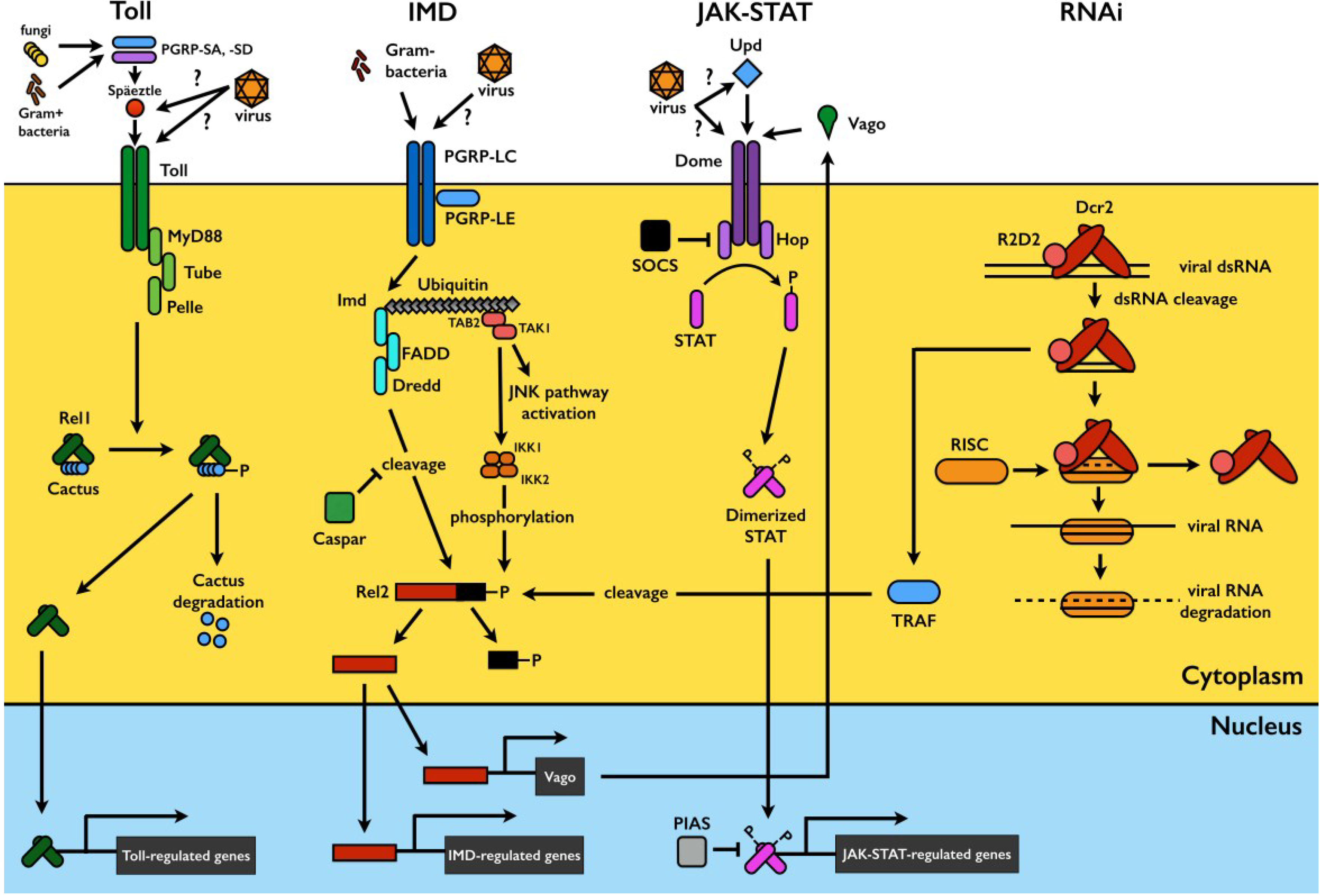
Mosquito Immunity against Arboviruses
Abstract
:1. Introduction
3. Arbovirus Interactions with Host Cell Processes and Host Factors
3.1. Apoptosis
3.2. Autophagy
3.3. The Vacuolar ATPase Complex
3.4. The Myeloid Differentiation 2-Related and Niemann-Pick Type C1 Proteins
4. Arbovirus Suppression of Insect Immune Responses
5. Natural Variation in Vector Competence for Arboviruses
6. Challenges and Future Directions
Acknowledgments
Author Contributions
Conflicts of Interest
References and Notes
- Bhatt, S.; Gething, P.W.; Brady, O.J.; Messina, J.P.; Farlow, A.W.; Moyes, C.L.; Drake, J.M.; Brownstein, J.S.; Hoen, A.G.; Sankoh, O.; et al. The global distribution and burden of dengue. Nature 2013, 496, 504–507. [Google Scholar] [CrossRef] [PubMed]
- Roehrig, J.T. West Nile Virus in the United States—A Historical Perspective. Viruses 2013, 5, 3088–3108. [Google Scholar] [CrossRef] [PubMed]
- Morrison, T.E. Re-emergence of chikungunya virus. J. Virol. 2014, 88, 11644–11647. [Google Scholar] [CrossRef] [PubMed]
- Lequime, S.; Lambrechts, L. Vertical transmission of arboviruses in mosquitoes: A historical perspective. Infect. Genet. Evol. 2014. [Google Scholar] [CrossRef]
- Salazar, M.I.; Richardson, J.H.; Sánchez-Vargas, I.; Olson, K.E.; Beaty, B.J. Dengue virus type 2: Replication and tropisms in orally infected Aedes aegypti mosquitoes. BMC Microbiol. 2007, 7, 9. [Google Scholar] [CrossRef] [PubMed]
- Gubler, D.J. Aedes aegypti and Aedes aegypti-borne disease control in the 1990s: Top down or bottom up. Am. J. Trop. Med. Hyg. 1989, 40, 571–578. [Google Scholar] [PubMed]
- Gratz, N.G.; Jany, W.C. What role for insecticides in vector control programs? Am. J. Trop. Med. Hyg. 1994, 50, 11–20. [Google Scholar] [PubMed]
- Nene, V.; Wortman, J.R.; Lawson, D.; Haas, B.; Kodira, C.; Tu, Z. (Jake); Loftus, B.; Xi, Z.; Megy, K.; Grabherr, M.; et al. Genome sequence of Aedes aegypti, a major arbovirus vector. Science 2007, 316, 1718–1723. [Google Scholar] [CrossRef] [PubMed]
- Arensburger, P.; Megy, K.; Waterhouse, R.M.; Abrudan, J.; Amedeo, P.; Antelo, B.; Bartholomay, L.; Bidwell, S.; Caler, E.; Camara, F.; et al. Sequencing of Culex quinquefasciatus Establishes a Platform for Mosquito Comparative Genomics. Science 2010, 330, 86–88. [Google Scholar] [CrossRef] [PubMed]
- Holt, R.A.; Subramanian, G.M.; Halpern, A.; Sutton, G.G.; Charlab, R.; Nusskern, D.R.; Wincker, P.; Clark, A.G.; Ribeiro, J.M.C.; Wides, R.; et al. The Genome Sequence of the Malaria Mosquito Anopheles gambiae. Science 2002, 298, 129–149. [Google Scholar] [CrossRef] [PubMed]
- Dimopoulos, G. Insect immunity and its implication in mosquito-malaria interactions. Cell Microbiol. 2003, 5, 3–14. [Google Scholar] [CrossRef] [PubMed]
- Lemaitre, B.; Hoffmann, J. The host defense of Drosophila melanogaster. Annu. Rev. Immunol. 2007, 25, 697–743. [Google Scholar] [CrossRef] [PubMed]
- Lemaitre, B.; Nicolas, E.; Michaut, L.; Reichhart, J.; Hoffmann, J. The dorsoventral regulatory gene cassette controls the potent antifungal response in Drosophila adults. Cell 1996, 86, 973–983. [Google Scholar] [CrossRef] [PubMed]
- Rutschmann, S.; Kilinc, A.; Ferrandon, D. Cutting edge: The Toll pathway is required for resistance to Gram-positive bacterial infections in Drosoph. J. Immunol. 2002, 168, 1542–1546. [Google Scholar] [CrossRef] [PubMed]
- Zambon, R.A.; Nandakumar, M.; Vakharia, V.N.; Wu, L.P. The Toll pathway is important for an antiviral response in Drosophila. Proc. Nat. Acad. Sci. USA 2005, 102, 7257–7262. [Google Scholar] [CrossRef] [PubMed]
- Michel, T.; Reichhart, J.-M.; Hoffmann, J.A.; Royet, J. Drosophila Toll is activated by Gram-positive bacteria through a circulating peptidoglycan recognition protein. Nature 2001, 414, 756–759. [Google Scholar] [CrossRef] [PubMed]
- Bischoff, V.; Vignal, C.; Boneca, I.G.; Michel, T.; Hoffmann, J.A.; Royet, J. Function of the Drosophila pattern-recognition receptor PGRP-SD in the detection of Gram-positive bacteria. Nat. Immunol. 2004, 5, 1175–1180. [Google Scholar] [CrossRef] [PubMed]
- DeLotto, Y.; DeLotto, R. Proteolytic processing of the Drosophila SpÃtzle protein by Easter generates a dimeric NGF-like molecule with ventralising activity. Mech. Dev. 1998, 72, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Weber, A.N.R.; Tauszig-Delamasure, S.; Hoffmann, J.A.; Lelievre, E.; Gascan, H.; Ray, K.P.; Morse, M.A.; Imler, J.-L.; Gay, N.J. Binding of the Drosophila cytokine Spatzle to Toll is direct and establishes signaling. Nat. Immunol. 2003, 4, 794–800. [Google Scholar] [CrossRef] [PubMed]
- Nicolas, E.; Reichhart, J.M.; Hoffmann, J.A.; Lemaitre, B. In vivo regulation of the Ikappa B homologue Cactus during the immune response of Drosophila. J. Biol. Chem. 1998, 273, 10463–10469. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, N.Q.; Grosshans, J.; Goltz, J.S.; Stein, D. Separable and redundant regulatory determinants in Cactus mediate its dorsal group dependent degradation. Development 2001, 128, 2963–2974. [Google Scholar] [PubMed]
- Hoffmann, J.A. The immune response of Drosophila. Nature 2003, 426, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Xi, Z.; Ramirez, J.L.; Dimopoulos, G. The Aedes aegypti Toll pathway controls dengue virus infection. PLoS Pathog. 2008, 4, e1000098. [Google Scholar] [CrossRef] [PubMed]
- Luplertlop, N.; Surasombatpattana, P.; Patramool, S.; Dumas, E.; Wasinpiyamongkol, L.; Saune, L.; Hamel, R.; Bernard, E.; Sereno, D.; Thomas, F.; et al. Induction of a peptide with activity against a broad spectrum of pathogens in the Aedes aegypti salivary gland, following infection with dengue virus. PLoS Pathog. 2011, 7, e1001252. [Google Scholar] [CrossRef] [PubMed]
- Sim, S.; Ramirez, J.L.; Dimopoulos, G. Dengue virus infection of the Aedes aegypti salivary gland and chemosensory apparatus induces genes that modulate infection and blood-feeding behavior. PLoS Pathog. 2012, 8, e1002631. [Google Scholar] [CrossRef] [PubMed]
- Ramirez, J.L.; Dimopoulos, G. The Toll immune signaling pathway control conserved anti-dengue defenses across diverse Ae. aegypti strains and against multiple dengue virus serotypes. Dev. Comp. Immunol. 2010, 34, 625–629. [Google Scholar] [CrossRef] [PubMed]
- Sim, S.; Jupatanakul, N.; Ramirez, J.L.; Kang, S.; Romero-Vivas, C.M.; Mohammed, H.; Dimopoulos, G. Transcriptomic Profiling of Diverse Aedes aegypti Strains Reveals Increased Basal-level Immune Activation in Dengue Virus-refractory Populations and Identifies Novel Virus-vector Molecular Interactions. PLoS Negl. Trop. Dis. 2013, 7, e2295. [Google Scholar] [CrossRef] [PubMed]
- Kambris, Z.; Cook, P.E.; Phuc, H.K.; Sinkins, S.P. Immune activation by life-shortening Wolbachia and reduced filarial competence in mosquitoes. Science 2009, 326, 134–136. [Google Scholar] [CrossRef] [PubMed]
- Kambris, Z.; Blagborough, A.M.; Pinto, S.B.; Blagrove, M.S.C.; Godfray, H.C.J.; Sinden, R.E.; Sinkins, S.P. Wolbachia Stimulates Immune Gene Expression and Inhibits Plasmodium Development in Anopheles gambiae. PLoS Pathog. 2010, 6, e1001143. [Google Scholar] [CrossRef] [PubMed]
- Moreira, L.A.; Iturbe-Ormaetxe, I.; Jeffery, J.A.; Lu, G.; Pyke, A.T.; Hedges, L.M.; Rocha, B.C.; Hall-Mendelin, S.; Day, A.; Riegler, M.; et al. A Wolbachia symbiont in Aedes aegypti limits infection with dengue, chikungunya, and Plasmodium. Cell 2009, 139, 1268–1278. [Google Scholar] [CrossRef] [PubMed]
- Bian, G.; Xu, Y.; Lu, P.; Xie, Y.; Xi, Z. The Endosymbiotic Bacterium Wolbachia Induces Resistance to Dengue Virus in Aedes aegypti. PLoS Pathog. 2010, 6, e1000833. [Google Scholar] [CrossRef] [PubMed]
- Iturbe-Ormaetxe, I.; Walker, T.; O’ Neill, S.L. Wolbachia and the biological control of mosquito-borne disease. EMBO Rep. 2011, 12, 508–518. [Google Scholar] [CrossRef] [PubMed]
- Pan, X.; Zhou, G.; Wu, J.; Bian, G.; Lu, P.; Raikhel, A.S.; Xi, Z. Wolbachia induces reactive oxygen species (ROS)-dependent activation of the Toll pathway to control dengue virus in the mosquito Aedes aegypti. Proc. Natl. Acad. Sci. USA 2012, 109, E23–E31. [Google Scholar] [CrossRef] [PubMed]
- Sanders, H.R.; Foy, B.D.; Evans, A.M.; Ross, L.S.; Beaty, B.J.; Olson, K.E.; Gill, S.S. Sindbis virus induces transport processes and alters expression of innate immunity pathway genes in the midgut of the disease vector, Aedes aegypti. Insect Biochem. Mol. Biol. 2005, 35, 1293–1307. [Google Scholar] [CrossRef] [PubMed]
- Bartholomay, L.C.; Waterhouse, R.M.; Mayhew, G.F.; Campbell, C.L.; Michel, K.; Zou, Z.; Ramirez, J.L.; Das, S.; Alvarez, K.; Arensburger, P.; et al. Pathogenomics of Culex quinquefasciatus and Meta-Analysis of Infection Responses to Diverse Pathogens. Science 2010, 330, 88–90. [Google Scholar] [CrossRef] [PubMed]
- Colpitts, T.M.; Cox, J.; Vanlandingham, D.L.; Feitosa, F.M.; Cheng, G.; Kurscheid, S.; Wang, P.; Krishnan, M.N.; Higgs, S.; Fikrig, E.; et al. Alterations in the Aedes aegypti Transcriptome during Infection with West Nile, Dengue and Yellow Fever Viruses. PLoS Pathog. 2011, 7, e1002189. [Google Scholar] [CrossRef] [PubMed]
- Georgel, P.; Naitza, S.; Kappler, C.; Ferrandon, D.; Zachary, D.; Swimmer, C.; Kopczynski, C.; Duyk, G.; Reichhart, J.-M.; Hoffmann, J.A.; et al. Drosophila Immune Deficiency (IMD) Is a Death Domain Protein that Activates Antibacterial Defense and Can Promote Apoptosis. Dev. Cell 2001, 1, 503–514. [Google Scholar] [CrossRef] [PubMed]
- Rutschmann, S.; Jung, A.C.; Zhou, R.; Silverman, N.; Hoffmann, J.A.; Ferrandon, D. Role of Drosophila IKKγ in a Toll-independent antibacterial immune response. Nat. Immunol. 2000, 1, 342–347. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Wu, L.P.; Anderson, K.V. The antibacterial arm of the Drosophila innate immune response requires an IκB kinase. Genes Dev. 2001, 15, 104–110. [Google Scholar] [CrossRef] [PubMed]
- Aggarwal, K.; Silverman, N. Positive and negative regulation of the Drosophila immune response. BMB Rep. 2008, 41, 267–277. [Google Scholar] [CrossRef] [PubMed]
- Choe, K.-M.; Lee, H.; Anderson, K.V. Drosophila peptidoglycan recognition protein LC (PGRP-LC) acts as a signal-transducing innate immune receptor. Proc. Natl. Acad. Sci. USA 2005, 102, 1122–1126. [Google Scholar] [CrossRef] [PubMed]
- Kleino, A.; Valanne, S.; Ulvila, J.; Kallio, J.; Myllymaki, H.; Enwald, H.; Stoven, S.; Poidevin, M.; Ueda, R.; Hultmark, D.; et al. Inhibitor of apoptosis 2 and TAK1-binding protein are components of the Drosophila Imd pathway. EMBO J. 2005, 24, 3423–3434. [Google Scholar] [CrossRef] [PubMed]
- Silverman, N.; Zhou, R.; Erlich, R.L.; Hunter, M.; Bernstein, E.; Schneider, D.; Maniatis, T. Immune Activation of NF-κB and JNK Requires Drosophila TAK. J. Biol. Chem. 2003, 278, 48928–48934. [Google Scholar] [CrossRef]
- Chen, W.; White, M.A.; Cobb, M.H. Stimulus-specific Requirements for MAP3 Kinases in Activating the JNK Pathway. J. Biol. Chem. 2002, 277, 49105–49110. [Google Scholar] [CrossRef] [PubMed]
- Sluss, H.K.; Han, Z.; Barrett, T.; Davis, R.J.; Ip, Y.T. A JNK signal transduction pathway that mediates morphogenesis and an immune response in Drosophila. Genes Dev. 1996, 10, 2745–2758. [Google Scholar] [CrossRef] [PubMed]
- Stöven, S.; Silverman, N.; Junell, A.; Hedengren-Olcott, M.; Erturk, D.; Engström, Y.; Maniatis, T.; Hultmark, D. Caspase-mediated processing of the Drosophila NF-κB factor Relish. PNAS 2003, 100, 5991–5996. [Google Scholar] [CrossRef] [PubMed]
- Leulier, F.; Rodriguez, A.; Khush, R.S.; Abrams, J.M.; Lemaitre, B. The Drosophila caspase Dredd is required to resist Gram-negative bacterial infection. EMBO Rep. 2000, 1, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Lee, J.H.; Lee, S.Y.; Kim, E.; Chung, J. Caspar, a suppressor of antibacterial immunity in Drosophila. PNAS 2006, 103, 16358–16363. [Google Scholar] [CrossRef] [PubMed]
- Meister, S.; Kanzok, S.M.; Zheng, X.; Luna, C.; Li, T.-R.; Hoa, N.T.; Clayton, J.R.; White, K.P.; Kafatos, F.C.; Christophides, G.K.; et al. Immune signaling pathways regulating bacterial and malaria parasite infection of the mosquito Anopheles gambiae. Proc. Natl. Acad. Sci. USA 2005, 102, 11420–11425. [Google Scholar] [CrossRef] [PubMed]
- Garver, L.S.; Dong, Y.; Dimopoulos, G. Caspar controls resistance to Plasmodium falciparum in diverse anopheline species. PLoS Pathog. 2009, 5, e1000335. [Google Scholar] [CrossRef] [PubMed]
- Garver, L.S.; Bahia, A.C.; Das, S.; Souza-Neto, J.A.; Shiao, J.; Dong, Y.; Dimopoulos, G. Anopheles Imd Pathway Factors and Effectors in Infection Intensity-Dependent Anti-Plasmodium Action. PLoS Pathog. 2012, 8, e1002737. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Das, S.; Cirimotich, C.; Souza-Neto, J.A.; McLean, K.J.; Dimopoulos, G. Engineered Anopheles immunity to Plasmodium infection. PLoS Pathog. 2011, 7, e1002458. [Google Scholar] [CrossRef] [PubMed]
- Costa, A.; Jan, E.; Sarnow, P.; Schneider, D. The Imd pathway is involved in antiviral immune responses in Drosophila. PLoS One 2009, 4, e7436. [Google Scholar] [CrossRef] [PubMed]
- Avadhanula, V.; Weasner, B.P.; Hardy, G.G.; Kumar, J.P.; Hardy, R.W. A novel system for the launch of alphavirus RNA synthesis reveals a role for the IMD pathway in arthropod antiviral response. PLoS Pathog. 2009, 5, e1000582. [Google Scholar] [CrossRef] [PubMed]
- Fragkoudis, R.; Chi, Y.; Siu, R.W.; Barry, G.; Attarzadeh-Yazdi, G.; Merits, A.; Nash, A.A.; Fazakerley, J.K.; Köhl, A. Semliki Forest virus strongly reduces mosquito host defence signaling. Insect Mol. Biol. 2008, 17, 647–656. [Google Scholar] [CrossRef] [PubMed]
- Fu, X.Y.; Schindler, C.; Improta, T.; Aebersold, R.; Darnell, J.E., Jr. The proteins of ISGF-3, the interferon alpha-induced transcriptional activator, define a gene family involved in signal transduction. Proc. Natl. Acad. Sci. USA 1992, 89, 7840–7843. [Google Scholar] [CrossRef] [PubMed]
- Schindler, C.; Fu, X.Y.; Improta, T.; Aebersold, R.; Darnell, J.E. Proteins of transcription factor ISGF-3: one gene encodes the 91-and 84-kDa ISGF-3 proteins that are activated by interferon alpha. Proc. Natl. Acad. Sci. USA 1992, 89, 7836–7839. [Google Scholar] [CrossRef] [PubMed]
- Dupuis, S.; Jouanguy, E.; Al-Hajjar, S.; Fieschi, C.; Al-Mohsen, I.Z.; Al-Jumaah, S.; Yang, K.; Chapgier, A.; Eidenschenk, C.; Eid, P.; et al. Impaired response to interferon-alpha/beta and lethal viral disease in human STAT1 deficiency. Nat. Genet. 2003, 33, 388–391. [Google Scholar] [CrossRef] [PubMed]
- Karst, S.M.; Wobus, C.E.; Lay, M.; Davidson, J.; Virgin, H.W. STAT1-dependent innate immunity to a Norwalk-like virus. Science 2003, 299, 1575–1578. [Google Scholar] [CrossRef] [PubMed]
- Arbouzova, N.I.; Zeidler, M.P. JAK/STAT signalling in Drosophila: Insights into conserved regulatory and cellular functions. Development 2006, 133, 2605–2616. [Google Scholar] [CrossRef] [PubMed]
- Hombria, J.C.-G.; Brown, S. The fertile field of Drosophila JAK/STAT Signalling. Curr. Biol. 2002, 12, R569–R575. [Google Scholar] [CrossRef] [PubMed]
- Barillas-Mury, C.; Han, Y.S.; Seeley, D.; Kafatos, F.C. Anopheles gambiae Ag-STAT, a new insect member of the STAT family, is activated in response to bacterial infection. EMBO J. 1999, 18, 959–967. [Google Scholar] [CrossRef] [PubMed]
- Dostert, C.; Jouanguy, E.; Irving, P.; Troxler, L.; Galiana-Arnoux, D.; Hetru, C.; Hoffmann, J.A.; Imler, J.-L. The Jak-STAT signaling pathway is required but not sufficient for the antiviral response of Drosophila. Nat. Immunol. 2005, 6, 946–953. [Google Scholar] [CrossRef] [PubMed]
- Souza-Neto, J.A.; Sim, S.; Dimopoulos, G. An evolutionary conserved function of the JAK-STAT pathway in anti-dengue defense. Proc. Natl. Acad. Sci. USA 2009, 106, 17841–17846. [Google Scholar] [CrossRef] [PubMed]
- Waldock, J.; Olson, K.E.; Christophides, G.K. Anopheles gambiae antiviral immune response to systemic O’nyong-nyong infection. PLoS Negl. Trop. Dis. 2012, 6, e1565. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.; Sim, C.; Byrd, B.D.; Collins, F.H.; Hong, Y.S. Ex vivo promoter analysis of antiviral heat shock cognate 70B gene in Anopheles gambiae. Virol. J. 2008, 5, 136. [Google Scholar] [CrossRef] [PubMed]
- Sim, C.; Hong, Y.S.; Tsetsarkin, K.A.; Vanlandingham, D.L.; Higgs, S.; Collins, F.H. Anopheles gambiae heat shock protein cognate 70B impedes o’nyong-nyong virus replication. BMC Genomics 2007, 8, 231. [Google Scholar] [CrossRef]
- Sim, C.; Hong, Y.S.; Vanlandingham, D.L.; Harker, B.W.; Christophides, G.K.; Kafatos, F.C.; Higgs, S.; Collins, F.H. Modulation of Anopheles gambiae gene expression in response to o’nyong-nyong virus infection. Insect Mol. Biol. 2005, 14, 475–481. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Vargas, I.; Travanty, E.A.; Keene, K.M.; Franz, A.W.; Beaty, B.J.; Blair, C.D.; Olson, K.E. RNA interference, arthropod-borne viruses, and mosquitoes. Virus Res. 2004, 102, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.-H.; Aliyari, R.; Li, W.-X.; Li, H.-W.; Kim, K.; Carthew, R.; Atkinson, P.; Ding, S.-W. RNA interference directs innate immunity against viruses in adult Drosophila. Science 2006, 312, 452–454. [Google Scholar] [CrossRef] [PubMed]
- Zambon, R.A.; Vakharia, V.N.; Wu, L.P. RNAi is an antiviral immune response against a dsRNA virus in Drosophila melanogaster. Cell. Microbiol. 2006, 8, 880–889. [Google Scholar] [CrossRef] [PubMed]
- Franz, A.W.E.; Sánchez-Vargas, I.; Adelman, Z.N.; Blair, C.D.; Beaty, B.J.; James, A.A.; Olson, K.E. Engineering RNA interference-based resistance to dengue virus type 2 in genetically modified Aedes aegypti. Proc. Natl. Acad. Sci. USA 2006, 103, 4198–4203. [Google Scholar] [CrossRef] [PubMed]
- Mathur, G.; Sanchez-Vargas, I.; Alvarez, D.; Olson, K.E.; Marinotti, O.; James, A.A. Transgene-mediated suppression of dengue viruses in the salivary glands of the yellow fever mosquito, Aedes aegypti. Insect Mol. Biol. 2010, 19, 753–763. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Vargas, I.; Scott, J.C.; Poole-Smith, B.K.; Franz, A.W.E.; Barbosa-Solomieu, V.; Wilusz, J.; Olson, K.E.; Blair, C.D. Dengue virus type 2 infections of Aedes aegypti are modulated by the mosquito’s RNA interference pathway. PLoS Pathog. 2009, 5, e1000299. [Google Scholar] [CrossRef] [PubMed]
- McFarlane, M.; Arias-Goeta, C.; Martin, E.; O’Hara, Z.; Lulla, A.; Mousson, L.; Rainey, S.M.; Misbah, S.; Schnettler, E.; Donald, C.L.; et al. Characterization of Aedes aegypti Innate-Immune Pathways that Limit Chikungunya Virus Replication. PLoS Negl. Trop. Dis. 2014, 8, e2994. [Google Scholar] [CrossRef] [PubMed]
- Campbell, C.L.; Keene, K.M.; Brackney, D.E.; Olson, K.E.; Blair, C.D.; Wilusz, J.; Foy, B.D. Aedes aegypti uses RNA interference in defense against Sindbis virus infection. BMC Microbiol. 2008, 8, 47. [Google Scholar] [CrossRef] [PubMed]
- Keene, K.M.; Foy, B.D.; Sanchez-Vargas, I.; Beaty, B.J.; Blair, C.D.; Olson, K.E. RNA interference acts as a natural antiviral response to Oʼnyong-nyong virus (Alphavirus; Togaviridae) infection of Anopheles gambiae. Proc. Natl. Acad. Sci. USA 2004, 101, 17240–17245. [Google Scholar] [CrossRef] [PubMed]
- Myles, K.M.; Wiley, M.R.; Morazzani, E.M.; Adelman, Z.N. Alphavirus-derived small RNAs modulate pathogenesis in disease vector mosquitoes. Proc. Natl. Acad. Sci. USA 2008, 105, 19938–19943. [Google Scholar] [CrossRef] [PubMed]
- Brackney, D.E.; Beane, J.E.; Ebel, G.D. RNAi targeting of West Nile virus in mosquito midguts promotes virus diversification. PLoS Pathog. 2009, 5, e1000502. [Google Scholar] [CrossRef] [PubMed]
- Scott, J.C.; Brackney, D.E.; Campbell, C.L.; Bondu-Hawkins, V.; Hjelle, B.; Ebel, G.D.; Olson, K.E.; Blair, C.D. Comparison of Dengue Virus Type 2-Specific Small RNAs from RNA Interference-Competent and –Incompetent Mosquito Cells. PLoS Negl. Trop. Dis. 2010, 4, e848. [Google Scholar] [CrossRef] [PubMed]
- Hess, A.M.; Prasad, A.N.; Ptitsyn, A.; Ebel, G.D.; Olson, K.E.; Barbacioru, C.; Monighetti, C.; Campbell, C.L. Small RNA profiling of Dengue virus-mosquito interactions implicates the PIWI RNA pathway in anti-viral defense. BMC Microbiol. 2011, 11, 45–45. [Google Scholar] [CrossRef] [PubMed]
- Morazzani, E.M.; Wiley, M.R.; Murreddu, M.G.; Adelman, Z.N.; Myles, K.M. Production of Virus-Derived Ping-Pong-Dependent piRNA-like Small RNAs in the Mosquito Soma. PLoS Pathog. 2012, 8, e1002470. [Google Scholar] [CrossRef] [PubMed]
- Schnettler, E.; Donald, C.L.; Human, S.; Watson, M.; Siu, R.W.C.; McFarlane, M.; Fazakerley, J.K.; Kohl, A.; Fragkoudis, R. Knockdown of piRNA pathway proteins results in enhanced Semliki Forest virus production in mosquito cells. J Gen. Virol. 2013, 94, 1680–1689. [Google Scholar] [CrossRef] [PubMed]
- Saleh, M.-C.; Tassetto, M.; van Rij, R.P.; Goic, B.; Gausson, V.; Berry, B.; Jacquier, C.; Antoniewski, C.; Andino, R. Antiviral immunity in Drosophila requires systemic RNA interference spread. Nature 2009, 458, 346–350. [Google Scholar] [CrossRef] [PubMed]
- Attarzadeh-Yazdi, G.; Fragkoudis, R.; Chi, Y.; Siu, R.W.C.; Ulper, L.; Barry, G.; Rodriguez-Andres, J.; Nash, A.A.; Bouloy, M.; Merits, A.; et al. Cell-to-cell spread of the RNA interference response suppresses Semliki Forest virus (SFV) infection of mosquito cell cultures and cannot be antagonized by SFV. J. Virol. 2009, 83, 5735–5748. [Google Scholar] [CrossRef] [PubMed]
- Goic, B.; Vodovar, N.; Mondotte, J.A.; Monot, C.; Frangeul, L.; Blanc, H.; Gausson, V.; Vera-Otarola, J.; Cristofari, G.; Saleh, M.-C.; et al. RNA-mediated interference and reverse transcription control the persistence of RNA viruses in the insect model Drosophila. Nat. Immunol. 2013, 14, 396–403. [Google Scholar] [CrossRef] [PubMed]
- Bonizzoni, M.; Britton, M.; Marinotti, O.; Dunn, W.A.; Fass, J.; James, A.A. Probing functional polymorphisms in the dengue vector, Aedes aegypti. BMC Genomics 2013, 14, 739. [Google Scholar] [CrossRef] [PubMed]
- Crochu, S.; Cook, S.; Attoui, H.; Charrel, R.N.; Chesse, R.D.; Belhouchet, M.; Lemasson, J.-J.; de Micco, P.; de Lamballerie, X. Sequences of flavivirus-related RNA viruses persist in DNA form integrated in the genome of Aedes spp. mosquitoes. J. Gen. Virol. 2004, 85, 1971–1980. [Google Scholar] [CrossRef] [PubMed]
- Fort, P.; Albertini, A.; Van-Hua, A.; Berthomieu, A.; Roche, S.; Delsuc, F.; Pasteur, N.; Capy, P.; Gaudin, Y.; Weill, M.; et al. Fossil Rhabdoviral Sequences Integrated into Arthropod Genomes: Ontogeny, Evolution, and Potential Functionality. Mol. Biol. Evol. 2012, 29, 381–390. [Google Scholar] [CrossRef] [PubMed]
- De Gregorio, E.; Spellman, P.T.; Tzou, P.; Rubin, G.M.; Lemaitre, B. The Toll and Imd pathways are the major regulators of the immune response in Drosophila. EMBO J. 2002, 21, 2568–2579. [Google Scholar] [CrossRef] [PubMed]
- Tanji, T.; Hu, X.; Weber, A.N.R.; Ip, Y.T. Toll and IMD pathways synergistically activate an innate immune response in Drosophila melanogaster. Mol. Cell. Biol. 2007, 27, 4578–4588. [Google Scholar] [CrossRef] [PubMed]
- Ramirez, J.L.; Souza-Neto, J.; Torres Cosme, R.; Rovira, J.; Ortiz, A.; Pascale, J.M.; Dimopoulos, G. Reciprocal tripartite interactions between the Aedes aegypti midgut microbiota, innate immune system and dengue virus influences vector competence. PLoS Negl. Trop. Dis. 2012, 6, e1561. [Google Scholar] [CrossRef] [PubMed]
- Deddouche, S.; Matt, N.; Budd, A.; Mueller, S.; Kemp, C.; Galiana-Arnoux, D.; Dostert, C.; Antoniewski, C.; Hoffmann, J.A.; Imler, J.-L.; et al. The DExD/H-box helicase Dicer-2 mediates the induction of antiviral activity in Drosophila. Nat. Immunol. 2008, 9, 1425–1432. [Google Scholar] [CrossRef] [PubMed]
- Paradkar, P.N.; Trinidad, L.; Voysey, R.; Duchemin, J.-B.; Walker, P.J. Secreted Vago restricts West Nile virus infection in Culex mosquito cells by activating the Jak-STAT pathway. PNAS 2012, 109, 18915–18920. [Google Scholar] [CrossRef] [PubMed]
- Paradkar, P.N.; Duchemin, J.-B.; Voysey, R.; Walker, P.J. Dicer-2-Dependent Activation of Culex Vago Occurs via the TRAF-Rel2 Signaling Pathway. PLoS Negl. Trop. Dis. 2014, 8, e2823. [Google Scholar] [CrossRef] [PubMed]
- Laosutthipong, C.; Kanthong, N.; Flegel, T.W. Novel, anionic, antiviral septapeptides from mosquito cells also protect monkey cells against dengue virus. Antivir. Res. 2013, 98, 449–456. [Google Scholar] [CrossRef] [PubMed]
- Kanthong, N.; Laosutthipong, C.; Flegel, T.W. Response to Dengue virus infections altered by cytokine-like substances from mosquito cell cultures. BMC Microbiol. 2010, 10, 290. [Google Scholar] [CrossRef] [PubMed]
- Fields, B.N.; Knipe, D.M.; Howley, P.M.; Griffin, D.E. Fields Virology, 4th ed.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2001. [Google Scholar]
- Acosta, E.G.; Castilla, V.; Damonte, E.B. Functional entry of dengue virus into Aedes albopictus mosquito cells is dependent on clathrin-mediated endocytosis. J. Gen. Virol. 2008, 89, 474–484. [Google Scholar] [CrossRef] [PubMed]
- Clyde, K.; Kyle, J.L.; Harris, E. Recent advances in deciphering viral and host determinants of dengue virus replication and pathogenesis. J. Virol. 2006, 80, 11418–11431. [Google Scholar] [CrossRef] [PubMed]
- Van der Schaar, H.M.; Rust, M.J.; Chen, C.; van der Ende-Metselaar, H.; Wilschut, J.; Zhuang, X.; Smit, J.M. Dissecting the cell entry pathway of dengue virus by single-particle tracking in living cells. PLoS Pathog. 2008, 4. [Google Scholar] [CrossRef]
- Smith, D.R. An update on mosquito cell expressed dengue virus receptor proteins. Insect Mol. Biol. 2012, 21, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Hardwick, J.M. Viral interference with apoptosis. Semin. Cell Dev. Biol. 1998, 9, 339–349. [Google Scholar] [CrossRef] [PubMed]
- Everett, H.; McFadden, G. Apoptosis: An innate immune response to virus infection. Trends Microbiol. 1999, 7, 160–165. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Behura, S.K.; Clem, R.J.; Schneemann, A.; Becnel, J.; Severson, D.W.; Zhou, L. P53-Mediated Rapid Induction of Apoptosis Conveys Resistance to Viral Infection in Drosophila melanogaster. PLoS Pathog. 2013, 9, e1003137. [Google Scholar] [CrossRef] [PubMed]
- Vaidyanathan, R.; Scott, T.W. Apoptosis in mosquito midgut epithelia associated with West Nile virus infection. Apoptosis 2006, 11, 1643–1651. [Google Scholar] [CrossRef] [PubMed]
- Girard, Y.A.; Schneider, B.S.; McGee, C.E.; Wen, J.; Han, V.C.; Popov, V.; Mason, P.W.; Higgs, S. Salivary gland morphology and virus transmission during long-term cytopathologic West Nile virus infection in Culex mosquitoes. Am. J. Trop. Med. Hyg. 2007, 76, 118–128. [Google Scholar] [PubMed]
- Ocampo, C.B.; Caicedo, P.A.; Jaramillo, G.; Ursic Bedoya, R.; Baron, O.; Serrato, I.M.; Cooper, D.M.; Lowenberger, C. Differential Expression of Apoptosis Related Genes in Selected Strains of Aedes aegypti with Different Susceptibilities to Dengue Virus. PLoS One 2013, 8, e61187. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Gort, T.; Boyle, D.L.; Clem, R.J. Effects of Manipulating Apoptosis on Sindbis Virus Infection of Aedes aegypti Mosquitoes. J. Virol. 2012, 86, 6546–6554. [Google Scholar] [CrossRef] [PubMed]
- Deretic, V.; Levine, B. Autophagy, Immunity, and Microbial Adaptations. Cell Host Microbe 2009, 5, 527–549. [Google Scholar] [CrossRef] [PubMed]
- Shelly, S.; Lukinova, N.; Bambina, S.; Berman, A.; Cherry, S. Autophagy Is an Essential Component of Drosophila Immunity against Vesicular Stomatitis Virus. Immunity 2009, 30, 588–598. [Google Scholar] [CrossRef] [PubMed]
- Moy, R.H.; Gold, B.; Molleston, J.M.; Schad, V.; Yanger, K.; Salzano, M.-V.; Yagi, Y.; Fitzgerald, K.A.; Stanger, B.Z.; Soldan, S.S.; et al. Antiviral Autophagy Restricts Rift Valley Fever Virus Infection and Is Conserved from Flies to Mammals. Immunity 2014, 40, 51–65. [Google Scholar] [CrossRef] [PubMed]
- Nakamoto, M.; Moy, R.H.; Xu, J.; Bambina, S.; Yasunaga, A.; Shelly, S.S.; Gold, B.; Cherry, S. Virus recognition by Toll-7 activates antiviral autophagy in Drosophila. Immunity 2012, 36, 658–667. [Google Scholar] [CrossRef] [PubMed]
- Patel, R.K.; Hardy, R.W. Role for the Phosphatidylinositol 3-Kinase-Akt-TOR Pathway during Sindbis Virus Replication in Arthropods. J. Virol. 2012, 86, 3595–3604. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.-R.; Lei, H.-Y.; Liu, M.-T.; Wang, J.-R.; Chen, S.-H.; Jiang-Shieh, Y.-F.; Lin, Y.-S.; Yeh, T.-M.; Liu, C.-C.; Liu, H.-S.; et al. Autophagic machinery activated by dengue virus enhances virus replication. Virology 2008, 374, 240–248. [Google Scholar] [CrossRef] [PubMed]
- Panyasrivanit, M.; Khakpoor, A.; Wikan, N.; Smith, D.R. Co-localization of constituents of the dengue virus translation and replication machinery with amphisomes. J. Gen. Virol. 2009, 90, 448–456. [Google Scholar] [CrossRef] [PubMed]
- Heaton, N.S.; Randall, G. Dengue Virus-Induced Autophagy Regulates Lipid Metabolism. Cell Host Microbe 2010, 8, 422–432. [Google Scholar] [CrossRef] [PubMed]
- Nishi, T.; Forgac, M. The vacuolar (H+)-ATPases—Nature’s most versatile proton pumps. Nat. Rev. Mol. Cell Biol. 2002, 3, 94–103. [Google Scholar] [CrossRef] [PubMed]
- Sessions, O.M.; Barrows, N.J.; Souza-Neto, J.A.; Robinson, T.J.; Hershey, C.L.; Rodgers, M.A.; Ramirez, J.L.; Dimopoulos, G.; Yang, P.L.; Pearson, J.L.; et al. Discovery of insect and human dengue virus host factors. Nature 2009, 458, 1047–1050. [Google Scholar] [CrossRef] [PubMed]
- Nawa, M. Effects of bafilomycin A1 on Japanese encephalitis virus in C6/36 mosquito cells. Arch. Virol. 1998, 143, 1555–1568. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.; Shields, A.R.; Jupatanakul, N.; Dimopoulos, G. Suppressing Dengue-2 Infection by Chemical Inhibition of Aedes aegypti Host Factors. PLoS Negl. Trop. Dis. 2014, 8, e3084. [Google Scholar] [CrossRef] [PubMed]
- Behura, S.K.; Gomez-Machorro, C.; Harker, B.W.; deBruyn, B.; Lovin, D.D.; Hemme, R.R.; Mori, A.; Romero-Severson, J.; Severson, D.W. Global Cross-Talk of Genes of the Mosquito Aedes aegypti in Response to Dengue Virus Infection. PLoS Negl. Trop. Dis. 2011, 5, e1385. [Google Scholar] [CrossRef] [PubMed]
- Horácková, J.; Rudenko, N.; Golovchenko, M.; Grubhoffer, L. Der-p2 (Dermatophagoides pteronyssinus) allergen-like protein from the hard tick Ixodes ricinus—A novel member of ML (MD-2-related lipid-recognition) domain protein family. Parasitology 2010, 137, 1139–1149. [Google Scholar] [CrossRef] [PubMed]
- Starostina, E.; Xu, A.; Lin, H.; Pikielny, C.W. A Drosophila Protein Family Implicated in Pheromone Perception Is Related to Tay-Sachs GM2-Activator Protein. J. Biol. Chem. 2009, 284, 585–594. [Google Scholar] [CrossRef] [PubMed]
- Garver, W.; Heidenreich, R. The Niemann-Pick C Proteins and Trafficking of Cholesterol through the Late Endosomal/Lysosomal System. Curr. Mol. Med. 2002, 2, 485–505. [Google Scholar] [CrossRef] [PubMed]
- Shimazu, R.; Akashi, S.; Ogata, H.; Nagai, Y.; Fukudome, K.; Miyake, K.; Kimoto, M. MD-2, a Molecule that Confers Lipopolysaccharide Responsiveness on Toll-like Receptor. J. Exp. Med. 1999, 189, 1777–1782. [Google Scholar] [CrossRef]
- Nagai, Y.; Akashi, S.; Nagafuku, M.; Ogata, M.; Iwakura, Y.; Akira, S.; Kitamura, T.; Kosugi, A.; Kimoto, M.; Miyake, K.; et al. Essential role of MD-2 in LPS responsiveness and TLR4 distribution. Nat. Immunol. 2002, 3, 667–672. [Google Scholar] [PubMed]
- Dong, Y.; Aguilar, R.; Xi, Z.; Warr, E.; Mongin, E.; Dimopoulos, G. Anopheles gambiae Immune Responses to Human and Rodent Plasmodium Parasite Species. PLoS Pathog. 2006, 2, e52. [Google Scholar] [CrossRef] [PubMed]
- Cote, M.; Misasi, J.; Ren, T.; Bruchez, A.; Lee, K.; Filone, C.M.; Hensley, L.; Li, Q.; Ory, D.; Chandran, K.; et al. Small molecule inhibitors reveal Niemann-Pick C1 is essential for Ebola virus infection. Nature 2011, 477, 344–348. [Google Scholar] [CrossRef] [PubMed]
- Carette, J.E.; Raaben, M.; Wong, A.C.; Herbert, A.S.; Obernosterer, G.; Mulherkar, N.; Kuehne, A.I.; Kranzusch, P.J.; Griffin, A.M.; Ruthel, G.; et al. Ebola virus entry requires the cholesterol transporter Niemann-Pick C. Nature 2011, 477, 340–343. [Google Scholar] [CrossRef] [PubMed]
- Jupatanakul, N.; Sim, S.; Dimopoulos, G. Aedes aegypti ML and Niemann-Pick type C family members are agonists of dengue virus infection. Dev. Comp. Immunol. 2014, 43, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Munoz-Jordan, J.L.; Laurent-Rolle, M.; Ashour, J.; Martínez-Sobrido, L.; Ashok, M.; Lipkin, I.W.; García-Sastre, A. Inhibition of alpha/beta interferon signaling by the NS4B protein of flaviviruses. J. Virol. 2005, 79, 8004–8013. [Google Scholar] [CrossRef] [PubMed]
- Munoz-Jordan, J.L.; Sánchez-Burgos, G.G.; Laurent-Rolle, M.; García-Sastre, A. Inhibition of interferon signaling by dengue virus. Proc. Natl. Acad. Sci. USA 2003, 100, 14333–14338. [Google Scholar] [CrossRef] [PubMed]
- Ashour, J.; Laurent-Rolle, M.; Shi, P.-Y.; García-Sastre, A. NS5 of dengue virus mediates STAT2 binding and degradation. J. Virol. 2009, 83, 5408–5418. [Google Scholar] [CrossRef] [PubMed]
- Lin, R.-J.; Chang, B.-L.; Yu, H.-P.; Liao, C.-L.; Lin, Y.-L. Blocking of interferon-induced Jak-Stat signaling by Japanese encephalitis virus NS5 through a protein tyrosine phosphatase-mediated mechanism. J. Virol. 2006, 80, 5908–5918. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.-W.; Cheng, C.-W.; Yang, T.-C.; Li, S.-W.; Cheng, M.-H.; Wan, L.; Lin, Y.-J.; Lai, C.-H.; Lin, W.-Y.; Kao, M.-C.; et al. Interferon antagonist function of Japanese encephalitis virus NS4A and its interaction with DEAD-box RNA helicase DDX. Virus Res. 2008, 137, 49–55. [Google Scholar] [CrossRef] [PubMed]
- Garmashova, N.; Gorchakov, R.; Frolova, E.; Frolov, I. Sindbis Virus Nonstructural Protein nsP2 Is Cytotoxic and Inhibits Cellular Transcription. J. Virol. 2006, 80, 5686–5696. [Google Scholar] [CrossRef] [PubMed]
- Garmashova, N.; Atasheva, S.; Kang, W.; Weaver, S.C.; Frolova, E.; Frolov, I. Analysis of Venezuelan Equine Encephalitis Virus Capsid Protein Function in the Inhibition of Cellular Transcription. J. Virol. 2007, 81, 13552–13565. [Google Scholar] [CrossRef] [PubMed]
- Breakwell, L.; Dosenovic, P.; Hedestam, G.B.K.; D’Amato, M.; Liljeström, P.; Fazakerley, J.; McInerney, G.M. Semliki Forest Virus Nonstructural Protein 2 Is Involved in Suppression of the Type I Interferon Response. J. Virol. 2007, 81, 8677–8684. [Google Scholar] [CrossRef] [PubMed]
- Sim, S.; Dimopoulos, G. Dengue Virus Inhibits Immune Responses in Aedes aegypti Cells. PLoS One 2010, 5, e10678. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.-C.; Chou, C.-M.; Hsu, Y.-L.; Lien, J.-C.; Wang, Y.-M.; Chen, S.-T.; Tsai, S.-C.; Hsiao, P.-W.; Huang, C.-J. Characterization of two mosquito STATs, AaSTAT and CtSTAT. J. Biol. Chem. 2004, 279, 3308–3317. [Google Scholar] [CrossRef] [PubMed]
- Aliyari, R.; Wu, Q.; Li, H.-W.; Wang, X.-H.; Li, F.; Green, L.D.; Han, C.S.; Li, W.-X.; Ding, S.-W. Mechanism of induction and suppression of antiviral immunity directed by virus-derived small RNAs in Drosoph. Cell Host Microbe 2008, 4, 387–397. [Google Scholar] [CrossRef] [PubMed]
- Nayak, A.; Berry, B.; Tassetto, M.; Kunitomi, M.; Acevedo, A.; Deng, C.; Krutchinsky, A.; Gross, J.; Antoniewski, C.; Andino, R.; et al. Cricket paralysis virus antagonizes Argonaute 2 to modulate antiviral defense in Drosophila. Nat. Struct. Mol. Biol. 2010, 17, 547–554. [Google Scholar] [CrossRef] [PubMed]
- Kakumani, P.K.; Ponia, S.S.; Rajgokul, K.S.; Sood, V.; Chinnappan, M.; Banerjea, A.C.; Medigeshi, G.; Malhotra, P.; Mukherjee, S.K.; Bhatnagar, R.K.; et al. Role of RNAi in dengue viral replication and identification of NS4B as a RNAi suppressor. J. Virol. 2013, 87, 8870–8883. [Google Scholar] [CrossRef] [PubMed]
- Cirimotich, C.M.; Scott, J.C.; Phillips, A.T.; Geiss, B.J.; Olson, K.E. Suppression of RNA interference increases alphavirus replication and virus-associated mortality in Aedes aegypti mosquitoes. BMC Microbiol. 2009, 9, 49. [Google Scholar] [CrossRef] [PubMed]
- Pijlman, G.P.; Funk, A.; Kondratieva, N.; Leung, J.; Torres, S.; van der Aa, L.; Liu, W.J.; Palmenberg, A.C.; Shi, P.-Y.; Hall, R.A.; et al. A Highly Structured, Nuclease-Resistant, Noncoding RNA Produced by Flaviviruses Is Required for Pathogenicity. Cell Host Microbe 2008, 4, 579–591. [Google Scholar] [CrossRef] [PubMed]
- Schnettler, E.; Sterken, M.G.; Leung, J.Y.; Metz, S.W.; Geertsema, C.; Goldbach, R.W.; Vlak, J.M.; Kohl, A.; Khromykh, A.A.; Pijlman, G.P.; et al. Noncoding Flavivirus RNA Displays RNA Interference Suppressor Activity in Insect and Mammalian Cells. J. Virol. 2012, 86, 13486–13500. [Google Scholar] [CrossRef] [PubMed]
- Schuessler, A.; Funk, A.; Lazear, H.M.; Cooper, D.A.; Torres, S.; Daffis, S.; Jha, B.K.; Kumagai, Y.; Takeuchi, O.; Hertzog, P.; et al. West Nile Virus Noncoding Subgenomic RNA Contributes to Viral Evasion of the Type I Interferon-Mediated Antiviral Response. J. Virol. 2012, 86, 5708–5718. [Google Scholar] [CrossRef] [PubMed]
- Chang, R.-Y.; Hsu, T.-W.; Chen, Y.-L.; Liu, S.-F.; Tsai, Y.-J.; Lin, Y.-T.; Chen, Y.-S.; Fan, Y.-H. Japanese encephalitis virus non-coding RNA inhibits activation of interferon by blocking nuclear translocation of interferon regulatory factor. Vet. Microbiol. 2013, 166, 11–21. [Google Scholar] [CrossRef] [PubMed]
- Bidet, K.; Dadlani, D.; Garcia-Blanco, M.A. G3BP1, G3BP2 and CAPRIN1 Are Required for Translation of Interferon Stimulated mRNAs and Are Targeted by a Dengue Virus Non-coding RNA. PLoS Pathog. 2014, 10, e1004242. [Google Scholar] [CrossRef] [PubMed]
- Campbell, C.L.; Harrison, T.; Hess, A.M.; Ebel, G.D. MicroRNA levels are modulated in Aedes aegypti after exposure to Dengue. Insect Mol. Biol. 2014, 23, 132–139. [Google Scholar] [CrossRef] [PubMed]
- Shrinet, J.; Jain, S.; Jain, J.; Bhatnagar, R.K.; Sunil, S. Next Generation Sequencing Reveals Regulation of Distinct Aedes microRNAs during Chikungunya Virus Development. PLoS Negl. Trop. Dis. 2014, 8, e2616. [Google Scholar] [CrossRef] [PubMed]
- Hussain, M.; Torres, S.; Schnettler, E.; Funk, A.; Grundhoff, A.; Pijlman, G.P.; Khromykh, A.A.; Asgari, S. West Nile virus encodes a microRNA-like small RNA in the 3? Untranslated region which up-regulates GATA4 mRNA and facilitates virus replication in mosquito cells. Nucleic Acids Res. 2012, 40, 2210–2223. [Google Scholar] [CrossRef] [PubMed]
- Bennett, K.E.; Olson, K.E.; Munoz Mde, L.; Fernandez-Salas, I.; Farfan-Ale, J.; Higgs, S.; Black, W.C., 4th; Beaty, B. Variation in vector competence for dengue 2 virus among 24 collections of Aedes aegypti from Mexico and the United States. Am. J. Trop. Med. Hyg. 2002, 67, 85–92. [Google Scholar] [PubMed]
- Lozano-Fuentes, S.; Fernández-Salas, I.; de Lourdes Muñoz, M.; Garcia-Rejon, J.; Olson, K.E.; Beaty, B.J.; Black, W.C., 4th. The neovolcanic axis is a barrier to gene flow among Aedes aegypti populations in mexico that differ in vector competence for dengue 2 virus. PLoS Negl. Trop. Dis. 2009, 3, e468. [Google Scholar] [CrossRef] [PubMed]
- Sylla, M.; Bosio, C.; Urdaneta-Marquez, L.; Ndiaye, M.; Black, W.C., 4th. Gene flow, subspecies composition, and dengue virus-2 susceptibility among Aedes aegypti collections in Senegal. PLoS Negl. Trop. Dis. 2009, 3, e408. [Google Scholar] [CrossRef] [PubMed]
- Black, W.C., 4th; Bennett, K.E.; Gorrochótegui-Escalante, N.; Barillas-Mury, C.V.; Fernández-Salas, I.; de Lourdes Muñoz, M.; Farfán-Alé, J.A.; Olson, K.E.; Beaty, B.J. Flavivirus susceptibility in Aedes aegypti. Arch. Med. Res. 2001, 33, 379–388. [Google Scholar]
- Bosio, C.; Beaty, B.; Black, W.C., 4th. Quantitative genetics of vector competence for dengue-2 virus in Aedes aegypti. Am. J. Trop. Med. Hyg. 1998, 59, 965–970. [Google Scholar] [PubMed]
- Bennett, K.E.; Beaty, B.J.; Black, W.C., 4th. Selection of D2S3, an Aedes aegypti (Diptera: Culicidae) strain with high oral susceptibility to dengue 2 virus and D2MEB, a strain with a midgut barrier to dengue 2 escape. J.Med. Entomol. 2005, 42, 110–119. [Google Scholar] [CrossRef] [PubMed]
- Bosio, C.F.; Fulton, R.E.; Salasek, M.L.; Beaty, B.J.; Black, W.C., 4th. Quantitative trait loci that control vector competence for dengue-2 virus in the mosquito Aedes aegypti. Genetics 2000, 156, 687–698. [Google Scholar] [PubMed]
- Gomez-Machorro, C.; Bennett, K.E.; del Lourdes Munoz, M.; Black, W.C., 4th. Quantitative trait loci affecting dengue midgut infection barriers in an advanced intercross line of Aedes aegypti. Insect Mol. Biol. 2004, 13, 637–648. [Google Scholar] [CrossRef] [PubMed]
- Bennett, K.E.; Flick, D.; Fleming, K.H.; Jochim, R.; Beaty, B.J.; Black, W.C., 4th. Quantitative trait loci that control dengue-2 virus dissemination in the mosquito Aedes aegypti. Genetics 2005, 170, 185–194. [Google Scholar] [CrossRef] [PubMed]
- Molina-Cruz, A.; Gupta, L.; Richardson, J.; Bennett, K.; Black, W.C., 4th; Barillas-Mury, C. Effect of mosquito midgut trypsin activity on dengue-2 virus infection and dissemination in Aedes aegypti. Am. J. Trop. Med. Hyg. 2005, 72, 631–637. [Google Scholar] [PubMed]
- Gorrochótegui-Escalante, N.; Lozano-Fuentes, S.; Bennett, K.E.; Molina-Cruz, A.; Beaty, B.J.; Black, W.C., 4th. Association mapping of segregating sites in the early trypsin gene and susceptibility to dengue-2 virus in the mosquito Aedes aegypti. Insect Biochem. Mol. Biol. 2005, 35, 771–788. [Google Scholar] [CrossRef] [PubMed]
- Lambrechts, L.; Chevillon, C.; Albright, R.; Thaisomboonsuk, B.; Richardson, J.; Jarman, R.; Scott, T. Genetic specificity and potential for local adaptation between dengue viruses and mosquito vectors. BMC Evol. Biol. 2009, 9, 160. [Google Scholar] [CrossRef] [PubMed]
- Fansiri, T.; Fontaine, A.; Diancourt, L.; Caro, V.; Thaisomboonsuk, B.; Richardson, J.H.; Jarman, R.G.; Ponlawat, A.; Lambrechts, L. Genetic Mapping of Specific Interactions between Aedes aegypti Mosquitoes and Dengue Viruses. PLoS Genet. 2013, 9, e1003621. [Google Scholar] [CrossRef] [PubMed]
- Lambrechts, L.; Quillery, E.; Noël, V.; Richardson, J.H.; Jarman, R.G.; Scott, T.W.; Chevillon, C. Specificity of resistance to dengue virus isolates is associated with genotypes of the mosquito antiviral gene Dicer. Proc. R. Soc. B 2013, 280. [Google Scholar] [CrossRef] [PubMed]
- Lambrechts, L. Quantitative genetics of Aedes aegypti vector competence for dengue viruses: Towards a new paradigm? Trends Parasitol. 2011, 27, 111–114. [Google Scholar] [CrossRef] [PubMed]
- Bonizzoni, M.; Dunn, W.A.; Campbell, C.L.; Olson, K.E.; Marinotti, O.; James, A.A. Strain Variation in the Transcriptome of the Dengue Fever Vector, Aedes aegypti. G3: Genes Genomes Genet. 2012, 2, 103–114. [Google Scholar]
- Cooper, T.F.; Rozen, D.E.; Lenski, R.E. Parallel changes in gene expression after 20,000 generations of evolution in Escherichia coli. Proc. Natl. Acad. Sci. USA 2003, 100, 1072–1077. [Google Scholar] [CrossRef] [PubMed]
- Aguilar, R.; Das, S.; Dong, Y.; Dimopoulos, G. Continuous exposure to Plasmodium results in decreased susceptibility and transcriptomic divergence of the Anopheles gambiae immune system. BMC Genomics 2007, 8, 451. [Google Scholar] [CrossRef] [PubMed]
- Ranz, J.M.; Castillo-Davis, C.I.; Meiklejohn, C.D.; Hartl, D.L. Sex-Dependent Gene Expression and Evolution of the Drosophila Transcriptome. Science 2003, 300, 1742–1745. [Google Scholar] [CrossRef] [PubMed]
- Aguilar, R.; Simard, F.; Kamdem, C.; Shields, T.; Glass, G.E.; Garver, L.S.; Dimopoulos, G. Genome-wide analysis of transcriptomic divergence between laboratory colony and field Anopheles gambiae mosquitoes of the M and S molecular forms. Insect Mol. Biol. 2010, 19, 695–705. [Google Scholar] [CrossRef] [PubMed]
- Rifkin, S.A.; Houle, D.; Kim, J.; White, K.P. A mutation accumulation assay reveals a broad capacity for rapid evolution of gene expression. Nature 2005, 438, 220–223. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Manfredini, F.; Dimopoulos, G. Implication of the mosquito midgut microbiota in the defense against malaria parasites. PLoS Pathog. 2009, 5, e1000423. [Google Scholar] [CrossRef] [PubMed]
- Jupatanakul, N.; Sim, S.; Dimopoulos, G. The insect microbiome modulates vector competence for arboviruses. Viruses 2014, 6, 4294–4313. [Google Scholar] [CrossRef] [PubMed]
- Bian, G.; Shin, S.W.; Cheon, H.M.; Kokoza, V.; Raikhel, A.S. Transgenic alteration of Toll immune pathway in the female mosquito Aedes aegypti. Proc. Natl. Acad. Sci. USA 2005, 102, 13568–13573. [Google Scholar] [CrossRef] [PubMed]
- Antonova, Y.; Alvarez, K.S.; Kim, Y.J.; Kokoza, V.; Raikhel, A.S. The role of NF-kappaB factor REL2 in the Aedes aegypti immune response. Insect Biochem. Mol. Biol. 2009, 39, 303–314. [Google Scholar] [CrossRef] [PubMed]
- Riehle, M.; Moreira, C.; Lampe, D.; Lauzon, C.; Jacobslorena, M. Using bacteria to express and display anti-Plasmodium molecules in the mosquito midgut. Int. J. Parasitol. 2007, 37, 595–603. [Google Scholar] [CrossRef] [PubMed]
- Bisi, D.C.; Lampe, D.J. Secretion of Anti-Plasmodium Effector Proteins from a Natural Pantoea agglomerans Isolate by Using PelB and HlyA Secretion Signals. Appl. Environ. Microbiol. 2011, 77, 4669–4675. [Google Scholar] [CrossRef] [PubMed]
- Fang, W.; Vega-Rodríguez, J.; Ghosh, A.K.; Jacobs-Lorena, M.; Kang, A.; St. Leger, R.J. Development of Transgenic Fungi That Kill Human Malaria Parasites in Mosquitoes. Science 2011, 331, 1074–1077. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sim, S.; Jupatanakul, N.; Dimopoulos, G. Mosquito Immunity against Arboviruses. Viruses 2014, 6, 4479-4504. https://doi.org/10.3390/v6114479
Sim S, Jupatanakul N, Dimopoulos G. Mosquito Immunity against Arboviruses. Viruses. 2014; 6(11):4479-4504. https://doi.org/10.3390/v6114479
Chicago/Turabian StyleSim, Shuzhen, Natapong Jupatanakul, and George Dimopoulos. 2014. "Mosquito Immunity against Arboviruses" Viruses 6, no. 11: 4479-4504. https://doi.org/10.3390/v6114479
APA StyleSim, S., Jupatanakul, N., & Dimopoulos, G. (2014). Mosquito Immunity against Arboviruses. Viruses, 6(11), 4479-4504. https://doi.org/10.3390/v6114479