Isolation of a Novel Fusogenic Orthoreovirus from Eucampsipoda africana Bat Flies in South Africa
Abstract
:1. Introduction
2. Materials and Methods
2.1. Ethics and Permit Statement
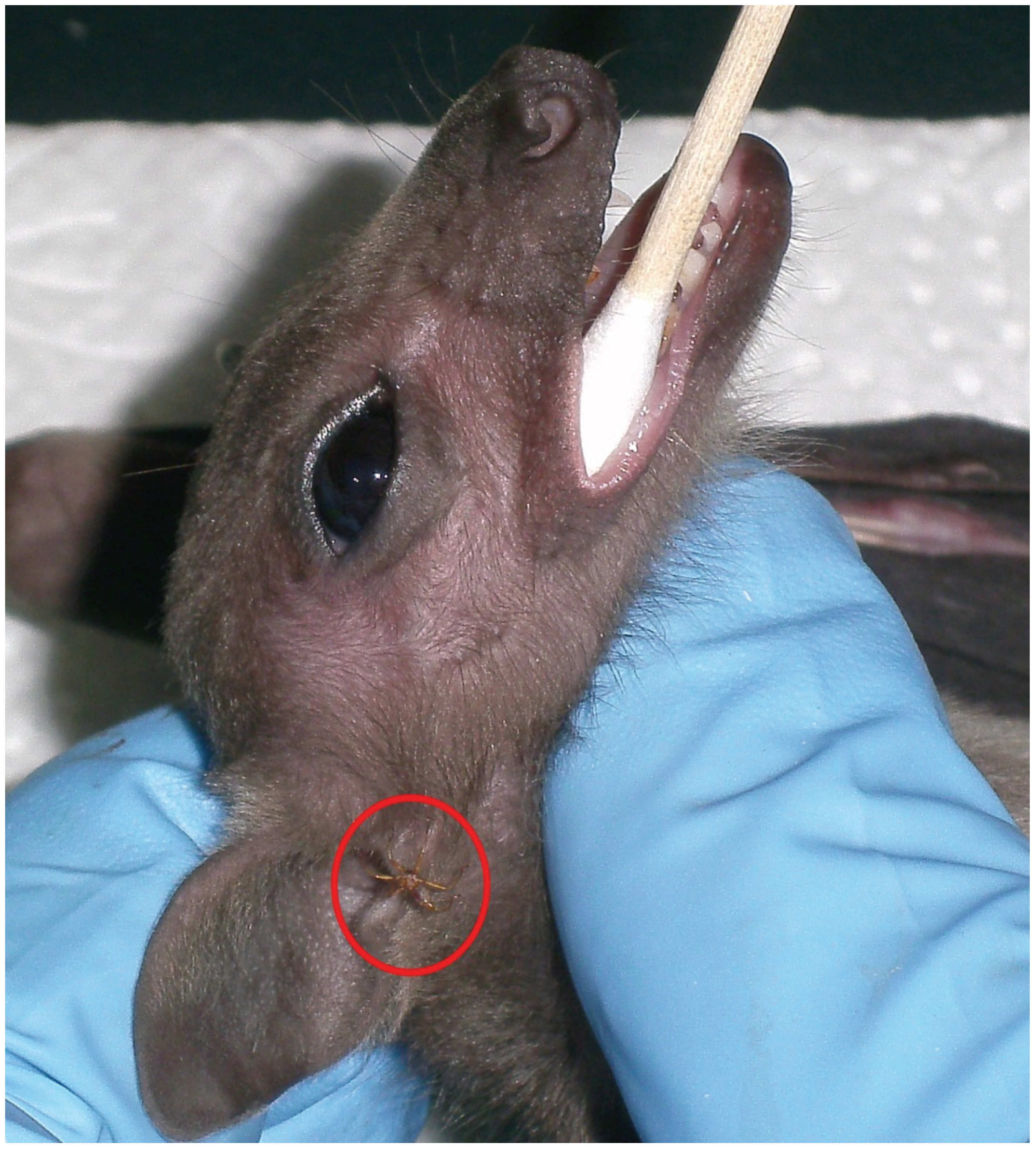
2.2. Sample Collection and Processing
2.3. Virus Isolation and Titration
2.4. Transmission Electron Microscopy (TEM)
2.5. Sequence-Independent Single-Primer Amplification (SISPA), Rapid Amplification of cDNA Ends (RACE), Next-Generation Sequencing (NGS) and Bioinformatics
2.6. Virus Phylogenetic, Genome and Protein Sequence Analysis
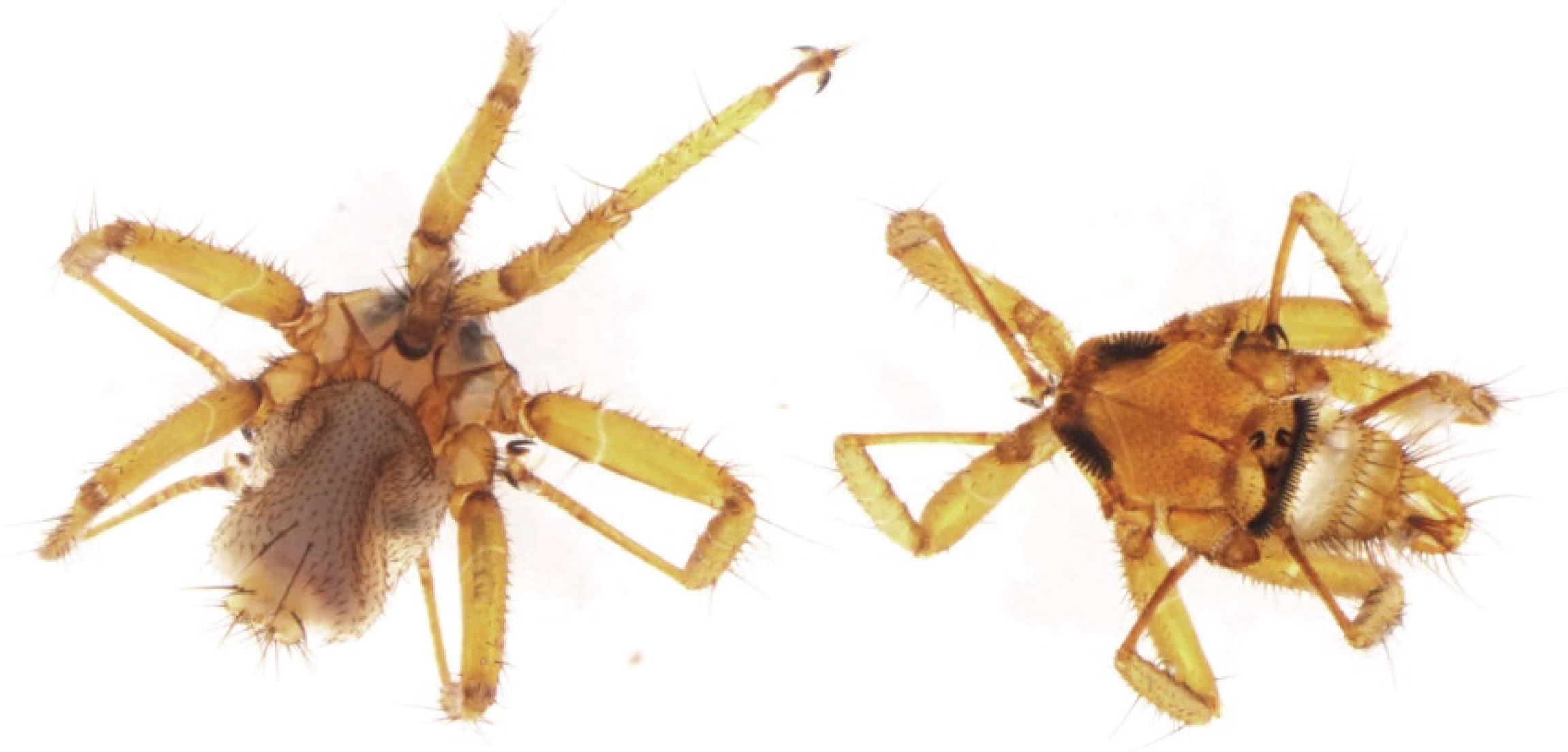
2.7. Phylogenetic Analysis of the Bat Fly
2.8. Virus Growth Curves
3. Results
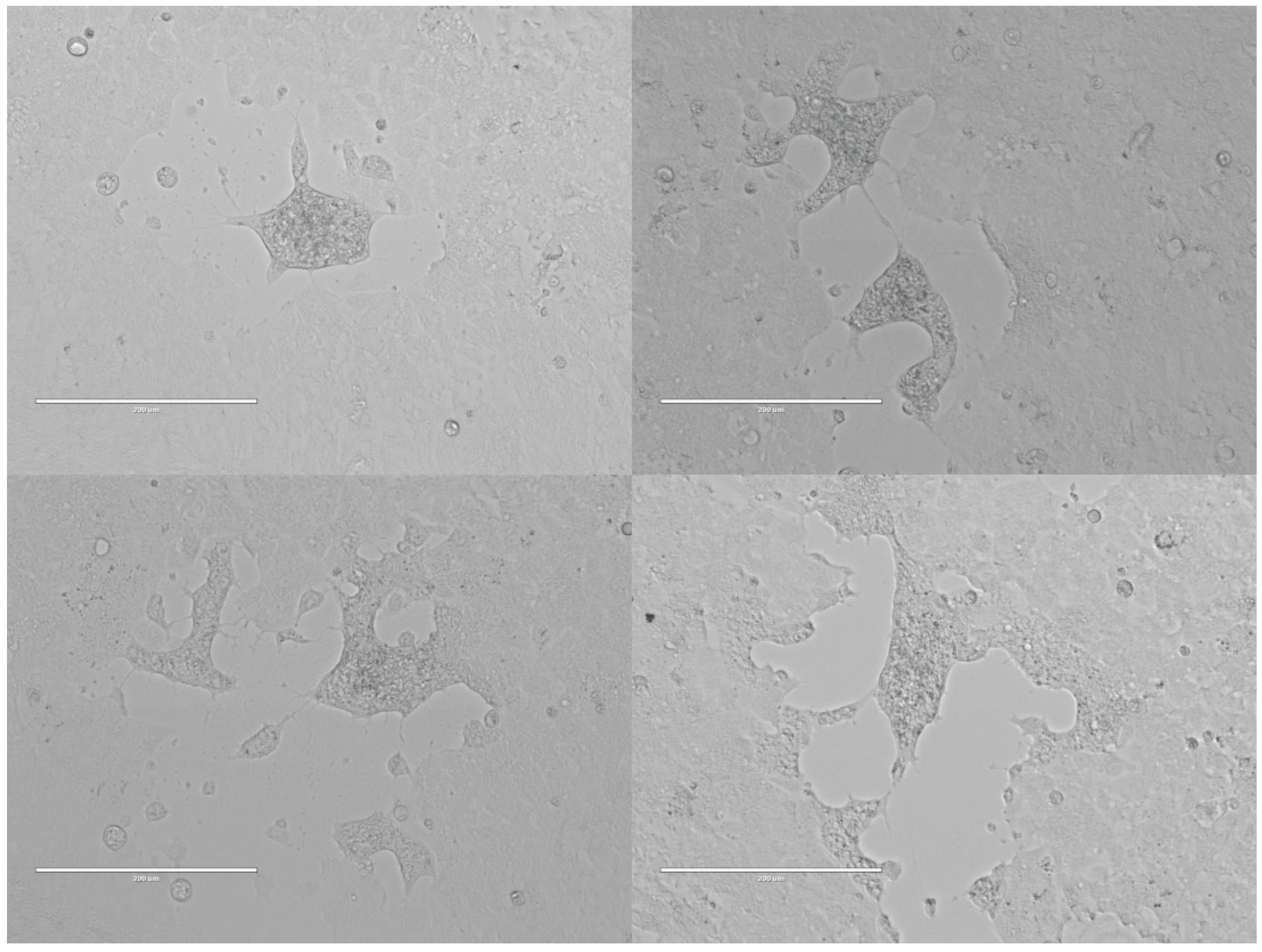
3.1. Isolation of a Syncytia-Forming Virus from Bat Ectoparasite Pools
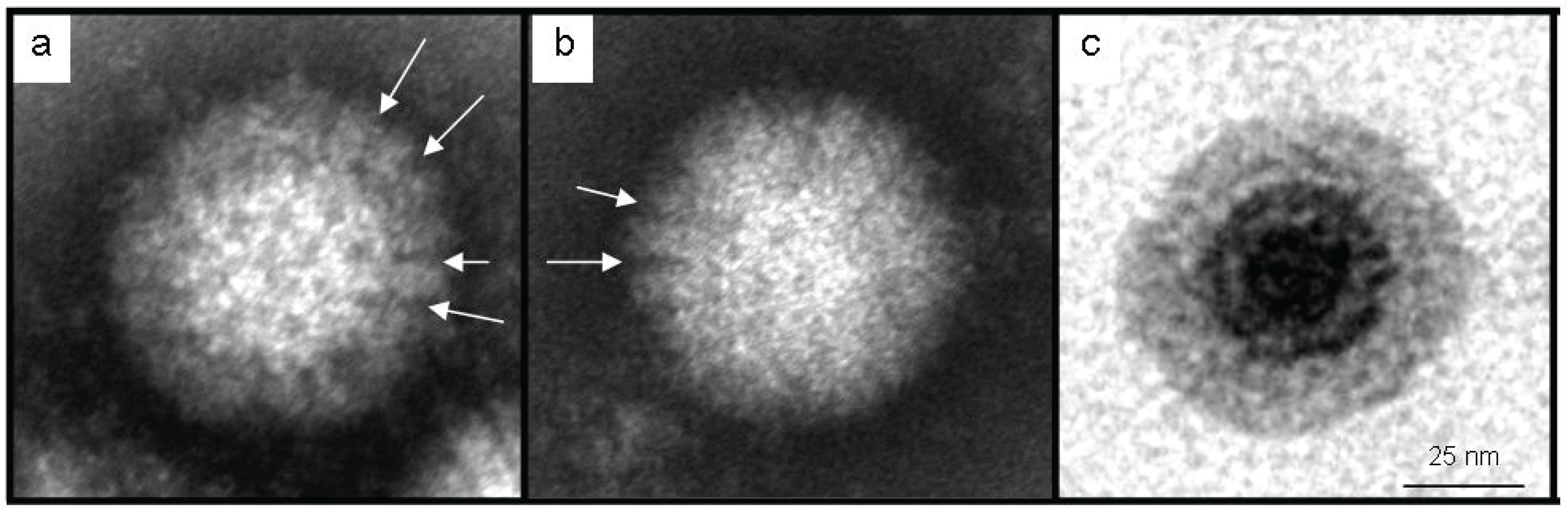
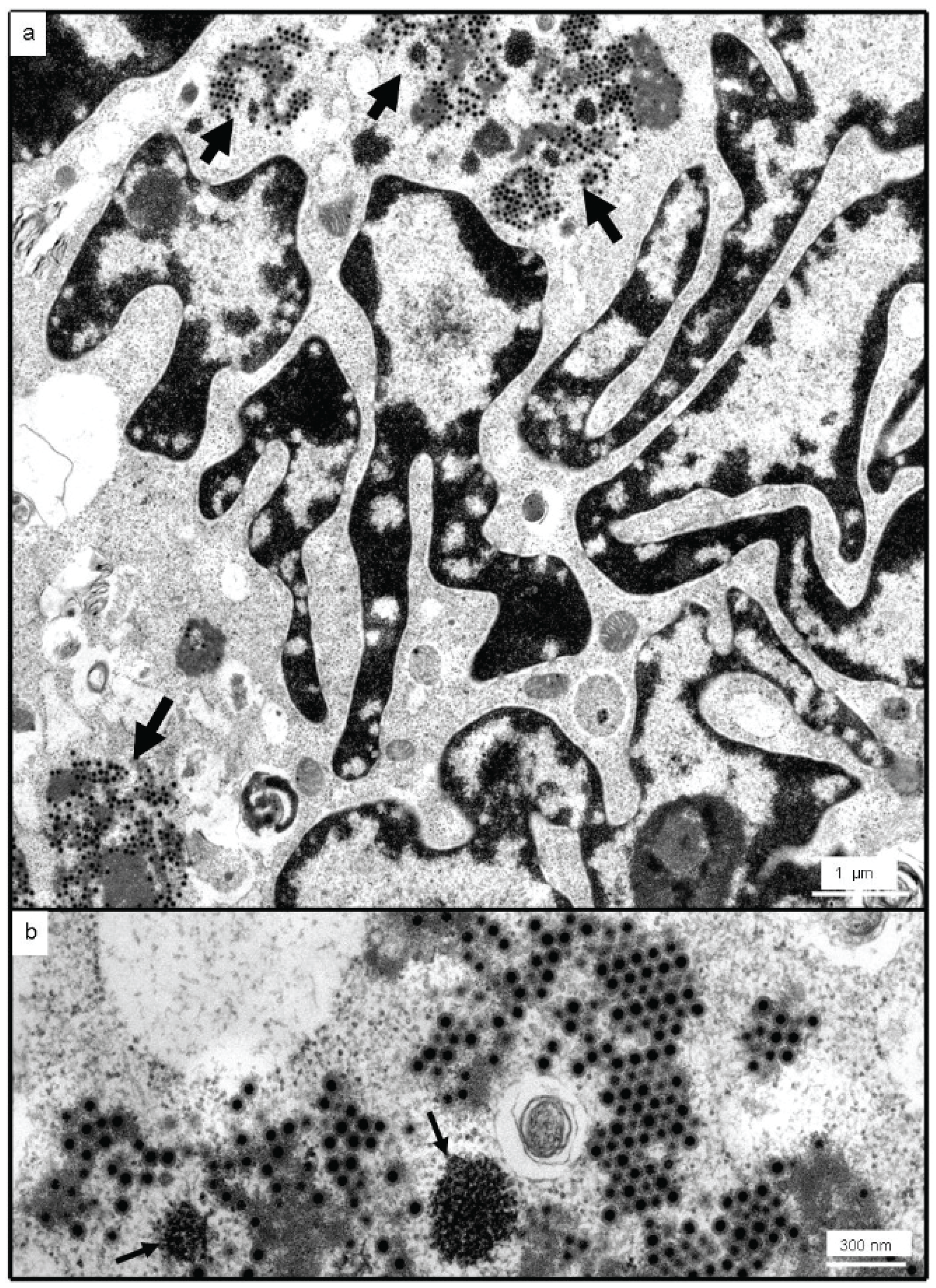
3.2. Identification as an Orthoreovirus by TEM and SISPA-NGS
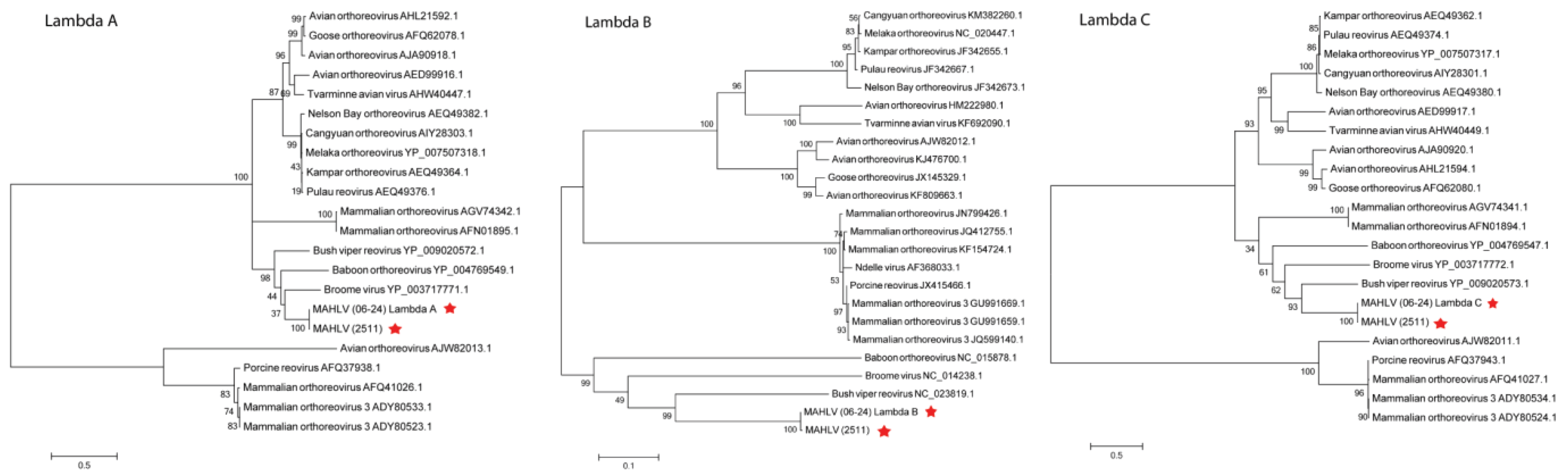
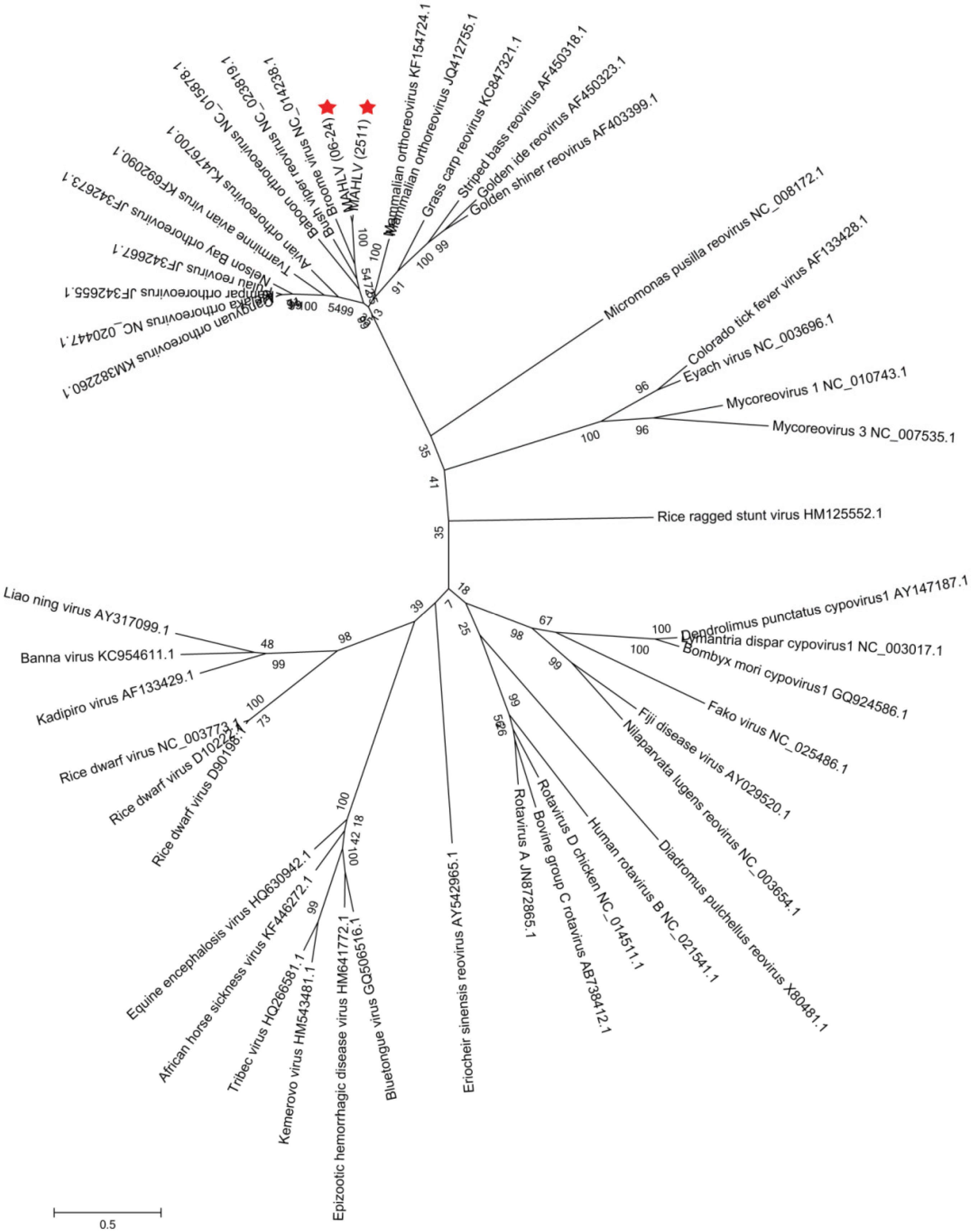
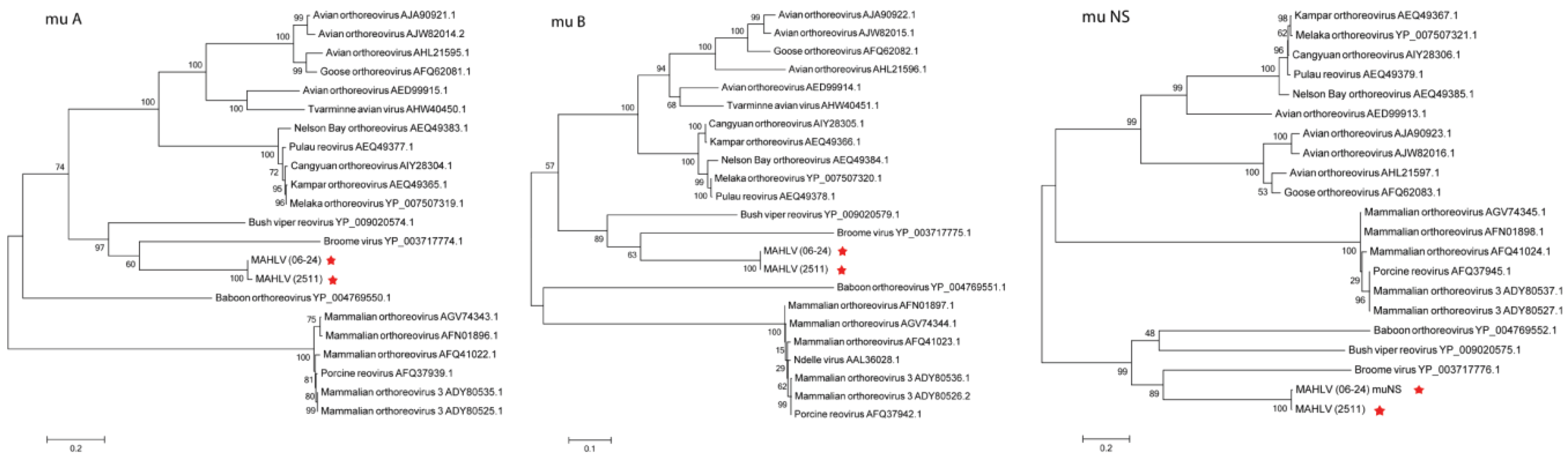
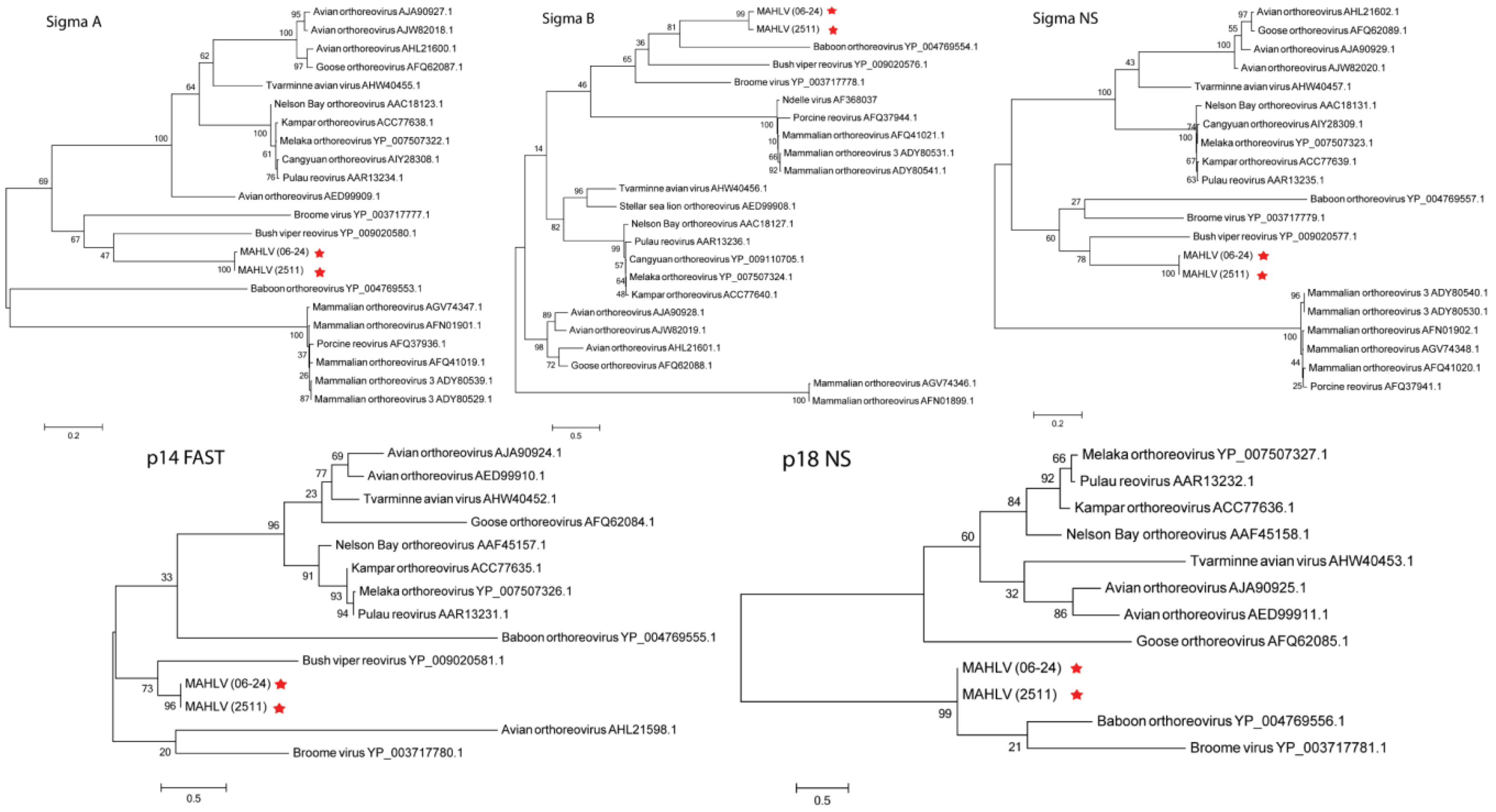
3.3. Genome Analysis and Phylogeny Reveals a Putative New African Orthoreovirus Species
3.4. Virus Growth in Mammalian and Insect Cells in Vitro
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Leroy, E.M.; Kumulungui, B.; Pourrut, X.; Rouquet, P.; Hassanin, A.; Yaba, P.; Délicat, A.; Paweska, J.T.; Gonzalez, J.P.; Swanepoel, R. Fruit bats as reservoirs of Ebola virus. Nature 2005, 438, 575–576. [Google Scholar] [CrossRef] [PubMed]
- Towner, J.S.; Pourrut, X.; Albariño, C.G.; Nkoque, C.N.; Bird, B.H.; Grard, G.; Ksiazek, T.G.; Gonzalez, J.P.; Nichol, S.T.; Leroy, E.M. Marburg virus infection detected in a common African bat. PLoS ONE 2007, 2, e764. [Google Scholar]
- Li, W.; Shi, Z.; Yu, M.; Ren, W.; Smith, C.; Epstein, J.H.; Wang, H.; Crameri, G.; Hu, Z.; Zhang, H.; et al. Bats are natural reservoirs of SARS-like coronaviruses. Science 2005, 310, 676–679. [Google Scholar] [CrossRef] [PubMed]
- Memish, Z.A.; Mishra, N.; Olival, K.J.; Fagbo, S.F.; Kapoor, V.; Epstein, J.H.; Alhakeem, R.; Durosinloun, A.; Al Asmari, M.; Islam, A.; et al. Middle East respiratory syndrome coronavirus in bats, Saudi Arabia. Emerg. Infect. Dis. 2013, 19, 1819–1823. [Google Scholar] [PubMed]
- Chua, K.B.; Bellini, W.J.; Rota, P.A.; Harcourt, B.H.; Tamin, A.; Lam, S.K.; Ksiazek, T.G.; Rollin, P.E.; Zaki, S.R.; Shieh, W.; et al. Nipah virus: A recently emergent deadly paramyxovirus. Science 2000, 288, 1432–1435. [Google Scholar] [CrossRef] [PubMed]
- Halpin, K.; Young, P.L.; Field, H.E.; Mackenzie, J.S. Isolation of Hendra virus from pteropid bats: A natural reservoir of Hendra virus. J. Gen. Virol. 2000, 81, 1927–1932. [Google Scholar] [CrossRef] [PubMed]
- Enright, J.B.; Sadler, W.W.; Moulton, J.E.; Constantine, D. Isolation of rabies virus from an insectivorous bat (Tadarida mexicana) in California. Proc. Soc. Exp. Biol. Med. 1955, 89, 94–96. [Google Scholar] [CrossRef] [PubMed]
- Van Thiel, P.P.; van den Hoek, J.A.; Eftimov, F.; Tepaske, R.; Zaaijer, H.J.; Spanjaard, L.; de Boer, H.E.; van Doornum, G.J.; Schutten, M.; Osterhaus, A.; et al. Fatal case of human rabies (Duvenhage virus) from a bat in Kenya: The Netherlands, December 2007. Eurosurveillance 2008, 13, 8007. Available online: http://www.eurosurveillance.org/ViewArticle.aspx?ArticleId=8007 (accessed on 25 February 2016). [Google Scholar] [PubMed]
- Sano, K.; Okazaki, S.; Taniguchi, S.; Masangkay, J.S.; Puentespina, R., Jr.; Eres, E.; Cosico, E.; Quibod, N.; Kondo, T.; Shimoda, H.; et al. Detection of a novel herpesvirus from bats in the Philippines. Virus Genes 2015. [Google Scholar] [CrossRef]
- Binger, T.; Annan, A.; Drexler, J.F.; Müller, M.A.; Kallies, R.; Adankwah, E.; Wollny, R.; Kopp, A.; Heidemann, H.; Dei, D.; et al. A novel rhabdovirus isolated from the straw-colored fruit bat Eidolon helvum, with signs of antibodies in swine and humans. J. Virol. 2015, 89, 4588–4597. [Google Scholar] [CrossRef] [PubMed]
- Kading, R.C.; Gilbert, A.T.; Mossel, E.C.; Crabtree, M.B.; Kuzmin, I.V.; Niezgoda, M.; Agwanda, B.; Markotter, W.; Weil, M.R.; Montgomery, J.M.; et al. Isolation and molecular characterization of Fikirini rhabdovirus, a novel virus from a Kenyan bat. J. Gen. Virol. 2013, 94, 2393–2398. [Google Scholar] [CrossRef] [PubMed]
- Calisher, C.H.; Childs, J.E.; Field, H.E.; Holmes, K.V.; Schountz, T. Bats: Important reservoir hosts of emerging viruses. Clin. Microbiol. Rev. 2006, 19, 531–545. [Google Scholar] [CrossRef] [PubMed]
- Tortosa, P.; Dsouli, N.; Gomard, Y.; Ramasindrazana, B.; Dick, C.W.; Goodman, S.M. Evolutionary history of Indian Ocean Nycteribiid bat flies mirroring the ecology of their hosts. PLoS ONE 2013, 8, e75215. [Google Scholar]
- Bertola, P.B.; Aires, C.C.; Favorito, S.E.; Graciolli, G.; Amaku, M.; Pinto-da-Rocha, R. Bat flies (Diptera: Streblidae, Nycteribiidae) parasitic on bats (Mammalia: Chiroptera) at Parque da Cantareira, Sao Paulo, Brazil: Parasitism rates and host-parasite associations. Mem. Inst. Oswaldo. Cruz. 2005, 100, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Dick, C.W.; Patterson, B.D. Bat flies–obligate ectoparasites of bats. In Micromammals and Macroparasites. From Evolutionary Ecology to Management; Morand, S., Krasnov, B.R., Poulin, R., Eds.; Springer: Tokyo, Japan, 2006; pp. 179–194. [Google Scholar]
- Brook, C.E.; Bai, Y.; Dobson, A.P.; Osikowicz, L.M.; Ranaivoson, H.C.; Zhu, Q.; Kosoy, M.Y.; Dittmar, K. Bartonella spp. in fruit bats and blood-feeding ectoparasites in Madagascar. PLoS Negl. Trop. Dis. 2015, 10, e0003532. [Google Scholar] [CrossRef] [PubMed]
- Morse, S.F.; Olival, K.J.; Kosoy, M.; Billeter, S.A.; Patterson, B.D.; Dick, C.W.; Dittmar, K. Global distribution and genetic diversity of Bartonella in bat flies (Hippoboscoidea, Streblidae, Nycteribiidae). Infect. Genet. Evol. 2012, 12, 1717–1723. [Google Scholar] [CrossRef] [PubMed]
- Aznar-Lopez, C.; Vasquez-Moron, S.; Marston, D.A.; Juste, J.; Ibáñez, C.; Miguel Berciano, J.; Salsamendi, E.; Aihartza, J.; Banyard, A.; McElhinney, L.; et al. Detection of rhabdovirus viral RNA in oropharyngeal swabs and ectoparasites of Spanish bats. J. Gen. Virol. 2013, 94, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Attoui, H.; Mertens, P.P.C.; Becnel, J.; Belaganahalli, S.; Bergoin, M.; Brussaard, C.P.; Chappell, J.D.; Ciarlet, M.; del Vas, M.; Dermody, T.S.; et al. Reoviridae. In Virus Taxonomy. Classification and Nomenclature of Viruses: Ninth Report of the International Committee on Taxonomy of Viruses; King, A.M.Q., Adams, M.J., Carstens, E.B., Lefkowitz, E.J., Eds.; Elsevier Academic Press: San Diego, CA, USA, 2012. [Google Scholar]
- Lorusso, A.; Teodori, L.; Leone, A.; Marcacci, M.; Mangone, I.; Orsini, M.; Capobianco-Dondona, A.; Camma, C.; Monaco, F.; Savini, G. A new member of the Pteropine Orthoreovirus species isolated from fruit bats imported to Italy. Inf. Gen. Evol. 2015, 30, 55–58. [Google Scholar]
- Gard, G.; Compans, R.W. Structure and cytopathic effects of Nelson Bay virus. J. Virol. 1970, 6, 100–106. [Google Scholar] [PubMed]
- Chua, K.B.; Crameri, G.; Hyatt, A.; Yu, M.; Tompang, M.R.; Rosli, J.; McEachern, J.; Crameri, S.; Kumurasamy, V.; Eaton, B.T.; et al. A previously unknown reovirus of bat origin is associated with an acute respiratory disease in humans. Proc. Nat. Acad. Sci. USA 2007, 104, 11424–11429. [Google Scholar] [CrossRef] [PubMed]
- Chua, K.B.; Voon, K.; Crameri, G.; Tan, H.S.; Rosli, J.; McEachern, J.; Suluraju, S.; Yu, M.; Wang, L. Identification and characterization of a new orthoreovirus from patients with acute respiratory infections. PLoS ONE 2008, 3, e3803. [Google Scholar]
- Kohl, C.; Lesnik, R.; Brinkmann, A.; Ebinger, A.; Radonić, A.; Nitsche, A.; Mühldorfer, K.; Wibbelt, G.; Kurth, A. Isolation and characterization of three mammalian orthoreoviruses from European bats. PLoS ONE 2012, 7, e43106. [Google Scholar]
- Wang, L.; Fu, S.; Cao, L.; Lei, W.; Cao, Y.; Song, J.; Tang, Q.; Zhang, H.; Feng, Y.; Yang, W.; et al. Isolation and identification of a natural reassortant mammalian orthoreovirus from Least Horseshoe bat in China. PLoS ONE 2015, 10, e0118598. [Google Scholar]
- Thalmann, C.M.; Cummins, D.M.; Yu, M.; Lunt, R.; Pritchard, L.I.; Hansson, E.; Crameri, S.; Hyatt, A.; Wang, L. Broome virus, a new fusogenic Orthoreovirus species isolated from an Australian fruit bat. Virology 2010, 402, 26–40. [Google Scholar] [CrossRef] [PubMed]
- Ogasawara, Y.; Ueda, H.; Kikuchi, N.; Kirisawa, R. Isolation and genomic characterization of a novel orthoreovirus from a brown-eared bulbul (Hypsipetes amaurotis) in Japan. J. Gen. Virol. 2015. [Google Scholar] [CrossRef]
- Palacios, G.; Wellehan, J.F.X., Jr.; Raverty, S.; Bussetti, A.V.; Hui, J.; Savji, N.; Nollens, H.N.; Lambourn, D.; Celone, C.; Hutchison, S.; et al. Discovery of an orthoreovirus in the aborted fetus of a Stellar sea lion (Eumetopias jubatus). J. Gen. Virol. 2011, 92, 2558–2565. [Google Scholar] [CrossRef] [PubMed]
- Jones, R.C. Avian reovirus infections. Rev. Sci. Tech. 2000, 19, 614–625. [Google Scholar] [PubMed]
- Steyer, A.; Gutiérrez-Aguire, I.; Kolenc, M.; Koren, S.; Kutnjak, D.; Pokorn, M.; Poljšak-Prijatelj, M.; Rački, N.; Ravnikar, M.; Sagadin, M.; et al. High similarity of novel orthoreovirus detected in a child hospitalized with acute gastroenteritis to mammalian orthoreoviruses found in bats in Europe. J. Clin. Microbiol. 2013, 51, 3818–3825. [Google Scholar] [CrossRef] [PubMed]
- Duncan, R.; Corcoran, J.; Shou, J.; Stoltz, D. Reptilian reovirus: A new fusogenic orthoreovirus species. Virology 2004, 319, 131–140. [Google Scholar] [CrossRef] [PubMed]
- Chua, K.B.; Voon, K.; Yu, M.; Keniscope, C.; Abdul Rasid, K.; Wang, L. Investigation of a potential zoonotic transmission of orthoreovirus associated with acute influenza-like illness in an adult patient. PLoS ONE 2011, 6, e25434. [Google Scholar]
- Yamanaka, A.; Iwakiri, A.; Yoshikawa, T.; Sakai, K.; Singh, H.; Himeji, D.; Kikuchi, I.; Ueda, A.; Yamamoto, S.; Miura, M.; et al. Imported case of acute respiratory tract infection associated with a member of species Nelson Bay orthoreovirus. PLoS ONE 2014, 9, e92777. [Google Scholar] [CrossRef] [PubMed]
- Ladner, J.T.; Beitzel, B.; Chain, P.S.G.; Davenport, M.G.; Donaldsen, E.F.; Frieman, M.; Kugelman, J.R.; Kuhn, J.H.; O’Rear, J.; Sabeti, P.C.; et al. Standards for sequencing viral genomes in the era of high-throughput sequencing. mBio 2014, 5. [Google Scholar] [CrossRef]
- Towner, J.S.; Amman, B.R.; Sealy, T.K.; Carrol, S.A.; Comer, J.A.; Kemp, A.; Swanepoel, R.; Paddock, C.D.; Balinandi, S.; Khristova, M.L.; et al. Isolation of genetically diverse Marburg viruses from Egyptian fruit bats. PLoS Pathog. 2009, 5, e1000536. [Google Scholar] [CrossRef] [PubMed]
- Paweska, J.T.; Jansen van Vuren, P.; Masumu, J.; Leman, P.A.; Grobbelaar, A.A.; Birkhead, M.; Clift, S.; Swanepoel, R.; Kemp, A. Virological and serological findings in Rousettus aegyptiacus experimentally inoculated with Vero cells-adapted Hogan strain of Marburg virus. PLoS ONE 2011, 7, e45479. [Google Scholar] [CrossRef] [PubMed]
- Morlan, J.D.; Qu, K.; Sinicropi, D.V. Selective depletion of rRNA enables whole transcriptome profiling of archival fixed tissue. PLoS ONE 2012, 7, e42882. [Google Scholar]
- Djikeng, A.; Halpin, R.; Kuzmickas, R.; DePasse, J.; Feldblyum, J.; Sengamalay, N.; Afonso, C.; Zhang, X.; Anderson, N.G.; Ghedin, E.; et al. Viral genome sequencing by random priming methods. BMC Genomics 2008, 9. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17. [Google Scholar] [CrossRef]
- Schmieder, R.; Edwards, R. Quality control and preprocessing of metagenomics datasets. Bioinformatics 2011, 27, 863–864. [Google Scholar] [CrossRef] [PubMed]
- Boisvert, S.; Raymond, F.; Godzaridis, E.; Laviolette, F.; Corbeil, J. Ray Meta: Scalable de novo metagenome assembly and profiling. Genome Biol. 2012, 13. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.T.; Taylor, W.R.; Thornton, J.M. The rapid generation of mutation data matrices from protein sequences. Comp. Appl. Biosci. 1992, 8, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Theodor, O. An Illustrated Catalogue of the Rothschild Collection of Nycteribiidae (Diptera) in the British Museum (Natural History), with Keys and Short Descriptions for the Identification of Subfamilies, Genera, Species and Subspecies; British Museum (Natural History): London, UK, 1967. [Google Scholar]
- Antczak, J.B.; Chmelo, R.; Pickup, D.J.; Joklik, W.K. Sequence at both termini of the 10 genes of reovirus serotype 3 (strain Dearing). Virology 1982, 121, 307–319. [Google Scholar] [CrossRef]
- Pigott, D.M.; Golding, N.; Mylne, A.; Huang, Z.; Weiss, D.J.; Brady, O.J.; Kraemer, M.U.; Hay, S.I. Mapping the zoonotic niche of Marburg virus disease in Africa. Trans. R. Soc. Trop. Med. Hyg. 2015, 109, 366–378. [Google Scholar] [CrossRef] [PubMed]



| Genbank Accession Number | Subfamily/ Family | Genus | Virus/Species (Isolate) | Gene Description (Encoding for Protein) | Nucleic Acid (NA) or Amino Acid (AA) |
|---|---|---|---|---|---|
| AF450323.1 | Spinareovirinae | Aquareovirus | Golden ide reovirus | RNA dependent RNA polymerase (RdRp) | NA |
| AF403399.1 | Spinareovirinae | Aquareovirus | Golden shiner reovirus | RdRp | NA |
| KC847321.1 | Spinareovirinae | Aquareovirus | Grass carp reovirus (HeNan988) | RdRp | NA |
| AF450318.1 | Spinareovirinae | Aquareovirus | Striped bass reovirus | RdRp | NA |
| AY542965.1 | Sedoreovirinae | Cardoreovirus | Eriocheir sinensis reovirus | RdRp | NA |
| AF133428.1 | Spinareovirinae | Coltivirus | Colorado tick fever virus | RdRp | NA |
| NC_003696.1 | Spinareovirinae | Coltivirus | Eyach virus | RdRp | NA |
| GQ924586.1 | Spinareovirinae | Cypovirus | Bombyx mori cypovirus1 | RdRp | NA |
| AY147187.1 | Spinareovirinae | Cypovirus | Dendrolimus punctatus cypovirus1 | RdRp | NA |
| NC_003017.1 | Spinareovirinae | Cypovirus | Lymantria dispar cypovirus1 | RdRp | NA |
| NC_025486.1 | Spinareovirinae | Dinovernavirus | Fako virus (CSW77) | RdRp | NA |
| AY029520.1 | Spinareovirinae | Fijivirus | Fiji disease virus | RdRp | NA |
| NC_003654.1 | Spinareovirinae | Fijivirus | Nilaparvata lugens reovirus | RdRp | NA |
| X80481.1 | Spinareovirinae | Idnoreovirus | Diadromus pulchellus reovirus | RdRp | NA |
| NC_008172.1 | Sedoreovirinae | Mimoreovirus | Micromonas pusilla reovirus | RdRp | NA |
| NC_010743.1 | Spinareovirinae | Mycoreovirus | Mycoreovirus 1 | RdRp | NA |
| NC_007535.1 | Spinareovirinae | Mycoreovirus | Mycoreovirus 3 | RdRp | NA |
| KF446272.1 | Sedoreovirinae | Orbivirus | African horse sickness virus (RSArrah/08) | RdRp | NA |
| GQ506516.1 | Sedoreovirinae | Orbivirus | Bluetongue virus (BT2-03) | RdRp | NA |
| HM641772.1 | Sedoreovirinae | Orbivirus | Epizootic hemorrhagic disease virus (CC304-06) | RdRp | NA |
| HQ630942.1 | Sedoreovirinae | Orbivirus | Equine encephalosis virus (Kyalami) | RdRp | NA |
| HM543481.1 | Sedoreovirinae | Orbivirus | Kemerovo virus (EgAn 1169-61) | RdRp | NA |
| HQ266581.1 | Sedoreovirinae | Orbivirus | Tribec virus | RdRp | NA |
| KJ476700.1 | Spinareovirinae | Orthoreovirus | Avian orthoreovirus (GX/2010/1) | RdRp | NA |
| NC_015878.1 | Spinareovirinae | Orthoreovirus | Baboon orthoreovirus | RdRp | NA |
| NC_014238.1 | Spinareovirinae | Orthoreovirus | Broome virus | RdRp | NA |
| NC_023819.1 | Spinareovirinae | Orthoreovirus | Bush viper reovirus | RdRp | NA |
| KM382260.1 | Spinareovirinae | Orthoreovirus | Cangyuan orthoreovirus | RdRp | NA |
| JF342655.1 | Spinareovirinae | Orthoreovirus | Kampar orthoreovirus | RdRp | NA |
| KF154724.1 | Spinareovirinae | Orthoreovirus | Mammalian orthoreovirus (SI-MRV01) | RdRp | NA |
| JQ412755.1 | Spinareovirinae | Orthoreovirus | Mammalian Orthoreovirus (T3/Bat/Germany/342/08) | RdRp | NA |
| NC_020447.1 | Spinareovirinae | Orthoreovirus | Melaka orthoreovirus | RdRp | NA |
| JF342673.1 | Spinareovirinae | Orthoreovirus | Nelson Bay orthoreovirus | RdRp | NA |
| JF342667.1 | Spinareovirinae | Orthoreovirus | Pulau reovirus | RdRp | NA |
| KF692090.1 | Spinareovirinae | Orthoreovirus | Tvarminne avian virus | RdRp | NA |
| HM125552.1 | Spinareovirinae | Oryzavirus | Rice ragged stunt virus | RdRp | NA |
| D90198.1 | Spinareovirinae | Oryzavirus | Rice dwarf virus | RdRp | NA |
| NC_003773.1 | Spinareovirinae | Oryzavirus | Rice dwarf virus | RdRp | NA |
| D10222.1 | Spinareovirinae | Oryzavirus | Rice dwarf virus | RdRp | NA |
| AB738412.1 | Sedoreovirinae | Rotavirus | Bovine group C rotavirus (Toyama) | RdRp | NA |
| NC_021541.1 | Sedoreovirinae | Rotavirus | Human rotavirus B (Bang373) | RdRp | NA |
| JN872865.1 | Sedoreovirinae | Rotavirus | Rotavirus A (RVA/Horse-wt/ARG/E4040/2008/G14P12) | RdRp | NA |
| NC_014511.1 | Sedoreovirinae | Rotavirus | Rotavirus D chicken (05V0049/DEU/2005) | RdRp | NA |
| KC954611.1 | Sedoreovirinae | Seadornavirus | Banna virus | RdRp | NA |
| AF133429.1 | Sedoreovirinae | Seadornavirus | Kadipiro virus | RdRp | NA |
| AY317099.1 | Sedoreovirinae | Seadornavirus | Liao ning virus (LNSV-NE97-31) | RdRp | NA |
| AJA90918.1 KJ476700.1 AJA90920.1 AJA90921.1 AJA90922.1 AJA90923.1 AJA90927.1 AJA90928.1 AJA90929.1 AJA90924.1 AJA90925.1 | Spinareovirinae | Orthoreovirus | Avian orthoreovirus (GX/2010/1) | Virus structural and non-structural proteins | AA |
| YP_004769549.1 NC_015878.1 YP_004769547.1 YP_004769550.1 YP_004769551.1 YP_004769552.1 YP_004769553.1 YP_004769554.1 YP_004769557.1 YP_004769555.1 YP_004769556.1 | Spinareovirinae | Orthoreovirus | Baboon orthoreovirus | Virus structural and non-structural proteins | AA |
| YP_003717771.1 NC_014238.1 YP_003717772.1 YP_003717774.1 YP_003717775.1 YP_003717776.1 YP_003717777.1 YP_003717778.1 YP_003717779.1 YP_003717780.1 YP_003717781.1 | Spinareovirinae | Orthoreovirus | Broome virus | Virus structural and non-structural proteins | AA |
| YP_009020572.1 NC_023819.1 YP_009020573.1 YP_009020574.1 YP_009020579.1 YP_009020575.1 YP_009020580.1 YP_009020576.1 YP_009020577.1 YP_009020581.1 | Spinareovirinae | Orthoreovirus | Bush viper reovirus | Virus structural and non-structural proteins | AA* |
| AIY28303.1 KM382260.1 AIY28301.1 AIY28304.1 AIY28305.1 AIY28306.1 AIY28308.1 YP_009110705.1 AIY28309.1 | Spinareovirinae | Orthoreovirus | Cangyuan orthoreovirus | Virus structural and non-structural proteins | AA* |
| AEQ49364.1 JF342655.1 AEQ49362.1 AEQ49365.1 AEQ49366.1 AEQ49367.1 ACC77638.1 ACC77640.1 ACC77639.1 ACC77635.1 ACC77636.1 | Spinareovirinae | Orthoreovirus | Kampar orthoreovirus | Virus structural and non-structural proteins | AA |
| AGV74342.1 KF154724.1 AGV74341.1 AGV74343.1 AGV74344.1 AGV74345.1 AGV74347.1 AGV74346.1 AGV74348.1 | Spinareovirinae | Orthoreovirus | Mammalian orthoreovirus (SI-MRV01) | Virus structural and non-structural proteins | AA* |
| AFN01895.1 JQ412755.1 AFN01894.1 AFN01896.1 AFN01897.1 AFN01898.1 AFN01901.1 AFN01899.1 AFN01902.1 | Spinareovirinae | Orthoreovirus | Mammalian Orthoreovirus (T3/Bat/Germany/342/08) | Virus structural and non-structural proteins | AA* |
| YP_007507318.1 NC_020447.1 YP_007507317.1 YP_007507319.1 YP_007507320.1 YP_007507321.1 YP_007507322.1 YP_007507324.1 YP_007507323.1 YP_007507326.1 YP_007507327.1 | Spinareovirinae | Orthoreovirus | Melaka orthoreovirus | Virus structural and non-structural proteins | AA |
| AEQ49382.1 JF342673.1 AEQ49380.1 AEQ49383.1 AEQ49384.1 AEQ49385.1 AAC18123.1 AAC18127.1 AAC18131.1 AAF45157.1 AAF45158.1 | Spinareovirinae | Orthoreovirus | Nelson Bay orthoreovirus | Virus structural and non-structural proteins | AA |
| AEQ49376.1 JF342667.1 AEQ49374.1 AEQ49377.1 AEQ49378.1 AEQ49379.1 AAR13234.1 AAR13236.1 AAR13235.1 AAR13231.1 AAR13232.1 | Spinareovirinae | Orthoreovirus | Pulau reovirus | Virus structural and non-structural proteins | AA |
| AHW40447.1 KF692090.1 AHW40449.1 AHW40450.1 AHW40451.1 AHW40455.1 AHW40456.1 AHW40457.1 AHW40452.1 AHW40453.1 | Spinareovirinae | Orthoreovirus | Tvarminne avian virus | Virus structural and non-structural proteins | AA* |
| AED99916.1 HM222980.1 AED99917.1 AED99915.1 AED99914.1 AED99913.1 AED99909.1 AED99908.1 AED99910.1 AED99911.1 | Spinareovirinae | Orthoreovirus | Stellar sea lion orthoreovirus | Virus structural and non-structural proteins | AA* |
| AHL21592.1 KF809663.1 AHL21594.1 AHL21595.1 AHL21596.1 AHL21597.1 AHL21600.1 AHL21601.1 AHL21602.1 AHL21598.1 | Spinareovirinae | Orthoreovirus | Avian orthoreovirus (D20/99) | Virus structural and non-structural proteins | AA* |
| AJW82013.1 AJW82012.1 AJW82011.1 AJW82014.2 AJW82015.1 AJW82016.1 AJW82018.1 AJW82019.1 AJW82020.1 | Spinareovirinae | Orthoreovirus | Avian orthoreovirus (PA/Turkey/22342/13) | Virus structural and non-structural proteins | AA* |
| AFQ62078.1 JX145329.1 AFQ62080.1 AFQ62081.1 AFQ62082.1 AFQ62083.1 AFQ62087.1 AFQ62088.1 AFQ62089.1 AFQ62084.1 AFQ62085.1 | Spinareovirinae | Orthoreovirus | Goose orthoreovirus (03G) | Virus structural and non-structural proteins | AA |
| ADY80533.1 GU991669.1 ADY80534.1 ADY80535.1 ADY80536.1 ADY80537.1 ADY80539.1 ADY80541.1 ADY80540.1 | Spinareovirinae | Orthoreovirus | Mammalian orthoreovirus (3jin-1) | Virus structural and non-structural proteins | AA* |
| ADY80523.1 GU991659.1 ADY80524.1 ADY80525.1 ADY80526.2 ADY80527.1 ADY80529.1 ADY80531.1 ADY80530.1 | Spinareovirinae | Orthoreovirus | Mammalian orthoreovirus 3 (R124) | Virus structural and non-structural proteins | AA* |
| JQ599140.1 | Spinareovirinae | Orthoreovirus | Mammalian orthoreovirus 3 (T3v1) | Virus structural and non-structural proteins | AA* |
| AFQ41026.1 JN799426.1 AFQ41027.1 AFQ41022.1 AFQ41023.1 AFQ41024.1 AFQ41019.1 AFQ41021.1 AFQ41020.1 | Spinareovirinae | Orthoreovirus | Mammalian orthoreovirus (729) | Virus structural and non-structural proteins | AA* |
| AF368033.1 AAL36028.1 AF368037 | Spinareovirinae | Orthoreovirus | Ndelle virus | Virus structural and non-structural proteins | AA* |
| AFQ37938.1 JX415466.1 AFQ37943.1 AFQ37939.1 AFQ37942.1 AFQ37945.1 AFQ37936.1 AFQ37944.1 AFQ37941.1 | Spinareovirinae | Orthoreovirus | Porcine reovirus (SHR-A) | Virus structural and non-structural proteins | AA* |
| KF021491.1 | Nycteribiidae | Eucampsipoda | africana (isolate N4) | COI | NA |
| KF021493.1 | Nycteribiidae | Eucampsipoda | inermis (isolate N8) | COI | NA |
| KF021494.1 | Nycteribiidae | Eucampsipoda | madagascarensis (isolate J43) | COI | NA |
| KF021495.1 | Nycteribiidae | Eucampsipoda | madagascarensis (isolate J48) | COI | NA |
| KF021500.1 | Nycteribiidae | Eucampsipoda | theodori (isolate 8B) | COI | NA |
| KF021496.1 | Nycteribiidae | Eucampsipoda | theodori (isolate 28DM) | COI | NA |
| KF021497.1 | Nycteribiidae | Eucampsipoda | theodori (isolate 29F) | COI | NA |
| KF021498.1 | Nycteribiidae | Eucampsipoda | theodori (isolate 30DM) | COI | NA |
| KF021499.1 | Nycteribiidae | Eucampsipoda | theodori (isolate 31DM) | COI | NA |
| AB632546.1 | Nycteribiidae | Nycteribia | allotopa (isolate NyAl10) | COI | NA |
| AB632547.1 | Nycteribiidae | Nycteribia | allotopa (isolate NyAl11) | COI | NA |
| KF021501.1 | Nycteribiidae | Nycteribia | parvula (isolate N16) | COI | NA |
| KF021503.1 | Nycteribiidae | Nycteribia | schmidlii (isolate N2) | COI | NA |
| KF021504.1 | Nycteribiidae | Nycteribia | schmidlii (isolate N3) | COI | NA |
| KF021502.1 | Nycteribiidae | Nycteribia | schmidlii (isolate N21) | COI | NA |
| KF021512.1 | Nycteribiidae | Nycteribia | stylidiopsis (isolate J32) | COI | NA |
| KF021513.1 | Nycteribiidae | Nycteribia | stylidiopsis (isolate J33) | COI | NA |
| KF021519.1 | Nycteribiidae | Penicillidia | fulvida (isolate GR21) | COI | NA |
| KF021518.1 | Nycteribiidae | Penicillidia | fulvida (isolate J68) | COI | NA |
| KF021520.1 | Nycteribiidae | Penicillidia | fulvida (isolate N20) | COI | NA |
| AB632562.1 | Nycteribiidae | Penicillidia | jenynsii (isolate PeJe6) | COI | NA |
| AB632563.1 | Nycteribiidae | Penicillidia | jenynsii (isolate PeJe7) | COI | NA |
| KF021525.1 | Nycteribiidae | Penicillidia | leptothrinax (isolate 35A) | COI | NA |
| KF021526.1 | Nycteribiidae | Penicillidia | leptothrinax (isolate 35B) | COI | NA |
| KF021527.1 | Nycteribiidae | Penicillidia | leptothrinax (isolate 36A) | COI | NA |
| KF021521.1 | Nycteribiidae | Penicillidia | leptothrinax (isolate GR7) | COI | NA |
| KF021529.1 | Nycteribiidae | Penicillidia | leptothrinax (isolate J34) | COI | NA |
| KF021533.1 | Nycteribiidae | Penicillidia | leptothrinax (isolate J66) | COI | NA |
| KF021535.1 | Nycteribiidae | Penicillidia | oceanica (isolate N17) | COI | NA |
| Virus (Isolate Number) | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | 21 | 22 | 23 | 24 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 99.8 | 56.6 | 53.6 | 58.0 | 66.3 | 55.6 | 55.5 | 54.6 | 54.7 | 55.6 | 55.2 | 55.6 | 57.7 | 57.5 | 57.2 | 56.6 | 56.6 | 54.5 | 54.5 | 54.4 | 54.5 | 54.4 | 54.4 | |
| 93.5 | 56.6 | 53.6 | 58.0 | 66.3 | 55.6 | 55.5 | 54.6 | 54.7 | 55.6 | 55.2 | 55.6 | 57.7 | 57.5 | 57.2 | 56.5 | 56.6 | 54.5 | 54.5 | 54.4 | 54.5 | 54.4 | 54.4 | |
| 57.0 | 56.5 | 50.0 | 50.9 | 55.4 | 70.6 | 70.5 | 54.8 | 54.9 | 70.5 | 70.2 | 70.5 | 71.3 | 72.1 | 91.4 | 95.2 | 91.2 | 54.8 | 54.8 | 54.8 | 54.9 | 54.8 | 54.8 | |
| 58.0 | 58.1 | 54.4 | 50.5 | 52.8 | 49.6 | 49.5 | 47.5 | 47.5 | 49.4 | 49.8 | 49.3 | 50.0 | 50.4 | 50.3 | 50.1 | 50.0 | 47.4 | 47.4 | 47.4 | 47.6 | 47.6 | 47.4 | |
| 60.2 | 60.7 | 55.0 | 58.9 | 56.0 | 50.5 | 50.4 | 50.7 | 50.7 | 50.6 | 50.0 | 50.4 | 51.4 | 50.4 | 51.4 | 51.5 | 51.2 | 50.7 | 50.7 | 50.6 | 50.4 | 50.4 | 50.8 | |
| 63.7 | 63.3 | 55.9 | 56.5 | 58.2 | 57.0 | 56.9 | 51.8 | 51.8 | 56.9 | 56.8 | 56.6 | 56.8 | 56.1 | 55.5 | 54.8 | 55.3 | 51.9 | 51.9 | 51.9 | 51.7 | 51.5 | 51.8 | |
| 56.6 | 56.6 | 63.8 | 52.9 | 53.4 | 56.8 | 99.4 | 53.3 | 53.5 | 99.4 | 95.9 | 98.8 | 73.3 | 72.5 | 71.3 | 69.8 | 71.3 | 53.4 | 53.3 | 53.3 | 53.5 | 53.3 | 53.4 | |
| 56.5 | 56.6 | 63.7 | 52.6 | 53.3 | 56.6 | 98.0 | 53.2 | 53.3 | 99.2 | 95.6 | 98.8 | 73.3 | 72.3 | 71.3 | 69.7 | 71.2 | 53.2 | 53.2 | 53.2 | 53.3 | 53.2 | 53.2 | |
| 56.8 | 57.4 | 54.6 | 53.7 | 55.2 | 55.6 | 53.0 | 52.8 | 99.4 | 53.3 | 52.4 | 53.1 | 53.4 | 53.1 | 54.4 | 54.4 | 53.9 | 98.9 | 98.8 | 98.6 | 98.7 | 98.3 | 99.1 | |
| 57.2 | 57.6 | 54.7 | 54.0 | 55.2 | 55.5 | 52.8 | 52.7 | 98.3 | 53.5 | 52.6 | 53.2 | 53.6 | 53.2 | 54.5 | 54.6 | 54.0 | 98.6 | 98.5 | 98.3 | 98.4 | 98.1 | 98.8 | |
| 56.5 | 56.4 | 63.6 | 52.8 | 53.5 | 56.5 | 97.9 | 98.6 | 52.6 | 52.5 | 95.6 | 98.6 | 73.2 | 72.1 | 71.2 | 69.7 | 71.1 | 53.4 | 53.3 | 53.3 | 53.3 | 53.3 | 53.4 | |
| 56.6 | 56.4 | 63.9 | 52.6 | 53.6 | 56.8 | 83.8 | 83.6 | 53.3 | 53.4 | 83.8 | 96.0 | 72.5 | 71.6 | 71.0 | 70.0 | 70.9 | 52.5 | 52.4 | 52.4 | 52.6 | 52.5 | 52.5 | |
| 55.9 | 55.9 | 63.6 | 52.5 | 53.2 | 56.6 | 94.0 | 94.6 | 53.2 | 53.0 | 94.6 | 83.6 | 73.2 | 72.1 | 71.2 | 69.7 | 71.1 | 53.2 | 53.1 | 53.1 | 53.2 | 53.1 | 53.2 | |
| 56.3 | 56.4 | 63.7 | 53.9 | 53.6 | 56.9 | 65.3 | 65.3 | 54.6 | 54.6 | 65.3 | 64.1 | 65.7 | 82.0 | 71.4 | 71.2 | 71.1 | 53.4 | 53.4 | 53.2 | 53.5 | 53.2 | 53.4 | |
| 57.5 | 57.4 | 64.2 | 52.7 | 53.6 | 56.1 | 65.3 | 65.3 | 54.9 | 54.8 | 65.4 | 64.6 | 65.1 | 70.2 | 72.1 | 72.1 | 71.5 | 53.2 | 53.2 | 53.1 | 53.2 | 52.7 | 53.2 | |
| 55.9 | 56.2 | 75.5 | 52.6 | 53.1 | 54.9 | 64.8 | 64.7 | 54.5 | 54.5 | 64.6 | 64.5 | 64.3 | 64.7 | 65.3 | 91.2 | 97.3 | 54.4 | 54.4 | 54.3 | 54.6 | 54.4 | 54.4 | |
| 56.8 | 56.8 | 83.6 | 52.6 | 54.5 | 54.8 | 63.1 | 63.0 | 54.3 | 53.9 | 63.1 | 63.8 | 63.3 | 63.6 | 64.7 | 75.4 | 90.7 | 54.5 | 54.5 | 54.4 | 54.6 | 54.4 | 54.5 | |
| 55.9 | 56.3 | 75.7 | 53.2 | 53.3 | 54.7 | 64.3 | 64.3 | 53.9 | 54.0 | 64.3 | 64.7 | 64.2 | 64.1 | 65.1 | 91.8 | 75.9 | 53.9 | 53.9 | 53.8 | 54.1 | 54.0 | 53.9 | |
| 56.6 | 56.9 | 54.7 | 53.6 | 55.6 | 55.4 | 54.0 | 53.8 | 89.8 | 89.8 | 53.7 | 53.2 | 54.0 | 54.8 | 54.5 | 54.0 | 54.6 | 53.8 | 99.8 | 99.7 | 98.3 | 97.8 | 99.7 | |
| 56.6 | 56.9 | 54.7 | 53.6 | 55.6 | 55.4 | 54.0 | 53.8 | 89.8 | 89.8 | 53.7 | 53.2 | 54.0 | 54.8 | 54.5 | 54.1 | 54.6 | 53.8 | 99.9 | 99.7 | 98.2 | 97.7 | 99.7 | |
| 56.5 | 56.8 | 54.7 | 53.6 | 55.6 | 55.4 | 53.9 | 53.7 | 89.8 | 89.8 | 53.7 | 53.2 | 53.9 | 54.7 | 54.4 | 54.0 | 54.6 | 53.8 | 99.9 | 99.8 | 98.1 | 97.5 | 99.5 | |
| 56.9 | 57.2 | 54.9 | 53.6 | 55.6 | 55.4 | 53.7 | 53.5 | 91.6 | 91.9 | 53.3 | 53.4 | 53.5 | 54.4 | 54.5 | 54.3 | 54.5 | 53.7 | 90.1 | 90.1 | 90.0 | 97.7 | 98.6 | |
| 57.0 | 57.0 | 54.9 | 53.1 | 54.9 | 55.2 | 53.9 | 53.6 | 89.8 | 90.2 | 53.7 | 53.0 | 53.5 | 54.6 | 54.5 | 54.2 | 54.3 | 54.0 | 89.4 | 89.4 | 89.4 | 90.4 | 98.0 | |
| 56.3 | 56.8 | 54.7 | 53.6 | 55.8 | 55.3 | 54.3 | 54.0 | 90.7 | 90.8 | 54.0 | 53.3 | 54.2 | 54.9 | 54.5 | 54.0 | 54.5 | 53.7 | 98.2 | 98.1 | 98.1 | 91.8 | 90.6 |
| Virus (Isolate) | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | 21 | 22 | 23 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 89.0 | 21.4 | 24.0 | 24.3 | 19.0 | 17.8 | 17.8 | 8.3 | 8.3 | 17.8 | 17.5 | 18.1 | 19.3 | 17.5 | 21.1 | 21.4 | 19.3 | 7.7 | 8.0 | 7.7 | 5.6 | 6.2 | |
| 80.4 | 21.7 | 24.3 | 23.1 | 20.2 | 17.5 | 17.5 | 5.6 | 5.9 | 17.5 | 17.8 | 17.5 | 18.7 | 18.4 | 21.7 | 21.1 | 19.6 | 5.6 | 5.9 | 8.6 | 5.0 | 7.1 | |
| 39.8 | 39.9 | 15.1 | 20.2 | 22.6 | 34.4 | 35.0 | 8.9 | 8.9 | 34.7 | 35.0 | 33.8 | 37.1 | 38.3 | 61.7 | 78.6 | 70.0 | 9.2 | 9.2 | 5.9 | 8.0 | 7.1 | |
| 41.7 | 41.7 | 35.8 | 19.3 | 17.2 | 17.5 | 16.9 | 7.1 | 7.1 | 17.2 | 16.9 | 16.0 | 17.2 | 17.5 | 16.3 | 14.8 | 15.7 | 6.5 | 6.8 | 7.7 | 7.1 | 6.8 | |
| 41.9 | 41.5 | 39.0 | 38.5 | 20.2 | 18.1 | 17.8 | 8.9 | 8.9 | 18.1 | 17.5 | 18.7 | 19.6 | 19.6 | 17.2 | 18.4 | 19.0 | 7.7 | 7.7 | 6.2 | 7.4 | 8.3 | |
| 38.7 | 39.3 | 39.5 | 37.3 | 35.8 | 18.4 | 18.1 | 3.9 | 3.9 | 18.4 | 18.1 | 17.5 | 17.8 | 18.1 | 22.6 | 21.7 | 22.0 | 3.6 | 3.6 | 5.3 | 5.0 | 6.2 | |
| 38.0 | 38.6 | 45.4 | 36.5 | 34.0 | 36.5 | 97.0 | 8.9 | 8.9 | 98.8 | 93.5 | 93.8 | 40.1 | 40.4 | 31.8 | 34.4 | 32.9 | 8.3 | 8.3 | 6.5 | 7.1 | 6.5 | |
| 37.8 | 38.7 | 45.4 | 37.0 | 33.9 | 36.0 | 92.5 | 9.5 | 9.5 | 97.6 | 91.7 | 92.0 | 40.1 | 40.4 | 32.3 | 35.0 | 34.1 | 8.6 | 8.6 | 6.5 | 7.7 | 6.8 | |
| 29.4 | 30.2 | 28.9 | 30.4 | 29.0 | 30.1 | 30.6 | 30.6 | 97.9 | 9.2 | 8.3 | 9.5 | 9.5 | 8.9 | 8.6 | 9.8 | 9.2 | 87.5 | 88.1 | 23.1 | 74.2 | 20.5 | |
| 36.5 | 36.9 | 48.7 | 35.0 | 34.8 | 38.0 | 50.3 | 49.7 | 30.8 | 9.2 | 8.3 | 9.8 | 9.8 | 9.2 | 8.9 | 9.8 | 9.5 | 87.2 | 87.8 | 23.1 | 73.3 | 21.4 | |
| 38.7 | 38.2 | 62.9 | 35.6 | 34.7 | 39.2 | 42.6 | 43.3 | 29.2 | 50.8 | 92.9 | 93.8 | 40.4 | 40.4 | 31.8 | 34.7 | 33.2 | 8.6 | 8.6 | 6.5 | 7.4 | 6.8 | |
| 40.1 | 39.8 | 74.7 | 37.4 | 38.7 | 40.2 | 47.2 | 46.6 | 29.9 | 50.4 | 64.2 | 88.4 | 39.5 | 39.8 | 32.0 | 34.4 | 33.8 | 7.7 | 7.7 | 5.9 | 6.5 | 6.2 | |
| 40.3 | 39.8 | 69.8 | 37.5 | 38.1 | 38.3 | 44.2 | 43.5 | 30.6 | 50.0 | 68.6 | 71.1 | 38.3 | 39.2 | 31.2 | 33.2 | 32.6 | 8.9 | 8.9 | 6.5 | 7.7 | 6.2 | |
| 30.1 | 30.5 | 31.6 | 31.8 | 31.2 | 32.2 | 31.3 | 32.2 | 80.3 | 30.8 | 31.5 | 32.0 | 32.4 | 56.1 | 36.2 | 37.7 | 35.9 | 10.1 | 9.8 | 8.0 | 10.1 | 6.2 | |
| 30.1 | 30.5 | 31.6 | 31.8 | 31.3 | 32.2 | 31.3 | 32.2 | 80.4 | 30.8 | 31.5 | 32.0 | 32.4 | 99.9 | 38.0 | 38.6 | 36.5 | 10.1 | 9.8 | 7.7 | 8.6 | 8.6 | |
| 29.6 | 29.4 | 31.8 | 31.0 | 28.0 | 31.9 | 31.7 | 31.6 | 43.8 | 31.3 | 28.8 | 31.7 | 32.0 | 43.4 | 43.4 | 63.2 | 70.3 | 8.6 | 8.6 | 5.6 | 8.3 | 8.0 | |
| 30.3 | 30.0 | 30.2 | 31.7 | 30.0 | 28.7 | 29.5 | 28.8 | 71.2 | 29.6 | 30.6 | 29.7 | 32.6 | 70.9 | 71.0 | 43.3 | 70.6 | 10.1 | 10.1 | 7.7 | 8.0 | 8.0 | |
| 28.7 | 29.9 | 31.3 | 31.2 | 28.5 | 30.6 | 29.7 | 29.5 | 44.1 | 28.7 | 27.6 | 30.3 | 30.1 | 43.3 | 43.4 | 58.0 | 41.7 | 8.6 | 8.6 | 5.9 | 7.1 | 7.1 | |
| 29.7 | 30.6 | 29.1 | 30.4 | 29.4 | 30.2 | 30.6 | 30.6 | 98.5 | 31.0 | 29.5 | 30.0 | 30.8 | 79.8 | 79.9 | 44.0 | 71.3 | 44.5 | 99.4 | 24.9 | 73.3 | 21.1 | |
| 38.0 | 38.7 | 46.1 | 36.9 | 34.0 | 37.3 | 96.2 | 95.5 | 30.5 | 50.8 | 42.4 | 47.2 | 43.9 | 32.0 | 32.0 | 32.2 | 29.5 | 29.8 | 30.6 | 24.9 | 73.6 | 21.1 | |
| 37.4 | 38.8 | 45.6 | 36.5 | 34.9 | 37.3 | 83.3 | 83.4 | 31.5 | 49.4 | 41.4 | 46.8 | 43.0 | 31.4 | 31.4 | 32.2 | 30.5 | 30.6 | 31.4 | 83.9 | 23.1 | 50.7 | |
| 36.9 | 37.7 | 45.1 | 36.4 | 34.3 | 36.9 | 93.1 | 91.0 | 31.1 | 49.2 | 41.6 | 45.5 | 43.1 | 32.4 | 32.4 | 32.4 | 30.6 | 30.3 | 31.2 | 94.2 | 82.3 | 20.8 | |
| 35.3 | 35.6 | 48.9 | 35.4 | 32.6 | 39.6 | 50.8 | 49.6 | 30.3 | 61.7 | 47.4 | 49.0 | 48.5 | 31.1 | 31.1 | 32.5 | 30.6 | 30.2 | 31.0 | 50.1 | 48.6 | 49.3 |
| Genome Segment | Gene | Gene Product | Public Sequence Database | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Size of Segment (nt) | 5' End Penta-Nucleotide Sequence | 3' End Penta-Nucleotide Sequence | 5' UTR (nt) | 3' UTR (nt) | Intergenic Region | Protein | Protein Size (aa) | Mw (kDa) | Proposed Function | Genbank Accession Numbers | ||||
| L1 | 3950 | 5'-GGUCA | UCAUC-3' | 14 | 51 | - | Lambda A | 1294 | 143.7 | Core structural protein, binds dsRNA, NTPase, helicase | KU198602 to KU198621 | |||
| L2 | 3925 | 5'-GGUCA | UCAUC-3' | 16 | 33 | - | Lambda C | 1292 | 146.8 | Guanylyl transferase, methyl transferase turret protein | ||||
| L3 | 3846 | 5'-GGUCA | UCAUC-3' | 13 | 35 | - | Lambda B | 1265 | 142.7 | Core protein, RNA-dependent RNA polymerase | ||||
| M1 | 2341 | 5'-GGUCA | UCAUC-3' | 16 | 39 | - | mu A | 761 | 87.2 | Core protein, transcription factor; ssRNA and dsRNA binding | ||||
| M2 | 2145 | 5'-GGUCA | UCAUC-3' | 28 | 86 | - | mu B | 676 | 73.4 | Outer capsid protein, membrane penetration during infection | ||||
| M3 | 2112 | 5'-GGUCA | UCAUC-3' | 34 | 77 | - | mu NS | 666 | 75.6 | Non-structural, virus inclusion | ||||
| S1 | 1322 | 5'-GGUCA | UCAUC-3' | 13 | 58 | - | Sigma A | 416 | 47.8 | Core protein, transcription factor; dsRNA binding; blocks interferon pathway | ||||
| S2 | 1282 | 5'-GGUCA | UCAUC-3' | 31 | 69 | - | Sigma B | 393 | 44.7 | Outer capsid protein | ||||
| S3 | 1209 | 5'-GGUCA | UCAUC-3' | 27 | 63 | - | Sigma NS | 372 | 41.5 | ssRNA binding; virus inclusion formation | ||||
| S4 | 1068 | 5'-GGUCA | UCAUC-3' | 27 | 48 | 156 | p14 | 125 | 13.9 | Non-structural, membrane fusion (FAST) | ||||
| p18 | 152 | 17.8 | Non-structural, unknown function | |||||||||||
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jansen van Vuren, P.; Wiley, M.; Palacios, G.; Storm, N.; McCulloch, S.; Markotter, W.; Birkhead, M.; Kemp, A.; Paweska, J.T. Isolation of a Novel Fusogenic Orthoreovirus from Eucampsipoda africana Bat Flies in South Africa. Viruses 2016, 8, 65. https://doi.org/10.3390/v8030065
Jansen van Vuren P, Wiley M, Palacios G, Storm N, McCulloch S, Markotter W, Birkhead M, Kemp A, Paweska JT. Isolation of a Novel Fusogenic Orthoreovirus from Eucampsipoda africana Bat Flies in South Africa. Viruses. 2016; 8(3):65. https://doi.org/10.3390/v8030065
Chicago/Turabian StyleJansen van Vuren, Petrus, Michael Wiley, Gustavo Palacios, Nadia Storm, Stewart McCulloch, Wanda Markotter, Monica Birkhead, Alan Kemp, and Janusz T. Paweska. 2016. "Isolation of a Novel Fusogenic Orthoreovirus from Eucampsipoda africana Bat Flies in South Africa" Viruses 8, no. 3: 65. https://doi.org/10.3390/v8030065
APA StyleJansen van Vuren, P., Wiley, M., Palacios, G., Storm, N., McCulloch, S., Markotter, W., Birkhead, M., Kemp, A., & Paweska, J. T. (2016). Isolation of a Novel Fusogenic Orthoreovirus from Eucampsipoda africana Bat Flies in South Africa. Viruses, 8(3), 65. https://doi.org/10.3390/v8030065






