Role of Pea Enation Mosaic Virus Coat Protein in the Host Plant and Aphid Vector
Abstract
:1. Introduction
2. Materials and Methods
2.1. CP Predicted Structure
2.2. Construction of CP Mutants
2.3. Plasmid Digestion and In Vitro Transcription of Viral RNAs
2.4. Plant Growth Conditions and Viral Inoculation
2.5. Virus Detection in Pea Plants
2.5.1. Quantification of Within-Host Accumulation by One-Step RT-PCR
2.5.2. Sanger Sequencing
2.5.3. CP Detection by Western Blot
2.5.4. Electron Microscopy
2.5.5. Aphid Transmission Assays
2.6. Statistical Analysis
3. Results
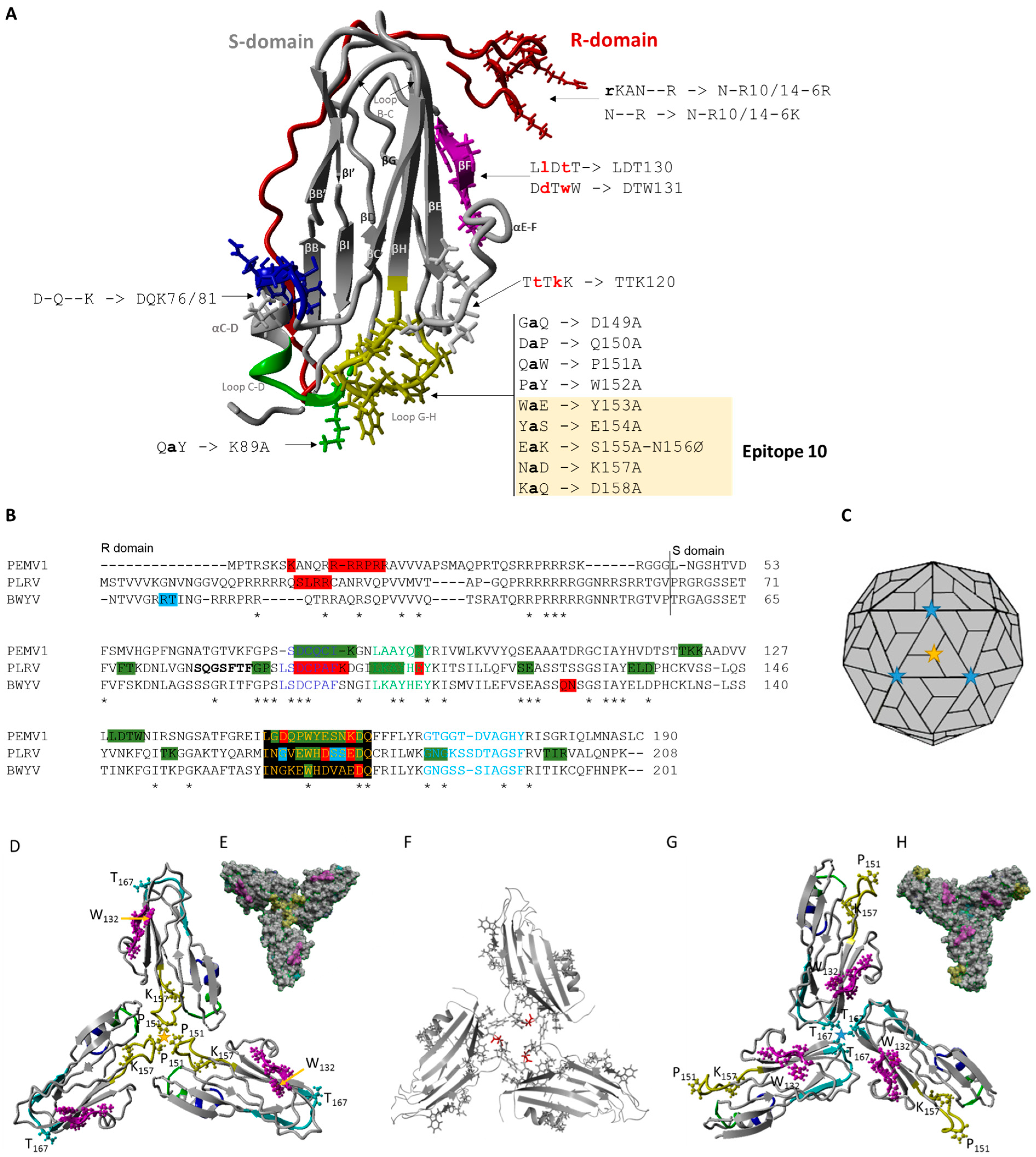
3.1. CP Predicted Structure
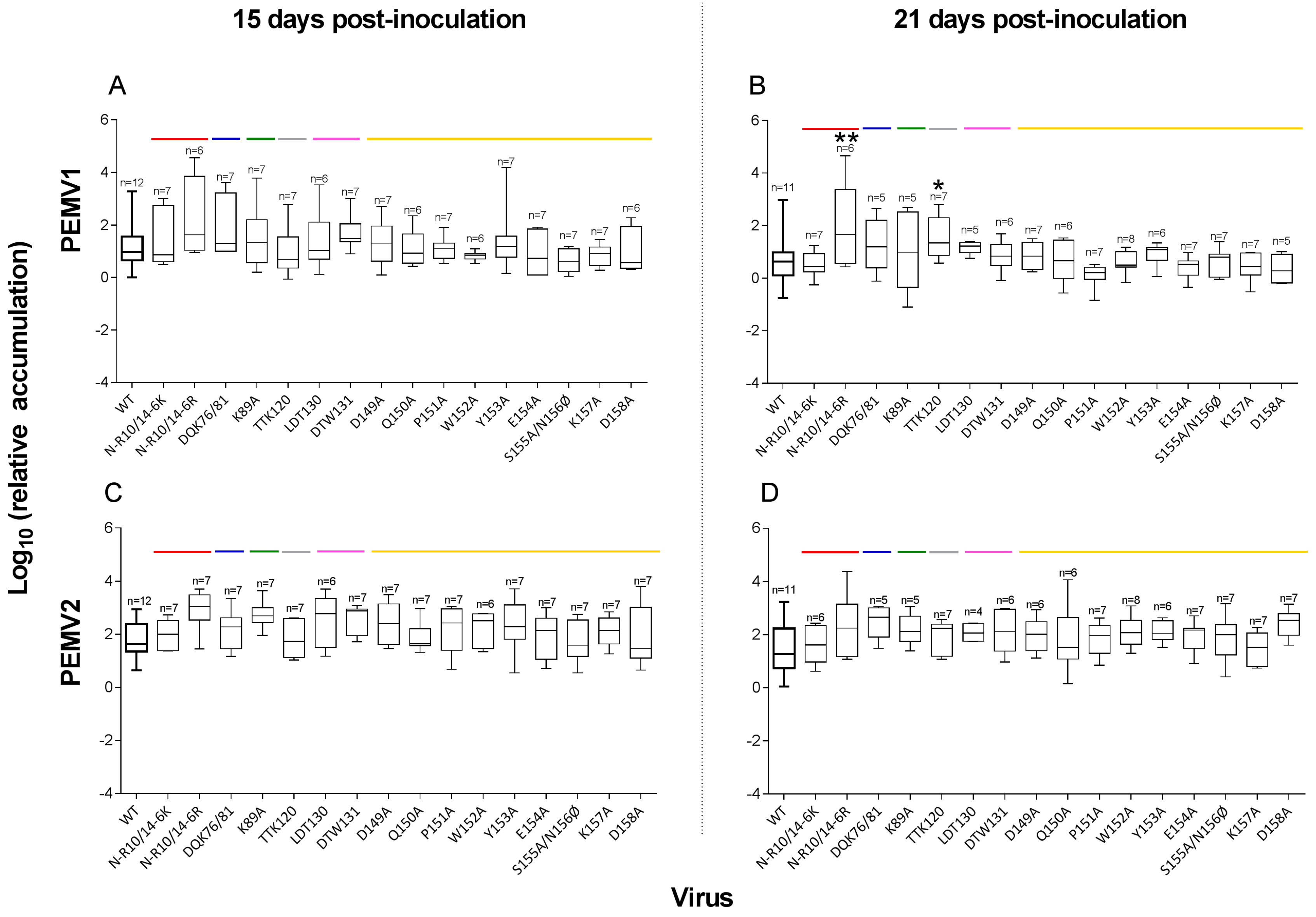
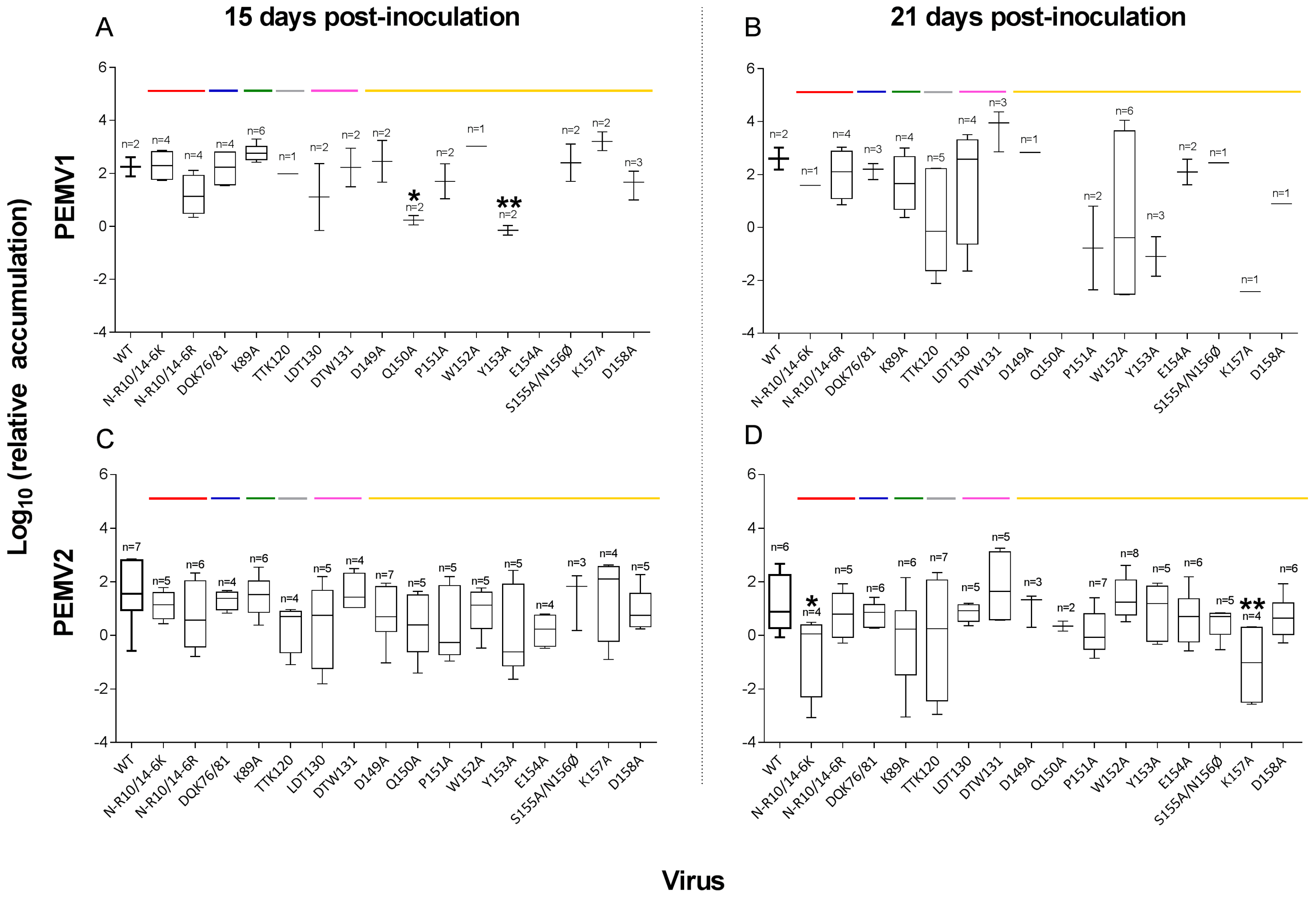
3.2. PEMV Replication and Systemic Infection
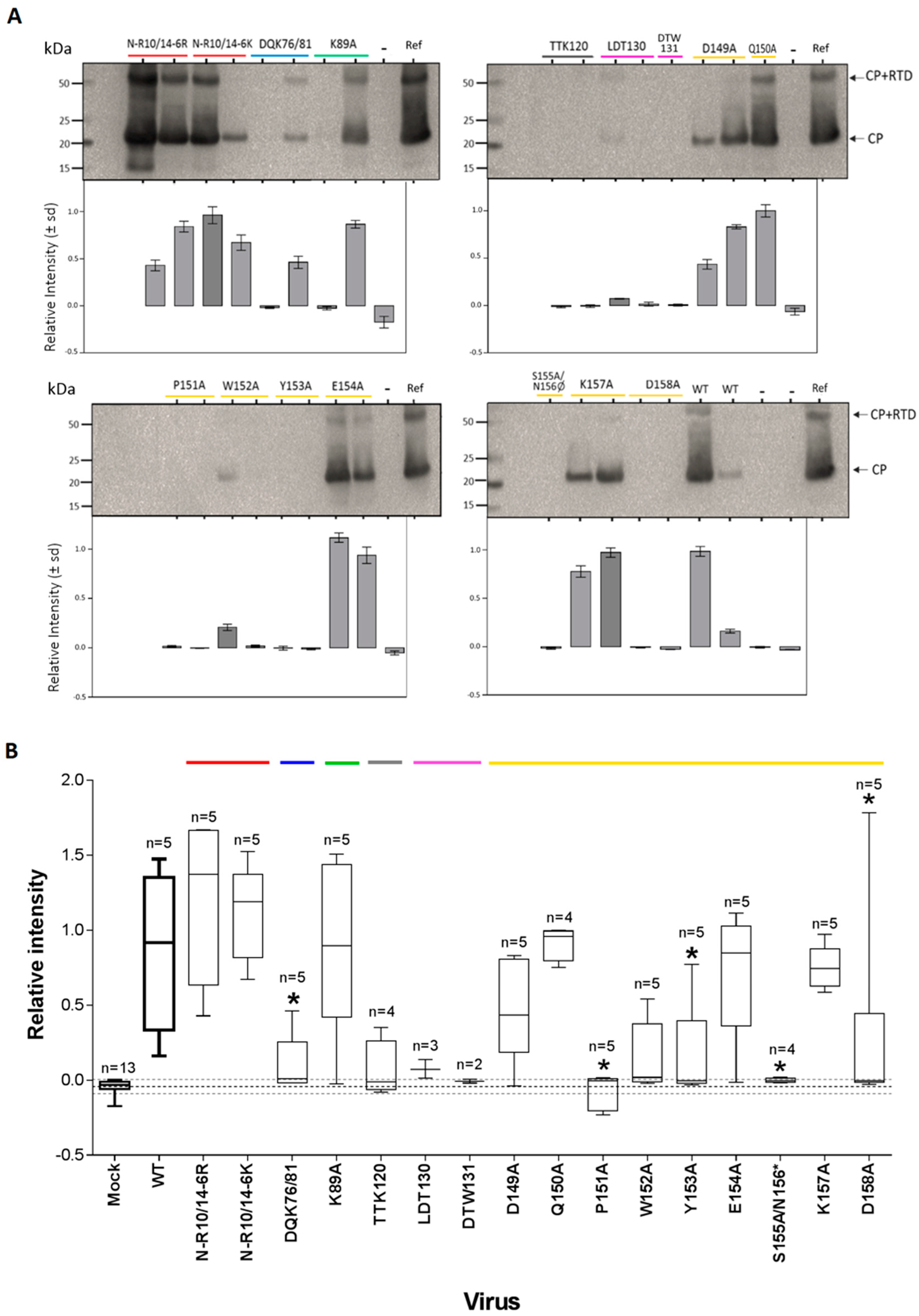
3.3. CP Expression and Virion Assembly
3.4. Aphid Transmission
4. Discussion
4.1. Replication in Inoculated Leaves
4.2. Abilities of Mutants to Move in Plants
4.3. Capacity of Assembly and Transmission by Aphids
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Fereres, A.; Raccah, B. Plant virus transmission by insects. eLS 2015. [Google Scholar] [CrossRef]
- Ng, J.C.; Perry, K.L. Transmission of plant viruses by aphid vectors. Mol. Plant Pathol. 2004, 5, 505–511. [Google Scholar] [CrossRef] [PubMed]
- Callaway, A.; Giesman-Cookmeyer, D.; Gillock, E.; Sit, T.L.; Lommel, S. The multifunctional capsid proteins of plant RNA viruses. Annu. Rev. Phytopathol. 2001, 39, 419–460. [Google Scholar] [CrossRef] [PubMed]
- Hogenhout, S.A.; Ammar, E.-D.; Whitfield, A.E.; Redinbaugh, M.G. Insect vector interactions with persistently transmitted viruses. Annu. Rev. Phytopathol. 2008, 46, 327–359. [Google Scholar] [CrossRef] [PubMed]
- Gray, S.; Gildow, F.E. Luteovirus-Aphid Interactions. Annu. Rev. Phytopathol. 2003, 41, 539–566. [Google Scholar] [CrossRef] [PubMed]
- Linz, L.B.; Liu, S.; Chougule, N.P.; Bonning, B.C. In vitro evidence supports membrane alanyl aminopeptidase N as a receptor for a plant virus in the pea aphid vector. J. Virol. 2015, 89, 11203–11212. [Google Scholar] [CrossRef] [PubMed]
- Demler, S.A.; de Zoeten, G.A. The nucleotide sequence and luteovirus-like nature of RNA 1 of an aphid non-transmissible strain of Pea enation mosaic virus. J. Gen. Virol. 1991, 72, 1819–1834. [Google Scholar] [CrossRef] [PubMed]
- Demler, S.A.; Rucker, D.G.; de Zoeten, G.A. The chimeric nature of the genome of Pea enation mosaic virus: The independent replication of RNA 2. J. Gen. Virol. 1993, 74, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Taliansky, M.E.; Robinson, D.J. Molecular biology of umbraviruses: Phantom warriors. J. Gen. Virol. 2003, 84, 1951–1960. [Google Scholar] [CrossRef] [PubMed]
- Demler, S.A.; Borkhsenious, O.N.; Rucker, D.G.; de Zoeten, G.A. Assessment of the autonomy of replicative and structural functions encoded by the luteo-phase of Pea enation mosaic virus. J. Gen. Virol. 1994, 75, 997–1007. [Google Scholar] [CrossRef] [PubMed]
- Shikata, E.; Maramorosch, K.; Granados, R.R. Electron microscopy of Pea enation mosaic virus in plants and aphid vectors. Virology 1966, 29, 426–436. [Google Scholar] [CrossRef]
- Harris, K.F.; Bath, J.E. The fate of pea enation mosaic virus in its pea aphid vector, Acyrthosiphon pisum (Harris). Virology 1972, 50, 778–790. [Google Scholar] [CrossRef]
- Demler, S.; De Zoeten, G.; Adam, G.; Harris, K. Pea enation mosaic enamovirus: Properties and aphid transmission. In The Plant Viruses; Springer: New York, NY, USA, 1996; pp. 303–344. [Google Scholar]
- Powell, C.A.; De Zoeten, G.A.; Gaard, G. The localization of Pea enation mosaic virus-induced RNA-dependent RNA polymerase in infected peas. Virology 1977, 78, 135–143. [Google Scholar] [CrossRef]
- Caspar, D.L.D.; Klug, A. Physical principles in the construction of regular viruses. Cold Spring Harb. Symp. Quant. Biol. 1962, 27, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Demler, S.A.; Rucker-Feeney, D.G.; Skaf, J.S.; de Zoeten, G.A. Expression and suppression of circulative aphid transmission in Pea enation mosaic virus. J. Gen. Virol. 1997, 78, 511–523. [Google Scholar] [CrossRef] [PubMed]
- Dolja, V.V.; Boyko, V.P.; Agranovsky, A.A.; Koonin, E.V. Phylogeny of capsid proteins of rod-shaped and filamentous RNA plant viruses: Two families with distinct patterns of sequence and probably structure conservation. Virology 1991, 184, 79–86. [Google Scholar] [CrossRef]
- Chapman, M.S.; Liljas, L. Structural folds of viral proteins. Adv. Protein Chem. 2003, 64, 125–196. [Google Scholar] [PubMed]
- Brault, V.; Bergdoll, M.; Mutterer, J.; Prasad, V.; Pfeffer, S.; Erdinger, M.; Richards, K.E.; Ziegler-Graff, V. Effects of point mutations in the major capsid protein of Beet western yellows virus on capsid formation, virus accumulation, and aphid transmission. J. Virol. 2003, 77, 3247–3256. [Google Scholar] [CrossRef] [PubMed]
- Terradot, L.; Souchet, M.; Tran, V.; Giblot Ducray-Bourdin, D. Analysis of a three-dimensional structure of potato leafroll virus coat protein obtained by homology modeling. Virology 2001, 286, 72–82. [Google Scholar] [CrossRef] [PubMed]
- Lee, L.; Kaplan, I.B.; Ripoll, D.R.; Liang, D.; Palukaitis, P.; Gray, S.M. A surface loop of the Potato leafroll virus coat protein is involved in virion assembly, systemic movement, and aphid transmission. J. Virol. 2005, 79, 1207–1214. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, I.B.; Lee, L.; Ripoll, D.R.; Palukaitis, P.; Gildow, F.; Gray, S.M. Point mutations in the Potato leafroll virus major capsid protein alter virion stability and aphid transmission. J. Gen. Virol. 2007, 88, 1821–1830. [Google Scholar] [CrossRef] [PubMed]
- Chavez, J.D.; Cilia, M.; Weisbrod, C.R.; Ju, H.-J.; Eng, J.K.; Gray, S.M.; Bruce, J.E. Cross-linking measurements of the Potato leafroll virus reveal protein interaction topologies required for virion stability, aphid transmission, and virus–plant interactions. J. Proteome Res. 2012, 11, 2968–2981. [Google Scholar] [CrossRef] [PubMed]
- DeBlasio, S.L.; Johnson, R.; Sweeney, M.M.; Karasev, A.; Gray, S.M.; MacCoss, M.J.; Cilia, M. Potato leafroll virus structural proteins manipulate overlapping, yet distinct protein interaction networks during infection. Proteomics 2015, 15, 2098–2112. [Google Scholar] [CrossRef] [PubMed]
- Commandeur, U.; Koenig, R.; Lesemann, D.-E.; Torrance, L.; Burgermeister, W.; Liu, Y.; Schots, A.; Alric, M.; Grassi, G. Epitope mapping on fragments of beet necrotic yellow vein virus coat protein. J. Gen. Virol. 1992, 73, 695–700. [Google Scholar] [CrossRef] [PubMed]
- Mayo, M.; Ziegler-Graff, V. Molecular biology of luteoviruses. Adv. Virus Res. 1996, 46, 4132460. [Google Scholar]
- Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinform. 2008, 9, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Wang, Y.; Zhang, Y. ResQ: An approach to unified estimation of B-factor and residue-specific error in protein structure prediction. J. Mol. Biol. 2016, 428, 693–701. [Google Scholar] [CrossRef] [PubMed]
- Schneidman-Duhovny, D.; Inbar, Y.; Nussinov, R.; Wolfson, H.J. PatchDock and SymmDock: Servers for rigid and symmetric docking. Nucleic Acids Res. 2005, 33, W363–W367. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef] [PubMed]
- Ruijter, J.; Ramakers, C.; Hoogaars, W.; Karlen, Y.; Bakker, O.; Van den Hoff, M.; Moorman, A. Amplification efficiency: Linking baseline and bias in the analysis of quantitative PCR data. Nucleic Acids Res. 2009, 37, e45. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Sivakumar, S.; Wang, Z.; Bonning, B.C.; Allen Miller, W. The readthrough domain of pea enation mosaic virus coat protein is not essential for virus stability in the hemolymph of the pea aphid. Arch. Virol. 2009, 154, 469–479. [Google Scholar] [CrossRef] [PubMed]
- Harrison, S.C.; Olson, A.J.; Schutt, C.E.; Winkler, F.K.; Bricogne, G. Tomato bushy stunt virus at 2.9 Å resolution. Nature 1978, 276, 368–373. [Google Scholar] [CrossRef] [PubMed]
- Skaf, J.S.; Rucker, D.G.; Demler, S.A.; Wobus, C.E.; de Zoeten, G.A. The coat protein is dispensable for the establishment of systemic infections by Pea enation mosaic enamovirus. Mol. Plant. Microbe Interact. 1997, 10, 929–932. [Google Scholar] [CrossRef]
- De Zoeten, G.A.; Powell, C.A.; Gaard, G.; German, T.L. In situ localization of Pea enation mosaic virus double-stranded ribonucleic acid. Virology 1976, 70, 459–469. [Google Scholar] [CrossRef]
- De Zoeten, G.A.; Gaard, G.; Diez, F.B. Nuclear vesiculation associated with Pea enation mosaic virus-infected plant tissue. Virology 1972, 48, 638–647. [Google Scholar] [CrossRef]
- Hipper, C.; Monsion, B.; Bortolamiol-Bécet, D.; Ziegler-Graff, V.; Brault, V. Formation of virions is strictly required for Turnip yellows virus long-distance movement in plants. J. Gen. Virol. 2014, 95, 496–505. [Google Scholar] [CrossRef] [PubMed]
- Ryabov, E.V.; Robinson, D.J.; Taliansky, M. Umbravirus-encoded proteins both stabilize heterologous viral RNA and mediate its systemic movement in some plant species. Virology 2001, 288, 391–400. [Google Scholar] [CrossRef] [PubMed]
- Ryabov, E.V.; Robinson, D.J.; Taliansky, M.E. A plant virus-encoded protein facilitates long-distance movement of heterologous viral RNA. Proc. Natl. Acad. Sci. USA 1999, 96, 1212–1217. [Google Scholar] [CrossRef] [PubMed]
- Taliansky, M.; Roberts, I.M.; Kalinina, N.; Ryabov, E.V.; Raj, S.K.; Robinson, D.J.; Oparka, K.J. An umbraviral protein, involved in long-distance RNA movement, binds viral RNA and forms unique, protective ribonucleoprotein complexes. J. Virol. 2003, 77, 3031–3040. [Google Scholar] [CrossRef] [PubMed]
- Yi, G.; Letteney, E.; Kim, C.-H.; Kao, C.C. Brome mosaic virus capsid protein regulates accumulation of viral replication proteins by binding to the replicase assembly RNA element. RNA 2009, 15, 615–626. [Google Scholar] [CrossRef] [PubMed]
- Besong-Ndika, J.; Ivanov, K.I.; Hafrèn, A.; Michon, T.; Mäkinen, K. Cotranslational coat protein-mediated inhibition of potyviral RNA translation. J. Virol. 2015, 89, 4237–4248. [Google Scholar] [CrossRef] [PubMed]
- Annamalai, P.; Apte, S.; Wilkens, S.; Rao, A. Deletion of highly conserved arginine-rich RNA binding motif in Cowpea chlorotic mottle virus capsid protein results in virion structural alterations and RNA packaging constraints. J. Virol. 2005, 79, 3277–3288. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.-K.; Hacker, D.L. In vitro analysis of an RNA binding site within the N-terminal 30 amino acids of the Southern cowpea mosaic virus coat protein. Virology 2001, 286, 317–327. [Google Scholar] [CrossRef] [PubMed]
- Tan, R.; Frankel, A.D. Structural variety of arginine-rich RNA-binding peptides. Proc. Natl. Acad. Sci. USA 1995, 92, 5282–5286. [Google Scholar] [CrossRef] [PubMed]
- Tan, R.; Chen, L.; Buettner, J.A.; Hudson, D.; Frankel, A.D. RNA recognition by an isolated α helix. Cell 1993, 73, 1031–1040. [Google Scholar] [CrossRef]
- Rouzé-Jouan, J.; Terradot, L.; Pasquer, F.; Tanguy, S.; Giblot Ducray-Bourdin, D. The passage of Potato leafroll virus through Myzus persicae gut membrane regulates transmission efficiency. J. Gen. Virol. 2001, 82, 17–23. [Google Scholar] [CrossRef] [PubMed]
- Plevka, P.; Tars, K.; Zeltins, A.; Balke, I.; Truve, E.; Liljas, L. The three-dimensional structure of ryegrass mottle virus at 2.9 Å resolution. Virology 2007, 369, 364–374. [Google Scholar] [CrossRef] [PubMed]
- Bhuvaneshwari, M.; Subramanya, H.; Gopinath, K.; Savithri, H.; Nayudu, M.; Murthy, M. Structure of sesbania mosaic virus at 3 Å resolution. Structure 1995, 3, 1021–1030. [Google Scholar] [CrossRef]
- Jolly, C.A.; Mayo, M.A. Changes in the amino acid sequence of the coat protein readthrough domain of Potato leafroll luteovirus affect the formation of an epitope and aphid transmission. Virology 1994, 201, 182–185. [Google Scholar] [CrossRef] [PubMed]
- Chay, C.A.; Gunasinge, U.B.; Dinesh-Kumar, S.P.; Miller, W.A.; Gray, S.M. Aphid transmission and systemic plant infection determinants of Barley yellow dwarf luteovirus-PAV are contained in the coat protein readthrough domain and 17-kDa protein, respectively. Virology 1996, 219, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Bruyère, A.; Brault, V.; Ziegler-Graff, V.; Simonis, M.-T.; Van den Heuvel, J.F.; Richards, K.; Guilley, H.; Jonard, G.; Herrbach, E. Effects of mutations in the Beet western yellows virus readthrough protein on its expression and packaging and on virus accumulation, symptoms, and aphid transmission. Virology 1997, 230, 323–334. [Google Scholar] [CrossRef] [PubMed]
- Van den Heuvel, J.F.; Bruyère, A.; Hogenhout, S.A.; Ziegler-Graff, V.; Brault, V.; Verbeek, M.; van der Wilk, F.; Richards, K. The N-terminal region of the luteovirus readthrough domain determines virus binding to Buchnera GroEL and is essential for virus persistence in the aphid. J. Virol. 1997, 71, 7258–7265. [Google Scholar] [PubMed]
- Brault, V.; van den Heuvel, J.F.; Verbeek, M.; Ziegler-Graff, V.; Reutenauer, A.; Herrbach, E.; Garaud, J.C.; Guilley, H.; Richards, K.; Jonard, G. Aphid transmission of Beet western yellows luteovirus requires the minor capsid read-through protein P74. EMBO J. 1995, 14, 650–659. [Google Scholar] [PubMed]
- Peter, K.A.; Liang, D.; Palukaitis, P.; Gray, S.M. Small deletions in the Potato leafroll virus readthrough protein affect particle morphology, aphid transmission, virus movement and accumulation. J. Gen. Virol. 2008, 89, 2037–2045. [Google Scholar] [CrossRef] [PubMed]


| (A) PEMV-1 Infectivity | |||||||||
| Location in CP | Inoculated Leaves | Upper Leaves 1 | |||||||
| 15 dpi | 21 dpi | 15 dpi | 21 dpi | 15 + 21 dpi | |||||
| Wild-Type | - | 12/12 | 11/12 | 2/12 | 2/12 | 4/24 | |||
| N-R10/14-6K | R-domain | 7/7 | 7/7 | 4/7 | # | 1/7 | 5/14 | ||
| N-R10/14-6R | R-domain | 6/6 | 6/6 | 4/7 | # | 4/6 | * | 8/13 | * |
| DQK76/81 | Helix αC-D | 7/7 | 5/5 | 4/7 | # | 3/5 | 7/12 | * | |
| K89A | Loop C-D | 7/7 | 5/5 | 6/7 | * | 4/5 | * | 10/12 | *** |
| TTK120 | Helix αE-F | 7/7 | 7/7 | 1/7 | 5/7 | * | 6/14 | ||
| LDT130 | Barrel βF | 6/7 | 5/5 | 2/6 | 5/5 | ** | 7/11 | * | |
| DTW131 | Barrel βF | 7/7 | 6/6 | 2/7 | 3/6 | 5/13 | |||
| D149A | Loop G-H | 7/7 | 6/7 | 2/7 | 1/6 | 3/13 | |||
| Q150A | Loop G-H | 6/6 | 6/6 | 2/6 | 0/6 | 2/12 | |||
| P151A | Loop G-H | 7/7 | 7/7 | 2/7 | 2/7 | 4/14 | |||
| W152A | Loop G-H | 6/6 | 8/8 | 1/6 | 6/8 | * | 7/14 | * | |
| Y153A | Epitope 10 | 7/7 | 6/6 | 2/7 | 3/6 | 5/13 | |||
| E154A | Epitope 10 | 7/7 | 7/7 | 0/7 | 2/7 | 2/14 | |||
| S155A/N156Ø | Epitope 10 | 7/7 | 7/7 | 2/7 | 1/7 | 3/14 | |||
| K157A | Epitope 10 | 7/7 | 7/7 | 2/7 | 1/7 | 3/14 | |||
| D158A | Epitope 10 | 6/7 | 6/7 | 3/6 | 1/6 | 4/12 | |||
| (B) PEMV-2 Infectivity | |||||||||
| Location in CP | Inoculated Leaves | Upper Leaves 1 | |||||||
| Co-Inoculated with: | 15 dpi | 21 dpi | 15 dpi | 21 dpi | 15 + 21 dpi | ||||
| PEMV-1 Wild-Type | - | 12/12 | 11/12 | 7/12 | 6/12 | 13/24 | |||
| N-R10/14-6K | R-domain | 7/7 | 7/7 | 5/7 | 4/7 | 9/14 | |||
| N-R10/14-6R | R-domain | 6/6 | 6/6 | 6/6 | # | 5/7 | 11/13 | # | |
| DQK76/81 | Helix αC-D | 7/7 | 5/5 | 4/7 | 6/6 | * | 10/13 | ||
| K89A | Loop C-D | 7/7 | 5/5 | 6/7 | 6/6 | * | 12/13 | * | |
| TTK120 | Helix αE-F | 7/7 | 7/7 | 4/7 | 7/7 | * | 11/14 | ||
| LDT130 | Barrel βF | 6/7 | 5/5 | 5/6 | 5/5 | * | 10/11 | * | |
| DTW131 | Barrel βF | 7/7 | 6/6 | 4/7 | 5/6 | 9/13 | |||
| D149A | Loop G-H | 7/7 | 7/7 | 7/7 | * | 3/7 | 10/14 | ||
| Q150A | Loop G-H | 6/6 | 6/6 | 5/6 | 2/6 | 7/12 | |||
| P151A | Loop G-H | 7/7 | 7/7 | 5/7 | 7/7 | * | 12/14 | * | |
| W152A | Loop G-H | 6/6 | 8/8 | 5/6 | 8/8 | * | 13/14 | * | |
| Y153A | Epitope 10 | 7/7 | 6/6 | 5/7 | 5/6 | 10/13 | |||
| E154A | Epitope 10 | 7/7 | 7/7 | 4/7 | 6/7 | 10/14 | |||
| S155A/N156Ø | Epitope 10 | 7/7 | 7/7 | 3/7 | 5/7 | 8/14 | |||
| K157A | Epitope 10 | 7/7 | 7/7 | 4/7 | 4/7 | 8/14 | |||
| D158A | Epitope 10 | 7/7 | 7/7 | 5/7 | 6/7 | 11/14 | |||
| Quantitative Trait | Leaf | Factor | 15 dpi | 21 dpi | ||||
|---|---|---|---|---|---|---|---|---|
| df | F | p 1 | df | F | p 1 | |||
| PEMV1 Viral Accumulation relative to actin | Inoculated | Clones | 16 | 1.3746 | 0.1694 | 16 | 2.2478 | 0.0082 * |
| upper | 13 | 4.1487 | 0.0011 ** | 10 | 1.3291 | 0.2652 | ||
| PEMV2 Viral Accumulation relative to actin | Inoculated | Clones | 16 | 1.4679 | 0.1258 | 16 | 0.9635 | 0.5021 |
| upper | 16 | 1.0486 | 0.4203 | 16 | 1.9346 | 0.0303 * | ||
| PEMV1 Viral Accumulation relative to β-tubulin | Inoculated | Clones | 16 | 1.1967 | 0.2838 | 16 | 2.1897 | 0.0109 * |
| upper | 13 | 3.1809 | 0.0063 * | 10 | 1.5485 | 0.1764 | ||
| PEMV2 Viral Accumulation relative to β-tubulin | Inoculated | Clones | 16 | 1.3484 | 0.1830 | 16 | 0.8879 | 0.5848 |
| upper | 16 | 0.8297 | 0.6483 | 16 | 2.1571 | 0.0141 * | ||
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Doumayrou, J.; Sheber, M.; Bonning, B.C.; Miller, W.A. Role of Pea Enation Mosaic Virus Coat Protein in the Host Plant and Aphid Vector. Viruses 2016, 8, 312. https://doi.org/10.3390/v8110312
Doumayrou J, Sheber M, Bonning BC, Miller WA. Role of Pea Enation Mosaic Virus Coat Protein in the Host Plant and Aphid Vector. Viruses. 2016; 8(11):312. https://doi.org/10.3390/v8110312
Chicago/Turabian StyleDoumayrou, Juliette, Melissa Sheber, Bryony C. Bonning, and W. Allen Miller. 2016. "Role of Pea Enation Mosaic Virus Coat Protein in the Host Plant and Aphid Vector" Viruses 8, no. 11: 312. https://doi.org/10.3390/v8110312
APA StyleDoumayrou, J., Sheber, M., Bonning, B. C., & Miller, W. A. (2016). Role of Pea Enation Mosaic Virus Coat Protein in the Host Plant and Aphid Vector. Viruses, 8(11), 312. https://doi.org/10.3390/v8110312







