Binding of RNA by the Nucleoproteins of Influenza Viruses A and B
Abstract
:1. Introduction
2. Materials and Methods
2.1. Protein Expression and Purification of Influenza A and B NPs
2.2. Fluorescence Anisotropy Measurements
2.3. Electron Microscopy
2.4. SEC-MALLS Experiments
3. Results
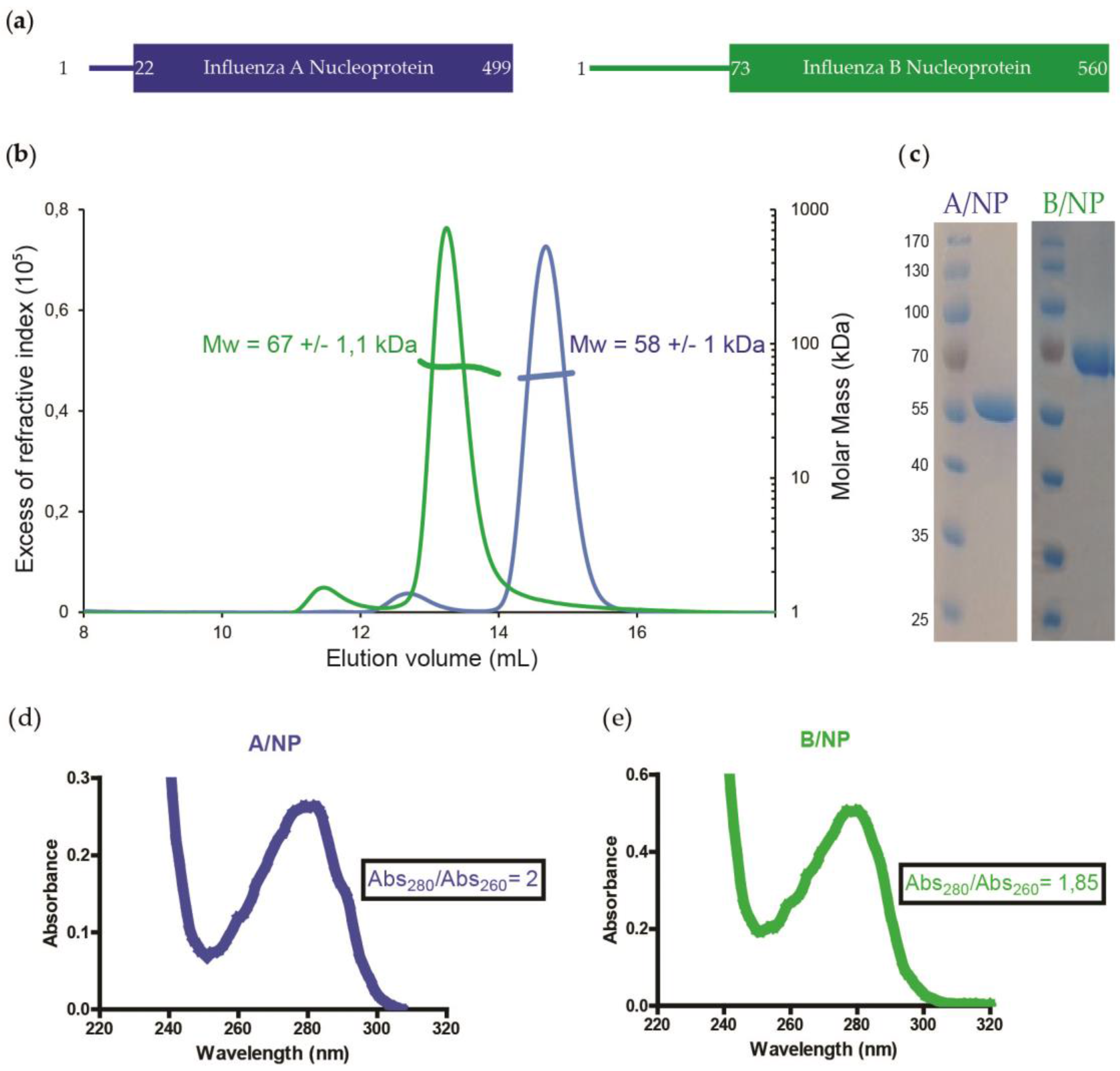
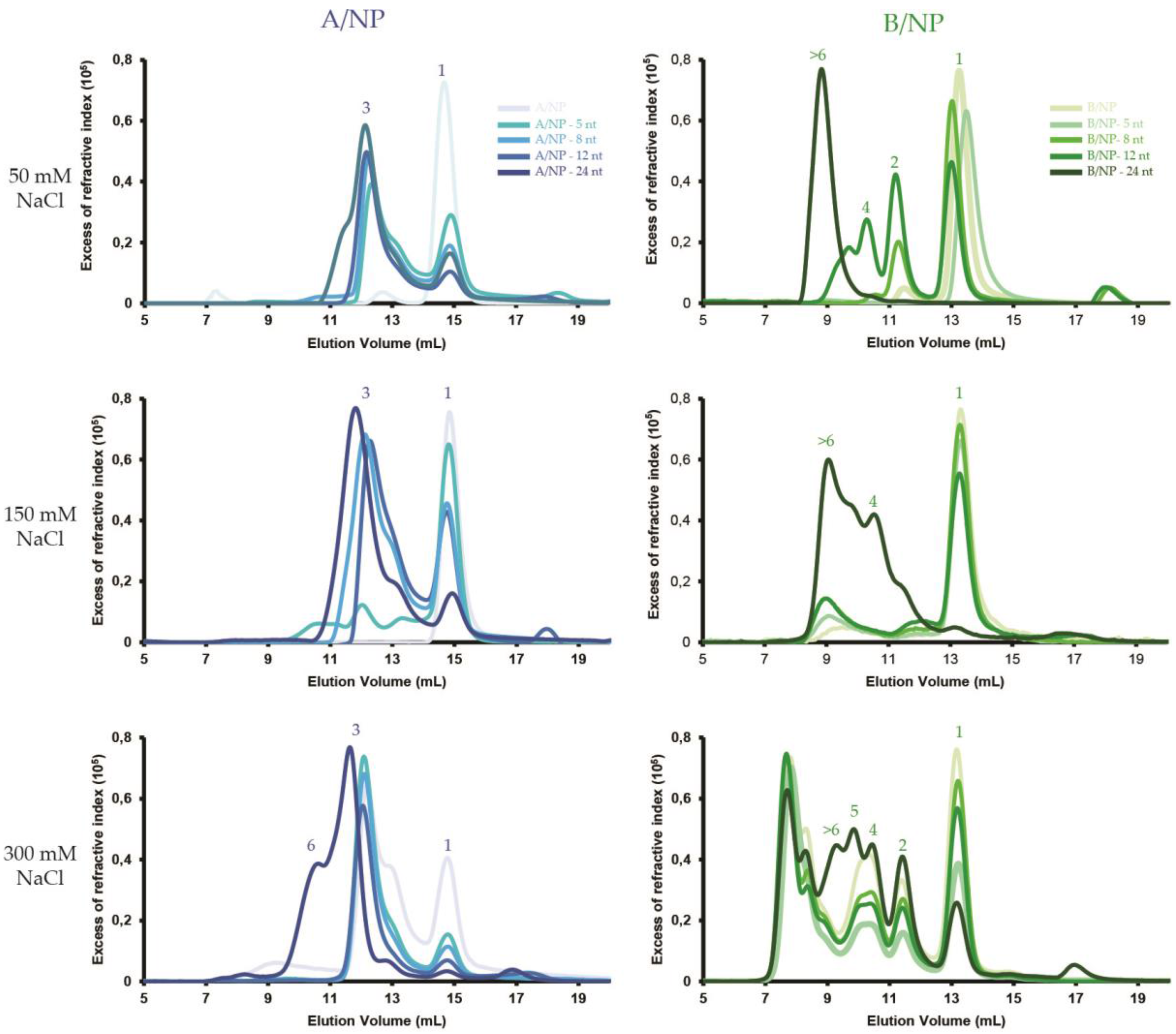
3.1. SEC-MALLS
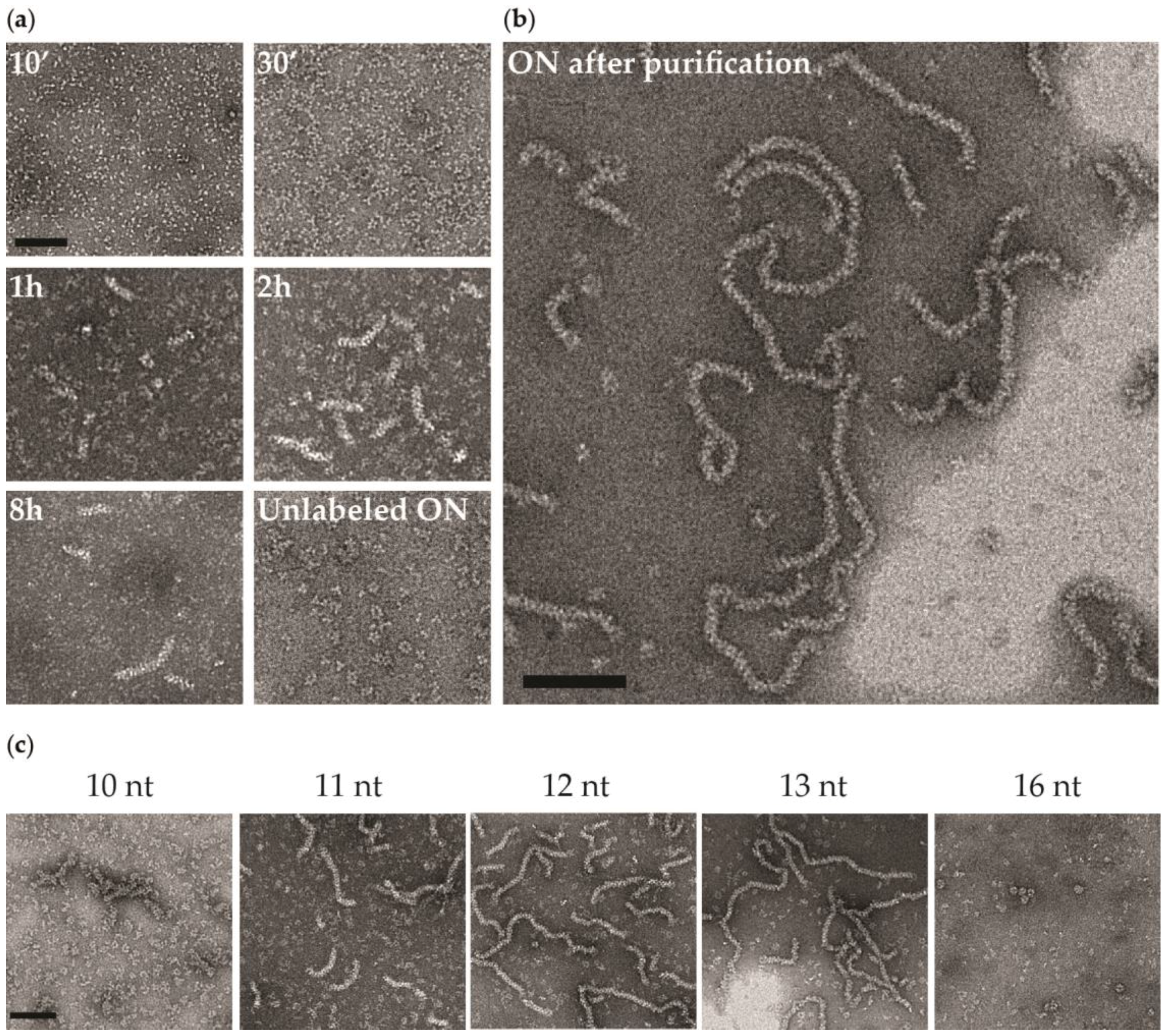
3.2. Negative Staining EM
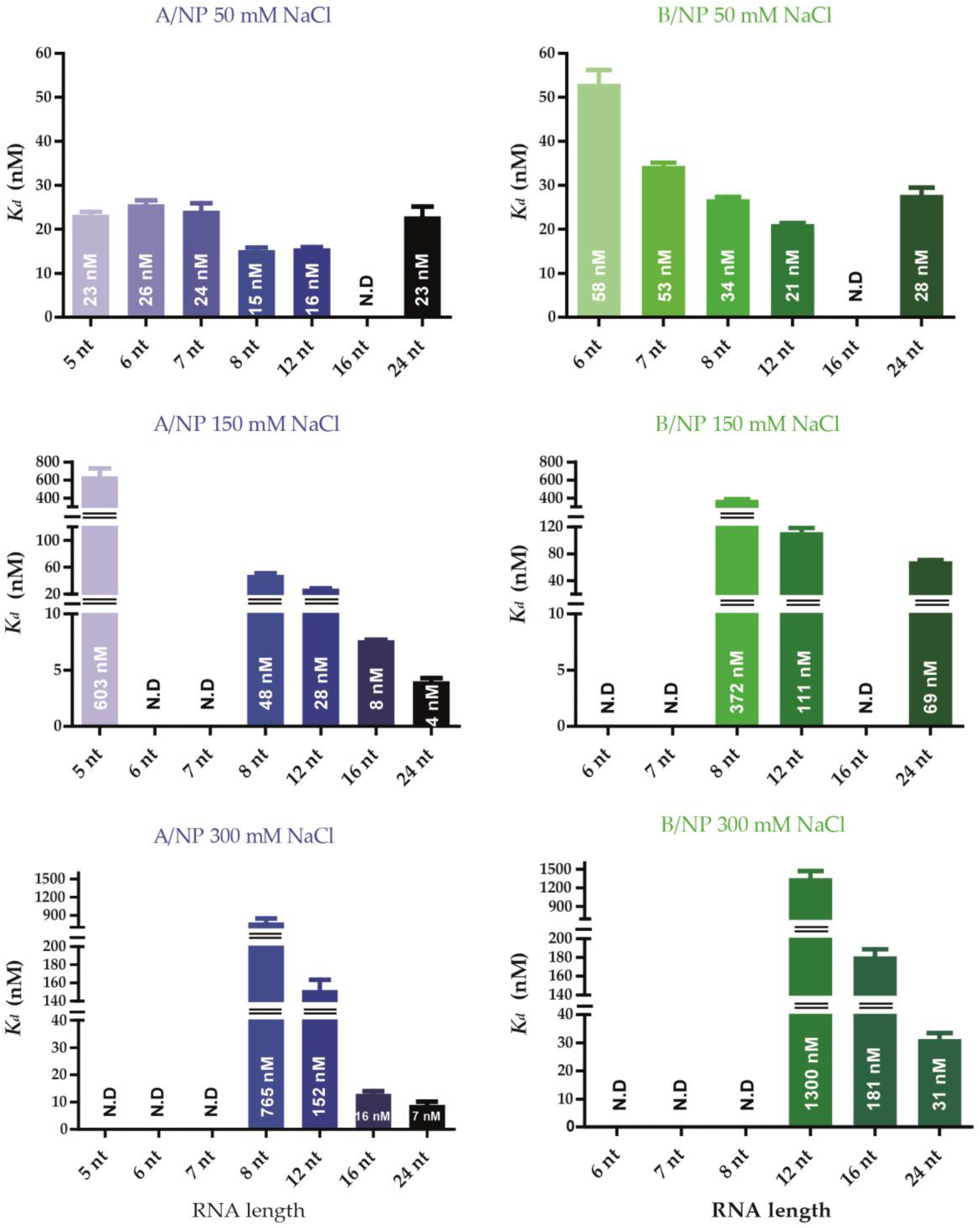
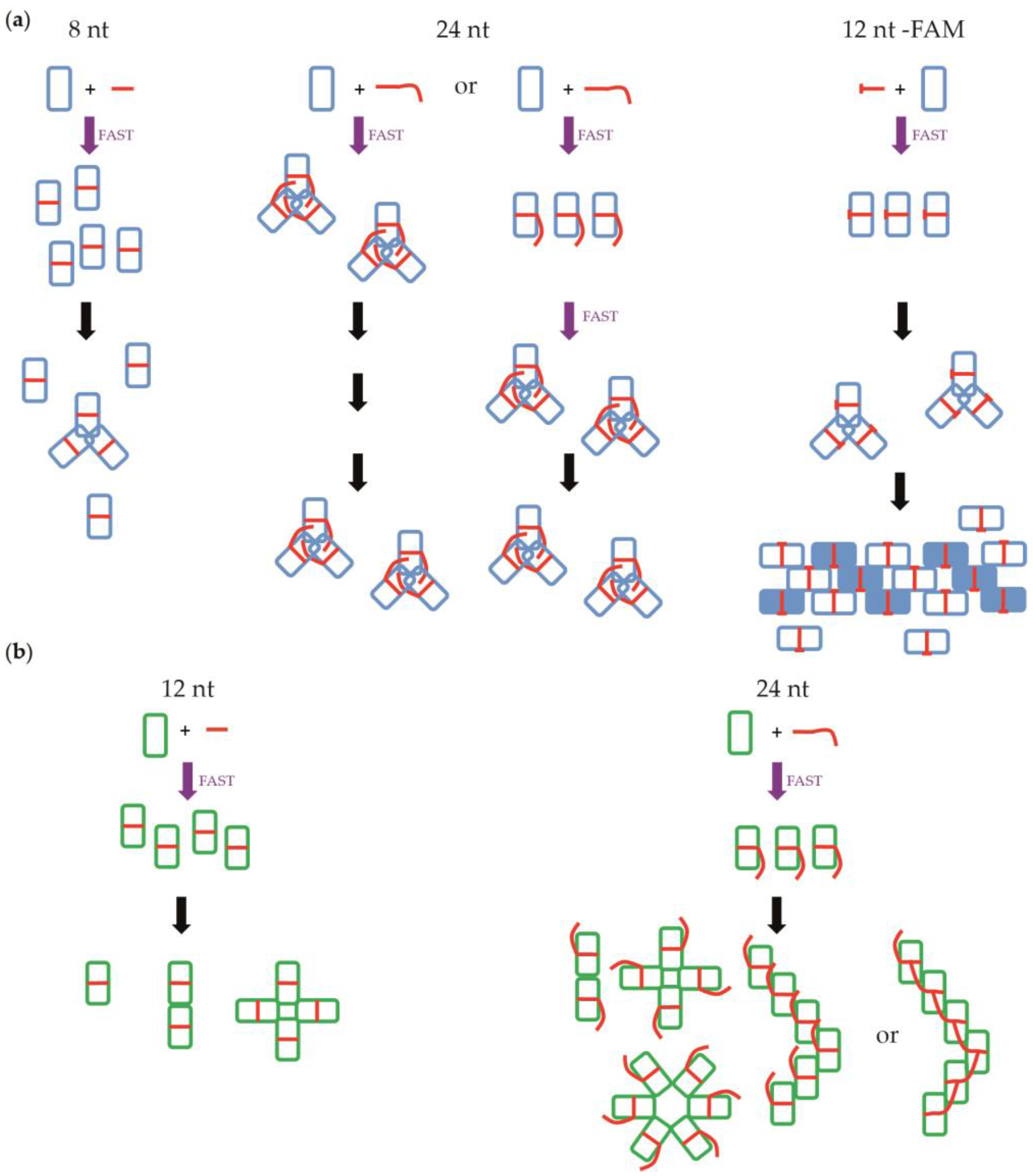
3.3. Fluorescence Anisotropy Measurements
4. Discussion
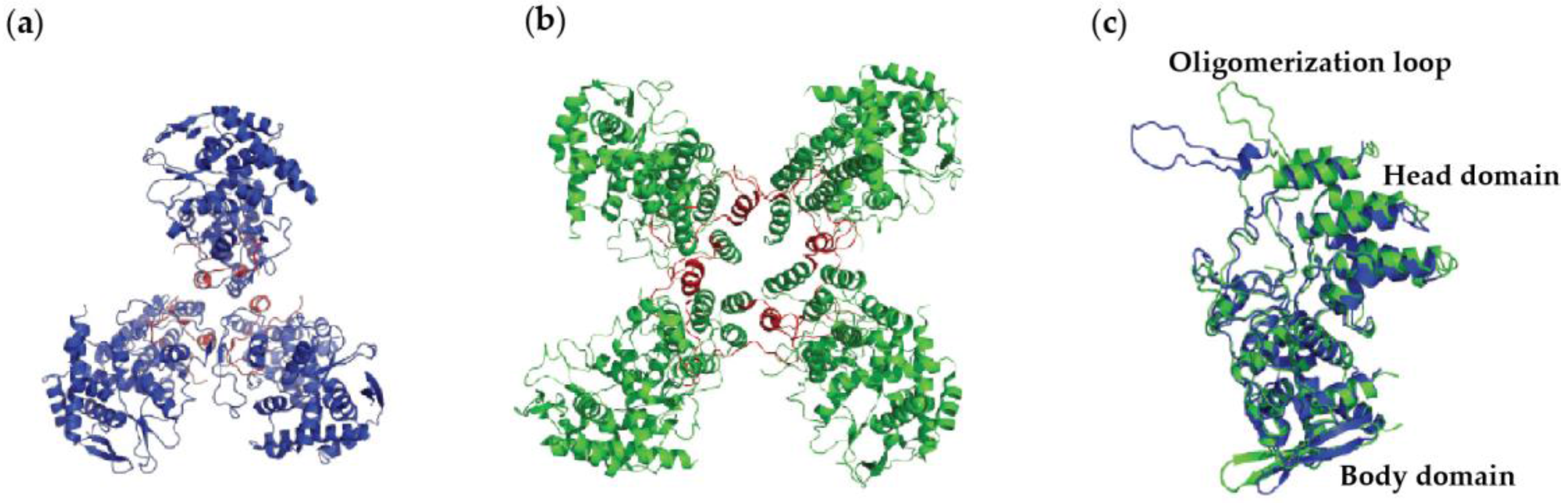
4.1. Nucleoprotein of A
4.2. Nucleoprotein of B
4.3. NP of Influenza A with 5′Phosphate-(UC)6-FAM3′
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Reguera, J.; Cusack, S.; Kolakofsky, D. Segmented negative strand RNA virus nucleoprotein structure. Curr. Opin. Virol. 2014, 5, 7–15. [Google Scholar]
- Ruigrok, R.W.; Crepin, T.; Kolakofsky, D. Nucleoproteins and nucleocapsids of negative-strand RNA viruses. Curr. Opin. Microbiol. 2011, 14, 504–510. [Google Scholar] [CrossRef] [PubMed]
- Albertini, A.A.; Wernimont, A.K.; Muziol, T.; Ravelli, R.B.; Clapier, C.R.; Schoehn, G.; Weissenhorn, W.; Ruigrok, R.W. Crystal structure of the rabies virus nucleoprotein-RNA complex. Science 2006, 313, 360–363. [Google Scholar] [CrossRef] [PubMed]
- Green, T.J.; Zhang, X.; Wertz, G.W.; Luo, M. Structure of the vesicular stomatitis virus nucleoprotein-RNA complex. Science 2006, 313, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Renner, M.; Bertinelli, M.; Leyrat, C.; Paesen, G.C.; Saraiva de Oliveira, L.F.; Huiskonen, J.T.; Grimes, J.M. Nucleocapsid assembly in pneumoviruses is regulated by conformational switching of the N protein. Elife 2016, 5, e12627. [Google Scholar] [CrossRef] [PubMed]
- Tawar, R.G.; Duquerroy, S.; Vonrhein, C.; Varela, P.F.; Damier-Piolle, L.; Castagne, N.; MacLellan, K.; Bedouelle, H.; Bricogne, G.; Bhella, D.; et al. Crystal structure of a nucleocapsid-like nucleoprotein-RNA complex of respiratory syncytial virus. Science 2009, 326, 1279–1283. [Google Scholar] [CrossRef] [PubMed]
- Gutsche, I.; Desfosses, A.; Effantin, G.; Ling, W.L.; Haupt, M.; Ruigrok, R.W.; Sachse, C.; Schoehn, G. Structural virology. Near-atomic cryo-EM structure of the helical measles virus nucleocapsid. Science 2015, 348, 704–707. [Google Scholar] [PubMed]
- Alayyoubi, M.; Leser, G.P.; Kors, C.A.; Lamb, R.A. Structure of the paramyxovirus parainfluenza virus 5 nucleoprotein-RNA complex. Proc. Natl. Acad. Sci. USA 2015, 112, E1792–E1799. [Google Scholar] [CrossRef] [PubMed]
- Calain, P.; Roux, L. The rule of six, a basic feature for efficient replication of Sendai virus defective interfering RNA. J. Virol. 1993, 67, 4822–4830. [Google Scholar] [PubMed]
- Egelman, E.H.; Wu, S.S.; Amrein, M.; Portner, A.; Murti, G. The Sendai virus nucleocapsid exists in at least four different helical states. J. Virol. 1989, 63, 2233–2243. [Google Scholar] [PubMed]
- Kolakofsky, D.; Pelet, T.; Garcin, D.; Hausmann, S.; Curran, J.; Roux, L. Paramyxovirus RNA synthesis and the requirement for hexamer genome length: The rule of six revisited. J. Virol. 1998, 72, 891–899. [Google Scholar] [PubMed]
- Hastie, K.M.; Kimberlin, C.R.; Zandonatti, M.A.; MacRae, I.J.; Saphire, E.O. Structure of the Lassa virus nucleoprotein reveals a dsRNA-specific 3′ to 5′ exonuclease activity essential for immune suppression. Proc. Natl. Acad. Sci. USA 2011, 108, 2396–2401. [Google Scholar] [CrossRef] [PubMed]
- Hastie, K.M.; Liu, T.; Li, S.; King, L.B.; Ngo, N.; Zandonatti, M.A.; Woods, V.L., Jr.; de la Torre, J.C.; Saphire, E.O. Crystal structure of the Lassa virus nucleoprotein-RNA complex reveals a gating mechanism for RNA binding. Proc. Natl. Acad. Sci. USA 2011, 108, 19365–19370. [Google Scholar] [CrossRef] [PubMed]
- Raymond, D.D.; Piper, M.E.; Gerrard, S.R.; Skiniotis, G.; Smith, J.L. Phleboviruses encapsidate their genomes by sequestering RNA bases. Proc. Natl. Acad. Sci. USA 2012, 109, 19208–19213. [Google Scholar] [CrossRef] [PubMed]
- Ariza, A.; Tanner, S.J.; Walter, C.T.; Dent, K.C.; Shepherd, D.A.; Wu, W.; Matthews, S.V.; Hiscox, J.A.; Green, T.J.; Luo, M.; et al. Nucleocapsid protein structures from orthobunyaviruses reveal insight into ribonucleoprotein architecture and RNA polymerization. Nucleic Acids Res. 2013, 41, 5912–5926. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Li, P.; Bottcher, B.; Elliott, R.M.; Dong, C. Crystal structure of Schmallenberg orthobunyavirus nucleoprotein-RNA complex reveals a novel RNA sequestration mechanism. RNA 2013, 19, 1129–1136. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Li, P.; Elliott, R.M.; Dong, C. Structure of Schmallenberg orthobunyavirus nucleoprotein suggests a novel mechanism of genome encapsidation. J. Virol. 2013, 87, 5593–5601. [Google Scholar] [CrossRef] [PubMed]
- Niu, F.; Shaw, N.; Wang, Y.E.; Jiao, L.; Ding, W.; Li, X.; Zhu, P.; Upur, H.; Ouyang, S.; Cheng, G.; Liu, Z.J. Structure of the Leanyer orthobunyavirus nucleoprotein-RNA complex reveals unique architecture for RNA encapsidation. Proc. Natl. Acad. Sci. USA 2013, 110, 9054–9059. [Google Scholar] [CrossRef] [PubMed]
- Reguera, J.; Malet, H.; Weber, F.; Cusack, S. Structural basis for encapsidation of genomic RNA by La Crosse Orthobunyavirus nucleoprotein. Proc. Natl. Acad. Sci. USA 2013, 110, 7246–7251. [Google Scholar] [CrossRef] [PubMed]
- Dong, S.; Yang, P.; Li, G.; Liu, B.; Wang, W.; Liu, X.; Xia, B.; Yang, C.; Lou, Z.; Guo, Y.; Rao, Z. Insight into the Ebola virus nucleocapsid assembly mechanism: Crystal structure of Ebola virus nucleoprotein core domain at 1.8 A resolution. Protein Cell 2015, 6, 351–362. [Google Scholar] [CrossRef] [PubMed]
- Dziubanska, P.J.; Derewenda, U.; Ellena, J.F.; Engel, D.A.; Derewenda, Z.S. The structure of the C-terminal domain of the Zaire ebolavirus nucleoprotein. Acta Crystallogr. D Biol. Crystallogr. 2014, 70, 2420–2429. [Google Scholar] [CrossRef] [PubMed]
- Kirchdoerfer, R.N.; Abelson, D.M.; Li, S.; Wood, M.R.; Saphire, E.O. Assembly of the Ebola Virus Nucleoprotein from a Chaperoned VP35 Complex. Cell Rep. 2015, 12, 140–149. [Google Scholar] [CrossRef] [PubMed]
- Leung, D.W.; Borek, D.; Luthra, P.; Binning, J.M.; Anantpadma, M.; Liu, G.; Harvey, I.B.; Su, Z.; Endlich-Frazier, A.; Pan, J.; et al. An Intrinsically Disordered Peptide from Ebola Virus VP35 Controls Viral RNA Synthesis by Modulating Nucleoprotein-RNA Interactions. Cell Rep. 2015, 11, 376–389. [Google Scholar] [CrossRef] [PubMed]
- Ng, A.K.; Lam, M.K.; Zhang, H.; Liu, J.; Au, S.W.; Chan, P.K.; Wang, J.; Shaw, P.C. Structural basis for RNA binding and homo-oligomer formation by influenza B virus nucleoprotein. J. Virol. 2012, 86, 6758–6767. [Google Scholar] [CrossRef] [PubMed]
- Ng, A.K.; Zhang, H.; Tan, K.; Li, Z.; Liu, J.H.; Chan, P.K.; Li, S.M.; Chan, W.Y.; Au, S.W.; Joachimiak, A.; et al. Structure of the influenza virus A H5N1 nucleoprotein: Implications for RNA binding, oligomerization, and vaccine design. FASEB J. 2008, 22, 3638–3647. [Google Scholar] [CrossRef] [PubMed]
- Rudolph, M.G.; Kraus, I.; Dickmanns, A.; Eickmann, M.; Garten, W.; Ficner, R. Crystal structure of the borna disease virus nucleoprotein. Structure 2003, 11, 1219–1226. [Google Scholar] [CrossRef] [PubMed]
- Ye, Q.; Krug, R.M.; Tao, Y.J. The mechanism by which influenza A virus nucleoprotein forms oligomers and binds RNA. Nature 2006, 444, 1078–1082. [Google Scholar] [CrossRef] [PubMed]
- Chenavas, S.; Estrozi, L.F.; Slama-Schwok, A.; Delmas, B.; Di Primo, C.; Baudin, F.; Li, X.; Crépin, T.; Ruigrok, R.W. Monomeric nucleoprotein of influenza A virus. PLoS Pathog. 2013, 9, e1003275. [Google Scholar] [CrossRef] [PubMed]
- Ruigrok, R.W.; Baudin, F. Structure of influenza virus ribonucleoprotein particles. II. Purified RNA-free influenza virus ribonucleoprotein forms structures that are indistinguishable from the intact influenza virus ribonucleoprotein particles. J. Gen. Virol. 1995, 76, 1009–1014. [Google Scholar] [PubMed]
- Compans, R.W.; Content, J.; Duesberg, P.H. Structure of the ribonucleoprotein of influenza virus. J. Virol. 1972, 10, 795–800. [Google Scholar] [PubMed]
- Duesberg, P.H. Distinct subunits of the ribonucleoprotein of influenza virus. J. Mol. Biol. 1969, 42, 485–499. [Google Scholar] [CrossRef]
- Kingsbury, D.W.; Webster, R.G. Some properties of influenza virus nucleocapsids. J. Virol. 1969, 4, 219–225. [Google Scholar] [PubMed]
- Pons, M.W.; Schulze, I.T.; Hirst, G.K.; Hauser, R. Isolation and characterization of the ribonucleoprotein of influenza virus. Virology 1969, 39, 250–259. [Google Scholar] [CrossRef]
- Ortega, J.; Martin-Benito, J.; Zurcher, T.; Valpuesta, J.M.; Carrascosa, J.L.; Ortin, J. Ultrastructural and functional analyses of recombinant influenza virus ribonucleoproteins suggest dimerization of nucleoprotein during virus amplification. J. Virol. 2000, 74, 156–163. [Google Scholar] [CrossRef] [PubMed]
- Hutchinson, E.C.; Charles, P.D.; Hester, S.S.; Thomas, B.; Trudgian, D.; Martinez-Alonso, M.; Fodor, E. Conserved and host-specific features of influenza virion architecture. Nat. Commun. 2014, 5, 4816. [Google Scholar] [CrossRef] [PubMed]
- Zheng, W.; Olson, J.; Vakharia, V.; Tao, Y.J. The crystal structure and RNA-binding of an orthomyxovirus nucleoprotein. PLoS Pathog. 2013, 9, e1003624. [Google Scholar] [CrossRef] [PubMed]
- Baudin, F.; Bach, C.; Cusack, S.; Ruigrok, R.W. Structure of influenza virus RNP. I. Influenza virus nucleoprotein melts secondary structure in panhandle RNA and exposes the bases to the solvent. EMBO J. 1994, 13, 3158–3165. [Google Scholar] [PubMed]
- Chan, W.H.; Ng, A.K.; Robb, N.C.; Lam, M.K.; Chan, P.K.; Au, S.W.; Wang, J.H.; Fodor, E.; Shaw, P.C. Functional analysis of the influenza virus H5N1 nucleoprotein tail loop reveals amino acids that are crucial for oligomerization and ribonucleoprotein activities. J. Virol. 2010, 84, 7337–7345. [Google Scholar] [CrossRef] [PubMed]
- Tarus, B.; Bakowiez, O.; Chenavas, S.; Duchemin, L.; Estrozi, L.F.; Bourdieu, C.; Lejal, N.; Bernard, J.; Moudjou, M.; Chevalier, C.; et al. Oligomerization paths of the nucleoprotein of influenza A virus. Biochimie 2012, 94, 776–785. [Google Scholar] [CrossRef] [PubMed]
- Elton, D.; Medcalf, L.; Bishop, K.; Harrison, D.; Digard, P. Identification of amino acid residues of influenza virus nucleoprotein essential for RNA binding. J. Virol. 1999, 73, 7357–7367. [Google Scholar] [PubMed]
- Boulo, S.; Akarsu, H.; Lotteau, V.; Muller, C.W.; Ruigrok, R.W.; Baudin, F. Human importin alpha and RNA do not compete for binding to influenza A virus nucleoprotein. Virology 2011, 409, 84–90. [Google Scholar] [CrossRef] [PubMed]
- Ye, Q.; Guu, T.S.; Mata, D.A.; Kuo, R.L.; Smith, B.; Krug, R.M.; Tao, Y.J. Biochemical and structural evidence in support of a coherent model for the formation of the double-helical influenza A virus ribonucleoprotein. MBio 2012, 4, e00467-12. [Google Scholar] [CrossRef] [PubMed]
- Fender, P.; Moriscot, C.; Ruigrok, R.W.; Schoehn, G. Electron Microscopy of Viruses: Techniques to Prepare Viruses and Viral Proteins for Observation by Electron Microscopy. In Reference Module in Biomedical Sciences; Elsevier: Amsterdam, The Netherlands, 2014. [Google Scholar]
- Valentine, R.C.; Wrigley, N.G.; Scrutton, M.C.; Irias, J.J.; Utter, M.F. Pyruvate carboxylase. 8. The subunit structure as examined by electron microscopy. Biochemistry 1966, 5, 3111–3116. [Google Scholar] [PubMed]
- Gerard, F.C.; Ribeiro Ede, A., Jr.; Leyrat, C.; Ivanov, I.; Blondel, D.; Longhi, S.; Ruigrok, R.W.; Jamin, M. Modular organization of rabies virus phosphoprotein. J. Mol. Biol. 2009, 388, 978–996. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Robert, X.; Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 2014, 42, W320–W324. [Google Scholar] [CrossRef] [PubMed]

| Virus | Protein | Method/Temp/[NaCl] | RNA | RNA Size (nt) | Kd (nM) | Reference |
|---|---|---|---|---|---|---|
| Influenza A | ||||||
| PR/8 | virus | FBA/37 °C/100 mM | segment 8 | 890 nt | 23 | [37] |
| PR/8 | virus | FBA/37 °C/100 mM | dsRNA | ds 20 nt | 71 | [37] |
| PR/8 | (MBP)-NP 1 | FBA/100 mM (KCl) | panhandle | 178 nt | 16 | [40] |
| PR/8 | His-NP | FBA/4 °C/100 mM | panhandle | 81 nt | 380 | [41] |
| WSN | His-NP 2 | SPR/25 °C/300 mM | 5′biotin-RNA | 24 nt | 30 | [39] |
| WSN | NP-His 2 | SPR/25 °C/300 mM | 5′biotin-RNA | 24 nt | 41 | [39] |
| WSN | His-NP | SPR/25 °C/300 mM | 5′biotin-RNA | 24 nt | 14 | [39] |
| WSN | NP-His | SPR/25 °C/300 mM | 5′biotin-poly | 8 nt | 70 | [39] |
| WSN | His-NP | SPR/25 °C/300 mM | 5′biotin-RNA | 40 nt | 17 | [39] |
| H5N1 | (MBP)-NP 1 | SPR/25 °C/100 mM | 2′O-methylated | 24 nt | 23 | [25] |
| H5N1 | (MBP)-NP 1 | SPR/25 °C | 2′O-methylated | 24 nt | 16 | [38] |
| WSN | (MBP)-NP 1 | Fluo-pol/RT/200 mM | 5′fluo-RNA | 20 nt | 3.6 | [42] |
| Influenza A mutants | ||||||
| PR/8 | E339A | FBA/4 °C/100 mM | panhandle | 81 nt | 1600 | [41] |
| PR/8 | R416A | FBA/4 °C/100 mM | panhandle | 81 nt | 2600 | [41] |
| WSN | R416A | SPR/25 °C/300 mM | 5′biotin-RNA | 24 nt | 10,000 | [39] |
| H5N1 | E339A | SPR/25 °C | 2′O-methylated | 24 nt | 858 | [38] |
| H5N1 | R416A | SPR/25 °C | 2′O-methylated | 24 nt | 975 | [38] |
| WSN | Δ402-429 | Fluo-pol/RT/200 mM | 5′fluo-RNA | 20 nt | 1.7 | [42] |
| Influenza B | ||||||
| B | MBP-NP | SPR/25 °C/150 mM | 2′O-methylated | 24 nt | 13 | [24] |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Labaronne, A.; Swale, C.; Monod, A.; Schoehn, G.; Crépin, T.; Ruigrok, R.W.H. Binding of RNA by the Nucleoproteins of Influenza Viruses A and B. Viruses 2016, 8, 247. https://doi.org/10.3390/v8090247
Labaronne A, Swale C, Monod A, Schoehn G, Crépin T, Ruigrok RWH. Binding of RNA by the Nucleoproteins of Influenza Viruses A and B. Viruses. 2016; 8(9):247. https://doi.org/10.3390/v8090247
Chicago/Turabian StyleLabaronne, Alice, Christopher Swale, Alexandre Monod, Guy Schoehn, Thibaut Crépin, and Rob W. H. Ruigrok. 2016. "Binding of RNA by the Nucleoproteins of Influenza Viruses A and B" Viruses 8, no. 9: 247. https://doi.org/10.3390/v8090247
APA StyleLabaronne, A., Swale, C., Monod, A., Schoehn, G., Crépin, T., & Ruigrok, R. W. H. (2016). Binding of RNA by the Nucleoproteins of Influenza Viruses A and B. Viruses, 8(9), 247. https://doi.org/10.3390/v8090247






