Exosomes: Implications in HIV-1 Pathogenesis
Abstract
:1. Introduction
2. Exosomes: Secreted Vesicles Loaded with Various Cargos
3. HIV-1 Exploits Exosomal Machinery: HIV-1 RNA and Protein Trafficking via Exosomes
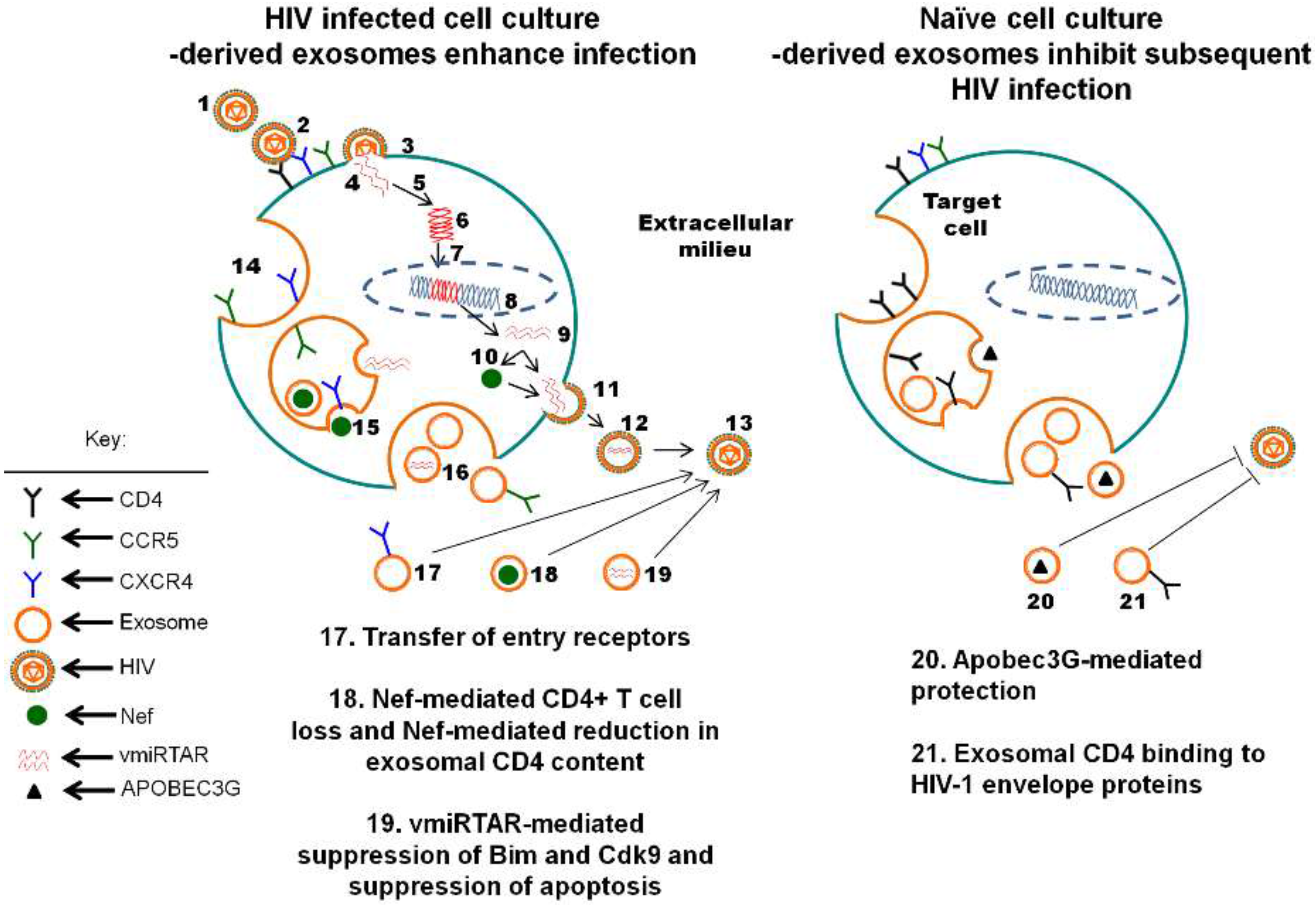
4. Exosomes Released by HIV-1-Infected Cultured Cells Contain HIV-1-Derived Virulence Factors and Influence Host Cell Infection
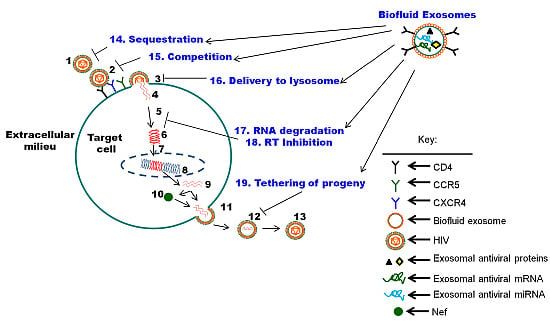
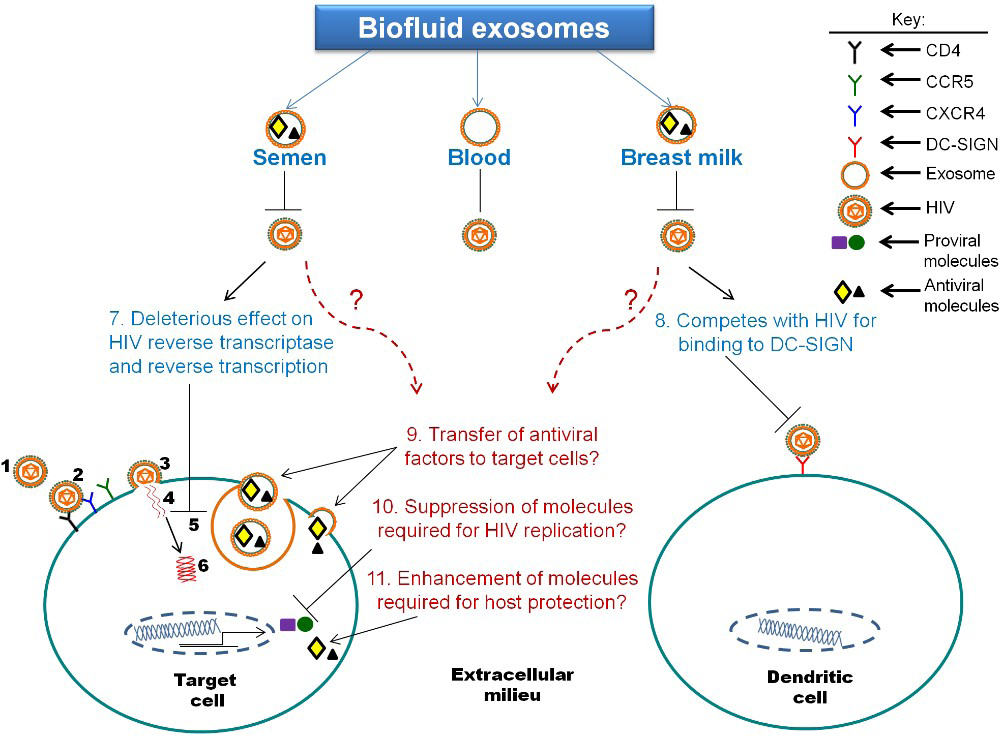
5. The Function of Biofluid Exosomes is Distinct from Cell Culture-Derived Exosomes: Intrinsic Protection from HIV-1
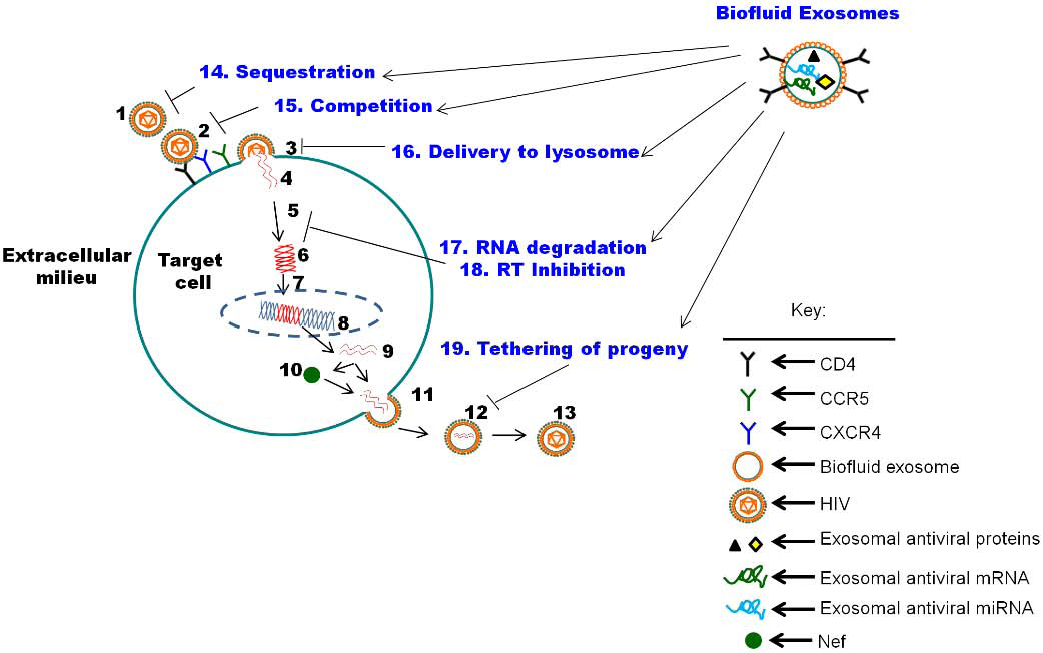
6. Exosome-Associated HIV-1 Host Restriction Factors, Cytokines, and miRNAs
| miRNA | Exosome Source | References |
|---|---|---|
| miR-28 | Dendritic cells, J77 T cells, semen, blood | [133,134] |
| miR-29a | Breast milk, semen, dendritic cells | [25,133] |
| miR-29b | Semen, blood | [25,133] |
| miR-125b | Dendritic cells semen, blood | [25,133,134] |
| miR-149 | Breast milk, semen dendritic cells, blood | [25,133] |
| miR-150 | J77 T cells, breast milk, blood, semen | [133] |
| miR-198 | Dendritic cells | [133] |
| miR-223 | Dendritic cells, serum, breast milk, blood, semen | [133,134] |
| miR-324 | Semen, blood, dendritic cells, | [25,133] |
| miR-378 | Breast milk, semen, blood, dendritic cells, | [25,133] |
| miR-382 | Semen, blood | [25,133] |
| vmiR88 | Primary alveolar macrophages, serum | [90] |
| vmiR99 | Primary alveolar macrophages, serum | [90] |
| vmiRTAR | J1.1 cells | [89] |
7. Exosomes and Their Promise for Diagnostic and Therapeutic Applications
8. Conclusions and Perspectives
Acknowledgements
Conflicts of Interest
References
- Harding, C.; Heuser, J.; Stahl, P. Receptor-mediated endocytosis of transferrin and recycling of the transferrin receptor in rat reticulocytes. J. Cell Biol. 1983, 97, 329–339. [Google Scholar] [CrossRef] [PubMed]
- Pan, B.T.; Teng, K.; Wu, C.; Adam, M.; Johnstone, R.M. Electron microscopic evidence for externalization of the transferrin receptor in vesicular form in sheep reticulocytes. J. Cell Biol. 1985, 101, 942–948. [Google Scholar] [CrossRef] [PubMed]
- Johnstone, R.M.; Adam, M.; Hammond, J.R.; Orr, L.; Turbide, C. Vesicle formation during reticulocyte maturation. Association of plasma membrane activities with released vesicles (exosomes). J. Biol. Chem. 1987, 262, 9412–9420. [Google Scholar] [PubMed]
- Raposo, G.; Nijman, H.W.; Stoorvogel, W.; Liejendekker, R.; Harding, C.V.; Melief, C.J.; Geuze, H.J. B lymphocytes secrete antigen-presenting vesicles. J. Exp. Med. 1996, 183, 1161–1172. [Google Scholar] [CrossRef] [PubMed]
- Valadi, H.; Ekstrom, K.; Bossios, A.; Sjostrand, M.; Lee, J.J.; Lotvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef] [PubMed]
- Simons, M.; Raposo, G. Exosomes—Vesicular carriers for intercellular communication. Curr. Opin. Cell Biol. 2009, 21, 575–581. [Google Scholar] [CrossRef] [PubMed]
- Bobrie, A.; Colombo, M.; Raposo, G.; Thery, C. Exosome secretion: Molecular mechanisms and roles in immune responses. Traffic 2011, 12, 1659–1668. [Google Scholar] [CrossRef] [PubMed]
- Thery, C.; Ostrowski, M.; Segura, E. Membrane vesicles as conveyors of immune responses. Nat. Rev. Immunol. 2009, 9, 581–593. [Google Scholar] [CrossRef] [PubMed]
- Lotvall, J.; Valadi, H. Cell to cell signalling via exosomes through esRNA. Cell Adhes. Migrat. 2007, 1, 156–158. [Google Scholar] [CrossRef]
- Caby, M.P.; Lankar, D.; Vincendeau-Scherrer, C.; Raposo, G.; Bonnerot, C. Exosomal-like vesicles are present in human blood plasma. Int. Immunol. 2005, 17, 879–887. [Google Scholar] [CrossRef] [PubMed]
- Pisitkun, T.; Shen, R.F.; Knepper, M.A. Identification and proteomic profiling of exosomes in human urine. Proc. Natl. Acad. Sci. USA 2004, 101, 13368–13373. [Google Scholar] [CrossRef] [PubMed]
- Palanisamy, V.; Sharma, S.; Deshpande, A.; Zhou, H.; Gimzewski, J.; Wong, D.T. Nanostructural and transcriptomic analyses of human saliva derived exosomes. PLoS ONE 2010, 5, e8577. [Google Scholar] [CrossRef] [PubMed]
- Admyre, C.; Johansson, S.M.; Qazi, K.R.; Filen, J.J.; Lahesmaa, R.; Norman, M.; Neve, E.P.; Scheynius, A.; Gabrielsson, S. Exosomes with immune modulatory features are present in human breast milk. J. Immunol. 2007, 179, 1969–1978. [Google Scholar] [CrossRef] [PubMed]
- Admyre, C.; Grunewald, J.; Thyberg, J.; Gripenback, S.; Tornling, G.; Eklund, A.; Scheynius, A.; Gabrielsson, S. Exosomes with major histocompatibility complex class II and co-stimulatory molecules are present in human BAL fluid. Eur. Respir. J. 2003, 22, 578–583. [Google Scholar] [CrossRef] [PubMed]
- Llorente, A.; de Marco, M.C.; Alonso, M.A. Caveolin-1 and MAL are located on prostasomes secreted by the prostate cancer PC-3 cell line. J. Cell Sci. 2004, 117, 5343–5351. [Google Scholar] [CrossRef] [PubMed]
- Sahlen, G.E.; Egevad, L.; Ahlander, A.; Norlen, B.J.; Ronquist, G.; Nilsson, B.O. Ultrastructure of the secretion of prostasomes from benign and malignant epithelial cells in the prostate. Prostate 2002, 53, 192–199. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, G.K.; Larsson, A.; Stavreus-Evers, A.; Ronquist, G. Prostasomes are heterogeneous regarding size and appearance but affiliated to one DNA-containing exosome family. Prostate 2012, 72, 1736–1745. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, R.; Saez, F.; Girouard, J.; Frenette, G. Role of exosomes in sperm maturation during the transit along the male reproductive tract. Blood Cells Mol. Dis. 2005, 35, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Frenette, G.; Girouard, J.; D’Amours, O.; Allard, N.; Tessier, L.; Sullivan, R. Characterization of two distinct populations of epididymosomes collected in the intraluminal compartment of the bovine cauda epididymis. Biol. Reprod. 2010, 83, 473–480. [Google Scholar] [CrossRef] [PubMed]
- Renneberg, H.; Albrecht, M.; Kurek, R.; Krause, E.; Lottspeich, F.; Aumuller, G.; Wilhelm, B. Identification and characterization of neutral endopeptidase (EC 3. 4. 24. 11) from human prostasomes—Localization in prostatic tissue and cell lines. Prostate 2001, 46, 173–183. [Google Scholar] [CrossRef]
- Renneberg, H.; Konrad, L.; Dammshauser, I.; Seitz, J.; Aumuller, G. Immunohistochemistry of prostasomes from human semen. Prostate 1997, 30, 98–106. [Google Scholar] [CrossRef]
- Skibinski, G.; Kelly, R.W.; James, K. Expression of a common secretory granule specific protein as a marker for the extracellular organelles (prostasomes) in human semen. Fertil. Steril. 1994, 61, 755–759. [Google Scholar] [PubMed]
- Stridsberg, M.; Fabiani, R.; Lukinius, A.; Ronquist, G. Prostasomes are neuroendocrine-like vesicles in human semen. Prostate 1996, 29, 287–295. [Google Scholar] [CrossRef]
- Madison, M.N.; Roller, R.J.; Okeoma, C.M. Human semen contains exosomes with potent anti-HIV-1 activity. Retrovirology 2014, 11, 102. [Google Scholar] [CrossRef] [PubMed]
- Vojtech, L.; Woo, S.; Hughes, S.; Levy, C.; Ballweber, L.; Sauteraud, R.P.; Strobl, J.; Westerberg, K.; Gottardo, R.; Tewari, M.; et al. Exosomes in human semen carry a distinctive repertoire of small non-coding RNAs with potential regulatory functions. Nucleic Acids Res. 2014, 42, 7290–7304. [Google Scholar] [CrossRef] [PubMed]
- Kaur, S.; Singh, S.P.; Elkahloun, A.G.; Wu, W.; Abu-Asab, M.S.; Roberts, D.D. CD47-dependent immunomodulatory and angiogenic activities of extracellular vesicles produced by T cells. Matrix Biol. 2014, 37, 49–59. [Google Scholar] [CrossRef] [PubMed]
- Madison, M.N.; Jones, P.H.; Okeoma, C.M. Exosomes in human semen restrict HIV-1 transmission by vaginal cells and block intravaginal replication of LP-BM5 murine AIDS virus complex. Virology 2015, 482, 189–201. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Liu, K.; Liu, Y.; Xu, Y.; Zhang, F.; Yang, H.; Liu, J.; Pan, T.; Chen, J.; Wu, M.; et al. Exosomes mediate the cell-to-cell transmission of IFN-α-induced antiviral activity. Nat. Immunol. 2013, 14, 793–803. [Google Scholar] [CrossRef] [PubMed]
- Naslund, T.I.; Paquin-Proulx, D.; Paredes, P.T.; Vallhov, H.; Sandberg, J.K.; Gabrielsson, S. Exosomes from breast milk inhibit HIV-1 infection of dendritic cells and subsequent viral transfer to CD4+ T cells. Aids 2014, 28, 171–180. [Google Scholar] [CrossRef] [PubMed]
- Arenaccio, C.; Chiozzini, C.; Columba-Cabezas, S.; Manfredi, F.; Affabris, E.; Baur, A.; Federico, M. Exosomes from human immunodeficiency virus type 1 (HIV-1)-infected cells license quiescent CD4+ T lymphocytes to replicate HIV-1 through a Nef- and ADAM17-dependent mechanism. J. Virol. 2014, 88, 11529–11539. [Google Scholar] [CrossRef] [PubMed]
- Arenaccio, C.; Chiozzini, C.; Columba-Cabezas, S.; Manfredi, F.; Federico, M. Cell activation and HIV-1 replication in unstimulated CD4+ T lymphocytes ingesting exosomes from cells expressing defective HIV-1. Retrovirology 2014, 11, 46. [Google Scholar] [CrossRef] [PubMed]
- Mahauad-Fernandez, W.D.; Jones, P.H.; Okeoma, C.M. Critical role for BST-2 in acute Chikungunya virus infection. J. Gen. Virol. 2014. [Google Scholar] [CrossRef]
- De Carvalho, J.V.; de Castro, R.O.; da Silva, E.Z.; Silveira, P.P.; da Silva-Januario, M.E.; Arruda, E.; Jamur, M.C.; Oliver, C.; Aguiar, R.S.; daSilva, L.L. Nef neutralizes the ability of exosomes from CD4+ T cells to act as decoys during HIV-1 infection. PLoS ONE 2014, 9, e113691. [Google Scholar] [CrossRef] [PubMed]
- Lenassi, M.; Cagney, G.; Liao, M.; Vaupotic, T.; Bartholomeeusen, K.; Cheng, Y.; Krogan, N.J.; Plemenitas, A.; Peterlin, B.M. HIV Nef is secreted in exosomes and triggers apoptosis in bystander CD4+ T cells. Traffic 2010, 11, 110–122. [Google Scholar] [CrossRef] [PubMed]
- Khatua, A.K.; Taylor, H.E.; Hildreth, J.E.; Popik, W. Inhibition of LINE-1 and Alu retrotransposition by exosomes encapsidating APOBEC3G and APOBEC3F. Virology 2010, 400, 68–75. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, D.L.; Nakayasu, E.S.; Joffe, L.S.; Guimaraes, A.J.; Sobreira, T.J.; Nosanchuk, J.D.; Cordero, R.J.; Frases, S.; Casadevall, A.; Almeida, I.C.; et al. Characterization of yeast extracellular vesicles: Evidence for the participation of different pathways of cellular traffic in vesicle biogenesis. PLoS ONE 2010, 5, e11113. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.C.; Lee, E.J.; Lee, J.H.; Jun, S.H.; Choi, C.W.; Kim, S.I.; Kang, S.S.; Hyun, S. Klebsiella pneumoniae secretes outer membrane vesicles that induce the innate immune response. FEMS Microbiol. Lett. 2012, 331, 17–24. [Google Scholar] [CrossRef] [PubMed]
- Thery, C.; Boussac, M.; Veron, P.; Ricciardi-Castagnoli, P.; Raposo, G.; Garin, J.; Amigorena, S. Proteomic analysis of dendritic cell-derived exosomes: A secreted subcellular compartment distinct from apoptotic vesicles. J. Immunol. 2001, 166, 7309–7318. [Google Scholar] [CrossRef] [PubMed]
- Van Niel, G.; Porto-Carreiro, I.; Simoes, S.; Raposo, G. Exosomes: A common pathway for a specialized function. J. Biochem. 2006, 140, 13–21. [Google Scholar] [CrossRef] [PubMed]
- Hemler, M.E. Tetraspanin proteins mediate cellular penetration, invasion, and fusion events and define a novel type of membrane microdomain. Ann. Rev. Cell Dev. Biol. 2003, 19, 397–422. [Google Scholar] [CrossRef] [PubMed]
- Zoller, M. Tetraspanins: Push and pull in suppressing and promoting metastasis. Nat. Rev. Cancer 2009, 9, 40–55. [Google Scholar] [CrossRef] [PubMed]
- Wubbolts, R.; Leckie, R.S.; Veenhuizen, P.T.; Schwarzmann, G.; Mobius, W.; Hoernschemeyer, J.; Slot, J.W.; Geuze, H.J.; Stoorvogel, W. Proteomic and biochemical analyses of human B cell-derived exosomes. Potential implications for their function and multivesicular body formation. J. Biol. Chem. 2003, 278, 10963–10972. [Google Scholar] [CrossRef] [PubMed]
- Thery, C.; Regnault, A.; Garin, J.; Wolfers, J.; Zitvogel, L.; Ricciardi-Castagnoli, P.; Raposo, G.; Amigorena, S. Molecular characterization of dendritic cell-derived exosomes. Selective accumulation of the heat shock protein HSC73. J. Cell. Biol. 1999, 147, 599–610. [Google Scholar] [CrossRef] [PubMed]
- Matsuo, H.; Chevallier, J.; Mayran, N.; le Blanc, I.; Ferguson, C.; Fauré, J.; Blanc, N.S.; Matile, S.; Dubochet, J.; Sadoul, R.; et al. Role of LBPA and Alix in multivesicular liposome formation and endosome organization. Science 2004, 303, 531–534. [Google Scholar] [CrossRef] [PubMed]
- Bellingham, S.A.; Coleman, B.M.; Hill, A.F. Small RNA deep sequencing reveals a distinct miRNA signature released in exosomes from prion-infected neuronal cells. Nucleic Acids Res. 2012, 40, 10937–10949. [Google Scholar] [CrossRef] [PubMed]
- Nolte-’t Hoen, E.N.; Buermans, H.P.; Waasdorp, M.; Stoorvogel, W.; Wauben, M.H.; ’t Hoen, P.A. Deep sequencing of RNA from immune cell-derived vesicles uncovers the selective incorporation of small non-coding RNA biotypes with potential regulatory functions. Nucleic Acids Res. 2012, 40, 9272–9285. [Google Scholar] [CrossRef] [PubMed]
- ExoCarta 2015. Available online: http://exocarta.org/index.html (accessed on 16 July 2015).
- Vesiclepedia 2015. Available online: http://www.microvesicles.org/ (accessed on 16 July 2015).
- Colombo, M.; Moita, C.; Van Niel, G.; Kowal, J.; Vigneron, J.; Benaroch, P.; Manel, N.; Moita, L.F.; Théry, C.; Raposo, G. Analysis of ESCRT functions in exosome biogenesis, composition and secretion highlights the heterogeneity of extracellular vesicles. J. Cell Sci. 2013, 126, 5553–5565. [Google Scholar] [CrossRef] [PubMed]
- Hoshino, D.; Kirkbride, K.; Costello, K.; Clark, E.; Sinha, S.; Grega-Larson, N.; Tyska, M.; Weaver, A. Exosome secretion is enhanced by invadopodia and drives invasive behavior. Cell Rep. 2013, 5, 1159–1168. [Google Scholar] [CrossRef] [PubMed]
- Gross, J.C.; Chaudhary, V.; Bartscherer, K.; Boutros, M. Active Wnt proteins are secreted on exosomes. Nat. Cell Biol. 2012, 14, 1036–1045. [Google Scholar] [CrossRef] [PubMed]
- Baietti, M.F.; Zhang, Z.; Mortier, E.; Melchior, A.; Degeest, G.; Geeraerts, A.; Ivarsson, Y.; Depoortere, F.; Coomans, C.; Vermeiren, E.; et al. Syndecan-syntenin-ALIX regulates the biogenesis of exosomes. Nat. Cell Biol. 2012, 14, 677–685. [Google Scholar] [CrossRef] [PubMed]
- Stuffers, S.; Sem Wegner, C.; Stenmark, H.; Brech, A. Multivesicular endosome biogenesis in the absence of ESCRTs. Traffic 2009, 10, 925–937. [Google Scholar] [CrossRef] [PubMed]
- Trajkovic, K.; Hsu, C.; Chiantia, S.; Rajendran, L.; Wenzel, D.; Wieland, F.; Schwille, P.; Brugger, B.; Simons, M. Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science 2008, 319, 1244–1247. [Google Scholar] [CrossRef] [PubMed]
- Van Niel, G.; Charrin, S.; Simoes, S.; Romao, M.; Rochin, L.; Saftig, P.; Marks, M.; Rubinstein, E.; Raposo, G. The Tetraspanin CD63 Regulates ESCRT-independent and -dependent endosomal sorting during melanogenesis. Dev. Cell 2011, 21, 708–721. [Google Scholar] [CrossRef] [PubMed]
- Perez-Hernandez, D.; Gutiérrez-Vázquez, C.; Jorge, I.; López-Martín, S.; Ursa, A.; Sánchez-Madrid, F.; Vázquez, J.; Yañez-Mó, M. The intracellular interactome of tetraspanin-enriched microdomains reveals their function as sorting machineries toward exosomes. J. Biol. Chem. 2013, 288, 11649–11661. [Google Scholar] [CrossRef] [PubMed]
- Ostrowski, M.; Carmo, N.B.; Krumeich, S.; Fanget, I.; Raposo, G.; Savina, A.; Moita, C.F.; Schauer, K.; Hume, A.N.; Freitas, R.P.; et al. Rab27a and Rab27b control different steps of the exosome secretion pathway. Nat. Cell Biol. 2010, 12 (suppl. 11–13), 19–30. [Google Scholar] [CrossRef] [PubMed]
- Savina, A.; Fader, C.M.; Damiani, M.T.; Colombo, M.I. Rab11 promotes docking and fusion of multivesicular bodies in a calcium-dependent manner. Traffic 2005, 6, 131–143. [Google Scholar] [CrossRef] [PubMed]
- Garrus, J.E.; Von Schwedler, U.K.; Pornillos, O.W.; Morham, S.G.; Zavitz, K.H.; Wang, H.E.; Wettstein, D.A.; Stray, K.M.; Côté, M.; Rich, R.L.; et al. Tsg101 and the vacuolar protein sorting pathway are essential for HIV-1 budding. Cell 2001, 107, 55–65. [Google Scholar] [CrossRef]
- Pornillos, O.; Higginson, D.S.; Stray, K.M.; Fisher, R.D.; Garrus, J.E.; Payne, M.; He, G.P.; Wang, H.E.; Morham, S.G.; Sundquist, W.I. HIV Gag mimics the Tsg101-recruiting activity of the human Hrs protein. J. Cell Biol. 2003, 162, 425–434. [Google Scholar] [CrossRef] [PubMed]
- Booth, A.M.; Fang, Y.; Fallon, J.K.; Yang, J.M.; Hildreth, J.E.K.; Gould, S.J.; Sandefur, S.; Varthakavi, V. Exosomes and HIV Gag bud from endosome-like domains of the T cell plasma membrane. J. Cell Biol. 2006, 172, 923–935. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.J.; Horton, R.; Varthakavi, V.; Spearman, P.; Ratner, L. Formation and release of virus-like particles by HIV-1 matrix protein. Aids 1999, 13, 281–283. [Google Scholar] [CrossRef] [PubMed]
- Cantin, R.; Diou, J.; Belanger, D.; Tremblay, A.M.; Gilbert, C. Discrimination between exosomes and HIV-1: Purification of both vesicles from cell-free supernatants. J. Immunol. Methods 2008, 338, 21–30. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Wu, N.; Gan, X.; Yan, W.; Morrell, J.C.; Gould, S.J. Higher-order oligomerization targets plasma membrane proteins and HIV gag to exosomes. PLoS Biol. 2007, 5, e158. [Google Scholar] [CrossRef] [PubMed]
- Usami, Y.; Popov, S.; Popova, E.; Inoue, M.; Weissenhorn, W.; Gottlinge, H.G. The ESCRT pathway and HIV-1 budding. Biochem. Soc. Trans. 2009, 37, 181–184. [Google Scholar] [CrossRef] [PubMed]
- Chertova, E.; Chertov, O.; Coren, L.V.; Roser, J.D.; Trubey, C.M.; Bess, J.W., Jr.; Sowder, R.C., 2nd; Barsov, E.; Hood, B.L.; Fisher, R.J.; et al. Proteomic and biochemical analysis of purified human immunodeficiency virus type 1 produced from infected monocyte-derived macrophages. J. Virol. 2006, 80, 9039–9052. [Google Scholar] [CrossRef] [PubMed]
- Gould, S.J.; Booth, A.M.; Hildreth, J.E. The Trojan exosome hypothesis. Proc. Natl. Acad. Sci. USA 2003, 100, 10592–10597. [Google Scholar] [CrossRef] [PubMed]
- Grigorov, B.; Attuil-Audenis, V.; Perugi, F.; Nedelec, M.; Watson, S.; Pique, C.; Darlix, J.L.; Conjeaud, H.; Muriaux, D. A role for CD81 on the late steps of HIV-1 replication in a chronically infected T cell line. Retrovirology 2009, 6. [Google Scholar] [CrossRef] [PubMed]
- Izquierdo-Useros, N.; Naranjo-Gomez, M.; Archer, J.; Hatch, S.C.; Erkizia, I.; Blanco, J.; Borras, F.E.; Puertas, M.C.; Connor, J.H.; Fernandez-Figueras, M.T.; et al. Capture and transfer of HIV-1 particles by mature dendritic cells converges with the exosome-dissemination pathway. Blood 2009, 113, 2732–2741. [Google Scholar] [CrossRef] [PubMed]
- Jolly, C.; Sattentau, Q.J. Human immunodeficiency virus type 1 assembly, budding, and cell-cell spread in T cells take place in tetraspanin-enriched plasma membrane domains. J. Virol. 2007, 81, 7873–7884. [Google Scholar] [CrossRef] [PubMed]
- Sato, K.; Aoki, J.; Misawa, N.; Daikoku, E.; Sano, K.; Tanaka, Y.; Koyanagi, Y. Modulation of human immunodeficiency virus type 1 infectivity through incorporation of tetraspanin proteins. J. Virol. 2008, 82, 1021–1033. [Google Scholar] [CrossRef] [PubMed]
- Izquierdo-Useros, N.; Lorizate, M.; Puertas, M.C.; Rodriguez-Plata, M.T.; Zangger, N.; Erikson, E.; Pino, M.; Erkizia, I.; Glass, B.; Clotet, B.; et al. Siglec-1 is a novel dendritic cell receptor that mediates HIV-1 trans-infection through recognition of viral membrane gangliosides. PLoS Biol. 2012, 10, e1001448. [Google Scholar] [CrossRef] [PubMed]
- Saunderson, S.C.; Dunn, A.C.; Crocker, P.R.; McLellan, A.D. CD169 mediates the capture of exosomes in spleen and lymph node. Blood 2014, 123, 208–216. [Google Scholar] [CrossRef] [PubMed]
- Izquierdo-Useros, N.; Blanco, J.; Erkizia, I.; Fernandez-Figueras, M.T.; Borras, F.E.; Naranjo-Gomez, M.; Bofill, M.; Ruiz, L.; Clotet, B.; Martinez-Picado, J. Maturation of blood-derived dendritic cells enhances human immunodeficiency virus type 1 capture and transmission. J. Virol. 2007, 81, 7559–7570. [Google Scholar] [CrossRef] [PubMed]
- Puryear, W.B.; Akiyama, H.; Geer, S.D.; Ramirez, N.P.; Yu, X.; Reinhard, B.M.; Gummuluru, S. Interferon-inducible mechanism of dendritic cell-mediated HIV-1 dissemination is dependent on Siglec-1/CD169. PLoS Pathog. 2013, 9, e1003291. [Google Scholar] [CrossRef] [PubMed]
- Raymond, A.D.; Campbell-Sims, T.C.; Khan, M.; Lang, M.; Huang, M.B.; Bond, V.C.; Powell, M.D. HIV Type 1 Nef is released from infected c ells in CD45+ microvesicles and is present in the plasma of HIV-infected individuals. AIDS Res. Hum. Retrovir. 2011, 27, 167–178. [Google Scholar] [CrossRef] [PubMed]
- Muratori, C.; Cavallin, L.E.; Kratzel, K.; Tinari, A.; de Milito, A.; Fais, S.; D’Aloja, P.; Federico, M.; Vullo, V.; Fomina, A.; et al. Massive secretion by T cells is caused by HIV Nef in infected cells and by Nef transfer to bystander cells. Cell Host Microbe 2009, 6, 218–230. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.A.; Huang, M.B.; Campbell, P.E.; Roth, W.W.; Campbell, T.; Khan, M.; Newman, G.; Villinger, F.; Powell, M.D.; Bond, V.C. Genetic characterization of HIV type 1 Nef-induced vesicle secretion. AIDS Res. Hum. Retrovir. 2010, 26, 173–192. [Google Scholar] [CrossRef] [PubMed]
- Lubben, N.B.; Sahlender, D.A.; Motley, A.M.; Lehner, P.J.; Benaroch, P.; Robinson, M.S. HIV-1 Nef-induced down-regulation of MHC class I requires AP-1 and clathrin but not PACS-1 and is impeded by AP-2. Mol. Biol. Cell 2007, 18, 3351–3365. [Google Scholar] [CrossRef] [PubMed]
- Schaefer, M.R.; Wonderlich, E.R.; Roeth, J.F.; Leonard, J.A.; Collins, K.L. HIV-1 Nef targets MHC-I and CD4 for degradation via a final common β-COP-dependent pathway in T cells. PLoS Pathog. 2008, 4, e1000131. [Google Scholar] [CrossRef] [PubMed]
- Da Silva, L.L.; Sougrat, R.; Burgos, P.V.; Janvier, K.; Mattera, R.; Bonifacino, J.S. Human immunodeficiency virus type 1 Nef protein targets CD4 to the multivesicular body pathway. J. Virol. 2009, 83, 6578–6590. [Google Scholar] [CrossRef] [PubMed]
- Roeth, J.F.; Williams, M.; Kasper, M.R.; Filzen, T.M.; Collins, K.L. HIV-1 Nef disrupts MHC-I trafficking by recruiting AP-1 to the MHC-I cytoplasmic tail. J. Cell Biol. 2004, 167, 903–913. [Google Scholar] [CrossRef] [PubMed]
- Anderson, S.J.; Lenburg, M.; Landau, N.R.; Garcia, J.V. The cytoplasmic domain of CD4 is sufficient for its down-regulation from the cell surface by human immunodeficiency virus type 1 Nef. J. Virol. 1994, 68, 3092–3101. [Google Scholar] [PubMed]
- Rhee, S.S.; Marsh, J.W. Human immunodeficiency virus type 1 Nef-induced down-modulation of CD4 is due to rapid internalization and degradation of surface CD4. J. Virol. 1994, 68, 5156–5163. [Google Scholar] [PubMed]
- Amorim, N.A.; da Silva, E.M.; de Castro, R.O.; da Silva-Januario, M.E.; Mendonca, L.M.; Bonifacino, J.S.; da Costa, L.J.; da Silva, L.L. Interaction of HIV-1 Nef protein with the host protein Alix promotes lysosomal targeting of CD4 receptor. J. Biol. Chem. 2014, 289, 27744–27756. [Google Scholar] [CrossRef] [PubMed]
- Campbell, T.D.; Khan, M.; Huang, M.B.; Bond, V.C.; Powell, M.D. HIV-1 Nef protein is secreted into vesicles that can fuse with target cells and virions. Ethn. Dis. 2008, 18, S2-14–S2-19. [Google Scholar] [PubMed]
- Columba Cabezas, S.; Federico, M. Sequences within RNA coding for HIV-1 Gag p17 are efficiently targeted to exosomes. Cell Microbiol. 2013, 15, 412–429. [Google Scholar] [CrossRef] [PubMed]
- Chivero, E.T.; Bhattarai, N.; Rydze, R.T.; Winters, M.A.; Holodniy, M.; Stapleton, J.T. Human pegivirus RNA is found in multiple blood mononuclear cells in vivo and serum-derived viral RNA-containing particles are infectious in vitro. J. Gen. Virol. 2014, 95, 1307–1319. [Google Scholar] [CrossRef] [PubMed]
- Narayanan, A.; Iordanskiy, S.; Das, R.; Van Duyne, R.; Santos, S.; Jaworski, E.; Guendel, I.; Sampey, G.; Dalby, E.; Iglesias-Ussel, M.; et al. Exosomes derived from HIV-1-infected cells contain trans-activation response element RNA. J. Biol. Chem. 2013, 288, 20014–20033. [Google Scholar] [CrossRef] [PubMed]
- Bernard, M.A.; Zhao, H.; Yue, S.C.; Anandaiah, A.; Koziel, H.; Tachado, S.D. Novel HIV-1 miRNAs stimulate TNFalpha release in human macrophages via TLR8 signaling pathway. PLoS ONE 2014, 9, e106006. [Google Scholar] [CrossRef] [PubMed]
- Kadiu, I.; Narayanasamy, P.; Dash, P.K.; Zhang, W.; Gendelman, H.E. Biochemical and biologic characterization of exosomes and microvesicles as facilitators of HIV-1 infection in macrophages. J. Immunol. 2012, 189, 744–754. [Google Scholar] [CrossRef] [PubMed]
- Mack, M.; Kleinschmidt, A.; Bruhl, H.; Klier, C.; Nelson, P.J.; Cihak, J.; Plachy, J.; Stangassinger, M.; Erfle, V.; Schlondorff, D. Transfer of the chemokine receptor CCR5 between cells by membrane-derived microparticles: A mechanism for cellular human immunodeficiency virus 1 infection. Nat. Med. 2000, 6, 769–775. [Google Scholar] [CrossRef] [PubMed]
- Rozmyslowicz, T.; Majka, M.; Kijowski, J.; Murphy, S.L.; Conover, D.O.; Poncz, M.; Ratajczak, J.; Gaulton, G.N.; Ratajczak, M.Z. Platelet- and megakaryocyte-derived microparticles transfer CXCR4 receptor to CXCR4-null cells and make them susceptible to infection by X4-HIV. AIDS 2003, 17, 33–42. [Google Scholar] [CrossRef] [PubMed]
- Chariot, P.; Dubreuil-Lemaire, M.L.; Zhou, J.Y.; Lamia, B.; Dume, L.; Larcher, B.; Monnet, I.; Levy, Y.; Astier, A.; Gherardi, R. Muscle involvement in human immunodeficiency virus-infected patients is associated with marked selenium deficiency. Muscle Nerve 1997, 20, 386–389. [Google Scholar] [CrossRef]
- Scadden, D.T.; Zeira, M.; Woon, A.; Wang, Z.; Schieve, L.; Ikeuchi, K.; Lim, B.; Groopman, J.E. Human immunodeficiency virus infection of human bone marrow stromal fibroblasts. Blood 1990, 76, 317–322. [Google Scholar] [PubMed]
- Permanyer, M.; Ballana, E.; Badia, R.; Pauls, E.; Clotet, B.; Este, J.A. Trans-infection but not infection from within endosomal compartments after cell-to-cell HIV-1 transfer to CD4+ T cells. J. Biol. Chem. 2012, 287, 32017–32026. [Google Scholar] [CrossRef] [PubMed]
- Peterlin, B.M.; Trono, D. Hide, shield and strike back: How HIV-infected cells avoid immune eradication. Nat. Rev. Immunol. 2003, 3, 97–107. [Google Scholar] [CrossRef] [PubMed]
- Costa, L.J.; Chen, N.; Lopes, A.; Aguiar, R.S.; Tanuri, A.; Plemenitas, A.; Peterlin, B.M. Interactions between Nef and AIP1 proliferate multivesicular bodies and facilitate egress of HIV-1. Retrovirology 2006, 3. [Google Scholar] [CrossRef] [PubMed]
- Madrid, R.; Janvier, K.; Hitchin, D.; Day, J.; Coleman, S.; Noviello, C.; Bouchet, J.; Benmerah, A.; Guatelli, J.; Benichou, S. Nef-induced alteration of the early/recycling endosomal compartment correlates with enhancement of HIV-1 infectivity. J. Biol. Chem. 2005, 280, 5032–5044. [Google Scholar] [CrossRef] [PubMed]
- Stumptner-Cuvelette, P.; Jouve, M.; Helft, J.; Dugast, M.; Glouzman, A.S.; Jooss, K.; Raposo, G.; Benaroch, P. Human immunodeficiency virus-1 Nef expression induces intracellular accumulation of multivesicular bodies and major histocompatibility complex class II complexes: Potential role of phosphatidylinositol 3-kinase. Mol. Biol. Cell 2003, 14, 4857–4870. [Google Scholar] [CrossRef] [PubMed]
- Admyre, C.; Johansson, S.M.; Paulie, S.; Gabrielsson, S. Direct exosome stimulation of peripheral human T cells detected by ELISPOT. Eur. J. Immunol. 2006, 36, 1772–1781. [Google Scholar] [CrossRef] [PubMed]
- Kesimer, M.; Scull, M.; Brighton, B.; deMaria, G.; Burns, K.; O’Neal, W.; Pickles, R.J.; Sheehan, J.K. Characterization of exosome-like vesicles released from human tracheobronchial ciliated epithelium: A possible role in innate defense. FASEB J. 2009, 23, 1858–1868. [Google Scholar] [CrossRef] [PubMed]
- Khatua, A.K.; Taylor, H.E.; Hildreth, J.E.; Popik, W. Exosomes packaging APOBEC3G confer human immunodeficiency virus resistance to recipient cells. J. Virol. 2009, 83, 512–521. [Google Scholar] [CrossRef] [PubMed]
- Xiang, X.; Liu, Y.; Zhuang, X.; Zhang, S.; Michalek, S.; Taylor, D.D.; Grizzle, W.; Zhang, H.G. TLR2-mediated expansion of MDSCs is dependent on the source of tumor exosomes. Am. J. Pathol. 2010, 177, 1606–1610. [Google Scholar] [CrossRef] [PubMed]
- Carlsson, L.; Pahlson, C.; Bergquist, M.; Ronquist, G.; Stridsberg, M. Antibacterial activity of human prostasomes. Prostate 2000, 44, 279–286. [Google Scholar] [CrossRef]
- Kitamura, M.; Namiki, M.; Matsumiya, K.; Tanaka, K.; Matsumoto, M.; Hara, T.; Kiyohara, H.; Okabe, M.; Okuyama, A.; Seya, T. Membrane cofactor protein (CD46) in seminal plasma is a prostasome-bound form with complement regulatory activity and measles virus neutralizing activity. Immunology 1995, 84, 626–632. [Google Scholar] [PubMed]
- Arienti, G.; Carlini, E.; Nicolucci, A.; Cosmi, E.V.; Santi, F.; Palmerini, C.A. The motility of human spermatozoa as influenced by prostasomes at various pH levels. Biol. Cell 1999, 91, 51–54. [Google Scholar] [CrossRef] [PubMed]
- Babiker, A.A.; Ronquist, G.; Nilsson, U.R.; Nilsson, B. Transfer of prostasomal CD59 to CD59-deficient red blood cells results in protection against complement-mediated hemolysis. Am. J. Reprod. Immunol. 2002, 47, 183–192. [Google Scholar] [CrossRef] [PubMed]
- Burden, H.P.; Holmes, C.H.; Persad, R.; Whittington, K. Prostasomes—Their effects on human male reproduction and fertility. Hum. Reprod. Update 2006, 12, 283–292. [Google Scholar] [CrossRef] [PubMed]
- Cross, N.L.; Mahasreshti, P. Prostasome fraction of human seminal plasma prevents sperm from becoming acrosomally responsive to the agonist progesterone. Arch. Androl. 1997, 39, 39–44. [Google Scholar] [CrossRef] [PubMed]
- Kelly, R.W. Immunosuppressive mechanisms in semen: Implications for contraception. Hum. Reprod. 1995, 10, 1686–1693. [Google Scholar] [PubMed]
- Palmerini, C.A.; Saccardi, C.; Carlini, E.; Fabiani, R.; Arienti, G. Fusion of prostasomes to human spermatozoa stimulates the acrosome reaction. Fertil. Steril. 2003, 80, 1181–1184. [Google Scholar] [CrossRef]
- Park, K.H.; Kim, B.J.; Kang, J.; Nam, T.S.; Lim, J.M.; Kim, H.T.; Park, J.K.; Kim, Y.G.; Chae, S.W.; Kim, U.H. Ca2+ signaling tools acquired from prostasomes are required for progesterone-induced sperm motility. Sci. Signal. 2011, 4, ra31. [Google Scholar] [CrossRef] [PubMed]
- Skibinski, G.; Kelly, R.W.; Harkiss, D.; James, K. Immunosuppression by human seminal plasma—Extracellular organelles (prostasomes) modulate activity of phagocytic cells. Am. J. Reprod. Immunol. 1992, 28, 97–103. [Google Scholar] [CrossRef] [PubMed]
- Tarazona, R.; Delgado, E.; Guarnizo, M.C.; Roncero, R.G.; Morgado, S.; Sanchez-Correa, B.; Gordillo, J.J.; Dejulian, J.; Casado, J.G. Human prostasomes express CD48 and interfere with NK cell function. Immunobiology 2011, 216, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Babiker, A.A.; Nilsson, B.; Ronquist, G.; Carlsson, L.; Ekdahl, K.N. Transfer of functional prostasomal CD59 of metastatic prostatic cancer cell origin protects cells against complement attack. Prostate 2005, 62, 105–114. [Google Scholar] [CrossRef] [PubMed]
- Jones, J.L.; Saraswati, S.; Block, A.S.; Lichti, C.F.; Mahadevan, M.; Diekman, A.B. Galectin-3 is associated with prostasomes in human semen. Glycoconj. J. 2010, 27, 227–236. [Google Scholar] [CrossRef] [PubMed]
- Kelly, R.W.; Holland, P.; Skibinski, G.; Harrison, C.; McMillan, L.; Hargreave, T.; James, K. Extracellular organelles (prostasomes) are immunosuppressive components of human semen. Clin. Exp. Immunol. 1991, 86, 550–556. [Google Scholar] [CrossRef] [PubMed]
- Bechoua, S.; Rieu, I.; Sion, B.; Grizard, G. Prostasomes as potential modulators of tyrosine phosphorylation in human spermatozoa. Syst. Biol. Reprod. Med. 2011, 57, 139–148. [Google Scholar] [CrossRef] [PubMed]
- Pons-Rejraji, H.; Artonne, C.; Sion, B.; Brugnon, F.; Canis, M.; Janny, L.; Grizard, G. Prostasomes: Inhibitors of capacitation and modulators of cellular signalling in human sperm. Int. J. Androl. 2011, 34, 568–580. [Google Scholar] [CrossRef] [PubMed]
- Saez, F.; Motta, C.; Boucher, D.; Grizard, G. Antioxidant capacity of prostasomes in human semen. Mol. Hum. Reprod. 1998, 4, 667–672. [Google Scholar] [CrossRef] [PubMed]
- Saez, F.; Motta, C.; Boucher, D.; Grizard, G. Prostasomes inhibit the NADPH oxidase activity of human neutrophils. Mol. Hum. Reprod. 2000, 6, 883–891. [Google Scholar] [CrossRef] [PubMed]
- Cross, N.L. Human seminal plasma prevents sperm from becoming acrosomally responsive to the agonist, progesterone: Cholesterol is the major inhibitor. Biol. Reprod. 1996, 54, 138–145. [Google Scholar] [CrossRef] [PubMed]
- Kelly, R.W.; Critchley, H.O. Immunomodulation by human seminal plasma: A benefit for spermatozoon and pathogen? Hum. Reprod. 1997, 12, 2200–2207. [Google Scholar] [CrossRef] [PubMed]
- Vojtech, L.; Hughes, S.; Levy, C.; Tewari, M.; Hladik, F. Exosomes in human semen impair antigen-presenting cell function and decrease antigen-specific T cell responses (MUC7P.767). J. Immunol. 2014, 192, 197.19. [Google Scholar]
- Konadu, K.A.; Chu, J.; Huang, M.B.; Amancha, P.K.; Armstrong, W.; Powell, M.D.; Villinger, F.; Bond, V.C. Association of Cytokines with Exosomes in the Plasma of HIV-1-Seropositive Individuals. J. Infect. Dis. 2014. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Ye, L.; Hou, W.; Zhou, Y.; Wang, Y.J.; Metzger, D.S.; Ho, W.Z. Cellular microRNA expression correlates with susceptibility of monocytes/macrophages to HIV-1 infection. Blood 2009, 113, 671–674. [Google Scholar] [CrossRef] [PubMed]
- Mantri, C.K.; Mantri, J.V.; Pandhare, J.; Dash, C. Methamphetamine inhibits HIV-1 replication in CD4+ T cells by modulating anti-HIV-1 miRNA expression. Am. J. Pathol. 2014, 184, 92–100. [Google Scholar] [CrossRef] [PubMed]
- Hariharan, M.; Scaria, V.; Pillai, B.; Brahmachari, S.K. Targets for human encoded microRNAs in HIV genes. Biochem. Biophys. Res. Commun. 2005, 337, 1214–1218. [Google Scholar] [CrossRef] [PubMed]
- Ahluwalia, J.K.; Khan, S.Z.; Soni, K.; Rawat, P.; Gupta, A.; Hariharan, M.; Scaria, V.; Lalwani, M.; Pillai, B.; Mitra, D.; et al. Human cellular microRNA hsa-miR-29a interferes with viral nef protein expression and HIV-1 replication. Retrovirology 2008, 5. [Google Scholar] [CrossRef] [PubMed]
- Mantri, C.K.; Pandhare Dash, J.; Mantri, J.V.; Dash, C.C. Cocaine enhances HIV-1 replication in CD4+ T cells by down-regulating miR-125b. PLoS ONE 2012, 7, e51387. [Google Scholar] [CrossRef] [PubMed]
- Sung, T.L.; Rice, A.P. miR-198 inhibits HIV-1 gene expression and replication in monocytes and its mechanism of action appears to involve repression of cyclin T1. PLoS Pathog. 2009, 5, e1000263. [Google Scholar] [CrossRef] [PubMed]
- Chevillet, J.R.; Kang, Q.; Ruf, I.K.; Briggs, H.A.; Vojtech, L.N.; Hughes, S.M.; Cheng, H.H.; Arroyo, J.D.; Meredith, E.K.; Gallichotte, E.N.; et al. Quantitative and stoichiometric analysis of the microRNA content of exosomes. Proc. Natl. Acad. Sci. USA 2014, 111, 14888–14893. [Google Scholar] [CrossRef] [PubMed]
- Mittelbrunn, M.; Gutierrez-Vazquez, C.; Villarroya-Beltri, C.; Gonzalez, S.; Sanchez-Cabo, F.; Gonzalez, M.A.; Bernad, A.; Sanchez-Madrid, F. Unidirectional transfer of microRNA-loaded exosomes from T cells to antigen-presenting cells. Nat. Commun. 2011, 2. [Google Scholar] [CrossRef] [PubMed]
- Conde-Vancells, J.; Rodriguez-Suarez, E.; Gonzalez, E.; Berisa, A.; Gil, D.; Embade, N.; Valle, M.; Luka, Z.; Elortza, F.; Wagner, C.; et al. Candidate biomarkers in exosome-like vesicles purified from rat and mouse urine samples. Proteomics 2010, 4, 416–425. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Madison, M.N.; Okeoma, C.M. Exosomes: Implications in HIV-1 Pathogenesis. Viruses 2015, 7, 4093-4118. https://doi.org/10.3390/v7072810
Madison MN, Okeoma CM. Exosomes: Implications in HIV-1 Pathogenesis. Viruses. 2015; 7(7):4093-4118. https://doi.org/10.3390/v7072810
Chicago/Turabian StyleMadison, Marisa N., and Chioma M. Okeoma. 2015. "Exosomes: Implications in HIV-1 Pathogenesis" Viruses 7, no. 7: 4093-4118. https://doi.org/10.3390/v7072810
APA StyleMadison, M. N., & Okeoma, C. M. (2015). Exosomes: Implications in HIV-1 Pathogenesis. Viruses, 7(7), 4093-4118. https://doi.org/10.3390/v7072810