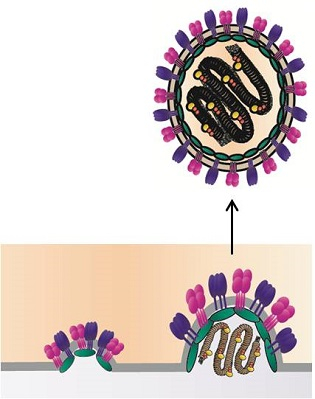
Paramyxovirus Glycoprotein Incorporation, Assembly and Budding: A Three Way Dance for Infectious Particle Production
Abstract
:1. Introduction
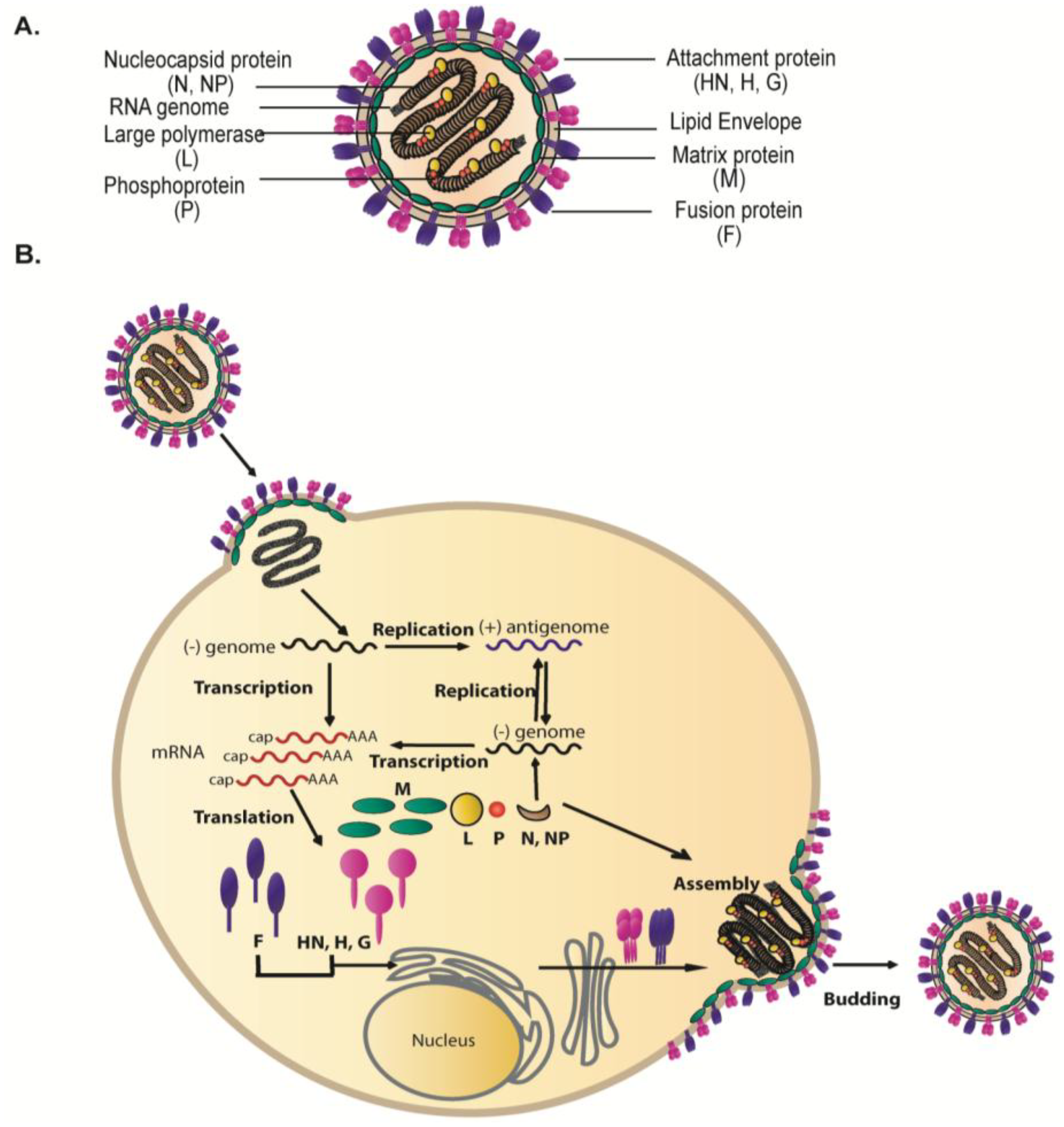
2. Interactions among the Viral Proteins are Critical for Glycoprotein Incorporation and Paramyxovirus Particle Assembly
2.1. Paramyxovirus Surface Glycoproteins
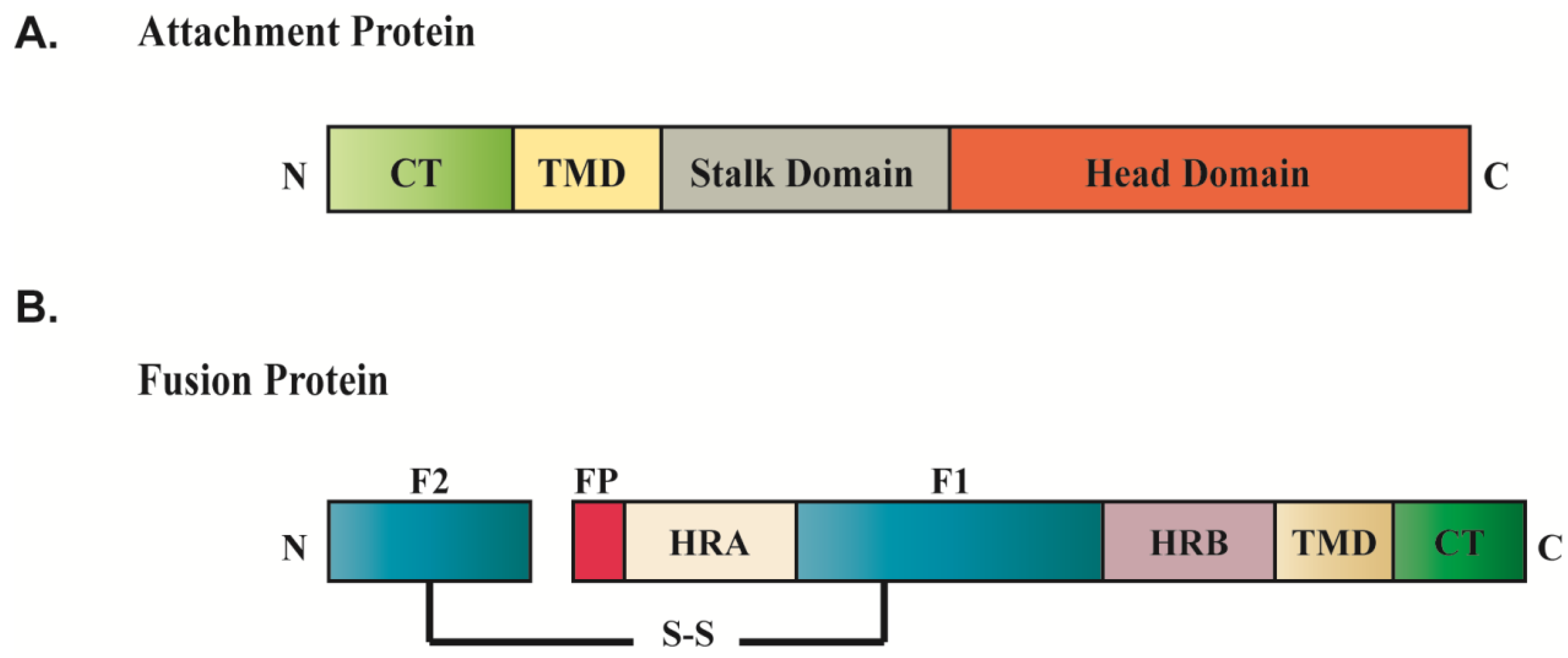
2.2. Matrix Proteins as Coordinators of Paramyxovirus Assembly and Budding
2.3. Interaction of Ribonucleoprotein Complex with Glycoproteins and M
2.4. Active Role of Glycoproteins in Paramyxovirus Particle Formation
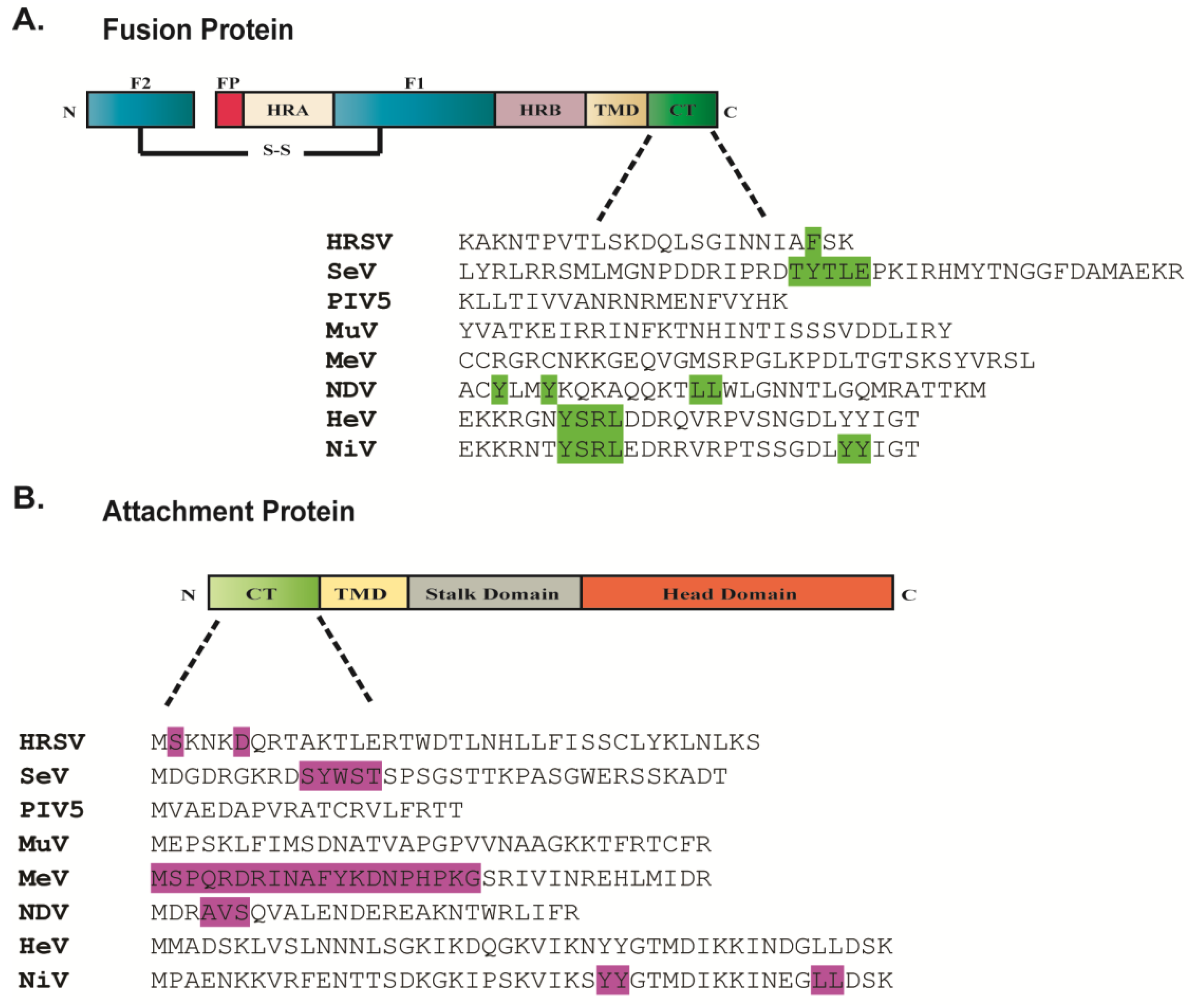
2.5. Role of the Cytoplasmic Tails of Glycoproteins in Particle Production
3. Intracellular Trafficking of Viral Components
3.1. Trafficking of Viral Glycoproteins
3.2. Intracellular Transport of Matrix Proteins and Ribonucleoproteins
4. Localization of Glycoproteins at Membrane Assembly Sites: A Role of Raft Microdomains
5. Role of Envelope Glycoproteins in Paramyxovirus Budding
6. Concluding Remarks
Acknowledgments
Author Contributions
Conflicts of Interest
References and Notes
- Lamb, R.A.; Parks, G.D. Paramyxoviridae: The Viruses and Their Replication, 5th ed.; Fields Virology; Knipe, D.M., Howley, P.M., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2007; Volume 1. [Google Scholar]
- Van den Hoogen, B.G.; de Jong, J.C.; Groen, J.; Kuiken, T.; de Groot, R.; Fouchier, R.A.; Osterhaus, A.D. A newly discovered human pneumovirus isolated from young children with respiratory tract disease. Nat. Med. 2001, 7, 719–724. [Google Scholar]
- Chua, K.B.; Bellini, W.J.; Rota, P.A.; Harcourt, B.H.; Tamin, A.; Lam, S.K.; Ksiazek, T.G.; Rollin, P.E.; Zaki, S.R.; Shieh, W.; et al. Nipah virus: A recently emergent deadly paramyxovirus. Science 2000, 288, 1432–1435. [Google Scholar] [CrossRef]
- Murray, K.; Selleck, P.; Hooper, P.; Hyatt, A.; Gould, A.; Gleeson, L.; Westbury, H.; Hiley, L.; Selvey, L.; Rodwell, B.; et al. A morbillivirus that caused fatal disease in horses and humans. Science 1995, 268, 94–97. [Google Scholar]
- Alexander, D.J.S.D. Newcastle Disease, Other Avian Paramyxoviruses, and Pneumovirus Infections. In Diseases of Poultry, 12th ed.; Saif, Y.M., Glisson, J.R., McDougald, L.R., Nolan, L.K., Swayne, D.E., Eds.; Blackwell Publishing: Ames, IA, USA, 2008. [Google Scholar]
- Govindarajan, D.; Buchholz, U.J.; Samal, S.K. Recovery of avian metapneumovirus subgroup C from cDNA: Cross-recognition of avian and human metapneumovirus support proteins. J. Virol. 2006, 80, 5790–5797. [Google Scholar] [CrossRef]
- Goldsmith, C.S.; Whistler, T.; Rollin, P.E.; Ksiazek, T.G.; Rota, P.A.; Bellini, W.J.; Daszak, P.; Wong, K.T.; Shieh, W.J.; Zaki, S.R. Elucidation of Nipah virus morphogenesis and replication using ultrastructural and molecular approaches. Virus Res. 2003, 92, 89–98. [Google Scholar] [CrossRef]
- Kalica, A.R.; Wright, P.F.; Hetrick, F.M.; Chanock, R.M. Electron microscopic studies of respiratory syncytial temperature-sensitive mutants. Arch. Gesamte Virusforsch 1973, 41, 248–258. [Google Scholar] [CrossRef]
- Yao, Q.; Compans, R.W. Filamentous particle formation by human parainfluenza virus type 2. J. Gen. Virol. 2000, 81, 1305–1312. [Google Scholar]
- Loo, L.H.; Jumat, M.R.; Fu, Y.; Ayi, T.C.; Wong, P.S.; Tee, N.W.; Tan, B.H.; Sugrue, R.J. Evidence for the interaction of the human metapneumovirus G and F proteins during virus-like particle formation. Virol. J. 2013, 10, 294. [Google Scholar] [CrossRef]
- Compans, R.W.; Holmes, K.V.; Dales, S.; Choppin, P.W. An electron microscopic study of moderate and virulent virus-cell interactions of the parainfluenza virus SV5. Virology 1966, 30, 411–426. [Google Scholar] [CrossRef]
- Peret, T.C.; Boivin, G.; Li, Y.; Couillard, M.; Humphrey, C.; Osterhaus, A.D.; Erdman, D.D.; Anderson, L.J. Characterization of human metapneumoviruses isolated from patients in North America. J. Infect. Dis. 2002, 185, 1660–1663. [Google Scholar] [CrossRef]
- Peret, T.C.; Boivin, G.; Li, Y.; Couillard, M.; Humphrey, C.; Osterhaus, A.D.; Erdman, D.D.; Anderson, L.J. Electron cryomicroscopy reveals different F1+F2 protein states in intact parainfluenza virions. J. Virol. 2008, 82, 3775–3781. [Google Scholar] [CrossRef]
- Li, M.; Schmitt, P.T.; Li, Z.; McCrory, T.S.; He, B.; Schmitt, A.P. Mumps virus matrix, fusion, and nucleocapsid proteins cooperate for efficient production of virus-like particles. J. Virol. 2009, 83, 7261–7272. [Google Scholar] [CrossRef]
- Li, Z.; Xu, J.; Patel, J.; Fuentes, S.; Lin, Y.; Anderson, D.; Sakamoto, K.; Wang, L.F.; He, B. Function of the small hydrophobic protein of J paramyxovirus. J. Virol. 2011, 85, 32–42. [Google Scholar] [CrossRef]
- Biacchesi, S.; Pham, Q.N.; Skiadopoulos, M.H.; Murphy, B.R.; Collins, P.L.; Buchholz, U.J. Infection of nonhuman primates with recombinant human metapneumovirus lacking the SH, G, or M2–2 protein categorizes each as a nonessential accessory protein and identifies vaccine candidates. J. Virol. 2005, 79, 12608–12613. [Google Scholar] [CrossRef]
- Takeuchi, K.; Tanabayashi, K.; Hishiyama, M.; Yamada, A. The mumps virus SH protein is a membrane protein and not essential for virus growth. Virology 1996, 225, 156–162. [Google Scholar] [CrossRef]
- He, B.; Leser, G.P.; Paterson, R.G.; Lamb, R.A. The paramyxovirus SV5 small hydrophobic (SH) protein is not essential for virus growth in tissue culture cells. Virology 1998, 250, 30–40. [Google Scholar] [CrossRef]
- Karron, R.A.; Buonagurio, D.A.; Georgiu, A.F.; Whitehead, S.S.; Adamus, J.E.; Clements-Mann, M.L.; Harris, D.O.; Randolph, V.B.; Udem, S.A.; Murphy, B.R.; et al. Respiratory syncytial virus (RSV) SH and G proteins are not essential for viral replication in vitro: Clinical evaluation and molecular characterization of a cold-passaged, attenuated RSV subgroup B mutant. Proc. Natl. Acad. Sci. USA 1997, 94, 13961–13966. [Google Scholar] [CrossRef]
- Harrison, M.S.; Sakaguchi, T.; Schmitt, A.P. Paramyxovirus assembly and budding: Building particles that transmit infections. Int. J. Biochem. Cell. Biol. 2010, 42, 1416–1429. [Google Scholar] [CrossRef]
- Takimoto, T.; Portner, A. Molecular mechanism of paramyxovirus budding. Virus Res. 2004, 106, 133–145. [Google Scholar] [CrossRef]
- Liljeroos, L.; Butcher, S.J. Matrix proteins as centralized organizers of negative-sense RNA virions. Front. Biosci. (Landmark Ed) 2013, 18, 696–715. [Google Scholar] [CrossRef]
- Villar, E.; Barroso, I.M. Role of sialic acid-containing molecules in paramyxovirus entry into the host cell: A minireview. Glycoconj. J. 2006, 23, 5–17. [Google Scholar] [CrossRef]
- Plemper, R.K.; Brindley, M.A.; Iorio, R.M. Structural and mechanistic studies of measles virus illuminate paramyxovirus entry. PLoS Pathog. 2011, 7, e1002058. [Google Scholar] [CrossRef]
- Plattet, P.; Plemper, R.K. Envelope protein dynamics in paramyxovirus entry. MBio 2013, 4, e00413-13. [Google Scholar]
- Lamb, R.A.; Jardetzky, T.S. Structural basis of viral invasion: Lessons from paramyxovirus F. Curr. Opin. Struct. Biol. 2007, 17, 427–436. [Google Scholar] [CrossRef]
- Bissonnette, M.L.; Connolly, S.A.; Young, D.F.; Randall, R.E.; Paterson, R.G.; Lamb, R.A. Analysis of the pH requirement for membrane fusion of different isolates of the paramyxovirus parainfluenza virus 5. J. Virol. 2006, 80, 3071–3077. [Google Scholar] [CrossRef]
- Baker, K.A.; Dutch, R.E.; Lamb, R.A.; Jardetzky, T.S. Structural basis for paramyxovirus-mediated membrane fusion. Mol. Cell 1999, 3, 309–319. [Google Scholar] [CrossRef]
- McGinnes, L.; Sergel, T.; Morrison, T. Mutations in the transmembrane domain of the HN protein of Newcastle disease virus affect the structure and activity of the protein. Virology 1993, 196, 101–110. [Google Scholar] [CrossRef]
- Biacchesi, S.; Skiadopoulos, M.H.; Yang, L.; Lamirande, E.W.; Tran, K.C.; Murphy, B.R.; Collins, P.L.; Buchholz, U.J. Recombinant human metapneumovirus lacking the small hydrophobic SH and/or attachment G glycoprotein: Deletion of G yields a promising vaccine candidate. J. Virol. 2004, 78, 12877–12887. [Google Scholar] [CrossRef]
- Teng, M.N.; Whitehead, S.S.; Collins, P.L. Contribution of the respiratory syncytial virus G glycoprotein and its secreted and membrane-bound forms to virus replication in vitro and in vivo. Virology 2001, 289, 283–296. [Google Scholar] [CrossRef]
- Schowalter, R.M.; Smith, S.E.; Dutch, R.E. Characterization of human metapneumovirus F protein-promoted membrane fusion: Critical roles for proteolytic processing and low pH. J. Virol. 2006, 80, 10931–10941. [Google Scholar] [CrossRef]
- Herfst, S.; Mas, V.; Ver, L.S.; Wierda, R.J.; Osterhaus, A.D.; Fouchier, R.A.; Melero, J.A. Low-pH-induced membrane fusion mediated by human metapneumovirus F protein is a rare, strain-dependent phenomenon. J. Virol. 2008, 82, 8891–8895. [Google Scholar] [CrossRef]
- Tayyari, F.; Marchant, D.; Moraes, T.J.; Duan, W.; Mastrangelo, P.; Hegele, R.G. Identification of nucleolin as a cellular receptor for human respiratory syncytial virus. Nat. Med. 2011, 17, 1132–1135. [Google Scholar]
- Cseke, G.; Maginnis, M.S.; Cox, R.G.; Tollefson, S.J.; Podsiad, A.B.; Wright, D.W.; Dermody, T.S.; Williams, J.V. Integrin alphavbeta1 promotes infection by human metapneumovirus. Proc. Natl. Acad. Sci. USA 2009, 106, 1566–1571. [Google Scholar] [CrossRef]
- Chang, A.; Masante, C.; Buchholz, U.J.; Dutch, R.E. Human metapneumovirus (HMPV) binding and infection are mediated by interactions between the HMPV fusion protein and heparan sulfate. J. Virol. 2012, 86, 3230–3243. [Google Scholar] [CrossRef]
- Wei, Y.; Zhang, Y.; Cai, H.; Mirza, A.M.; Iorio, R.M.; Peeples, M.E.; Niewiesk, S.; Li, J. Roles of the putative integrin-binding motif of the human metapneumovirus fusion (F) protein in cell-cell fusion, viral infectivity, and pathogenesis. J. Virol. 2014, 88, 4338–4352. [Google Scholar] [CrossRef]
- Colf, L.A.; Juo, Z.S.; Garcia, K.C. Structure of the measles virus hemagglutinin. Nat. Struct. Mol. Biol. 2007, 14, 1227–1228. [Google Scholar] [CrossRef]
- Hashiguchi, T.; Kajikawa, M.; Maita, N.; Takeda, M.; Kuroki, K.; Sasaki, K.; Kohda, D.; Yanagi, Y.; Maenaka, K. Crystal structure of measles virus hemagglutinin provides insight into effective vaccines. Proc. Natl. Acad. Sci. USA 2007, 104, 19535–19540. [Google Scholar] [CrossRef]
- Lawrence, M.C.; Borg, N.A.; Streltsov, V.A.; Pilling, P.A.; Epa, V.C.; Varghese, J.N.; McKimm-Breschkin, J.L.; Colman, P.M. Structure of the haemagglutinin-neuraminidase from human parainfluenza virus type III. J. Mol. Biol. 2004, 335, 1343–1357. [Google Scholar] [CrossRef]
- Yuan, P.; Paterson, R.G.; Leser, G.P.; Lamb, R.A.; Jardetzky, T.S. Structure of the ulster strain Newcastle disease virus hemagglutinin-neuraminidase reveals auto-inhibitory interactions associated with low virulence. PLoS Pathog. 2012, 8, e1002855. [Google Scholar] [CrossRef]
- Yuan, P.; Swanson, K.A.; Leser, G.P.; Paterson, R.G.; Lamb, R.A.; Jardetzky, T.S. Structure of the Newcastle disease virus hemagglutinin-neuraminidase (HN) ectodomain reveals a four-helix bundle stalk. Proc. Natl. Acad. Sci. USA 2011, 108, 14920–14925. [Google Scholar]
- Yuan, P.; Thompson, T.B.; Wurzburg, B.A.; Paterson, R.G.; Lamb, R.A.; Jardetzky, T.S. Structural studies of the parainfluenza virus 5 hemagglutinin-neuraminidase tetramer in complex with its receptor, sialyllactose. Structure 2005, 13, 803–815. [Google Scholar] [CrossRef]
- Xu, K.; Rajashankar, K.R.; Chan, Y.P.; Himanen, J.P.; Broder, C.C.; Nikolov, D.B. Host cell recognition by the henipaviruses: Crystal structures of the Nipah G attachment glycoprotein and its complex with ephrin-B3. Proc. Natl. Acad. Sci. USA 2008, 105, 9953–9958. [Google Scholar] [CrossRef]
- Bowden, T.A.; Crispin, M.; Harvey, D.J.; Jones, E.Y.; Stuart, D.I. Dimeric architecture of the Hendra virus attachment glycoprotein: Evidence for a conserved mode of assembly. J. Virol. 2010, 84, 6208–6217. [Google Scholar] [CrossRef]
- Bowden, T.A.; Crispin, M.; Harvey, D.J.; Aricescu, A.R.; Grimes, J.M.; Jones, E.Y.; Stuart, D.I. Crystal structure and carbohydrate analysis of Nipah virus attachment glycoprotein: A template for antiviral and vaccine design. J. Virol. 2008, 82, 11628–11636. [Google Scholar] [CrossRef]
- Brindley, M.A.; Plemper, R.K. Blue native PAGE and biomolecular complementation reveal a tetrameric or higher-order oligomer organization of the physiological measles virus attachment protein H. J. Virol. 2010, 84, 12174–12184. [Google Scholar] [CrossRef]
- Xu, K.; Chan, Y.P.; Rajashankar, K.R.; Khetawat, D.; Yan, L.; Kolev, M.V.; Broder, C.C.; Nikolov, D.B. New insights into the Hendra virus attachment and entry process from structures of the virus G glycoprotein and its complex with Ephrin-B2. PLoS One 2012, 7, e48742. [Google Scholar] [CrossRef]
- Bose, S.; Welch, B.D.; Kors, C.A.; Yuan, P.; Jardetzky, T.S.; Lamb, R.A. Structure and mutagenesis of the parainfluenza virus 5 hemagglutinin-neuraminidase stalk domain reveals a four-helix bundle and the role of the stalk in fusion promotion. J. Virol. 2011, 85, 12855–12866. [Google Scholar] [CrossRef]
- Deng, R.; Wang, Z.; Mahon, P.J.; Marinello, M.; Mirza, A.; Iorio, R.M. Mutations in the Newcastle disease virus hemagglutinin-neuraminidase protein that interfere with its ability to interact with the homologous F protein in the promotion of fusion. Virology 1999, 253, 43–54. [Google Scholar] [CrossRef]
- Melanson, V.R.; Iorio, R.M. Amino acid substitutions in the F-specific domain in the stalk of the Newcastle disease virus HN protein modulate fusion and interfere with its interaction with the F protein. J. Virol. 2004, 78, 13053–13061. [Google Scholar] [CrossRef]
- Paal, T.; Brindley, M.A.; St Clair, C.; Prussia, A.; Gaus, D.; Krumm, S.A.; Snyder, J.P.; Plemper, R.K. Probing the spatial organization of measles virus fusion complexes. J. Virol. 2009, 83, 10480–10493. [Google Scholar] [CrossRef]
- Lee, J.K.; Prussia, A.; Paal, T.; White, L.K.; Snyder, J.P.; Plemper, R.K. Functional interaction between paramyxovirus fusion and attachment proteins. J. Biol. Chem. 2008, 283, 16561–16572. [Google Scholar]
- Stone-Hulslander, J.; Morrison, T.G. Mutational analysis of heptad repeats in the membrane-proximal region of Newcastle disease virus HN protein. J. Virol. 1999, 73, 3630–3637. [Google Scholar]
- Yao, Q.; Hu, X.; Compans, R.W. Association of the parainfluenza virus fusion and hemagglutinin-neuraminidase glycoproteins on cell surfaces. J. Virol. 1997, 71, 650–656. [Google Scholar]
- Plemper, R.K.; Hammond, A.L.; Gerlier, D.; Fielding, A.K.; Cattaneo, R. Strength of envelope protein interaction modulates cytopathicity of measles virus. J. Virol. 2002, 76, 5051–5061. [Google Scholar] [CrossRef]
- Aguilar, H.C.; Ataman, Z.A.; Aspericueta, V.; Fang, A.Q.; Stroud, M.; Negrete, O.A.; Kammerer, R.A.; Lee, B. A novel receptor-induced activation site in the Nipah virus attachment glycoprotein (G) involved in triggering the fusion glycoprotein (F). J. Biol. Chem. 2009, 284, 1628–1635. [Google Scholar]
- Dutch, R.E.; Jardetzky, T.S.; Lamb, R.A. Virus membrane fusion proteins: Biological machines that undergo a metamorphosis. Biosci. Rep. 2000, 20, 597–612. [Google Scholar] [CrossRef]
- Murakami, M.; Towatari, T.; Ohuchi, M.; Shiota, M.; Akao, M.; Okumura, Y.; Parry, M.A.; Kido, H. Mini-plasmin found in the epithelial cells of bronchioles triggers infection by broad-spectrum influenza A viruses and Sendai virus. Eur. J. Biochem. 2001, 268, 2847–2855. [Google Scholar] [CrossRef]
- Garten, W.; Hallenberger, S.; Ortmann, D.; Schafer, W.; Vey, M.; Angliker, H.; Shaw, E.; Klenk, H.D. Processing of viral glycoproteins by the subtilisin-like endoprotease furin and its inhibition by specific peptidylchloroalkylketones. Biochimie 1994, 76, 217–225. [Google Scholar] [CrossRef]
- Watanabe, M.; Hirano, A.; Stenglein, S.; Nelson, J.; Thomas, G.; Wong, T.C. Engineered serine protease inhibitor prevents furin-catalyzed activation of the fusion glycoprotein and production of infectious measles virus. J. Virol. 1995, 69, 3206–3210. [Google Scholar]
- Gotoh, B.; Ohnishi, Y.; Inocencio, N.M.; Esaki, E.; Nakayama, K.; Barr, P.J.; Thomas, G.; Nagai, Y. Mammalian subtilisin-related proteinases in cleavage activation of the paramyxovirus fusion glycoprotein: Superiority of furin/PACE to PC2 or PC1/PC3. J. Virol. 1992, 66, 6391–6397. [Google Scholar]
- Pager, C.T.; Craft, W.W.; Patch, J.; Dutch, R.E. A mature and fusogenic form of the Nipah virus fusion protein requires proteolytic processing by cathepsin L. Virology 2006, 346, 251–257. [Google Scholar] [CrossRef]
- Pager, C.T.; Dutch, R.E. Cathepsin L is involved in proteolytic processing of the Hendra virus fusion protein. J. Virol. 2005, 79, 12714–12720. [Google Scholar] [CrossRef]
- Meulendyke, K.A.; Wurth, M.A.; McCann, R.O.; Dutch, R.E. Endocytosis plays a critical role in proteolytic processing of the Hendra virus fusion protein. J. Virol. 2005, 79, 12643–12649. [Google Scholar] [CrossRef]
- Diederich, S.; Thiel, L.; Maisner, A. Role of endocytosis and cathepsin-mediated activation in Nipah virus entry. Virology 2008, 375, 391–400. [Google Scholar] [CrossRef]
- Krzyzaniak, M.A.; Zumstein, M.T.; Gerez, J.A.; Picotti, P.; Helenius, A. Host cell entry of respiratory syncytial virus involves macropinocytosis followed by proteolytic activation of the F protein. PLoS Pathog. 2013, 9, e1003309. [Google Scholar] [CrossRef]
- Jardetzky, T.S.; Lamb, R.A. Activation of paramyxovirus membrane fusion and virus entry. Curr. Opin. Virol. 2014, 5C, 24–33. [Google Scholar] [CrossRef]
- Chang, A.; Dutch, R.E. Paramyxovirus fusion and entry: Multiple paths to a common end. Viruses 2012, 4, 613–636. [Google Scholar] [CrossRef]
- Smith, E.C.; Popa, A.; Chang, A.; Masante, C.; Dutch, R.E. Viral entry mechanisms: The increasing diversity of paramyxovirus entry. FEBS J. 2009, 276, 7217–7227. [Google Scholar] [CrossRef]
- Perez, M.; Garcia-Barreno, B.; Melero, J.A.; Carrasco, L.; Guinea, R. Membrane permeability changes induced in Escherichia coli by the SH protein of human respiratory syncytial virus. Virology 1997, 235, 342–351. [Google Scholar] [CrossRef]
- Gan, S.W.; Ng, L.; Lin, X.; Gong, X.; Torres, J. Structure and ion channel activity of the human respiratory syncytial virus (hRSV) small hydrophobic protein transmembrane domain. Protein Sci. 2008, 17, 813–820. [Google Scholar] [CrossRef]
- Kochva, U.; Leonov, H.; Arkin, I.T. Modeling the structure of the respiratory syncytial virus small hydrophobic protein by silent-mutation analysis of global searching molecular dynamics. Protein Sci. 2003, 12, 2668–2674. [Google Scholar] [CrossRef]
- Gan, S.W.; Tan, E.; Lin, X.; Yu, D.; Wang, J.; Tan, G.M.; Vararattanavech, A.; Yeo, C.Y.; Soon, C.H.; Soong, T.W.; Pervushin, K.; Torres, J. The small hydrophobic protein of the human respiratory syncytial virus forms pentameric ion channels. J. Biol. Chem. 2012, 287, 24671–24689. [Google Scholar] [CrossRef]
- Collins, P.L.; Mottet, G. Membrane orientation and oligomerization of the small hydrophobic protein of human respiratory syncytial virus. J. Gen. Virol. 1993, 74, 1445–1450. [Google Scholar] [CrossRef]
- Carter, S.D.; Dent, K.C.; Atkins, E.; Foster, T.L.; Verow, M.; Gorny, P.; Harris, M.; Hiscox, J.A.; Ranson, N.A.; Griffin, S.; Barr, J.N. Direct visualization of the small hydrophobic protein of human respiratory syncytial virus reveals the structural basis for membrane permeability. FEBS Lett. 2010, 584, 2786–2790. [Google Scholar] [CrossRef]
- Masante, C.; el Najjar, F.; Chang, A.; Jones, A.; Moncman, C.L.; Dutch, R.E. The human metapneumovirus small hydrophobic protein has properties consistent with those of a viroporin and can modulate viral fusogenic activity. J. Virol. 2014, 88, 6423–6433. [Google Scholar] [CrossRef]
- Bao, X.; Kolli, D.; Liu, T.; Shan, Y.; Garofalo, R.P.; Casola, A. Human metapneumovirus small hydrophobic protein inhibits NF-kappaB transcriptional activity. J. Virol. 2008, 82, 8224–8229. [Google Scholar] [CrossRef]
- Wilson, R.L.; Fuentes, S.M.; Wang, P.; Taddeo, E.C.; Klatt, A.; Henderson, A.J.; He, B. Function of small hydrophobic proteins of paramyxovirus. J. Virol. 2006, 80, 1700–1709. [Google Scholar] [CrossRef]
- Le Nouen, C.; Hillyer, P.; Brock, L.G.; Winter, C.C.; Rabin, R.L.; Collins, P.L.; Buchholz, U.J. Human Metapneumovirus SH and G glycoproteins inhibit macropinocytosis-mediated entry into human dendritic cells and reduce CD4+ t cell activation. J. Virol. 2014, 88, 6453–6469. [Google Scholar] [CrossRef]
- Jin, H.; Cheng, X.; Zhou, H.Z.; Li, S.; Seddiqui, A. Respiratory syncytial virus that lacks open reading frame 2 of the M2 gene (M2–2) has altered growth characteristics and is attenuated in rodents. J. Virol. 2000, 74, 74–82. [Google Scholar] [CrossRef]
- Ling, R.; Sinkovic, S.; Toquin, D.; Guionie, O.; Eterradossi, N.; Easton, A.J. Deletion of the SH gene from avian metapneumovirus has a greater impact on virus production and immunogenicity in turkeys than deletion of the G gene or M2–2 open reading frame. J. Gen. Virol. 2008, 89, 525–533. [Google Scholar] [CrossRef]
- Bukreyev, A.; Whitehead, S.S.; Murphy, B.R.; Collins, P.L. Recombinant respiratory syncytial virus from which the entire SH gene has been deleted grows efficiently in cell culture and exhibits site-specific attenuation in the respiratory tract of the mouse. J. Virol. 1997, 71, 8973–8982. [Google Scholar]
- Whitehead, S.S.; Bukreyev, A.; Teng, M.N.; Firestone, C.Y.; St Claire, M.; Elkins, W.R.; Collins, P.L.; Murphy, B.R. Recombinant respiratory syncytial virus bearing a deletion of either the NS2 or SH gene is attenuated in chimpanzees. J. Virol. 1999, 73, 3438–3442. [Google Scholar]
- Yoshida, T.; Nagai, Y.; Maeno, K.; Iinuma, M.; Hamaguchi, M.; Matsumoto, T.; Nagayoshi, S.; Hoshino, M. Studies on the role of M protein in virus assembly using a ts mutant of HVJ (Sendai virus). Virology 1979, 92, 139–154. [Google Scholar] [CrossRef]
- Kondo, T.; Yoshida, T.; Miura, N.; Nakanishi, M. Temperature-sensitive phenotype of a mutant Sendai virus strain is caused by its insufficient accumulation of the M protein. J. Biol. Chem. 1993, 268, 21924–21930. [Google Scholar]
- Baczko, K.; Liebert, U.G.; Billeter, M.; Cattaneo, R.; Budka, H.; ter Meulen, V. Expression of defective measles virus genes in brain tissues of patients with subacute sclerosing panencephalitis. J. Virol. 1986, 59, 472–478. [Google Scholar]
- Hirano, A.; Ayata, M.; Wang, A.H.; Wong, T.C. Functional analysis of matrix proteins expressed from cloned genes of measles virus variants that cause subacute sclerosing panencephalitis reveals a common defect in nucleocapsid binding. J. Virol. 1993, 67, 1848–1853. [Google Scholar]
- Cathomen, T.; Mrkic, B.; Spehner, D.; Drillien, R.; Naef, R.; Pavlovic, J.; Aguzzi, A.; Billeter, M.A.; Cattaneo, R. A matrix-less measles virus is infectious and elicits extensive cell fusion: Consequences for propagation in the brain. EMBO J. 1998, 17, 3899–3908. [Google Scholar] [CrossRef]
- Mitra, R.; Baviskar, P.; Duncan-Decocq, R.R.; Patel, D.; Oomens, A.G. The human respiratory syncytial virus matrix protein is required for maturation of viral filaments. J. Virol. 2012, 86, 4432–4443. [Google Scholar] [CrossRef]
- Takimoto, T.; Murti, K.G.; Bousse, T.; Scroggs, R.A.; Portner, A. Role of matrix and fusion proteins in budding of Sendai virus. J. Virol. 2001, 75, 11384–11391. [Google Scholar] [CrossRef]
- Sugahara, F.; Uchiyama, T.; Watanabe, H.; Shimazu, Y.; Kuwayama, M.; Fujii, Y.; Kiyotani, K.; Adachi, A.; Kohno, N.; Yoshida, T.; Sakaguchi, T. Paramyxovirus Sendai virus-like particle formation by expression of multiple viral proteins and acceleration of its release by C protein. Virology 2004, 325, 1–10. [Google Scholar] [CrossRef]
- Pohl, C.; Duprex, W.P.; Krohne, G.; Rima, B.K.; Schneider-Schaulies, S. Measles virus M and F proteins associate with detergent-resistant membrane fractions and promote formation of virus-like particles. J. Gen. Virol. 2007, 88, 1243–1250. [Google Scholar] [CrossRef]
- Runkler, N.; Pohl, C.; Schneider-Schaulies, S.; Klenk, H.D.; Maisner, A. Measles virus nucleocapsid transport to the plasma membrane requires stable expression and surface accumulation of the viral matrix protein. Cell Microbiol. 2007, 9, 1203–1214. [Google Scholar] [CrossRef]
- Ciancanelli, M.J.; Basler, C.F. Mutation of YMYL in the Nipah virus matrix protein abrogates budding and alters subcellular localization. J. Virol. 2006, 80, 12070–12078. [Google Scholar] [CrossRef]
- Patch, J.R.; Crameri, G.; Wang, L.F.; Eaton, B.T.; Broder, C.C. Quantitative analysis of Nipah virus proteins released as virus-like particles reveals central role for the matrix protein. Virol. J. 2007, 4, 1. [Google Scholar] [CrossRef]
- Coronel, E.C.; Murti, K.G.; Takimoto, T.; Portner, A. Human parainfluenza virus type 1 matrix and nucleoprotein genes transiently expressed in mammalian cells induce the release of virus-like particles containing nucleocapsid-like structures. J. Virol. 1999, 73, 7035–7038. [Google Scholar]
- Pantua, H.D.; McGinnes, L.W.; Peeples, M.E.; Morrison, T.G. Requirements for the assembly and release of Newcastle disease virus-like particles. J. Virol. 2006, 80, 11062–11073. [Google Scholar] [CrossRef]
- Battisti, A.J.; Meng, G.; Winkler, D.C.; McGinnes, L.W.; Plevka, P.; Steven, A.C.; Morrison, T.G.; Rossmann, M.G. Structure and assembly of a paramyxovirus matrix protein. Proc. Natl. Acad. Sci. USA 2012, 109, 13996–14000. [Google Scholar] [CrossRef]
- Money, V.A.; McPhee, H.K.; Mosely, J.A.; Sanderson, J.M.; Yeo, R.P. Surface features of a Mononegavirales matrix protein indicate sites of membrane interaction. Proc. Natl. Acad. Sci. USA 2009, 106, 4441–4446. [Google Scholar]
- Leyrat, C.; Renner, M.; Harlos, K.; Huiskonen, J.T.; Grimes, J.M. Structure and self-assembly of the calcium binding matrix protein of human metapneumovirus. Structure 2014, 22, 136–148. [Google Scholar] [CrossRef]
- Dessen, A.; Volchkov, V.; Dolnik, O.; Klenk, H.D.; Weissenhorn, W. Crystal structure of the matrix protein VP40 from Ebola virus. EMBO J. 2000, 19, 4228–4236. [Google Scholar] [CrossRef]
- Neumann, P.; Lieber, D.; Meyer, S.; Dautel, P.; Kerth, A.; Kraus, I.; Garten, W.; Stubbs, M.T. Crystal structure of the Borna disease virus matrix protein (BDV-M) reveals ssRNA binding properties. Proc. Natl. Acad. Sci. USA 2009, 106, 3710–3715. [Google Scholar]
- Hewitt, J.A.; Nermut, M.V. A morphological study of the M-protein of Sendai virus. J. Gen. Virol. 1977, 34, 127–136. [Google Scholar] [CrossRef]
- Heggeness, M.H.; Smith, P.R.; Choppin, P.W. In vitro assembly of the nonglycosylated membrane protein (M) of Sendai virus. Proc. Natl. Acad. Sci. USA 1982, 79, 6232–6236. [Google Scholar] [CrossRef]
- McPhee, H.K.; Carlisle, J.L.; Beeby, A.; Money, V.A.; Watson, S.M.; Yeo, R.P.; Sanderson, J.M. Influence of lipids on the interfacial disposition of respiratory syncytial virus matrix protein. Langmuir 2011, 27, 304–311. [Google Scholar] [CrossRef]
- Jasenosky, L.D.; Neumann, G.; Lukashevich, I.; Kawaoka, Y. Ebola virus VP40-induced particle formation and association with the lipid bilayer. J. Virol. 2001, 75, 5205–5214. [Google Scholar] [CrossRef]
- Stricker, R.; Mottet, G.; Roux, L. The Sendai virus matrix protein appears to be recruited in the cytoplasm by the viral nucleocapsid to function in viral assembly and budding. J. Gen. Virol. 1994, 75, 1031–1042. [Google Scholar] [CrossRef]
- Riedl, P.; Moll, M.; Klenk, H.D.; Maisner, A. Measles virus matrix protein is not cotransported with the viral glycoproteins but requires virus infection for efficient surface targeting. Virus Res. 2002, 83, 1–12. [Google Scholar] [CrossRef]
- Subhashri, R.; Shaila, M.S. Characterization of membrane association of Rinderpest virus matrix protein. Biochem. Biophys. Res. Commun. 2007, 355, 1096–1101. [Google Scholar] [CrossRef]
- Ye, Z.; Sun, W.; Suryanarayana, K.; Justice, P.; Robinson, D.; Wagner, R.R. Membrane-binding domains and cytopathogenesis of the matrix protein of vesicular stomatitis virus. J. Virol. 1994, 68, 7386–7396. [Google Scholar]
- Shnyrova, A.V.; Ayllon, J.; Mikhalyov, I.I.; Villar, E.; Zimmerberg, J.; Frolov, V.A. Vesicle formation by self-assembly of membrane-bound matrix proteins into a fluidlike budding domain. J. Cell Biol. 2007, 179, 627–633. [Google Scholar] [CrossRef]
- Liljeroos, L.; Krzyzaniak, M.A.; Helenius, A.; Butcher, S.J. Architecture of respiratory syncytial virus revealed by electron cryotomography. Proc. Natl. Acad. Sci. USA 2013, 110, 11133–11138. [Google Scholar] [CrossRef]
- Loney, C.; Mottet-Osman, G.; Roux, L.; Bhella, D. Paramyxovirus ultrastructure and genome packaging: Cryo-electron tomography of Sendai virus. J. Virol. 2009, 83, 8191–8197. [Google Scholar] [CrossRef]
- Kiss, G.; Holl, J.M.; Williams, G.M.; Alonas, E.; Vanover, D.; Lifland, A.W.; Gudheti, M.; Guerrero-Ferreira, R.C.; Nair, V.; Yi, H.; et al. Structural analysis of respiratory syncytial virus reveals the position of m2–1 between the matrix protein and the ribonucleoprotein complex. J. Virol. 2014, 88, 7602–7617. [Google Scholar] [CrossRef]
- Liljeroos, L.; Huiskonen, J.T.; Ora, A.; Susi, P.; Butcher, S.J. Electron cryotomography of measles virus reveals how matrix protein coats the ribonucleocapsid within intact virions. Proc. Natl. Acad. Sci. USA 2011, 108, 18085–18090. [Google Scholar]
- Tawar, R.G.; Duquerroy, S.; Vonrhein, C.; Varela, P.F.; Damier-Piolle, L.; Castagne, N.; MacLellan, K.; Bedouelle, H.; Bricogne, G.; Bhella, D.; et al. Crystal structure of a nucleocapsid-like nucleoprotein-RNA complex of respiratory syncytial virus. Science 2009, 326, 1279–1283. [Google Scholar] [CrossRef]
- Errington, W.; Emmerson, P.T. Assembly of recombinant Newcastle disease virus nucleocapsid protein into nucleocapsid-like structures is inhibited by the phosphoprotein. J. Gen. Virol. 1997, 78, 2335–2339. [Google Scholar]
- Schmitt, A.P.; Leser, G.P.; Waning, D.L.; Lamb, R.A. Requirements for budding of paramyxovirus simian virus 5 virus-like particles. J. Virol. 2002, 76, 3952–3964. [Google Scholar] [CrossRef]
- Spehner, D.; Kirn, A.; Drillien, R. Assembly of nucleocapsidlike structures in animal cells infected with a vaccinia virus recombinant encoding the measles virus nucleoprotein. J. Virol. 1991, 65, 6296–6300. [Google Scholar]
- Rager, M.; Vongpunsawad, S.; Duprex, W.P.; Cattaneo, R. Polyploid measles virus with hexameric genome length. EMBO J. 2002, 21, 2364–2372. [Google Scholar] [CrossRef]
- Goff, P.H.; Gao, Q.; Palese, P. A majority of infectious Newcastle disease virus particles contain a single genome, while a minority contain multiple genomes. J. Virol. 2012, 86, 10852–10856. [Google Scholar] [CrossRef]
- Coronel, E.C.; Takimoto, T.; Murti, K.G.; Varich, N.; Portner, A. Nucleocapsid incorporation into parainfluenza virus is regulated by specific interaction with matrix protein. J. Virol. 2001, 75, 1117–1123. [Google Scholar] [CrossRef]
- Mottet, G.; Roux, L. Budding efficiency of Sendai virus nucleocapsids: Influence of size and ends of the RNA. Virus Res. 1989, 14, 175–187. [Google Scholar] [CrossRef]
- Kolakofsky, D.; Bruschi, A. Antigenomes in Sendai virions and Sendai virus-infected cells. Virology 1975, 66, 185–191. [Google Scholar] [CrossRef]
- Iwasaki, M.; Takeda, M.; Shirogane, Y.; Nakatsu, Y.; Nakamura, T.; Yanagi, Y. The matrix protein of measles virus regulates viral RNA synthesis and assembly by interacting with the nucleocapsid protein. J. Virol. 2009, 83, 10374–10383. [Google Scholar] [CrossRef]
- Schmitt, P.T.; Ray, G.; Schmitt, A.P. The C-terminal end of parainfluenza virus 5 NP protein is important for virus-like particle production and M-NP protein interaction. J. Virol. 2010, 84, 12810–12823. [Google Scholar] [CrossRef]
- Li, D.; Jans, D.A.; Bardin, P.G.; Meanger, J.; Mills, J.; Ghildyal, R. Association of respiratory syncytial virus M protein with viral nucleocapsids is mediated by the M2–1 protein. J. Virol. 2008, 82, 8863–8870. [Google Scholar] [CrossRef]
- Hamaguchi, M.; Yoshida, T.; Nishikawa, K.; Naruse, H.; Nagai, Y. Transcriptive complex of Newcastle disease virus. I. Both L and P proteins are required to constitute an active complex. Virology 1983, 128, 105–117. [Google Scholar] [CrossRef]
- Rodriguez, L.; Cuesta, I.; Asenjo, A.; Villanueva, N. Human respiratory syncytial virus matrix protein is an RNA-binding protein: Binding properties, location and identity of the RNA contact residues. J. Gen. Virol. 2004, 85, 709–719. [Google Scholar] [CrossRef]
- Essaidi-Laziosi, M.; Shevtsova, A.; Gerlier, D.; Roux, L. Mutation of the TYTLE motif in the cytoplasmic tail of the sendai virus fusion protein deeply affects viral assembly and particle production. PLoS One 2013, 8, e78074. [Google Scholar] [CrossRef]
- Ali, A.; Nayak, D.P. Assembly of Sendai virus: M protein interacts with F and HN proteins and with the cytoplasmic tail and transmembrane domain of F protein. Virology 2000, 276, 289–303. [Google Scholar] [CrossRef]
- Sanderson, C.M.; Wu, H.H.; Nayak, D.P. Sendai virus M protein binds independently to either the F or the HN glycoprotein in vivo. J. Virol. 1994, 68, 69–76. [Google Scholar]
- Fouillot-Coriou, N.; Roux, L. Structure-function analysis of the Sendai virus F and HN cytoplasmic domain: Different role for the two proteins in the production of virus particle. Virology 2000, 270, 464–475. [Google Scholar] [CrossRef]
- Mottet-Osman, G.; Iseni, F.; Pelet, T.; Wiznerowicz, M.; Garcin, D.; Roux, L. Suppression of the Sendai virus M protein through a novel short interfering RNA approach inhibits viral particle production but does not affect viral RNA synthesis. J. Virol. 2007, 81, 2861–2868. [Google Scholar] [CrossRef]
- Gosselin-Grenet, A.S.; Mottet-Osman, G.; Roux, L. Sendai virus particle production: Basic requirements and role of the SYWST motif present in HN cytoplasmic tail. Virology 2010, 405, 439–447. [Google Scholar] [CrossRef]
- Schmitt, A.P.; He, B.; Lamb, R.A. Involvement of the cytoplasmic domain of the hemagglutinin-neuraminidase protein in assembly of the paramyxovirus simian virus 5. J. Virol. 1999, 73, 8703–8712. [Google Scholar]
- Waning, D.L.; Schmitt, A.P.; Leser, G.P.; Lamb, R.A. Roles for the cytoplasmic tails of the fusion and hemagglutinin-neuraminidase proteins in budding of the paramyxovirus simian virus 5. J. Virol. 2002, 76, 9284–9297. [Google Scholar] [CrossRef]
- Batonick, M.; Wertz, G.W. Requirements for human respiratory syncytial virus glycoproteins in assembly and egress from infected cells. Adv. Virol. 2011, 2011, 343408:1–343408:8. [Google Scholar]
- Ghildyal, R.; Li, D.; Peroulis, I.; Shields, B.; Bardin, P.G.; Teng, M.N.; Collins, P.L.; Meanger, J.; Mills, J. Interaction between the respiratory syncytial virus G glycoprotein cytoplasmic domain and the matrix protein. J. Gen. Virol. 2005, 86, 1879–1884. [Google Scholar] [CrossRef]
- Tahara, M.; Takeda, M.; Yanagi, Y. Altered interaction of the matrix protein with the cytoplasmic tail of hemagglutinin modulates measles virus growth by affecting virus assembly and cell-cell fusion. J. Virol. 2007, 81, 6827–6836. [Google Scholar] [CrossRef]
- Stone, R.; Takimoto, T. Critical role of the fusion protein cytoplasmic tail sequence in parainfluenza virus assembly. PLoS One 2013, 8, e61281. [Google Scholar] [CrossRef]
- Shaikh, F.Y.; Cox, R.G.; Lifland, A.W.; Hotard, A.L.; Williams, J.V.; Moore, M.L.; Santangelo, P.J.; Crowe, J.E., Jr. A critical phenylalanine residue in the respiratory syncytial virus fusion protein cytoplasmic tail mediates assembly of internal viral proteins into viral filaments and particles. MBio 2012, 3, e00270-11. [Google Scholar]
- Moll, M.; Klenk, H.D.; Maisner, A. Importance of the cytoplasmic tails of the measles virus glycoproteins for fusogenic activity and the generation of recombinant measles viruses. J. Virol. 2002, 76, 7174–7186. [Google Scholar] [CrossRef]
- Cathomen, T.; Naim, H.Y.; Cattaneo, R. Measles viruses with altered envelope protein cytoplasmic tails gain cell fusion competence. J. Virol. 1998, 72, 1224–1234. [Google Scholar]
- Baviskar, P.S.; Hotard, A.L.; Moore, M.L.; Oomens, A.G. The respiratory syncytial virus fusion protein targets to the perimeter of inclusion bodies and facilitates filament formation by a cytoplasmic tail-dependent mechanism. J. Virol. 2013, 87, 10730–10741. [Google Scholar] [CrossRef]
- Jin, H.; Leser, G.P.; Zhang, J.; Lamb, R.A. Influenza virus hemagglutinin and neuraminidase cytoplasmic tails control particle shape. EMBO J. 1997, 16, 1236–1247. [Google Scholar] [CrossRef]
- Mebatsion, T.; Konig, M.; Conzelmann, K.K. Budding of rabies virus particles in the absence of the spike glycoprotein. Cell 1996, 84, 941–951. [Google Scholar] [CrossRef]
- Kim, S.H.; Yan, Y.; Samal, S.K. Role of the cytoplasmic tail amino acid sequences of Newcastle disease virus hemagglutinin-neuraminidase protein in virion incorporation, cell fusion, and pathogenicity. J. Virol. 2009, 83, 10250–10255. [Google Scholar] [CrossRef]
- Weise, C.; Erbar, S.; Lamp, B.; Vogt, C.; Diederich, S.; Maisner, A. Tyrosine residues in the cytoplasmic domains affect sorting and fusion activity of the Nipah virus glycoproteins in polarized epithelial cells. J. Virol. 2010, 84, 7634–7641. [Google Scholar] [CrossRef]
- Mattera, R.; Farias, G.G.; Mardones, G.A.; Bonifacino, J.S. Co-assembly of viral envelope glycoproteins regulates their polarized sorting in neurons. PLoS Pathog. 2014, 10, e1004107. [Google Scholar] [CrossRef]
- Samal, S.; Khattar, S.K.; Paldurai, A.; Palaniyandi, S.; Zhu, X.; Collins, P.L.; Samal, S.K. Mutations in the cytoplasmic domain of the Newcastle disease virus fusion protein confer hyperfusogenic phenotypes modulating viral replication and pathogenicity. J. Virol. 2013, 87, 10083–10093. [Google Scholar] [CrossRef]
- Takimoto, T.; Bousse, T.; Coronel, E.C.; Scroggs, R.A.; Portner, A. Cytoplasmic domain of Sendai virus HN protein contains a specific sequence required for its incorporation into virions. J. Virol. 1998, 72, 9747–9754. [Google Scholar]
- Popa, A.; Carter, J.R.; Smith, S.E.; Hellman, L.; Fried, M.G.; Dutch, R.E. Residues in the Hendra virus fusion protein transmembrane domain are critical for endocytic recycling. J. Virol. 2012, 86, 3014–3026. [Google Scholar] [CrossRef]
- Stone-Hulslander, J.; Morrison, T.G. Detection of an interaction between the HN and F proteins in Newcastle disease virus-infected cells. J. Virol. 1997, 71, 6287–6295. [Google Scholar]
- Plemper, R.K.; Hammond, A.L.; Cattaneo, R. Measles virus envelope glycoproteins hetero-oligomerize in the endoplasmic reticulum. J. Biol. Chem. 2001, 276, 44239–44246. [Google Scholar] [CrossRef]
- Tong, S.; Compans, R.W. Alternative mechanisms of interaction between homotypic and heterotypic parainfluenza virus HN and F proteins. J. Gen. Virol. 1999, 80, 107–115. [Google Scholar]
- Paterson, R.G.; Johnson, M.L.; Lamb, R.A. Paramyxovirus fusion (F) protein and hemagglutinin-neuraminidase (HN) protein interactions: Intracellular retention of F and HN does not affect transport of the homotypic HN or F protein. Virology 1997, 237, 1–9. [Google Scholar] [CrossRef]
- Whitman, S.D.; Smith, E.C.; Dutch, R.E. Differential rates of protein folding and cellular trafficking for the Hendra virus F and G proteins: Implications for F-G complex formation. J. Virol. 2009, 83, 8998–9001. [Google Scholar] [CrossRef]
- West, D.S.; Sheehan, M.S.; Segeleon, P.K.; Dutch, R.E. Role of the simian virus 5 fusion protein N-terminal coiled-coil domain in folding and promotion of membrane fusion. J. Virol. 2005, 79, 1543–1551. [Google Scholar] [CrossRef]
- Sergel-Germano, T.; McQuain, C.; Morrison, T. Mutations in the fusion peptide and heptad repeat regions of the Newcastle disease virus fusion protein block fusion. J. Virol. 1994, 68, 7654–7658. [Google Scholar]
- Sergel, T.A.; McGinnes, L.W.; Morrison, T.G. Mutations in the fusion peptide and adjacent heptad repeat inhibit folding or activity of the Newcastle disease virus fusion protein. J. Virol. 2001, 75, 7934–7943. [Google Scholar] [CrossRef]
- Parks, G.D.; Lamb, R.A. Defective assembly and intracellular transport of mutant paramyxovirus hemagglutinin-neuraminidase proteins containing altered cytoplasmic domains. J. Virol. 1990, 64, 3605–3616. [Google Scholar]
- Smith, E.C.; Smith, S.E.; Carter, J.R.; Webb, S.R.; Gibson, K.M.; Hellman, L.M.; Fried, M.G.; Dutch, R.E. Trimeric transmembrane domain interactions in paramyxovirus fusion proteins: Roles in protein folding, stability, and function. J. Biol. Chem. 2013, 288, 35726–35735. [Google Scholar] [CrossRef]
- Yao, Q.; Compans, R.W. Differences in the role of the cytoplasmic domain of human parainfluenza virus fusion proteins. J. Virol. 1995, 69, 7045–7053. [Google Scholar]
- Bagai, S.; Lamb, R.A. Individual roles of N-linked oligosaccharide chains in intracellular transport of the paramyxovirus SV5 fusion protein. Virology 1995, 209, 250–256. [Google Scholar] [CrossRef]
- McGinnes, L.; Sergel, T.; Reitter, J.; Morrison, T. Carbohydrate modifications of the NDV fusion protein heptad repeat domains influence maturation and fusion activity. Virology 2001, 283, 332–342. [Google Scholar] [CrossRef]
- Segawa, H.; Yamashita, T.; Kawakita, M.; Taira, H. Functional analysis of the individual oligosaccharide chains of Sendai virus fusion protein. J. Biochem. 2000, 128, 65–72. [Google Scholar] [CrossRef]
- Carter, J.R.; Pager, C.T.; Fowler, S.D.; Dutch, R.E. Role of N-linked glycosylation of the Hendra virus fusion protein. J. Virol. 2005, 79, 7922–7925. [Google Scholar] [CrossRef]
- Moll, M.; Kaufmann, A.; Maisner, A. Influence of N-glycans on processing and biological activity of the nipah virus fusion protein. J. Virol. 2004, 78, 7274–7278. [Google Scholar] [CrossRef]
- Sawatsky, B.; von Messling, V. Canine distemper viruses expressing a hemagglutinin without N-glycans lose virulence but retain immunosuppression. J. Virol. 2010, 84, 2753–2761. [Google Scholar] [CrossRef]
- Biering, S.B.; Huang, A.; Vu, A.T.; Robinson, L.R.; Bradel-Tretheway, B.; Choi, E.; Lee, B.; Aguilar, H.C. N-Glycans on the Nipah virus attachment glycoprotein modulate fusion and viral entry as they protect against antibody neutralization. J. Virol. 2012, 86, 11991–12002. [Google Scholar] [CrossRef]
- Collins, P.L.; Mottet, G. Post-translational processing and oligomerization of the fusion glycoprotein of human respiratory syncytial virus. J. Gen. Virol. 1991, 72, 3095–3101. [Google Scholar] [CrossRef]
- Zimmer, G.; Trotz, I.; Herrler, G. N-glycans of F protein differentially affect fusion activity of human respiratory syncytial virus. J. Virol. 2001, 75, 4744–4751. [Google Scholar] [CrossRef]
- Moll, M.; Klenk, H.D.; Herrler, G.; Maisner, A. A single amino acid change in the cytoplasmic domains of measles virus glycoproteins H and F alters targeting, endocytosis, and cell fusion in polarized Madin-Darby canine kidney cells. J. Biol. Chem. 2001, 276, 17887–17894. [Google Scholar]
- Leser, G.P.; Ector, K.J.; Ng, D.T.; Shaughnessy, M.A.; Lamb, R.A. The signal for clathrin-mediated endocytosis of the paramyxovirus SV5 HN protein resides at the transmembrane domain-ectodomain boundary region. Virology 1999, 262, 79–92. [Google Scholar] [CrossRef]
- Whitman, S.D.; Dutch, R.E. Surface density of the Hendra G protein modulates Hendra F protein-promoted membrane fusion: Role for Hendra G protein trafficking and degradation. Virology 2007, 363, 419–429. [Google Scholar] [CrossRef]
- Vogt, C.; Eickmann, M.; Diederich, S.; Moll, M.; Maisner, A. Endocytosis of the Nipah virus glycoproteins. J. Virol. 2005, 79, 3865–3872. [Google Scholar] [CrossRef]
- Leser, G.P.; Ector, K.J.; Lamb, R.A. The paramyxovirus simian virus 5 hemagglutinin-neuraminidase glycoprotein, but not the fusion glycoprotein, is internalized via coated pits and enters the endocytic pathway. Mol. Biol. Cell 1996, 7, 155–172. [Google Scholar] [CrossRef]
- Diederich, S.; Moll, M.; Klenk, H.D.; Maisner, A. The Nipah virus fusion protein is cleaved within the endosomal compartment. J. Biol. Chem. 2005, 280, 29899–29903. [Google Scholar] [CrossRef]
- Robach, J.G.; Lamb, R.A. Analysis of parainfluenza virus-5 hemagglutinin-neuraminidase protein mutants that are blocked in internalization and degradation. Virology 2010, 406, 189–201. [Google Scholar] [CrossRef]
- Spielhofer, P.; Bachi, T.; Fehr, T.; Christiansen, G.; Cattaneo, R.; Kaelin, K.; Billeter, M.A.; Naim, H.Y. Chimeric measles viruses with a foreign envelope. J. Virol. 1998, 72, 2150–2159. [Google Scholar]
- Roux, L.; Beffy, P.; Portner, A. Three variations in the cell surface expression of the haemagglutinin-neuraminidase glycoprotein of Sendai virus. J. Gen. Virol. 1985, 66, 987–1000. [Google Scholar] [CrossRef]
- Mellman, I.; Nelson, W.J. Coordinated protein sorting, targeting and distribution in polarized cells. Nat. Rev. Mol. Cell Biol. 2008, 9, 833–845. [Google Scholar] [CrossRef]
- Bonifacino, J.S.; Dell’Angelica, E.C. Molecular bases for the recognition of tyrosine-based sorting signals. J. Cell. Biol. 1999, 145, 923–926. [Google Scholar] [CrossRef]
- Runkler, N.; Dietzel, E.; Carsillo, M.; Niewiesk, S.; Maisner, A. Sorting signals in the measles virus wild-type glycoproteins differently influence virus spread in polarized epithelia and lymphocytes. J. Gen. Virol. 2009, 90, 2474–2482. [Google Scholar] [CrossRef]
- Moll, M.; Pfeuffer, J.; Klenk, H.D.; Niewiesk, S.; Maisner, A. Polarized glycoprotein targeting affects the spread of measles virus in vitro and in vivo. J. Gen. Virol. 2004, 85, 1019–1027. [Google Scholar] [CrossRef]
- Muth, T.R.; Caplan, M.J. Transport protein trafficking in polarized cells. Annu. Rev. Cell Dev. Biol. 2003, 19, 333–366. [Google Scholar] [CrossRef]
- Rodriguez-Boulan, E.; Kreitzer, G.; Musch, A. Organization of vesicular trafficking in epithelia. Nat. Rev. Mol. Cell Biol. 2005, 6, 233–247. [Google Scholar] [CrossRef]
- Cosson, P.; Perrin, J.; Bonifacino, J.S. Anchors aweigh: Protein localization and transport mediated by transmembrane domains. Trends Cell Biol. 2013, 23, 511–517. [Google Scholar] [CrossRef]
- Sanger, C.; Muhlberger, E.; Ryabchikova, E.; Kolesnikova, L.; Klenk, H.D.; Becker, S. Sorting of Marburg virus surface protein and virus release take place at opposite surfaces of infected polarized epithelial cells. J. Virol. 2001, 75, 1274–1283. [Google Scholar] [CrossRef]
- Barman, S.; Adhikary, L.; Kawaoka, Y.; Nayak, D.P. Influenza A virus hemagglutinin containing basolateral localization signal does not alter the apical budding of a recombinant influenza A virus in polarized MDCK cells. Virology 2003, 305, 138–152. [Google Scholar] [CrossRef]
- Zimmer, G.; Zimmer, K.P.; Trotz, I.; Herrler, G. Vesicular stomatitis virus glycoprotein does not determine the site of virus release in polarized epithelial cells. J. Virol. 2002, 76, 4103–4107. [Google Scholar] [CrossRef]
- Kolesnikova, L.; Ryabchikova, E.; Shestopalov, A.; Becker, S. Basolateral budding of Marburg virus: VP40 retargets viral glycoprotein GP to the basolateral surface. J. Infect. Dis. 2007, 196, S232–S236. [Google Scholar] [CrossRef]
- Maisner, A.; Klenk, H.; Herrler, G. Polarized budding of measles virus is not determined by viral surface glycoproteins. J. Virol. 1998, 72, 5276–5278. [Google Scholar]
- Naim, H.Y.; Ehler, E.; Billeter, M.A. Measles virus matrix protein specifies apical virus release and glycoprotein sorting in epithelial cells. EMBO J. 2000, 19, 3576–3585. [Google Scholar] [CrossRef]
- Lamp, B.; Dietzel, E.; Kolesnikova, L.; Sauerhering, L.; Erbar, S.; Weingartl, H.; Maisner, A. Nipah virus entry and egress from polarized epithelial cells. J. Virol. 2013, 87, 3143–3154. [Google Scholar] [CrossRef]
- Batonick, M.; Oomens, A.G.; Wertz, G.W. Human respiratory syncytial virus glycoproteins are not required for apical targeting and release from polarized epithelial cells. J. Virol. 2008, 82, 8664–8672. [Google Scholar] [CrossRef]
- Lin, S.; Naim, H.Y.; Rodriguez, A.C.; Roth, M.G. Mutations in the middle of the transmembrane domain reverse the polarity of transport of the influenza virus hemagglutinin in MDCK epithelial cells. J. Cell Biol. 1998, 142, 51–57. [Google Scholar] [CrossRef]
- Kundu, A.; Avalos, R.T.; Sanderson, C.M.; Nayak, D.P. Transmembrane domain of influenza virus neuraminidase, a type II protein, possesses an apical sorting signal in polarized MDCK cells. J. Virol. 1996, 70, 6508–6515. [Google Scholar]
- Brock, S.C.; Goldenring, J.R.; Crowe, J.E., Jr. Apical recycling systems regulate directional budding of respiratory syncytial virus from polarized epithelial cells. Proc. Natl. Acad. Sci. USA 2003, 100, 15143–15148. [Google Scholar] [CrossRef]
- Schuck, S.; Simons, K. Polarized sorting in epithelial cells: Raft clustering and the biogenesis of the apical membrane. J. Cell Sci. 2004, 117, 5955–5964. [Google Scholar] [CrossRef]
- Scheiffele, P.; Peranen, J.; Simons, K. N-glycans as apical sorting signals in epithelial cells. Nature 1995, 378, 96–98. [Google Scholar] [CrossRef]
- Yeaman, C.; Le Gall, A.H.; Baldwin, A.N.; Monlauzeur, L.; Le Bivic, A.; Rodriguez-Boulan, E. The O-glycosylated stalk domain is required for apical sorting of neurotrophin receptors in polarized MDCK cells. J. Cell Biol. 1997, 139, 929–940. [Google Scholar] [CrossRef]
- Potter, B.A.; Hughey, R.P.; Weisz, O.A. Role of N- and O-glycans in polarized biosynthetic sorting. Am. J. Physiol. Cell Physiol. 2006, 290, C1–C10. [Google Scholar]
- Sanderson, C.M.; McQueen, N.L.; Nayak, D.P. Sendai virus assembly: M protein binds to viral glycoproteins in transit through the secretory pathway. J. Virol. 1993, 67, 651–663. [Google Scholar]
- Santangelo, P.J.; Bao, G. Dynamics of filamentous viral RNPs prior to egress. Nucleic Acids Res. 2007, 35, 3602–3611. [Google Scholar] [CrossRef]
- Nakatsu, Y.; Ma, X.; Seki, F.; Suzuki, T.; Iwasaki, M.; Yanagi, Y.; Komase, K.; Takeda, M. Intracellular transport of the measles virus ribonucleoprotein complex is mediated by Rab11A-positive recycling endosomes and drives virus release from the apical membrane of polarized epithelial cells. J. Virol. 2013, 87, 4683–4693. [Google Scholar] [CrossRef]
- Chambers, R.; Takimoto, T. Trafficking of Sendai virus nucleocapsids is mediated by intracellular vesicles. PLoS One 2010, 5, e10994. [Google Scholar] [CrossRef]
- Utley, T.J.; Ducharme, N.A.; Varthakavi, V.; Shepherd, B.E.; Santangelo, P.J.; Lindquist, M.E.; Goldenring, J.R.; Crowe, J.E., Jr. Respiratory syncytial virus uses a Vps4-independent budding mechanism controlled by Rab11-FIP2. Proc. Natl. Acad. Sci. USA 2008, 105, 10209–10214. [Google Scholar] [CrossRef]
- Ghildyal, R.; Mills, J.; Murray, M.; Vardaxis, N.; Meanger, J. Respiratory syncytial virus matrix protein associates with nucleocapsids in infected cells. J. Gen. Virol. 2002, 83, 753–757. [Google Scholar]
- Qi, M.; Williams, J.A.; Chu, H.; Chen, X.; Wang, J.J.; Ding, L.; Akhirome, E.; Wen, X.; Lapierre, L.A.; Goldenring, J.R.; Spearman, P. Rab11-FIP1C and Rab14 direct plasma membrane sorting and particle incorporation of the HIV-1 envelope glycoprotein complex. PLoS Pathog. 2013, 9, e1003278. [Google Scholar] [CrossRef]
- Ghildyal, R.; Ho, A.; Dias, M.; Soegiyono, L.; Bardin, P.G.; Tran, K.C.; Teng, M.N.; Jans, D.A. The respiratory syncytial virus matrix protein possesses a Crm1-mediated nuclear export mechanism. J. Virol. 2009, 83, 5353–5362. [Google Scholar] [CrossRef]
- Wang, Y.E.; Park, A.; Lake, M.; Pentecost, M.; Torres, B.; Yun, T.E.; Wolf, M.C.; Holbrook, M.R.; Freiberg, A.N.; Lee, B. Ubiquitin-regulated nuclear-cytoplasmic trafficking of the Nipah virus matrix protein is important for viral budding. PLoS Pathog. 2010, 6, e1001186. [Google Scholar] [CrossRef]
- Peeples, M.E. Differential detergent treatment allows immunofluorescent localization of the Newcastle disease virus matrix protein within the nucleus of infected cells. Virology 1988, 162, 255–259. [Google Scholar] [CrossRef]
- Peeples, M.E.; Wang, C.; Gupta, K.C.; Coleman, N. Nuclear entry and nucleolar localization of the Newcastle disease virus (NDV) matrix protein occur early in infection and do not require other NDV proteins. J. Virol. 1992, 66, 3263–3269. [Google Scholar]
- Duan, Z.; Li, Q.; He, L.; Zhao, G.; Chen, J.; Hu, S.; Liu, X. Application of green fluorescent protein-labeled assay for the study of subcellular localization of Newcastle disease virus matrix protein. J. Virol. Methods 2013, 194, 118–122. [Google Scholar] [CrossRef]
- Duan, Z.; Song, Q.; Wang, Y.; He, L.; Chen, J.; Zhu, Y.; Hu, S.; Liu, X. Characterization of signal sequences determining the nuclear export of Newcastle disease virus matrix protein. Arch. Virol. 2013, 158, 2589–2595. [Google Scholar] [CrossRef]
- Duan, Z.; Chen, J.; Xu, H.; Zhu, J.; Li, Q.; He, L.; Liu, H.; Hu, S.; Liu, X. The nucleolar phosphoprotein B23 targets Newcastle disease virus matrix protein to the nucleoli and facilitates viral replication. Virology 2014, 452–453, 212–222. [Google Scholar] [CrossRef]
- Duan, Z.; Hu, Z.; Zhu, J.; Xu, H.; Chen, J.; Liu, H.; Hu, S.; Liu, X. Mutations in the FPIV motif of Newcastle disease virus matrix protein attenuate virus replication and reduce virus budding. Arch. Virol. 2014, 159, 1813–1819. [Google Scholar] [CrossRef]
- Yuan, H.; Puckett, S.; Lyles, D.S. Inhibition of host transcription by vesicular stomatitis virus involves a novel mechanism that is independent of phosphorylation of TATA-binding protein (TBP) or association of TBP with TBP-associated factor subunits. J. Virol. 2001, 75, 4453–4458. [Google Scholar] [CrossRef]
- Rajani, K.R.; Pettit Kneller, E.L.; McKenzie, M.O.; Horita, D.A.; Chou, J.W.; Lyles, D.S. Complexes of vesicular stomatitis virus matrix protein with host Rae1 and Nup98 involved in inhibition of host transcription. PLoS Pathog. 2012, 8, e1002929. [Google Scholar] [CrossRef]
- Bian, T.; Gibbs, J.D.; Orvell, C.; Imani, F. Respiratory syncytial virus matrix protein induces lung epithelial cell cycle arrest through a p53 dependent pathway. PLoS One 2012, 7, e38052. [Google Scholar]
- Lyles, D.S. Assembly and budding of negative-strand RNA viruses. Adv. Virus Res. 2013, 85, 57–90. [Google Scholar] [CrossRef]
- Schmitt, A.P.; Lamb, R.A. Influenza virus assembly and budding at the viral budozone. Adv. Virus Res. 2005, 64, 383–416. [Google Scholar] [CrossRef]
- Brown, D.A.; London, E. Structure and function of sphingolipid- and cholesterol-rich membrane rafts. J. Biol. Chem. 2000, 275, 17221–17224. [Google Scholar] [CrossRef]
- Chazal, N.; Gerlier, D. Virus entry, assembly, budding, and membrane rafts. Microbiol. Mol. Biol. Rev. 2003, 67, 226–237. [Google Scholar] [CrossRef]
- Takahashi, T.; Suzuki, T. Function of membrane rafts in viral lifecycles and host cellular response. Biochem. Res. Int. 2011, 2011, 245090. [Google Scholar]
- Rossman, J.S.; Lamb, R.A. Influenza virus assembly and budding. Virology 2011, 411, 229–236. [Google Scholar] [CrossRef]
- Campbell, S.M.; Crowe, S.M.; Mak, J. Lipid rafts and HIV-1: From viral entry to assembly of progeny virions. J. Clin. Virol. 2001, 22, 217–227. [Google Scholar] [CrossRef]
- Bavari, S.; Bosio, C.M.; Wiegand, E.; Ruthel, G.; Will, A.B.; Geisbert, T.W.; Hevey, M.; Schmaljohn, C.; Schmaljohn, A.; Aman, M.J. Lipid raft microdomains: A gateway for compartmentalized trafficking of Ebola and Marburg viruses. J. Exp. Med. 2002, 195, 593–602. [Google Scholar]
- Vincent, S.; Gerlier, D.; Manie, S.N. Measles virus assembly within membrane rafts. J. Virol. 2000, 74, 9911–9915. [Google Scholar] [CrossRef]
- Manie, S.N.; de Breyne, S.; Vincent, S.; Gerlier, D. Measles virus structural components are enriched into lipid raft microdomains: A potential cellular location for virus assembly. J. Virol. 2000, 74, 305–311. [Google Scholar] [CrossRef]
- Caballero, M.; Carabana, J.; Ortego, J.; Fernandez-Munoz, R.; Celma, M.L. Measles virus fusion protein is palmitoylated on transmembrane-intracytoplasmic cysteine residues which participate in cell fusion. J. Virol. 1998, 72, 8198–8204. [Google Scholar]
- Sanderson, C.M.; Avalos, R.; Kundu, A.; Nayak, D.P. Interaction of Sendai viral F, HN, and M proteins with host cytoskeletal and lipid components in Sendai virus-infected BHK cells. Virology 1995, 209, 701–707. [Google Scholar] [CrossRef]
- Laliberte, J.P.; McGinnes, L.W.; Morrison, T.G. Incorporation of functional HN-F glycoprotein-containing complexes into newcastle disease virus is dependent on cholesterol and membrane lipid raft integrity. J. Virol. 2007, 81, 10636–10648. [Google Scholar] [CrossRef]
- Oomens, A.G.; Bevis, K.P.; Wertz, G.W. The cytoplasmic tail of the human respiratory syncytial virus F protein plays critical roles in cellular localization of the F protein and infectious progeny production. J. Virol. 2006, 80, 10465–10477. [Google Scholar] [CrossRef]
- Gosselin-Grenet, A.S.; Marq, J.B.; Abrami, L.; Garcin, D.; Roux, L. Sendai virus budding in the course of an infection does not require Alix and VPS4A host factors. Virology 2007, 365, 101–112. [Google Scholar] [CrossRef]
- Kerviel, A.; Thomas, A.; Chaloin, L.; Favard, C.; Muriaux, D. Virus assembly and plasma membrane domains: Which came first? Virus Res. 2013, 171, 332–340. [Google Scholar] [CrossRef]
- Leser, G.P.; Lamb, R.A. Influenza virus assembly and budding in raft-derived microdomains: A quantitative analysis of the surface distribution of HA, NA and M2 proteins. Virology 2005, 342, 215–227. [Google Scholar]
- Chen, B.J.; Leser, G.P.; Jackson, D.; Lamb, R.A. The influenza virus M2 protein cytoplasmic tail interacts with the M1 protein and influences virus assembly at the site of virus budding. J. Virol. 2008, 82, 10059–10070. [Google Scholar] [CrossRef]
- Chang, T.H.; Segovia, J.; Sabbah, A.; Mgbemena, V.; Bose, S. Cholesterol-rich lipid rafts are required for release of infectious human respiratory syncytial virus particles. Virology 2012, 422, 205–213. [Google Scholar] [CrossRef]
- Gosselin-Grenet, A.S.; Mottet-Osman, G.; Roux, L. From assembly to virus particle budding: Pertinence of the detergent resistant membranes. Virology 2006, 344, 296–303. [Google Scholar] [CrossRef]
- Rossman, J.S.; Lamb, R.A. Viral membrane scission. Annu. Rev. Cell Dev. Biol. 2013, 29, 551–569. [Google Scholar] [CrossRef]
- McDonald, B.; Martin-Serrano, J. No strings attached: The ESCRT machinery in viral budding and cytokinesis. J. Cell Sci. 2009, 122, 2167–2177. [Google Scholar] [CrossRef]
- Schmitt, A.P.; Leser, G.P.; Morita, E.; Sundquist, W.I.; Lamb, R.A. Evidence for a new viral late-domain core sequence, FPIV, necessary for budding of a paramyxovirus. J. Virol. 2005, 79, 2988–2997. [Google Scholar] [CrossRef]
- Chen, B.J.; Lamb, R.A. Mechanisms for enveloped virus budding: Can some viruses do without an ESCRT? Virology 2008, 372, 221–232. [Google Scholar] [CrossRef]
- Patch, J.R.; Han, Z.; McCarthy, S.E.; Yan, L.; Wang, L.F.; Harty, R.N.; Broder, C.C. The YPLGVG sequence of the Nipah virus matrix protein is required for budding. Virol. J. 2008, 5, 137. [Google Scholar] [CrossRef]
- Salditt, A.; Koethe, S.; Pohl, C.; Harms, H.; Kolesnikova, L.; Becker, S.; Schneider-Schaulies, S. Measles virus M protein-driven particle production does not involve the endosomal sorting complex required for transport (ESCRT) system. J. Gen. Virol. 2010, 91, 1464–1472. [Google Scholar] [CrossRef]
- Weng, Y.; Lu, W.; Harmon, A.; Xiang, X.; Deng, Q.; Song, M.; Wang, D.; Yu, Q.; Li, F. The cellular endosomal sorting complex required for transport pathway is not involved in avian metapneumovirus budding in a virus-like-particle expression system. J. Gen. Virol. 2011, 92, 1205–1213. [Google Scholar] [CrossRef]
- Sabo, Y.; Ehrlich, M.; Bacharach, E. The conserved YAGL motif in human metapneumovirus is required for higher-order cellular assemblies of the matrix protein and for virion production. J. Virol. 2011, 85, 6594–6609. [Google Scholar] [CrossRef]
- Bruce, E.A.; Digard, P.; Stuart, A.D. The Rab11 pathway is required for influenza a virus budding and filament formation. J. Virol. 2010, 84, 5848–5859. [Google Scholar] [CrossRef]
- Acosta, D.; Adelman, J.; Affolder, T.; Akimoto, T.; Albrow, M.G.; Ambrose, D.; Amerio, S.; Amidei, D.; Anastassov, A.; Anikeev, K.; et al. Evidence for B0 s-->phiphi decay and measurements of branching ratio and A(CP) for B+ -->phiK+. Phys. Rev. Lett. 2005, 95, 031801. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
El Najjar, F.; Schmitt, A.P.; Dutch, R.E. Paramyxovirus Glycoprotein Incorporation, Assembly and Budding: A Three Way Dance for Infectious Particle Production. Viruses 2014, 6, 3019-3054. https://doi.org/10.3390/v6083019
El Najjar F, Schmitt AP, Dutch RE. Paramyxovirus Glycoprotein Incorporation, Assembly and Budding: A Three Way Dance for Infectious Particle Production. Viruses. 2014; 6(8):3019-3054. https://doi.org/10.3390/v6083019
Chicago/Turabian StyleEl Najjar, Farah, Anthony P. Schmitt, and Rebecca Ellis Dutch. 2014. "Paramyxovirus Glycoprotein Incorporation, Assembly and Budding: A Three Way Dance for Infectious Particle Production" Viruses 6, no. 8: 3019-3054. https://doi.org/10.3390/v6083019
APA StyleEl Najjar, F., Schmitt, A. P., & Dutch, R. E. (2014). Paramyxovirus Glycoprotein Incorporation, Assembly and Budding: A Three Way Dance for Infectious Particle Production. Viruses, 6(8), 3019-3054. https://doi.org/10.3390/v6083019