Immune Memory to Sudan Virus: Comparison between Two Separate Disease Outbreaks
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Ethics Statement
2.3. Sample Collection
2.4. Antigens and Stimulants
2.5. Internal Control Sera
2.6. Specific IgG Detection Assays
2.7. Normalization of Raw Data and Selection of Cut-off Values
2.8. Plaque Reduction Neutralization Test
2.9. Whole Blood Stimulation from Ebolavirus Survivors and Healthy Volunteers
2.10. Cytokine Detection Using Q-Plex™ ELISA-Based Chemiluminescent Assay
2.11. Statistical Analysis
3. Results
3.1. Cohorts and Blood Samples
3.2. Humoral Response Profiles against SUDV Proteins
| Serology | PRNT50 | ||||||
|---|---|---|---|---|---|---|---|
| VP30 | VP35 | VP40 | NP | GP1–649 1 | SUDV Whole Ag | SUDV | |
| S-1 | +++ | - | ++ | +++ | +++ | +++ | +++ |
| S-2 | ++ | - | +++ | +++ | +++ | +++ | +++ |
| S-3 | - | + | ++ | ++ | ++ | ++ | ++ |
| S-4 | - | - | +++ | - | - | - | + |
| S-5 | ++ | + | ++ | +++ | +++ | +++ | +++ |
| N-1 | - | - | - | - | - | - | - |
| N-2 | - | - | - | - | - | - | - |
| N-3 | - | - | - | - | - | - | - |
| N-4 | - | - | - | - | - | - | - |
| N-5 | - | - | - | - | - | - | - |
| Serology | PRNT50 | ||||||
|---|---|---|---|---|---|---|---|
| VP30 | VP35 | VP40 | NP | GP1–649 1 | SUDV Whole Ag | SUDV | |
| S-1 | ++ | - | + | +++ | +++ | +++ | +++ |
| S-2 | + | - | + | +++ | +++ | +++ | + |
| S-3 | + | + | - | +++ | +++ | ++ | +++ |
| S-4 | +++ | - | - | + | +++ | +++ | +++ |
| S-5 | ++ | - | + | +++ | +++ | ++ | - |
| S-6 | +++ | - | +++ | +++ | +++ | +++ | ++ |
| N-1 | - | - | - | - | - | - | - |
| N-2 | - | - | - | - | - | - | - |
| N-3 | - | - | - | - | - | - | - |
| N-4 | - | - | - | - | - | - | - |
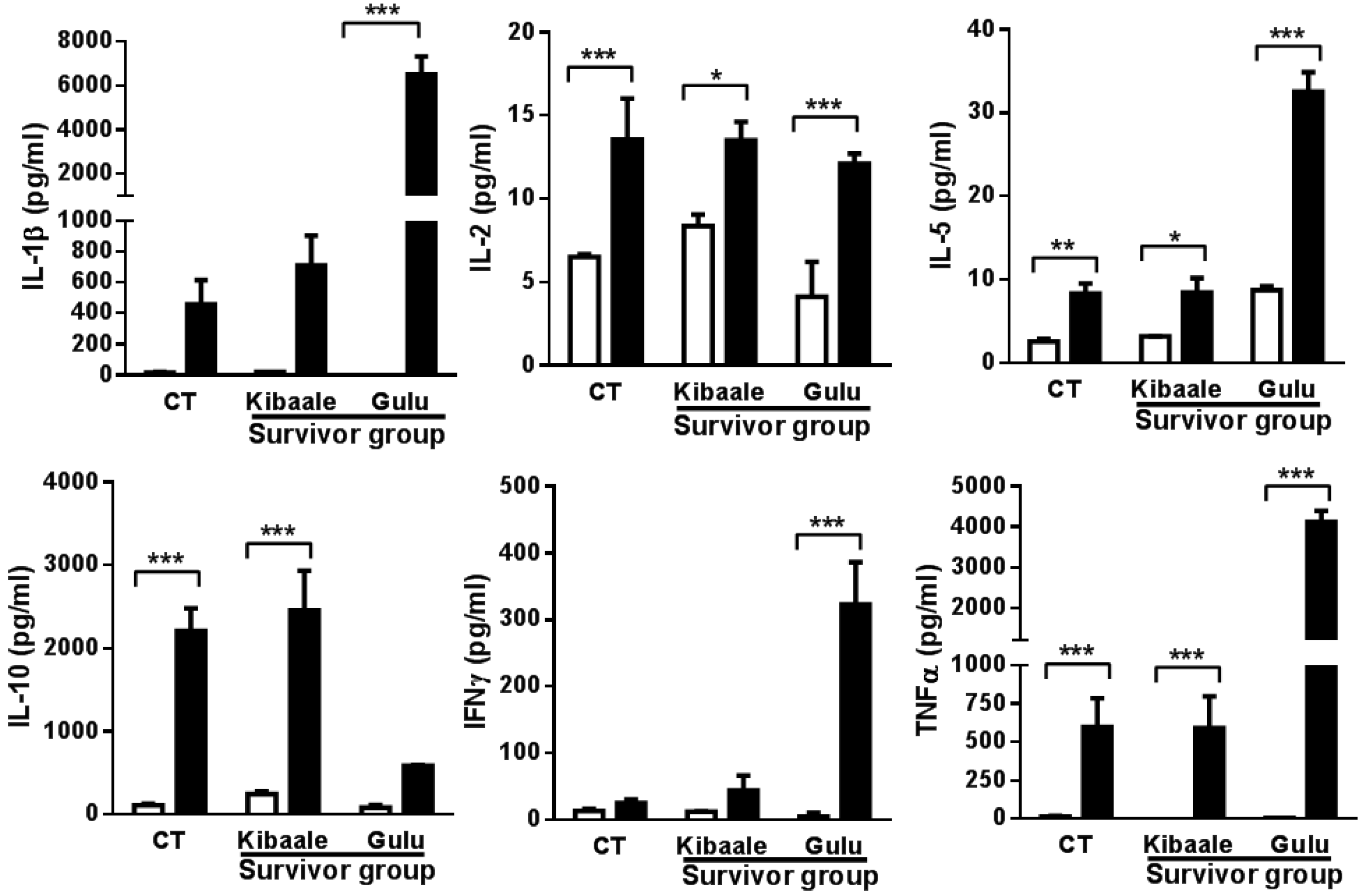
3.3. SUDV-Induced Cytokine Levels in Whole Blood Stimulation Assay

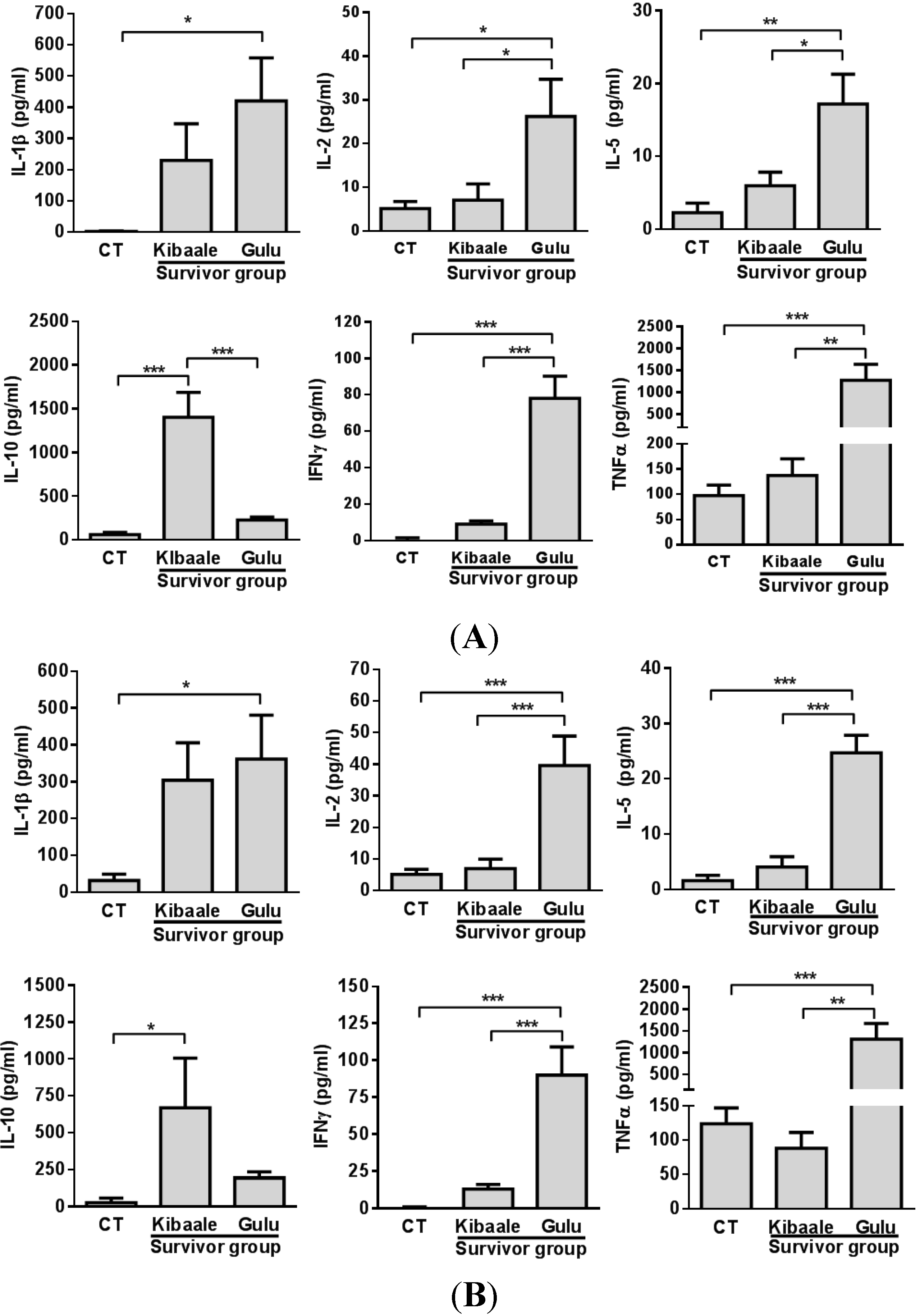
3.4. Humoral, Cell-Mediated and Neutralization Cross Reactivity to EBOV GP1–649 in SUDV Survivors
| Kibaale Group | Gulu Group | ||||||
|---|---|---|---|---|---|---|---|
| EBOV Whole Ag | EBOV GP1–649 | PRNT50 | EBOV Whole Ag | EBOV GP1–649 | PRNT50 | ||
| S-1 | - | + | - | S-1 | - | +++ | - |
| S-2 | +++ | +++ | - | S-2 | - | - | - |
| S-3 | - | - | - | S-3 | - | + | - |
| S-4 | - | - | - | S-4 | - | - | - |
| S-5 | + | ++ | - | S-5 | - | - | - |
| N-1 | - | - | - | S-6 | ++ | ++ | - |
| N-2 | - | - | - | N-1 | - | - | - |
| N-3 | - | - | - | N-2 | - | - | - |
| N-4 | - | - | - | N-3 | - | - | - |
| N-5 | - | - | - | N-4 | - | - | - |
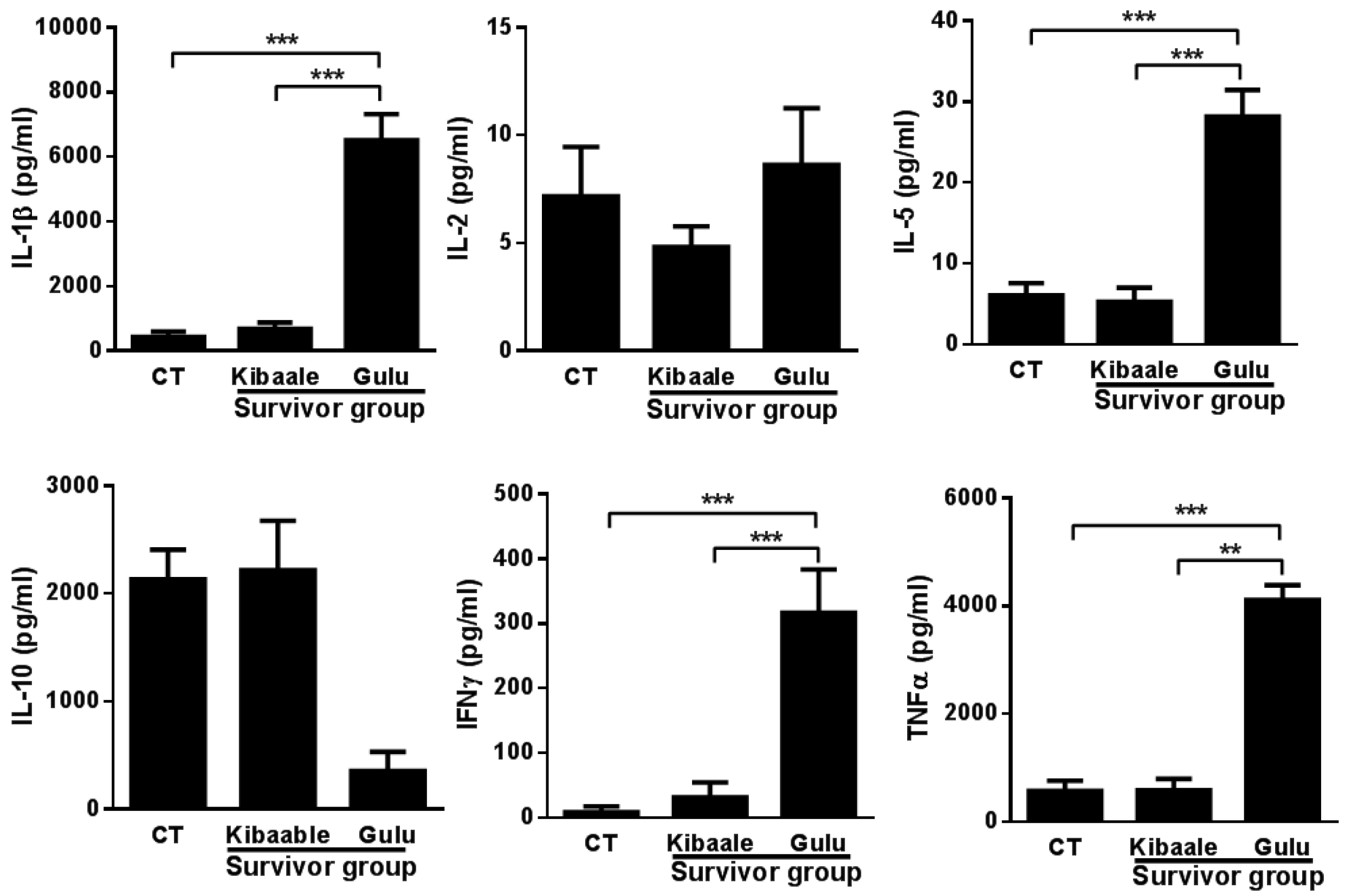
4. Discussion
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kuhn, J.H. Filoviruses. A compendium of 40 years of epidemiological, clinical, and laboratory studies. Arch. Virol. Suppl. 2008, 20, 13–360. [Google Scholar] [PubMed]
- Ascenzi, P.; Bocedi, A.; Heptonstall, J.; Capobianchi, M.R.; di Caro, A.; Mastrangelo, E.; Bolognesi, M.; Ippolito, G. Ebolavirus and marburgvirus: Insight the filoviridae family. Mol. Aspects Med. 2008, 29, 151–185. [Google Scholar] [CrossRef] [PubMed]
- Feldmann, H.; Geisbert, T.W. Ebola haemorrhagic fever. Lancet 2011, 377, 849–862. [Google Scholar] [CrossRef] [PubMed]
- Dolnik, O.; Kolesnikova, L.; Becker, S. Filoviruses: Interactions with the host cell. Cell. Mol. Life Sci. 2008, 65, 756–776. [Google Scholar] [CrossRef] [PubMed]
- Albarino, C.G.; Shoemaker, T.; Khristova, M.L.; Wamala, J.F.; Muyembe, J.J.; Balinandi, S.; Tumusiime, A.; Campbell, S.; Cannon, D.; Gibbons, A.; et al. Genomic analysis of filoviruses associated with four viral hemorrhagic fever outbreaks in uganda and the democratic republic of the congo in 2012. Virology 2013, 442, 97–100. [Google Scholar] [CrossRef] [PubMed]
- Anonymous. Outbreak of Ebola Hemorrhagic Fever—Uganda, August 2000–January 2001. Can. Commun. Dis. Rep. 2001, 27, 49–53. [Google Scholar]
- Mohamadzadeh, M.; Chen, L.; Schmaljohn, A.L. How ebola and marburg viruses battle the immune system. Nat. Rev. Immunol. 2007, 7, 556–567. [Google Scholar] [CrossRef] [PubMed]
- Kondratowicz, A.S.; Maury, W.J. Ebolavirus: A brief review of novel therapeutic targets. Future Microbiol. 2012, 7, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Gupta, M.; Goldsmith, C.S.; Metcalfe, M.G.; Spiropoulou, C.F.; Rollin, P.E. Reduced virus replication, proinflammatory cytokine production, and delayed macrophage cell death in human pbmcs infected with the newly discovered bundibugyo ebolavirus relative to zaire ebolavirus. Virology 2010, 402, 203–208. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, A.; Lukwiya, M.; Bausch, D.; Mahanty, S.; Sanchez, A.J.; Wagoner, K.D.; Rollin, P.E. Analysis of human peripheral blood samples from fatal and nonfatal cases of Ebola (Sudan) hemorrhagic fever: Cellular responses, virus load, and nitric oxide levels. J. Virol. 2004, 78, 10370–10377. [Google Scholar] [CrossRef] [PubMed]
- Wauquier, N.; Becquart, P.; Gasquet, C.; Leroy, E.M. Immunoglobulin G in Ebola outbreak survivors, Gabon. Emerg. Infect. Dis. 2009, 15, 1136–1137. [Google Scholar] [CrossRef] [PubMed]
- Sobarzo, A.; Ochayon, D.E.; Lutwama, J.J.; Balinandi, S.; Guttman, O.; Marks, R.S.; Kuehne, A.I.; Dye, J.M.; Yavelsky, V.; Lewis, E.C.; et al. Persistent immune responses after ebola virus infection. N. Engl. J. Med. 2013, 369, 492–493. [Google Scholar] [CrossRef] [PubMed]
- Lamunu, M.; Lutwama, J.J.; Kamugisha, J.; Opio, A.; Nambooze, J.; Ndayimirije, N.; Okware, S. Containing a haemorrhagic fever epidemic: The ebola experience in uganda (October 2000–January 2001). Int. J. Infect. Dis. 2004, 8, 27–37. [Google Scholar] [CrossRef] [PubMed]
- Sobarzo, A.; Groseth, A.; Dolnik, O.; Becker, S.; Lutwama, J.J.; Perelman, E.; Yavelsky, V.; Muhammad, M.; Kuehne, A.I.; Marks, R.S.; et al. Profile and persistence of the virus-specific neutralizing humoral immune response in human survivors of sudan ebolavirus (gulu). J. Infect. Dis. 2013, 208, 299–309. [Google Scholar] [CrossRef] [PubMed]
- Sobarzo, A.; Perelman, E.; Groseth, A.; Dolnik, O.; Becker, S.; Lutwama, J.J.; Dye, J.M.; Yavelsky, V.; Lobel, L.; Marks, R.S. Profiling the native specific human humoral immune response to Sudan Ebola virus strain gulu by chemiluminescence enzyme-linked immunosorbent assay. Clin. Vaccine Immunol. 2012, 19, 1844–1852. [Google Scholar] [CrossRef] [PubMed]
- Dias, J.M.; Kuehne, A.I.; Abelson, D.M.; Bale, S.; Wong, A.C.; Halfmann, P.; Muhammad, M.A.; Fusco, M.L.; Zak, S.E.; Kang, E.; et al. A shared structural solution for neutralizing ebolaviruses. Nat. Struct. Mol. Biol. 2011, 18, 1424–1427. [Google Scholar] [CrossRef] [PubMed]
- Kohavi, R. A study of cross. In Proceedings of the 14th International Joint Conference on Artifcial Intelligence (IJCAI); Morgan Kaufmann Publishers Inc.: San Francisco, CA, USA, 1995. [Google Scholar]
- Paweska, J.T.; Burt, F.J.; Swanepoel, R. Validation of igg-sandwich and igm-capture elisa for the detection of antibody to rift valley fever virus in humans. J. Virol. Methods 2005, 124, 173–181. [Google Scholar] [CrossRef] [PubMed]
- Swenson, D.L.; Wang, D.; Luo, M.; Warfield, K.L.; Woraratanadharm, J.; Holman, D.H.; Dong, J.Y.; Pratt, W.D. Vaccine to confer to nonhuman primates complete protection against multistrain ebola and marburg virus infections. Clin. Vaccine Immunol. 2008, 15, 460–467. [Google Scholar] [CrossRef] [PubMed]
- Cavelti-Weder, C.; Babians-Brunner, A.; Keller, C.; Stahel, M.A.; Kurz-Levin, M.; Zayed, H.; Solinger, A.M.; Mandrup-Poulsen, T.; Dinarello, C.A.; Donath, M.Y. Effects of gevokizumab on glycemia and inflammatory markers in type 2 diabetes. Diabetes Care 2012, 35, 1654–1662. [Google Scholar] [CrossRef] [PubMed]
- Dye, J.M.; Herbert, A.S.; Kuehne, A.I.; Barth, J.F.; Muhammad, M.A.; Zak, S.E.; Ortiz, R.A.; Prugar, L.I.; Pratt, W.D. Postexposure antibody prophylaxis protects nonhuman primates from filovirus disease. PNAS 2012, 109, 5034–5039. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, N.J.; Martin, J.E.; Graham, B.S.; Nabel, G.J. Correlates of protective immunity for Ebola vaccines: Implications for regulatory approval by the animal rule. Nat. Rev. Microbiol. 2009, 7, 393–400. [Google Scholar] [CrossRef] [PubMed]
- Shedlock, D.J.; Aviles, J.; Talbott, K.T.; Wong, G.; Wu, S.J.; Villarreal, D.O.; Myles, D.J.; Croyle, M.A.; Yan, J.; Kobinger, G.P.; et al. Induction of broad cytotoxic t cells by protective DNA vaccination against marburg and Ebola. Mol. Ther. 2013, 21, 1432–1444. [Google Scholar] [CrossRef] [PubMed]
- Marzi, A.; Yoshida, R.; Miyamoto, H.; Ishijima, M.; Suzuki, Y.; Higuchi, M.; Matsuyama, Y.; Igarashi, M.; Nakayama, E.; Kuroda, M.; et al. Protective efficacy of neutralizing monoclonal antibodies in a nonhuman primate model of Ebola hemorrhagic fever. PLoS One 2012, 7, e36192. [Google Scholar] [CrossRef] [PubMed]
- Baize, S.; Leroy, E.M.; Georges-Courbot, M.C.; Capron, M.; Lansoud-Soukate, J.; Debre, P.; Fisher-Hoch, S.P.; McCormick, J.B.; Georges, A.J. Defective humoral responses and extensive intravascular apoptosis are associated with fatal outcome in ebola virus-infected patients. Nat. Med. 1999, 5, 423–426. [Google Scholar] [CrossRef] [PubMed]
- Wong, G.; Richardson, J.S.; Pillet, S.; Patel, A.; Qiu, X.; Alimonti, J.; Hogan, J.; Zhang, Y.; Takada, A.; Feldmann, H.; et al. Immune parameters correlate with protection against ebola virus infection in rodents and nonhuman primates. Sci. Transl. Med. 2012, 4, 158ra146. [Google Scholar] [CrossRef] [PubMed]
- Wong, G.; Kobinger, G.P.; Qiu, X. Characterization of host immune responses in ebola virus infections. Expert Rev. Clin. Immunol. 2014, 10, 781–790. [Google Scholar] [CrossRef] [PubMed]
- Hoenen, T.; Groseth, A.; Feldmann, H. Current Ebola vaccines. Expert Opin. Biol. Ther. 2012, 12, 859–872. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, N.J.; Hensley, L.; Asiedu, C.; Geisbert, T.W.; Stanley, D.; Johnson, J.; Honko, A.; Olinger, G.; Bailey, M.; Geisbert, J.B.; et al. Cd8+ cellular immunity mediates rad5 vaccine protection against ebola virus infection of nonhuman primates. Nat. Med. 2011, 17, 1128–1131. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sobarzo, A.; Eskira, Y.; Herbert, A.S.; Kuehne, A.I.; Stonier, S.W.; Ochayon, D.E.; Fedida-Metula, S.; Balinandi, S.; Kislev, Y.; Tali, N.; et al. Immune Memory to Sudan Virus: Comparison between Two Separate Disease Outbreaks. Viruses 2015, 7, 37-51. https://doi.org/10.3390/v7010037
Sobarzo A, Eskira Y, Herbert AS, Kuehne AI, Stonier SW, Ochayon DE, Fedida-Metula S, Balinandi S, Kislev Y, Tali N, et al. Immune Memory to Sudan Virus: Comparison between Two Separate Disease Outbreaks. Viruses. 2015; 7(1):37-51. https://doi.org/10.3390/v7010037
Chicago/Turabian StyleSobarzo, Ariel, Yael Eskira, Andrew S. Herbert, Ana I. Kuehne, Spencer W. Stonier, David E. Ochayon, Shlomit Fedida-Metula, Steven Balinandi, Yaara Kislev, Neta Tali, and et al. 2015. "Immune Memory to Sudan Virus: Comparison between Two Separate Disease Outbreaks" Viruses 7, no. 1: 37-51. https://doi.org/10.3390/v7010037
APA StyleSobarzo, A., Eskira, Y., Herbert, A. S., Kuehne, A. I., Stonier, S. W., Ochayon, D. E., Fedida-Metula, S., Balinandi, S., Kislev, Y., Tali, N., Lewis, E. C., Lutwama, J. J., Dye, J. M., Yavelsky, V., & Lobel, L. (2015). Immune Memory to Sudan Virus: Comparison between Two Separate Disease Outbreaks. Viruses, 7(1), 37-51. https://doi.org/10.3390/v7010037






