Molecular Determinants of Human T-lymphotropic Virus Type 1 Transmission and Spread
Abstract
:1. Introduction
2. HTLV-1 Associated Diseases
| Disease or Syndrome | Clinical Characteristics and Pathologic Outcomes |
|---|---|
| Adult T-cell leukemia/ lymphoma (ATL) |
|
| HTLV-1-Associated Myelopathy/ Tropical Spastic Paraparesis (HAM/TSP) |
|
| HTLV-1-associated Dermatitis |
|
| Ocular Lesions |
|
| Inflammatory Arthropathy and Polymyositis |
|
3. Replication and Organization of the HTLV-1 Genome
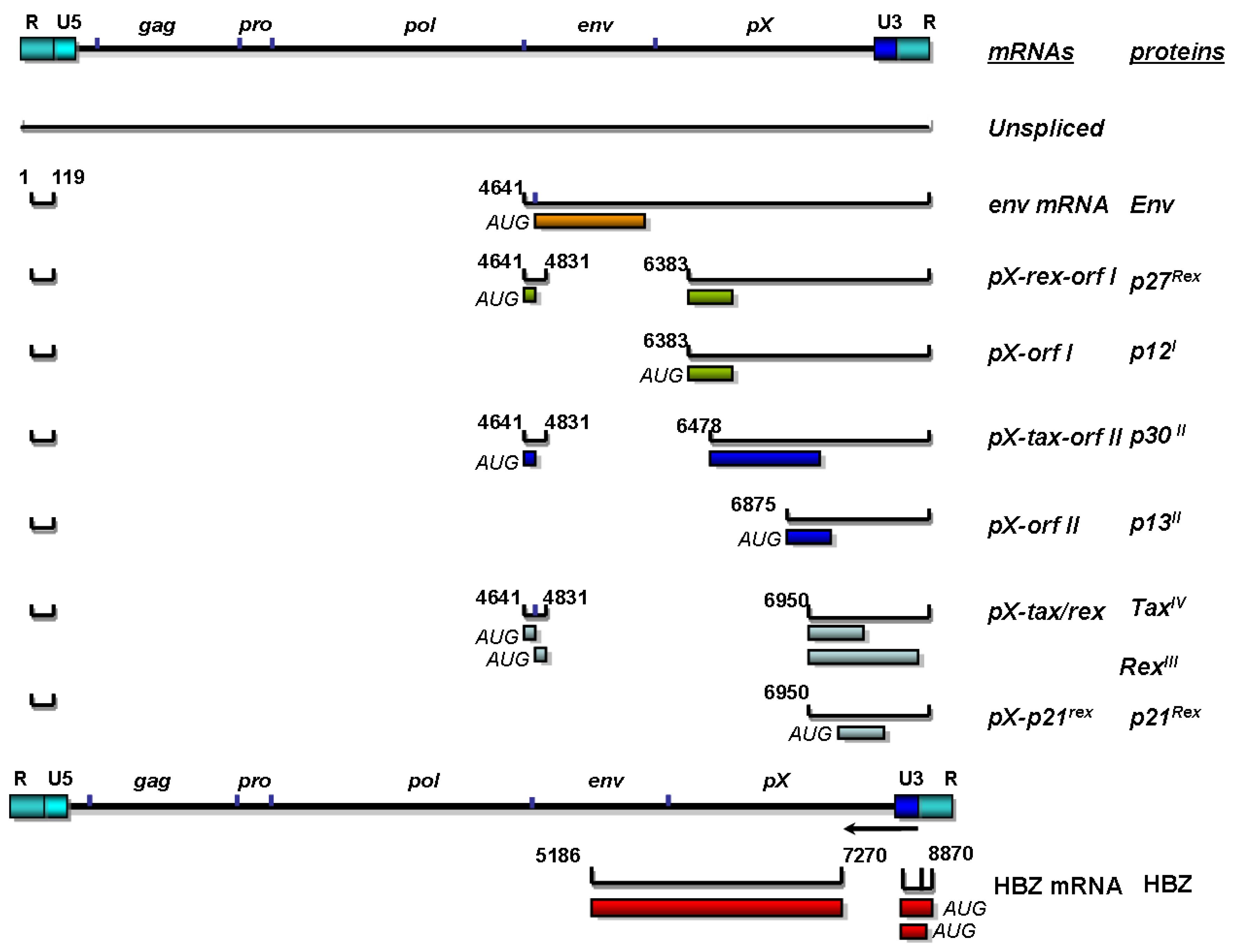
4. Structural Proteins of HTLV-1 and Their Influence on Viral Particle Assembly and Transmission

5. Regulatory Proteins of HTLV-1
5.1. Tax
5.2. Rex
6. Nonstructural Proteins of HTLV-1: Essential Role in Viral Spread and Transmission
6.1. p12 and p8
6.2. p13
6.3. p30
6.4. HBZ
7. Animal Models to Evaluate Viral Determinants of HTLV-1 Transmission and Spread
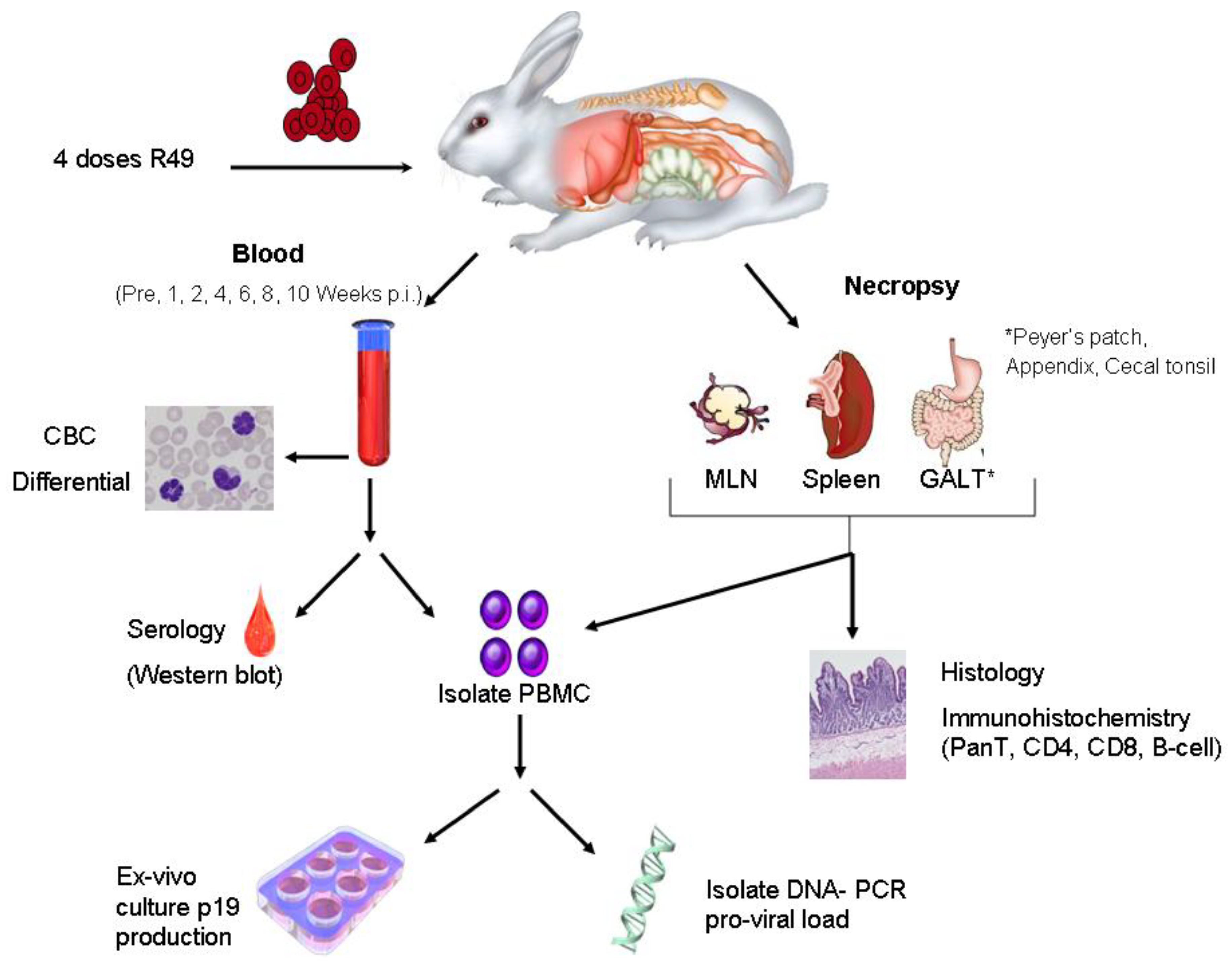
8. Conclusions
Acknowledgments
Conflicts of Interest
References and Notes
- Sonoda, S.; Li, H.C.; Tajima, K. Ethnoepidemiology of HTLV-1 related diseases: Ethnic determinants of HTLV-1 susceptibility and its worldwide dispersal. Cancer Sci. 2011, 102, 295–301. [Google Scholar] [CrossRef]
- Anderson, D.W.; Epstein, J.S.; Lee, T.H.; Lairmore, M.D.; Saxinger, C.; Kalyanaraman, V.S.; Slamon, D.; Parks, W.; Poiesz, B.J.; Pierik, L.T. Serological confirmation of human T-lymphotropic virus type I infection in healthy blood and plasma donors. Blood 1989, 74, 2585–2591. [Google Scholar] [CrossRef] [PubMed]
- Khabbaz, R.F.; Onorato, I.M.; Cannon, R.O.; Hartley, T.M.; Roberts, B.; Hosein, B.; Kaplan, J.E. Seroprevalence of HTLV-1 and HTLV-2 among intravenous drug users and persons in clinics for sexually transmitted diseases. N. Engl. J. Med. 1992, 326, 375–380. [Google Scholar] [CrossRef] [PubMed]
- Hino, S.; Yamaguchi, K.; Katamine, S.; Sugiyama, H.; Amagasaki, T.; Kinoshita, K.; Yoshida, Y.; Doi, H.; Tsuji, Y.; Miyamoto, T. Mother-to-child transmission of human T-cell leukemia virus type-I. Jpn. J. Cancer Res. 1985, 76, 474–480. [Google Scholar] [PubMed]
- Kinoshita, K.; Amagasaki, T.; Hino, S.; Doi, H.; Yamanouchi, K.; Ban, N.; Momita, S.; Ikeda, S.; Kamihira, S.; Ichimaru, M. Milk-borne transmission of HTLV-I from carrier mothers to their children. Jpn. J. Cancer Res. 1987, 78, 674–680. [Google Scholar]
- Minagawa, H.; Mora, C.A.; Asher, D.M.; Stone, G.A.; Liberski, P.P.; Gibbs, C.J. Transmission of human T-cell leukemia virus type 1 from a patient with HTLV-1 associated myelopathy/tropical spastic paraparesis and an asymptomatic carrier to rabbits. Arch. Virol. 1991, 118, 235–245. [Google Scholar] [CrossRef]
- Kaul, D.R.; Taranto, S.; Alexander, C.; Covington, S.; Marvin, M.; Nowicki, M.; Orlowski, J.; Pancoska, C.; Pruett, T.L.; Ison, M.G. Donor screening for human T-cell lymphotrophic virus 1/2: Changing paradigms for changing testing capacity. Am. J. Transplant. 2010, 10, 207–213. [Google Scholar] [CrossRef]
- Hino, S.; Katamine, S.; Kawase, K.; Miyamoto, T.; Doi, H.; Tsuji, Y.; Yamabe, T. Intervention of maternal transmission of HTLV-1 in Nagasaki, Japan. Leukemia 1994, 8, S68–S70. [Google Scholar]
- Eshima, N.; Tabata, M.; Okada, T.; Karukaya, S. Population dynamics of HTLV-I infection: A discrete-time mathematical epidemic model approach. Math. Med. Biol. 2003, 20, 29–45. [Google Scholar] [CrossRef]
- Jones, K.S.; Petrow-Sadowski, C.; Huang, Y.K.; Bertolette, D.C.; Ruscetti, F.W. Cell-free HTLV-1 infects dendritic cells leading to transmission and transformation of CD4(+) T cells. Nat. Med. 2008, 14, 429–436. [Google Scholar] [CrossRef]
- Igakura, T.; Stinchcombe, J.C.; Goon, P.K.; Taylor, G.P.; Weber, J.N.; Griffiths, G.M.; Tanaka, Y.; Osame, M.; Bangham, C.R. Spread of HTLV-I between lymphocytes by virus-induced polarization of the cytoskeleton. Science 2003, 299, 1713–1716. [Google Scholar] [CrossRef] [PubMed]
- Majorovits, E.; Nejmeddine, M.; Tanaka, Y.; Taylor, G.P.; Fuller, S.D.; Bangham, C.R. Human T-lymphotropic virus-1 visualized at the virological synapse by electron tomography. PLoS ONE 2008, 3, e2251. [Google Scholar] [CrossRef]
- Nejmeddine, M.; Barnard, A.L.; Tanaka, Y.; Taylor, G.P.; Bangham, C.R. HTLV-1 tax protein triggers microtubule reorientation in the virological synapse. J. Biol. Chem. 2005, 280, 29653–29660. [Google Scholar] [CrossRef] [PubMed]
- Van, P.N.; Gold, H.; Andresen, V.; Schwartz, O.; Jones, K.; Ruscetti, F.; Lockett, S.; Gudla, P.; Venzon, D.; Franchini, G. Human T-cell leukemia virus type 1 p8 protein increases cellular conduits and virus transmission. Proc. Natl. Acad. Sci. U. S. A. 2010. [CrossRef]
- Pais-Correia, A.M.; Sachse, M.; Guadagnini, S.; Robbiati, V.; Lasserre, R.; Gessain, A.; Gout, O.; Alcover, A.; Thoulouze, M.I. Biofilm-like extracellular viral assemblies mediate HTLV-1 cell-to-cell transmission at virological synapses. Nat. Med. 2010, 16, 83–89. [Google Scholar] [CrossRef]
- Taylor, G.P.; Matsuoka, M. Natural history of adult T-cell leukemia/lymphoma and approaches to therapy. Oncogene 2005, 24, 6047–6057. [Google Scholar] [CrossRef] [PubMed]
- Ohshima, K. Pathological features of diseases associated with human T-cell leukemia virus type I. Cancer Sci. 2007, 98, 772–778. [Google Scholar] [CrossRef]
- Yasunaga, J.; Matsuoka, M. Human T-cell leukemia virus type I induces adult T-cell leukemia: From clinical aspects to molecular mechanisms. Cancer Control 2007, 14, 133–140. [Google Scholar] [CrossRef]
- Ishitsuka, K.; Tamura, K. Treatment of adult T-cell leukemia/lymphoma: Past, present, and future. Eur. J. Haematol. 2008, 80, 185–196. [Google Scholar] [CrossRef]
- Goncalves, D.U.; Proietti, F.A.; Ribas, J.G.; Araujo, M.G.; Pinheiro, S.R.; Guedes, A.C.; Carneiro-Proietti, A.B. Epidemiology, treatment, and prevention of human T-cell leukemia virus type 1-associated diseases. Clin. Microbiol. Rev. 2010, 23, 577–589. [Google Scholar] [CrossRef]
- Yasunaga, J.; Matsuoka, M. Leukaemogenic mechanism of human T-cell leukaemia virus type I. Rev. Med. Virol. 2007, 17, 301–311. [Google Scholar] [CrossRef]
- Mahieux, R.; Gessain, A. HTLV-1 and associated adult T-cell leukemia/lymphoma. Rev. Clin. Exp. Hematol. 2003, 7, 336–361. [Google Scholar] [PubMed]
- Cavrois, M.; Leclercq, I.; Gout, O.; Gessain, A.; Wainhobson, S.; Wattel, E. Persistent oligoclonal expansion of human T-cell leukemia virus type 1 infected circulating cells in patients with Tropical spastic paraparesis/HTLV-1 associated myelopathy. Oncogene 1998, 17, 77–82. [Google Scholar] [CrossRef]
- Mortreux, F.; Kazanji, M.; Gabet, A.S.; de Thoisy, B.; Wattel, E. Two-step nature of human T-cell leukemia virus type 1 replication in experimentally infected squirrel monkeys (Saimiri sciureus). J. Virol. 2001, 75, 1083–1089. [Google Scholar] [CrossRef]
- Matsuoka, M.; Jeang, K.T. Human T-cell leukemia virus type 1 (HTLV-1) and leukemic transformation: viral infectivity, Tax, HBZ and therapy. Oncogene 2011, 30, 1379–1389. [Google Scholar] [CrossRef]
- Matsuoka, M.; Green, P.L. The HBZ gene, a key player in HTLV-1 pathogenesis. Retrovirology 2009, 6, 71. [Google Scholar] [CrossRef]
- Gessain, A.; Barin, F.; Vernant, J.; Gout, O.; Maurs, L.; Calender, A.; Dethe, G. Antibodies to human T lymphotropic virus type 1 in patients with tropical spastic paresis. Lancet 1985, 2, 407–410. [Google Scholar] [CrossRef]
- Osame, M.; Usuku, K.; Izumo, S.; Ijichi, N.; Amitani, H.; Igata, A.; Matsumoto, M.; Tara, M. HTLV-I associated myelopathy, a new clinical entity. Lancet 1986, i, 1031–1032. [Google Scholar] [CrossRef]
- Uchiyama, T. Human T cell leukemia virus type I (HTLV-I) and human diseases. [Review] [134 refs]. Annual. Rev. Immunol. 1997, 15, 15–37. [Google Scholar] [CrossRef]
- Cooper, S.A.; van der Loeff, M.S.; Taylor, G.P. The neurology of HTLV-1 infection. Pract. Neurol. 2009, 9, 16–26. [Google Scholar] [CrossRef]
- Nakamura, T. Immunopathogenesis of HTLV-I-associated myelopathy/tropical spastic paraparesis. Ann. Med. 2000, 32, 600–607. [Google Scholar] [CrossRef]
- Buggage, R.R. Ocular manifestations of human T-cell lymphotropic virus type 1 infection. Curr. Opin. Ophthalmol. 2003, 14, 420–425. [Google Scholar] [CrossRef] [PubMed]
- Bittencourt, A.L.; de Oliveira, M.F. Cutaneous manifestations associated with HTLV-1 infection. Int. J. Dermatol. 2010, 49, 1099–1110. [Google Scholar] [CrossRef] [PubMed]
- Satou, Y.; Matsuoka, M. HTLV-1 and the host immune system: How the virus disrupts immune regulation, leading to HTLV-1 associated diseases. J. Clin. Exp. Hematop. 2010, 50, 1–8. [Google Scholar] [CrossRef]
- Kashanchi, F.; Brady, J.N. Transcriptional and post-transcriptional gene regulation of HTLV-1. Oncogene 2005, 24, 5938–5951. [Google Scholar] [CrossRef]
- Giam, C.Z.; Jeang, K.T. HTLV-1 Tax and adult T-cell leukemia. Front Biosci. 2007, 12, 1496–1507. [Google Scholar] [CrossRef]
- Nyborg, J.K.; Egan, D.; Sharma, N. The HTLV-1 Tax protein: Revealing mechanisms of transcriptional activation through histone acetylation and nucleosome disassembly. Biochim. Biophys. Acta 2010, 1799, 266–274. [Google Scholar] [CrossRef]
- Ciminale, V.; D'Agostino, D.; Zotti, L.; Franchini, G.; Felber, B.K.; Chieco-Bianchi, L. Expression and characterization of proteins produced by mRNAs spliced into the X region of the human T-cell leukemia/lymphotropic virus type II. Virology 1995, 209, 445–456. [Google Scholar] [CrossRef]
- Ciminale, V.; D'Agostino, D.M.; Zotti, L.; Chieco-Bianchi, L. Coding potential of the X region of human T-cell leukemia/lymphotropic virus type II. J. Acquir. Immune. Defic. Syndr. Hum. Retrovirol. 1996, 13, S220–S227. [Google Scholar] [CrossRef]
- Murata, K.; Hayashibara, T.; Sugahara, K.; Uemura, A.; Yamaguchi, T.; Harasawa, H.; Hasegawa, H.; Tsuruda, K.; Okazaki, T.; Koji, T.; et al. A novel alternative splicing isoform of human T-cell leukemia virus type 1 bZIP factor (HBZ-SI) targets distinct subnuclear localization. J. Virol. 2006, 80, 2495–2505. [Google Scholar] [CrossRef]
- Rende, F.; Cavallari, I.; Corradin, A.; Silic-Benussi, M.; Toulza, F.; Toffolo, G.M.; Tanaka, Y.; Jacobson, S.; Taylor, G.P.; D'Agostino, D.M.; et al. Kinetics and intracellular compartmentalization of HTLV-1 gene expression: nuclear retention of HBZ mRNA. Blood 2011, 117, 4855–4859. [Google Scholar] [CrossRef] [PubMed]
- Inlora, J.; Chukkapalli, V.; Derse, D.; Ono, A. Gag localization and virus-like particle release mediated by the matrix domain of human T-lymphotropic virus type 1 gag are less dependent on phosphatidylinositol-(4,5)-bisphosphate than those mediated by the matrix domain of HIV-1 gag. J. Virol. 2011, 85, 3802–3810. [Google Scholar] [CrossRef] [PubMed]
- Heidecker, G.; Lloyd, P.A.; Fox, K.; Nagashima, K.; Derse, D. Late assembly motifs of human T-cell leukemia virus type 1 and their relative roles in particle release. J. Virol. 2004, 78, 6636–6648. [Google Scholar] [CrossRef]
- Dorweiler, I.J.; Ruone, S.J.; Wang, H.; Burry, R.W.; Mansky, L.M. Role of the human T-cell leukemia virus type 1 PTAP motif in Gag targeting and particle release. J. Virol. 2006, 80, 3634–3643. [Google Scholar] [CrossRef]
- Heidecker, G.; Lloyd, P.A.; Soheilian, F.; Nagashima, K.; Derse, D. The role of WWP1-Gag interaction and Gag ubiquitination in assembly and release of human T-cell leukemia virus type 1. J. Virol. 2007, 81, 9769–9777. [Google Scholar] [CrossRef]
- Leblanc, I.; Rosenberg, A.R.; Dokhelar, M.C. Multiple functions for the basic amino acids of the human T-cell leukemia virus type 1 matrix protein in viral transmission. J. Virol. 1999, 73, 1860–1867. [Google Scholar] [CrossRef]
- Hemonnot, B.; Molle, D.; Bardy, M.; Gay, B.; Laune, D.; Devaux, C.; Briant, L. Phosphorylation of the HTLV-1 matrix L-domain-containing protein by virus-associated ERK-2 kinase. Virology 2006, 349, 430–439. [Google Scholar] [CrossRef]
- Kimata, J.T.; Wong, F.; Wang, J.; Ratner, L. Construction and characterization of infectious human T-cell leukemia virus type 1 molecular clones. Virology 1994, 204, 656–664. [Google Scholar] [CrossRef]
- Stewart-Maynard, K.M.; Cruceanu, M.; Wang, F.; Vo, M.N.; Gorelick, R.J.; Williams, M.C.; Rouzina, I.; Musier-Forsyth, K. Retroviral nucleocapsid proteins display non-equivalent levels of nucleic acid chaperone activity. J. Virol. 2008, 82, 10129–10142. [Google Scholar] [CrossRef]
- Qualley, D.F.; Stewart-Maynard, K.M.; Wang, F.; Mitra, M.; Gorelick, R.J.; Rouzina, I.; Williams, M.C.; Musier-Forsyth, K. C-terminal domain modulates the nucleic acid chaperone activity of human T-cell leukemia virus type 1 nucleocapsid protein via an electrostatic mechanism. J. Biol. Chem. 2010, 285, 295–307. [Google Scholar] [CrossRef]
- Derse, D.; Hill, S.A.; Princler, G.; Lloyd, P.; Heidecker, G. Resistance of human T cell leukemia virus type 1 to APOBEC3G restriction is mediated by elements in nucleocapsid. Proc. Natl. Acad. Sci. U. S. A. 2007, 104, 2915–2920. [Google Scholar] [CrossRef] [PubMed]
- Naka, H.; Teruya, K.; Bang, J.K.; Aimoto, S.; Tatsumi, T.; Konno, H.; Nosaka, K.; Akaji, K. Evaluations of substrate specificity and inhibition at PR/p3 cleavage site of HTLV-1 protease. Bioorg. Med. Chem. Lett. 2006, 16, 3761–3764. [Google Scholar] [CrossRef]
- Tozser, J.; Weber, I.T. The protease of human T-cell leukemia virus type-1 is a potential therapeutic target. Curr. Pharm. Des. 2007, 13, 1285–1294. [Google Scholar] [CrossRef]
- Derse, D.; Crise, B.; Li, Y.; Princler, G.; Lum, N.; Stewart, C.; McGrath, C.F.; Hughes, S.H.; Munroe, D.J.; Wu, X. Human T-cell leukemia virus type 1 integration target sites in the human genome: comparison with those of other retroviruses. J. Virol. 2007, 81, 6731–6741. [Google Scholar] [CrossRef]
- Seiki, M.; Hattori, S.; Hirayama, Y.; Yoshida, M. Human adult T-cell leukemia virus: Complete nucleotide sequence of the provirus genome integrated in leukemia cell DNA. Proc. Natl. Acad. Sci. U. S. A. 1983, 80, 3618–3622. [Google Scholar] [CrossRef]
- Tsujimoto, A.; Teruuchi, T.; Imamura, J.; Shimotohno, K.; Miyoshi, I.; Miwa, M. Nucleotide sequence analysis of a provirus derived from HTLV-1-associated myelopathy (HAM). Mol. Biol. Med. 1988, 5, 29–42. [Google Scholar]
- Pique, C.; Dokhelar, M.C. The HTLV-1 Envelope: An overview. In Genetic Structure and Regulation of HIV; Haseltine, W.A., Wongstaal, F., Eds.; Raven Press, Ltd.: New York, NY, USA, 1991; pp. 499–510. [Google Scholar]
- Hattori, S.; Kiyokawa, T.; Imagawa, K.; Shimizu, F.; Hashimura, E.; Seiki, M.; Yoshida, M. Identification of gag and env gene products of human T-cell leukemia virus (HTLV). Virology 1984, 136, 338–347. [Google Scholar] [CrossRef]
- Lee, T.H.; Coligan, J.E.; Homma, T.; McLane, M.F.; Tachibana, N.; Essex, M. Human T-cell leukemia virus-associated membrane antigens: identity of the major antigens recognized after virus infection. Proc. Natl. Acad. Sci. U. S. A. 1984, 81, 3856–3860. [Google Scholar] [CrossRef]
- Delamarre, L.; Rosenberg, A.R.; Pique, C.; Pham, D.; Callebaut, I.; Dokhelar, M.C. The HTLV-I envelope glycoproteins: Structure and functions. J. Acquir. Immune. Defic. Syndr. Hum. Retrovirol. 1996, 13, S85–S91. [Google Scholar] [CrossRef]
- Astier-Gin, T.; Portail, J.P.; Londos-Gagliardi, D.; Moynet, D.; Blanchard, S.; Dalibart, R.; Pouliquen, J.F.; Georges-Courbot, M.C.; Hajjar, C.; et al. Neutralizing activity and antibody reactivity toward immunogenic regions of the human T cell leukemia virus type I surface glycoprotein in sera of infected patients with different clinical states. J. Infect. Dis. 1997, 175, 716–719. [Google Scholar] [CrossRef]
- Tanaka, Y.; Tanaka, R.; Terada, E.; Koyanagi, Y.; Miyanokurosaki, N.; Yamamoto, N.; Baba, E.; Nakamura, M.; Shida, H. Induction of antibody responses that neutralize human T- cell leukemia virus type I infection in vitro and in vivo by peptide immunization. J. Virol. 1994, 68, 6323–6331. [Google Scholar] [CrossRef]
- Delamarre, L.; Pique, C.; Pham, D.; Tursz, T.; Dokhelar, M.C. Identification of functional regions in the human T-cell leukemia virus type I SU glycoprotein. J. Virol. 1994, 68, 3544–3549. [Google Scholar] [CrossRef] [PubMed]
- Le, B., I; Grange, M.P.; Delamarre, L.; Rosenberg, A.R.; Blot, V.; Pique, C.; Dokhelar, M.C. HTLV-1 structural proteins. Virus Res. 2001, 78, 5–16. [Google Scholar]
- Delamarre, L.; Rosenberg, A.R.; Pique, C.; Pham, D.; Dokhelar, M.C. A novel human T-Leukemia virus type 1 cell-to-cell transmission assay permits definition of SU glycoprotein amino acids important for infectivity. J. Virol. 1997, 71, 259–266. [Google Scholar] [CrossRef]
- Rosenberg, J.R.; Delamarre, L.; Pique, C.; Pham, D.; Dokhelar, M.C. The ectodomain of the human T-cell leukemia virus type 1 TM glycoprotein is involved in postfusion events. J. Virol. 1997, 71, 7180–7186. [Google Scholar] [CrossRef]
- Ilinskaya, A.; Heidecker, G.; Derse, D. Opposing effects of a tyrosine-based sorting motif and a PDZ-binding motif regulate human T-lymphotropic virus type 1 envelope trafficking. J. Virol. 2010, 84, 6995–7004. [Google Scholar] [CrossRef]
- Blot, V.; Delamarre, L.; Perugi, F.; Pham, D.; Benichou, S.; Benarous, R.; Hanada, T.; Chishti, A.H.; Dokhelar, M.C.; Pique, C. Human Dlg protein binds to the envelope glycoproteins of human T-cell leukemia virus type 1 and regulates envelope mediated cell-cell fusion in T lymphocytes. J. Cell Sci. 2004, 117, 3983–3993. [Google Scholar] [CrossRef]
- Yoshida, S.; Higuchi, M.; Shoji, T.; Yoshita, M.; Ishioka, K.; Takahashi, M.; Oie, M.; Tanaka, Y.; Uchiyama, M.; Fujii, M. Knockdown of synapse-associated protein Dlg1 reduces syncytium formation induced by human T-cell leukemia virus type 1. Virus Genes 2008, 37, 9–15. [Google Scholar] [CrossRef]
- Tsukahara, T.; Wielgosz, M.M.; Ratner, L. Characterization of envelope glycoprotein mutants for human T-cell leukemia virus type 1 infectivity and immortalization. J. Virol. 2001, 75, 9553–9559. [Google Scholar] [CrossRef]
- Silverman, L.R.; Phipps, A.J.; Montgomery, A.; Fernandez, S.; Tsukahara, T.; Ratner, L.; Lairmore, M.D. In vivo analysis of replication and immunogenicity of proviral clones of human T-lymphotropic virus type 1 with selective envelope surface-unit mutations. Blood 2005, 106, 3602–3608. [Google Scholar] [CrossRef]
- Jassal, S.R.; Pohler, R.G.; Brighty, D.W. Human T-cell leukemia virus type 1 receptor expression among syncytium- resistant cell lines revealed by a novel surface glycoprotein- immunoadhesin. J. Virol. 2001, 75, 8317–8328. [Google Scholar] [CrossRef] [PubMed]
- Hague, B.F.; Zhao, T.M.; Kindt, T.J. Binding of HTLV-1 virions to T cells occurs by a temperature and calcium-dependent process and is blocked by certain type 2 adenosine receptor antagonists. Virus Res. 2003, 93, 31–39. [Google Scholar] [CrossRef]
- Manel, N.; Kinet, S.; Battini, J.L.; Kim, F.J.; Taylor, N.; Sitbon, M. The HTLV receptor is an early T-cell activation marker whose expression requires de novo protein synthesis. Blood 2003, 101, 1913–1918. [Google Scholar] [CrossRef]
- Pinon, J.D.; Klasse, P.J.; Jassal, S.R.; Welson, S.; Weber, J.; Brighty, D.W.; Sattentau, Q.J. Human T-cell leukemia virus type 1 envelope glycoprotein gp46 interacts with cell surface heparan sulfate proteoglycans. J. Virol. 2003, 77, 9922–9930. [Google Scholar] [CrossRef]
- Sagara, Y.; Inoue, Y.; Tsujimura, M.; Kojima, E.; Shiraki, H.; Kashiwagi, S. Novel biomarker of HTLV-1-associated disease: Specific appearance of antibody recognizing the receptor-binding site on HTLV-1 envelope protein. Cancer Sci. 2004, 95, 835–839. [Google Scholar] [CrossRef]
- Manel, N.; Battini, J.L.; Taylor, N.; Sitbon, M. HTLV-1 tropism and envelope receptor. Oncogene 2005, 24, 6016–6025. [Google Scholar] [CrossRef]
- Ceccaldi, P.E.; Delebecque, F.; Prevost, M.C.; Moris, A.; Abastado, J.P.; Gessain, A.; Schwartz, O.; Ozden, S. DC-SIGN facilitates fusion of dendritic cells with human T-cell leukemia virus Type 1-infected cells. J. Virol. 2006, 80, 4771–4780. [Google Scholar] [CrossRef]
- Jones, K.S.; Fugo, K.; Petrow-Sadowski, C.; Huang, Y.; Bertolette, D.C.; Lisinski, I.; Cushman, S.W.; Jacobson, S.; Ruscetti, F.W. Human T-cell leukemia virus type 1 (HTLV-1) and HTLV-2 use different receptor complexes to enter T cells. J. Virol. 2006, 80, 8291–8302. [Google Scholar] [CrossRef]
- Ghez, D.; Lepelletier, Y.; Lambert, S.; Fourneau, J.M.; Blot, V.; Janvier, S.; Arnulf, B.; van Endert, P.M.; Heveker, N.; Pique, C.; et al. Neuropilin-1 is involved in human T-cell lymphotropic virus type 1 entry. J. Virol. 2006, 80, 6844–6854. [Google Scholar] [CrossRef]
- Jin, Q.; Agrawal, L.; Vanhorn-Ali, Z.; Alkhatib, G. GLUT-1-independent infection of the glioblastoma/astroglioma U87 cells by the human T cell leukemia virus type 1. Virology 2006, 353, 99–110. [Google Scholar] [CrossRef]
- Takenouchi, N.; Jones, K.S.; Lisinski, I.; Fugo, K.; Yao, K.; Cushman, S.W.; Ruscetti, F.W.; Jacobson, S. GLUT1 is not the primary binding receptor but is associated with cell-to-cell transmission of human T-cell leukemia virus type 1. J. Virol. 2007, 81, 1506–1510. [Google Scholar] [CrossRef] [PubMed]
- Jones, K.S.; Petrow-Sadowski, C.; Bertolette, D.C.; Huang, Y.; Ruscetti, F.W. Heparan sulfate proteoglycans mediate attachment and entry of human T-cell leukemia virus type 1 virions into CD4+ T cells. J. Virol. 2005, 79, 12692–12702. [Google Scholar] [CrossRef] [PubMed]
- Higuchi, M.; Fujii, M. Distinct functions of HTLV-1 Tax1 from HTLV-2 Tax2 contribute key roles to viral pathogenesis. Retrovirology 2009, 6, 117. [Google Scholar] [CrossRef] [PubMed]
- Hiscott, J.; Petropoulos, L.; Lacoste, J. Molecular interactions between HTLV-1 Tax protein and the NF-kappa B/kappa B transcription complex. [Review] [77 refs]. Virology 1995, 214, 3–11. [Google Scholar] [CrossRef] [PubMed]
- Peloponese, J.M., Jr.; Kinjo, T.; Jeang, K.T. Human T-cell leukemia virus type 1 Tax and cellular transformation. Int. J. Hematol. 2007, 86, 101–106. [Google Scholar] [CrossRef]
- Cann, A.J.; Rosenblatt, J.D.; Wachsman, W.; Shah, N.P.; Chen, I.S. Identification of the gene responsible for human T-cell leukaemia virus transcriptional regulation. Nature 1985, 318, 571–574. [Google Scholar] [CrossRef]
- Felber, B.K.; Paskalis, H.; Kleinmanewing, C. The pX Protein of HTLV-1 is a Transcriptional Activator of its Long Terminal Repeats. Science 1985, 229, 675–679. [Google Scholar] [CrossRef]
- Fujisawa, J.; Seiki, M.; Kiyokawa, T.; Yoshida, M. Functional activation of the long terminal repeat of human T-cell leukemia virus type I by a trans-acting factor. Proc. Natl. Acad. Sci. U. S. A. 1985, 82, 2277–2281. [Google Scholar] [CrossRef]
- Sodroski, J.G.; Rosen, C.A.; Haseltine, W.A. Trans-acting transcriptional activation of the long terminal repeat of human T lymphotropic viruses in infected cells. Science 1984, 223, 381–385. [Google Scholar] [CrossRef]
- Harrod, R.; Tang, Y.; Nicot, C.; Lu, H.S.; Vassilev, A.; Nakatani, Y.; Giam, C.Z. An exposed KID-like domain in human T-cell lymphotropic virus type 1 Tax is responsible for the recruitment of coactivators CBP/p300. Mol. Cell. Biol. 1998, 18, 5052–5061. [Google Scholar] [CrossRef]
- Zhao, L.J.; Giam, C.Z. Human T-cell lymphotropic virus type I (HTLV-I) transcriptional activator, Tax, enhances CREB binding to HTLV-I 21-base-pair repeats by protein-protein interaction. Proc. Natl. Acad. Sci. U. S. A. 1992, 89, 7070–7074. [Google Scholar] [CrossRef]
- Harrod, R.; Kuo, Y.L.; Tang, Y.; Yao, Y.; Vassilev, A.; Nakatani, Y.; Giam, C.Z. p300 and p300/cAMP-responsive element-binding protein associated factor interact with human T-cell lymphotropic virus type-1 Tax in a multi- histone acetyltransferase/activator-enhancer complex. J. Biol. Chem. 2000, 275, 11852–11857. [Google Scholar] [CrossRef]
- Jiang, H.; Lu, H.; Schiltz, R.L.; Pise-Masison, C.A.; Ogryzko, V.V.; Nakatani, Y.; Brady, J.N. PCAF interacts with tax and stimulates tax transactivation in a histone acetyltransferase-independent manner. Mol. Cell. Biol. 1999, 19, 8136–8145, Erratum: 2000, 20, 1897. [Google Scholar] [CrossRef]
- Perini, G.; Wagner, S.; Green, M.R. Recognition of bZIP proteins by the human T-cell leukaemia virus transactivator Tax. Nature 1995, 376, 602–605. [Google Scholar] [CrossRef]
- Maruyama, M.; Shibuya, H.; Harada, H.; Hatakeyama, M.; Seiki, M.; Fujita, T.; Inoue, J.; Yoshida, M.; Taniguchi, T. Evidence for aberrant activation of the interleukin-2 autocrine loop by HTLV-1-encoded p40x and T3/Ti complex triggering. Cell 1987, 48, 343–350. [Google Scholar] [CrossRef]
- Duyao, M.P.; Kessler, D.J.; Spicer, D.B.; Bartholomew, C.; Cleveland, J.L.; Siekevitz, M.; Sonenshein, G.E. Transactivation of the c-myc promoter by human T cell leukemia virus type 1 tax is mediated by NFkB. J. Biol. Chem. 1992, 267, 16288–16291. [Google Scholar] [CrossRef]
- Lemasson, I.; Roberthebmann, V.; Hamaia, S.; Dodon, M.D.; Gazzolo, L.; Devaux, C. Transrepression of lck gene expression by human T-cell leukemia virus type 1-encoded p40(tax). J. Virol. 1997, 71, 1975–1983. [Google Scholar] [CrossRef]
- Nicot, C.; Mahieux, R.; Takemoto, S.; Franchini, G. Bcl-X(L) is up-regulated by HTLV-I and HTLV-II in vitro and in ex vivo ATLL samples. Blood 2000, 96, 275–281. [Google Scholar] [CrossRef]
- Tsukahara, T.; Kannagi, M.; Ohashi, T.; Kato, H.; Arai, M.; Nunez, G.; Iwanaga, Y.; Yamamoto, N.; Ohtani, K.; Nakamura, M.; Fujii, M. Induction of Bcl-x(L) expression by human T-cell leukemia virus type 1 Tax through NF-kappaB in apoptosis-resistant T-cell transfectants with Tax. J. Virol. 1999, 73, 7981–7987. [Google Scholar] [CrossRef]
- Murakami, T.; Hirai, H.; Suzuki, T.; Fujisawa, J.I.; Yoshida, M. HTLV-1 tax enhances NF-kappa B2 expression and binds to the products p52 and p100, but does not suppress the inhibitory function of p100. Virology 1995, 206, 1066–1074. [Google Scholar] [CrossRef]
- Yoshida, M. Mechanism of transcriptional activation of viral and cellular genes by oncogenic protein of HTLV-1. Leukemia 1994, 8, S51–S53. [Google Scholar] [PubMed]
- Sun, S.C.; Elwood, J.; Beraud, C.; Greene, W.C. Human T-cell leukemia virus type I tax activation of NF- kappa B/Rel involves phosphorylation and degradation of I kappa B alpha and RelA (p65)-mediated induction of the c- rel gene. Mol. Cell. Biol. 1994, 14, 7377–7384. [Google Scholar] [CrossRef] [PubMed]
- Moriuchi, M.; Inoue, H.; Moriuchi, H. Reciprocal interactions between human T-lymphotropic virus type 1 and prostaglandins: implications for viral transmission. J. Virol. 2001, 75, 192–198. [Google Scholar] [CrossRef] [PubMed]
- Hieshima, K.; Nagakubo, D.; Nakayama, T.; Shirakawa, A.K.; Jin, Z.; Yoshie, O. Tax-inducible production of CC chemokine ligand 22 by human T cell leukemia virus type 1 (HTLV-1)-infected T cells promotes preferential transmission of HTLV-1 to CCR4-expressing CD4+ T cells. J. Immunol. 2008, 180, 931–939. [Google Scholar] [CrossRef] [PubMed]
- Barnard, A.L.; Igakura, T.; Tanaka, Y.; Taylor, G.P.; Bangham, C.R. Engagement of specific T cell surface molecules regulates cytoskeletal polarization in HTLV-1-infected lymphocytes. Blood 2005, 106, 988–995. [Google Scholar] [CrossRef]
- Moriuchi, M.; Moriuchi, H. Induction of lactoferrin gene expression in myeloid or mammary gland cells by human T-cell leukemia virus type 1 (HTLV-1) tax: Implications for milk-borne transmission of HTLV-1. J. Virol. 2006, 80, 7118–7126. [Google Scholar] [CrossRef]
- Inoue, J.; Itoh, M.; Akizawa, T.; Toyoshima, H.; Yoshida, M. HTLV-1 Rex protein accumulates unspliced RNA in the nucleus as well as in cytoplasm. Oncogene 1991, 6, 1753–1757. [Google Scholar]
- Nagashima, K.; Yoshida, M.; Seiki, M. A single species of pX mRNA of human T-cell leukemia virus type 1 encodes trans-activator p40x and two other phosphoproteins. J. Virol. 1986, 60, 394–399. [Google Scholar] [CrossRef]
- Younis, I.; Green, P.L. The human T-cell leukemia virus rex protein. Front. Biosci. 2005, 10, 431–445. [Google Scholar] [CrossRef]
- Inoue, J.; Seiki, M.; Yoshida, M. The second pX product p27 chi-III of HTLV-1 is required for gag gene expression. FEBS Lett. 1986, 209, 187–190. [Google Scholar] [CrossRef]
- Ballaun, C.; Farrington, G.K.; Dobrovnik, M.; Rusche, J.; Hauber, J.; Bohnlein, E. Functional analysis of human T-cell leukemia virus type 1 rex-response element: Direct RNA binding of Rex protein correlates with in vivo activity. J. Virol. 1991, 65, 4408–4413. [Google Scholar] [CrossRef]
- Bogerd, H.P.; Echarri, A.; Ross, T.M.; Cullen, B.R. Inhibition of human immunodeficiency virus Rev and human T- cell leukemia virus Rex function, but not Mason-Pfizer monkey virus constitutive transport element activity, by a mutant human nucleoporin targeted to Crm1. J. Virol. 1998, 72, 8627–8635. [Google Scholar] [CrossRef]
- Hakata, Y.; Umemoto, T.; Matsushita, S.; Shida, H. Involvement of human CRM1 (exportin 1) in the export and multimerization of the Rex protein of human T-cell leukemia virus type 1. J. Virol. 1998, 72, 6602–6607. [Google Scholar] [CrossRef]
- Kubota, S.; Hatanaka, M.; Pomerantz, R.J. Nucleo-cytoplasmic redistribution of the HTLV-I rex protein: Alterations by coexpression of the HTLV-I p21(x) protein. Virology 1996, 220, 502–507. [Google Scholar] [CrossRef]
- Ye, J.; Silverman, L.; Lairmore, M.D.; Green, P.L. HTLV-1 Rex is required for viral spread and persistence in vivo but is dispensable for cellular immortalization in vitro. Blood 2003, 102, 3963–3969. [Google Scholar] [CrossRef]
- Ye, J.; Xie, L.; Green, P.L. Tax and overlapping rex sequences do not confer the distinct transformation tropisms of human T-cell leukemia virus types 1 and 2. J. Virol. 2003, 77, 7728–7735. [Google Scholar] [CrossRef] [PubMed]
- Berneman, Z.N.; Gartenhaus, R.B.; Reitz, M.S.; Blattner, W.A.; Manns, A.; Hanchard, B.; Ikehara, O.; Gallo, R.C.; Klotman, M.E. Expression of alternatively spliced human T-lymphotropic virus type 1 pX mRNA in infected cell lines and in primary uncultured cells from patients with adult T-cell leukemia/lymphoma and healthy carriers. Proc. Natl. Acad. Sci. U. S. A. 1992, 89, 3005–3009. [Google Scholar] [CrossRef]
- Koralnik, I.J.; Gessain, A.; Klotman, M.E.; Lo, M.A.; Berneman, Z.N.; Franchini, G. Protein isoforms encoded by the pX region of human T-cell leukemia/lymphotropic virus type I. Proc. Natl. Acad. Sci U. S. A. 1992, 89, 8813–8817. [Google Scholar] [CrossRef]
- Derse, D.; Mikovits, J.; Ruscetti, F. X-I and X-II open reading frames of HTLV-I are not required for virus replication or for immortalization of primary T-cells in vitro. Virology 1997, 237, 123–128. [Google Scholar] [CrossRef]
- Collins, N.D.; Newbound, G.C.; Albrecht, B.; Beard, J.L.; Ratner, L.; Lairmore, M.D. Selective ablation of human T-cell lymphotropic virus type 1 p12I reduces viral infectivity in vivo. Blood 1998, 91, 4701–4707. [Google Scholar] [CrossRef]
- Bartoe, J.T.; Albrecht, B.; Collins, N.D.; Robek, M.D.; Ratner, L.; Green, P.L.; Lairmore, M.D. Functional role of pX open reading frame II of human T- lymphotropic virus type 1 in maintenance of viral loads in vivo. J. Virol. 2000, 74, 1094–1100. [Google Scholar] [CrossRef] [PubMed]
- Albrecht, B.; Collins, N.D.; Burniston, M.T.; Nisbet, J.W.; Ratner, L.; Green, P.L.; Lairmore, M.D. Human T-lymphotropic virus type 1 open reading frame I p12(I) is required for efficient viral infectivity in primary lymphocytes. J. Virol. 2000, 74, 9828–9835. [Google Scholar] [CrossRef] [PubMed]
- Silverman, L.R.; Phipps, A.J.; Montgomery, A.; Ratner, L.; Lairmore, M.D. Human T-cell lymphotropic virus type 1 open reading frame II-encoded p30II is required for in vivo replication: evidence of in vivo reversion. J. Virol. 2004, 78, 3837–3845. [Google Scholar] [CrossRef] [PubMed]
- Salahuddin, S.Z.; Markham, P.D.; Wong-Staal, F.; Franchini, G.; Kalyanaraman, V.S.; Gallo, R.C. Restricted expression of human T-cell leukemia—Lymphoma virus (HTLV) in transformed human umbilical cord blood lymphocytes. Virology 1983, 129, 51–64. [Google Scholar] [CrossRef] [PubMed]
- Valeri, V.W.; Hryniewicz, A.; Andresen, V.; Jones, K.; Fenizia, C.; Bialuk, I.; Chung, H.K.; Fukumoto, R.; Washington, P.R.; Ferrari, M.G.; et al. Requirement of the human T-cell leukemia virus p12 and p30 genes for infectivity of human dendritic cells and macaques but not rabbits. Blood 2010, 116, 3809–3817. [Google Scholar] [CrossRef]
- Kim, S.J.; Nair, A.M.; Fernandez, S.; Mathes, L.; Lairmore, M.D. Enhancement of LFA-1-mediated T cell adhesion by human T lymphotropic virus type 1 p12I1. J. Immunol. 2006, 176, 5463–5470. [Google Scholar] [CrossRef]
- Albrecht, B.; D'Souza, C.D.; Ding, W.; Tridandapani, S.; Coggeshall, K.M.; Lairmore, M.D. Activation of nuclear factor of activated T cells by human T-lymphotropic virus type 1 accessory protein p12(I). J. Virol. 2002, 76, 3493–3501. [Google Scholar] [CrossRef]
- Ding, W.; Albrecht, B.; Kelley, R.E.; Muthusamy, N.; Kim, S.J.; Altschuld, R.A.; Lairmore, M.D. Human T-cell lymphotropic virus type 1 p12(I) expression increases cytoplasmic calcium to enhance the activation of nuclear factor of activated T cells. J. Virol. 2002, 76, 10374–10382. [Google Scholar] [CrossRef]
- Zhang, W.; Nisbet, J.W.; Albrecht, B.; Ding, W.; Kashanchi, F.; Bartoe, J.T.; Lairmore, M.D. Human T-lymphotropic virus type 1 p30(II) regulates gene transcription by binding CREB binding protein/p300. J. Virol. 2001, 75, 9885–9895. [Google Scholar] [CrossRef]
- Lairmore, M.D.; Albrecht, B.; D'Souza, C.; Nisbet, J.W.; Ding, W.; Bartoe, J.T.; Green, P.L.; Zhang, W. In vitro and in vivo functional analysis of human T cell lymphotropic virus type 1 pX open reading frames I and II. AIDS Res. Hum. Retroviruses 2000, 16, 1757–1764. [Google Scholar] [CrossRef]
- Zhang, W.; Nisbet, J.W.; Bartoe, J.T.; Ding, W.; Lairmore, M.D. Human T-lymphotropic virus type 1 p30(II) functions as a transcription factor and differentially modulates CREB-responsive promoters. J. Virol. 2000, 74, 11270–11277. [Google Scholar] [CrossRef]
- Ciminale, V.; Pavlakis, G.N.; Derse, D.; Cunningham, C.P.; Felber, B.K. Complex splicing in the human T-cell leukemia virus (HTLV) family of retroviruses: Novel mRNAs and proteins produced by HTLV type I. J. Virol. 1992, 66, 1737–1745. [Google Scholar] [CrossRef]
- Chen, Y.M.; Chen, S.H.; Fu, C.Y.; Chen, J.Y.; Osame, M. Antibody reactivities to tumor-suppressor protein p53 and HTLV-I Tof, Rex and Tax in HTLV-I-infected people with differing clinical status. Int. J. Cancer 1997, 71, 196–202. [Google Scholar] [CrossRef]
- Pique, C.; Dokhelar, M.C. In vivo production of Rof and Tof proteins of HTLV type 1: Evidence from cytotoxic T lymphocytes. AIDS Res. Hum. Retroviruses 2000, 16, 1783–1786. [Google Scholar] [CrossRef]
- Princler, G.L.; Julias, J.G.; Hughes, S.H.; Derse, D. Roles of viral and cellular proteins in the expression of alternatively spliced HTLV-1 pX mRNAs. Virology 2003, 317, 136–145. [Google Scholar] [CrossRef]
- Trovato, R.; Mulloy, J.C.; Johnson, J.M.; Takemoto, S.; de Oliveira, M.P.; Franchini, G. A lysine-to-arginine change found in natural alleles of the human T-cell lymphotropic/leukemia virus type 1 p12(I) protein greatly influences its stability. J. Virol. 1999, 73, 6460–6467. [Google Scholar] [CrossRef]
- Ding, W.; Albrecht, B.; Luo, R.; Zhang, W.; Stanley, J.R.; Newbound, G.C.; Lairmore, M.D. Endoplasmic reticulum and cis-Golgi localization of human T- lymphotropic virus type 1 p12(I): Association with calreticulin and calnexin. J. Virol. 2001, 75, 7672–7682. [Google Scholar] [CrossRef]
- Koralnik, I.J.; Fullen, J.; Franchini, G. The p12I, p13II, and p30II proteins encoded by human T-cell leukemia/lymphotropic virus type I open reading frames I and II are localized in three different cellular compartments. J. Virol. 1993, 67, 2360–2366. [Google Scholar] [CrossRef]
- Franchini, G. Molecular mechanisms of human T-cell leukemia/lymphotropic virus type I infection. Blood 1995, 86, 3619–3639. [Google Scholar] [CrossRef]
- Kim, S.J.; Ding, W.; Albrecht, B.; Green, P.L.; Lairmore, M.D. A conserved calcineurin-binding motif in human T lymphotropic virus type 1 p12I functions to modulate nuclear factor of activated T cell activation. J. Biol. Chem. 2003, 278, 15550–15557. [Google Scholar] [CrossRef]
- Ding, W.; Kim, S.J.; Nair, A.M.; Michael, B.; Boris-Lawrie, K.; Tripp, A.; Feuer, G.; Lairmore, M.D. Human T-cell lymphotropic virus type 1 p12(I) enhances interleukin-2 production during T-cell activation. J. Virol. 2003, 77, 11027–11039. [Google Scholar] [CrossRef] [PubMed]
- Trovato, R.; Mulloy, J.C.; Fu, K.; Fullen, J.; Leonard, W.J.; Franchini, G. Mapping of binding sites between HTLV-p12(I) and beta- and gamma(c) chains of the interleukin 2 receptor (IL-2R). In Proceedings of the Cold Spring Harbor Retroviruses Meeting, Cold Spring Harbor, NY, USA, May,1997; Abstract book. p. 231. [Google Scholar]
- Nicot, C.; Mulloy, J.C.; Ferrari, M.G.; Johnson, J.M.; Fu, K.; Fukumoto, R.; Trovato, R.; Fullen, J.; Leonard, W.J.; Franchini, G. HTLV-1 p12(I) protein enhances STAT5 activation and decreases the interleukin-2 requirement for proliferation of primary human peripheral blood mononuclear cells. Blood 2001, 98, 823–829. [Google Scholar] [CrossRef] [PubMed]
- Franchini, G.; Mulloy, J.C.; Koralnik, I.J.; Lo, M.A.; Sparkowski, J.J.; Andresson, T.; Goldstein, D.J.; Schlegel, R. The human T-cell leukemia/lymphotropic virus type I p12I protein cooperates with the E5 oncoprotein of bovine papillomavirus in cell transformation and binds the 16-kilodalton subunit of the vacuolar H+ ATPase. J. Virol. 1993, 67, 7701–7704. [Google Scholar] [CrossRef] [PubMed]
- Koralnik, I.; Mulloy, J.C.; Andresson, T.; Fullen, J.; Franchini, G. Mapping of the intermolecular association of the human T-cell leukemia/lymphotropic virus type 1 p12I and the vacuolar H+ATPase 16 kDa subunit protein. J. Gen. Virol. 1995, 76, 1909–1916. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.M.; Nicot, C.; Fullen, J.; Ciminale, V.; Casareto, L.; Mulloy, J.C.; Jacobson, S.; Franchini, G. Free major histocompatibility complex class I heavy chain Is preferentially targeted for degradation by human T-cell leukemia/lymphotropic virus type 1 p12(I) protein. J. Virol. 2001, 75, 6086–6094. [Google Scholar] [CrossRef]
- Taylor, J.M.; Brown, M.; Nejmeddine, M.; Kim, K.J.; Ratner, L.; Lairmore, M.; Nicot, C. Novel role for interleukin-2 receptor-Jak signaling in retrovirus transmission. J. Virol. 2009, 83, 11467–11476. [Google Scholar] [CrossRef]
- Fukumoto, R.; Dundr, M.; Nicot, C.; Adams, A.; Valeri, V.W.; Samelson, L.E.; Franchini, G. Inhibition of T-cell receptor signal transduction and viral expression by the linker for activation of T cells-interacting p12(I) protein of human T-cell leukemia/lymphoma virus type 1. J. Virol. 2007, 81, 9088–9099. [Google Scholar] [CrossRef]
- Wake, A.; Tanaka, Y.; Nakatsuka, K.; Misago, M.; Oda, S.; Morimoto, I.; Eto, S. Calcium-dependent homotypic adhesion through leukocyte function-associated antigen-1/intracellular adhesion molecule-1 induces interleukin-1 and parathyroid hormone-related protein production on adult T-cell leukemia cells in vitro. Blood 1995, 86, 2257–2267. [Google Scholar] [CrossRef]
- Stewart, M.P.; Mcdowall, A.; Hogg, N. LFA-1-mediated adhesion is regulated by cytoskeletal restraint and by a Ca2+-dependent protease, calpain. J. Cell Biol. 1998, 140, 699–707. [Google Scholar] [CrossRef]
- Kaplan, M.J.; Beretta, L.; Yung, R.L.; Richardson, B.C. LFA-1 overexpression and T cell autoreactivity: mechanisms. Immunol. Invest. 2000, 29, 427–442. [Google Scholar]
- Basbous, J.; Arpin, C.; Gaudray, G.; Piechaczyk, M.; Devaux, C.; Mesnard, J.M. The HBZ factor of human T-cell leukemia virus type I dimerizes with transcription factors JunB and c-Jun and modulates their transcriptional activity. J. Biol. Chem. 2003, 278, 43620–43627. [Google Scholar] [CrossRef]
- Arnold, J.; Yamamoto, B.; Li, M.; Phipps, A.J.; Younis, I.; Lairmore, M.D.; Green, P.L. Enhancement of infectivity and persistence in vivo by HBZ, a natural antisense coded protein of HTLV-1. Blood 2006, 107, 3976–3982. [Google Scholar] [CrossRef]
- Koralnik, I.J.; Gessain, A.; Klotman, M.E.; Lo, M.A.; Berneman, Z.N.; Franchini, G. Protein isoforms encoded by the pX region of human T-cell leukemia/lymphotropic virus type I. Proc. Natl. Acad. Sci. U. S. A. 1992, 89, 8813–8817. [Google Scholar] [CrossRef]
- D'Agostino, D.M.; Ciminale, V.; Zotti, L.; Rosato, A.; Chieco-Bianchi, L. The human T-cell lymphotropic virus type 1 Tof protein contains a bipartite nuclear localization signal that is able to functionally replace the amino-terminal domain of Rex. J. Virol. 1997, 71, 75–83. [Google Scholar] [CrossRef]
- D'Agostino, D.M.; Zotti, L.; Ferro, T.; Cavallori, I.; Silic-Benussi, M.; Chieco-Bianchi, L.; Ciminale, V. Expression and functional properties of proteins encoded in the x-II ORF of HTLV-I. Virus Res. 2001, 78, 35–43. [Google Scholar] [CrossRef]
- Ciminale, V.; Zotti, L.; Dagostino, D.M.; Ferro, T.; Casareto, L.; Franchini, G.; Bernardi, P.; Chiecobianchi, L. Mitochondrial targeting of the p13(II) protein coded by the x-II ORF of human T-cell leukemia/lymphotropic virus type I (HTLV-I). Oncogene 1999, 18, 4505–4514. [Google Scholar] [CrossRef]
- D'Agostino, D.M.; Ranzato, L.; Arrigoni, G.; Cavallari, I.; Belleudi, F.; Torrisi, M.R.; Silic-Benussi, M.; Ferro, T.; Petronilli, V.; Marin, O.; Chieco-Bianchi, L.; Bernardi, P.; Ciminale, V. Mitochondrial alterations induced by the p13II protein of human T-cell leukemia virus type 1. Critical role of arginine residues. J. Biol. Chem. 2002, 277, 34424–34433. [Google Scholar] [CrossRef]
- Silic-Benussi, M.; Cavallari, I.; Zorzan, T.; Rossi, E.; Hiraragi, H.; Rosato, A.; Horie, K.; Saggioro, D.; Lairmore, M.D.; Willems, L.; Chieco-Bianchi, L.; D'Agostino, D.M.; Ciminale, V. Suppression of tumor growth and cell proliferation by p13II, a mitochondrial protein of human T cell leukemia virus type 1. Proc. Natl. Acad. Sci U. S. A. 2004, 101, 6629–6634. [Google Scholar] [CrossRef]
- Hiraragi, H.; Michael, B.; Nair, A.; Silic-Benussi, M.; Ciminale, V.; Lairmore, M. Human T-lymphotropic virus type 1 mitochondrion-localizing protein p13II sensitizes Jurkat T cells to Ras-mediated apoptosis. J. Virol. 2005, 79, 9449–9457. [Google Scholar] [CrossRef]
- Hiraragi, H.; Kim, S.J.; Phipps, A.J.; Silic-Benussi, M.; Ciminale, V.; Ratner, L.; Green, P.L.; Lairmore, M.D. Human T-lymphotropic virus type 1 mitochondrion-localizing protein p13II is required for viral infectivity in vivo. J. Virol. 2006, 80, 3469–3476. [Google Scholar] [CrossRef]
- Silic-Benussi, M.; Cavallari, I.; Vajente, N.; Vidali, S.; Chieco-Bianchi, L.; Di, L.F.; Saggioro, D.; D'Agostino, D.M.; Ciminale, V. Redox regulation of T-cell turnover by the p13 protein of human T-cell leukemia virus type 1: Distinct effects in primary versus transformed cells. Blood 2010, 116, 54–62. [Google Scholar] [CrossRef] [PubMed]
- Lefebvre, L.; Vanderplasschen, A.; Ciminale, V.; Heremans, H.; Dangoisse, O.; Jauniaux, J.C.; Toussaint, J.F.; Zelnik, V.; Burny, A.; Kettmann, R.; Willems, L. Oncoviral bovine leukemia virus G4 and human T-cell leukemia virus type 1 p13(II) accessory proteins interact with farnesyl pyrophosphate synthetase. J. Virol. 2002, 76, 1400–1414. [Google Scholar] [CrossRef] [PubMed]
- Lefebvre, L.; Ciminale, V.; Vanderplasschen, A.; D'Agostino, D.; Burny, A.; Willems, L.; Kettmann, R. Subcellular localization of the bovine leukemia virus R3 and G4 accessory proteins. J. Virol. 2002, 76, 7843–7854. [Google Scholar] [CrossRef] [PubMed]
- Ghorbel, S.; Sinha-Datta, U.; Dundr, M.; Brown, M.; Franchini, G.; Nicot, C. HTLV-I p30 nuclear/nucleolar retention is mediated through interactions with RNA and a constituent of the 60s ribosomal subunit. J. Biol. Chem. 2006, 281, 37150–37158. [Google Scholar] [CrossRef]
- Nicot, C.; Dundr, M.; Johnson, J.M.; Fullen, J.R.; Alonzo, N.; Fukumoto, R.; Princler, G.L.; Derse, D.; Misteli, T.; Franchini, G. HTLV-1-encoded p30II is a post-transcriptional negative regulator of viral replication. Nat. Med. 2004, 10, 197–201. [Google Scholar] [CrossRef]
- Younis, I.; Boris-Lawrie, K.; Green, P.L. Human T-cell leukemia virus open reading frame II encodes a posttranscriptional repressor that is recruited at the level of transcription. J. Virol. 2006, 80, 181–191. [Google Scholar] [CrossRef]
- Sinha-Datta, U.; Datta, A.; Ghorbel, S.; Duc, D.M.; Nicot, C. HTLV-I Rex and p30 interactions govern the switch between virus latency and replication. J. Biol. Chem. 2007, 282, 14608–14615. [Google Scholar] [CrossRef]
- Younis, I.; Khair, L.; Dundr, M.; Lairmore, M.D.; Franchini, G.; Green, P.L. Repression of human T-cell leukemia virus type 1 and type 2 replication by a viral mRNA-encoded posttranscriptional regulator. J. Virol. 2004, 78, 11077–11083. [Google Scholar] [CrossRef]
- Datta, A.; Silverman, L.; Phipps, A.J.; Hiraragi, H.; Ratner, L.; Lairmore, M.D. Human T-lymphotropic virus type-1 p30 alters cell cycle G2 regulation of T lymphocytes to enhance cell survival. Retrovirology 2007, 4, 49. [Google Scholar] [CrossRef]
- Anupam, R.; Datta, A.; Kesic, M.; Green-Church, K.; Shkriabai, N.; Kvaratskhelia, M.; Lairmore, M.D. Human T-lymphotropic virus type 1 p30 interacts with REG{gamma} and modulates ataxia telangiectasia mutated to promote cell survival. J. Biol. Chem. 2011, 286, 7661–7668. [Google Scholar] [CrossRef]
- Awasthi, S.; Sharma, A.; Wong, K.; Zhang, J.; Matlock, E.F.; Rogers, L.; Motloch, P.; Takemoto, S.; Taguchi, H.; Cole, M.D.; et al. A human T-cell lymphotropic virus type 1 enhancer of Myc transforming potential stabilizes Myc-TIP60 transcriptional interactions. Mol. Cell. Biol. 2005, 25, 6178–6198. [Google Scholar] [CrossRef]
- Baydoun, H.; Pancewicz, J.; Nicot, C. Human T-cell leukemia virus p30 inhibits homologous recombination and favors unfaithful DNA repair. Blood 2011. [CrossRef]
- Robek, M.D.; Wong, F.H.; Ratner, L. Human T-Cell leukemia virus type 1 pX-I and pX-II open reading frames are dispensable for the immortalization of primary lymphocytes. J. Virol. 1998, 72, 4458–4462. [Google Scholar] [CrossRef]
- Larocca, D.; Chao, L.A.; Seto, M.H.; Brunck, T.K. Human T-cell leukemia virus minus strand transcription in infected T-cells. Biochem. Biophys. Res. Commun. 1989, 163, 1006–1013. [Google Scholar] [CrossRef]
- Gaudray, G.; Gachon, F.; Basbous, J.; Biard-Piechaczyk, M.; Devaux, C.; Mesnard, J.M. The complementary strand of the human T-cell leukemia virus type 1 RNA genome encodes a bZIP transcription factor that down-regulates viral transcription. J. Virol. 2002, 76, 12813–12822. [Google Scholar] [CrossRef]
- Hivin, P.; Frederic, M.; rpin-Andre, C.; Basbous, J.; Gay, B.; Thebault, S.; Mesnard, J.M. Nuclear localization of HTLV-I bZIP factor (HBZ) is mediated by three distinct motifs. J. Cell Sci. 2005, 118, 1355–1362. [Google Scholar] [CrossRef]
- Arnold, J.; Zimmerman, B.; Li, M.; Lairmore, M.D.; Green, P.L. Human T-cell Leukemia virus type-1 antisense-encoded gene, Hbz, promotes T lymphocyte proliferation. Blood 2008, 112, 3788–3797. [Google Scholar] [CrossRef] [PubMed]
- Thebault, S.; Basbous, J.; Hivin, P.; Devaux, C.; Mesnard, J.M. HBZ interacts with JunD and stimulates its transcriptional activity. FEBS Lett. 2004, 562, 165–170. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, J.; Ohshima, T.; Isono, O.; Shimotohno, K. HTLV-1 HBZ suppresses AP-1 activity by impairing both the DNA-binding ability and the stability of c-Jun protein. Oncogene 2005, 24, 1001–1010. [Google Scholar] [CrossRef] [PubMed]
- Satou, Y.; Yasunaga, J.; Zhao, T.; Yoshida, M.; Miyazato, P.; Takai, K.; Shimizu, K.; Ohshima, K.; Green, P.L.; Ohkura, N.; et al. HTLV-1 bZIP factor induces T-cell lymphoma and systemic inflammation in vivo. PLoS Pathog. 2011, 7, e1001274. [Google Scholar] [CrossRef]
- Lairmore, M.D.; Silverman, L.; Ratner, L. Animal models for human T-lymphotropic virus type 1 (HTLV-1) infection and transformation. Oncogene 2005, 24, 6005–6015. [Google Scholar] [CrossRef]
- Akagi, T.; Takeda, I.; Oka, T.; Ohtsuki, Y.; Yano, S.; Miyoshi, I. Experimental infection of rabbits with human T-cell leukemia virus type 1. Jpn. J. Cancer Res. 1985, 76, 86–94. [Google Scholar]
- Lairmore, M.D.; Roberts, B.; Frank, D.; Rovnak, J.; Weiser, M.G.; Cockerell, G.L. Comparative biological responses of rabbits infected with human T-lymphotropic virus Type I isolates from patients with lymphoproliferative and neurodegenerative disease. Int. J. Cancer 1992, 50, 124–130. [Google Scholar] [CrossRef]
- Murata, N.; Hakoda, E.; Machida, H.; Ikezoe, T.; Sawada, T.; Hoshino, H.; Miyoshi, I. Prevention of human T cell lymphotropic virus type 1 infection in Japanese macaques by passive immunization. Leukemia 1996, 10, 1971–1974. [Google Scholar]
- Nakamura, H.; Hayami, M.; Ohta, Y.; Ishikawa, K.; Tsujimoto, H.; Kiyokawa, T.; Yoshida, M.; Sasagawa, A.; Honjo, S. Protection of cynomolgus monkeys against infection by human T-cell leukemia virus type-1 by immunization with viral env gene products produced in escerichia coli. Int. J. Cancer 1987, 40, 403–407. [Google Scholar] [CrossRef]
- Ibrahim, F.; Fiette, L.; Gessain, A.; Buisson, N.; Dethe, G.; Bomford, R. Infection of rats with human T-cell leukemia virus type-1: Susceptibility of inbred strains, antibody response and provirus location. Int. J. Cancer 1994, 58, 446–451. [Google Scholar] [CrossRef]
- Suga, T.; Kameyama, T.; Shimotohno, K.; Matsumura, M.; Tanaka, H.; Kushida, S.; Ami, Y.; Uchida, M.; Uchida, K.; Miwa, M. Infectiion of Rats with HTLV-1: A Small-Animal Model for HTLV-1 Carriers. Int. J. Cancer 1991, 49, 764–769. [Google Scholar] [CrossRef]
- Miyoshi, I.; Yoshimoto, S.; Kubonishi, I.; Fujishita, M.; Ohtsuki, Y.; Yamashita, M.; Yamato, K.; Hirose, S.; Taguchi, H.; Niiya, K. Infectious transmission of human T-cell leukemia virus to rabbits. Int. J. Cancer 1985, 35, 81–85. [Google Scholar] [CrossRef]
- Kotani, S.; Yoshimoto, S.; Yamato, K.; Fujishita, M.; Yamashita, M.; Ohtsuki, Y.; Taguchi, H.; Miyoshi, I. Serial transmission of human T-cell leukemia virus type 1 by blood transfusion in rabbits and its prevention by use of X-irradiated stored blood. Int. J. Cancer 1986, 37, 843–847. [Google Scholar] [CrossRef]
- Iwahara, Y.; Takehara, N.; Kataoka, R.; Sawada, T.; Ohtsuki, Y.; Nakachi, H.; Maehama, T.; Okayama, T.; Miyoshi, I. Transmission of HTLV-1 to rabbits via semen and breast milk from seropositive healthy persons. Int. J. Cancer 1990, 45, 980–983. [Google Scholar] [CrossRef]
- Kataoka, R.; Takehara, N.; Iwahara, Y.; Sawada, T.; Ohtsuki, Y.; Dawei, Y.; Hoshino, H.; Miyoshi, I. Transmission of HTLV-I by blood transfusion and its prevention by passive immunization in rabbits. Blood 1990, 76, 1657–1661. [Google Scholar] [CrossRef] [PubMed]
- Uemura, Y.; Kotani, S.; Yoshimoto, S.; Fujishita, M.; Yamashita, M.; Ohtsuki, Y.; Taguchi, H.; Miyoshi, I. Mother-to-offspring transmission of human T cell leukemia virus type I in rabbits. Blood 1987, 69, 1255–1258. [Google Scholar] [CrossRef] [PubMed]
- Uemura, Y.; Kotani, S.; Yoshimoto, S.; Fujishita, M.; Yano, S.; Ohtsuki, Y.; Miyoshi, I. Oral transmission of human T-cell leukemia virus type-1 in the rabbit. Jpn. J. Cancer Res. 1986, 77, 970–973. [Google Scholar] [PubMed]
- Hirose, S.; Kotani, S.; Uemura, Y.; Fujishita, M.; Taguchi, H.; Ohtsuki, Y.; Miyoshi, I. Milk-borne transmission of human T-cell leukemia virus type I in rabbits. Virology 1988, 162, 487–489. [Google Scholar] [CrossRef]
- Takehara, N.; Iwahara, Y.; Uemura, Y.; Sawada, T.; Ohtsuki, Y.; Iwai, H.; Hoshino, H.; Miyoshi, I. Effect of Immunization on HTLV-1 Infection in Rabbits. Int. J. Cancer 1989, 44, 332–336. [Google Scholar] [CrossRef]
- Sawada, T.; Iwahara, Y.; Ishii, K.; Taguchi, H.; Hoshino, H.; Miyoshi, I. Immunoglobulin prophylaxis against milkborne transmission of human T cell leukemia virus type 1 in rabbits. J. Infect. Dis. 1991, 164, 1193–1196. [Google Scholar] [CrossRef]
- Miyoshi, I.; Takehara, N.; Sawada, T.; Iwahara, Y.; Kataoka, R.; Yang, D.; Hoshino, H. Immunoglobulin prophylaxis against HTLV-I in a rabbit model. Leukemia 1992, 6, 24–26. [Google Scholar]
- Cockerell, G.L.; Lairmore, M.D.; De, B.; Rovnak, J.; Hartley, T.; Miyoshi, I. Persistent infection of rabbits with HTLV-I: Patterns of anti-viral reactivity and detection of virus by gene amplification. Int. J. Cancer 1990, 45, 127–130. [Google Scholar] [CrossRef]
- Lal, R.; Rudolph, D.; Palker, T.; Colligan, J.; Folks, T. A synthetic peptide elicits antibodies reactive with the external glycoprotein of HTLV-I. J. Gen. Virol. 1991, 72, 2321–2324. [Google Scholar] [CrossRef]
- Tanaka, Y.; Zeng, L.; Shiraki, H.; Shida, H.; Tozawa, H. Identification of a neutralization epitope on the envelope gp46 antigen of human T-cell leukemia virus type I and induction of neutralizing antibody by peptide immunization. J. Immunol. 1991, 147, 354–360. [Google Scholar] [CrossRef]
- Chen, Y.; Lee, T.; Samuel, K.; Okayama, A.; Tachibana, N.; Miyoshi, I.; Papas, T.; Essex, M. Delineation of type-specific regions on the envelope glycoproteins of human T cell leukemia viruses. J. Immunol. 1991, 147, 2368–2376. [Google Scholar] [CrossRef]
- Haynes, R.A.; Zimmerman, B.; Millward, L.; Ware, E.; Premanandan, C.; Yu, L.; Phipps, A.J.; Lairmore, M.D. Early spatial and temporal events of human T-lymphotropic virus type 1 spread following blood-borne transmission in a rabbit model of infection. J. Virol. 2010, 84, 5124–5130. [Google Scholar] [CrossRef]
- Derse, D.; Mikovits, J.; Polianova, M.; Felber, B.K.; Ruscetti, F. Virions released from cells transfected with a molecular clone of human T-cell leukemia virus type I give rise to primary and secondary infections of T cells. J. Virol. 1995, 69, 1907–1912. [Google Scholar] [CrossRef]
- Zhao, T.M.; Robinson, M.A.; Bowers, F.S.; Kindt, T.J. Characterization of an infectious molecular clone of human T- cell leukemia virus type I. J. Virol. 1995, 69, 2024–2030. [Google Scholar] [CrossRef]
- Collins, N.D.; Newbound, G.C.; Ratner, L.; Lairmore, M.D. In vitro CD4(+) lymphocyte transformation and infection in a rabbit model with a molecular clone of human T-cell lymphotropic virus type 1. J. Virol. 1996, 70, 7241–7246. [Google Scholar] [CrossRef]
- Haynes, R.A.; Ware, E.; Premanandan, C.; Zimmerman, B.; Yu, L.; Phipps, A.J.; Lairmore, M.D. Cyclosporine-induced immune suppression alters establishment of HTLV-1 infection in a rabbit model. Blood 2010, 115, 815–823. [Google Scholar] [CrossRef]
- Bangham, C.R. CTL quality and the control of human retroviral infections. Eur. J. Immunol. 2009, 39, 1700–1712. [Google Scholar] [CrossRef]
- MacNamara, A.; Rowan, A.; Hilburn, S.; Kadolsky, U.; Fujiwara, H.; Suemori, K.; Yasukawa, M.; Taylor, G.; Bangham, C.R.; Asquith, B. HLA class I binding of HBZ determines outcome in HTLV-1 infection. PLoS Pathog. 2010, 6, e1001117. [Google Scholar] [CrossRef]
- Hilburn, S.; Rowan, A.; Demontis, M.A.; MacNamara, A.; Asquith, B.; Bangham, C.R.; Taylor, G.P. In vivo expression of human T-lymphotropic virus type 1 basic leucine-zipper protein generates specific CD8+ and CD4+ T-lymphocyte responses that correlate with clinical outcome. J. Infect. Dis. 2011, 203, 529–536. [Google Scholar] [CrossRef]
- Zimmerman, B.; Niewiesk, S.; Lairmore, M.D. Mouse models of human T lymphotropic virus type-1-associated adult T-cell leukemia/lymphoma. Vet. Pathol. 2010, 47, 677–689. [Google Scholar] [CrossRef] [PubMed]
- Nitta, T.; Tanaka, M.; Sun, B.; Hanai, S.; Miwa, M. The genetic background as a determinant of human T-cell leukemia virus type 1 proviral load. Biochem. Biophys. Res. Commun. 2003, 309, 161–165. [Google Scholar] [CrossRef] [PubMed]
- Furuta, R.A.; Sugiura, K.; Kawakita, S.; Inada, T.; Ikehara, S.; Matsuda, T.; Fujisawa, J.J. Mouse model for the equilibration interaction between the host immune system and human T-cell leukemia virus type 1 gene expression. J. Virol. 2002, 76, 2703–2713. [Google Scholar] [CrossRef]
- Feng, R.; Kabayama, A.; Uchida, K.; Hoshino, H.; Miwa, M. Cell-free entry of human t-cell leukemia virus type 1 to mouse cells. Jpn. J. Cancer Res. 2001, 92, 410–416. [Google Scholar] [CrossRef] [PubMed]
- Fang, J.; Kushida, S.; Feng, R.; Tanaka, M.; Kawamura, T.; Abe, H.; Maeda, N.; Onobori, M.; Hori, M.; Uchida, K.; Miwa, M. Transmission of human T-cell leukemia virus type 1 to mice. J. Virol. 1998, 72, 3952–3957. [Google Scholar] [CrossRef]
- Gabet, A.S.; Gessain, A.; Wattel, E. High simian T-cell leukemia virus type 1 proviral loads combined with genetic stability as a result of cell-associated provirus replication in naturally infected, asymptomatic monkeys. Int. J. Cancer 2003, 107, 74–83. [Google Scholar] [CrossRef]
- Takemura, T.; Yamashita, M.; Shimada, M.K.; Ohkura, S.; Shotake, T.; Ikeda, M.; Miura, T.; Hayami, M. High prevalence of simian T-lymphotropic virus type L in wild ethiopian baboons. J. Virol. 2002, 76, 1642–1648. [Google Scholar] [CrossRef]
- Gessain, A.; Mahieux, R.; De, T.G. Genetic variability and molecular epidemiology of human and simian T cell leukemia/lymphoma virus type I. J. Acquir. Immune. Defic. Syndr. Hum. Retrovirol. 1996, 13, S132–S145. [Google Scholar] [CrossRef]
- Kazanji, M.; Tartaglia, J.; Franchini, G.; de Thoisy, B.; Talarmin, A.; Contamin, H.; Gessain, A.; de The, G. Immunogenicity and protective efficacy of recombinant human T-cell leukemia/lymphoma virus type 1 NYVAC and naked DNA vaccine candidates in squirrel monkeys (Saimiri sciureus). J. Virol. 2001, 75, 5939–5948. [Google Scholar] [CrossRef]
- Kazanji, M. HTLV type 1 infection in squirrel monkeys (Saimiri sciureus): A promising animal model for HTLV type 1 human infection. AIDS Res. Hum. Retroviruses 2000, 16, 1741–1746. [Google Scholar] [CrossRef]
- Sundaram, R.; Lynch, M.P.; Rawale, S.V.; Sun, Y.; Kazanji, M.; Kaumaya, P.T. De novo design of peptide immunogens that mimic the coiled coil region of human T-cell leukemia virus type-1 glycoprotein 21 transmembrane subunit for induction of native protein reactive neutralizing antibodies. J. Biol. Chem. 2004, 279, 24141–24151. [Google Scholar] [CrossRef]
- Hakata, Y.; Yamada, M.; Shida, H. Rat CRM1 is responsible for the poor activity of human T-cell leukemia virus type 1 Rex protein in rat cells. J. Virol. 2001, 75, 11515–11525. [Google Scholar] [CrossRef] [PubMed]
- Kannagi, M.; Ohashi, T.; Hanabuchi, S.; Kato, H.; Koya, Y.; Hasegawa, A.; Masuda, T.; Yoshiki, T. Immunological aspects of rat models of HTLV type 1-infected T lymphoproliferative disease. AIDS Res. Hum. Retroviruses 2000, 16, 1737–1740. [Google Scholar] [CrossRef]
- Kasai, T.; Ikeda, H.; Tomaru, U.; Yamashita, I.; Ohya, O.; Morita, K.; Wakisaka, A.; Matsuoka, E.; Moritoyo, T.; Hashimoto, K.; et al. A rat model of human T lymphocyte virus type I (HTLV-I) infection: In situ detection of HTLV-I provirus DNA in microglia/macrophages in affected spinal cords of rats with HTLV-1-induced chronic progressive myeloneuropathy. Acta Neuropathol. 1999, 97, 107–112. [Google Scholar] [CrossRef]
- Sun, B.; Fang, J.; Yagami, K.; Kushida, S.; Tanaka, M.; Uchida, K.; Miwa, M. Age-dependent paraparesis in WKA rats: evaluation of MHC k-haplotype and HTLV-1 infection. J. Neurol. Sci. 1999, 167, 16–21. [Google Scholar] [CrossRef]
- Ishiguro, N.; Abe, M.; Seto, K.; Sakurai, H.; Ikeda, H.; Wakisaka, A.; Togashi, T.; Tateno, M.; Yoshiki, T. A rat model of human T lymphocyte virus type 1 (HTLV-1) infection. 1. Humoral antibody response, provirus integration, and YTLV-1-associated myelopathy/tropical spastic paraparesis-like myelopathy in seronegative HTLV-1 carrier rats. J. Exp. Med. 1992, 176, 981–989. [Google Scholar] [CrossRef]
- Hasegawa, A.; Ohashi, T.; Hanabuchi, S.; Kato, H.; Takemura, F.; Masuda, T.; Kannagi, M. Expansion of human T-cell leukemia virus type 1 (HTLV-1) reservoir in orally infected rats: Inverse correlation with HTLV-1-specific cellular immune response. J. Virol. 2003, 77, 2956–2963. [Google Scholar] [CrossRef]
- Willems, L.; Burny, A.; Collete, D.; Dangoisse, O.; Dequiedt, F.; Gatot, J.S.; Kerkhofs, P.; Lefebvre, L.; Merezak, C.; Peremans, T.; et al. Genetic determinants of bovine leukemia virus pathogenesis. AIDS Res. Hum. Retroviruses 2000, 16, 1787–1795. [Google Scholar] [CrossRef]
- Gillet, N.; Florins, A.; Boxus, M.; Burteau, C.; Nigro, A.; Vandermeers, F.; Balon, H.; Bouzar, A.B.; Defoiche, J.; Burny, A.; et al. Mechanisms of leukemogenesis induced by bovine leukemia virus: prospects for novel anti-retroviral therapies in human. Retrovirology 2007, 4, 18. [Google Scholar] [CrossRef]
- Florins, A.; Gillet, N.; Asquith, B.; Boxus, M.; Burteau, C.; Twizere, J.C.; Urbain, P.; Vandermeers, F.; Debacq, C.; Sanchez-Alcaraz, M.T.; et al. Cell dynamics and immune response to BLV infection: a unifying model. Front. Biosci. 2007, 12, 1520–1531. [Google Scholar] [CrossRef]
- Kushida, S.; Matsumura, M.; Tanaka, H.; Ami, Y.; Hori, M.; Kobayashi, M.; Uchida, K.; Yagami, K.; Kameyama, T.; Yoshizawa, T.; et al. HTLV-1-associated myelopathy/tropical spastic paraparesis-like rats. Jpn. J. Cancer Res. 1993, 84, 831–833. [Google Scholar] [CrossRef]
- Yoshiki, T. Chronic progressive myeloneuropathy in WKAH rats induced by HTLV- I infection as an animal model for HAM/TSP in humans. Intervirology 1995, 38, 229–237. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, N.; Hayami, M.; Komuro, A.; Schneider, J.; Hunsmann, G.; Okada, M.; Hinuma, Y. Experimental infection of cynomolgus monkeys with a human retrovirus, adult T-cell leukemia virus. Med. Microbiol. Immunol. (Berl.) 1984, 173, 57–64. [Google Scholar] [CrossRef]
- Miyoshi, I.; Yoshimoto, S.; Fujishita, M.; Kubonishi, I.; Taguchi, H.; Ohtsuki, Y. Infectious transmission of human T-cell leukemia virus to animals. Princess Takamatsu Symp. 1984, 15, 121–127. [Google Scholar] [PubMed]
- Nakamura, H.; Tanaka, Y.; Tsujimoto, A.K.; Ishikawa, K.; Takadaya, K.I.; Tozawa, H.; Tsujimoto, H.; Honjo, S.; Hayami, M. Experimental inoculation of monkeys with autologous lymphoid cell lines immortalized by and producing human T-cell leukemia virus type-I. Int. J. Cancer 1986, 38, 867–875. [Google Scholar] [CrossRef] [PubMed]
- Kazanji, M.; Moreau, J.P.; Mahieux, R.; Bonnemains, B.; Bomford, R.; Gessain, A.; Dethe, G. HTLV-I infection in squirrel monkeys (Saimiri sciureus) using autologous, homologous, or heterologous HTLV-I- transformed cell lines. Virology 1997, 231, 258–266. [Google Scholar] [CrossRef]
- Kazanji, M.; Ureta-Vidal, A.; Ozden, S.; Tangy, F.; de Thoisy, B.; Fiette, L.; Talarmin, A.; Gessain, A.; de The, G. Lymphoid organs as a major reservoir for human T-cell leukemia virus type 1 in experimentally infected squirrel monkeys (Saimiri sciureus): Provirus expression, persistence, and humoral and cellular immune responses. J. Virol. 2000, 74, 4860–4867. [Google Scholar] [CrossRef]
- Beilke, M.A.; Traina-Dorge, V.; England, J.D.; Blanchard, J.L. Polymyositis, arthritis, and uveitis in a macaque experimentally infected with human T lymphotropic virus type I. Arthritis Rheumatism. 1996, 39, 610–615. [Google Scholar] [CrossRef]
- McGinn, T.M.; Tao, B.; Cartner, S.; Schoeb, T.; Davis, I.; Ratner, L.; Fultz, P.N. Association of primate T-cell lymphotropic virus infection of pig-tailed macaques with high mortality. Virology 2002, 304, 364–378. [Google Scholar] [CrossRef]
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Lairmore, M.D.; Anupam, R.; Bowden, N.; Haines, R.; Haynes II, R.A.H.; Ratner, L.; Green, P.L. Molecular Determinants of Human T-lymphotropic Virus Type 1 Transmission and Spread. Viruses 2011, 3, 1131-1165. https://doi.org/10.3390/v3071131
Lairmore MD, Anupam R, Bowden N, Haines R, Haynes II RAH, Ratner L, Green PL. Molecular Determinants of Human T-lymphotropic Virus Type 1 Transmission and Spread. Viruses. 2011; 3(7):1131-1165. https://doi.org/10.3390/v3071131
Chicago/Turabian StyleLairmore, Michael D., Rajaneesh Anupam, Nadine Bowden, Robyn Haines, Rashade A. H. Haynes II, Lee Ratner, and Patrick L. Green. 2011. "Molecular Determinants of Human T-lymphotropic Virus Type 1 Transmission and Spread" Viruses 3, no. 7: 1131-1165. https://doi.org/10.3390/v3071131
APA StyleLairmore, M. D., Anupam, R., Bowden, N., Haines, R., Haynes II, R. A. H., Ratner, L., & Green, P. L. (2011). Molecular Determinants of Human T-lymphotropic Virus Type 1 Transmission and Spread. Viruses, 3(7), 1131-1165. https://doi.org/10.3390/v3071131




