Bacterial Virus Ontology; Coordinating across Databases
Abstract
:1. Introduction
2. Materials and Methods
2.1. Creation of the Bacterial Virus Vocabulary and ViralZone Pages
2.2. Mapping of Viral Life-Cycle Processes to GO
2.3. Creation of New UniProtKB Keywords
3. Results
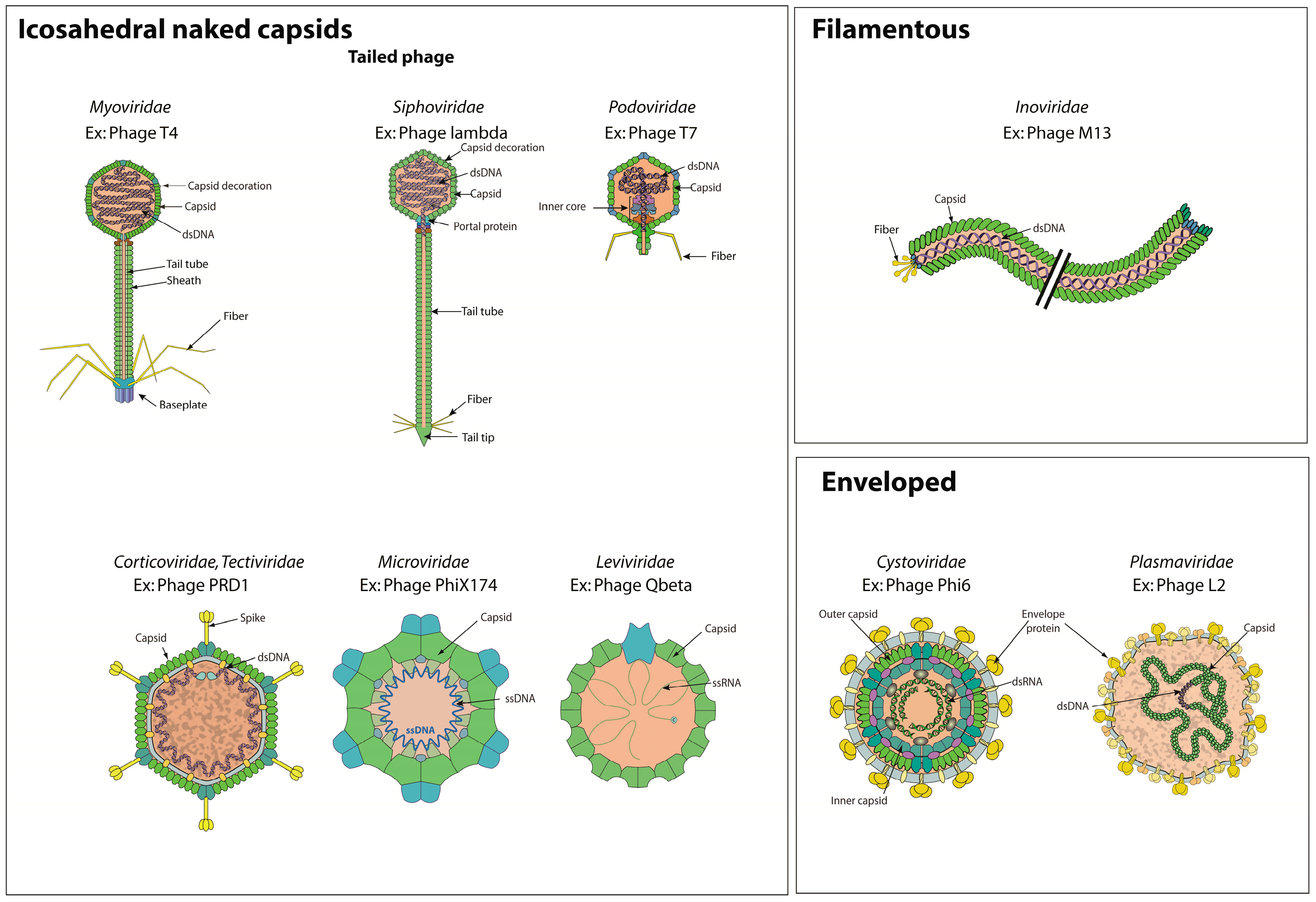
3.1. Bacterial Virions
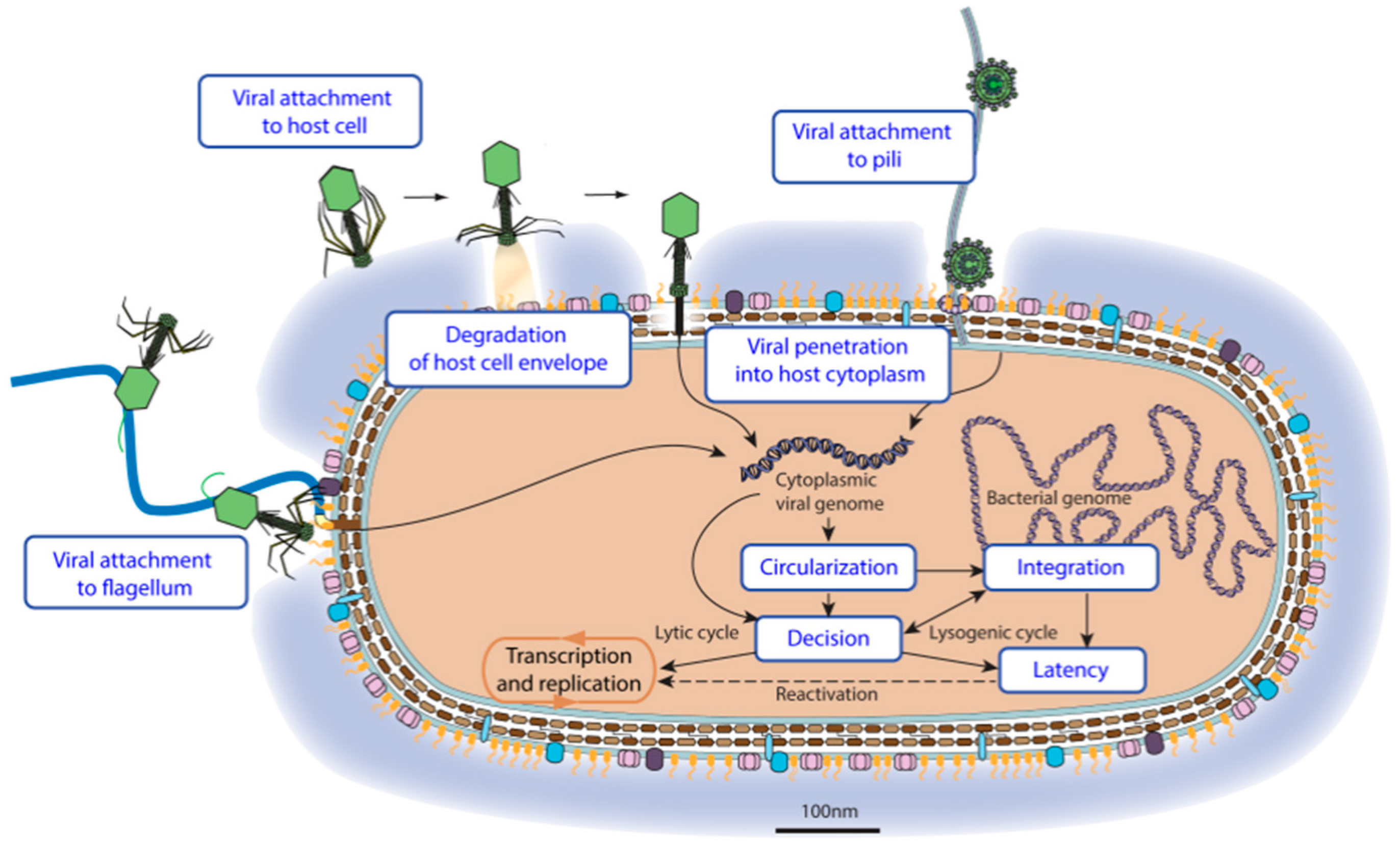
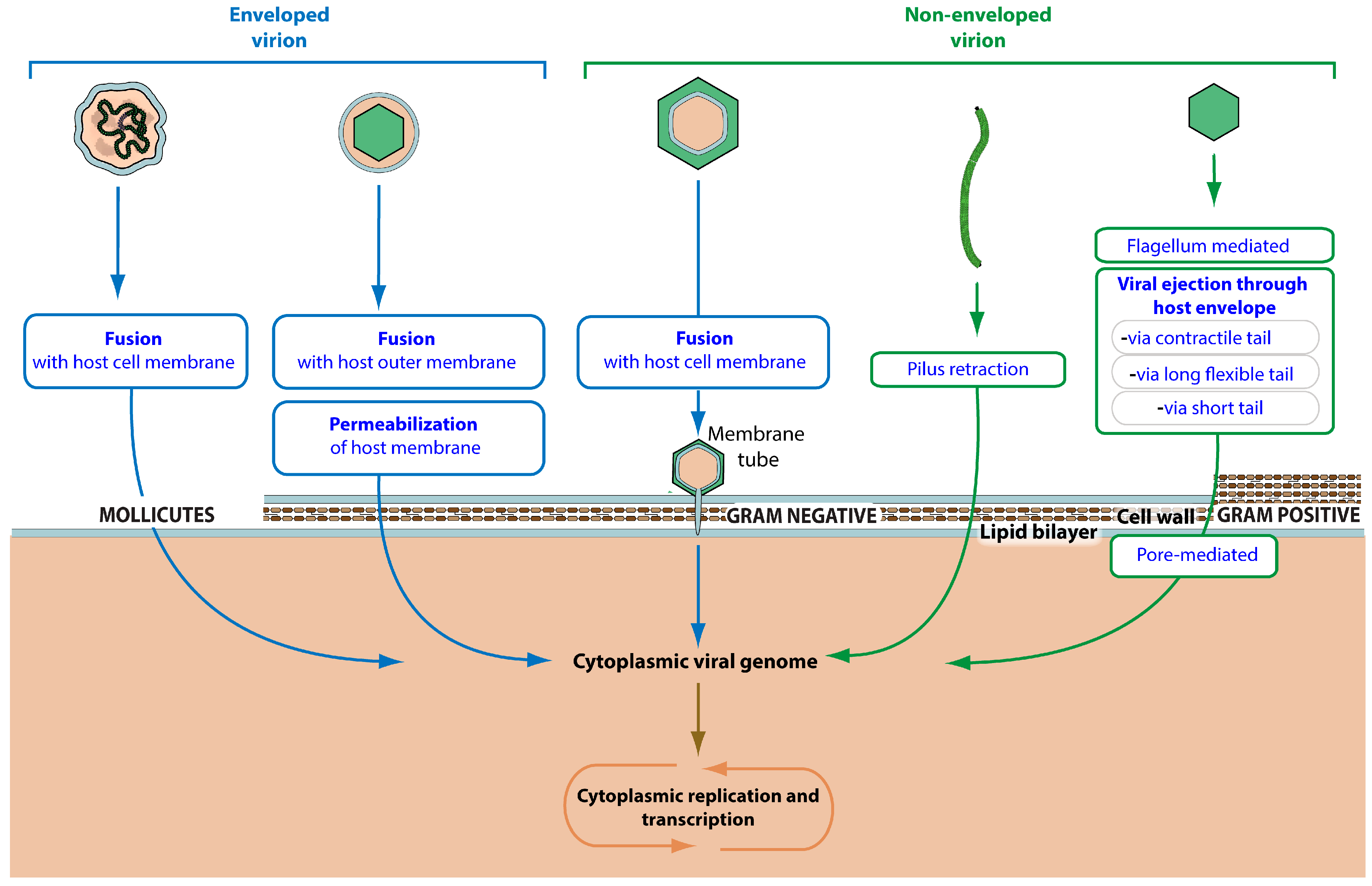
3.2. Virus Entry
3.3. Latency
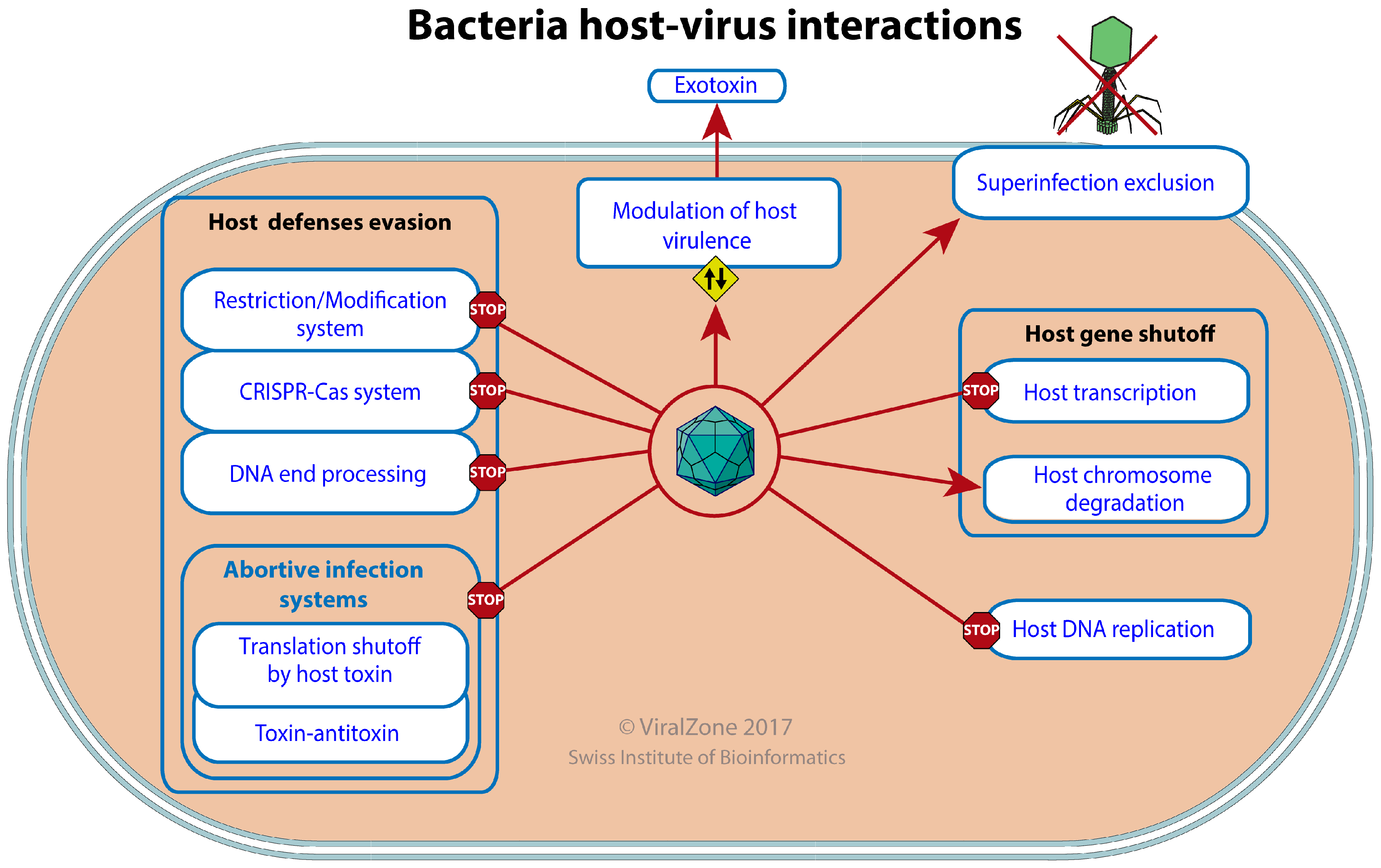
3.4. Host-Virus Interactions
3.5. Viral Replication
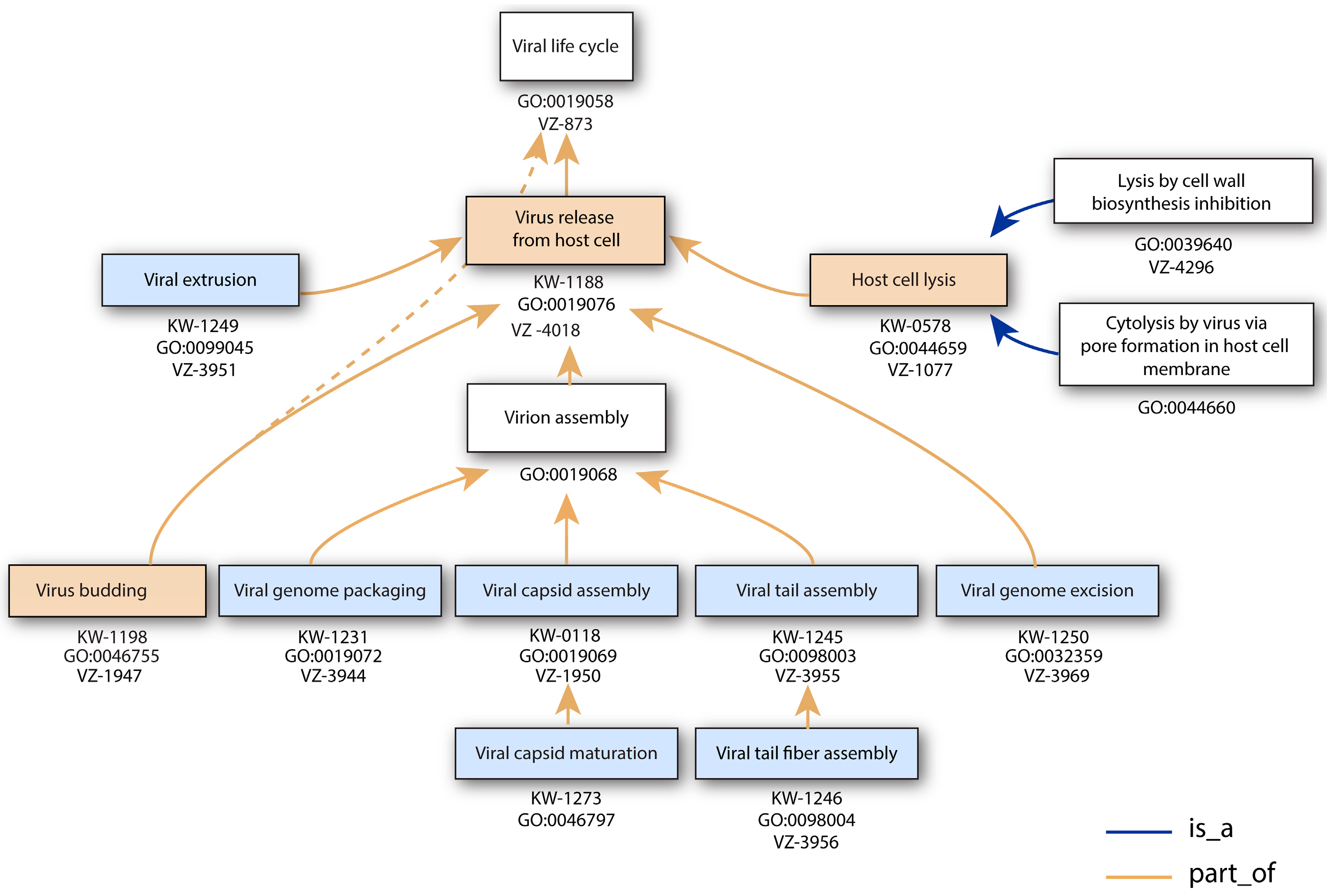
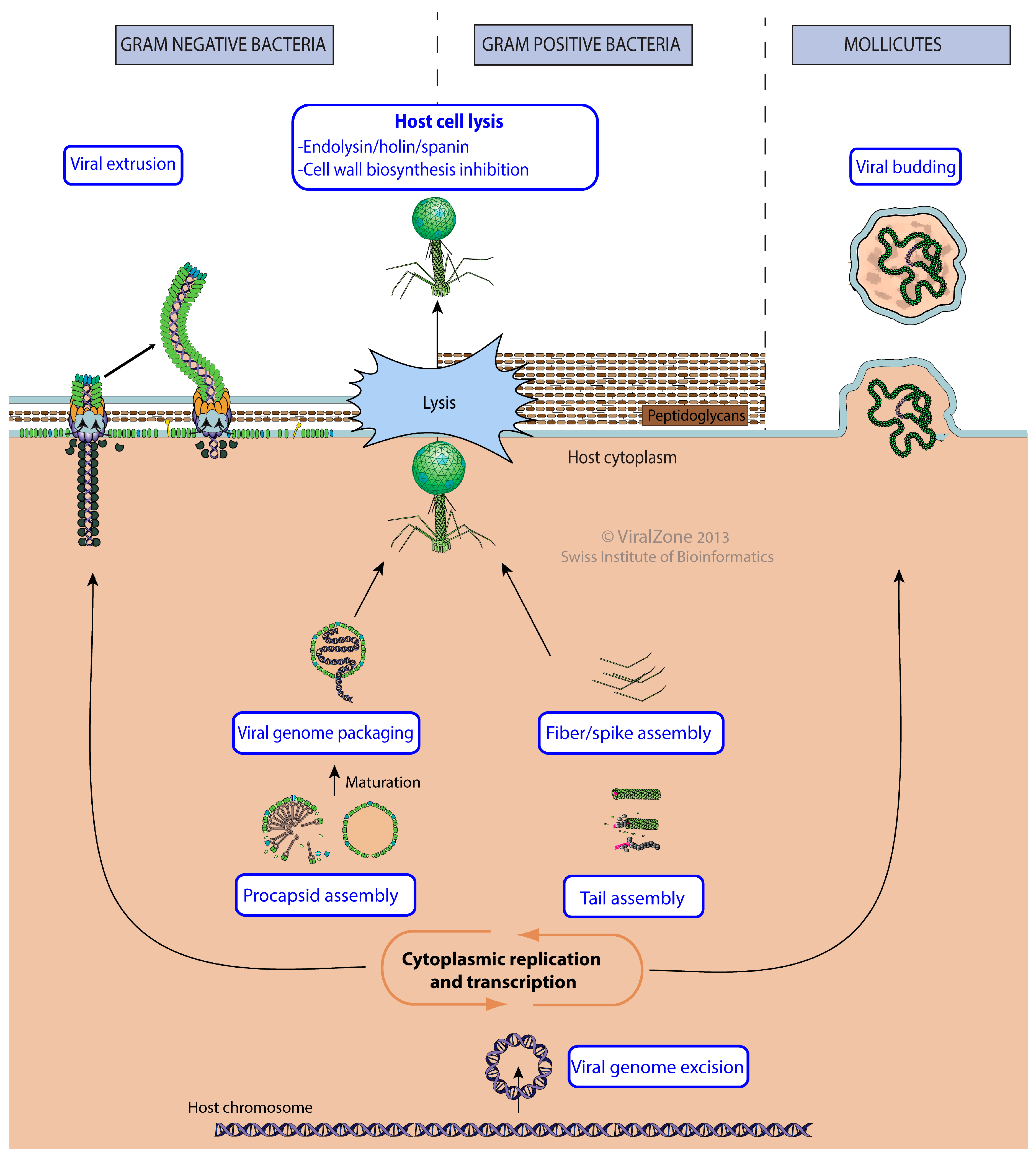
3.6. Virus Release from Host Cell
4. Discussion
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Hershey, A.D.; Chase, M. Independent functions of viral protein and nucleic acid in growth of bacteriophage. J. Gen. Physiol. 1952, 36, 39–56. [Google Scholar] [CrossRef] [PubMed]
- Haq, I.U.; Chaudhry, W.N.; Akhtar, M.N.; Andleeb, S.; Qadri, I. Bacteriophages and their implications on future biotechnology: A review. Virol. J. 2012, 9, 9. [Google Scholar] [CrossRef] [PubMed]
- Henry, M.; Debarbieux, L. Tools from viruses: Bacteriophage successes and beyond. Virology 2012, 434, 151–161. [Google Scholar] [CrossRef] [PubMed]
- Sharan, S.K.; Thomason, L.C.; Kuznetsov, S.G.; Court, D.L. Recombineering: A homologous recombination-based method of genetic engineering. Nat. Protoc. 2009, 4, 206–223. [Google Scholar] [CrossRef] [PubMed]
- Cisek, A.A.; Dąbrowska, I.; Gregorczyk, K.P.; Wyżewski, Z. Phage Therapy in Bacterial Infections Treatment: One Hundred Years after the Discovery of Bacteriophages. Curr. Microbiol. 2017, 74, 277–283. [Google Scholar] [CrossRef] [PubMed]
- Drulis-Kawa, Z.; Majkowska-Skrobek, G.; Maciejewska, B.; Delattre, A.-S.; Lavigne, R. Learning from bacteriophages-advantages and limitations of phage and phage-encoded protein applications. Curr. Protein Pept. Sci. 2012, 13, 699–722. [Google Scholar] [CrossRef] [PubMed]
- Snyder, J.C.; Bolduc, B.; Young, M.J. 40 Years of archaeal virology: Expanding viral diversity. Virology 2015, 479–480, 369–378. [Google Scholar] [CrossRef] [PubMed]
- Mann, N.H. The third age of phage. PLoS Biol. 2005, 3, e182. [Google Scholar] [CrossRef] [PubMed]
- Wagner, P.L.; Waldor, M.K. Bacteriophage control of bacterial virulence. Infect. Immun. 2002, 70, 3985–3993. [Google Scholar] [CrossRef] [PubMed]
- Jover, L.F.; Effler, T.C.; Buchan, A.; Wilhelm, S.W.; Weitz, J.S. The elemental composition of virus particles: Implications for marine biogeochemical cycles. Nat. Rev. Microbiol. 2014, 12, 519–528. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Valera, F.; Martin-Cuadrado, A.-B.; Rodriguez-Brito, B.; Pasić, L.; Thingstad, T.F.; Rohwer, F.; Mira, A. Explaining microbial population genomics through phage predation. Nat. Rev. Microbiol. 2009, 7, 828–836. [Google Scholar] [CrossRef] [PubMed]
- Nelson, D. Phage taxonomy: We agree to disagree. J. Bacteriol. 2004, 186, 7029–7031. [Google Scholar] [CrossRef] [PubMed]
- Hulo, C.; de Castro, E.; Masson, P.; Bougueleret, L.; Bairoch, A.; Xenarios, I.; Le Mercier, P. ViralZone: A knowledge resource to understand virus diversity. Nucleic Acids Res. 2011, 39, 576–582. [Google Scholar] [CrossRef] [PubMed]
- UniProt Consortium. UniProt: A hub for protein information. Nucleic Acids Res. 2015, 43, 204–212. [Google Scholar]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene ontology: Tool for the unification of biology. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Gene Ontology Consortium. The Gene Ontology in 2010: Extensions and refinements. Nucleic Acids Res. 2010, 38, D331–D335. [Google Scholar]
- Masson, P.; Hulo, C.; de Castro, E.; Foulger, R.; Poux, S.; Bridge, A.; Lomax, J.; Bougueleret, L.; Xenarios, I.; Le Mercier, P. An integrated ontology resource to explore and study host-virus relationships. PLoS ONE 2014, 9, e108075. [Google Scholar] [CrossRef] [PubMed]
- Calendar, R. The Bacteriophages, 2th ed.; Oxford University Press: Oxford, UK, 2006. [Google Scholar]
- Toussaint, A.; Lima-Mendez, G.; Leplae, R. PhiGO, a phage ontology associated with the ACLAME database. Res. Microbiol. 2007, 158, 567–571. [Google Scholar] [CrossRef] [PubMed]
- Leiman, P.G.; Shneider, M.M. Contractile tail machines of bacteriophages. Adv. Exp. Med. Biol. 2012, 726, 93–114. [Google Scholar] [PubMed]
- Casjens, S.R.; Molineux, I.J. Short noncontractile tail machines: Adsorption and DNA delivery by podoviruses. Adv. Exp. Med. Biol. 2012, 726, 143–179. [Google Scholar] [PubMed]
- Davidson, A.R.; Cardarelli, L.; Pell, L.G.; Radford, D.R.; Maxwell, K.L. Long noncontractile tail machines of bacteriophages. Adv. Exp. Med. Biol. 2012, 726, 115–142. [Google Scholar] [PubMed]
- Büttner, C.R.; Wu, Y.; Maxwell, K.L.; Davidson, A.R. Baseplate assembly of phage Mu: Defining the conserved core components of contractile-tailed phages and related bacterial systems. Proc. Natl. Acad. Sci. USA 2016, 113, 10174–10179. [Google Scholar] [CrossRef] [PubMed]
- Taylor, N.M.I.; Prokhorov, N.S.; Guerrero-Ferreira, R.C.; Shneider, M.M.; Browning, C.; Goldie, K.N.; Stahlberg, H.; Leiman, P.G. Structure of the T4 baseplate and its function in triggering sheath contraction. Nature 2016, 533, 346–352. [Google Scholar] [CrossRef] [PubMed]
- Holland, S.J.; Sanz, C.; Perham, R.N. Identification and specificity of pilus adsorption proteins of filamentous bacteriophages infecting Pseudomonas aeruginosa. Virology 2006, 345, 540–548. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Shin, H.; Lee, J.-H.; Ryu, S. Identification and characterization of a novel flagellum-dependent Salmonella-infecting bacteriophage, iEPS5. Appl. Environ. Microbiol. 2013, 79, 4829–4837. [Google Scholar] [CrossRef] [PubMed]
- Grahn, A.M.; Daugelavicius, R.; Bamford, D.H. Sequential model of phage PRD1 DNA delivery: Active involvement of the viral membrane. Mol. Microbiol. 2002, 46, 1199–1209. [Google Scholar] [CrossRef] [PubMed]
- Rakonjac, J.; Bennett, N.J.; Spagnuolo, J.; Gagic, D.; Russel, M. Filamentous bacteriophage: Biology, phage display and nanotechnology applications. Curr. Issues Mol. Biol. 2011, 13, 51–76. [Google Scholar] [PubMed]
- Puspurs, A.H.; Trun, N.J.; Reeve, J.N. Bacteriophage Mu DNA circularizes following infection of Escherichia coli. EMBO J. 1983, 2, 345–352. [Google Scholar] [PubMed]
- Mardanov, A.V.; Ravin, N.V. Conversion of linear DNA with hairpin telomeres into a circular molecule in the course of phage N15 lytic replication. J. Mol. Biol. 2009, 391, 261–268. [Google Scholar] [CrossRef] [PubMed]
- Roldan, L.A.; Baker, T.A. Differential role of the Mu B protein in phage Mu integration vs. replication: Mechanistic insights into two transposition pathways. Mol. Microbiol. 2001, 40, 141–155. [Google Scholar] [CrossRef] [PubMed]
- Farruggio, A.P.; Chavez, C.L.; Mikell, C.L.; Calos, M.P. Efficient reversal of phiC31 integrase recombination in mammalian cells. Biotechnol. J. 2012, 7, 1332–1336. [Google Scholar] [CrossRef] [PubMed]
- Pope, W.H.; Bowman, C.A.; Russell, D.A.; Jacobs-Sera, D.; Asai, D.J.; Cresawn, S.G.; Jacobs, W.R.; Hendrix, R.W.; Lawrence, J.G.; Hatfull, G.F. Whole genome comparison of a large collection of mycobacteriophages reveals a continuum of phage genetic diversity. eLife 2015, 4, e06416. [Google Scholar] [CrossRef] [PubMed]
- Labrie, S.J.; Samson, J.E.; Moineau, S. Bacteriophage resistance mechanisms. Nat. Rev. Microbiol. 2010, 8, 317–327. [Google Scholar] [CrossRef] [PubMed]
- Samson, J.E.; Magadán, A.H.; Sabri, M.; Moineau, S. Revenge of the phages: Defeating bacterial defences. Nat. Rev. Microbiol. 2013, 11, 675–687. [Google Scholar] [CrossRef] [PubMed]
- Roberts, R.J. Restriction and modification enzymes and their recognition sequences. Nucleic Acids Res. 1981, 9, 167–204. [Google Scholar] [CrossRef]
- Studier, F.W. Gene 0.3 of bacteriophage T7 acts to overcome the DNA restriction system of the host. J. Mol. Biol. 1975, 94, 283–295. [Google Scholar] [CrossRef]
- Berkner, K.L.; Folk, W.R. The effects of substituted pyrimidines in DNAs on cleavage by sequence-specific endonucleases. J. Biol. Chem. 1979, 254, 2551–2560. [Google Scholar] [PubMed]
- Dillingham, M.S.; Kowalczykowski, S.C. RecBCD enzyme and the repair of double-stranded DNA breaks. Microbiol. Mol. Biol. Rev. MMBR 2008, 72, 642–671. [Google Scholar] [CrossRef] [PubMed]
- Lipinska, B.; Rao, A.S.; Bolten, B.M.; Balakrishnan, R.; Goldberg, E.B. Cloning and identification of bacteriophage T4 gene 2 product Gp2 and action of Gp2 on infecting DNA in vivo. J. Bacteriol. 1989, 171, 488–497. [Google Scholar] [CrossRef] [PubMed]
- Murphy, K.C. The lambda Gam protein inhibits RecBCD binding to dsDNA ends. J. Mol. Biol. 2007, 371, 19–24. [Google Scholar] [CrossRef] [PubMed]
- Chopin, M.-C.; Chopin, A.; Bidnenko, E. Phage abortive infection in lactococci: Variations on a theme. Curr. Opin. Microbiol. 2005, 8, 473–479. [Google Scholar] [CrossRef] [PubMed]
- Otsuka, Y.; Yonesaki, T. Dmd of bacteriophage T4 functions as an antitoxin against Escherichia coli LsoA and RnlA toxins. Mol. Microbiol. 2012, 83, 669–681. [Google Scholar] [CrossRef] [PubMed]
- Barrangou, R.; Marraffini, L.A. CRISPR-Cas systems: Prokaryotes upgrade to adaptive immunity. Mol. Cell 2014, 54, 234–244. [Google Scholar] [CrossRef] [PubMed]
- Bondy-Denomy, J.; Pawluk, A.; Maxwell, K.L.; Davidson, A.R. Bacteriophage genes that inactivate the CRISPR/Cas bacterial immune system. Nature 2013, 493, 429–432. [Google Scholar] [CrossRef] [PubMed]
- Seed, K.D.; Lazinski, D.W.; Calderwood, S.B.; Camilli, A. A bacteriophage encodes its own CRISPR/Cas adaptive response to evade host innate immunity. Nature 2013, 494, 489–491. [Google Scholar] [CrossRef] [PubMed]
- Bae, B.; Davis, E.; Brown, D.; Campbell, E.A.; Wigneshweraraj, S.; Darst, S.A. Phage T7 Gp2 inhibition of Escherichia coli RNA polymerase involves misappropriation of σ70 domain 1.1. Proc. Natl. Acad. Sci. USA 2013, 110, 19772–19777. [Google Scholar] [CrossRef] [PubMed]
- Souther, A.; Bruner, R.; Elliott, J. Degradation of Escherichia coli chromosome after infection by bacteriophage T4: Role of bacteriophage gene D2a. J. Virol. 1972, 10, 979–984. [Google Scholar] [PubMed]
- Powell, I.B.; Tulloch, D.L.; Hillier, A.J.; Davidson, B.E. Phage DNA synthesis and host DNA degradation in the life cycle of Lactococcus lactis bacteriophage c6A. J. Gen. Microbiol. 1992, 138, 945–950. [Google Scholar] [CrossRef] [PubMed]
- Yano, S.T.; Rothman-Denes, L.B. A phage-encoded inhibitor of Escherichia coli DNA replication targets the DNA polymerase clamp loader. Mol. Microbiol. 2011, 79, 1325–1338. [Google Scholar] [CrossRef] [PubMed]
- Belley, A.; Callejo, M.; Arhin, F.; Dehbi, M.; Fadhil, I.; Liu, J.; McKay, G.; Srikumar, R.; Bauda, P.; Bergeron, D.; et al. Competition of bacteriophage polypeptides with native replicase proteins for binding to the DNA sliding clamp reveals a novel mechanism for DNA replication arrest in Staphylococcus aureus. Mol. Microbiol. 2006, 62, 1132–1143. [Google Scholar] [CrossRef] [PubMed]
- Bondy-Denomy, J.; Qian, J.; Westra, E.R.; Buckling, A.; Guttman, D.S.; Davidson, A.R.; Maxwell, K.L. Prophages mediate defense against phage infection through diverse mechanisms. ISME J. 2016, 10, 2854–2866. [Google Scholar] [CrossRef] [PubMed]
- Berngruber, T.W.; Weissing, F.J.; Gandon, S. Inhibition of superinfection and the evolution of viral latency. J. Virol. 2010, 84, 10200–10208. [Google Scholar] [CrossRef] [PubMed]
- Braun, V.; Killmann, H.; Herrmann, C. Inactivation of FhuA at the cell surface of Escherichia coli K-12 by a phage T5 lipoprotein at the periplasmic face of the outer membrane. J. Bacteriol. 1994, 176, 4710–4717. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.J.; Henning, U. Superinfection exclusion by T-even-type coliphages. Trends Microbiol. 1994, 2, 137–139. [Google Scholar] [CrossRef]
- Abedon, S.T.; Lejeune, J.T. Why bacteriophage encode exotoxins and other virulence factors. Evol. Bioinform. Online 2007, 1, 97–110. [Google Scholar] [PubMed]
- Boyd, E.F.; Brüssow, H. Common themes among bacteriophage-encoded virulence factors and diversity among the bacteriophages involved. Trends Microbiol. 2002, 10, 521–529. [Google Scholar] [CrossRef]
- Mann, N.H.; Clokie, M.R.J.; Millard, A.; Cook, A.; Wilson, W.H.; Wheatley, P.J.; Letarov, A.; Krisch, H.M. The genome of S-PM2, a “photosynthetic” T4-type bacteriophage that infects marine Synechococcus strains. J. Bacteriol. 2005, 187, 3188–3200. [Google Scholar] [CrossRef] [PubMed]
- Maluf, N.K.; Gaussier, H.; Bogner, E.; Feiss, M.; Catalano, C.E. Assembly of bacteriophage lambda terminase into a viral DNA maturation and packaging machine. Biochemistry 2006, 45, 15259–15268. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Kottadiel, V.I.; Vafabakhsh, R.; Dai, L.; Chemla, Y.R.; Ha, T.; Rao, V.B. A promiscuous DNA packaging machine from bacteriophage T4. PLoS Biol. 2011, 9, e1000592. [Google Scholar] [CrossRef] [PubMed]
- Narajczyk, M.; Barańska, S.; Wegrzyn, A.; Wegrzyn, G. Switch from theta to sigma replication of bacteriophage lambda DNA: Factors involved in the process and a model for its regulation. Mol. Genet. Genom. 2007, 278, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Salas, M.; Holguera, I.; Redrejo-Rodríguez, M.; de Vega, M. DNA-binding proteins essential for protein-primed bacteriophage Φ29 DNA replication. Front. Mol. Biosci. 2016, 3, 37. [Google Scholar] [CrossRef] [PubMed]
- Hulo, C.; Masson, P.; Le Mercier, P.; Toussaint, A. A structured annotation frame for the transposable phages: A new proposed family “Saltoviridae” within the Caudovirales. Virology 2015, 477, 155–163. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, J.A. Molecular model for the transposition and replication of bacteriophage Mu and other transposable elements. Proc. Natl. Acad. Sci. USA 1979, 76, 1933–1937. [Google Scholar] [CrossRef] [PubMed]
- Lundström, K.H.; Bamford, D.H.; Palva, E.T.; Lounatmaa, K. Lipid-containing bacteriophage PR4: Structure and life cycle. J. Gen. Virol. 1979, 43, 583–592. [Google Scholar] [CrossRef] [PubMed]
- Johnson, M.D.; Mindich, L. Plasmid-directed assembly of the lipid-containing membrane of bacteriophage phi 6. J. Bacteriol. 1994, 176, 4124–4132. [Google Scholar] [CrossRef] [PubMed]
- Young, R. Phage lysis: Three steps, three choices, one outcome. J. Microbiol. Seoul Korea 2014, 52, 243–258. [Google Scholar] [CrossRef] [PubMed]
- Young, R. Phage lysis: Do we have the hole story yet? Curr. Opin. Microbiol. 2013, 16, 790–797. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Struck, D.K.; Young, R. Purification and functional characterization of phiX174 lysis protein E. Biochemistry 2009, 48, 4999–5006. [Google Scholar] [CrossRef] [PubMed]
- Karnik, S.; Billeter, M. The lysis function of RNA bacteriophage Qbeta is mediated by the maturation (A2) protein. EMBO J. 1983, 2, 1521–1526. [Google Scholar] [PubMed]
- Marvin, D.A.; Symmons, M.F.; Straus, S.K. Structure and assembly of filamentous bacteriophages. Prog. Biophys. Mol. Biol. 2014, 114, 80–122. [Google Scholar] [CrossRef] [PubMed]
- Poddar, S.K.; Cadden, S.P.; Das, J.; Maniloff, J. Heterogeneous progeny viruses are produced by a budding enveloped phage. Intervirology 1985, 23, 208–221. [Google Scholar] [CrossRef] [PubMed]
- Burge, S.; Kelly, E.; Lonsdale, D.; Mutowo-Muellenet, P.; McAnulla, C.; Mitchell, A.; Sangrador-Vegas, A.; Yong, S.-Y.; Mulder, N.; Hunter, S. Manual GO annotation of predictive protein signatures: The InterPro approach to GO curation. Database J. Biol. Databases Curation 2012, 2012. [Google Scholar] [CrossRef] [PubMed]
- Pedruzzi, I.; Rivoire, C.; Auchincloss, A.H.; Coudert, E.; Keller, G.; de Castro, E.; Baratin, D.; Cuche, B.A.; Bougueleret, L.; Poux, S.; et al. HAMAP in 2015: Updates to the protein family classification and annotation system. Nucleic Acids Res. 2015, 43, D1064–D1070. [Google Scholar] [CrossRef] [PubMed]
- Delattre, H.; Souiai, O.; Fagoonee, K.; Guerois, R.; Petit, M.-A. Phagonaute: A web-based interface for phage synteny browsing and protein function prediction. Virology 2016, 496, 42–50. [Google Scholar] [CrossRef] [PubMed]
- Category: CACAO-GONUTS. Available online: https://gowiki.tamu.edu/wiki/index.php/Category:CACAO (accessed on 16 May 2017).
| UniProt Keywords | GO Terms | UniProt KW | ViralZone Pages | UniProt Entries | |
|---|---|---|---|---|---|
| Virion | GO:0019012 | KW-0946 | * | VZ-885 | 457 |
| Tailed Bacterial virus | VZ-4076 | NA | |||
| Capsid protein | GO:0046728 | KW-0167 | * | 166 | |
| Capsid decoration protein | GO:0098021 | KW-1232 | 24 | ||
| Viral tail protein | GO:0098015 | KW-1227 | 138 | ||
| Viral tail sheath protein | GO:0098027 | KW-1229 | 11 | ||
| Viral tail tube protein | GO:0098026 | KW-1228 | 21 | ||
| Viral baseplate protein | GO:0098025 | KW-1226 | 33 | ||
| Viral tail fiber protein | GO:0098024 | KW-1230 | 42 | ||
| Capsid inner membrane protein | GO:0039641 | KW-1231 | 15 | ||
| Virus Entry into Host Cell | GO:0046718 | KW-1160 | * | VZ-3996 | 296 |
| Viral attachment to host cell | GO:0019062 | KW-1161 | * | VZ-956 | 72 |
| Viral attachment to host adhesion receptor | GO:0098671 | KW-1233 | * | VZ-3943 | 29 |
| Viral attachment to host entry receptor | GO:0098670 | KW-1234 | * | VZ-3942 | 16 |
| Viral attachment to host cell pilus | GO:0039666 | KW-1175 | VZ-981 | 15 | |
| Viral attachment to host cell flagellum | GO:0098931 | KW-1240 | VZ-3949 | 0 | |
| Degradation of host cell envelope components during virus entry | GO:0098994 | KW-1235 | VZ-3938 | 29 | |
| Degradation of host peptidoglycans during virus entry | GO:0098932 | KW-1236 | VZ-3940 | 19 | |
| Degradation of host lipopolysaccharides during virus entry | GO:0098995 | KW-1237 | VZ-3939 | 3 | |
| Degradation of host capsule during virus entry | GO:0098996 | KW-1238 | VZ-3896 | 4 | |
| Viral penetration into host cytoplasm | GO:0046718 | KW-1162 | * | VZ-4016 | 161 |
| Fusion of viral membrane with host outer membrane | GO:0098997 | KW-1239 | VZ-3941 | 1 | |
| Pore-mediated penetration of viral genome into host cell | GO:0044694 | KW-1172 | * | VZ-979 | 7 |
| Viral genome ejection through host cell envelope | GO:0039678 | KW-1171 | VZ-986 | 130 | |
| Viral contractile tail ejection system | GO:0099000 | KW-1242 | VZ-3950 | 30 | |
| Viral long flexible tail ejection system | GO:0099001 | KW-1243 | VZ-3952 | 41 | |
| Viral short tail ejection system | GO:0099002 | KW-1244 | VZ-3954 | 32 | |
| Viral penetration into host cell via pilus retraction | GO:0039667 | KW-1241 | VZ-3953 | 17 | |
| Viral penetration via permeabilization of host membrane | GO:0099008 | KW-1173 | * | VZ-985 | 0 |
| Viral genome circularization | GO:0099009 | KW-1253 | VZ-3968 | 8 | |
| Viral genome integration | GO:0044826 | KW-1179 | * | VZ-980 | 14 |
| Viral receptor tropism switching | GO:0098678 | KW-1264 | VZ-4498 | 10 | |
| Viral Latency | GO:0019042 | KW-1251 | * | VZ-3970 | 6 |
| Latency-replication decision | GO:0098689 | KW-1252 | VZ-3964 | 4 | |
| Viral reactivation from latency | GO:0019046 | KW-1272 | 35 | ||
| Host–Virus Interaction | GO:0019048 | KW-0945 | * | VZ-3756 | 154 |
| Host defense evasion | GO:0044413 | 0 | |||
| Restriction-modification system evasion by virus | GO:0099018 | KW-1258 | VZ-3966 | 16 | |
| CRISPR-Cas system evasion by virus | GO:0098672 | KW-1257 | VZ-3962 | 3 | |
| DNA end degradation evasion by virus | GO:0099016 | KW-1256 | VZ-3963 | 6 | |
| Evasion of bacteria-mediated translation shutoff by virus | KW-1259 | VZ-3961 | 3 | ||
| Evasion of toxin-antitoxin system | VZ-4077 | 0 | |||
| Host gene expression shutoff by virus | GO:0039657 | KW-1190 | * | 12 | |
| Bacterial host gene expression shutoff by virus | KW-1261 | VZ-4496 | 12 | ||
| Bacterial host transcription shutoff by virus | KW-1263 | VZ-4497 | 4 | ||
| Degradation of host chromosome by virus | GO:0099015 | KW-1247 | VZ-3947 | 8 | |
| Inhibition of host DNA replication by virus | GO:0098673 | KW-1248 | VZ-3948 | 9 | |
| Modulation of host virulence by virus | GO:0098676 | KW-1254 | * | VZ-3965 | 9 |
| Viral exotoxin | KW-1255 | VZ-3967 | 9 | ||
| Superinfection exclusion | GO:0098669 | KW-1260 | VZ-3971 | 3 | |
| Viral Replication | * | VZ-915 | NA | ||
| Viral DNA replication | GO:0039693 | KW-0235 | * | 65 | |
| dsDNA bidirectional replication | * | VZ-1939 | 0 | ||
| dsDNA rolling circle replication | * | VZ-2676 | 0 | ||
| DNA strand displacement replication | * | VZ-1940 | 0 | ||
| Replicative transposition | * | VZ-4017 | 0 | ||
| Viral RNA replication | GO:0039694 | KW-0693 | * | 8 | |
| Virus Release from Host Cell | GO:0019076 | KW-1188 | * | VZ-4018 | 322 |
| Viral genome packaging | GO:0019072 | KW-1231 | * | VZ-3944 | 15 |
| Host cell lysis by virus | GO:0044659 | KW-0578 | * | VZ-1077 | 79 |
| Lysis by cell wall biosynthesis inhibition | GO:0039640 | VZ-4296 | 0 | ||
| Cytolysis by virus via pore formation in host cell membrane | GO:0044660 | * | 223 | ||
| Holin/endolysin/spanin cell lysis by virus | VZ-4056 | 0 | |||
| Viral extrusion | GO:0099045 | KW-1249 | VZ-3951 | 22 | |
| Viral genome excision | GO:0032359 | KW-1250 | VZ-3969 | 20 | |
| Viral capsid assembly | GO:0019069 | KW-0118 | * | VZ-1950 | 85 |
| Viral capsid maturation | GO:0046797 | KW-1273 | 0 | ||
| Viral budding | GO:0046755 | KW-1198 | * | VZ-1947 | 0 |
| Viral tail assembly | GO:0098003 | KW-1245 | VZ-3955 | 90 | |
| Viral tail fiber assembly | GO:0098004 | KW-1246 | VZ-3956 | 9 | |
| TOTAL | 3072 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hulo, C.; Masson, P.; Toussaint, A.; Osumi-Sutherland, D.; De Castro, E.; Auchincloss, A.H.; Poux, S.; Bougueleret, L.; Xenarios, I.; Le Mercier, P. Bacterial Virus Ontology; Coordinating across Databases. Viruses 2017, 9, 126. https://doi.org/10.3390/v9060126
Hulo C, Masson P, Toussaint A, Osumi-Sutherland D, De Castro E, Auchincloss AH, Poux S, Bougueleret L, Xenarios I, Le Mercier P. Bacterial Virus Ontology; Coordinating across Databases. Viruses. 2017; 9(6):126. https://doi.org/10.3390/v9060126
Chicago/Turabian StyleHulo, Chantal, Patrick Masson, Ariane Toussaint, David Osumi-Sutherland, Edouard De Castro, Andrea H. Auchincloss, Sylvain Poux, Lydie Bougueleret, Ioannis Xenarios, and Philippe Le Mercier. 2017. "Bacterial Virus Ontology; Coordinating across Databases" Viruses 9, no. 6: 126. https://doi.org/10.3390/v9060126
APA StyleHulo, C., Masson, P., Toussaint, A., Osumi-Sutherland, D., De Castro, E., Auchincloss, A. H., Poux, S., Bougueleret, L., Xenarios, I., & Le Mercier, P. (2017). Bacterial Virus Ontology; Coordinating across Databases. Viruses, 9(6), 126. https://doi.org/10.3390/v9060126






