Emerging Viruses in the Felidae: Shifting Paradigms
Abstract
:1. Introduction
| Feline Virus | Human Homologue |
|---|---|
| Feline Leukemia Virus (FeLV) [5] | Human T-Cell Leukemia Virus (HTLV) [6] |
| Feline Immunodeficiency Virus (FIV) [7] | Human Immunodeficiency Virus (HIV-AIDS) [8] |
| Feline Coronavirus (FCoV) [9] | SARS-Coronavirus (Severe acute respiratory syndrome) [10] |
| Feline Sarcoma Virus (FSV) [11] | ~20 Human Oncogenes [12] |
| Avian H5N1 Influenza [13] | Avian H5N1 Influenza [14] |
| Feline Herpes Virus (FHV) [15] | Cytomegalovirus (CMV-retinitis) |
| Feline Foamy Virus (FFV) [16] | Human Foamy Virus (No pathology) [17] |
| Feline Calicivirus (FCV) [18] | Human Calicivirus (Diarrhea, vomiting) [19] |
| Feline Parvovirus (FPV) [20] | Human B19 Parvovirus (Fifth disease) [21] |
| Feline Morbillivirus (CDV) [22] | Human Morbillivirus (Measles) [23] |
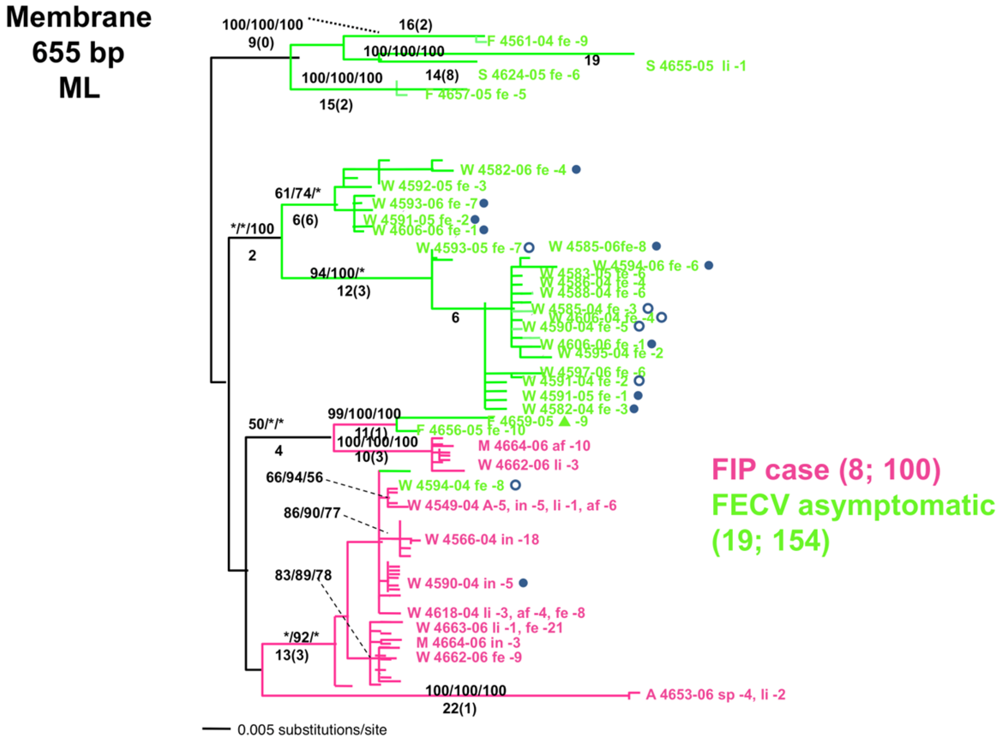
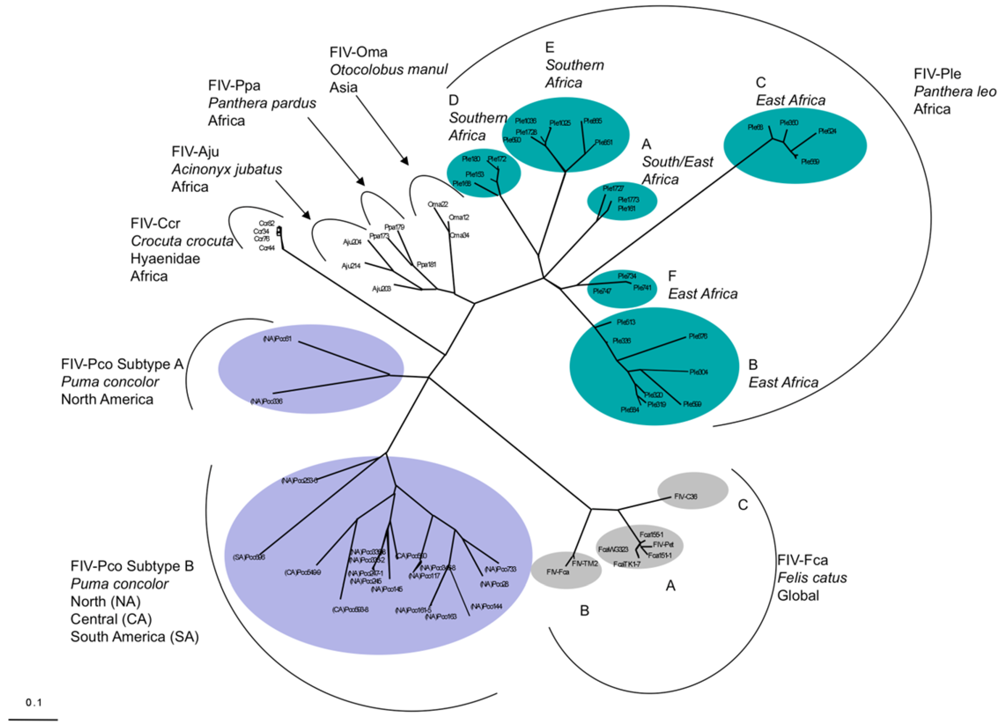
3. Feline Immunodeficiency Virus: FIV Pathogenesis in Felidae Species
| Species | Common Name | Free Ranging | Captive | Citation | ||
|---|---|---|---|---|---|---|
| AB+ | PCR+ | AB+ | PCR+ | |||
| Felis silvestris | European wild cat | 5/125 | 0/3 | 4/13 | [7,69,70,71,72] | |
| F. libyca | African wild cat | 1/16 | 0/1 | [7] | ||
| F. bieti | Chinese desert cat | |||||
| F. margarita | Desert cat | 0/14 | 6/13 | 0/7 | [7,69] | |
| F. nigripes | Black-footed cat | 3/11 | 0/4 | [7] | ||
| F. chaus | Jungle cat | 5/17 | 0/6 | [7] | ||
| Otocolobus manul | Pallas cat | 10/27 | 7/26 | 12/19 | 2/2 | [7,43] |
| Prionailurus rubiginosis | Rusty spotted cat | 0/1 | [7] | |||
| P. bengalensis | Asian leopard cat | 1/12 | 1/1 | 0/81 | [7,73,74] | |
| P. viverrinus | Fishing cat | 1/25 | 0/2 | [7] | ||
| P. planiceps | Flat-headed cat | 0/2 | 1/9 | [7] | ||
| Puma concolor | Puma | 150/360 | 61/123 | 45/166 | [7,61,75,76,77] | |
| P. yagouaroundi | Jaguarundi | 9/40 | 1/8 | [7,78] | ||
| Acinonyx jubatus | Cheetah | 22/303 | 7/10 | 6/242 | [7,75,79,80] | |
| Lynx pardinus | Iberian lynx | 7/74 | 0/75 | [7,81] | ||
| L. lynx | Eurasian lynx | 0/10 | [7,75] | |||
| L. canadensis | Canada lynx | 0/92 | 1/2 | 0/1 | [7,82] | |
| L. rufus | Bobcat | 32/115 | 17/32 | 1/8 | 0/1 | [7,83] |
| Leopardus pardalis | Ocelot | 8/26 | 1/14 | 10/88 | 0/6 | [7,75,78] |
| L. wiedii | Margay | 1/5 | 1/1 | 4/88 | 1/3 | [7,78] |
| L. jacobita | Andean mountain cat | |||||
| L. colocolo | Pampas cat | 1/12 | [7] | |||
| L. geoffroyi | Geoffroy's cat | 1/6 | 0/1 | 8/45 | 0/7 | [7] |
| L. guigna | Kodkod | 0/2 | [7] | |||
| L. tigrinus | Tigrina | 3/40 | 0/2 | [7,78] | ||
| Caracal caracal | Caracal | 0/3 | 0/22 | [7,79] | ||
| C. aurata | African golden cat | 0/2 | [7] | |||
| C. serval | Serval | 0/4 | [7,75] | |||
| Pardofelis badia | Bay cat | 0/1 | [7] | |||
| P. temminckii | Asian golden cat | 0/1 | 3/29 | 0/2 | [7] | |
| P. marmorata | Marbled cat | 2/10 | 0/3 | [7] | ||
| Panthera leo | Lion | 212/321 | 72/132 | 1/1 | [7,75,80,84,85,86] | |
| P. onca | Jaguar | 0/2 | 8/42 | 0/7 | [7,75,78] | |
| P. pardus | Leopard | 7/10 | 1/96 | 0/1 | [7,75,79,80] | |
| P. tigris | Tiger | 0/1 | 25/217 | 1/12 | [7,75] | |
| P. unca | Snow leopard | 3/77 | 1/2 | [7,75] | ||
| Neofelis nebulosa | Clouded leopard | 4/59 | 0/2 | [7] | ||
| Medical Condition | FIVPle Negative | FIVPle Positive | Odds Ratio | P Value | ||
|---|---|---|---|---|---|---|
| % Affected | # Individuals | % Affected | # Individuals | |||
| Immunodeficiency | ||||||
| CD4 depletion Absolute number of CD4+ T-cells /mL in peripheral whole blood ±s.e. | 0 | 5 | 100 | 8 | NA | 0.00015 |
| Oral manifestations | ||||||
| Gingivitis | 40 | 15 | 88.4 | 43 | 11.4 | 0.00016 |
| Papillomavirus | 14.3 | 14 | 53.19 | 47 | 6.82 | 0.01009 |
| Chronic Inflammatory Response | ||||||
| Lymphadenopathy | 41.67 | 12 | 76.6 | 47 | 4.58 | 0.01900 |
| Hyperglobulinemia | 0 | 14 | 85.71 | 46 | NA | <2 × 10−9 |
| Erythrocyte sedimentation rate | 13.33 | 15 | 64.86 | 37 | 12 | 0.00076 |
| (> 2 s.d. above mean) | ||||||
| Dehydration (> 4%) | 26.67 | 15 | 63.04 | 46 | 4.69 | 0.01408 |
| Loss of Condition and Under Nutrition | ||||||
| Hair and coat abnormalities | 13.3 | 15 | 52.27 | 44 | 7.12 | 0.00840 |
| Hypoalbuminemia (marker of cachexia) (serum albumin > 2 s.d. below mean) | 0 | 14 | 46.94 | 46 | NA | 0.00129 |
| Anemia (hemoglobin / PCV >2 s.d. below mean) | 11.11 | 18 | 55.77 | 52 | 10.09 | 0.00101 |
| Cachexia/unexplained weight loss | Not documented | Observed in 3 FIV+ populations | NA | NA | ||
| Lymphoid response evidence | ||||||
| Histopathologic evidence: Lymphoid activation | Not documented | Yes | NA | NA | ||
| Histopathologic evidence: Lymphoid atrophy & depletion | Not documented | Yes | NA | NA | ||
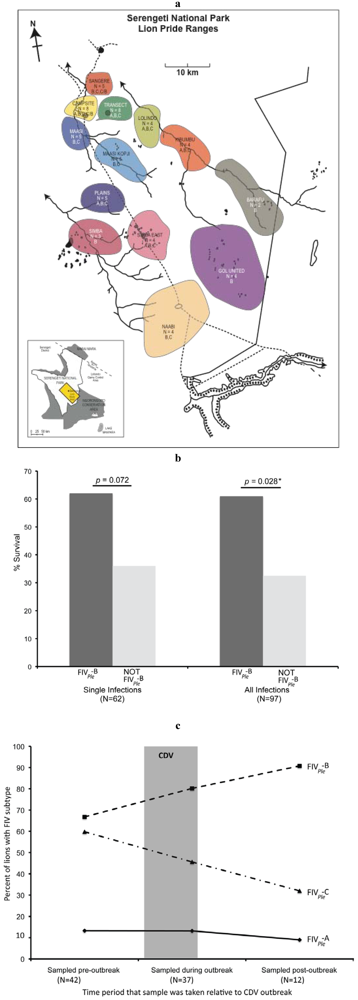
4. FeLV Outbreak in Free Ranging Florida Pumas
5. Conclusions
Conflict of Interest
References and Notes
- Driscoll, C.A.; Macdonald, D.W.; O'Brien, S.J. From wild animals to domestic pets, an evolutionary view of domestication. Proc. Natl. Acad. Sci. U. S. A. 2009, 106, 9971–9978. [Google Scholar]
- WHO. Summary of probable SARS cases with onset of illness from 1 November 2002 to 31 July 2003. Available online: http://www.who.int/csr/sars/country/table2004_2004_2021/en/index.html (accessed on 16 January 2012).
- Hardy, W.D., Jr.; McClelland, A.J.; Zuckerman, E.E.; Snyder, H.W., Jr.; MacEwen, E.G.; Francis, D.; Essex, M. Development of virus non-producer lymphosarcomas in pet cats exposed to FeLv. Nature 1980, 288, 90–92. [Google Scholar]
- Pedersen, N.C.; Ho, E.W.; Brown, M.L.; Yamamoto, J.K. Isolation of a T-lymphotropic virus from domestic cats with an immunodeficiency-like syndrome. Science 1987, 235, 790–793. [Google Scholar]
- Mullins, J.I.; Hoover, E.A. Molecular aspects of feline leukemia virus pathogenesis. In Retrovirus Biology and Human Disease; Gallo, R.C., Wong-Staal, F., Eds.; Dekker: New York, NY, USA, 1990; pp. 87–116. [Google Scholar]
- Slattery, J.P.; Franchini, G.; Gessain, A. Genomic evolution, patterns of global dissemination, and interspecies transmission of human and simian T-cell leukemia/lymphotropic viruses. Genome Res. 1999, 9, 525–540. [Google Scholar]
- Troyer, J.L.; Pecon-Slattery, J.; Roelke, M.E.; Johnson, W.; VandeWoude, S.; Vazquez-Salat, N.; Brown, M.; Frank, L.; Woodroffe, R.; Winterbach, C.; et al. Seroprevalence and genomic divergence of circulating strains of feline immunodeficiency virus among Felidae and Hyaenidae species. J. Virol. 2005, 79, 8282–8294. [Google Scholar]
- O'Brien, S.J.; Nelson, G.W. Human genes that limit AIDS. Nat. Genet. 2004, 36, 565–574. [Google Scholar]
- Pedersen, N.C. A review of feline infectious peritonitis virus infection: 1963–2008. J. Feline Med. Surg. 2009, 11, 225–258. [Google Scholar]
- Eickmann, M.; Becker, S.; Klenk, H.D.; Doerr, H.W.; Stadler, K.; Censini, S.; Guidotti, S.; Masignani, V.; Scarselli, M.; Mora, M.; et al. Phylogeny of the SARS coronavirus. Science 2003, 302, 1504–1505. [Google Scholar]
- Ruscetti, S.K.; Turek, L.P.; Sherr, C.J. Three independent isolates of feline sarcoma virus code for three distinct gag-x polyproteins. J. Virol. 1980, 35, 259–264. [Google Scholar]
- Maeda, N.; Fan, H.; Yoshikai, Y. Oncogenesis by retroviruses: Old and new paradigms. Rev. Med. Virol. 2008, 18, 387–405. [Google Scholar]
- Harder, T.C.; Vahlenkamp, T.W. Influenza virus infections in dogs and cats. Vet. Immunol. Immunopathol. 2009, 134, 54–60. [Google Scholar]
- Van Kerkhove, M.D.; Mumford, E.; Mounts, A.W.; Bresee, J.; Ly, S.; Bridges, C.B.; Otte, J. Highly pathogenic avian influenza (H5N1): Pathways of exposure at the animal-human interface, a systematic review. PLoS One 2011, 6, e14582. [Google Scholar]
- Thiry, E.; Addie, D.; Belak, S.; Boucraut-Baralon, C.; Egberink, H.; Frymus, T.; Gruffydd-Jones, T.; Hartmann, K.; Hosie, M.J.; Lloret, A.; et al. Feline herpesvirus infection. ABCD guidelines on prevention and management. J. Feline Med. Surg. 2009, 11, 547–555. [Google Scholar]
- Winkler, I.G.; Flugel, R.M.; Lochelt, M.; Flower, R.L. Detection and molecular characterisation of feline foamy virus serotypes in naturally infected cats. Virology 1998, 247, 144–151. [Google Scholar]
- Meiering, C.D.; Linial, M.L. Historical perspective of foamy virus epidemiology and infection. Clin. Microbiol. Rev. 2001, 14, 165–176. [Google Scholar]
- Radford, A.D.; Coyne, K.P.; Dawson, S.; Porter, C.J.; Gaskell, R.M. Feline calicivirus. Vet. Res. 2007, 38, 319–335. [Google Scholar]
- Blanton, L.H.; Adams, S.M.; Beard, R.S.; Wei, G.; Bulens, S.N.; Widdowson, M.A.; Glass, R.I.; Monroe, S.S. Molecular and epidemiologic trends of caliciviruses associated with outbreaks of acute gastroenteritis in the United States, 2000–2004. J. Infect. Dis. 2006, 193, 413–421. [Google Scholar]
- Ikeda, Y.; Nakamura, K.; Miyazawa, T.; Takahashi, E.; Mochizuki, M. Feline host range of canine parvovirus: Recent emergence of new antigenic types in cats. Emerg. Infect. Dis. 2002, 8, 341–346. [Google Scholar]
- Brown, K.E. The expanding range of parvoviruses which infect humans. Rev. Med. Virol. 2010, 20, 231–244. [Google Scholar]
- Munson, L. Feline morbillivirus infection. In Infectious Diseases of Wild Animals; Williams, E.S., Barker, I.K., Eds.; Iowa State University Press: Ames, IA, USA, 2001; pp. 59–62. [Google Scholar]
- Rota, P.A.; Brown, K.; Mankertz, A.; Santibanez, S.; Shulga, S.; Muller, C.P.; Hubschen, J.M.; Siqueira, M.; Beirnes, J.; Ahmed, H.; et al. Global distribution of measles genotypes and measles molecular epidemiology. J. Infect. Dis. 2011, 204, S514–S523. [Google Scholar]
- Masters, P.S. The molecular biology of coronaviruses. Adv. Virus Res. 2006, 66, 193–292. [Google Scholar]
- Pedersen, N.C.; Evermann, J.F.; McKeirnan, A.J.; Ott, R.L. Pathogenicity studies of feline coronavirus isolates 79-1146 and 79-1683. Am. J. Vet. Res. 1984, 45, 2580–2585. [Google Scholar]
- de Groot, R.J. Feline infectous peritonitis. In The Coronoviridae; Siddell, S.G., Ed.; Plenum Press: New York, NY, USA, 1995; pp. 293–309. [Google Scholar]
- Addie, D.D. Clustering of feline coronaviruses in multicat households. Vet. J. 2000, 159, 8–9. [Google Scholar]
- Addie, D.D.; Jarrett, O. A study of naturally occurring feline coronavirus infections in kittens. Vet. Rec. 1992, 130, 133–137. [Google Scholar]
- Kennedy, M.; Citino, S.; McNabb, A.H.; Moffatt, A.S.; Gertz, K.; Kania, S. Detection of feline coronavirus in captive Felidae in the USA. J. Vet. Diagn. Invest. 2002, 14, 520–522. [Google Scholar]
- Foley, J.E.; Poland, A.; Carlson, J.; Pedersen, N.C. Patterns of feline coronavirus infection and fecal shedding from cats in multiple-cat environments. J. Am. Vet. Med. Assoc. 1997, 210, 1307–1312. [Google Scholar]
- Foley, J.E.; Poland, A.; Carlson, J.; Pedersen, N.C. Risk factors for feline infectious peritonitis among cats in multiple-cat environments with endemic feline enteric coronavirus. J. Am. Vet. Med. Assoc. 1997, 210, 1313–1318. [Google Scholar]
- Weiss, R.C.; Scott, F.W. Pathogenesis of feline infectious peritonitis: Nature and development of viremia. Am. J. Vet. Res. 1981, 42, 382–390. [Google Scholar]
- Kipar, A.; Kohler, K.; Leukert, W.; Reinacher, M. A comparison of lymphatic tissues from cats with spontaneous feline infectious peritonitis (FIP), cats with FIP virus infection but no FIP, and cats with no infection. J. Comp. Pathol. 2001, 125, 182–191. [Google Scholar] [CrossRef]
- Kipar, A.; Meli, M.L.; Failing, K.; Euler, T.; Gomes-Keller, M.A.; Schwartz, D.; Lutz, H.; Reinacher, M. Natural feline coronavirus infection: Differences in cytokine patterns in association with the outcome of infection. Vet. Immunol. Immunopathol. 2006, 112, 141–155. [Google Scholar]
- Rottier, P.J. The Coronavirus Membrane Glycoprotein. In The Coronaviridae; Siddell, S.G., Ed.; Plenum Press: New York, NY, USA, 1995; pp. 115–140. [Google Scholar]
- Poland, A.M.; Vennema, H.; Foley, J.E.; Pedersen, N.C. Two related strains of feline infectious peritonitis virus isolated from immunocompromised cats infected with a feline enteric coronavirus. J. Clin. Microbiol. 1996, 34, 3180–3184. [Google Scholar]
- Vennema, H.; Poland, A.; Foley, J.; Pedersen, N.C. Feline infectious peritonitis viruses arise by mutation from endemic feline enteric coronaviruses. Virology 1998, 243, 150–157. [Google Scholar]
- Addie, D.D.; Jarrett, J.O. Feline coronavirus antibodies in cats. Vet. Rec. 1992, 131, 202–203. [Google Scholar]
- Pedersen, N.C.; Black, J.W.; Boyle, J.F.; Evermann, J.F.; McKeirnan, A.J.; Ott, R.L. Pathogenic differences between various feline coronavirus isolates. Adv. Exp. Med. Biol. 1984, 173, 365–380. [Google Scholar]
- Pedersen, N.; Liu, H.; Dodd, K.A.; Pesavento, P.A. Significance of coronavirus mutants in feces and diseased tissues of cats suffering from feline infectious peritonitis. Viruses 2009, 1, 166–184. [Google Scholar] [CrossRef]
- Anishchenko, M.; Bowen, R.A.; Paessler, S.; Austgen, L.; Greene, I.P.; Weaver, S.C. Venezuelan encephalitis emergence mediated by a phylogenetically predicted viral mutation. Proc. Natl. Acad. Sci. U. S. A. 2006, 103, 4994–4999. [Google Scholar]
- Mongkolsapaya, J.; Dejnirattisai, W.; Xu, X.N.; Vasanawathana, S.; Tangthawornchaikul, N.; Chairunsri, A.; Sawasdivorn, S.; Duangchinda, T.; Dong, T.; Rowland-Jones, S.; et al. Original antigenic sin and apoptosis in the pathogenesis of dengue hemorrhagic fever. Nat. Med. 2003, 9, 921–927. [Google Scholar]
- Brown, M.A.; Troyer, J.L.; Pecon-Slattery, J.; Roelke, M.E.; O'Brien, S.J. Genetics and pathogenesis of feline infectious peritonitis virus. Emerg. Infect. Dis. 2009, 15, 1445–1452. [Google Scholar]
- Chang, H.W.; Egberink, H.F.; Rottier, P.J. Sequence analysis of feline coronaviruses and the circulating virulent/avirulent theory. Emerg. Infect. Dis. 2011, 17, 744–746. [Google Scholar]
- Norris, J.M.; Bosward, K.L.; White, J.D.; Baral, R.M.; Catt, M.J.; Malik, R. Clinicopathological findings associated with feline infectious peritonitis in Sydney, Australia: 42 cases (1990–2002). Aust. Vet. 2005, 83, 666–673. [Google Scholar] [CrossRef]
- Pesteanu-Somogyi, L.D.; Radzai, C.; Pressler, B.M. Prevalence of feline infectious peritonitis in specific cat breeds. J. Feline Med. Surg. 2006, 8, 1–5. [Google Scholar] [CrossRef]
- Olmsted, R.A.; Barnes, A.K.; Yamamoto, J.K.; Hirsch, V.M.; Purcell, R.H.; Johnson, P.R. Molecular cloning of feline immunodeficiency virus. Proc. Natl. Acad. Sci. U. S. A. 1989, 86, 2448–2452. [Google Scholar]
- Bendinelli, M.; Pistello, M.; Lombardi, S.; Poli, A.; Garzelli, C.; Matteucci, D.; Ceccherini-Nelli, L.; Malvaldi, G.; Tozzini, F. Feline immunodeficiency virus: An interesting model for AIDS studies and an important cat pathogen. Clin. Microbiol. Rev. 1995, 8, 87–112. [Google Scholar]
- Willett, B.J.; Flynn, J.N.; Hosie, M.J. FIV infection of the domestic cat: An animal model for AIDS. Immunol. Today 1997, 18, 182–189. [Google Scholar]
- Burkhard, M.J.; Dean, G.A. Transmission and immunopathogenesis of FIV in cats as a model for HIV. Curr. HIV Res. 2003, 1, 15–29. [Google Scholar]
- Sodora, D.L.; Shpaer, E.G.; Kitchell, B.E.; Dow, S.W.; Hoover, E.A.; Mullins, J.I. Identification of three feline immunodeficiency virus (FIV) env gene subtypes and comparison of the FIV and human immunodeficiency virus type 1 evolutionary patterns. J. Virol. 1994, 68, 2230–2238. [Google Scholar]
- Kakinuma, S.; Motokawa, K.; Hohdatsu, T.; Yamamoto, J.K.; Koyama, H.; Hashimoto, H. Nucleotide sequence of feline immunodeficiency virus: Classification of Japanese isolates into two subtypes which are distinct from non-Japanese subtypes. J. Virol. 1995, 69, 3639–3646. [Google Scholar]
- Carpenter, M.A.; Brown, E.W.; MacDonald, D.W.; O'Brien, S.J. Phylogeographic patterns of feline immunodeficiency virus genetic diversity in the domestic cat. Virology 1998, 251, 234–243. [Google Scholar]
- Hayward, J.J.; Rodrigo, A.G. Molecular epidemiology of feline immunodeficiency virus in the domestic cat (Felis catus). Vet. Immunol. Immunopathol. 2009, 134, 68–74. [Google Scholar]
- Pecoraro, M.R.; Tomonaga, K.; Miyazawa, T.; Kawaguchi, Y.; Sugita, S.; Tohya, Y.; Kai, C.; Etcheverrigaray, M.E.; Mikami, T. Genetic diversity of Argentine isolates of feline immunodeficiency virus. J. Gen. Virol. 1996, 77, 2031–2035. [Google Scholar]
- Johnson, W.E.; Eizirik, E.; Pecon-Slattery, J.; Murphy, W.J.; Antunes, A.; Teeling, E.; O'Brien, S.J. The late Miocene radiation of modern Felidae: A genetic assessment. Science 2006, 311, 73–77. [Google Scholar]
- Antunes, A.; Troyer, J.L.; Roelke, M.E.; Pecon-Slattery, J.; Packer, C.; Winterbach, C.; Winterbach, H.; Hemson, G.; Frank, L.; Stander, P.; et al. The evolutionary dynamics of the lion Panthera leo revealed by host and viral population genomics. PLoS Genet. 2008, 4, e1000251. [Google Scholar]
- Pecon-Slattery, J.; Troyer, J.L.; Johnson, W.E.; O'Brien, S.J. Evolution of feline immunodeficiency virus in Felidae: Implications for human health and wildlife ecology. Vet. Immunol. Immunopathol. 2008, 123, 32–44. [Google Scholar]
- Olmsted, R.A.; Langley, R.; Roelke, M.E.; Goeken, R.M.; Adger-Johnson, D.; Goff, J.P.; Albert, J.P.; Packer, C.; Laurenson, M.K.; Caro, T.M.; et al. Worldwide prevalence of lentivirus infection in wild feline species: Epidemiologic and phylogenetic aspects. J. Virol. 1992, 66, 6008–6018. [Google Scholar]
- Carpenter, M.A.; O'Brien, S.J. Coadaptation and immunodeficiency virus: Lessons from the Felidae. Curr. Opin. Genet. Dev. 1995, 5, 739–745. [Google Scholar]
- Carpenter, M.A.; Brown, E.W.; Culver, M.; Johnson, W.E.; Pecon-Slattery, J.; Brousset, D.; O'Brien, S.J. Genetic and phylogenetic divergence of feline immunodeficiency virus in the puma (Puma concolor). J. Virol. 1996, 70, 6682–6693. [Google Scholar]
- Franklin, S.P.; Troyer, J.L.; Terwee, J.A.; Lyren, L.M.; Boyce, W.M.; Riley, S.P.; Roelke, M.E.; Crooks, K.R.; Vandewoude, S. Frequent transmission of immunodeficiency viruses among bobcats and pumas. J. Virol. 2007, 81, 10961–10969. [Google Scholar]
- VandeWoude, S.; Apetrei, C. Going wild: Lessons from naturally occurring T-lymphotropic lentiviruses. Clin. Microbiol. Rev. 2006, 19, 728–762. [Google Scholar]
- Troyer, J.L.; Vandewoude, S.; Pecon-Slattery, J.; McIntosh, C.; Franklin, S.; Antunes, A.; Johnson, W.; O'Brien, S.J. FIV cross-species transmission: An evolutionary prospective. Vet. Immunol. Immunopathol. 2008, 123, 159–166. [Google Scholar]
- Pecon-Slattery, J.; McCracken, C.L.; Troyer, J.L.; VandeWoude, S.; Roelke, M.; Sondgeroth, K.; Winterbach, C.; Winterbach, H.; O'Brien, S.J. Genomic organization, sequence divergence, and recombination of feline immunodeficiency virus from lions in the wild. BMC Genom. 2008, 9, 66. [Google Scholar]
- Olmsted, R.A.; Hirsch, V.M.; Purcell, R.H.; Johnson, P.R. Nucleotide sequence analysis of feline immunodeficiency virus: Genome organization and relationship to other lentiviruses. Proc. Natl. Acad. Sci. U. S. A. 1989, 86, 8088–8092. [Google Scholar]
- Langley, R.J.; Hirsch, V.M.; O'Brien, S.J.; Adger-Johnson, D.; Goeken, R.M.; Olmsted, R.A. Nucleotide sequence analysis of puma lentivirus (PLV-14): Genomic organization and relationship to other lentiviruses. Virology 1994, 202, 853–864. [Google Scholar]
- Barr, M.C.; Zou, L.; Long, F.; Hoose, W.A.; Avery, R.J. Proviral organization and sequence analysis of feline immunodeficiency virus isolated from a Pallas' cat. Virology 1997, 228, 84–91. [Google Scholar]
- Ostrowski, S.; Van Vuuren, M.; Lenain, D.M.; Durand, A. A serologic survey of wild felids from central west Saudi Arabia. J. Wildl. Dis. 2003, 39, 696–701. [Google Scholar]
- Leutenegger, C.M.; Hofmann-Lehmann, R.; Riols, C.; Liberek, M.; Worel, G.; Lups, P.; Fehr, D.; Hartmann, M.; Weilenmann, P.; Lutz, H. Viral infections in free-living populations of the European wildcat. J. Wildl. Dis. 1999, 35, 678–686. [Google Scholar]
- Daniels, M.J.; Golder, M.C.; Jarrett, O.; MacDonald, D.W. Feline viruses in wildcats from Scotland. J. Wildl. Dis. 1999, 35, 121–124. [Google Scholar]
- Fromont, E.; Sager, A.; Leger, F.; Bourguemestre, F.; Jouquelet, E.; Stahl, P.; Pontier, D.; Artois, M. Prevalence and pathogenicity of retroviruses in wildcats in France. Vet. Rec. 2000, 146, 317–319. [Google Scholar]
- Nishimura, Y.; Goto, Y.; Yoneda, K.; Endo, Y.; Mizuno, T.; Hamachi, M.; Maruyama, H.; Kinoshita, H.; Koga, S.; Komori, M.; et al. Interspecies transmission of feline immunodeficiency virus from the domestic cat to the Tsushima cat (Felis bengalensis euptilura) in the wild. J. Virol. 1999, 73, 7916–7921. [Google Scholar]
- Nakamura, K.; Miyazawa, T.; Ikeda, Y.; Sato, E.; Nishimura, Y.; Nguyen, N.T.; Takahashi, E.; Mochizuki, M.; Mikami, T. Contrastive prevalence of feline retrovirus infections between northern and southern Vietnam. J. Vet. Med. Sci. 2000, 62, 921–923. [Google Scholar]
- Lutz, H.; Isenbugel, E.; Lehmann, R.; Sabapara, R.H.; Wolfensberger, C. Retrovirus infections in non-domestic felids: Serological studies and attempts to isolate a lentivirus. Vet. Immunol. Immunopathol. 1992, 35, 215–224. [Google Scholar]
- Franklin, S.P.; Troyer, J.L.; Terwee, J.A.; Lyren, L.M.; Kays, R.W.; Riley, S.P.; Boyce, W.M.; Crooks, K.R.; Vandewoude, S. Variability in assays used for detection of lentiviral infection in bobcats (Lynx rufus), pumas (Puma concolor), and ocelots (Leopardus pardalis). J. Wildl. Dis. 2007, 43, 700–710. [Google Scholar]
- Biek, R.; Rodrigo, A.G.; Holley, D.; Drummond, A.; Anderson, C.R., Jr.; Ross, H.A.; Poss, M. Epidemiology, genetic diversity, and evolution of endemic feline immunodeficiency virus in a population of wild cougars. J. Virol. 2003, 77, 9578–9589. [Google Scholar]
- Blanco, K.; Peña, R.; Hernández, C.; Jiménez, M.; Araya, L.N.; Romero, J.J.; Dolz, G. Serological detection of viral infections in captive wild cats from Costa Rica. Vet. Med. Int. 2011, 2011, 879029. [Google Scholar]
- Thalwitzer, S.; Wachter, B.; Robert, N.; Wibbelt, G.; Muller, T.; Lonzer, J.; Meli, M.L.; Bay, G.; Hofer, H.; Lutz, H. Seroprevalences to viral pathogens in free-ranging and captive cheetahs (Acinonyx jubatus) on Namibian Farmland. Clin. Vaccine Immunol. 2009, 17, 232–238. [Google Scholar]
- Osofsky, S.A.; Hirsch, K.J.; Zuckerman, E.E.; et al. Feline lentivirus and feline oncovirus status of free-ranging lions (Panthera leo), leopards (Panthera pardus), and cheetahs (Acinonyx jubatus) in Botswana: A regional perspective. J. Zoo wildl. Med. 1996, 27, 453–467. [Google Scholar]
- Meli, M.L.; Cattori, V.; Martinez, F.; Lopez, G.; Vargas, A.; Simon, M.A.; Zorrilla, I.; Munoz, A.; Palomares, F.; Lopez-Bao, J.V.; et al. Feline leukemia virus and other pathogens as important threats to the survival of the critically endangered Iberian lynx (Lynx pardinus). PLoS One 2009, 4, e4744. [Google Scholar]
- Biek, R.; Zarnke, R.L.; Gillin, C.; Wild, M.; Squires, J.R.; Poss, M. Serologic survey for viral and bacterial infections in western populations of Canada lynx (Lynx canadensis). J. Wildl. Dis. 2002, 38, 840–845. [Google Scholar]
- Franklin, S.P.; Kays, R.W.; Moreno, R.; TerWee, J.A.; Troyer, J.L.; VandeWoude, S. Ocelots on Barro Colorado Island are infected with feline immunodeficiency virus but not other common feline and canine viruses. J. Wildl. Dis. 2008, 44, 760–765. [Google Scholar]
- Brennan, G.; Podell, M.D.; Wack, R.; Kraft, S.; Troyer, J.L.; Bielefeldt-Ohmann, H.; VandeWoude, S. Neurologic disease in captive lions (Panthera leo) with low-titer lion lentivirus infection. J. Clin. Microbiol. 2006, 44, 4345–4352. [Google Scholar]
- Driciru, M.; Siefert, L.; Prager, K.C.; Dubovi, E.; Sande, R.; Princee, F.; Friday, T.; Munson, L. A serosurvey of viral infections in lions (Panthera leo), from Queen Elizabeth National Park, Uganda. J. Wildl. Dis. 2006, 42, 667–671. [Google Scholar]
- Adams, H.; van Vuuren, M.; Kania, S.; Bosman, A.M.; Keet, D.; New, J.; Kennedy, M. Sensitivity and specificity of a nested polymerase chain reaction for detection of lentivirus infection in lions (Panthera leo). J. Zoo Wildl. Med. 2011, 41, 608–615. [Google Scholar]
- Packer, C.; Altizer, S.; Appel, M.; Brown, E.; Martenson, J.; O’Brien, S.J.; Roelke-Parker, M.; Hofmann-Lehmann, R.; Lutz, H. Viruses of the Serengeti: Patterns of infection and mortality in African lions. J. Anim. Ecol. 1999, 68, 1161–1178. [Google Scholar]
- Roelke, M.E.; Brown, M.A.; Troyer, J.L.; Winterbach, H.; Winterbach, C.; Hemson, G.; Smith, D.; Johnson, R.C.; Pecon-Slattery, J.; Roca, A.L.; et al. Pathological manifestations of feline immunodeficiency virus (FIV) infection in wild African lions. Virology 2009, 390, 1–12. [Google Scholar]
- Roelke, M.E.; Pecon-Slattery, J.; Taylor, S.; Citino, S.; Brown, E.; Packer, C.; Vandewoude, S.; O'Brien, S.J. T-lymphocyte profiles in FIV-infected wild lions and pumas reveal CD4 depletion. J. Wildl. Dis. 2006, 42, 234–248. [Google Scholar]
- Keele, B.F.; Jones, J.H.; Terio, K.A.; Estes, J.D.; Rudicell, R.S.; Wilson, M.L.; Li, Y.; Learn, G.H.; Beasley, T.M.; Schumacher-Stankey, J.; et al. Increased mortality and AIDS-like immunopathology in wild chimpanzees infected with SIVcpz. Nature 2009, 460, 515–519. [Google Scholar]
- Roelke-Parker, M.E.; Munson, L.; Packer, C.; Kock, R.; Cleaveland, S.; Carpenter, M.; O'Brien, S.J.; Pospischil, A.; Hofmann-Lehmann, R.; Lutz, H.; et al. A canine distemper virus epidemic in Serengeti lions (Panthera leo). Nature 1996, 379, 441–445. [Google Scholar]
- Brown, E.W.; Yuhki, N.; Packer, C.; O'Brien, S.J. A lion lentivirus related to feline immunodeficiency virus: Epidemiologic and phylogenetic aspects. J. Virol. 1994, 68, 5953–5968. [Google Scholar]
- Troyer, J.L.; Pecon-Slattery, J.; Roelke, M.E.; Black, L.; Packer, C.; O'Brien, S.J. Patterns of feline immunodeficiency virus multiple infection and genome divergence in a free-ranging population of African lions. J. Virol. 2004, 78, 3777–3791. [Google Scholar]
- Troyer, J.L.; Roelke, M.E.; Jespersen, J.M.; Baggett, N.; Buckley-Beason, V.; Macnulty, D.; Craft, M.; Packer, C.; Pecon-Slattery, J.; O'Brien, S.J. FIV diversity: FIV(Ple) subtype composition may influence disease outcome in African lions. Vet. Immunol. Immunopathol. 2011, 143, 338–346. [Google Scholar]
- VandeWoude, S.; Hageman, C.A.; O'Brien, S.J.; Hoover, E.A. Nonpathogenic lion and puma lentiviruses impart resistance to superinfection by virulent feline immunodeficiency virus. J. Acquir. Immune Defic. Syndr. 2002, 29, 1–10. [Google Scholar]
- VandeWoude, S.; Hageman, C.L.; Hoover, E.A. Domestic cats infected with lion or puma lentivirus develop anti-feline immunodeficiency virus immune responses. J. Acquir. Immune Defic. Syndr. 2003, 34, 20–31. [Google Scholar]
- Terwee, J.A.; Carlson, J.K.; Sprague, W.S.; Sondgeroth, K.S.; Shropshire, S.B.; Troyer, J.L.; VandeWoude, S. Prevention of immunodeficiency virus induced CD4+ T-cell depletion by prior infection with a non-pathogenic virus. Virology 2008, 377, 63–70. [Google Scholar]
- Jarrett, W.F.; Crawford, E.M.; Martin, W.B.; Davie, F. A Virus-Like Particle Associated with Leukemia (Lymphosarcoma). Nature 1964, 202, 567–569. [Google Scholar]
- Hardy, W.D. The virology, immunology and epidemiology of the feline leukemia virus. In Feline Leukemia Virus; Hardy, W.D., Essex, M., McClelland, A.J., Eds.; Elsevier/North-Holland: New York, NY, USA, 1980; pp. 33–78. [Google Scholar]
- Overbaugh, J.; Bangham, C.R. Selection forces and constraints on retroviral sequence variation. Science 2001, 292, 1106–1109. [Google Scholar]
- Phipps, A.J.; Chen, H.; Hayes, K.A.; Roy-Burman, P.; Mathes, L.E. Differential pathogenicity of two feline leukemia virus subgroup A molecular clones, pFRA and pF6A. J. Virol. 2000, 74, 5796–5801. [Google Scholar]
- Neil, J.C.; Fulton, R.; Rigby, M.; Stewart, M. Feline leukaemia virus: Generation of pathogenic and oncogenic variants. Curr. Top. Microbiol. Immunol. 1991, 171, 67–93. [Google Scholar]
- Stewart, M.A.; Warnock, M.; Wheeler, A.; Wilkie, N.; Mullins, J.I.; Onions, D.E.; Neil, J.C. Nucleotide sequences of a feline leukemia virus subgroup A envelope gene and long terminal repeat and evidence for the recombinational origin of subgroup B viruses. J. Virol. 1986, 58, 825–834. [Google Scholar]
- Benveniste, R.E.; Sherr, C.J.; Todaro, G.J. Evolution of type C viral genes: Origin of feline leukemia virus. Science 1975, 190, 886–888. [Google Scholar]
- Cunningham, M.W.; Brown, M.A.; Shindle, D.B.; Terrell, S.P.; Hayes, K.A.; Ferree, B.C.; McBride, R.T.; Blankenship, E.L.; Jansen, D.; Citino, S.B.; et al. Epizootiology and management of feline leukemia virus in the Florida puma. J. Wildl. Dis. 2008, 44, 537–552. [Google Scholar]
- Kennedy-Stoskopf, S. Emerging viral infections in large cats. In Zoo and Wild Animal Medicine; Fowler, M.E., Miller, R.E., Eds.; W.B. Saunders: Philadelphia, PA, USA, 1999; pp. 401–410. [Google Scholar]
- Brown, M.A.; Cunningham, M.W.; Roca, A.L.; Troyer, J.L.; Johnson, W.E.; O'Brien, S.J. Genetic characterization of feline leukemia virus from Florida panthers. Emerg. Infect. Dis. 2008, 14, 252–259. [Google Scholar]
- Young, S.P.; Goldman, A. The Puma, Mysterious American Cat; Dover Publications: New York, NY, USA, 1946; p. 358. [Google Scholar]
- Nowak, R.M.; McBride, R. Status survey of the Florida panther. In World Wildlife Fund Yearbook 1973–1974; Danbury Press: Danbury, CT, USA, 1974; pp. 112–113. [Google Scholar]
- Johnson, W.E.; Onorato, D.P.; Roelke, M.E.; Land, E.D.; Cunningham, M.; Belden, R.C.; McBride, R.; Jansen, D.; Lotz, M.; Shindle, D.; et al. Genetic restoration of the Florida panther. Science 2010, 329, 1641–1645. [Google Scholar]
- Seal, U.S. A Plan for Genetic Restoration and Management of the Florida Panther (Felis concolor coryi).; White Oak Conservation Center: Yulee, FL, USA, 1994; p. 23. [Google Scholar]
- McBride, R. The Documented Panther Population (DPP) and Its Current Distribution from July 1, 2002 to June 30, 2003; Livestock Protection Company: Alpine, TA, USA, 2003; p. 11. [Google Scholar]
- Benveniste, R.E.; Todaro, G.J. Segregation of RD-114 AND FeL-V-related sequences in crosses between domestic cat and leopard cat. Nature 1975, 257, 506–508. [Google Scholar]
- Chandhasin, C.; Coan, P.N.; Pandrea, I.; Grant, C.K.; Lobelle-Rich, P.A.; Puetter, A.; Levy, L.S. Unique long terminal repeat and surface glycoprotein gene sequences of feline leukemia virus as determinants of disease outcome. J. Virol. 2005, 79, 5278–5287. [Google Scholar]
- Chandhasin, C.; Coan, P.N.; Levy, L.S. Subtle mutational changes in the SU protein of a natural feline leukemia virus subgroup A isolate alter disease spectrum. J. Virol. 2005, 79, 1351–1360. [Google Scholar]
- Lee, I.T.; Levy, J.K.; Gorman, S.P.; Crawford, P.C.; Slater, M.R. Prevalence of feline leukemia virus infection and serum antibodies against feline immunodeficiency virus in unowned free-roaming cats. J. Am. Vet. Med. Assoc. 2002, 220, 620–622. [Google Scholar]
- O'Connor, T.P., Jr.; Tonelli, Q.J.; Scarlett, J.M. Report of the National FeLV/FIV Awareness Project. J. Am. Vet. Med. Assoc. 1991, 199, 1348–1353. [Google Scholar]
- Ishida, T.; Washizu, T.; Toriyabe, K.; Motoyoshi, S.; Tomoda, I.; Pedersen, N.C. Feline immunodeficiency virus infection in cats of Japan. J. Am. Vet. Med. Assoc. 1989, 194, 221–225. [Google Scholar]
- O'Connor, T.P., Jr.; Tanguay, S.; Steinman, R.; Smith, R.; Barr, M.C.; Yamamoto, J.K.; Pedersen, N.C.; Andersen, P.R.; Tonelli, Q.J. Development and evaluation of immunoassay for detection of antibodies to the feline T-lymphotropic lentivirus (feline immunodeficiency virus). J. Clin. Microbiol. 1989, 27, 474–479. [Google Scholar]
- Yamamoto, J.K.; Hansen, H.; Ho, E.W.; Morishita, T.Y.; Okuda, T.; Sawa, T.R.; Nakamura, R.M.; Pedersen, N.C. Epidemiologic and clinical aspects of feline immunodeficiency virus infection in cats from the continental United States and Canada and possible mode of transmission. J. Am. Vet. Med. Assoc. 1989, 194, 213–220. [Google Scholar]
- Cohen, N.D.; Carter, C.N.; Thomas, M.A.; Lester, T.L.; Eugster, A.K. Epizootiologic association between feline immunodeficiency virus infection and feline leukemia virus seropositivity. J. Am. Vet. Med. Assoc. 1990, 197, 220–225. [Google Scholar]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
O’Brien, S.J.; Troyer, J.L.; Brown, M.A.; Johnson, W.E.; Antunes, A.; Roelke, M.E.; Pecon-Slattery, J. Emerging Viruses in the Felidae: Shifting Paradigms. Viruses 2012, 4, 236-257. https://doi.org/10.3390/v4020236
O’Brien SJ, Troyer JL, Brown MA, Johnson WE, Antunes A, Roelke ME, Pecon-Slattery J. Emerging Viruses in the Felidae: Shifting Paradigms. Viruses. 2012; 4(2):236-257. https://doi.org/10.3390/v4020236
Chicago/Turabian StyleO’Brien, Stephen J., Jennifer L. Troyer, Meredith A. Brown, Warren E. Johnson, Agostinho Antunes, Melody E. Roelke, and Jill Pecon-Slattery. 2012. "Emerging Viruses in the Felidae: Shifting Paradigms" Viruses 4, no. 2: 236-257. https://doi.org/10.3390/v4020236
APA StyleO’Brien, S. J., Troyer, J. L., Brown, M. A., Johnson, W. E., Antunes, A., Roelke, M. E., & Pecon-Slattery, J. (2012). Emerging Viruses in the Felidae: Shifting Paradigms. Viruses, 4(2), 236-257. https://doi.org/10.3390/v4020236






