Correspondence of Neutralizing Humoral Immunity and CD4 T Cell Responses in Long Recovered Sudan Virus Survivors
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Ethics Statement
2.3. Sample Collection
2.4. Antigens and Stimulations
2.5. Internal Control Sera
2.6. Specific IgG Detection Assays
2.7. Normalization of Raw Data and Selection of Cut-off Values
2.8. Plaque Reduction Neutralization Test
2.9. Whole Blood Stimulation from SUDV Survivors and Healthy Volunteers
2.10. Cytokine and Chemokines Detection Using Q-Plex™ ELISA-Based Chemiluminescent Assay
2.11. Flow Cytometry Analysis
2.12. Statistical Analysis
3. Results
3.1. Cohorts and Blood Samples
3.2. Humoral Immune Responses to SUDV Proteins and Neutralization Profiles
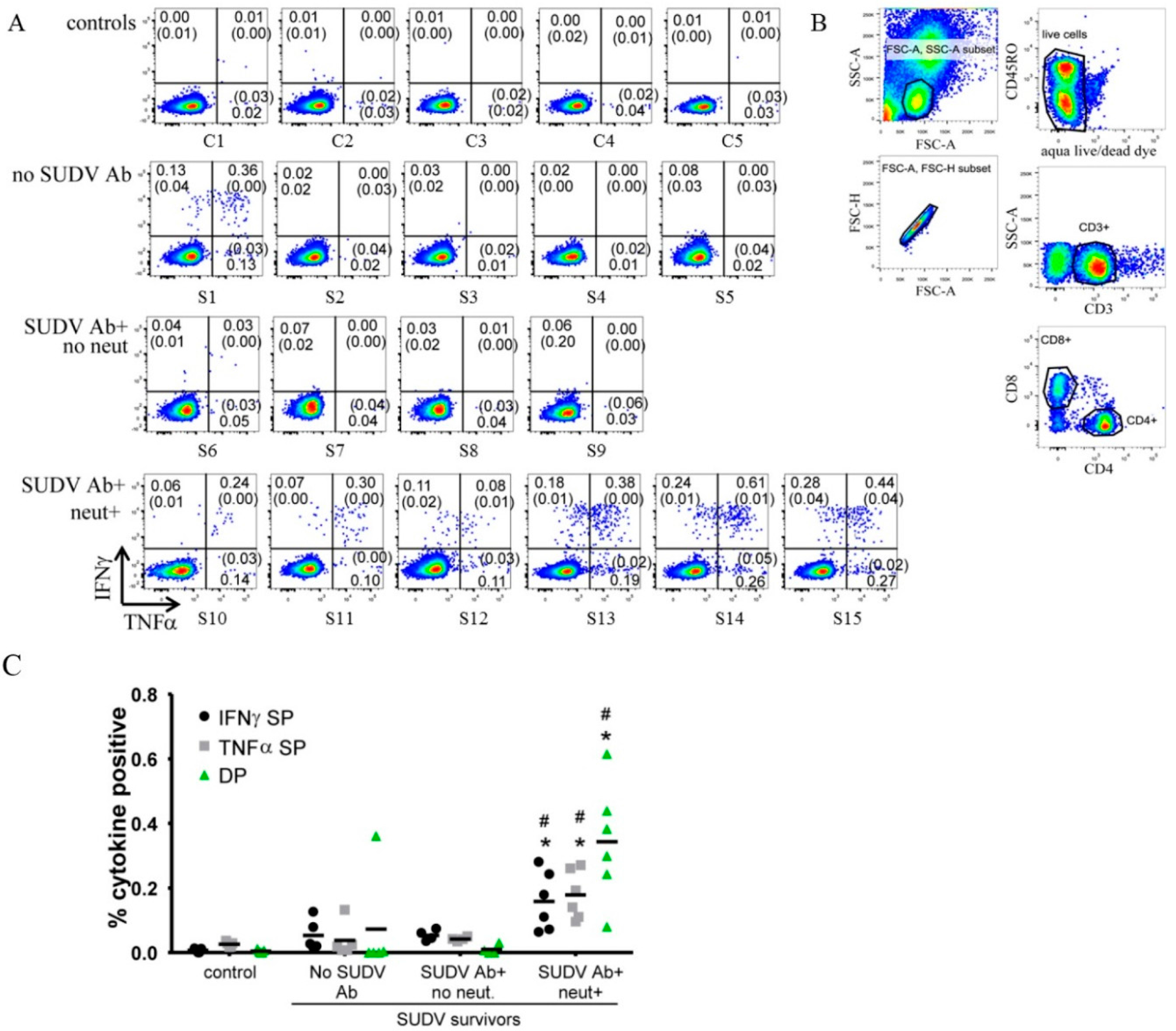
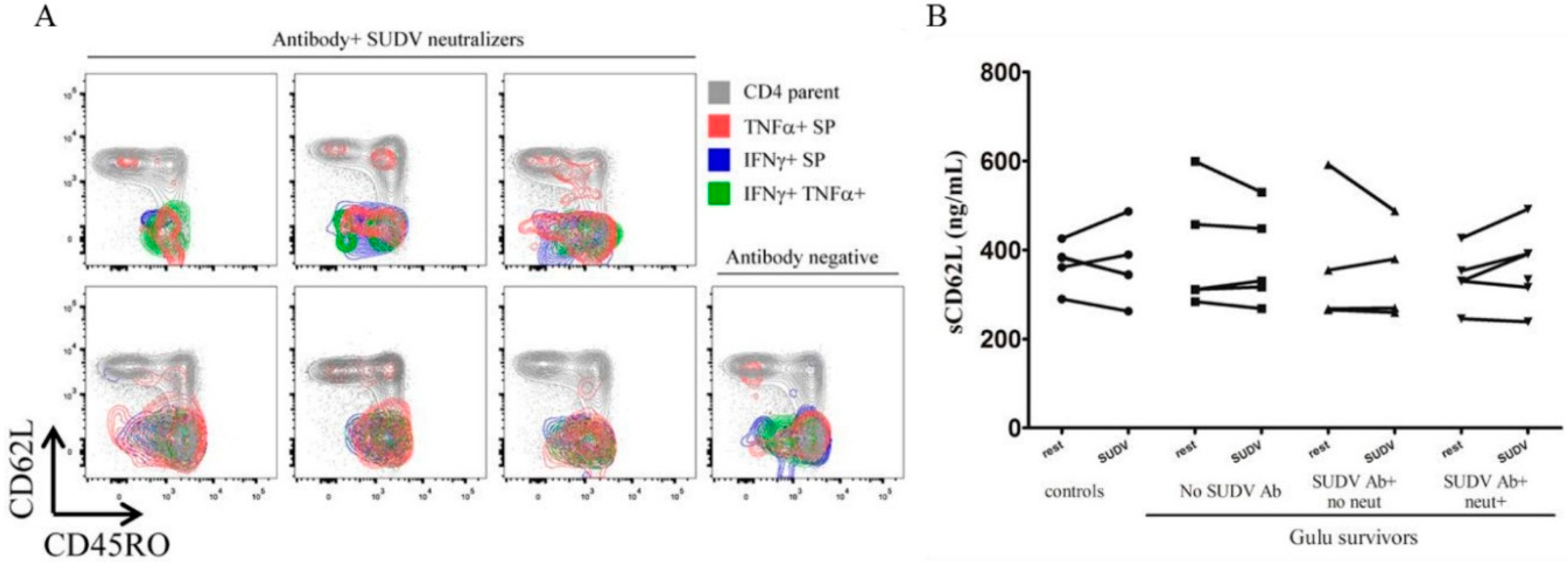
3.3. Flow Cytometry
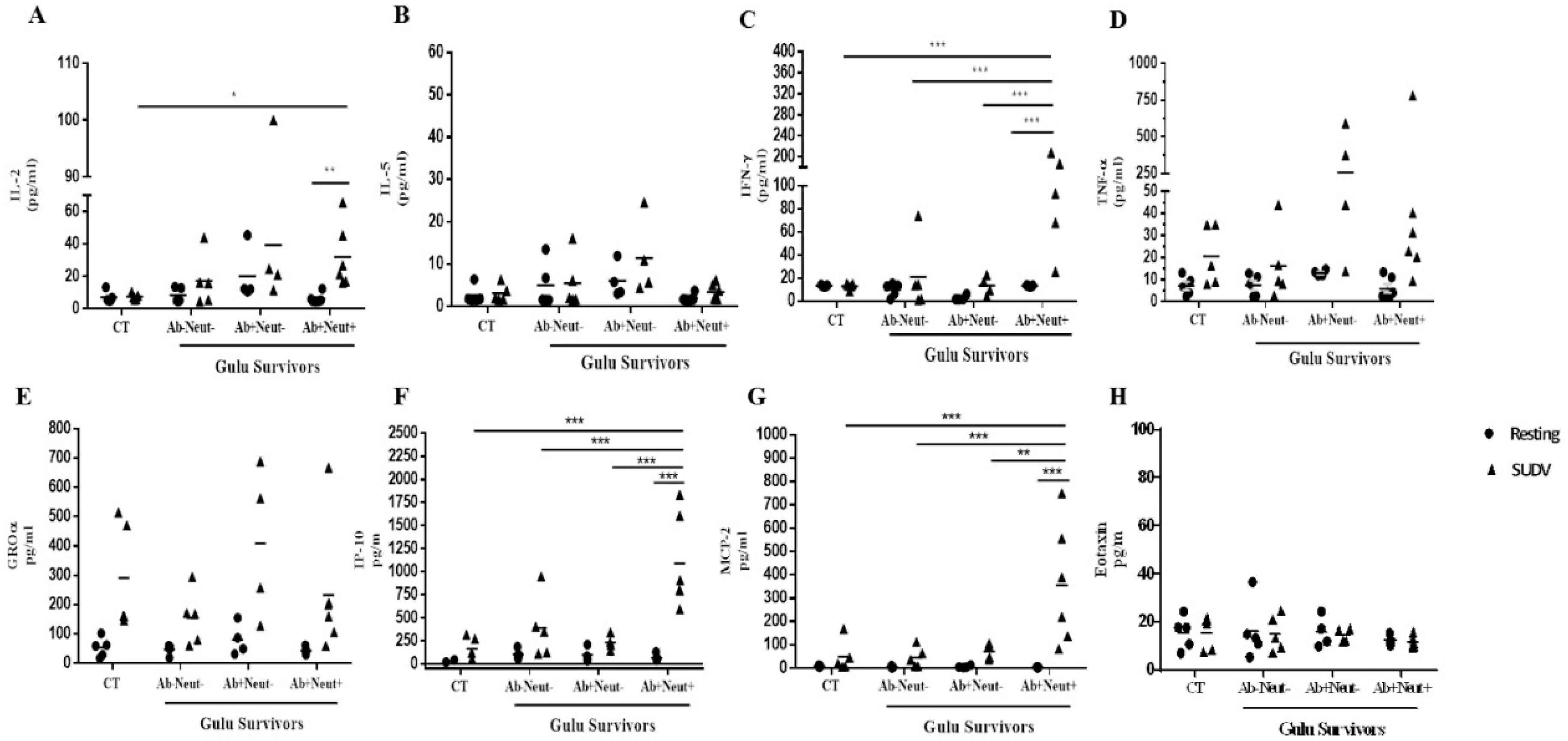
3.4. Cytokine and Chemokine Levels in Whole Blood Stimulation by Multiplex ELISA
4. Discussion
5. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kuhn, J.H. Filoviruses. A Compendium of 40 Years of Epidemiological, Clinical, and Laboratory Studies; Springer-Verlag Wien: New York, NY, USA, 2008. [Google Scholar]
- Mohamadzadeh, M.; Chen, L.; Olinger, G.G.; Pratt, W.D.; Schmaljohn, A.L. Filoviruses and the balance of innate, adaptive, and inflammatory responses. Viral Immunol. 2006, 19, 602–612. [Google Scholar] [CrossRef] [PubMed]
- Mohamadzadeh, M.; Chen, L.; Schmaljohn, A.L. How Ebola and Marburg viruses battle the immune system. Nat. Rev. Immunol. 2007, 7, 556–567. [Google Scholar] [CrossRef] [PubMed]
- Wong, G.; Kobinger, G.P.; Qiu, X. Characterization of host immune responses in Ebola virus infections. Expert Rev. Clin. Immunol. 2014, 10, 781–790. [Google Scholar] [CrossRef] [PubMed]
- McElroy, A.K.; Akondy, R.S.; Davis, C.W.; Ellebedy, A.H.; Mehta, A.K.; Kraft, C.S.; Lyon, G.M.; Ribner, B.S.; Varkey, J.; Sidney, J.; et al. Human Ebola virus infection results in substantial immune activation. Proc. Natl. Acad. Sci. USA 2015, 112, 4719–4724. [Google Scholar] [CrossRef] [PubMed]
- Kondratowicz, A.S.; Maury, W.J. Ebolavirus: A brief review of novel therapeutic targets. Future Microbiol. 2012, 7, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Baize, S.; Leroy, E.M.; Georges, A.J.; Georges-Courbot, M.C.; Capron, M.; Bedjabaga, I.; Lansoud-Soukate, J.; Mavoungou, E. Inflammatory responses in Ebola virus-infected patients. Clin. Exp. Immunol. 2002, 128, 163–168. [Google Scholar] [CrossRef] [PubMed]
- Hutchinson, K.L.; Rollin, P.E. Cytokine and chemokine expression in humans infected with Sudan Ebola Virus. J. Infect. Dis. 2007, 196, S357–S363. [Google Scholar] [CrossRef] [PubMed]
- Bellan, S.E.; Pulliam, J.R.; Dushoff, J.; Meyers, L.A. Ebola control: Effect of asymptomatic infection and acquired immunity. Lancet 2014, 384, 1499–1500. [Google Scholar] [CrossRef]
- Leroy, E.M.; Baize, S.; Volchkov, V.E.; Fisher-Hoch, S.P.; Georges-Courbot, M.C.; Lansoud-Soukate, J.; Capron, M.; Debre, P.; McCormick, J.B.; Georges, A.J. Human asymptomatic Ebola infection and strong inflammatory response. Lancet 2000, 355, 2210–2215. [Google Scholar] [CrossRef]
- Dixon, M.G.; Schafer, I.J. Ebola viral disease outbreak—West Africa, 2014. MMWR Morb. Mortal. Wkly. Rep. 2014, 63, 548–551. [Google Scholar] [PubMed]
- Labouba, I.; Leroy, E.M. Ebola outbreaks in 2014. J. Clin. Virol. 2015, 64, 109–110. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, J.H.; Dodd, L.E.; Wahl-Jensen, V.; Radoshitzky, S.R.; Bavari, S.; Jahrling, P.B. Evaluation of perceived threat differences posed by filovirus variants. Biosecur. Bioterror. 2011, 9, 361–371. [Google Scholar] [CrossRef] [PubMed]
- Sobarzo, A.; Ochayon, D.E.; Lutwama, J.J.; Balinandi, S.; Guttman, O.; Marks, R.S.; Kuehne, A.I.; Dye, J.M.; Yavelsky, V.; Lewis, E.C.; et al. Persistent immune responses after Ebola virus infection. N. Engl. J. Med. 2013, 369, 492–493. [Google Scholar] [CrossRef] [PubMed]
- Sobarzo, A.; Eskira, Y.; Herbert, A.S.; Kuehne, A.I.; Stonier, S.W.; Ochayon, D.E.; Fedida-Metula, S.; Balinandi, S.; Kislev, Y.; Tali, N.; et al. Immune memory to Sudan virus: Comparison between two separate disease outbreaks. Viruses 2015, 7, 37–51. [Google Scholar] [CrossRef] [PubMed]
- Sobarzo, A.; Groseth, A.; Dolnik, O.; Becker, S.; Lutwama, J.J.; Perelman, E.; Yavelsky, V.; Muhammad, M.; Kuehne, A.I.; Marks, R.S.; et al. Profile and persistence of the virus-specific neutralizing humoral immune response in human survivors of Sudan Ebolavirus (Gulu). J. Infect. Dis. 2013, 208, 299–309. [Google Scholar] [CrossRef] [PubMed]
- Lamunu, M.; Lutwama, J.J.; Kamugisha, J.; Opio, A.; Nambooze, J.; Ndayimirije, N.; Okware, S. Containing a haemorrhagic fever epidemic: The Ebola experience in Uganda (October 2000–January 2001). Int. J. Infect. Dis. 2004, 8, 27–37. [Google Scholar] [CrossRef] [PubMed]
- Sobarzo, A.; Perelman, E.; Groseth, A.; Dolnik, O.; Becker, S.; Lutwama, J.J.; Dye, J.M.; Yavelsky, V.; Lobel, L.; Marks, R.S. Profiling the native specific human humoral immune response to Sudan Ebola virus strain Gulu by chemiluminescence enzyme-linked immunosorbent assay. Clin. Vaccine Immunol. 2012, 19, 1844–1852. [Google Scholar] [CrossRef] [PubMed]
- Dias, J.M.; Kuehne, A.I.; Abelson, D.M.; Bale, S.; Wong, A.C.; Halfmann, P.; Muhammad, M.A.; Fusco, M.L.; Zak, S.E.; Kang, E.; et al. A shared structural solution for neutralizing Ebolaviruses. Nat. Struct. Mol. Biol. 2011, 18, 1424–1427. [Google Scholar] [CrossRef] [PubMed]
- Kohavi, R. A study of cross. In Proceedings of the Appears in the International Joint Conference on Artifcial Intelligence (IJCAI), Montreal, QC, Canada, 20–25 August 1995.
- Paweska, J.T.; Burt, F.J.; Swanepoel, R. Validation of IgG-sandwich and IgM-capture ELISA for the detection of antibody to Rift Valley fever virus in humans. J. Virol. Methods 2005, 124, 173–181. [Google Scholar] [CrossRef] [PubMed]
- Swenson, D.L.; Wang, D.; Luo, M.; Warfield, K.L.; Woraratanadharm, J.; Holman, D.H.; Dong, J.Y.; Pratt, W.D. Vaccine to confer to nonhuman primates complete protection against multistrain Ebola and Marburg virus infections. Clin. Vaccine Immunol. 2008, 15, 460–467. [Google Scholar] [CrossRef] [PubMed]
- Cavelti-Weder, C.; Babians-Brunner, A.; Keller, C.; Stahel, M.A.; Kurz-Levin, M.; Zayed, H.; Solinger, A.M.; Mandrup-Poulsen, T.; Dinarello, C.A.; Donath, M.Y. Effects of gevokizumab on glycemia and inflammatory markers in type 2 diabetes. Diabetes Care 2012, 35, 1654–1662. [Google Scholar] [CrossRef] [PubMed]
- Feldmann, H.; Geisbert, T.W. Ebola haemorrhagic fever. Lancet 2011, 377, 849–862. [Google Scholar] [CrossRef]
- Clark, D.V.; Kibuuka, H.; Millard, M.; Wakabi, S.; Lukwago, L.; Taylor, A.; Eller, M.A.; Eller, L.A.; Michael, N.L.; Honko, A.N.; et al. Long-term sequelae after Ebola virus disease in Bundibugyo, Uganda: A retrospective cohort study. Lancet Infect. Dis. 2015, 15, 905–912. [Google Scholar] [CrossRef]
- Varkey, J.B.; Shantha, J.G.; Crozier, I.; Kraft, C.S.; Lyon, G.M.; Mehta, A.K.; Kumar, G.; Smith, J.R.; Kainulainen, M.H.; Whitmer, S.; et al. Persistence of Ebola virus in ocular fluid during convalescence. N. Engl. J. Med. 2015, 372, 2423–2427. [Google Scholar] [CrossRef] [PubMed]
- Griffith, J.W.; Sokol, C.L.; Luster, A.D. Chemokines and chemokine receptors: Positioning cells for host defense and immunity. Annu. Rev. Immunol. 2014, 32, 659–702. [Google Scholar] [CrossRef] [PubMed]
- Mackay, C.R. Chemokine receptors and T cell chemotaxis. J. Exp. Med. 1996, 184, 799–802. [Google Scholar] [CrossRef] [PubMed]

| Serology | ||||||
|---|---|---|---|---|---|---|
| NP | VP40 | VP30 | GP1–649 1 | SUDV | PRNT80 | |
| C1 | − | − | − | − | − | − |
| C2 | − | − | − | − | − | − |
| C3 | − | − | − | − | − | − |
| C4 | − | − | − | − | − | − |
| C5 | − | − | − | − | − | − |
| S1 | ++ | − | + | − | − | − |
| S2 | − | + | − | − | − | − |
| S3 | − | − | − | − | − | − |
| S4 | − | − | − | − | − | − |
| S5 | − | − | − | − | − | − |
| S6 | +++ | − | + | − | +++ | − |
| S7 | + | − | + | + | − | − |
| S8 | +++ | + | − | +++ | − | − |
| S9 | ++ | + | − | +++ | +++ | − |
| S10 | +++ | +++ | − | +++ | +++ | +++ |
| S11 | +++ | ++ | − | +++ | − | + |
| S12 | +++ | +++ | − | +++ | + | + |
| S13 | +++ | +++ | + | +++ | +++ | +++ |
| S14 | +++ | + | − | +++ | +++ | +++ |
| S15 | +++ | +++ | − | +++ | − | ++ |
| Multiplex ELISA | Flow | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| IL-2 | IL-5 | IFNγ | TNFα | GROα | IP-10 | MCP-2 | Eotaxin | IFNγ | TNFα | TNFα + IFNγ | ||
| Spearman r | 0.4672 | 0.0704 | 0.6892 | 0.2141 | 0.0778 | 0.6999 | 0.6860 | 0.2577 | 0.6714 | 0.7672 | 0.7561 | |
| p value (two side) | 0.0378 | 0.7679 | 0.0011 | 0.3646 | 0.7442 | 0.0006 | 0.0008 | 0.2727 | 0.0012 | <0.0001 | 0.0001 | |
| Cytokine Secretion | Chemokine Secretion | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| IL-2 | IL-5 | IFNγ | TNFα | GROα | IP-10 | MCP-2 | Eotaxin | ||
| C1 | − | − | − | − | − | − | ++ | − | |
| C2 | − | − | − | − | − | − | − | − | |
| C3 | − | − | − | − | − | − | − | − | |
| C4 | − | − | − | − | − | − | − | − | |
| C5 | − | − | − | − | − | − | − | − | |
| S1 | ++ | − | ++ | − | − | ++ | ++ | + | |
| S2 | − | − | − | − | − | − | − | − | |
| S3 | − | − | − | − | − | − | − | − | |
| S4 | + | − | − | − | − | − | + | − | |
| S5 | + | ++ | − | − | − | − | − | − | |
| S6 | − | − | − | − | − | − | − | − | |
| S7 | + | ++ | − | +++ | + | − | − | − | |
| S8 | +++ | + | + | +++ | − | − | + | − | |
| S9 | + | − | − | − | − | − | + | − | |
| S10 | + | − | + | − | − | + | ++ | − | |
| S11 | + | − | +++ | − | − | + | ++ | − | |
| S12 | +++ | − | +++ | − | − | + | + | − | |
| S13 | ++ | − | +++ | − | − | ++ | +++ | ++ | |
| S14 | ++ | − | ++ | +++ | − | +++ | +++ | − | |
| S15 | + | − | ND | − | + | ++ | +++ | − | |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sobarzo, A.; Stonier, S.W.; Herbert, A.S.; Ochayon, D.E.; Kuehne, A.I.; Eskira, Y.; Fedida-Metula, S.; Tali, N.; Lewis, E.C.; Egesa, M.; et al. Correspondence of Neutralizing Humoral Immunity and CD4 T Cell Responses in Long Recovered Sudan Virus Survivors. Viruses 2016, 8, 133. https://doi.org/10.3390/v8050133
Sobarzo A, Stonier SW, Herbert AS, Ochayon DE, Kuehne AI, Eskira Y, Fedida-Metula S, Tali N, Lewis EC, Egesa M, et al. Correspondence of Neutralizing Humoral Immunity and CD4 T Cell Responses in Long Recovered Sudan Virus Survivors. Viruses. 2016; 8(5):133. https://doi.org/10.3390/v8050133
Chicago/Turabian StyleSobarzo, Ariel, Spencer W. Stonier, Andrew S. Herbert, David E. Ochayon, Ana I. Kuehne, Yael Eskira, Shlomit Fedida-Metula, Neta Tali, Eli C. Lewis, Moses Egesa, and et al. 2016. "Correspondence of Neutralizing Humoral Immunity and CD4 T Cell Responses in Long Recovered Sudan Virus Survivors" Viruses 8, no. 5: 133. https://doi.org/10.3390/v8050133
APA StyleSobarzo, A., Stonier, S. W., Herbert, A. S., Ochayon, D. E., Kuehne, A. I., Eskira, Y., Fedida-Metula, S., Tali, N., Lewis, E. C., Egesa, M., Cose, S., Lutwama, J. J., Yavelsky, V., Dye, J. M., & Lobel, L. (2016). Correspondence of Neutralizing Humoral Immunity and CD4 T Cell Responses in Long Recovered Sudan Virus Survivors. Viruses, 8(5), 133. https://doi.org/10.3390/v8050133






