Deficiency of IFNAR1 Increases the Production of Influenza Vaccine Viruses in MDCK Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells and Virus
2.2. IFNAR1 Knockdown in MDCK Cells
2.3. RNA Extraction and Reverse Transcription
2.4. Quantitative Real-Time PCR (qPCR)
2.5. Construction of Plasmids Targeting Canine IFNAR1
2.6. Generation of IFNAR1-KO Cell Lines
2.7. PCR and Sequencing
2.8. Cell Density Curve Assay
2.9. Western Blot
2.10. Determination of Virus Titer
2.11. RNA-Seq and Data Analysis
2.12. Statistical Analysis
3. Results
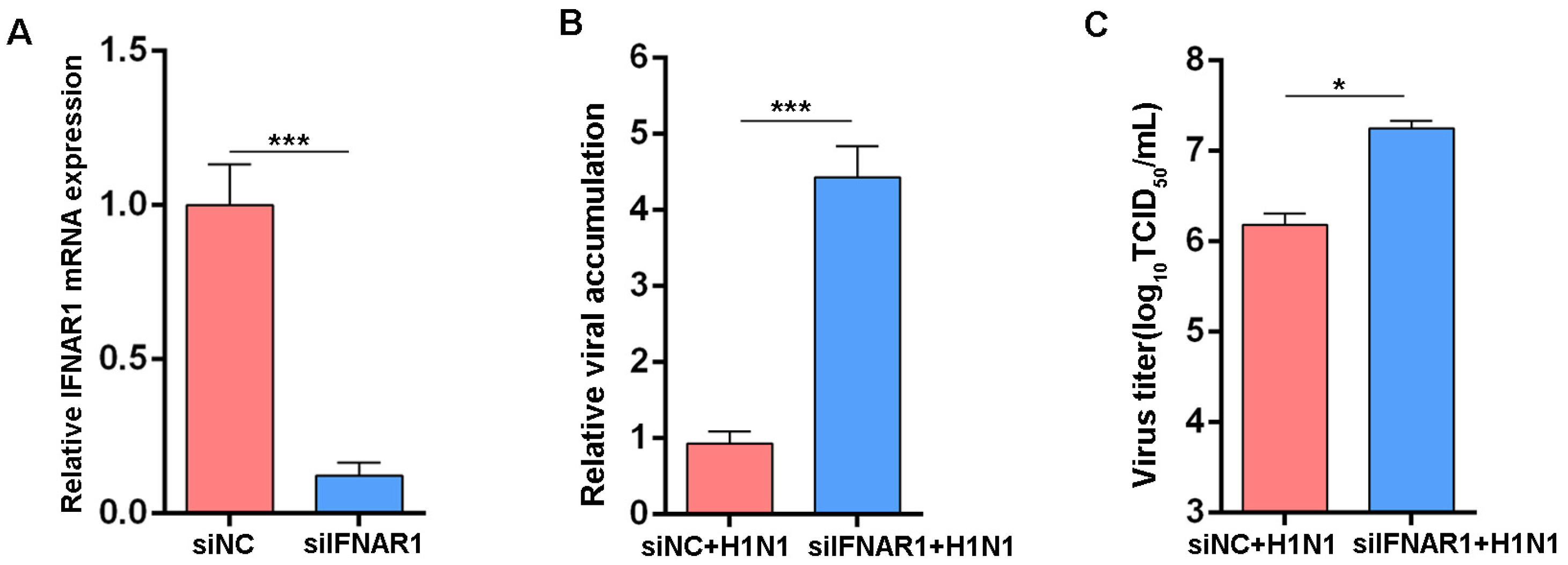
3.1. Knockdown of IFNAR1 Enhances Influenza Virus Replication in MDCK Cells
3.2. Generation of IFNAR1-KO MDCK Cell Line
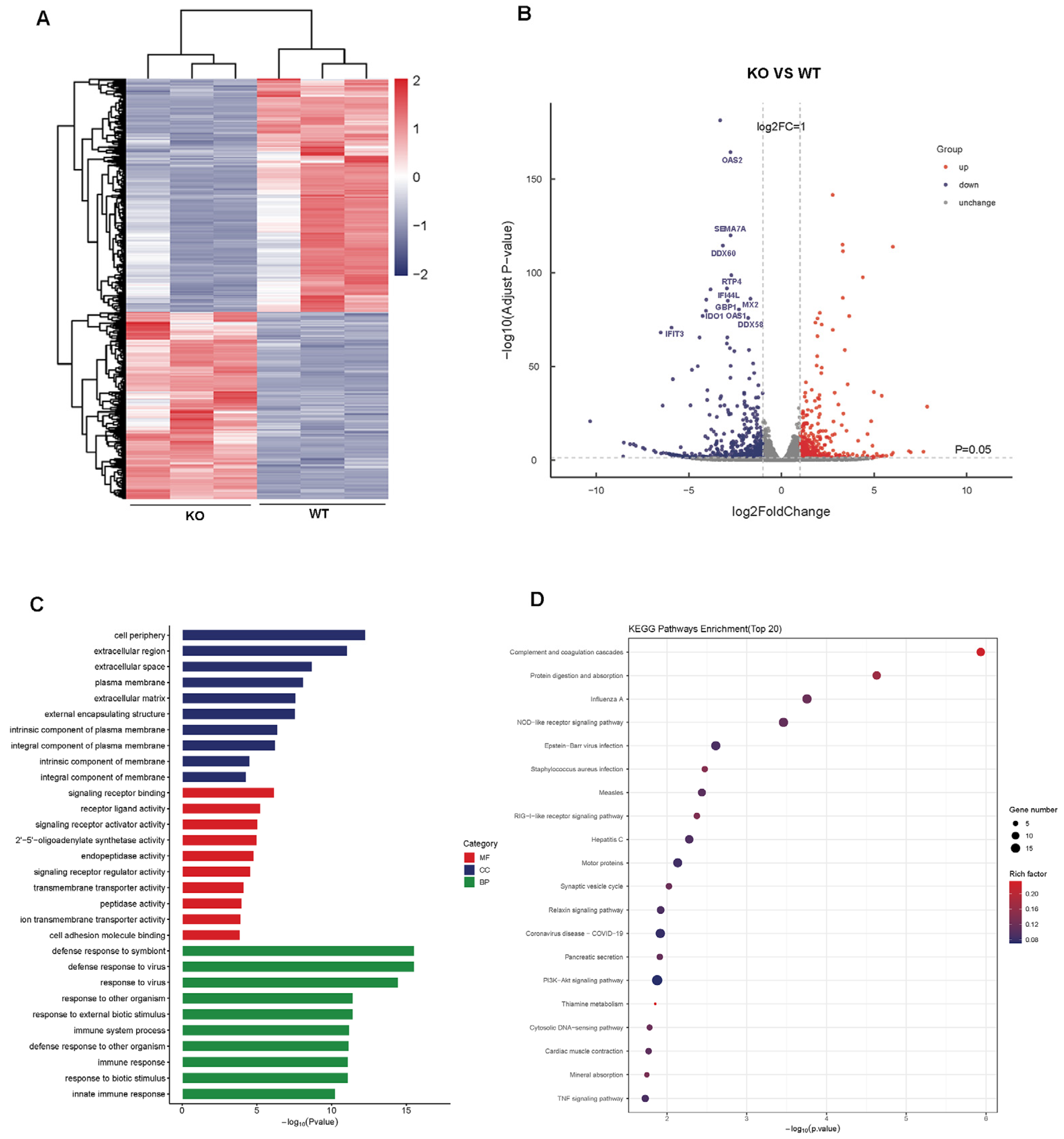
3.3. Transcriptomic Analysis of KO vs. WT Cells
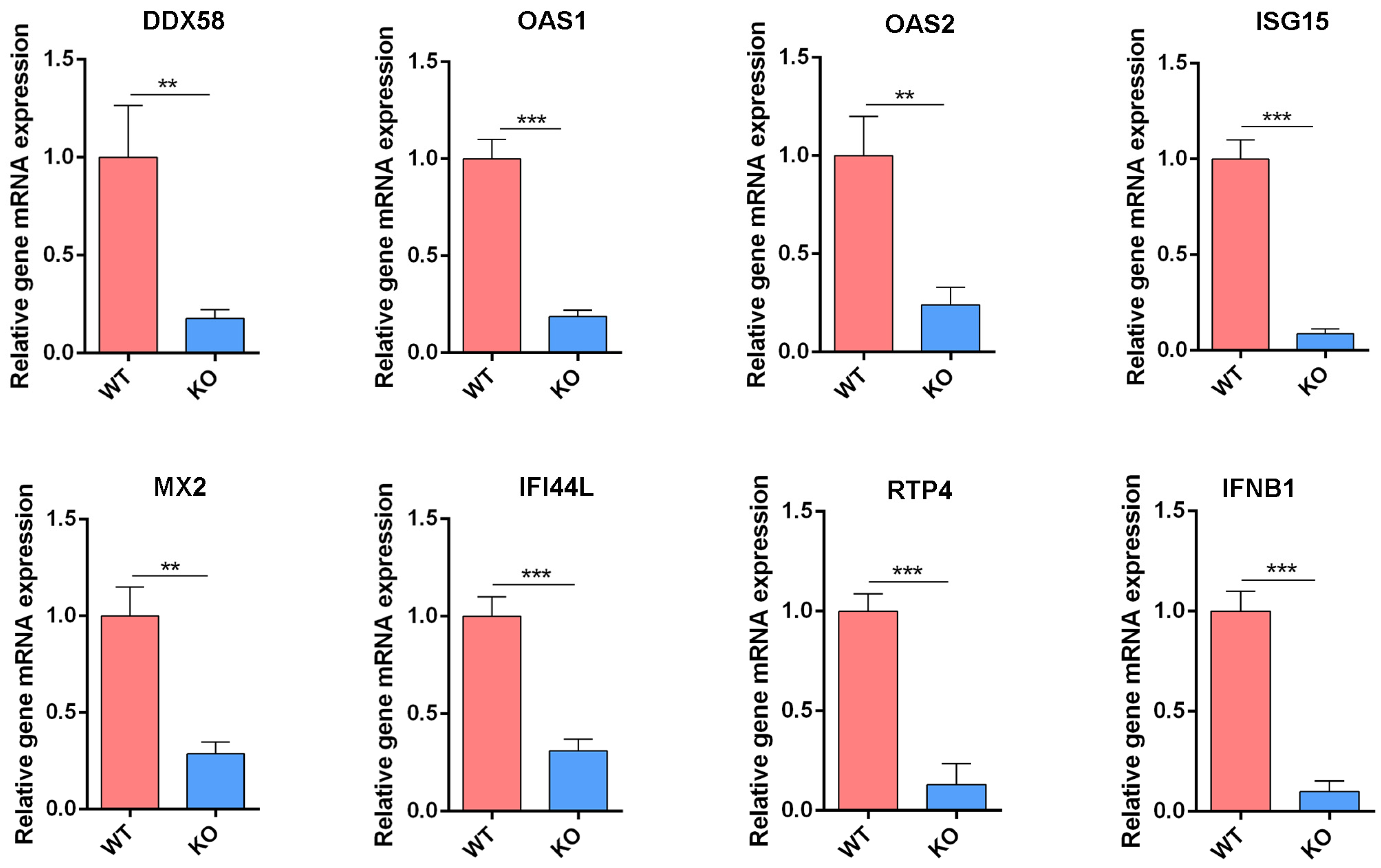
3.4. Reduced Expression of IFN-Related Genes in IFNAR1-KO Cells
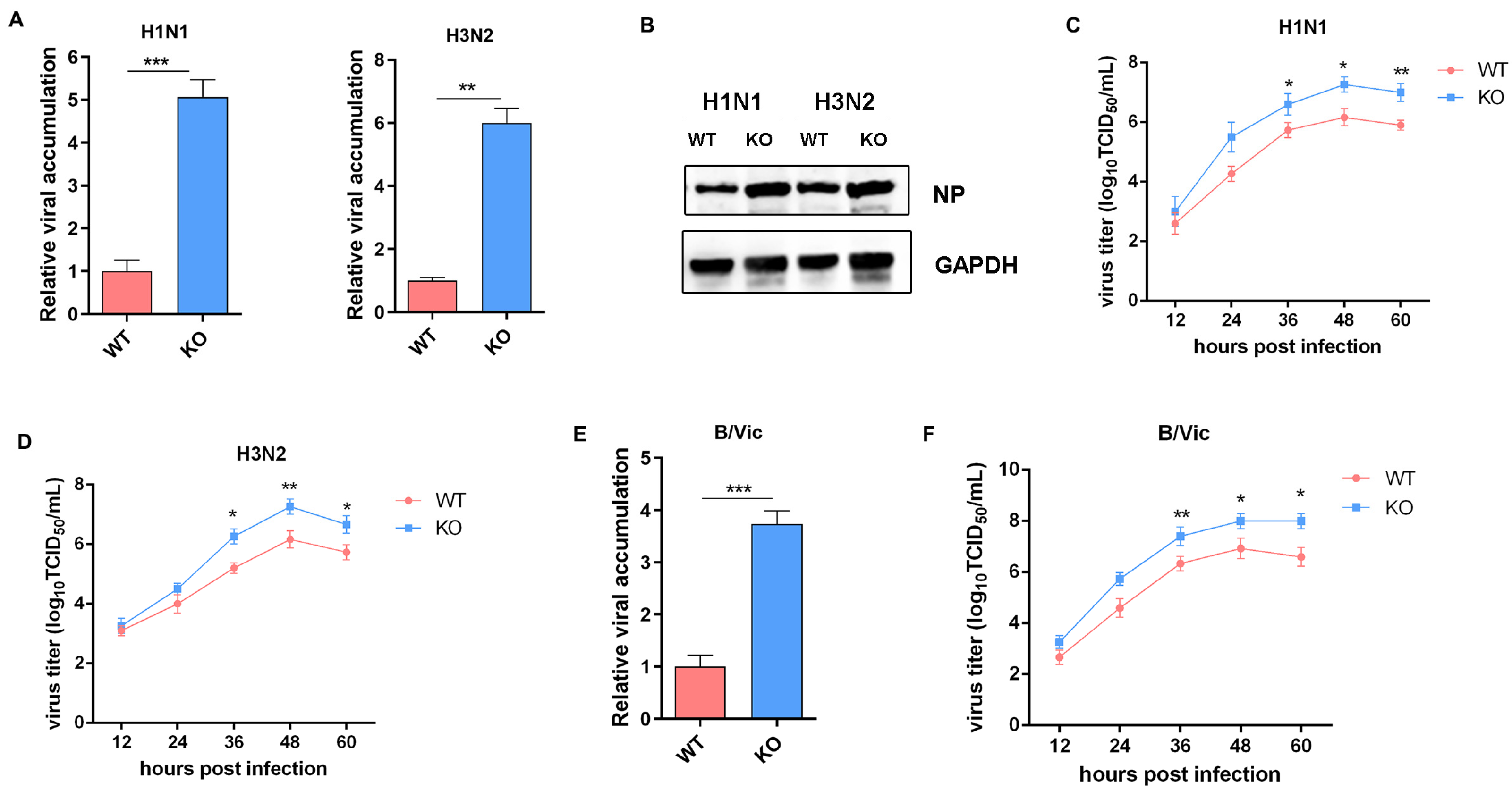
3.5. Increased Viral Titers in IFNAR1-KO Cells
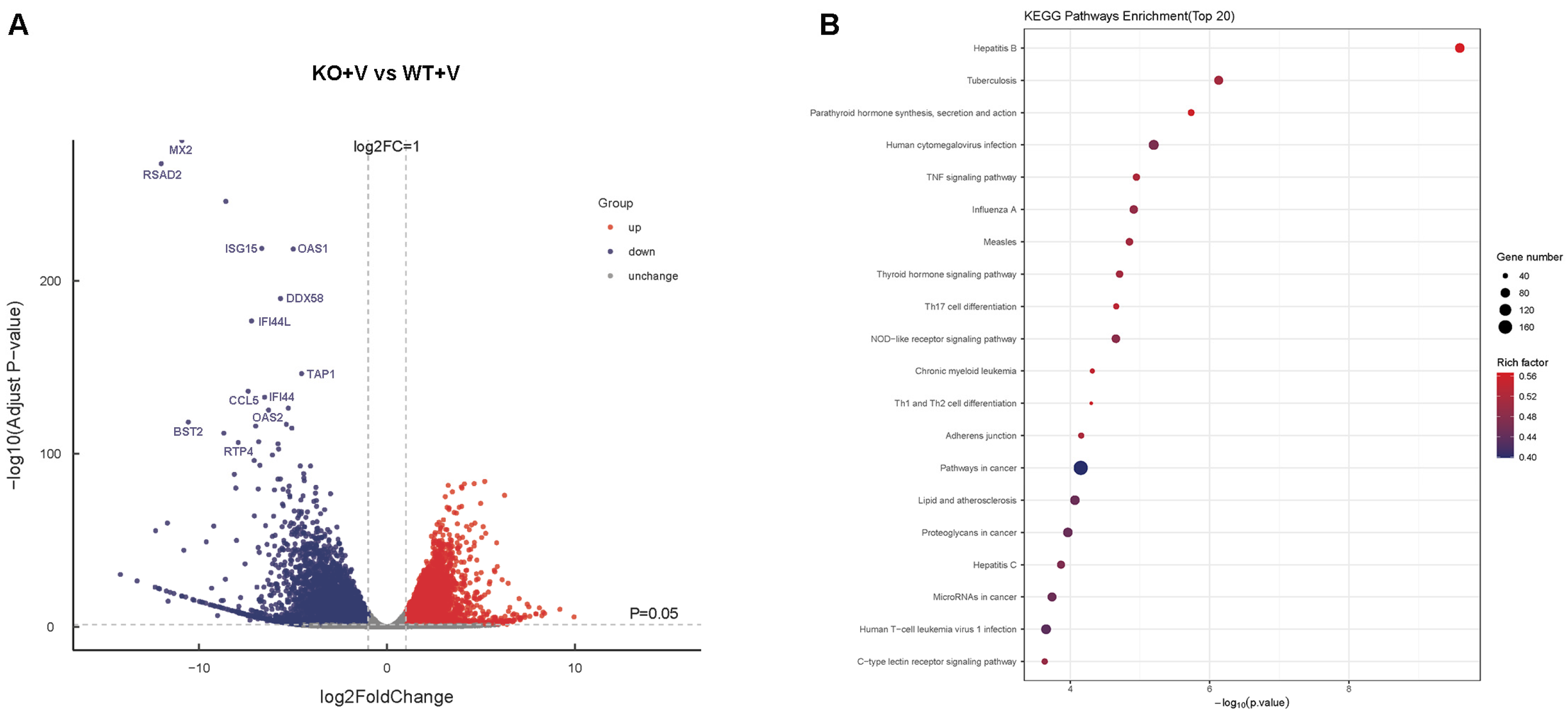
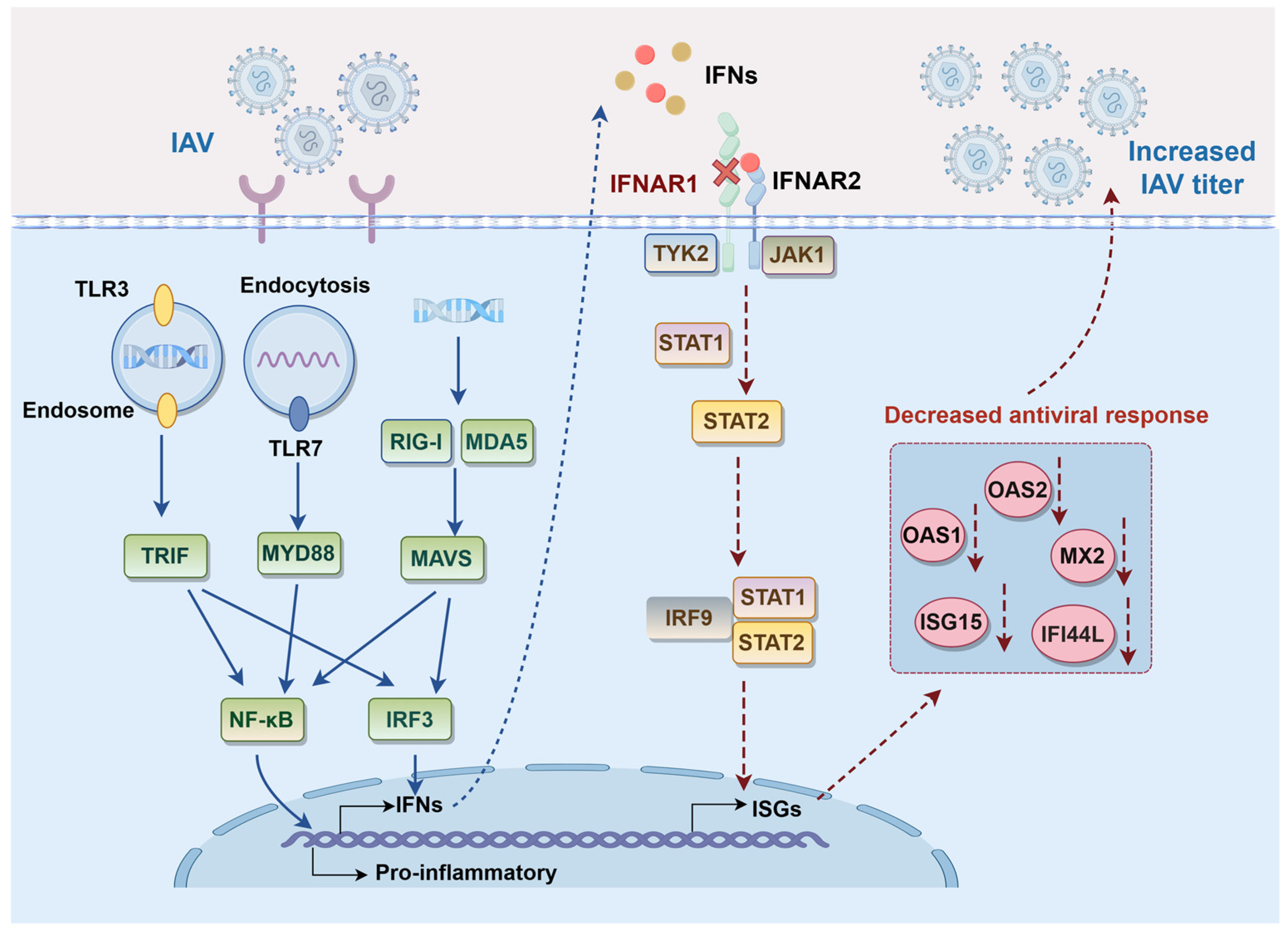
3.6. Transcriptomic Analysis of IFNAR1-KO vs. WT Cells Following Viral Infection
3.7. Validation of Gene Expression Following Viral Infection
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
| Primer Name | Sequence (5′-3′) |
|---|---|
| F1 | CACCGAAGGAAACAACACTTCTCCG |
| R1 | AAACCGGAGAAGTGTTGTTTCCTTC |
| Primer Name | Sequence (5′-3′) |
|---|---|
| F2 | CTTGGTTAAAATTGCCTGGGTG |
| R2 | GCCCTTATGGTAACAGTAAACGAG |
References
- Krammer, F.; Smith, G.J.D.; Fouchier, R.A.M.; Peiris, M.; Kedzierska, K.; Doherty, P.C.; Palese, P.; Shaw, M.L.; Treanor, J.; Webster, R.G.; et al. Influenza. Nat. Rev. Dis. Primers 2018, 4, 3. [Google Scholar] [CrossRef]
- Neumann, G.; Chen, H.; Gao, G.F.; Shu, Y.; Kawaoka, Y. H5N1 influenza viruses: Outbreaks and biological properties. Cell Res. 2010, 20, 51–61. [Google Scholar] [CrossRef]
- Iuliano, A.D.; Roguski, K.M.; Chang, H.H.; Muscatello, D.J.; Palekar, R.; Tempia, S.; Cohen, C.; Gran, J.M.; Schanzer, D.; Cowling, B.J.; et al. Estimates of global seasonal influenza-associated respiratory mortality: A modelling study. Lancet 2018, 391, 1285–1300. [Google Scholar] [CrossRef]
- Shi, J.; Zeng, X.; Cui, P.; Yan, C.; Chen, H. Alarming situation of emerging H5 and H7 avian influenza and effective control strategies. Emerg. Microbes Infect. 2022, 12, 2155072. [Google Scholar] [CrossRef]
- Gao, F.; Wang, Q.; Qiu, C.; Luo, J.; Li, X. Pandemic preparedness of effective vaccines for the outbreak of newly H5N1 highly pathogenic avian influenza virus. Virol. Sin. 2024, 39, 981–985. [Google Scholar] [CrossRef]
- Sun, H.; Xiao, Y.; Liu, J.; Wang, D.; Li, F.; Wang, C.; Li, C.; Zhu, J.; Song, J.; Sun, H.; et al. Prevalent Eurasian avian-like H1N1 swine influenza virus with 2009 pandemic viral genes facilitating human infection. Proc. Natl. Acad. Sci. USA 2020, 117, 17204–17210. [Google Scholar] [CrossRef]
- McLean, H.Q.; Belongia, E.A. Influenza Vaccine Effectiveness: New Insights and Challenges. Cold Spring Harb. Perspect. Med. 2021, 11, a038315. [Google Scholar] [CrossRef]
- Wong, S.-S.; Webby, R.J. Traditional and New Influenza Vaccines. Clin. Microbiol. Rev. 2013, 26, 476–492. [Google Scholar] [CrossRef]
- Parker, L.; Wharton, S.A.; Martin, S.R.; Cross, K.; Lin, Y.; Liu, Y.; Feizi, T.; Daniels, R.S.; McCauley, J.W. Effects of egg-adaptation on receptor-binding and antigenic properties of recent influenza A (H3N2) vaccine viruses. J. Gen. Virol. 2016, 97, 1333–1344. [Google Scholar] [CrossRef]
- Liang, W.; Tan, T.J.C.; Wang, Y.; Lv, H.; Sun, Y.; Bruzzone, R.; Mok, C.K.P.; Wu, N.C. Egg-adaptive mutations of human influenza H3N2 virus are contingent on natural evolution. PLoS Pathog. 2022, 18, e1010875. [Google Scholar] [CrossRef]
- Manini, I.; Trombetta, C.M.; Lazzeri, G.; Pozzi, T.; Rossi, S.; Montomoli, E. Egg-Independent Influenza Vaccines and Vaccine Candidates. Vaccines 2017, 5, 18. [Google Scholar] [CrossRef]
- Tree, J.A.; Richardson, C.; Fooks, A.R.; Clegg, J.C.; Looby, D. Comparison of large-scale mammalian cell culture systems with egg culture for the production of influenza virus A vaccine strains. Vaccine 2001, 19, 3444–3450. [Google Scholar] [CrossRef]
- Milián, E.; Kamen, A.A. Current and Emerging Cell Culture Manufacturing Technologies for Influenza Vaccines. BioMed Res. Int. 2015, 2015, 504831. [Google Scholar] [CrossRef]
- Palache, A.M.; Brands, R.; Scharrenburg, G.J.M.v. Immunogenicity and Reactogenicity of Influenza Subunit Vaccines Produced in MDCK Cells or Fertilized Chicken Eggs. J. Infect. Dis. 1997, 176, S20–S23. [Google Scholar] [CrossRef]
- Gregersen, J.-P.; Heinz-Josef, S.; Heidi, T.; Bröker, M. Safety of MDCK Cell Culture-Based Influenza Vaccines. Future Microbiol. 2011, 6, 143–152. [Google Scholar] [CrossRef]
- Rockman, S.; Laurie, K.; Ong, C.; Rajaram, S.; McGovern, I.; Tran, V.; Youhanna, J. Cell-Based Manufacturing Technology Increases Antigenic Match of Influenza Vaccine and Results in Improved Effectiveness. Vaccines 2022, 11, 52. [Google Scholar] [CrossRef] [PubMed]
- Taub, M.; Chuman, L.; Saier, M.H.; Sato, G. Growth of Madin-Darby canine kidney epithelial cell (MDCK) line in hormone-supplemented, serum-free medium. Proc. Natl. Acad. Sci. USA 1979, 76, 3338–3342. [Google Scholar] [CrossRef] [PubMed]
- Hütter, J.; Rödig, J.V.; Höper, D.; Seeberger, P.H.; Reichl, U.; Rapp, E.; Lepenies, B. Toward Animal Cell Culture–Based Influenza Vaccine Design: Viral Hemagglutinin N-Glycosylation Markedly Impacts Immunogenicity. J. Immunol. 2013, 190, 220–230. [Google Scholar] [CrossRef]
- Zhang, J.; Nian, X.; Liu, B.; Zhang, Z.; Zhao, W.; Han, X.; Ma, Y.; Jin, D.; Ma, H.; Zhang, Q.; et al. Development of MDCK-based quadrivalent split seasonal influenza virus vaccine with high safety and immunoprotection: A preclinical study. Antivir. Res. 2023, 216, 105639. [Google Scholar] [CrossRef]
- Wang, Q.; Wang, J.; Xu, Y.; Li, Z.; Wang, B.; Li, Y. The Interaction of Influenza A NS1 and Cellular TRBP Protein Modulates the Function of RNA Interference Machinery. Front. Microbiol. 2022, 13, 859420. [Google Scholar] [CrossRef] [PubMed]
- Killip, M.J.; Fodor, E.; Randall, R.E. Influenza virus activation of the interferon system. Virus Res. 2015, 209, 11–22. [Google Scholar] [CrossRef]
- Iwasaki, A.; Pillai, P.S. Innate immunity to influenza virus infection. Nat. Rev. Immunol. 2014, 14, 315–328. [Google Scholar] [CrossRef]
- Liu, G.; Lu, Y.; Liu, Q.; Zhou, Y. Inhibition of Ongoing Influenza A Virus Replication Reveals Different Mechanisms of RIG-I Activation. J. Virol. 2019, 93, e02066-18. [Google Scholar] [CrossRef]
- Tripathi, S.; White, M.R.; Hartshorn, K.L. The amazing innate immune response to influenza A virus infection. Innate Immun. 2015, 21, 73–98. [Google Scholar] [CrossRef]
- Shemesh, M.; Lochte, S.; Piehler, J.; Schreiber, G. IFNAR1 and IFNAR2 play distinct roles in initiating type I interferon–induced JAK-STAT signaling and activating STATs. Sci. Signal. 2021, 14, eabe4627. [Google Scholar] [CrossRef] [PubMed]
- Schneider, W.M.; Chevillotte, M.D.; Rice, C.M. Interferon-stimulated genes: A complex web of host defenses. Annu. Rev. Immunol. 2014, 32, 513–545. [Google Scholar] [CrossRef] [PubMed]
- Mayuramart, O.; Poomipak, W.; Rattanaburi, S.; Khongnomnan, K.; Anuntakarun, S.; Saengchoowong, S.; Chavalit, T.; Chantaravisoot, N.; Payungporn, S. IRF7-deficient MDCK cell based on CRISPR/Cas9 technology for enhancing influenza virus replication and improving vaccine production. PeerJ 2022, 10, e13989. [Google Scholar] [CrossRef]
- Wang, Q.; Luo, J.; Li, B.; Ye, Q.; Xu, W.; Gao, F.; Zhou, L.; Lu, W.; Tan, W.-S.; Li, X. Reduction in Interferon-Stimulated Genes Contributes to High-Yield Production of Influenza Virus in Suspension MDCK Cells. Vaccines 2024, 12, 287. [Google Scholar] [CrossRef]
- Kalbfuss, B.; Knöchlein, A.; Kröber, T.; Reichl, U. Monitoring influenza virus content in vaccine production: Precise assays for the quantitation of hemagglutination and neuraminidase activity. Biologicals 2008, 36, 145–161. [Google Scholar] [CrossRef]
- Reed, L.J.; Muench, H. A SIMPLE METHOD OF ESTIMATING FIFTY PER CENT ENDPOINTS. Am. J. Epidemiol. 1938, 27, 493–497. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef]
- Kanehisa, M.; Araki, M.; Goto, S.; Hattori, M.; Hirakawa, M.; Itoh, M.; Katayama, T.; Kawashima, S.; Okuda, S.; Tokimatsu, T.; et al. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 2007, 36, D480–D484. [Google Scholar] [CrossRef]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene Ontology: Tool for the unification of biology. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef]
- Kim, Y.H.; Hong, K.J.; Kim, H.; Nam, J.H. Influenza vaccines: Past, present, and future. Rev. Med. Virol. 2021, 32, e2243. [Google Scholar] [CrossRef]
- Ye, Q.; Phan, T.; Hu, W.-S.; Liu, X.; Fan, L.; Tan, W.-S.; Zhao, L. Transcriptomic Characterization Reveals Attributes of High Influenza Virus Productivity in MDCK Cells. Viruses 2021, 13, 2200. [Google Scholar] [CrossRef]
- Meyers, C.; Huang, D.; Peng, W.-J.; Ye, Q.; Liu, X.-P.; Zhao, L.; Fan, L.; Xia-Hou, K.; Jia, H.-J.; Luo, J.; et al. Serum-Free Suspension Culture of MDCK Cells for Production of Influenza H1N1 Vaccines. PLoS ONE 2015, 10, e0141686. [Google Scholar] [CrossRef]
- Tapia, F.; Vogel, T.; Genzel, Y.; Behrendt, I.; Hirschel, M.; Gangemi, J.D.; Reichl, U. Production of high-titer human influenza A virus with adherent and suspension MDCK cells cultured in a single-use hollow fiber bioreactor. Vaccine 2014, 32, 1003–1011. [Google Scholar] [CrossRef]
- Le, Y.; Zhang, J.; Gong, Z.; Zhang, Z.; Nian, X.; Li, X.; Yu, D.; Ma, N.; Zhou, R.; Zhang, G.; et al. TRAF3 deficiency in MDCK cells improved sensitivity to the influenza A virus. Heliyon 2023, 9, e19246. [Google Scholar] [CrossRef] [PubMed]
- Cai, C.; Wang, X.; Zhao, Y.; Yi, C.; Jin, Z.; Zhang, A.; Han, L. Construction of a mavs-inactivated MDCK cell line for facilitating the propagation of canine distemper virus (CDV). Mol Immunol. 2019, 114, 133–138. [Google Scholar] [CrossRef] [PubMed]
- Eshchenko, N.; Sergeeva, M.; Zhuravlev, E.; Kudria, K.; Goncharova, E.; Komissarov, A.; Stepanov, G. A Knockout of the IFITM3 Gene Increases the Sensitivity of WI-38 VA13 Cells to the Influenza A Virus. Int. J. Mol. Sci. 2024, 25, 625. [Google Scholar] [CrossRef] [PubMed]
- Walter, M.R. The Role of Structure in the Biology of Interferon Signaling. Front. Immunol. 2020, 11, 606489. [Google Scholar] [CrossRef]
- Haji Abdolvahab, M.; Mofrad, M.R.K.; Schellekens, H. Chapter Eight—Interferon Beta: From Molecular Level to Therapeutic Effects. In International Review of Cell and Molecular Biology; Jeon, K.W., Galluzzi, L., Eds.; Academic Press: Cambridge, MA, USA, 2016; Volume 326, pp. 343–372. [Google Scholar]
- Xin, P.; Xu, X.; Deng, C.; Liu, S.; Wang, Y.; Zhou, X.; Ma, H.; Wei, D.; Sun, S. The role of JAK/STAT signaling pathway and its inhibitors in diseases. Int. Immunopharmacol. 2020, 80, 106210. [Google Scholar] [CrossRef] [PubMed]
- Hu, Q.; Bian, Q.; Rong, D.; Wang, L.; Song, J.; Huang, H.-S.; Zeng, J.; Mei, J.; Wang, P.-Y. JAK/STAT pathway: Extracellular signals, diseases, immunity, and therapeutic regimens. Front. Bioeng. Biotechnol. 2023, 11, 1110765. [Google Scholar] [CrossRef] [PubMed]
- Geng, Y.; Jiang, C.; Yang, H.; Xia, Q.; Xu, X.; Yang, K.; Yuan, X.; Chen, J.; Chen, Y.; Chen, X.; et al. Construction of an IFNAR1 knockout MDBK cell line using CRISPR/Cas9 and its effect on bovine virus replication. Front. Immunol. 2024, 15, 1404649. [Google Scholar] [CrossRef] [PubMed]
- Nicolson, C.; Harvey, R.; Engelhardt, O.G.; Robertson, J.S. The Ability of a Non-Egg Adapted (Cell-Like) A(H1N1)pdm09 Virus to Egg-Adapt at HA Loci Other than 222 and 223 and Its Effect on the Yield of Viral Protein. PLoS ONE 2016, 11, e0166761. [Google Scholar] [CrossRef]
- Barman, S.; Franks, J.; Turner, J.C.; Yoon, S.-W.; Webster, R.G.; Webby, R.J. Egg-adaptive mutations in H3N2v vaccine virus enhance egg-based production without loss of antigenicity or immunogenicity. Vaccine 2015, 33, 3186–3192. [Google Scholar] [CrossRef] [PubMed]
- Ma, G.-L.; Qiao, Z.-L.; He, D.; Wang, J.; Kong, Y.-Y.; Xin, X.-Y.; Wen, F.-Q.; Bao, S.-J.; Ma, Z.-R.; Wang, F.-S.; et al. Establishment of a low-tumorigenic MDCK cell line and study of differential molecular networks. Biologicals 2020, 68, 112–121. [Google Scholar] [CrossRef]
- Ye, Q.; Yao, H.; Xiao, Z.; Zhao, L.; Tan, W.-S. Insights into IAV Replication and Lipid Metabolism in Suspension-Adapted MDCK-STAT1-KO Cells. Vaccines 2025, 13, 106. [Google Scholar] [CrossRef]
- Ye, Q.; Xiao, Z.; Bai, C.; Yao, H.; Zhao, L.; Tan, W.-S. Unveiling the multi-characteristic potential of hyper-productive suspension MDCK cells for advanced influenza A virus propagation. Vaccine 2025, 52, 126900. [Google Scholar] [CrossRef]


| Primer Name | Sequence (5′-3′) |
|---|---|
| cIFNAR1 mRNA-F | GAGCAGAGGAAGGAAACAAC |
| cIFNAR1 mRNA-R | CAGGCAGGGAGATGTTTATTATG |
| cRIG-I mRNA-F | AGAGCAAGTTCAGTCAACTGTG |
| cRIG-I mRNA-R | ACTGAAGGTGGACATGGGTC |
| cOAS1 mRNA-F | TCTTCAGGCAAAGGCACGAC |
| cOAS1 mRNA-R | ACTCTGGACCTCAAAGTACAC |
| cOAS2 mRNA-F | GGTGAAGTATTGGTACCAACAGTG |
| cOAS2 mRNA-R | CTGCTCCTGCTGTCTGATTAG |
| cISG15 mRNA-F | CATGGCTGGGAACCTGACTG |
| cISG15 mRNA-R | GAGATCCCATCCTGCAGCAC |
| cMX2 mRNA-F | ACTTGGCATCAGCCATGAGC |
| cMX2 mRNA-R | TGATCGTCTCTTGCTTCTGGATG |
| cIFI44L mRNA-F | ACCAGCATTACTGAGCAGTATAG |
| cIFI44L mRNA-R | TGTAGGGTATGTCTTCCATGC |
| cRTP4 mRNA-F | CTGAGGTCACGAAACAACACAAC |
| cRTP4 mRNA-R | TGGGTCTCAGTAGTGGTATGGC |
| cIFNβ mRNA-F | GAGCAACGACTTGCTTCGATC |
| cIFNβ mRNA-F | CTGGAACTGGCGTGATTTCTC |
| cGAPDH mRNA-F | CCAAGAGGGTCATCATCTCTGC |
| cGAPDH mRNA-R | TGCCGAAGTGGTCATGGATG |
| IVR-215 NP-F | CCAAGCAAACAATGGCGAAG |
| IVR-215 NP-R | CTTTTACTGCAGCACCTGC |
| SAN-010 NP-F | GTGTGGATGGCATGCCATTC |
| SAN-010 NP-R | CTCCGCTCCTGGTCCTTATG |
| BVR-26 NP-F | CAGAGATAAAGAAGAGCGTCTAC |
| BVR-26 NP-R | TTCTTGTCATCAGTGGCAGC |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Q.; Chen, T.; Feng, M.; Zheng, M.; Gao, F.; Qiu, C.; Luo, J.; Li, X. Deficiency of IFNAR1 Increases the Production of Influenza Vaccine Viruses in MDCK Cells. Viruses 2025, 17, 1097. https://doi.org/10.3390/v17081097
Wang Q, Chen T, Feng M, Zheng M, Gao F, Qiu C, Luo J, Li X. Deficiency of IFNAR1 Increases the Production of Influenza Vaccine Viruses in MDCK Cells. Viruses. 2025; 17(8):1097. https://doi.org/10.3390/v17081097
Chicago/Turabian StyleWang, Qi, Tuanjie Chen, Mengru Feng, Mei Zheng, Feixia Gao, Chenchen Qiu, Jian Luo, and Xiuling Li. 2025. "Deficiency of IFNAR1 Increases the Production of Influenza Vaccine Viruses in MDCK Cells" Viruses 17, no. 8: 1097. https://doi.org/10.3390/v17081097
APA StyleWang, Q., Chen, T., Feng, M., Zheng, M., Gao, F., Qiu, C., Luo, J., & Li, X. (2025). Deficiency of IFNAR1 Increases the Production of Influenza Vaccine Viruses in MDCK Cells. Viruses, 17(8), 1097. https://doi.org/10.3390/v17081097






