The Heterologous Expression of the p22 RNA Silencing Suppressor of the Crinivirus Tomato Chlorosis Virus from Tobacco Rattle Virus and Potato Virus X Enhances Disease Severity but Does Not Complement Suppressor-Defective Mutant Viruses
Abstract
1. Introduction
2. Materials and Methods
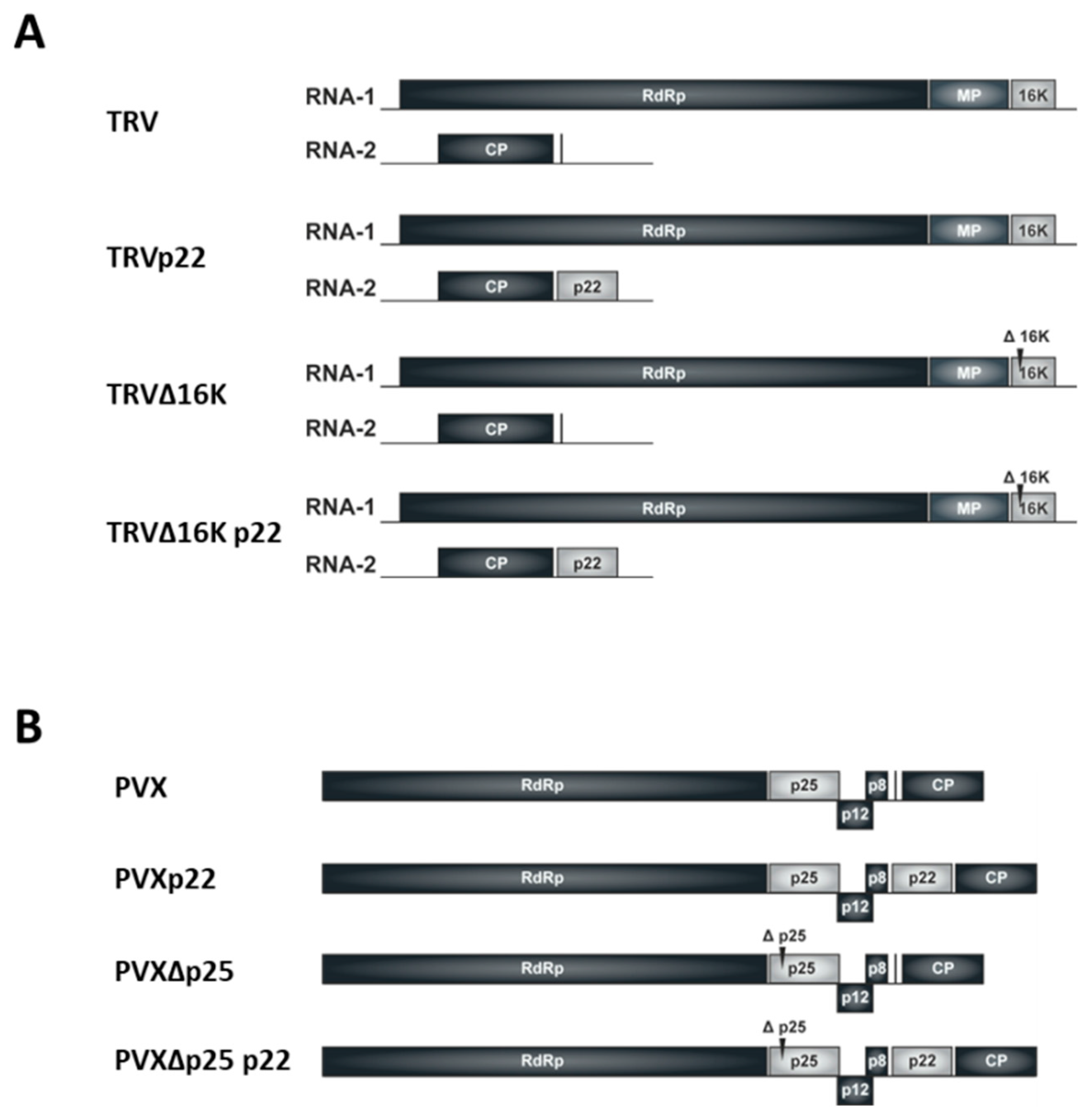
2.1. Plasmid Constructs
2.2. Plant Material and Agroinfiltration
2.3. RNA Analysis
3. Results
3.1. Effect of the Heterologous Expression of the ToCV p22 Suppressor on TRV Virulence
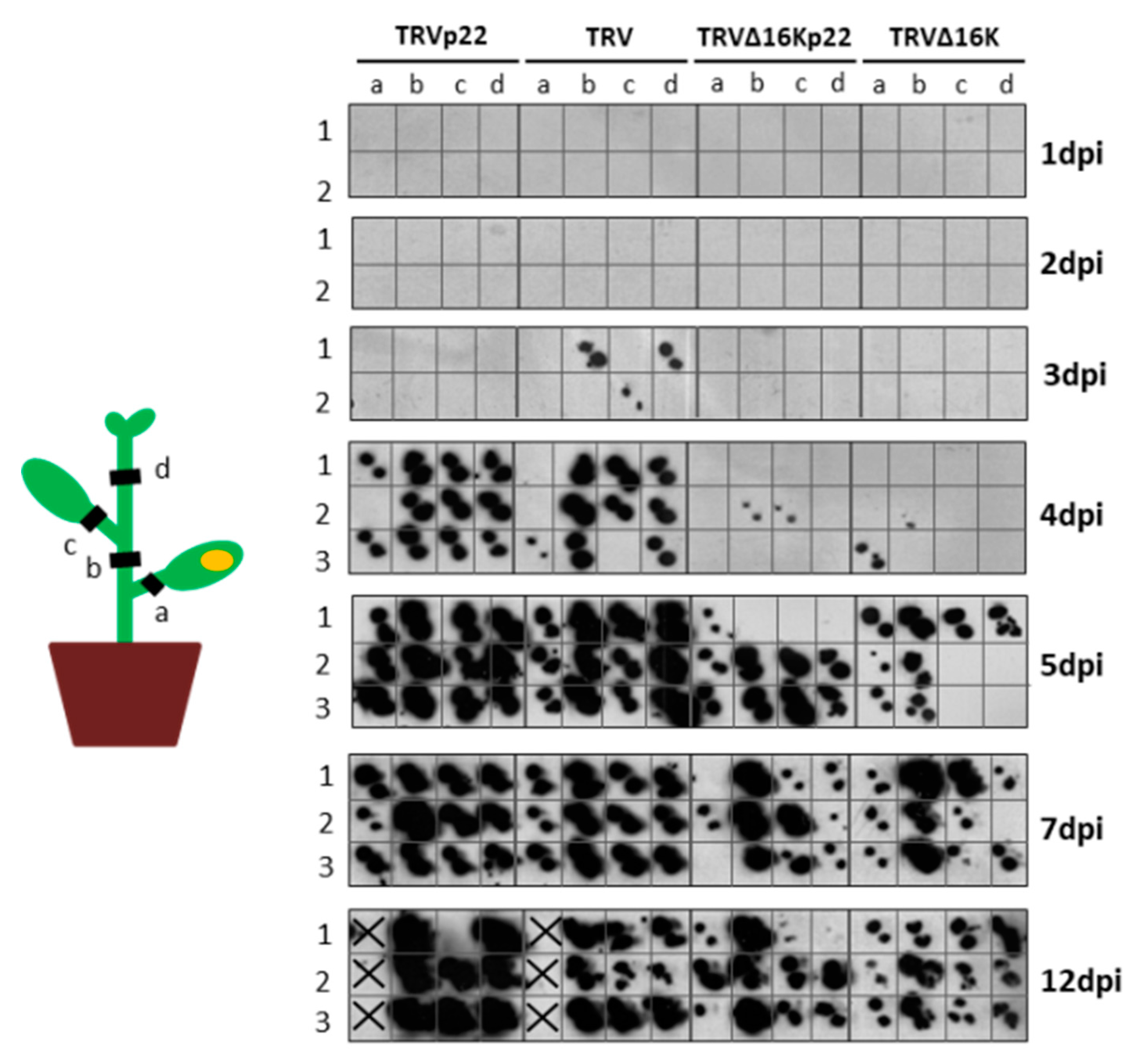
3.2. Enhanced Virulence with Heterologous Expression of the p22 Suppressor from a TRV Vector Is Not Linked to Faster Systemic Spread
3.3. Effect of the Heterologous Expression of the p22 Suppressor on PVX Virulence
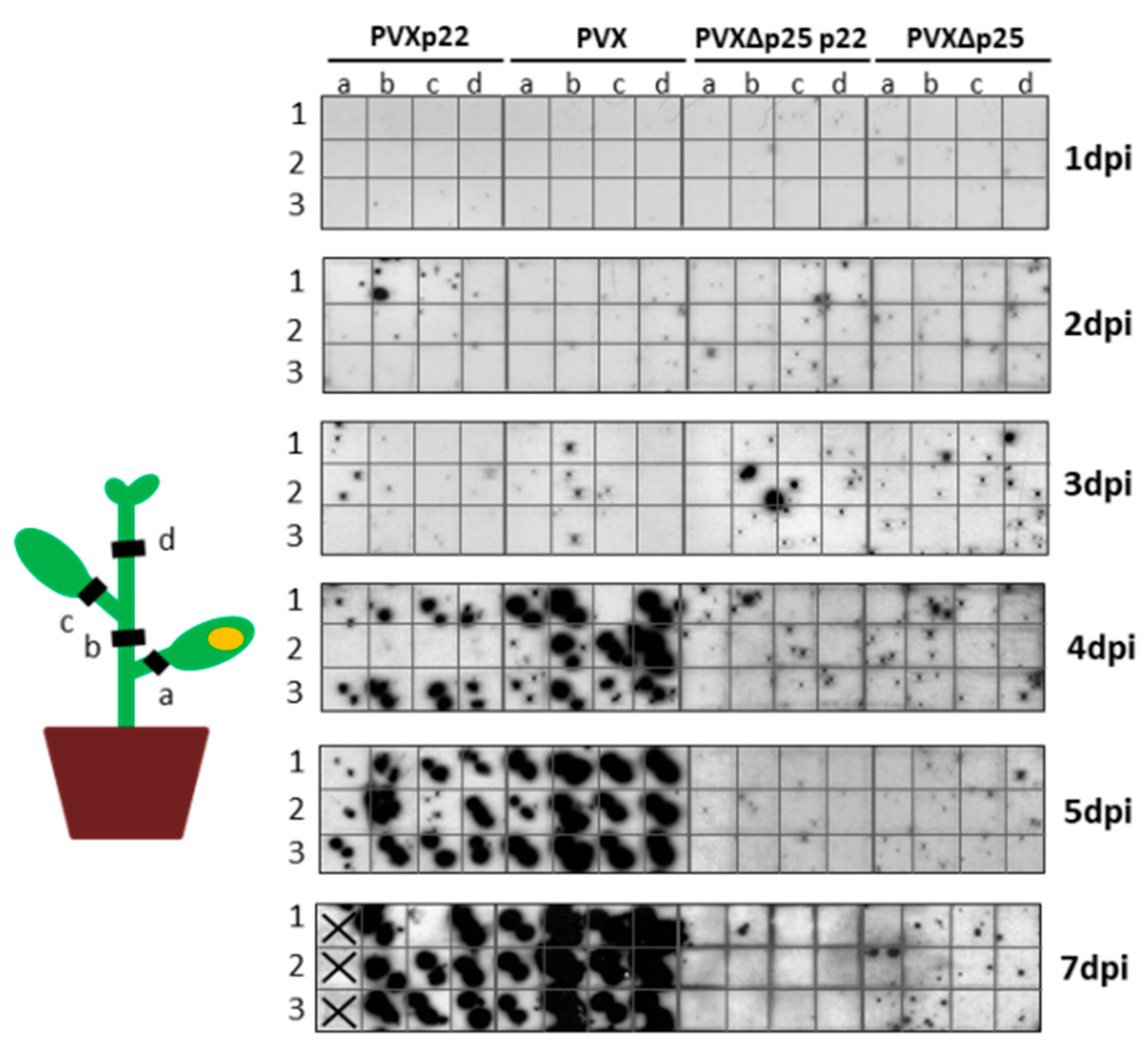
3.4. The Increased Virulence of the Recombinant Virus PVXp22 Is Not Linked to Faster Systemic Spread
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Baulcombe, D.C. RNA silencing in plants. Nature 2004, 431, 356–363. [Google Scholar] [CrossRef] [PubMed]
- Ding, S.W.; Shi, B.J.; Li, W.X.; Symons, R.H. An interspecies hybrid RNA virus is significantly more virulent than either parental virus. Proc. Natl. Acad. Sci. USA 1996, 93, 7470–7474. [Google Scholar] [CrossRef] [PubMed]
- Pruss, G.; Ge, X.; Shi, X.M.; Carrington, J.C.; Vance, V.B. Plant viral synergism: The potyviral genome encodes a broad-range pathogenicity enhancer that transactivates replication of heterologous viruses. Plant Cell 1997, 9, 859–868. [Google Scholar] [CrossRef] [PubMed]
- Vance, V.B.; Berger, P.H.; Carrington, J.C.; Hunt, A.G.; Shi, X.M. 5′ proximal potyviral sequences mediate Potato virus X/potyviral synergistic disease in transgenic tobacco. Virology 1995, 206, 583–590. [Google Scholar] [CrossRef]
- Goodman, R.M.; Ross, A.F. Enhancement by Potato virus Y of Potato virus X synthesis in doubly infected tobacco depends on the timing of invasion by the viruses. Virology 1974, 58, 16–24. [Google Scholar] [CrossRef]
- Damirdagh, I.S.; Ross, A.F. A marked synergistic interaction of Potato virus X and Y in inoculated leaves of tobacco. Virology 1967, 31, 296–307. [Google Scholar] [CrossRef]
- Vance, V.B. Replication of Potato virus X RNA is altered in coinfections with Potato virus Y. Virology 1991, 182, 486–494. [Google Scholar] [PubMed]
- González-Jara, P.; Tenllado, F.; Martínez-García, B.; Atencio, F.A.; Barajas, D.; Vargas, M.; Díaz-Ruiz, J.; Díaz-Ruiz, J.R. Host-dependent differences during synergistic infection by potyviruses with Potato virus X. Mol. Plant Pathol. 2004, 5, 29–35. [Google Scholar] [CrossRef] [PubMed]
- Moissiard, G.; Voinnet, O. Viral suppression of RNA silencing in plants. Mol. Plant Pathol. 2004, 5, 71–82. [Google Scholar] [CrossRef] [PubMed]
- Scholthof, H.B.; Scholthof, K.B.G.; Jackson, A.O. Identification of Tomato bushy stunt virus host-specific symptom determinants by expression of individual genes from a Potato virus X vector. Plant Cell 1995, 7, 1157–1172. [Google Scholar] [CrossRef] [PubMed]
- Dolja, V.V.; Kreuze, J.F.; Valkonen, J.P.T. Comparative and functional genomics of closteroviruses. Virus Res. 2006, 117, 38–51. [Google Scholar] [CrossRef] [PubMed]
- Lozano, G.; Moriones, E.; Navas-Castillo, J. Complete nucleotide sequence of the RNA2 of the crinivirus Tomato chlorosis virus. Arch. Virol. 2006, 151, 581–587. [Google Scholar] [CrossRef] [PubMed]
- Lozano, G.; Moriones, E.; Navas-Castillo, J. Complete sequence of the RNA1 of a European isolate of Tomato chlorosis virus. Arch. Virol. 2007, 152, 839–841. [Google Scholar] [CrossRef] [PubMed]
- Wintermantel, W.M.; Wisler, G.C.; Anchieta, A.G.; Liu, H.Y.; Karasev, A.V.; Tzanetakis, I.E. The complete nucleotide sequence and genome organization of Tomato chlorosis virus. Arch. Virol. 2005, 150, 2287–2298. [Google Scholar] [CrossRef] [PubMed]
- Wisler, G.C.; Li, R.H.; Liu, H.Y.; Lowry, D.S.; Duffus, J.E. Tomato chlorosis virus: A new whitefly-transmitted, phloem-limited, bipartite closterovirus of tomato. Phytopathology 1998, 88, 402–409. [Google Scholar] [CrossRef] [PubMed]
- Cañizares, M.C.; Navas-Castillo, J.; Moriones, E. Multiple suppressors of RNA silencing encoded by both genomic RNAs of the crinivirus, Tomato chlorosis virus. Virology 2008, 379, 168–174. [Google Scholar] [CrossRef] [PubMed]
- Landeo-Ríos, Y.; Navas-Castillo, J.; Moriones, E.; Cañizares, M.C. The p22 RNA silencing suppressor of the crinivirus Tomato chlorosis virus preferentially binds long dsRNAs preventing them from cleavage. Virology 2016, 488, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Robertson, D. VIGS vectors for gene silencing: Many targets, many tools. Annu. Rev. Plant Biol. 2004, 55, 495–519. [Google Scholar] [CrossRef] [PubMed]
- MacFarlane, S.A.; Popovich, A.H. Efficient expression of foreign proteins in roots from tobravirus vectors. Virology 2000, 267, 29–35. [Google Scholar] [CrossRef] [PubMed]
- Mallory, A.C.; Parks, G.; Endres, M.W.; Baulcombe, D.; Bowman, L.H.; Pruss, G.J.; Vance, V.B. The amplicon-plus system for high-level expression of transgenes in plants. Nat. Biotechnol. 2002, 20, 622–625. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, W.D.O.; Baulcombe, D.C. Infectious RNA produced by in vitro transcription of a full-length Tobacco rattle virus RNA1 cDNA. J. Gen. Virol. 1989, 70, 963–968. [Google Scholar] [CrossRef]
- Deng, X.; Kelloniemi, J.; Haikonen, T.; Vuorinen, A.L.; Elomaa, P.; Teeri, T.H.; Valkonen, J.P. Modification of Tobacco rattle virus RNA1 to serve as a VIGS vector reveals that the 29K movement protein is an RNA silencing suppressor of the virus. Mol. Plant-Microbe Interact. 2013, 26, 503–514. [Google Scholar] [CrossRef] [PubMed]
- Ghazala, W.; Waltermann, A.; Pilot, R.; Winter, S.; Varrelmann, M. Functional characterization and subcellular localization of the 16K cysteine-rich suppressor of gene silencing protein of Tobacco rattle virus. J. Gen. Virol. 2008, 89, 1748–1758. [Google Scholar] [CrossRef] [PubMed]
- Martín-Hernández, A.M.; Baulcombe, D.C. Tobacco rattle virus 16-kilodalton protein encodes a suppressor of RNA silencing that allows transient viral entry in meristems. J. Virol. 2008, 82, 4064–4071. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Priego, L.; Donaire, L.; Barajas, D.; Llave, C. Silencing suppressor activity of the Tobacco rattle virus-encoded 16-kDa protein and interference with endogenous small RNA-guided regulatory pathways. Virology 2008, 376, 346–356. [Google Scholar] [CrossRef] [PubMed]
- Reavy, B.; Dawson, S.; Canto, T.; MacFarlane, S.A. Heterologous expression of plant virus genes that suppress post-transcriptional gene silencing results in suppression of RNA interference in Drosophila cells. BMC Biotechnol. 2004, 4, 18. [Google Scholar] [CrossRef] [PubMed]
- MacFarlane, S.A. Molecular biology of the tobraviruses. J. Gen. Virol. 1999, 80, 2799–2807. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Reavy, B.; Swanson, M.; MacFarlane, S.A. Functional replacement of the Tobacco rattle virus cysteine-rich protein by pathogenicity proteins from unrelated plant viruses. Virology 2002, 298, 232–239. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.L.; Schiff, M.; Marathe, R.; Dinesh-Kumar, S.P. Tobacco Rar1, EDS1 and NPR1/NIM1 like genes are required for N-mediated resistance to Tobacco mosaic virus. Plant J. 2002, 30, 415–429. [Google Scholar] [CrossRef] [PubMed]
- Ratcliff, F.; Martín-Hernández, A.M.; Baulcombe, D.C. Tobacco rattle virus as a vector for analysis of gene function by silencing. Plant J. 2001, 25, 237–245. [Google Scholar] [CrossRef] [PubMed]
- Ruiz, M.T.; Voinnet, O.; Baulcombe, D.C. Initiation and maintenance of virus-induced gene silencing. Plant Cell 1998, 10, 937–946. [Google Scholar] [CrossRef] [PubMed]
- Swanson, M.; Barker, H.; MacFarlane, S.A. Rapid vascular movement of tobraviruses does not require coat protein: Evidence from mutated and wild-type viruses. Ann. Appl. Biol. 2002, 141, 259–266. [Google Scholar] [CrossRef]
- Verchot-Lubicz, J.; Ye, C.M.; Bamunusinghe, D. Molecular biology of potexviruses: Recent advances. J. Gen. Virol. 2007, 88, 1643–1655. [Google Scholar] [CrossRef] [PubMed]
- Angell, S.M.; Davies, C.; Baulcombe, D.C. Cell-to-cell movement of Potato virus X is associated with a change in the size exclusion limit of plasmodesmata in trichome cells of Nicotiana clevelandii. Virology 1996, 215, 197–201. [Google Scholar] [CrossRef] [PubMed]
- Bayne, E.H.; Rakitina, D.V.; Morozov, S.Y.; Baulcombe, D.C. Cell-to-cell movement of Potato potexvirus X is dependent on suppression of RNA silencing. Plant J. 2005, 44, 471–482. [Google Scholar] [CrossRef] [PubMed]
- Jones, L.; Hamilton, A.J.; Voinnet, O.; Thomas, C.L.; Maule, A.J.; Baulcombe, D.C. RNA-DNA interactions and DNA methylation in post-transcriptional gene silencing. Plant Cell 1999, 11, 2291–2301. [Google Scholar] [CrossRef] [PubMed]
- Noris, E.; Accotto, G.P.; Tavazza, R.; Brunetti, A.; Crespi, S.; Tavazza, M. Resistance to tomato yellow leaf curl geminivirus in Nicotiana benthamiana plants transformed with a truncated viral C1 gene. Virology 1996, 225, 130–138. [Google Scholar] [CrossRef] [PubMed]
- Karyeija, R.F.; Kreuze, J.F.; Gibson, R.W.; Valkonen, J.P. Synergistic interactions of a potyvirus and a phloem-limited crinivirus in sweet potato plants. Virology 2000, 269, 26–36. [Google Scholar] [CrossRef] [PubMed]
- Mukasa, S.B.; Rubaihayo, P.R.; Valkonen, J.P.T. Interactions between a crinivirus, an ipomoirus and a potyvirus in coinfected sweetpotato plants. Plant Pathol. 2006, 55, 458–467. [Google Scholar] [CrossRef]
- García-Cano, E.; Resende, R.O.; Fernández-Muñoz, R.; Moriones, E. Synergistic interaction between Tomato chlorosis virus and Tomato spotted wilt virus results in breakdown of resistance in tomato. Phytopathology 2006, 96, 1263–1269. [Google Scholar] [CrossRef] [PubMed]
- Wintermantel, W.M.; Cortez, A.A.; Anchieta, A.G.; Gulati-Sakhuja, A.; Hladky, L.L. Co-infection by two criniviruses alters accumulation of each virus in a host-specific manner and influences efficiency of virus transmission. Phytopathology 2008, 98, 1340–1345. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Turina, M.; Medina, V.; Falk, B.W. Synergistic interaction between the Potyvirus, Turnip mosaic virus and the Crinivirus, Lettuce infectious yellows virus in plants and protoplasts. Virus Res. 2009, 144, 163–170. [Google Scholar] [CrossRef] [PubMed]
- MacDiarmid, R. RNA silencing in productive virus infections. Annu. Rev. Phytopathol. 2005, 43, 523–544. [Google Scholar] [CrossRef] [PubMed]
- Tian, J.; Pei, H.; Zhang, S.; Chen, J.; Chen, W.; Yang, R.; Meng, Y.; You, J.; Gao, J.; Ma, N. TRV-GFP: A modified Tobacco rattle virus vector for efficient and visualizable analysis of gene function. J. Exp. Bot. 2014, 65, 311–322. [Google Scholar] [CrossRef] [PubMed]
- Aguilar, E.; Almendral, D.; Allende, L.; Pacheco, R.; Chung, B.N.; Canto, T.; Tenllado, F. The P25 protein of Potato virus X (PVX) is the main pathogenicity determinant responsible for systemic necrosis in PVX-associated synergisms. J. Virol. 2015, 89, 2090–2103. [Google Scholar] [CrossRef] [PubMed]
- Landeo-Ríos, Y.; Navas-Castillo, J.; Moriones, E.; Cañizares, M.C. The p22 RNA silencing suppressor of the crinivirus Tomato chlorosis virus is dispensable for local viral replication but important for counteracting an antiviral RDR6-mediated response during systemic infection. Viruses 2016, 8, 182. [Google Scholar] [CrossRef] [PubMed]
- Scholthof, H.B.; Scholthof, K.B.G.; Kibbert, M.; Jackson, A.O. Tomato bushy stunt virus spread is regulated by two nested genes that function in cell-to-cell movement and host-dependent systemic invasion. Virology 1995, 213, 425–438. [Google Scholar] [CrossRef] [PubMed]
- Cronin, S.; Verchot, J.; Haldeman-Cahill, R.; Schaad, M.C.; Carrington, J.C. Long-distance movement factor: A transport function of the potyvirus helper component proteinase. Plant Cell 1995, 7, 549–559. [Google Scholar] [CrossRef] [PubMed]
- Ding, S.W.; Li, W.X.; Symons, R.H. A novel naturally occurring hybrid-gene encoded by a plant RNA virus facilitates long distance virus movement. EMBO J. 1995, 14, 5762–5772. [Google Scholar] [PubMed]
- Klein, P.G.; Klein, R.R.; Rodríguez-Cerezo, E.; Hunt, A.G.; Shaw, A.G. Mutational analysis of the Tobacco vein mottling virus genome. Virology 1994, 204, 759–769. [Google Scholar] [CrossRef] [PubMed]


© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Landeo-Ríos, Y.; Navas-Castillo, J.; Moriones, E.; Cañizares, M.C. The Heterologous Expression of the p22 RNA Silencing Suppressor of the Crinivirus Tomato Chlorosis Virus from Tobacco Rattle Virus and Potato Virus X Enhances Disease Severity but Does Not Complement Suppressor-Defective Mutant Viruses. Viruses 2017, 9, 358. https://doi.org/10.3390/v9120358
Landeo-Ríos Y, Navas-Castillo J, Moriones E, Cañizares MC. The Heterologous Expression of the p22 RNA Silencing Suppressor of the Crinivirus Tomato Chlorosis Virus from Tobacco Rattle Virus and Potato Virus X Enhances Disease Severity but Does Not Complement Suppressor-Defective Mutant Viruses. Viruses. 2017; 9(12):358. https://doi.org/10.3390/v9120358
Chicago/Turabian StyleLandeo-Ríos, Yazmín, Jesús Navas-Castillo, Enrique Moriones, and M. Carmen Cañizares. 2017. "The Heterologous Expression of the p22 RNA Silencing Suppressor of the Crinivirus Tomato Chlorosis Virus from Tobacco Rattle Virus and Potato Virus X Enhances Disease Severity but Does Not Complement Suppressor-Defective Mutant Viruses" Viruses 9, no. 12: 358. https://doi.org/10.3390/v9120358
APA StyleLandeo-Ríos, Y., Navas-Castillo, J., Moriones, E., & Cañizares, M. C. (2017). The Heterologous Expression of the p22 RNA Silencing Suppressor of the Crinivirus Tomato Chlorosis Virus from Tobacco Rattle Virus and Potato Virus X Enhances Disease Severity but Does Not Complement Suppressor-Defective Mutant Viruses. Viruses, 9(12), 358. https://doi.org/10.3390/v9120358





