A Novel Adenovirus in Chinstrap Penguins (Pygoscelis antarctica) in Antarctica
Abstract
:1. Introduction
2. Results and Discussion
| Samples No. | Tissues tested | Positives | Designation | Accession No. |
|---|---|---|---|---|
| CSP09-1 | Lu, Li, K, Col, LN, Sp, Br | Lu | CSPAdVno1 | KC593379 |
| CSP09-2 | W-B | - | - | - |
| CSP10-1 | Lu, Li, K, Ht, Int, Tr | Lu, Li, K, Ht, Int | CSPAdVno2 | KC593380 |
| CSP10-2 | Li, K | CSPAdVno3 | KC593381 | |
| CSP10-3 | Lu, Li, K, Int | CSPAdVno4 | KC593382 | |
| CSP10-4 | - | - | - | |
| CSP10-5 | Lu, K, Int, Tr | CSPAdVno5 | KC593383 | |
| CPS10-6 | Lu, Li, K, Ht, Int, Tr | CSPAdVno6 | KC593384 | |
| CSP10-7 | Ht, Tr | CSPAdVno7 | KC593385 | |
| CSP10-8 | Lu, K, Int, Tr | CSPAdVno8 | KC593386 |
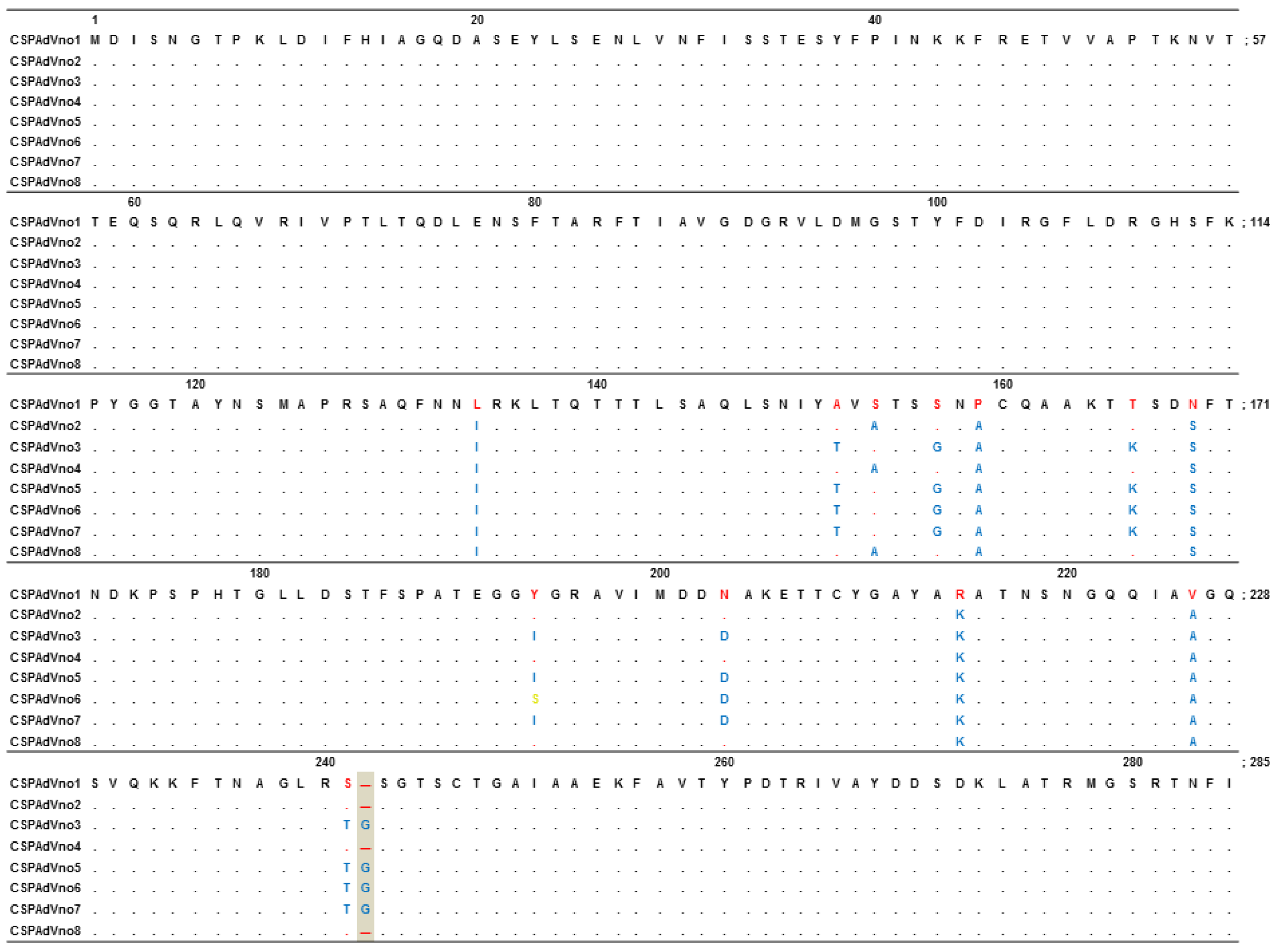

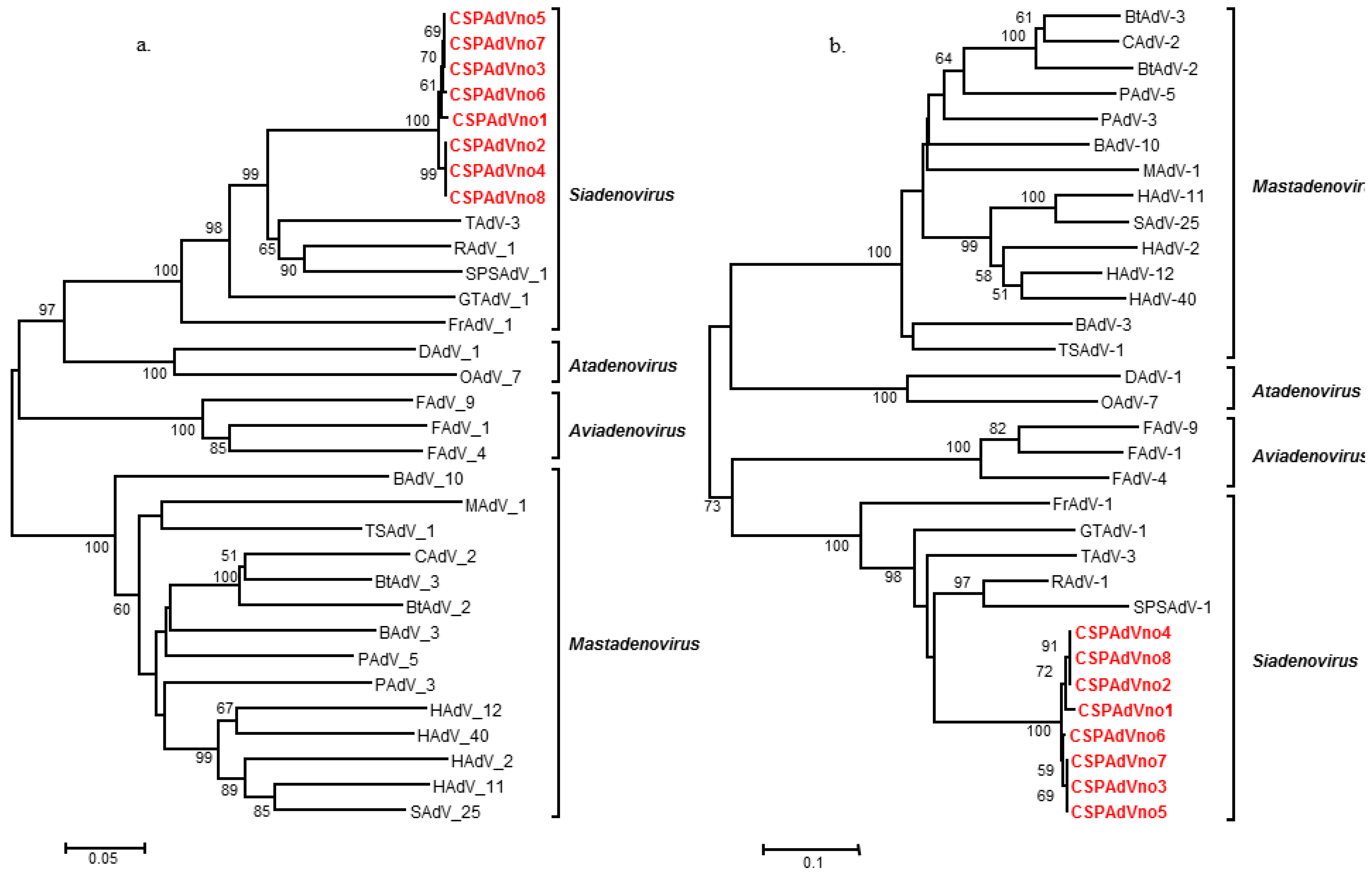
| Virus strain | Sequence identity (%) | |
|---|---|---|
| nucleotide | amino acid | |
| CSPAdVno2 | 98.7 | 97.9 |
| CSPAdVno3 | 97.5 | 96.1 |
| CSPAdVno4 | 98.7 | 97.9 |
| CSPAdVno5 | 97.5 | 96.1 |
| CSPAdVno6 | 97.8 | 96.1 |
| CSPAdVno7 | 97.5 | 96.1 |
| CSPAdVno8 | 98.7 | 97.9 |
| SPSAdV-1 | 71.8 | 65.5 |
| RAdV-1 | 71.0 | 70.0 |
| TAdV-3 | 71.4 | 67.6 |
| GTAdV-1 | 69.4 | 66.5 |
| FrAdV-1 | 61.0 | 61.6 |
3. Experimental Section
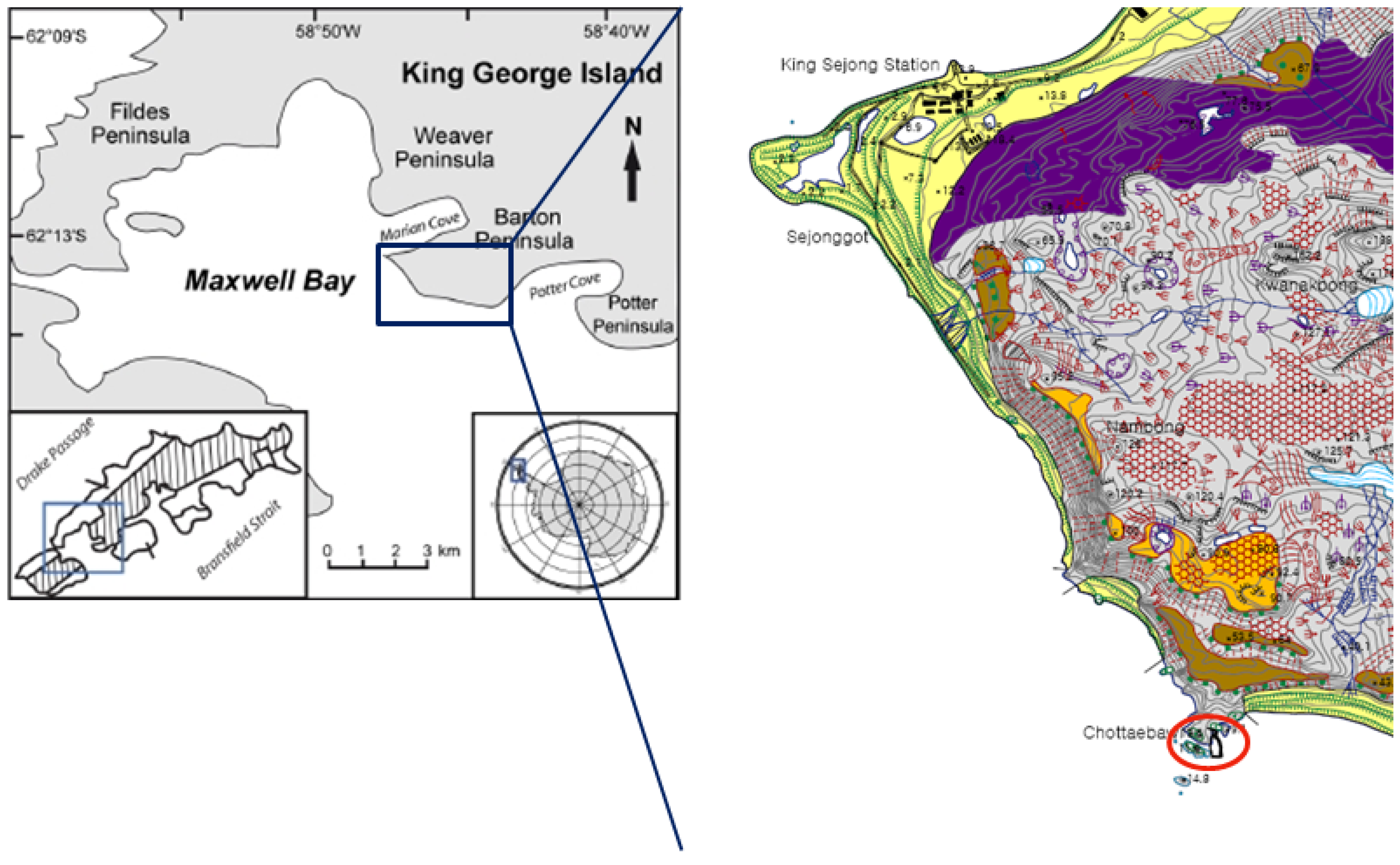
3.1. Sample Collection
3.2. DNA Extraction and PCR
3.3. Sequencing and Sequence Analysis
3.4. Isolation Attempts
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References and Notes
- Davison, A.J.; Benkö, M.; Harrach, B. Genetic content and evolution of adenoviruses. J. Gen. Virol. 2003, 84, 2895–2908. [Google Scholar] [CrossRef]
- David, M.K.; Peter, M.H.; Diane, E.G.; Robert, A.L.; Malcolm, A.M.; Bernard, R.; Stephen, E.S. Adenoviruses. In Fields Virology, 4th ed.; David, M.K., Peter, M.H., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2001; Volume 2, pp. 2404–2408. [Google Scholar]
- Harrach, B.; Benkö, M.; Both, G.W.; Brown, M.; Davison, A.J.; Echavarría, M.; Hess, M.; Jones, M.S.; Kajon, A.; Lehmkuhl, H.D.; et al. Family adenoviridae. In Virus Taxonomy: IXth Report of the International Committee on Taxonomy of Viruses; King, A.M.Q., Lefkowitz, E., Adams, M.J., Carstens, E.B., Eds.; Elsevier: New York, NY, USA, 2011; Volume 9, pp. 125–141. [Google Scholar]
- Kovács, G.M.; Davison, A.J.; Zakhartchouk, A.N.; Harrach, B. Analysis of the first complete genome sequence of an Old World monkey adenovirus reveals a lineage distinct from the six human adenovirus species. J. Gen. Virol. 2004, 85, 2799–2807. [Google Scholar] [CrossRef]
- Morrison, M.D.; Onions, D.E.; Nicolson, L. Complete DNA sequence of canine adenovirus type 1. J. Gen. Virol. 1997, 78, 873–878. [Google Scholar]
- Rusvai, M.; Harrach, B.; Bánrévi, A.; Evans, P.S.; Benkö, M. Identification and sequence analysis of the core protein genes of bovine adenovirus 2. Virus Res. 2000, 70, 25–30. [Google Scholar] [CrossRef]
- Aggarwal, N.; Mittal, S.K. Sequence analysis of procine adenovirus type 3 E1 region, pIX and pIVa2 genes, and two novel open reading frames. Intervirology 2000, 43, 6–12. [Google Scholar] [CrossRef]
- Klempa, B.; Krüger, D.H.; Auste, B.; Stanko, M.; Krawczyk, A.; Nickel, K.F.; Uberta, K.; Stang, A. A novel cardiotropic murine adenovirus representing a distinct species of mastadenoviruses. J. Virol. 2009, 83, 5749–5759. [Google Scholar] [CrossRef]
- Maeda, K.; Hondo, E.; Terakawa, J.; Kiso, Y.; Nakaichi, N.; Endoh, D.; Sakai, K.; Morikawa, S.; Mizutani, T. Isolation of novel adenovirus from fruit bat (Pteropus dasymallus yayeyamae). Emerg. Infect. Dis. 2008, 14, 347–249. [Google Scholar] [CrossRef]
- Sonntag, M.; Mühldorfer, K.; Speck, S.; Wibbelt, G.; Kurth, A. New adenovirus in bats, Germany. Emerg. Infect. Dis. 2009, 15, 2052–2055. [Google Scholar] [CrossRef]
- Schrenzel, M.; Oaks, J.L.; Rotstein, D.; Maalouf, G.; Snook, E.; Sandfort, C.; Rideout, B. Characterization of a new species of adenovirus in falcons. J. Clin. Microbiol. 2005, 43, 3402–3412. [Google Scholar] [CrossRef]
- Chiocca, S.; Kurzbauer, R.; Schaffner, G.; Baker, A.; Mautner, V.; Cotton, M. The complete DNA sequence and genomic organization of the avian adenovirus CELO. J. Virol. 1996, 70, 2939–2949. [Google Scholar]
- Dán, A.; Ruzsics, Z.; Russell, W.C.; Benkö, M.; Harrach, B. Analysis of the hexon gene sequence of bovine adenovirus type 4 provides further support for a new adenovirus genus (Atadenovirus). J. Gen. Virol. 1998, 79, 1453–1460. [Google Scholar]
- Farkas, S.L.; Harrach, B.; Benkö, M. Completion of the genome analysis of snake adenovirus type 1, a representative of the reptilian lineage within the novel genus Atadenovirus. Virus Res. 2008, 132, 132–139. [Google Scholar] [CrossRef]
- Hess, M.; Blöcker, H.; Brandt, P. The complete nucleotide sequence of the egg drop syndrome virus: An intermediate between mastadenoviruses and aviadenoviruses. Virology 1997, 238, 145–156. [Google Scholar] [CrossRef]
- Davison, A.J.; Wright, K.M.; Harrach, B. DNA sequence of frog adenovirus. J. Gen. Virol. 2000, 81, 2431–2439. [Google Scholar]
- Pitcovski, J.; Mualem, M.; Rei-Koren, Z.; Krispel, S.; Shmueli, E.; Peretz, Y.; Gutter, B.; Gallili, G.E.; Michael, A.; Goldberg, D. The complete DNA sequence and genome organization of the avian adenovirus, hemorrhagic enteritis virus. Virology 1998, 249, 307–315. [Google Scholar] [CrossRef]
- Kovács, E.R.; Benkö, M. Complete sequence of raptor adenovirus 1 confirms the characteristic genome organization of siadenoviruses. Infect. Genet. Evol. 2011, 5, 1058–1065. [Google Scholar] [CrossRef]
- Kovács, E.R.; Jánoska, M.; Harrach, B.; Benkö, M. Recognition and partial genome characterization by non-specific DNA amplification and PCR of a new siadenovirus species in a sample originating from Parus major, a great tit. J. Virol. Methods 2010, 163, 262–268. [Google Scholar] [CrossRef]
- Kovács, G.M.; LaPatra, S.E.; D’Halluin, J.C.; Benkö, M. Phylogenetic analysis of the hexon and protease genes of a fish adenovirus isolated from white sturgeon (Acipenser transmontanus) supports the proposal for a new adenovirus genus. Virus Res. 2003, 98, 27–34. [Google Scholar] [CrossRef]
- Park, Y.M.; Kim, J.H.; Gu, S.H.; Lee, S.Y.; Lee, M.G.; Kang, Y.K.; Kang, S.H.; Kim, H.J.; Song, J.W. Full genome analysis of a novel adenovirus from the South Polar skua (Catharacta maccormicki) in Antarctica. Virology 2012, 422, 144–150. [Google Scholar] [CrossRef]
- Yogui, G.T.; Sericano, J.L. Levels and pattern of polybrominated diphenyl ethers in eggs of Antarctic seabirds: Endemic versus migratory species. Environ. Pollut. 2009, 157, 975–980. [Google Scholar] [CrossRef]
- Raue, R.; Gerlach, H.; Müller, H. Phylogenetic analysis of the hexon loop 1 region of an adenovirus from psittacine birds supports the existence of a new psittacine adenovirus (PsAdV). Arch. Virol. 2005, 150, 1933–1943. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular Evolutionary Genetics Analysis using Maximum Likelihood, Evolutionaly Distance, and Maximum Parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef]
- PAUP*4.0: Phylogenetic Analysis Using Parsimony, version 4.0; Sinauer Associates, Inc: Sunderland, MA, USA, 2002.
- MEGA: Molucular Evolutionary Genetics Analysis. Available online: http://www.megasoftware.net/ (accessed on 8 March 2013).
- Nazerian, K.; Fadly, A.M. Propagation of virulent and avirulent turkey hemorrhagic enteritis virus in cell culture. Avian. Dis. 1982, 26, 816–827. [Google Scholar] [CrossRef]
- Zsivanovits, P.; Monks, D.J.; Forbes, N.A.; Ursu, K.; Raue, R.; Benkö, M. Presumptive identification of a novel adenovirus in a Harris hawk (Parabuteo unicinctus), a Bengal eagle owl (Bubo bengalensis), and a Verreaux’s eagle owl (Bubo lacteus). J. Avian Med. Sug. 2006, 20, 105–112. [Google Scholar] [CrossRef]
- Kovács, E.R.; Benkö, M. Confirmation of a novel siadenovirus species detected in raptors: partial sequence and phylogenetic analysis. Virus Res. 2009, 140, 64–70. [Google Scholar] [CrossRef]
- Barbosa, A.; Palacios, M.J. Health of Antarctic birds: A review of their parasites, pathogens and diseases. Polar Biol. 2009, 32, 1095–1115. [Google Scholar] [CrossRef]
- Pearce, D.A.; Wilson, W.H. Viruses in Antarctic ecosystems. Antarctic Sci. 2003, 15, 319–331. [Google Scholar] [CrossRef]
- Alexander, D.J.; Manvell, R.J.; Collins, M.S.; Brockman, S.J.; Westbury, H.A.; Morgan, I.; Austin, F.J. Characterization of paramyxoviruses isolated from penguins in Antarctica and sub-Antarctica during 1976–1979. Arch. Virol. 1989, 109, 135–143. [Google Scholar] [CrossRef]
- Austin, F.J.; Webster, R.G. Evidence of ortho and paramyxoviruses in fauna from Antarctica. J. Wild. Dis. 1993, 29, 568–571. [Google Scholar] [CrossRef]
- Thomazelli, L.M.; Araujo, J.; Oliveira, D.B.; Sanfilippo, L.; Ferreira, C.S.; Brentano, L.; Pelizari, V.H.; Nakayama, C.; Duarte, R.; Hurtado, R.; et al. Newcastle disease virus in penguins from King George Island on the Antarctic region. Vet. Microbiol. 2010, 146, 155–160. [Google Scholar]
- Wallensten, A.; Munster, V.J.; Osterhaus, A.D.M.E.; Waldenström, J.; Bonnedahl, J.; Broman, T.; Fouchier, R.A.M.; Olson, B. Mounting evidence for the presence of influenza A virus in the avifauna of the Antarctic region. Antarctic Sci. 2006, 18, 353–356. [Google Scholar]
- Chang, C.M.; Lebarbenchon, C.; Gauthier-Clerc, M.; Bohec, C.L.; Beaune, D.; Maho, Y.L.; Werf, S. Molecular surveillance for avian influenza A virus in king penguins (Aptenodytes patagonicus). Polar Biol. 2009, 32, 663–665. [Google Scholar] [CrossRef]
- Abad, F.X.; Busquets, N.; Sanchez, A.; Ryan, P.G.; Majö, N.; Gonalez-Solís, J. Serological and virological surveys of the influenza A viruses in Antarctic and sub-Antarctic penguins. Antarctic Sci. 2013, 25, 339–344. [Google Scholar] [CrossRef]
- Kerry, K.; Riddle, M.; Clarke, J. Diseases of Antarctic wildlife; 18; Australian Antartic Division: Channel Highway, Kingston, Australia, August 1998; pp. 89–91.
- Chown, S.L.; Lee, J.E.; Hughes, K.A.; Barrett, P.J.; Bergstrom, D.M.; Convey, P.; Cowan, D.A.; Crosbie, K.; Dyer, G.; Frenot, Y.; et al. Challenges to the future conservation of the Antarctica. Science 2012, 337, 158–159. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Lee, S.-Y.; Kim, J.-H.; Park, Y.M.; Shin, O.S.; Kim, H.; Choi, H.-G.; Song, J.-W. A Novel Adenovirus in Chinstrap Penguins (Pygoscelis antarctica) in Antarctica. Viruses 2014, 6, 2052-2061. https://doi.org/10.3390/v6052052
Lee S-Y, Kim J-H, Park YM, Shin OS, Kim H, Choi H-G, Song J-W. A Novel Adenovirus in Chinstrap Penguins (Pygoscelis antarctica) in Antarctica. Viruses. 2014; 6(5):2052-2061. https://doi.org/10.3390/v6052052
Chicago/Turabian StyleLee, Sook-Young, Jeong-Hoon Kim, Yon Mi Park, Ok Sarah Shin, Hankyeom Kim, Han-Gu Choi, and Jin-Won Song. 2014. "A Novel Adenovirus in Chinstrap Penguins (Pygoscelis antarctica) in Antarctica" Viruses 6, no. 5: 2052-2061. https://doi.org/10.3390/v6052052
APA StyleLee, S.-Y., Kim, J.-H., Park, Y. M., Shin, O. S., Kim, H., Choi, H.-G., & Song, J.-W. (2014). A Novel Adenovirus in Chinstrap Penguins (Pygoscelis antarctica) in Antarctica. Viruses, 6(5), 2052-2061. https://doi.org/10.3390/v6052052






