The Roles of Syncytin-Like Proteins in Ruminant Placentation
Abstract
:1. Introduction
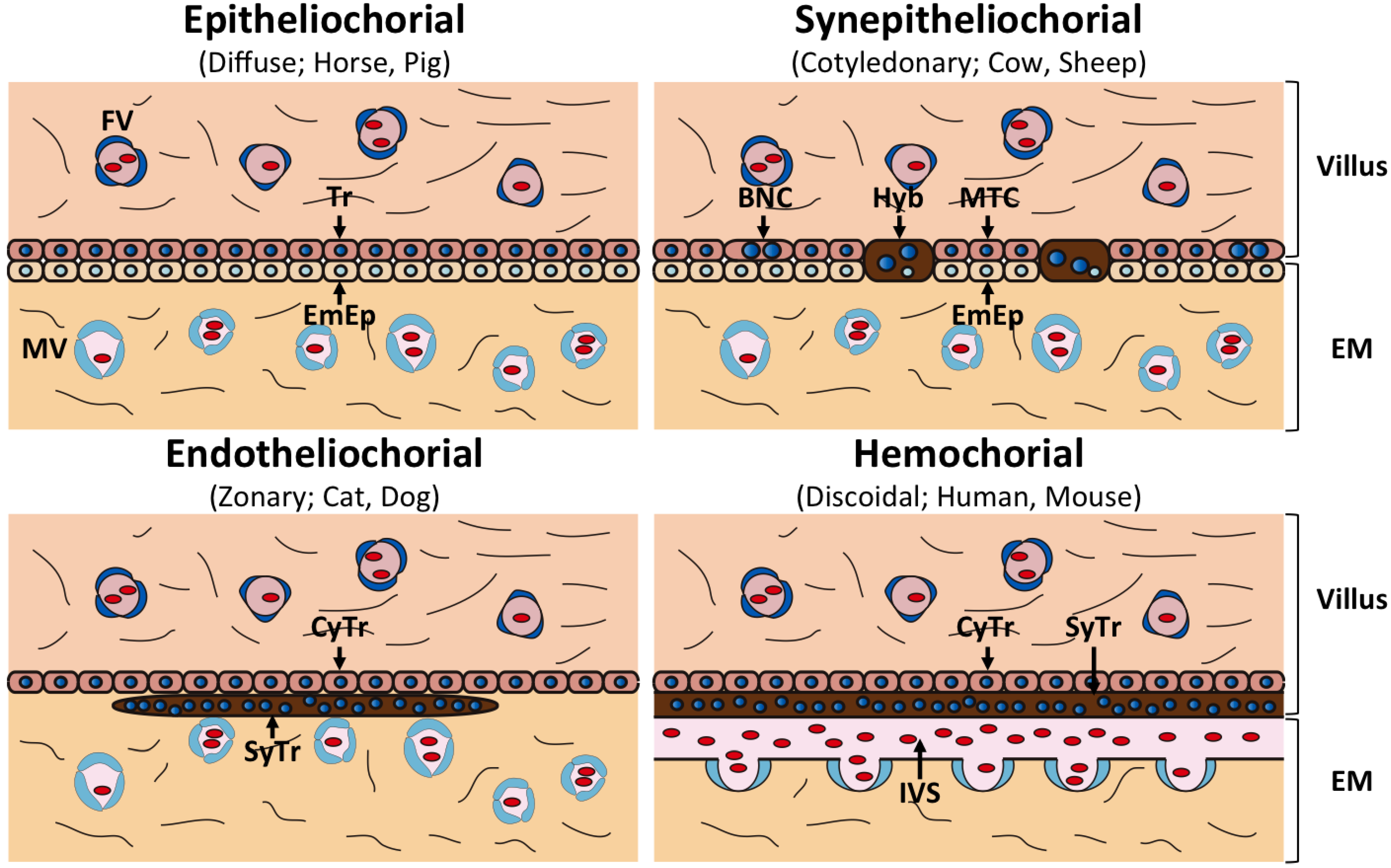
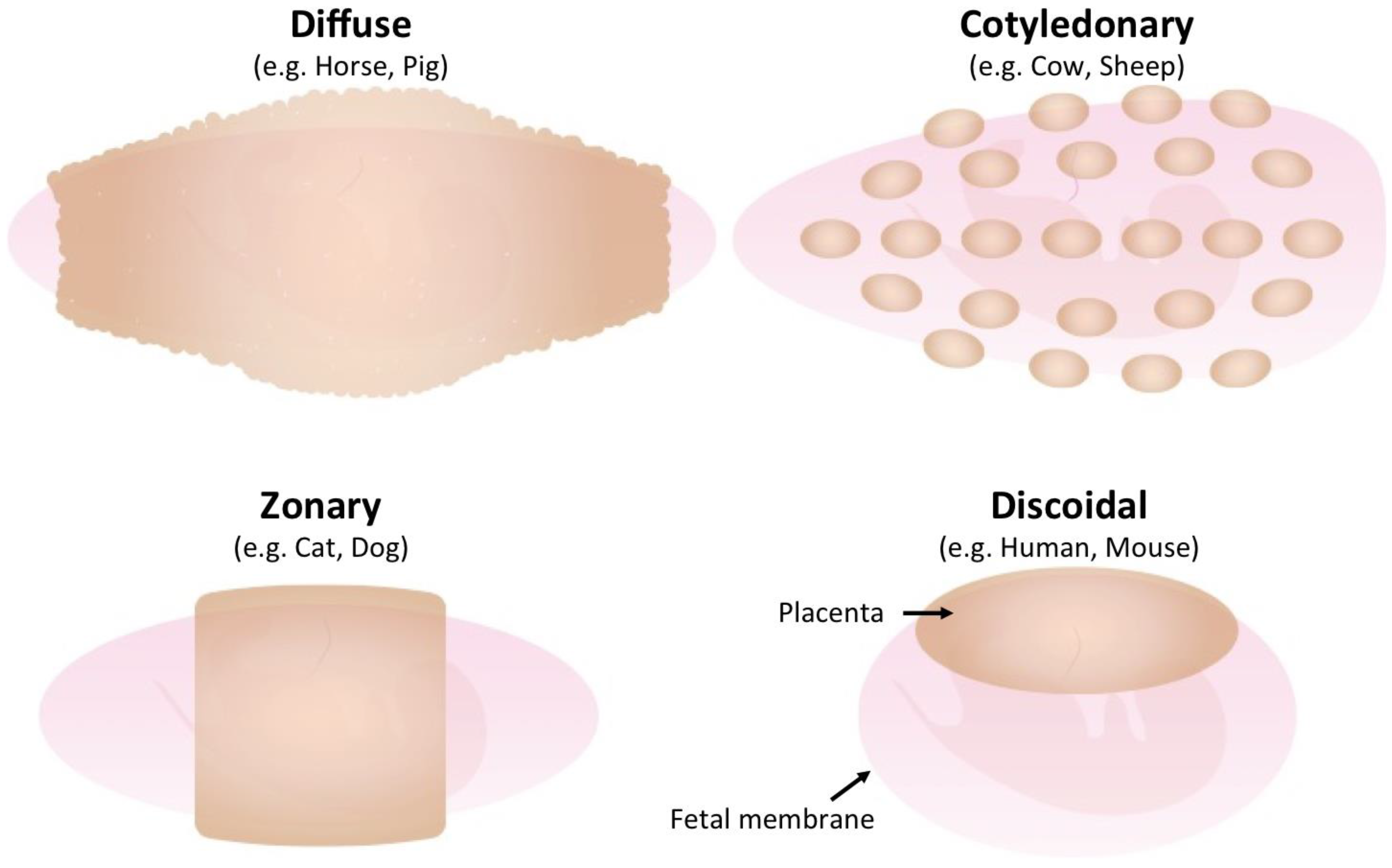
2. Diversity of Mammalian Placenta
3. ERVs Involved in Placentation
3.2. Discovery of Envs Derived from ERVs in Human Placenta
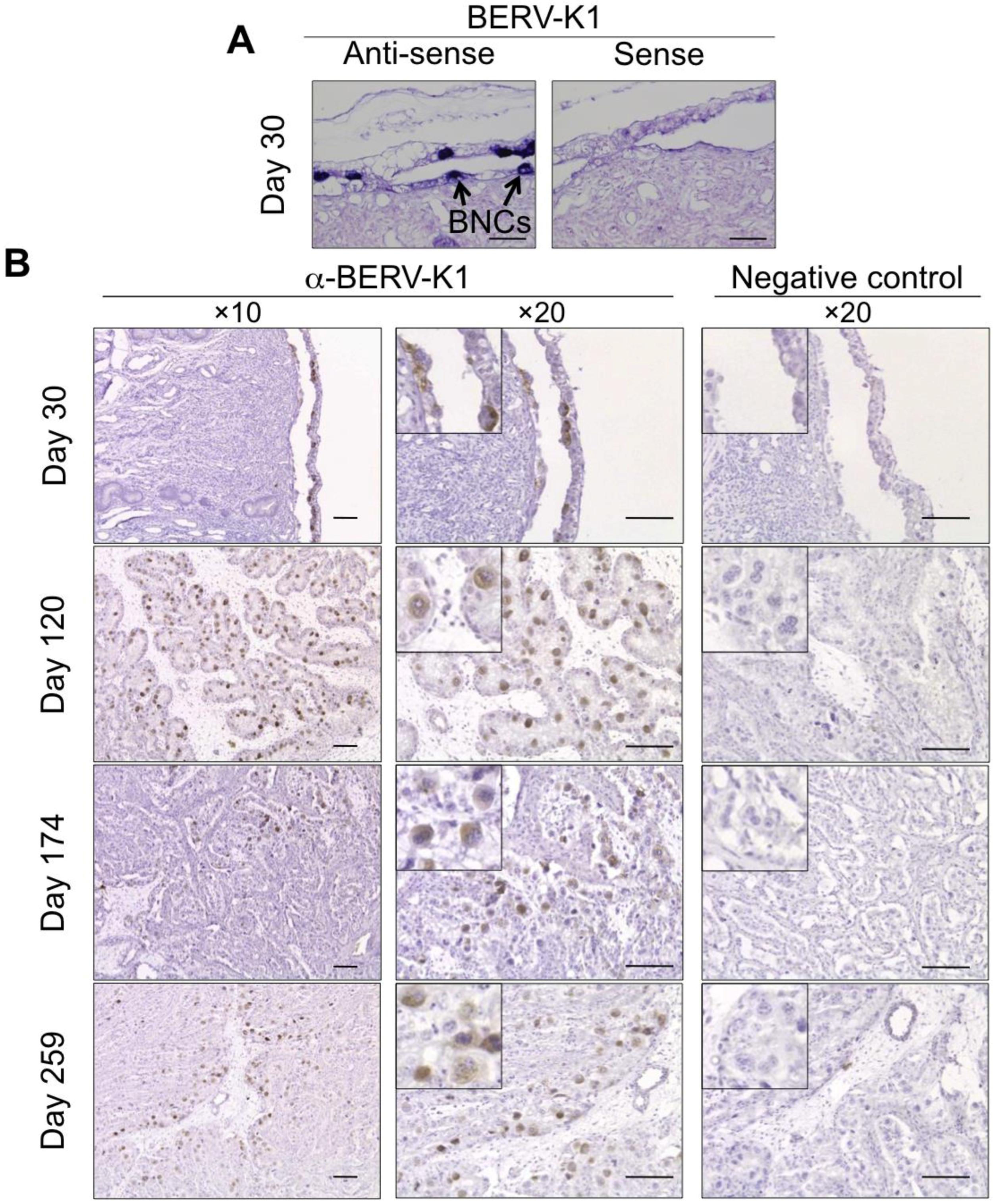
3.3. ERV Envs Expressed in Ruminant Placenta
| Name of ERV | Species | Expressed Tissues | Expressed Period | Appearance in Host Genome |
|---|---|---|---|---|
| Syncytin-Rum1 | Most ruminants (except for Tragulidae) | Placenta (BNCs) | Through gestation | Over 30 MYA |
| BERV-K1 (Fematrin-1) | Subfamily Bovinae | Placenta (BNCs) | Through gestation | 25.4 to 18.3 MYA |
| bERVE-A | Bos taurus | Placenta (BNCs) | Through gestation | Not examined |
| BERV-P | Genus Bos | Trophectoderm | Periimplantation | 16.9 to 7 MYA |
| enJSRV | Subfamily Caprinae | Placenta (BNCs), Endometrium | Through gestation | Within 7 MYA |
4. Perspective on the Relationship between Fematrin-1 and Syncytin-Rum1
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gifford, R.; Tristem, M. The evolution, distribution and diversity of endogenous retroviruses. Virus Genes 2003, 26, 291–315. [Google Scholar] [CrossRef] [PubMed]
- Stoye, J.P. Studies of endogenous retroviruses reveal a continuing evolutionary saga. Nat. Rev. Microbiol. 2012, 10, 395–406. [Google Scholar] [CrossRef] [PubMed]
- Tarlinton, R.E.; Meers, J.; Young, P.R. Retroviral invasion of the koala genome. Nature 2006, 442, 79–81. [Google Scholar] [CrossRef] [PubMed]
- Mi, S.; Lee, X.; Li, X.; Veldman, G.M.; Finnerty, H.; Racie, L.; LaVallie, E.; Tang, X.Y.; Edouard, P.; Howes, S.; et al. Syncytin is a captive retroviral envelope protein involved in human placental morphogenesis. Nature 2000, 403, 785–789. [Google Scholar] [PubMed]
- Blaise, S.; de Parseval, N.; Bénit, L.; Heidmann, T. Genomewide screening for fusogenic human endogenous retrovirus envelopes identifies syncytin 2, a gene conserved on primate evolution. Proc. Natl. Acad. Sci. USA 2003, 100, 13013–13018. [Google Scholar] [CrossRef] [PubMed]
- Dupressoir, A.; Marceau, G.; Vernochet, C.; Bénit, L.; Kanellopoulos, C.; Sapin, V.; Heidmann, T. Syncytin-A and syncytin-B, two fusogenic placenta-specific murine envelope genes of retroviral origin conserved in Muridae. Proc. Natl. Acad. Sci. USA 2005, 102, 725–730. [Google Scholar] [CrossRef] [PubMed]
- Heidmann, O.; Vernochet, C.; Dupressoir, A.; Heidmann, T. Identification of an endogenous retroviral envelope gene with fusogenic activity and placenta-specific expression in the rabbit: A new “syncytin” in a third order of mammals. Retrovirology 2009, 6, e107. [Google Scholar] [CrossRef] [PubMed]
- Cornelis, G.; Heidmann, O.; Bernard-Stoecklin, S.; Reynaud, K.; Véron, G.; Mulot, B.; Dupressoir, A.; Heidmann, T. Ancestral capture of syncytin-Car1, a fusogenic endogenous retroviral envelope gene involved in placentation and conserved in Carnivora. Proc. Natl. Acad. Sci. USA 2012, 109, E432–E441. [Google Scholar] [CrossRef] [PubMed]
- Dunlap, K.A.; Palmarini, M.; Varela, M.; Burghardt, R.C.; Hayashi, K.; Farmer, J.L.; Spencer, T.E. Endogenous retroviruses regulate periimplantation placental growth and differentiation. Proc. Natl. Acad. Sci. USA 2006, 103, 14390–14395. [Google Scholar] [CrossRef] [PubMed]
- Baba, K.; Nakaya, Y.; Shojima, T.; Muroi, Y.; Kizaki, K.; Hashizume, K.; Imakawa, K.; Miyazawa, T. Identification of novel endogenous betaretroviruses which are transcribed in the bovine placenta. J. Virol. 2011, 85, 1237–1245. [Google Scholar] [CrossRef] [PubMed]
- Nakaya, Y.; Koshi, K.; Nakagawa, S.; Hashizume, K.; Miyazawa, T. Fematrin-1 is involved in fetomaternal cell-to-cell fusion in Bovinae placenta and has contributed to diversity of ruminant placentation. J. Virol. 2013, 87, 10563–10572. [Google Scholar] [CrossRef] [PubMed]
- Cornelis, G.; Heidmann, O.; Degrelle, S.A.; Vernochet, C.; Lavialle, C.; Letzelter, C.; Bernard-Stoecklin, S.; Hassanin, A.; Mulot, B.; Guillomot, M.; et al. Captured retroviral envelope syncytin gene associated with the unique placental structure of higher ruminants. Proc. Natl. Acad. Sci. USA 2013, 110, E828–E837. [Google Scholar] [CrossRef] [PubMed]
- Wildman, D.E.; Chen, C.; Erez, O.; Grossman, L.I.; Goodman, M.; Romero, R. Evolution of the mammalian placenta revealed by phylogenetic analysis. Proc. Natl. Acad. Sci. USA 2006, 103, 3203–3208. [Google Scholar] [CrossRef] [PubMed]
- Yamada, O.; Todoroki, J.; Kizaki, K.; Takahashi, T.; Imai, K.; Patel, O.V.; Schuler, L.A.; Hashizume, K. Expression of prolactin-related protein I at the fetomaternal interface during the implantation period in cows. Reproduction 2002, 124, 427–437. [Google Scholar] [CrossRef] [PubMed]
- Yamada, O.; Todoroki, J.; Takahashi, T.; Hashizume, K. The dynamic expression of extracellular matrix in the bovine endometrium at implantation. J. Vet. Med. Sci. 2002, 64, 207–214. [Google Scholar] [CrossRef] [PubMed]
- Kizaki, K.; Ushizawa, K.; Takahashi, T.; Yamada, O.; Todoroki, J.; Sato, T.; Ito, A.; Hashizume, K. Gelatinase (MMP-2 and -9) expression profiles during gestation in the bovine endometrium. Reprod. Biol. Endocrinol. 2008, 6, e66. [Google Scholar] [CrossRef] [PubMed]
- Klisch, K.; Pfarrer, C.; Schuler, G.; Hoffmann, B.; Leiser, R. Tripolar acytokinetic mitosis and formation of feto-maternal syncytia in the bovine placentome: Different modes of the generation of multinuclear cells. Anat. Embryol. 1999, 200, 229–237. [Google Scholar] [CrossRef] [PubMed]
- Klisch, K.; Hecht, W.; Pfarrer, C.; Schuler, G.; Hoffmann, B.; Leiser, R. DNA content and ploidy level of bovine placentomal trophoblast giant cells. Placenta 1999, 20, 451–458. [Google Scholar] [CrossRef] [PubMed]
- Hallenberger, S.; Bosch, V.; Angliker, H.; Shaw, E.; Klenk, H.D.; Garten, W. Inhibition of furin-mediated cleavage activation of HIV-1 glycoprotein gp160. Nature 1992, 360, 358–361. [Google Scholar] [CrossRef] [PubMed]
- Schultz, A.; Rein, A. Different recombinant murine leukemia viruses use different cell surface receptors. Virology 1985, 145, 335–339. [Google Scholar] [CrossRef]
- Chang, C.; Chen, P.T.; Chang, G.D.; Huang, C.J.; Chen, H. Functional characterization of the placental fusogenic membrane protein syncytin. Biol. Reprod. 2004, 71, 1956–1962. [Google Scholar]
- Chen, C.P.; Chen, L.F.; Yang, S.R.; Chen, C.Y.; Ko, C.C.; Chang, G.D.; Chen, H. Functional characterization of the human placental fusogenic membrane protein syncytin 2. Biol. Reprod. 2008, 79, 815–823. [Google Scholar] [CrossRef] [PubMed]
- Wallin, M.; Ekström, M.; Garoff, H. Isomerization of the intersubunit disulphide-bond in ENV controls retrovirus fusion. EMBO J. 2004, 23, 54–65. [Google Scholar] [CrossRef] [PubMed]
- Henzy, J.E.; Coffin, J.M. Betaretroviral Envelope Subunits Are Noncovalently Associated and Restricted to the Mammalian Class. J. Virol. 2013, 87, 1937–1946. [Google Scholar] [CrossRef] [PubMed]
- Cheynet, V.; Oriol, G.; Mallet, F. Identification of the hASCT2-binding domain of the ENV ERVWE1/syncytin-1 fusogenic glycoprotein. Retrovirology 2006, 3, e41. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, R.; Miyazawa, T.; Matsuura, Y. Cell-binding properties of the envelope proteins of porcine endogenous retroviruses. Microbes Infect. 2005, 7, 658–665. [Google Scholar] [CrossRef] [PubMed]
- Argaw, T.; Figueroa, M.; Salomon, D.R.; Wilson, C.A. Identification of residues outside of the receptor binding domain that influence the infectivity and tropism of porcine endogenous retrovirus. J. Virol. 2008, 82, 7483–7491. [Google Scholar] [CrossRef] [PubMed]
- Kim, F.J.; Manel, N.; Garrido, E.N.; Valle, C.; Sitbon, M.; Battini, J.L. HTLV-1 and -2 envelope SU subdomains and critical determinants in receptor binding. Retrovirology 2004, 1, e41. [Google Scholar] [CrossRef] [PubMed]
- Jones, K.S.; Lambert, S.; Bouttier, M.; Bénit, L.; Ruscetti, F.W.; Hermine, O.; Pique, C. Molecular aspects of HTLV-1 entry: Functional domains of the HTLV-1 surface subunit (SU) and their relationships to the entry receptors. Viruses 2011, 3, 794–810. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Rassa, J.C.; deObaldia, M.E.; Albritton, L.M.; Ross, S.R. Identification of the receptor binding domain of the mouse mammary tumor virus envelope protein. J. Virol. 2003, 77, 10468–10478. [Google Scholar] [CrossRef] [PubMed]
- Checkley, M.A.; Luttge, B.G.; Freed, E.O. HIV-1 envelope glycoprotein biosynthesis, trafficking, and incorporation. J. Mol. Biol. 2011, 410, 582–608. [Google Scholar] [CrossRef] [PubMed]
- Bobkova, M.; Stitz, J.; Engelstädter, M.; Cichutek, K.; Buchholz, C.J. Identification of R-peptides in envelope proteins of C-type retroviruses. J. Gen. Virol. 2002, 83, 2241–2246. [Google Scholar] [PubMed]
- Mangeney, M.; Renard, M.; Schlecht-Louf, G.; Bouallaga, I.; Heidmann, O.; Letzelter, C.; Richaud, A.; Ducos, B.; Heidmann, T. Placental syncytins: Genetic disjunction between the fusogenic and immunosuppressive activity of retroviral envelope proteins. Proc. Natl. Acad. Sci. USA 2007, 104, 20534–20539. [Google Scholar] [CrossRef] [PubMed]
- Blond, J.L.; Lavillette, D.; Cheynet, V.; Bouton, O.; Oriol, G.; Chapel-Fernandes, S.; Mandrand, B.; Mallet, F.; Cosset, F.L. An envelope glycoprotein of the human endogenous retrovirus HERV-W is expressed in the human placenta and fuses cells expressing the type D mammalian retrovirus receptor. J. Virol. 2000, 74, 3321–3329. [Google Scholar] [CrossRef] [PubMed]
- Lyden, T.W.; Johnson, P.M.; Mwenda, J.M.; Rote, N.S. Expression of endogenous HIV-1 crossreactive antigens within normal human extravillous trophoblast cells. J. Reprod. Immunol. 1995, 28, 233–245. [Google Scholar] [CrossRef]
- Lin, L.; Xu, B.; Rote, N.S. Expression of endogenous retrovirus ERV-3 induces differentiation in BeWo, a choriocarcinoma model of human placental trophoblast. Placenta 1999, 20, 109–118. [Google Scholar] [CrossRef] [PubMed]
- Boyd, M.T.; Bax, C.M.; Bax, B.E.; Bloxam, D.L.; Weiss, R.A. The human endogenous retrovirus ERV-3 is upregulated in differentiating placental trophoblast cells. Virology 1993, 196, 905–909. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Xu, B.; Rote, N.S. The cellular mechanism by which the human endogenous retrovirus ERV-3 env gene affects proliferation and differentiation in a human placental trophoblast model, BeWo. Placenta 2000, 21, 73–78. [Google Scholar] [CrossRef] [PubMed]
- Langat, D.K.; Johnson, P.M.; Rote, N.S.; Wango, E.O.; Owiti, G.O.; Isahakia, M.A.; Mwenda, J.M. Characterization of antigens expressed in normal baboon trophoblast and cross-reactive with HIV/SIV antibodies. J. Reprod. Immunol. 1999, 42, 41–58. [Google Scholar] [CrossRef]
- Cáceres, M.; Thomas, J.W. The gene of retroviral origin Syncytin 1 is specific to hominoids and is inactive in Old World monkeys. J. Hered. 2006, 97, 100–106. [Google Scholar] [CrossRef] [PubMed]
- Esnault, C.; Cornelis, G.; Heidmann, O.; Heidmann, T. Differential evolutionary fate of an ancestral primate endogenous retrovirus envelope gene, the EnvV syncytin, captured for a function in placentation. PLoS Genet. 2013, 9, e1003400. [Google Scholar] [CrossRef] [PubMed]
- Lee, X.; Keith, J.C., Jr.; Stumm, N.; Moutsatsos, I.; McCoy, J.M.; Crum, C.P.; Genest, D.; Chin, D.; Ehrenfels, C.; Pijnenborg, R.; et al. Downregulation of placental syncytin expression and abnormal protein localization in pre-eclampsia. Placenta 2001, 22, 808–812. [Google Scholar] [CrossRef] [PubMed]
- Malassiné, A.; Blaise, S.; Handschuh, K.; Lalucque, H.; Dupressoir, A.; Evain-Brion, D.; Heidmann, T. Expression of the fusogenic HERV-FRD Env glycoprotein (syncytin 2) in human placenta is restricted to villous cytotrophoblastic cells. Placenta 2007, 28, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Hayward, M.D.; Pötgens, A.J.; Drewlo, S.; Kaufmann, P.; Rasko, J.E. Distribution of human endogenous retrovirus type W receptor in normal human villous placenta. Pathology 2007, 39, 406–412. [Google Scholar] [CrossRef] [PubMed]
- Esnault, C.; Priet, S.; Ribet, D.; Vernochet, C.; Bruls, T.; Lavialle, C.; Weissenbach, J.; Heidmann, T. A placenta-specific receptor for the fusogenic, endogenous retrovirus-derived, human syncytin-2. Proc. Natl. Acad. Sci. USA 2008, 105, 17532–17537. [Google Scholar] [CrossRef] [PubMed]
- Cianciolo, G.J.; Copeland, T.D.; Oroszlan, S.; Snyderman, R. Inhibition of lymphocyte proliferation by a synthetic peptide homologous to retroviral envelope proteins. Science 1985, 230, 453–455. [Google Scholar] [CrossRef] [PubMed]
- Sonigo, P.; Barker, C.; Hunter, E.; Wain-Hobson, S. Nucleotide sequence of Mason-Pfizer monkey virus: An immunosuppressive D-type retrovirus. Cell 1986, 45, 375–385. [Google Scholar] [CrossRef]
- Tolosa, J.M.; Parsons, K.S.; Hansbro, P.M.; Smith, R.; Wark, P.A. The placental protein syncytin-1 impairs antiviral responses and exaggerates inflammatory responses to influenza. PLoS ONE 2015, 10, e0118629. [Google Scholar] [CrossRef] [PubMed]
- Tolosa, J.M.; Schjenken, J.E.; Clifton, V.L.; Vargas, A.; Barbeau, B.; Lowry, P.; Maiti, K.; Smith, R. The endogenous retroviral envelope protein syncytin-1 inhibits LPS/PHA-stimulated cytokine responses in human blood and is sorted into placental exosomes. Placenta 2012, 33, 933–941. [Google Scholar] [CrossRef] [PubMed]
- Lavialle, C.; Cornelis, G.; Dupressoir, A.; Esnault, C.; Heidmann, O.; Vernochet, C.; Heidmann, T. Paleovirology of “syncytins”, retroviral ENV genes exapted for a role in placentation. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2013, 368, 20120507. [Google Scholar] [CrossRef] [PubMed]
- Nakaya, Y.; Kizaki, K.; Takahashi, T.; Patel, O.V.; Hashizume, K. The characterization of DNA methylation-mediated regulation of bovine placental lactogen and bovine prolactin-related protein-1 genes. BMC Mol. Biol. 2009, 10, e19. [Google Scholar] [CrossRef] [PubMed]
- Su, J.; Wang, Y.; Xing, X.; Liu, J.; Zhang, Y. Genome-wide analysis of DNA methylation in bovine placentas. BMC Genomics 2014, 15, e12. [Google Scholar] [CrossRef] [PubMed]
- Klisch, K.; Mess, A. Evolutionary differentiation of cetartiodactyl placentae in the light of the viviparity-driven conflict hypothesis. Placenta 2007, 28, 353–360. [Google Scholar] [CrossRef] [PubMed]
- Wooding, F.B.; Hobbs, T.; Morgan, G.; Heap, R.B.; Flint, A.P. Cellular dynamics of growth in sheep and goat synepitheliochorial placentomes: An autoradiographic study. J. Reprod. Fertil. 1993, 98, 275–283. [Google Scholar] [CrossRef] [PubMed]
- Koshi, K.; Suzuki, Y.; Nakaya, Y.; Imai, K.; Hosoe, M.; Takahashi, T.; Kizaki, K.; Miyazawa, T.; Hashizume, K. Bovine trophoblastic cell differentiation and binucleation involves enhanced endogenous retrovirus element expression. Reprod. Biol. Endocrinol. 2012, 10, e41. [Google Scholar] [CrossRef] [PubMed]
- York, D.F.; Vigne, R.; Verwoerd, D.W.; Querat, G. Nucleotide sequence of the jaagsiekte retrovirus, an exogenous and endogenous type D and B retrovirus of sheep and goats. J. Virol. 1992, 66, 4930–4939. [Google Scholar] [PubMed]
- Carlson, J.; Lyon, M.; Bishop, J.; Vaiman, A.; Cribiu, E.; Mornex, J.F.; Brown, S.; Knudson, D.; DeMartini, J.; Leroux, C. Chromosomal distribution of endogenous Jaagsiekte sheep retrovirus proviral sequences in the sheep genome. J. Virol. 2003, 77, 9662–9668. [Google Scholar] [CrossRef] [PubMed]
- Arnaud, F.; Caporale, M.; Varela, M.; Biek, R.; Chessa, B.; Alberti, A.; Golder, M.; Mura, M.; Zhang, Y.P.; Yu, L.; et al. A paradigm for virus-host coevolution: Sequential counter-adaptations between endogenous and exogenous retroviruses. PLoS Pathog. 2007, 3, e170. [Google Scholar] [CrossRef] [PubMed]
- Mura, M.; Murcia, P.; Caporale, M.; Spencer, T.E.; Nagashima, K.; Rein, A.; Palmarini, M. Late viral interference induced by transdominant Gag of an endogenous retrovirus. Proc. Natl. Acad. Sci. USA 2004, 101, 11117–11122. [Google Scholar] [CrossRef] [PubMed]
- Murcia, P.R.; Arnaud, F.; Palmarini, M. The transdominant endogenous retrovirus enJS56A1 associates with and blocks intracellular trafficking of Jaagsiekte sheep retrovirus Gag. J. Virol. 2007, 81, 1762–1772. [Google Scholar] [CrossRef] [PubMed]
- Arnaud, F.; Murcia, P.R.; Palmarini, M. Mechanisms of late restriction induced by an endogenous retrovirus. J. Virol. 2007, 81, 11441–11451. [Google Scholar] [CrossRef] [PubMed]
- Dunlap, K.A.; Palmarini, M.; Adelson, D.L.; Spencer, T.E. Sheep endogenous betaretroviruses (enJSRVs) and the hyaluronidase 2 (HYAL2) receptor in the ovine uterus and conceptus. Biol. Reprod. 2005, 73, 271–279. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, S.; Bai, H.; Sakurai, T.; Nakaya, Y.; Konno, T.; Miyazawa, T.; Gojobori, T.; Imakawa, K. Dynamic evolution of endogenous retrovirus-derived genes expressed in bovine conceptuses during the period of placentation. Genome. Biol. Evol. 2013, 5, 296–306. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Etxebarria, K.; Jugo, B.M. Genome-wide detection and characterization of endogenous retroviruses in Bos taurus. J. Virol. 2010, 84, 10852–10862. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Etxebarria, K.; Jugo, B.M. Evolutionary history of bovine endogenous retroviruses in the Bovidae family. BMC Evol. Biol. 2013, 13, e256. [Google Scholar] [CrossRef] [PubMed]
- Xiao, R.; Park, K.; Lee, H.; Kim, J.; Park, C. Identification and classification of endogenous retroviruses in cattle. J. Virol. 2008, 82, 582–587. [Google Scholar] [CrossRef] [PubMed]
- Nakaya, Y.; Miyazawa, T. Dysfunction of bovine endogenous retrovirus K2 envelope glycoprotein is related to unsuccessful intracellular trafficking. J. Virol. 2014, 88, 6896–6905. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, Y.; Imakawa, K. Retroviral endogenization and its role in the genital tract during mammalian evolution. J. Mamm. Ova Res. 2011, 28, 203–218. [Google Scholar] [CrossRef]
- Wooding, F.B.; Kimura, J.; Fukuta, K.; Forhead, A.J. A light and electron microscopical study of the Tragulid (mouse deer) placenta. Placenta 2007, 28, 1039–1048. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nakaya, Y.; Miyazawa, T. The Roles of Syncytin-Like Proteins in Ruminant Placentation. Viruses 2015, 7, 2928-2942. https://doi.org/10.3390/v7062753
Nakaya Y, Miyazawa T. The Roles of Syncytin-Like Proteins in Ruminant Placentation. Viruses. 2015; 7(6):2928-2942. https://doi.org/10.3390/v7062753
Chicago/Turabian StyleNakaya, Yuki, and Takayuki Miyazawa. 2015. "The Roles of Syncytin-Like Proteins in Ruminant Placentation" Viruses 7, no. 6: 2928-2942. https://doi.org/10.3390/v7062753
APA StyleNakaya, Y., & Miyazawa, T. (2015). The Roles of Syncytin-Like Proteins in Ruminant Placentation. Viruses, 7(6), 2928-2942. https://doi.org/10.3390/v7062753