Genotypic Analysis of Kaposi’s Sarcoma-Associated Herpesvirus from Patients with Kaposi’s Sarcoma in Xinjiang, China
Abstract
:1. Introduction
2. Materials and Methods
2.1. Tissue Specimens
2.2. Preparation of DNA
2.3. PCR Amplification
2.4. DNA Sequencing
2.5. Phylogenetic Tree Analysis
2.6. Statistical Analysis
3. Results
3.1. KS Cases Characterization
| KS Groups | M/F Ratio | Age | ||||||
|---|---|---|---|---|---|---|---|---|
| Mean (Years) | SD (Years) | CV (%) | P25 | P75 | Median (Years) | Range (Years) | ||
| Classic KS (23) | 6.67 * | 54.39 | 13.94 | 25.64 | 40.00 | 65.50 | 53.00 § | 32–73 |
| AIDS-associated KS (5) | 4.00 * | 23.80 | 9.65 | 40.56 | 18.00 | 29.00 | 28.00 § | 10–34 |
| Total (28) | 6.00 | 48.93 | 17.74 | 36.25 | 37.00 | 63.00 | 50.50 | 10–73 |
3.2. ORF-K1 Amplification
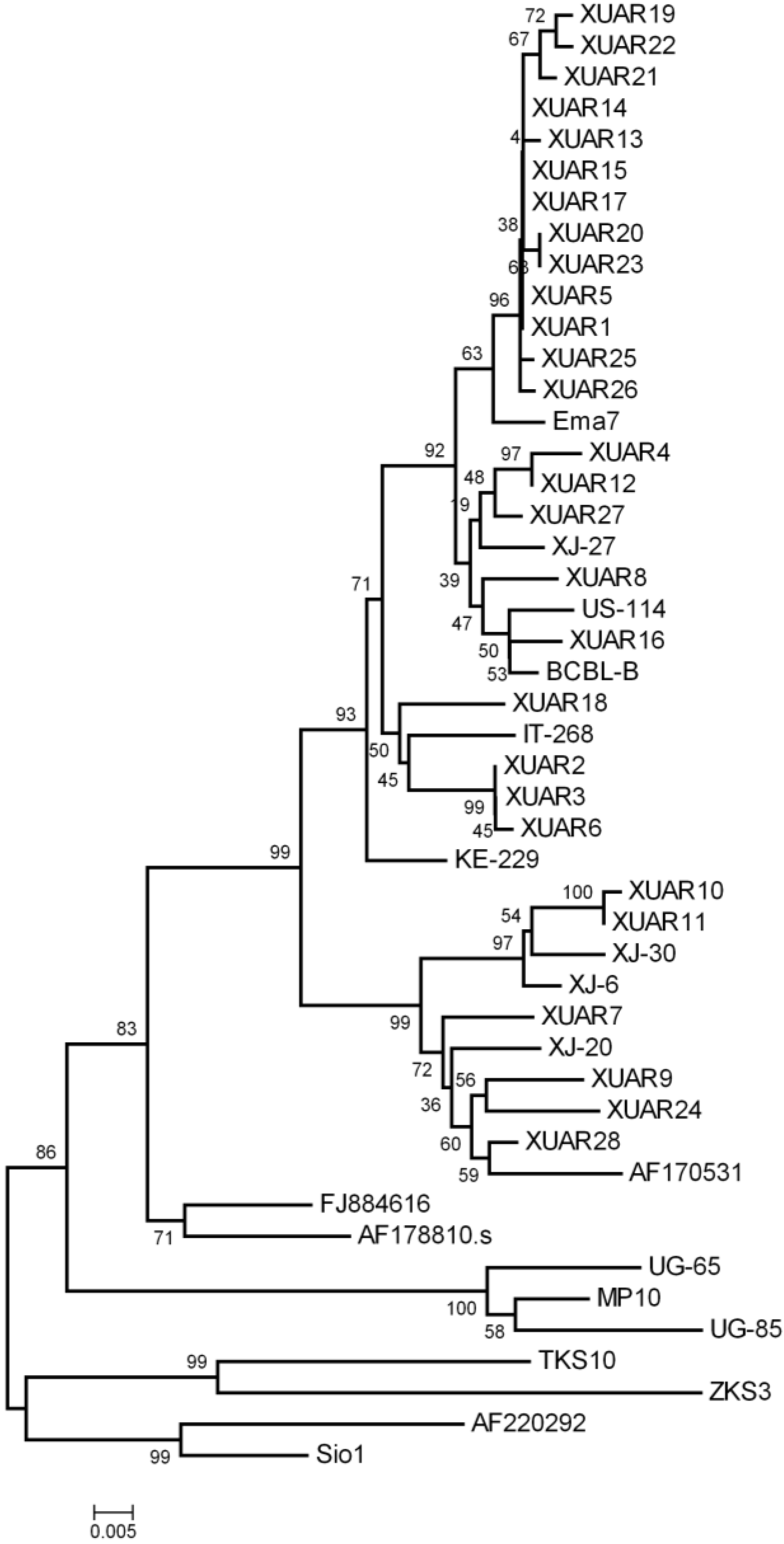
3.3. Phylogenetic Analysis
| KS Subtype | K1 Subtype * | ||
|---|---|---|---|
| Classic KS | |||
| A (19) | A1 | 2 | |
| A2 | 11 | ||
| A3 | 4 | ||
| A4 | 2 | ||
| C (4) | C2 | 2 | |
| C3 | 2 | ||
| AIDS-associated KS | |||
| A (4) | A1 | 1 | |
| A2 | 3 | ||
| C (1) | C3 | 1 | |
| KSHV Isolates | GenBank Accession Numbers | Ethnicity | Subtypes |
|---|---|---|---|
| XUAR1 | KM582680 | Uygur | A2 |
| XUAR2 | KM582681 | Kazakh | A3 |
| XUAR3 | KM582682 | Uygur | A3 |
| XUAR4 | KM582683 | Uygur | A1 |
| XUAR5 | KM582684 | Uygur | A2 |
| XUAR 6 | KM582685 | Uygur | A3 |
| XUAR7 | KM582687 | Uygur | C3 |
| XUAR8 | KM582688 | Kazakh | A4 |
| XUAR9 | KM582689 | Uygur | C3 |
| XUAR10 | KM582690 | Uygur | C2 |
| XUAR11 | KM582691 | Kazakh | C2 |
| XUAR12 | KM582692 | Kazakh | A1 |
| XUAR13 | KM582693 | Uygur | A2 |
| XUAR14 | KM582694 | Hui | A2 |
| XUAR15 | KM582695 | Uygur | A2 |
| XUAR16 | KM582696 | Han | A4 |
| XUAR17 | KM582697 | Uygur | A2 |
| XUAR18 | KM582698 | Kazakh | A3 |
| XUAR19 | KM582699 | Uygur | A2 |
| XUAR20 | KM582700 | Kazakh | A2 |
| XUAR21 | KM582701 | Kazakh | A2 |
| XUAR22 | KM582702 | Uygur | A2 |
| XUAR23 | KM582703 | Kazakh | A2 |
| XUAR24 | KM582704 | Uygur | C3 |
| XUAR25 | KM582705 | Uygur | A2 |
| XUAR26 | KM582706 | Uygur | A2 |
| XUAR27 | KM582707 | Uygur | A1 |
| XUAR28 | KM582708 | Uygur | C3 |
4. Discussion
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References and Notes
- Kaposi, M. Idiopathisches multiples pigmentsarkom der haut. Arch. Dermatol. Syph. 1872, 4, 265–273. [Google Scholar] [CrossRef]
- Beral, V.; Peterman, T.A.; Berkelman, R.L.; Jaffe, H.W. Kaposi’s sarcoma among persons with aids: A sexually transmitted infection? Lancet 1990, 335, 123–128. [Google Scholar] [CrossRef] [PubMed]
- Antman, K.; Chang, Y. Kaposi’s sarcoma. N. Engl. J. Med. 2000, 342, 1027–1038. [Google Scholar] [CrossRef] [PubMed]
- Friedman-Kien, A.E.; Saltzman, B.R. Clinical manifestations of classical, endemic african, and epidemic aids-associated kaposi’s sarcoma. J. Am. Acad. Dermatol. 1990, 22, 1237–1250. [Google Scholar] [CrossRef]
- Iscovich, J.; Boffetta, P.; Winkelmann, R.; Brennan, P.; Azizi, E. Classic kaposi’s sarcoma in jews living in israel, 1961–1989: A population-based incidence study. Aids 1998, 12, 2067–2072. [Google Scholar] [PubMed]
- Guo, W.X.; Antakly, T. Aids-related kaposi’s sarcoma: Evidence for direct stimulatory effect of glucocorticoid on cell proliferation. Am. J. Pathol. 1995, 146, 727–734. [Google Scholar] [PubMed]
- Fatahzadeh, M. Kaposi sarcoma: Review and medical management update. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. 2012, 113, 2–16. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.; Cesarman, E.; Pessin, M.S.; Lee, F.; Culpepper, J.; Knowles, D.M.; Moore, P.S. Identification of herpesvirus-like DNA sequences in aids-associated kaposi’s sarcoma. Science 1994, 266, 1865–1869. [Google Scholar] [CrossRef] [PubMed]
- Cesarman, E.; Chang, Y.; Moore, P.S.; Said, J.W.; Knowles, D.M. Kaposi’s sarcoma-associated herpesvirus-like DNA sequences in aids-related body-cavity-based lymphomas. N. Engl. J. Med. 1995, 332, 1186–1191. [Google Scholar] [CrossRef] [PubMed]
- Soulier, J.; Grollet, L.; Oksenhendler, E.; Cacoub, P.; Cazals-Hatem, D.; Babinet, P.; d’Agay, M.F.; Clauvel, J.P.; Raphael, M.; Degos, L.; et al. Kaposi’s sarcoma-associated herpesvirus-like DNA sequences in multicentric castleman’s disease. Blood 1995, 86, 1276–1280. [Google Scholar] [PubMed]
- Russo, J.J.; Bohenzky, R.A.; Chien, M.C.; Chen, J.; Yan, M.; Maddalena, D.; Parry, J.P.; Peruzzi, D.; Edelman, I.S.; Chang, Y.; et al. Nucleotide sequence of the kaposi sarcoma-associated herpesvirus (hhv8). Proc. Natl. Acad. Sci. USA 1996, 93, 14862–14867. [Google Scholar] [CrossRef] [PubMed]
- Kanno, T.; Sato, Y.; Nakamura, T.; Sakamoto, K.; Sata, T.; Katano, H. Genotypic and clinicopathological characterization of kaposi’s sarcoma-associated herpesvirus infection in japan. J. Med. Virol. 2010, 82, 400–406. [Google Scholar] [CrossRef] [PubMed]
- Kajumbula, H.; Wallace, R.G.; Zong, J.C.; Hokello, J.; Sussman, N.; Simms, S.; Rockwell, R.F.; Pozos, R.; Hayward, G.S.; Boto, W. Ugandan kaposi’s sarcoma-associated herpesvirus phylogeny: Evidence for cross-ethnic transmission of viral subtypes. Intervirology 2006, 49, 133–143. [Google Scholar] [CrossRef] [PubMed]
- Hayward, G.S.; Zong, J.C. Modern evolutionary history of the human kshv genome. Curr. Top. Microbiol. Immunol. 2007, 312, 1–42. [Google Scholar] [PubMed]
- Biggar, R.J.; Whitby, D.; Marshall, V.; Linhares, A.C.; Black, F. Human herpesvirus 8 in brazilian amerindians: A hyperendemic population with a new subtype. J. Infect. Dis. 2000, 181, 1562–1568. [Google Scholar] [CrossRef] [PubMed]
- Meng, Y.X.; Sata, T.; Stamey, F.R.; Voevodin, A.; Katano, H.; Koizumi, H.; Deleon, M.; de Cristofano, M.A.; Galimberti, R.; Pellett, P.E. Molecular characterization of strains of human herpesvirus 8 from japan, argentina and kuwait. J. Gen. Virol. 2001, 82, 499–506. [Google Scholar] [PubMed]
- Kasolo, F.C.; Monze, M.; Obel, N.; Anderson, R.A.; French, C.; Gompels, U.A. Sequence analyses of human herpesvirus-8 strains from both african human immunodeficiency virus-negative and -positive childhood endemic kaposi’s sarcoma show a close relationship with strains identified in febrile children and high variation in the k1 glycoprotein. J. Gen. Virol. 1998, 79, 3055–3065. [Google Scholar] [PubMed]
- Fu, B.; Sun, F.; Li, B.; Yang, L.; Zeng, Y.; Sun, X.; Xu, F.; Rayner, S.; Guadalupe, M.; Gao, S.J.; et al. Seroprevalence of kaposi’s sarcoma-associated herpesvirus and risk factors in xinjiang, china. J. Med. Virol. 2009, 81, 1422–1431. [Google Scholar] [CrossRef] [PubMed]
- Dilnur, P.; Katano, H.; Wang, Z.H.; Osakabe, Y.; Kudo, M.; Sata, T.; Ebihara, Y. Classic type of kaposi’s sarcoma and human herpesvirus 8 infection in xinjiang, china. Pathol. Int. 2001, 51, 845–852. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.Z.; Pu, X.M.; Wu, W.D.; Jin, Y.; Juhear, M.; Wu, X.J. Genotypic analysis on the orf-k1 gene of human herpesvirus 8 from patients with kaposi’s sarcoma in xinjiang, china. J. Genet. Genomics 2008, 35, 657–663. [Google Scholar] [CrossRef] [PubMed]
- Hayward, G.S. Kshv strains: The origins and global spread of the virus. Semin. Cancer Biol. 1999, 9, 187–199. [Google Scholar] [CrossRef] [PubMed]
- Cook, P.M.; Whitby, D.; Calabro, M.L.; Luppi, M.; Kakoola, D.N.; Hjalgrim, H.; Ariyoshi, K.; Ensoli, B.; Davison, A.J.; Schulz, T.F. Variability and evolution of kaposi’s sarcoma-associated herpesvirus in europe and africa. International collaborative group. Aids 1999, 13, 1165–1176. [Google Scholar] [PubMed]
- Zong, J.C.; Ciufo, D.M.; Alcendor, D.J.; Wan, X.; Nicholas, J.; Browning, P.J.; Rady, P.L.; Tyring, S.K.; Orenstein, J.M.; Rabkin, C.S.; et al. High-level variability in the orf-k1 membrane protein gene at the left end of the kaposi’s sarcoma-associated herpesvirus genome defines four major virus subtypes and multiple variants or clades in different human populations. J. Virol. 1999, 73, 4156–4170. [Google Scholar] [PubMed]
- White, T.; Hagen, M.; Gudza, I.; White, I.E.; Ndemera, B.; Gwanzura, L.; Borok, M.; Campbell, T.B. Genetic diversity of the kaposi’s sarcoma herpesvirus k1 protein in aids-ks in zimbabwe. J. Clin. Virol. 2008, 42, 165–171. [Google Scholar] [PubMed]
- Whitby, D.; Marshall, V.A.; Bagni, R.K.; Wang, C.D.; Gamache, C.J.; Guzman, J.R.; Kron, M.; Ebbesen, P.; Biggar, R.J. Genotypic characterization of kaposi’s sarcoma-associated herpesvirus in asymptomatic infected subjects from isolated populations. J. Gen. Virol. 2004, 85, 155–163. [Google Scholar] [CrossRef] [PubMed]
- Su, I.J.; Hsu, Y.S.; Chang, Y.C.; Wang, I.W. Herpesvirus-like DNA sequence in kaposi’s sarcoma from aids and non-aids patients in taiwan. Lancet 1995, 345, 722–723. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ouyang, X.; Zeng, Y.; Fu, B.; Wang, X.; Chen, W.; Fang, Y.; Luo, M.; Wang, L. Genotypic Analysis of Kaposi’s Sarcoma-Associated Herpesvirus from Patients with Kaposi’s Sarcoma in Xinjiang, China. Viruses 2014, 6, 4800-4810. https://doi.org/10.3390/v6124800
Ouyang X, Zeng Y, Fu B, Wang X, Chen W, Fang Y, Luo M, Wang L. Genotypic Analysis of Kaposi’s Sarcoma-Associated Herpesvirus from Patients with Kaposi’s Sarcoma in Xinjiang, China. Viruses. 2014; 6(12):4800-4810. https://doi.org/10.3390/v6124800
Chicago/Turabian StyleOuyang, Xinxing, Yan Zeng, Bishi Fu, Xiaowu Wang, Wei Chen, Yuan Fang, Minhua Luo, and Linding Wang. 2014. "Genotypic Analysis of Kaposi’s Sarcoma-Associated Herpesvirus from Patients with Kaposi’s Sarcoma in Xinjiang, China" Viruses 6, no. 12: 4800-4810. https://doi.org/10.3390/v6124800
APA StyleOuyang, X., Zeng, Y., Fu, B., Wang, X., Chen, W., Fang, Y., Luo, M., & Wang, L. (2014). Genotypic Analysis of Kaposi’s Sarcoma-Associated Herpesvirus from Patients with Kaposi’s Sarcoma in Xinjiang, China. Viruses, 6(12), 4800-4810. https://doi.org/10.3390/v6124800




