Biol. 2014, 142, 577–591. [Google Scholar] [CrossRef]
- Chong, R.H.; Gonzalez-Gonzalez, E.; Lara, M.F.; Speaker, T.J.; Contag, C.H.; Kaspar, R.L.; Coulman, S.A.; Hargest, R.; Birchall, J.C. Gene silencing following siRNA delivery to skin via coated steel microneedles: In vitro and in vivo proof-of-concept. J. Control. Release 2013, 166, 211–219. [Google Scholar] [CrossRef]
- Walther, W.; Stein, U.; Fichtner, I.; Kobelt, D.; Aumann, J.; Arlt, F.; Schlag, P.M. Nonviral jet-injection gene transfer for efficient in vivo cytosine deaminase suicide gene therapy of colon carcinoma. Mol. Ther. 2005, 12, 1176–1184. [Google Scholar] [CrossRef]
- Villemejane, J.; Mir, L.M. Physical methods of nucleic acid transfer: General concepts and applications. Br J. Pharmacol. 2009, 157, 207–219. [Google Scholar] [CrossRef]
- McLenachan, S.; Zhang, D.; Palomo, A.B.; Edel, M.J.; Chen, F.K. mRNA transfection of mouse and human neural stem cell cultures. PLoS ONE 2013, 8, e83596. [Google Scholar] [CrossRef]
- Kigasawa, K.; Kajimoto, K.; Hama, S.; Saito, A.; Kanamura, K.; Kogure, K. Noninvasive delivery of siRNA into the epidermis by iontophoresis using an atopic dermatitis-like model rat. Int. J. Pharm. 2010, 383, 157–160. [Google Scholar] [CrossRef]
- Ryu, Y.C.; Kim, D.I.; Kim, S.H.; Wang, H.-M.D.; Hwang, B.H. Synergistic Transdermal Delivery of Biomacromolecules Using Sonophoresis after Microneedle Treatment. Biotechnol. Bioprocess Eng. 2018, 23, 286–292. [Google Scholar] [CrossRef]
- Su, X.; Fricke, J.; Kavanagh, D.G.; Irvine, D.J. In vitro and in vivo mRNA delivery using lipid-enveloped pH-responsive polymer nanoparticles. Mol. Pharm. 2011, 8, 774–787. [Google Scholar] [CrossRef] [PubMed]
- Michel, T.; Luft, D.; Abraham, M.K.; Reinhardt, S.; Salinas Medina, M.L.; Kurz, J.; Schaller, M.; Avci-Adali, M.; Schlensak, C.; Peter, K.; et al. Cationic Nanoliposomes Meet mRNA: Efficient Delivery of Modified mRNA Using Hemocompatible and Stable Vectors for Therapeutic Applications. Mol. Ther. Nucleic Acids 2017, 8, 459–468. [Google Scholar] [CrossRef] [PubMed]
- Dewitte, H.; Van Lint, S.; Heirman, C.; Thielemans, K.; De Smedt, S.C.; Breckpot, K.; Lentacker, I. The potential of antigen and TriMix sonoporation using mRNA-loaded microbubbles for ultrasound-triggered cancer immunotherapy. J. Control. Release 2014, 194, 28–36. [Google Scholar] [CrossRef]
- Gan, L.M.; Lagerstrom-Fermer, M.; Carlsson, L.G.; Arfvidsson, C.; Egnell, A.C.; Rudvik, A.; Kjaer, M.; Collen, A.; Thompson, J.D.; Joyal, J.; et al. Intradermal delivery of modified mRNA encoding VEGF-A in patients with type 2 diabetes. Nat. Commun. 2019, 10, 871. [Google Scholar] [CrossRef]
- Sun, N.; Ning, B.; Hansson, K.M.; Bruce, A.C.; Seaman, S.A.; Zhang, C.; Rikard, M.; DeRosa, C.A.; Fraser, C.L.; Wagberg, M.; et al. Modified VEGF-A mRNA induces sustained multifaceted microvascular response and accelerates diabetic wound healing. Sci. Rep. 2018, 8, 17509. [Google Scholar] [CrossRef]
- Ita, K. Transdermal Delivery of Drugs with Microneedles-Potential and Challenges. Pharmaceutics 2015, 7, 90–105. [Google Scholar] [CrossRef]
- Trepotec, Z.; Lichtenegger, E.; Plank, C.; Aneja, M.K.; Rudolph, C. Delivery of mRNA Therapeutics for the Treatment of Hepatic Diseases. Mol. Ther. 2019, 27, 794–802. [Google Scholar] [CrossRef]
- Chou, J.Y.; Mansfield, B.C. Recombinant AAV-directed gene therapy for type I glycogen storage diseases. Expert Opin. Biol. Ther. 2011, 11, 1011–1024. [Google Scholar] [CrossRef]
- Ehrengruber, M.U.; Lundstrom, K. Alphaviruses: Semliki Forest Virus and Sindbis Virus Vectors for Gene Transfer into Neurons. Curr. Protoc. Neurosci. 2007, 57, 4.22.21–4.22.27. [Google Scholar] [CrossRef]
- Rozovics, J.M.; Chase, A.J.; Cathcart, A.L.; Chou, W.; Gershon, P.D.; Palusa, S.; Wilusz, J.; Semler, B.L. Picornavirus modification of a host mRNA decay protein. MBio 2012, 3, e00431-12. [Google Scholar] [CrossRef] [PubMed]
- Schott, J.W.; Morgan, M.; Galla, M.; Schambach, A. Viral and Synthetic RNA Vector Technologies and Applications. Mol. Ther. 2016, 24, 1513–1527. [Google Scholar] [CrossRef]
- Strauss, J.H.; Strauss, E.G. The alphaviruses: Gene expression, replication, and evolution. Microbiol. Rev. 1994, 58, 491–562. [Google Scholar] [CrossRef]
- Vanlandingham, D.L.; Tsetsarkin, K.; Hong, C.; Klingler, K.; McElroy, K.L.; Lehane, M.J.; Higgs, S. Development and characterization of a double subgenomic chikungunya virus infectious clone to express heterologous genes in Aedes aegypti mosquitoes. Insect Biochem. Mol. Biol. 2005, 35, 1162–1170. [Google Scholar] [CrossRef] [PubMed]
- Lundstrom, K. RNA Viruses as Tools in Gene Therapy and Vaccine Development. Genes 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Lundstrom, K. Alphavirus vectors for vaccine production and gene therapy. Expert Rev. Vaccines 2003, 2, 447–459. [Google Scholar] [CrossRef] [PubMed]
- Nakanishi, M.; Otsu, M. Development of Sendai virus vectors and their potential applications in gene therapy and regenerative medicine. Curr. Gene Ther. 2012, 12, 410–416. [Google Scholar] [CrossRef]
- Bounds, C.E.; Terry, F.E.; Moise, L.; Hannaman, D.; Martin, W.D.; De Groot, A.S.; Suschak, J.J.; Dupuy, L.C.; Schmaljohn, C.S. An immunoinformatics-derived DNA vaccine encoding human class II T cell epitopes of Ebola virus, Sudan virus, and Venezuelan equine encephalitis virus is immunogenic in HLA transgenic mice. Hum. Vaccin Immunother. 2017, 13, 2824–2836. [Google Scholar] [CrossRef]
- Taylor, K.; Kolokoltsova, O.; Ronca, S.E.; Estes, M.; Paessler, S. Live, Attenuated Venezuelan Equine Encephalitis Virus Vaccine (TC83) Causes Persistent Brain Infection in Mice with Non-functional alphabeta T-Cells. Front. Microbiol. 2017, 8, 81. [Google Scholar] [CrossRef]
- Tezel, A.; Dokka, S.; Kelly, S.; Hardee, G.E.; Mitragotri, S. Topical delivery of anti-sense oligonucleotides using low-frequency sonophoresis. Pharm. Res. 2004, 21, 2219–2225. [Google Scholar] [CrossRef]
- Koch, G. Interaction of poliovirus-specific RNAs with HeLa cells and E. coli. Curr. Top Microbiol. Immunol. 1973, 62, 89–138. [Google Scholar] [CrossRef] [PubMed]
- Malone, R.W.; Felgner, P.L.; Verma, I.M. Cationic liposome-mediated RNA transfection. Proc. Natl. Acad. Sci. USA 1989, 86, 6077–6081. [Google Scholar] [CrossRef] [PubMed]
- Pollard, C.; Rejman, J.; De Haes, W.; Verrier, B.; Van Gulck, E.; Naessens, T.; De Smedt, S.; Bogaert, P.; Grooten, J.; Vanham, G.; et al. Type I IFN counteracts the induction of antigen-specific immune responses by lipid-based delivery of mRNA vaccines. Mol. Ther. 2013, 21, 251–259. [Google Scholar] [CrossRef] [PubMed]
- Ulkoski, D.; Bak, A.; Wilson, J.T.; Krishnamurthy, V.R. Recent advances in polymeric materials for the delivery of RNA therapeutics. Expert Opin. Drug Deliv. 2019, 16, 1149–1167. [Google Scholar] [CrossRef]
- Gary, D.J.; Lee, H.; Sharma, R.; Lee, J.S.; Kim, Y.; Cui, Z.Y.; Jia, D.; Bowman, V.D.; Chipman, P.R.; Wan, L.; et al. Influence of nano-carrier architecture on in vitro siRNA delivery performance and in vivo biodistribution: Polyplexes vs micelleplexes. ACS Nano 2011, 5, 3493–3505. [Google Scholar] [CrossRef]
- Yu, H.; Guo, C.; Feng, B.; Liu, J.; Chen, X.; Wang, D.; Teng, L.; Li, Y.; Yin, Q.; Zhang, Z.; et al. Triple-Layered pH-Responsive Micelleplexes Loaded with siRNA and Cisplatin Prodrug for NF-Kappa B Targeted Treatment of Metastatic Breast Cancer. Theranostics 2016, 6, 14–27. [Google Scholar] [CrossRef]
- Tros de Ilarduya, C.; Sun, Y.; Duzgunes, N. Gene delivery by lipoplexes and polyplexes. Eur. J. Pharm. Sci. 2010, 40, 159–170. [Google Scholar] [CrossRef]
- Rosenkranz, A.A.; Sobolev, A.S. Polyethylenimine-based polyplex nanoparticles and features of their behavior in cells and tissues. Russian Chem. Bull. 2016, 64, 2749–2755. [Google Scholar] [CrossRef]
- Lungwitz, U.; Breunig, M.; Blunk, T.; Gopferich, A. Polyethylenimine-based non-viral gene delivery systems. Eur. J. Pharm. Biopharm. 2005, 60, 247–266. [Google Scholar] [CrossRef]
- Cheng, C.; Convertine, A.J.; Stayton, P.S.; Bryers, J.D. Multifunctional triblock copolymers for intracellular messenger RNA delivery. Biomaterials 2012, 33, 6868–6876. [Google Scholar] [CrossRef]
- Gary, D.J.; Puri, N.; Won, Y.Y. Polymer-based siRNA delivery: Perspectives on the fundamental and phenomenological distinctions from polymer-based DNA delivery. J. Control. Release 2007, 121, 64–73. [Google Scholar] [CrossRef]
- Lu, Y.; Kawakami, S.; Yamashita, F.; Hashida, M. Development of an antigen-presenting cell-targeted DNA vaccine against melanoma by mannosylated liposomes. Biomaterials 2007, 28, 3255–3262. [Google Scholar] [CrossRef] [PubMed]
- Zohra, F.T.; Chowdhury, E.H.; Tada, S.; Hoshiba, T.; Akaike, T. Effective delivery with enhanced translational activity synergistically accelerates mRNA-based transfection. Biochem. Biophys. Res. Commun. 2007, 358, 373–378. [Google Scholar] [CrossRef] [PubMed]
- Michanek, A.; Kristen, N.; Hook, F.; Nylander, T.; Sparr, E. RNA and DNA interactions with zwitterionic and charged lipid membranes - a DSC and QCM-D study. Biochim. Biophys. Acta Biomembr. 2010, 1798, 829–838. [Google Scholar] [CrossRef] [PubMed]
- Du, Z.; Munye, M.M.; Tagalakis, A.D.; Manunta, M.D.; Hart, S.L. The role of the helper lipid on the DNA transfection efficiency of lipopolyplex formulations. Sci. Rep. 2014, 4, 7107. [Google Scholar] [CrossRef] [PubMed]
- Arthur, J.F.; Butterfield, L.H.; Roth, M.D.; Bui, L.A.; Kiertscher, S.M.; Lau, R.; Dubinett, S.; Glaspy, J.; McBride, W.H.; Economou, J.S. A comparison of gene transfer methods in human dendritic cells. Cancer Gene Ther. 1997, 4, 17–25. [Google Scholar]
- Granot, Y.; Peer, D. Delivering the right message: Challenges and opportunities in lipid nanoparticles-mediated modified mRNA therapeutics-An innate immune system standpoint. Semin. Immunol. 2017, 34, 68–77. [Google Scholar] [CrossRef]
- Akbarzadeh, A.; Rezaei-Sadabady, R.; Davaran, S.; Joo, S.W.; Zarghami, N.; Hanifehpour, Y.; Samiei, M.; Kouhi, M.; Nejati-Koshki, K. Liposome: Classification, preparation, and applications. Nanoscale Res. Lett. 2013, 8, 102. [Google Scholar] [CrossRef]
- Wang, Y.; Su, H.H.; Yang, Y.; Hu, Y.; Zhang, L.; Blancafort, P.; Huang, L. Systemic delivery of modified mRNA encoding herpes simplex virus 1 thymidine kinase for targeted cancer gene therapy. Mol. Ther. 2013, 21, 358–367. [Google Scholar] [CrossRef]
- Parnaste, L.; Arukuusk, P.; Langel, K.; Tenson, T.; Langel, U. The Formation of Nanoparticles between Small Interfering RNA and Amphipathic Cell-Penetrating Peptides. Mol. Ther. Nucleic Acids 2017, 7, 1–10. [Google Scholar] [CrossRef]
- Viger-Gravel, J.; Schantz, A.; Pinon, A.C.; Rossini, A.J.; Schantz, S.; Emsley, L. Structure of Lipid Nanoparticles Containing siRNA or mRNA by Dynamic Nuclear Polarization-Enhanced NMR Spectroscopy. J. Phys. Chem. B 2018, 122, 2073–2081. [Google Scholar] [CrossRef] [PubMed]
- Filion, M.C.; Phillips, N.C. Toxicity and immunomodulatory activity of liposomal vectors formulated with cationic lipids toward immune effector cells. Biochim. Biophys. Acta Biomembr. 1997, 1329, 345–356. [Google Scholar] [CrossRef]
- Zhao, Y.N.; Piao, Y.Z.; Zhang, C.M.; Jiang, Y.M.; Liu, A.; Cui, S.H.; Zhi, D.F.; Zhen, Y.H.; Zhang, S.B. Replacement of quaternary ammonium headgroups by tri-ornithine in cationic lipids for the improvement of gene delivery in vitro and in vivo. J. Mater. Chem. B 2017, 5, 7963–7973. [Google Scholar] [CrossRef]
- Landesman-Milo, D.; Peer, D. Toxicity profiling of several common RNAi-based nanomedicines: A comparative study. Drug Deliv. Transl. Res. 2014, 4, 96–103. [Google Scholar] [CrossRef] [PubMed]
- Ma, Z.; Li, J.; He, F.; Wilson, A.; Pitt, B.; Li, S. Cationic lipids enhance siRNA-mediated interferon response in mice. Biochem. Biophys. Res. Commun. 2005, 330, 755–759. [Google Scholar] [CrossRef]
- Platanias, L.C. Mechanisms of type-I- and type-II-interferon-mediated signalling. Nat. Rev. Immunol. 2005, 5, 375–386. [Google Scholar] [CrossRef]
- Lv, H.; Zhang, S.; Wang, B.; Cui, S.; Yan, J. Toxicity of cationic lipids and cationic polymers in gene delivery. J. Control. Release 2006, 114, 100–109. [Google Scholar] [CrossRef]
- Hecker, J.G. Non-Viral, Lipid-Mediated DNA and mRNA Gene Therapy of the Central Nervous System (CNS): Chemical-Based Transfection; Humana Press: New York, NY, USA, 2016; pp. 307–324. [Google Scholar]
- Fenton, O.S.; Kauffman, K.J.; McClellan, R.L.; Appel, E.A.; Dorkin, J.R.; Tibbitt, M.W.; Heartlein, M.W.; DeRosa, F.; Langer, R.; Anderson, D.G. Bioinspired Alkenyl Amino Alcohol Ionizable Lipid Materials for Highly Potent In Vivo mRNA Delivery. Adv. Mater. 2016, 28, 2939–2943. [Google Scholar] [CrossRef]
- de Groot, A.M.; Thanki, K.; Gangloff, M.; Falkenberg, E.; Zeng, X.; van Bijnen, D.C.J.; van Eden, W.; Franzyk, H.; Nielsen, H.M.; Broere, F.; et al. Immunogenicity Testing of Lipidoids In Vitro and In Silico: Modulating Lipidoid-Mediated TLR4 Activation by Nanoparticle Design. Mol. Ther. Nucleic Acids 2018, 11, 159–169. [Google Scholar] [CrossRef]
- McKinlay, C.J.; Benner, N.L.; Haabeth, O.A.; Waymouth, R.M.; Wender, P.A. Enhanced mRNA delivery into lymphocytes enabled by lipid-varied libraries of charge-altering releasable transporters. Proc. Natl. Acad. Sci. USA 2018, 115, E5859–E5866. [Google Scholar] [CrossRef]
- Fenton, O.S.; Kauffman, K.J.; Kaczmarek, J.C.; McClellan, R.L.; Jhunjhunwala, S.; Tibbitt, M.W.; Zeng, M.D.; Appel, E.A.; Dorkin, J.R.; Mir, F.F.; et al. Synthesis and Biological Evaluation of Ionizable Lipid Materials for the In Vivo Delivery of Messenger RNA to B Lymphocytes. Adv. Mater. 2017, 29. [Google Scholar] [CrossRef] [PubMed]
- Phua, K.K.; Nair, S.K.; Leong, K.W. Messenger RNA (mRNA) nanoparticle tumour vaccination. Nanoscale 2014, 6, 7715–7729. [Google Scholar] [CrossRef] [PubMed]
- Kauffman, K.J.; Dorkin, J.R.; Yang, J.H.; Heartlein, M.W.; DeRosa, F.; Mir, F.F.; Fenton, O.S.; Anderson, D.G. Optimization of Lipid Nanoparticle Formulations for mRNA Delivery in Vivo with Fractional Factorial and Definitive Screening Designs. Nano Lett. 2015, 15, 7300–7306. [Google Scholar] [CrossRef]
- Thanki, K.; Zeng, X.; Justesen, S.; Tejlmann, S.; Falkenberg, E.; Van Driessche, E.; Morck Nielsen, H.; Franzyk, H.; Foged, C. Engineering of small interfering RNA-loaded lipidoid-poly(DL-lactic-co-glycolic acid) hybrid nanoparticles for highly efficient and safe gene silencing: A quality by design-based approach. Eur. J. Pharm. Biopharm. 2017, 120, 22–33. [Google Scholar] [CrossRef] [PubMed]
- Jansen, M.A.A.; Klausen, L.H.; Thanki, K.; Lyngso, J.; Skov Pedersen, J.; Franzyk, H.; Nielsen, H.M.; van Eden, W.; Dong, M.; Broere, F.; et al. Lipidoid-polymer hybrid nanoparticles loaded with TNF siRNA suppress inflammation after intra-articular administration in a murine experimental arthritis model. Eur. J. Pharm. Biopharm. 2019, 142, 38–48. [Google Scholar] [CrossRef]
- Zhao, W.; Zhang, C.; Li, B.; Zhang, X.; Luo, X.; Zeng, C.; Li, W.; Gao, M.; Dong, Y. Lipid Polymer Hybrid Nanomaterials for mRNA Delivery. Cell. Mol. Bioeng. 2018, 11, 397–406. [Google Scholar] [CrossRef] [PubMed]
- Kaczmarek, J.C.; Patel, A.K.; Kauffman, K.J.; Fenton, O.S.; Webber, M.J.; Heartlein, M.W.; DeRosa, F.; Anderson, D.G. Polymer-Lipid Nanoparticles for Systemic Delivery of mRNA to the Lungs. Angew Chem. Int. Ed. Engl. 2016, 55, 13808–13812. [Google Scholar] [CrossRef]
- Ball, R.L.; Hajj, K.A.; Vizelman, J.; Bajaj, P.; Whitehead, K.A. Lipid Nanoparticle Formulations for Enhanced Co-delivery of siRNA and mRNA. Nano Lett. 2018, 18, 3814–3822. [Google Scholar] [CrossRef]
- Colombo, S.; Cun, D.; Remaut, K.; Bunker, M.; Zhang, J.; Martin-Bertelsen, B.; Yaghmur, A.; Braeckmans, K.; Nielsen, H.M.; Foged, C. Mechanistic profiling of the siRNA delivery dynamics of lipid-polymer hybrid nanoparticles. J. Control. Release 2015, 201, 22–31. [Google Scholar] [CrossRef]
- van den Brand, D.; Gorris, M.A.J.; van Asbeck, A.H.; Palmen, E.; Ebisch, I.; Dolstra, H.; Hällbrink, M.; Massuger, L.F.A.G.; Brock, R. Peptide-mediated delivery of therapeutic mRNA in ovarian cancer. Eur. J. Pharm. Biopharm. 2019, 141, 180–190. [Google Scholar] [CrossRef]
- Udhayakumar, V.K.; De Beuckelaer, A.; McCaffrey, J.; McCrudden, C.M.; Kirschman, J.L.; Vanover, D.; Van Hoecke, L.; Roose, K.; Deswarte, K.; De Geest, B.G.; et al. Arginine-Rich Peptide-Based mRNA Nanocomplexes Efficiently Instigate Cytotoxic T Cell Immunity Dependent on the Amphipathic Organization of the Peptide. Adv. Healthc. Mater. 2017, 6. [Google Scholar] [CrossRef] [PubMed]
- Kang, Z.; Meng, Q.; Liu, K. Peptide-based gene delivery vectors. J. Mater. Chem. B 2019, 7, 1824–1841. [Google Scholar] [CrossRef]
- Lacroix, C.; Humanes, A.; Coiffier, C.; Gigmes, D.; Verrier, B.; Trimaille, T. Polylactide-Based Reactive Micelles as a Robust Platform for mRNA Delivery. Pharm. Res. 2020, 37, 30. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Dorkin, J.R.; Wang, W.; Chang, P.H.; Webber, M.J.; Tang, B.C.; Yang, J.; Abutbul-Ionita, I.; Danino, D.; DeRosa, F.; et al. Poly(glycoamidoamine) Brushes Formulated Nanomaterials for Systemic siRNA and mRNA Delivery in Vivo. Nano Lett. 2016, 16, 842–848. [Google Scholar] [CrossRef]
- Zhao, M.; Li, M.; Zhang, Z.; Gong, T.; Sun, X. Induction of HIV-1 gag specific immune responses by cationic micelles mediated delivery of gag mRNA. Drug Deliv. 2016, 23, 2596–2607. [Google Scholar] [CrossRef]
- Sunshine, J.C.; Sunshine, S.B.; Bhutto, I.; Handa, J.T.; Green, J.J. Poly(beta-amino ester)-nanoparticle mediated transfection of retinal pigment epithelial cells in vitro and in vivo. PLoS ONE 2012, 7, e37543. [Google Scholar] [CrossRef]
- Islam, M.A.; Reesor, E.K.; Xu, Y.; Zope, H.R.; Zetter, B.R.; Shi, J. Biomaterials for mRNA delivery. Biomater. Sci. 2015, 3, 1519–1533. [Google Scholar] [CrossRef]
- Siewert, C.; Haas, H.; Nawroth, T.; Ziller, A.; Nogueira, S.S.; Schroer, M.A.; Blanchet, C.E.; Svergun, D.I.; Radulescu, A.; Bates, F.; et al. Investigation of charge ratio variation in mRNA - DEAE-dextran polyplex delivery systems. Biomaterials 2019, 192, 612–620. [Google Scholar] [CrossRef]
- Miao, L.; Li, L.; Huang, Y.; Delcassian, D.; Chahal, J.; Han, J.; Shi, Y.; Sadtler, K.; Gao, W.; Lin, J.; et al. Delivery of mRNA vaccines with heterocyclic lipids increases anti-tumor efficacy by STING-mediated immune cell activation. Nat. Biotechnol. 2019, 37, 1174–1185. [Google Scholar] [CrossRef]
- Rybakova, Y.; Kowalski, P.S.; Huang, Y.; Gonzalez, J.T.; Heartlein, M.W.; DeRosa, F.; Delcassian, D.; Anderson, D.G. mRNA Delivery for Therapeutic Anti-HER2 Antibody Expression In Vivo. Mol. Ther. 2019, 27, 1415–1423. [Google Scholar] [CrossRef]
- Sedic, M.; Senn, J.J.; Lynn, A.; Laska, M.; Smith, M.; Platz, S.J.; Bolen, J.; Hoge, S.; Bulychev, A.; Jacquinet, E.; et al. Safety Evaluation of Lipid Nanoparticle-Formulated Modified mRNA in the Sprague-Dawley Rat and Cynomolgus Monkey. Vet. Pathol. 2018, 55, 341–354. [Google Scholar] [CrossRef] [PubMed]
- Persano, S.; Guevara, M.L.; Li, Z.; Mai, J.; Ferrari, M.; Pompa, P.P.; Shen, H. Lipopolyplex potentiates anti-tumor immunity of mRNA-based vaccination. Biomaterials 2017, 125, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Kranz, L.M.; Diken, M.; Haas, H.; Kreiter, S.; Loquai, C.; Reuter, K.C.; Meng, M.; Fritz, D.; Vascotto, F.; Hefesha, H.; et al. Systemic RNA delivery to dendritic cells exploits antiviral defence for cancer immunotherapy. Nature 2016, 534, 396–401. [Google Scholar] [CrossRef]
- Veiga, N.; Goldsmith, M.; Granot, Y.; Rosenblum, D.; Dammes, N.; Kedmi, R.; Ramishetti, S.; Peer, D. Cell specific delivery of modified mRNA expressing therapeutic proteins to leukocytes. Nat. Comm. 2018, 9, 4493. [Google Scholar] [CrossRef]
- Hossain, M.K.; Wall, K.A. Use of Dendritic Cell Receptors as Targets for Enhancing Anti-Cancer Immune Responses. Cancers 2019, 11, 418. [Google Scholar] [CrossRef]
- Midoux, P.; Pichon, C. Lipid-based mRNA vaccine delivery systems. Expert Rev. Vaccines 2015, 14, 221–234. [Google Scholar] [CrossRef]
- Wang, F.; Xiao, W.; Elbahnasawy, M.A.; Bao, X.; Zheng, Q.; Gong, L.; Zhou, Y.; Yang, S.; Fang, A.; Farag, M.M.S.; et al. Optimization of the Linker Length of Mannose-Cholesterol Conjugates for Enhanced mRNA Delivery to Dendritic Cells by Liposomes. Front. Pharmacol. 2018, 9, 980. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).